This report serves to describe the mutational landscape and properties of a given individual set, as well as rank genes and genesets according to mutational significance. MutSig v1.5 was used to generate the results found in this report.
Working with individual set: BRCA.
Number of patients in set: 507
The input for this pipeline is a set of individuals with the following files associated for each:
1. An annotated .maf file describing the mutations called for the respective individual, and their properties.
2. A .wig file that contains information about the coverage of the sample.
Significantly mutated genes (q ≤ 0.1): 71
Mutations seen in COSMIC: 0
Significantly mutated genes in COSMIC territory: 0
Genes with clustered mutations (&le 3 aa apart): 2
Significantly mutated genesets: 210
Significantly mutated genesets: (excluding sig. mutated genes): 16
Table 1. Get Full Table Table representing breakdown of mutations by type.
| type | count |
|---|---|
| Frame_Shift_Del | 951 |
| Frame_Shift_Ins | 313 |
| In_Frame_Del | 377 |
| In_Frame_Ins | 71 |
| Indel | 86 |
| Missense_Mutation | 13573 |
| Nonsense_Mutation | 965 |
| Nonstop_Mutation | 23 |
| Silent | 4910 |
| Splice_Site | 444 |
| Total | 21713 |
Table 2. Get Full Table A breakdown of mutation rates per category discovered for this individual set.
| category | n | N | rate | rate_per_mb | relative_rate |
|---|---|---|---|---|---|
| A->T | 3221 | 7065210282 | 4.6e-07 | 0.46 | 0.38 |
| C->(A/T) | 4896 | 7052709183 | 6.9e-07 | 0.69 | 0.58 |
| A->(C/G) | 3991 | 7065210282 | 5.6e-07 | 0.56 | 0.47 |
| C->G | 1465 | 7052709183 | 2.1e-07 | 0.21 | 0.17 |
| indel+null | 3077 | 14117919972 | 2.2e-07 | 0.22 | 0.18 |
| double_null | 153 | 14117919972 | 1.1e-08 | 0.011 | 0.0091 |
| Total | 16803 | 14117919972 | 1.2e-06 | 1.2 | 1 |
The x axis represents the samples. The y axis represents the exons, one row per exon, and they are sorted by average coverage across samples. For exons with exactly the same average coverage, they are sorted next by the %GC of the exon. (The secondary sort is especially useful for the zero-coverage exons at the bottom).
Figure 1.
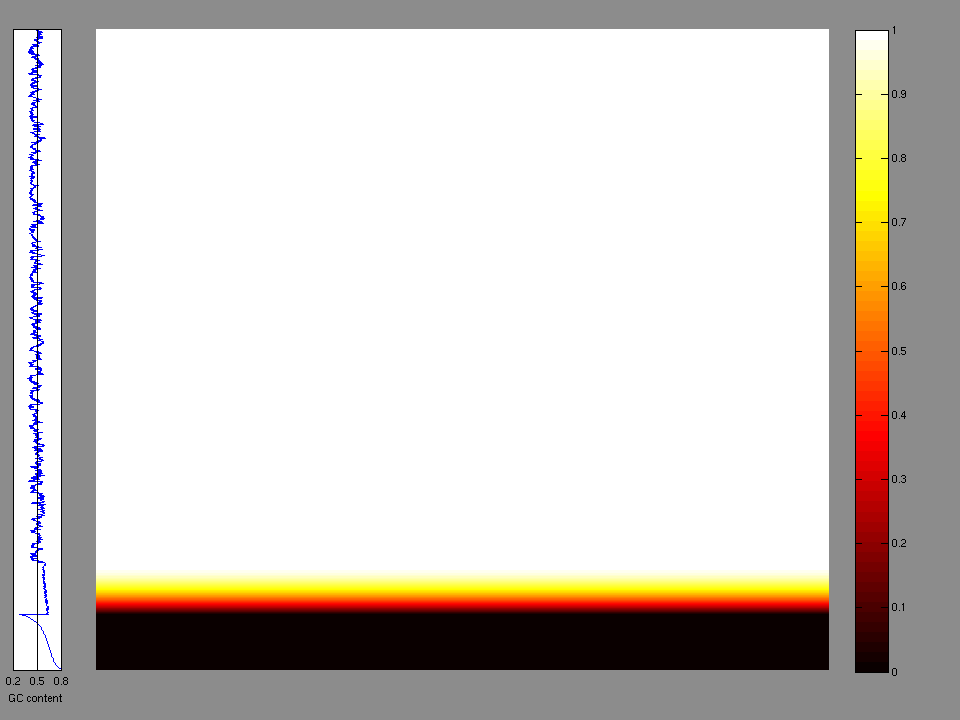

Figure 2.
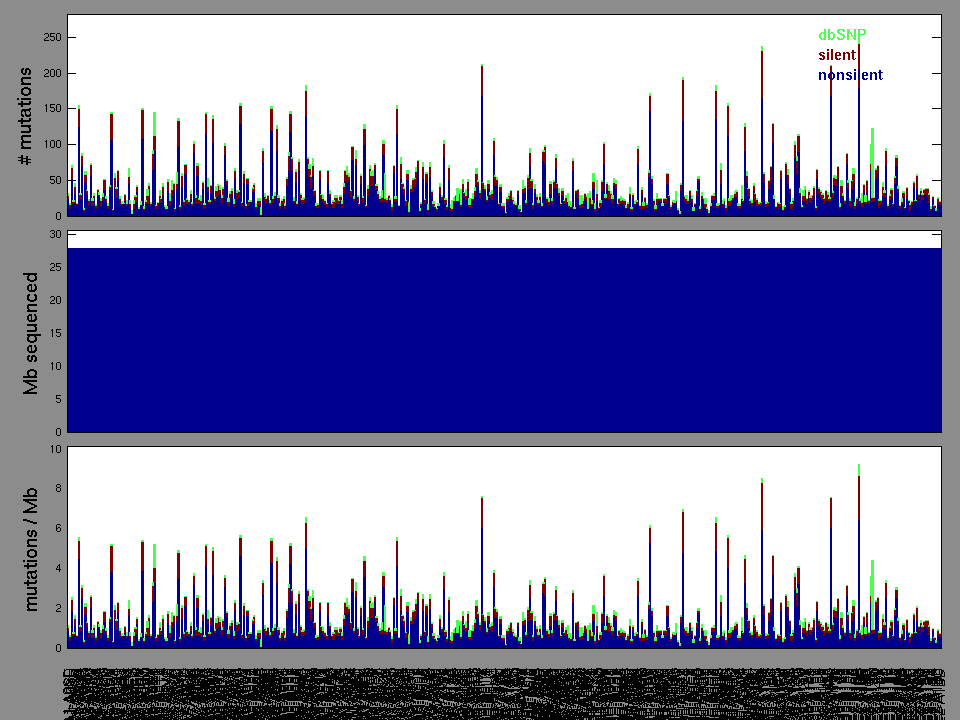
Table 3. Get Full Table A Ranked List of Significantly Mutated Genes. Number of significant genes found: 71. Number of genes displayed: 35
| rank | gene | description | N | n | npat | nsite | nsil | n1 | n2 | n3 | n4 | n5 | n6 | p | q |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | PIK3CA | phosphoinositide-3-kinase, catalytic, alpha polypeptide | 1666509 | 168 | 158 | 29 | 3 | 54 | 17 | 88 | 1 | 8 | 0 | <1.00e-15 | <8.64e-12 |
| 2 | TP53 | tumor protein p53 | 664677 | 142 | 139 | 94 | 3 | 15 | 33 | 18 | 7 | 69 | 0 | <1.00e-15 | <8.64e-12 |
| 3 | MAP3K1 | mitogen-activated protein kinase kinase kinase 1 | 2095431 | 53 | 36 | 52 | 1 | 1 | 1 | 1 | 0 | 22 | 28 | 1.45e-14 | 5.25e-11 |
| 4 | MLL3 | myeloid/lymphoid or mixed-lineage leukemia 3 | 7507149 | 30 | 29 | 30 | 2 | 4 | 1 | 3 | 0 | 20 | 2 | 1.45e-14 | 5.25e-11 |
| 5 | GATA3 | GATA binding protein 3 | 562770 | 55 | 53 | 32 | 1 | 1 | 1 | 0 | 0 | 51 | 2 | 1.52e-14 | 5.25e-11 |
| 6 | CDH1 | cadherin 1, type 1, E-cadherin (epithelial) | 1288794 | 31 | 31 | 29 | 0 | 0 | 4 | 0 | 0 | 27 | 0 | 1.89e-14 | 5.43e-11 |
| 7 | MAP2K4 | mitogen-activated protein kinase kinase 4 | 570375 | 18 | 18 | 17 | 0 | 4 | 0 | 2 | 0 | 12 | 0 | 5.94e-14 | 1.13e-10 |
| 8 | TBX3 | T-box 3 (ulnar mammary syndrome) | 601302 | 11 | 11 | 11 | 0 | 0 | 0 | 1 | 0 | 10 | 0 | 6.42e-14 | 1.13e-10 |
| 9 | RUNX1 | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 373659 | 14 | 14 | 13 | 1 | 0 | 0 | 3 | 0 | 11 | 0 | 6.91e-14 | 1.13e-10 |
| 10 | PTEN | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 632736 | 15 | 14 | 15 | 0 | 0 | 0 | 3 | 0 | 10 | 2 | 6.99e-14 | 1.13e-10 |
| 11 | PIK3R1 | phosphoinositide-3-kinase, regulatory subunit 1 (alpha) | 1199055 | 14 | 14 | 13 | 1 | 1 | 2 | 1 | 1 | 9 | 0 | 7.23e-14 | 1.13e-10 |
| 12 | AKT1 | v-akt murine thymoma viral oncogene homolog 1 | 732108 | 12 | 12 | 2 | 1 | 0 | 11 | 1 | 0 | 0 | 0 | 7.93e-12 | 1.14e-08 |
| 13 | CTCF | CCCTC-binding factor (zinc finger protein) | 1127568 | 11 | 11 | 10 | 2 | 1 | 1 | 3 | 0 | 6 | 0 | 8.11e-11 | 1.08e-07 |
| 14 | NCOR1 | nuclear receptor co-repressor 1 | 3741153 | 16 | 15 | 16 | 0 | 0 | 0 | 2 | 0 | 12 | 2 | 8.20e-10 | 1.01e-06 |
| 15 | RPGR | retinitis pigmentosa GTPase regulator | 1546350 | 10 | 10 | 10 | 0 | 1 | 3 | 0 | 0 | 6 | 0 | 8.59e-08 | 0.000099 |
| 16 | RB1 | retinoblastoma 1 (including osteosarcoma) | 1396278 | 9 | 9 | 9 | 0 | 0 | 0 | 1 | 0 | 8 | 0 | 2.57e-07 | 0.00028 |
| 17 | CBFB | core-binding factor, beta subunit | 226122 | 5 | 5 | 5 | 1 | 2 | 0 | 1 | 0 | 2 | 0 | 3.67e-07 | 0.00037 |
| 18 | ZFP36L1 | zinc finger protein 36, C3H type-like 1 | 519675 | 6 | 6 | 6 | 0 | 0 | 0 | 2 | 0 | 4 | 0 | 4.04e-07 | 0.00039 |
| 19 | CDKN1B | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 276315 | 5 | 5 | 4 | 0 | 0 | 0 | 0 | 0 | 5 | 0 | 4.33e-07 | 0.00039 |
| 20 | FOXA1 | forkhead box A1 | 490269 | 7 | 7 | 6 | 0 | 1 | 3 | 2 | 0 | 1 | 0 | 4.72e-07 | 0.00041 |
| 21 | PRRX1 | paired related homeobox 1 | 287976 | 5 | 5 | 5 | 0 | 1 | 2 | 1 | 0 | 1 | 0 | 4.09e-06 | 0.0034 |
| 22 | MUC4 | mucin 4, cell surface associated | 2554266 | 16 | 13 | 16 | 9 | 5 | 5 | 3 | 3 | 0 | 0 | 4.78e-06 | 0.0038 |
| 23 | AFF2 | AF4/FMR2 family, member 2 | 2015325 | 11 | 11 | 11 | 1 | 1 | 5 | 1 | 1 | 3 | 0 | 7.93e-06 | 0.0060 |
| 24 | DSPP | dentin sialophosphoprotein | 852774 | 7 | 6 | 7 | 0 | 3 | 1 | 0 | 0 | 3 | 0 | 9.29e-06 | 0.0067 |
| 25 | ATN1 | atrophin 1 | 1634568 | 8 | 8 | 6 | 0 | 0 | 4 | 0 | 0 | 4 | 0 | 0.000011 | 0.0077 |
| 26 | MYB | v-myb myeloblastosis viral oncogene homolog (avian) | 1177761 | 7 | 7 | 7 | 0 | 0 | 2 | 0 | 1 | 4 | 0 | 0.000014 | 0.0096 |
| 27 | GPS2 | G protein pathway suppressor 2 | 469482 | 5 | 5 | 5 | 1 | 0 | 0 | 0 | 0 | 5 | 0 | 0.000022 | 0.014 |
| 28 | DALRD3 | DALR anticodon binding domain containing 3 | 596739 | 6 | 6 | 6 | 0 | 0 | 2 | 1 | 0 | 3 | 0 | 0.000022 | 0.014 |
| 29 | SF3B1 | splicing factor 3b, subunit 1, 155kDa | 2047773 | 10 | 10 | 6 | 0 | 0 | 2 | 7 | 0 | 1 | 0 | 0.000032 | 0.017 |
| 30 | NKAIN4 | Na+/K+ transporting ATPase interacting 4 | 104949 | 3 | 3 | 3 | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 0.000032 | 0.017 |
| 31 | PIWIL1 | piwi-like 1 (Drosophila) | 1351662 | 8 | 8 | 8 | 1 | 2 | 3 | 2 | 0 | 1 | 0 | 0.000032 | 0.017 |
| 32 | FAM166A | 236769 | 4 | 4 | 4 | 0 | 0 | 1 | 0 | 0 | 3 | 0 | 0.000032 | 0.017 | |
| 33 | SAAL1 | serum amyloid A-like 1 | 676338 | 5 | 5 | 5 | 0 | 0 | 1 | 0 | 0 | 4 | 0 | 0.000039 | 0.020 |
| 34 | TMEM82 | transmembrane protein 82 | 161733 | 4 | 4 | 4 | 0 | 1 | 1 | 1 | 0 | 1 | 0 | 0.000043 | 0.022 |
| 35 | HIST1H2BC | histone cluster 1, H2bc | 195195 | 4 | 4 | 4 | 1 | 0 | 2 | 1 | 0 | 1 | 0 | 0.000049 | 0.023 |
Note:
N - number of sequenced bases in this gene across the individual set.
n - number of (nonsilent) mutations in this gene across the individual set.
npat - number of patients (individuals) with at least one nonsilent mutation.
nsite - number of unique sites having a non-silent mutation.
nsil - number of silent mutations in this gene across the individual set.
n1 - number of nonsilent mutations of type: A->T .
n2 - number of nonsilent mutations of type: C->(A/T) .
n3 - number of nonsilent mutations of type: A->(C/G) .
n4 - number of nonsilent mutations of type: C->G .
n5 - number of nonsilent mutations of type: indel+null .
null - mutation category that includes nonsense, frameshift, splice-site mutations
p_classic = p-value for the observed amount of nonsilent mutations being elevated in this gene
p_ns_s = p-value for the observed nonsilent/silent ratio being elevated in this gene
p = p-value (overall)
q = q-value, False Discovery Rate (Benjamini-Hochberg procedure)
In this analysis, COSMIC is used as a filter to increase power by restricting the territory of each gene. Cosmic version: v48.
Table 4. Get Full Table Significantly mutated genes (COSMIC territory only). To access the database please go to: COSMIC. Number of significant genes found: 0. Number of genes displayed: 10
| rank | gene | description | n | cos | n_cos | N_cos | cos_ev | p | q |
|---|---|---|---|---|---|---|---|---|---|
| 1 | A4GNT | alpha-1,4-N-acetylglucosaminyltransferase | 0 | 0 | 0 | 0 | 0 | 1 | 1 |
| 2 | AACS | acetoacetyl-CoA synthetase | 0 | 0 | 0 | 0 | 0 | 1 | 1 |
| 3 | ABCA9 | ATP-binding cassette, sub-family A (ABC1), member 9 | 3 | 0 | 0 | 0 | 0 | 1 | 1 |
| 4 | ABCC10 | ATP-binding cassette, sub-family C (CFTR/MRP), member 10 | 2 | 0 | 0 | 0 | 0 | 1 | 1 |
| 5 | ABCF2 | ATP-binding cassette, sub-family F (GCN20), member 2 | 3 | 0 | 0 | 0 | 0 | 1 | 1 |
| 6 | ABHD2 | abhydrolase domain containing 2 | 2 | 0 | 0 | 0 | 0 | 1 | 1 |
| 7 | ABHD4 | abhydrolase domain containing 4 | 0 | 0 | 0 | 0 | 0 | 1 | 1 |
| 8 | ACADS | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 0 | 0 | 0 | 0 | 0 | 1 | 1 |
| 9 | ACOT11 | acyl-CoA thioesterase 11 | 2 | 0 | 0 | 0 | 0 | 1 | 1 |
| 10 | ACRBP | acrosin binding protein | 1 | 0 | 0 | 0 | 0 | 1 | 1 |
Note:
n - number of (nonsilent) mutations in this gene across the individual set.
cos = number of unique mutated sites in this gene in COSMIC
n_cos = overlap between n and cos.
N_cos = number of individuals times cos.
cos_ev = total evidence: number of reports in COSMIC for mutations seen in this gene.
p = p-value for seeing the observed amount of overlap in this gene)
q = q-value, False Discovery Rate (Benjamini-Hochberg procedure)
Table 5. Get Full Table Genes with Clustered Mutations
| num | gene | desc | n | mindist | npairs3 | npairs12 |
|---|---|---|---|---|---|---|
| 2757 | FER1L6 | fer-1-like 6 (C. elegans) | 2 | 0 | 1 | 1 |
| 1917 | CYP11B2 | cytochrome P450, family 11, subfamily B, polypeptide 2 | 3 | 2 | 1 | 1 |
| 18 | ABCA13 | ATP-binding cassette, sub-family A (ABC1), member 13 | 13 | 30 | 0 | 0 |
| 4758 | MXRA5 | matrix-remodelling associated 5 | 5 | 42 | 0 | 0 |
| 7739 | TRANK1 | 3 | 139 | 0 | 0 | |
| 7552 | TLN1 | talin 1 | 7 | 220 | 0 | 0 |
| 6611 | SEC31B | SEC31 homolog B (S. cerevisiae) | 5 | 263 | 0 | 0 |
| 4747 | MUC6 | mucin 6, oligomeric mucus/gel-forming | 4 | 301 | 0 | 0 |
| 2 | A2BP1 | 3 | Inf | 0 | 0 | |
| 3 | A2M | alpha-2-macroglobulin | 3 | Inf | 0 | 0 |
Note:
n - number of mutations in this gene in the individual set.
mindist - distance (in aa) between closest pair of mutations in this gene
npairs3 - how many pairs of mutations are within 3 aa of each other.
npairs12 - how many pairs of mutations are within 12 aa of each other.
Table 6. Get Full Table A Ranked List of Significantly Mutated Genesets. (Source: MSigDB GSEA Cannonical Pathway Set).Number of significant genesets found: 210. Number of genesets displayed: 10
| rank | geneset | description | genes | N_genes | mut_tally | N | n | npat | nsite | nsil | n1 | n2 | n3 | n4 | n5 | n6 | p | q |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HSA04510_FOCAL_ADHESION | Genes involved in focal adhesion | ACTB, ACTG1, ACTN1, ACTN2, ACTN3, ACTN4, AKT1, AKT2, AKT3, ARHGAP5, BAD, BCAR1, BCL2, BIRC2, BIRC3, BIRC4, BRAF, CAPN2, CAV1, CAV2, CAV3, CCND1, CCND2, CCND3, CDC42, CHAD, COL11A1, COL11A2, COL1A1, COL1A2, COL2A1, COL3A1, COL4A1, COL4A2, COL4A4, COL4A6, COL5A1, COL5A2, COL5A3, COL6A1, COL6A2, COL6A3, COL6A6, COMP, CRK, CRKL, CTNNB1, DIAPH1, DOCK1, EGF, EGFR, ELK1, ERBB2, FARP2, FIGF, FLNA, FLNB, FLNC, FLT1, FN1, FYN, GRB2, GRLF1, GSK3B, HGF, HRAS, IBSP, IGF1, IGF1R, ILK, ITGA1, ITGA10, ITGA11, ITGA2, ITGA2B, ITGA3, ITGA4, ITGA5, ITGA6, ITGA7, ITGA8, ITGA9, ITGAV, ITGB1, ITGB3, ITGB4, ITGB5, ITGB6, ITGB7, ITGB8, JUN, KDR, LAMA1, LAMA2, LAMA3, LAMA4, LAMA5, LAMB1, LAMB2, LAMB3, LAMB4, LAMC1, LAMC2, LAMC3, LOC653852, MAP2K1, MAPK1, MAPK10, MAPK3, MAPK8, MAPK9, MET, MLCK, MRCL3, MRLC2, MYL2, MYL5, MYL7, MYL8P, MYL9, MYLC2PL, MYLK, MYLK2, MYLPF, PAK1, PAK2, PAK3, PAK4, PAK6, PAK7, PARVA, PARVB, PARVG, PDGFA, PDGFB, PDGFC, PDGFD, PDGFRA, PDGFRB, PDPK1, PGF, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PIP5K1C, PPP1CA, PPP1CB, PPP1CC, PPP1R12A, PRKCA, PRKCB1, PRKCG, PTEN, PTK2, PXN, RAC1, RAC2, RAC3, RAF1, RAP1A, RAP1B, RAPGEF1, RELN, RHOA, ROCK1, ROCK2, SHC1, SHC2, SHC3, SHC4, SOS1, SOS2, SPP1, SRC, THBS1, THBS2, THBS3, THBS4, TLN1, TLN2, TNC, TNN, TNR, TNXB, VASP, VAV1, VAV2, VAV3, VCL, VEGFA, VEGFB, VEGFC, VTN, VWF, ZYX | 185 | ACTB(2), ACTG1(1), ACTN2(4), ACTN3(1), ACTN4(4), AKT1(12), AKT2(1), AKT3(2), ARHGAP5(3), BAD(1), BRAF(2), CAPN2(2), CAV1(1), CCND1(1), CCND3(5), COL11A1(2), COL11A2(1), COL1A1(6), COL1A2(3), COL2A1(3), COL3A1(1), COL4A1(4), COL4A2(2), COL4A4(1), COL4A6(3), COL5A1(2), COL5A2(4), COL5A3(4), COL6A3(5), COL6A6(4), COMP(3), CRKL(1), EGF(2), EGFR(4), ELK1(2), ERBB2(4), FARP2(3), FIGF(1), FLNA(4), FLNB(7), FLNC(8), FLT1(2), FN1(3), GRB2(1), GRLF1(4), HGF(3), IGF1(1), IGF1R(1), ILK(2), ITGA1(3), ITGA10(1), ITGA11(4), ITGA2(2), ITGA2B(2), ITGA4(1), ITGA6(3), ITGA7(5), ITGA8(2), ITGAV(3), ITGB1(2), ITGB3(2), ITGB4(4), ITGB5(2), ITGB8(1), JUN(1), KDR(2), LAMA1(6), LAMA2(7), LAMA3(6), LAMA4(4), LAMB1(3), LAMB2(3), LAMB3(4), LAMB4(6), LAMC1(2), LAMC2(1), LAMC3(1), MAP2K1(1), MAPK1(1), MAPK10(1), MAPK3(1), MAPK8(2), MET(4), MYL7(1), MYLK(5), PAK1(2), PAK2(1), PAK3(1), PAK7(1), PARVA(1), PDGFB(1), PDGFRA(3), PDGFRB(2), PIK3CA(168), PIK3CB(3), PIK3CG(2), PIK3R1(14), PIK3R3(1), PIP5K1C(3), PPP1R12A(1), PRKCG(1), PTEN(15), PTK2(1), PXN(1), RAF1(1), RAPGEF1(3), RELN(10), RHOA(2), ROCK1(1), ROCK2(2), SHC1(1), SHC3(1), SHC4(2), SOS1(3), SOS2(1), SPP1(1), THBS1(3), THBS3(1), THBS4(1), TLN1(7), TLN2(4), TNC(3), TNN(3), TNR(1), TNXB(3), VAV1(1), VAV3(1), VCL(3), VWF(6) | 263403231 | 529 | 325 | 378 | 116 | 112 | 131 | 163 | 29 | 86 | 8 | <1.00e-15 | <5.98e-15 |
| 2 | HSA04210_APOPTOSIS | Genes involved in apoptosis | AIFM1, AKT1, AKT2, AKT3, APAF1, ATM, BAD, BAX, BCL2, BCL2L1, BID, BIRC2, BIRC3, BIRC4, CAPN1, CAPN2, CASP10, CASP3, CASP6, CASP7, CASP8, CASP9, CFLAR, CHP, CHUK, CSF2RB, CYCS, DFFA, DFFB, ENDOG, FADD, FAS, FASLG, IKBKB, IKBKG, IL1A, IL1B, IL1R1, IL1RAP, IL3, IL3RA, IRAK1, IRAK2, IRAK3, IRAK4, MAP3K14, MYD88, NFKB1, NFKB2, NFKBIA, NGFB, NTRK1, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PRKACA, PRKACB, PRKACG, PRKAR1A, PRKAR1B, PRKAR2A, PRKAR2B, RELA, RIPK1, TNF, TNFRSF10A, TNFRSF10B, TNFRSF10C, TNFRSF10D, TNFRSF1A, TNFSF10, TP53, TRADD, TRAF2 | 79 | AKT1(12), AKT2(1), AKT3(2), APAF1(1), ATM(9), BAD(1), BID(4), CAPN1(3), CAPN2(2), CASP6(1), CASP8(2), CSF2RB(2), DFFB(2), FAS(1), FASLG(1), IKBKB(2), IL1A(1), IL1R1(1), IL1RAP(2), IL3RA(1), IRAK1(1), IRAK2(2), IRAK3(1), IRAK4(2), NFKB1(1), NFKB2(4), NTRK1(1), PIK3CA(168), PIK3CB(3), PIK3CG(2), PIK3R1(14), PIK3R3(1), PPP3CA(3), PPP3CB(2), PRKACA(1), PRKAR1A(1), PRKAR2A(1), PRKAR2B(1), RELA(1), RIPK1(3), TNFRSF10D(1), TNFRSF1A(1), TP53(142) | 59436624 | 408 | 310 | 210 | 20 | 82 | 90 | 118 | 15 | 100 | 3 | <1.00e-15 | <5.98e-15 |
| 3 | HSA04010_MAPK_SIGNALING_PATHWAY | Genes involved in MAPK signaling pathway | ACVR1B, ACVR1C, AKT1, AKT2, AKT3, ARRB1, ARRB2, ATF2, ATF4, BDNF, BRAF, CACNA1A, CACNA1B, CACNA1C, CACNA1D, CACNA1E, CACNA1F, CACNA1G, CACNA1H, CACNA1I, CACNA1S, CACNA2D1, CACNA2D2, CACNA2D3, CACNA2D4, CACNB1, CACNB2, CACNB3, CACNB4, CACNG1, CACNG2, CACNG3, CACNG4, CACNG5, CACNG6, CACNG7, CACNG8, CASP3, CD14, CDC25B, CDC42, CHP, CHUK, CRK, CRKL, DAXX, DDIT3, DUSP1, DUSP10, DUSP14, DUSP16, DUSP2, DUSP3, DUSP4, DUSP5, DUSP6, DUSP7, DUSP8, DUSP9, ECSIT, EGF, EGFR, ELK1, ELK4, EVI1, FAS, FASLG, FGF1, FGF10, FGF11, FGF12, FGF13, FGF14, FGF16, FGF17, FGF18, FGF19, FGF2, FGF20, FGF21, FGF22, FGF23, FGF3, FGF4, FGF5, FGF6, FGF7, FGF8, FGF9, FGFR1, FGFR2, FGFR3, FGFR4, FLNA, FLNB, FLNC, FOS, GADD45A, GADD45B, GADD45G, GNA12, GNG12, GRB2, HRAS, IKBKB, IKBKG, IL1A, IL1B, IL1R1, IL1R2, JUN, JUND, KRAS, LOC653852, MAP2K1, MAP2K1IP1, MAP2K2, MAP2K3, MAP2K4, MAP2K5, MAP2K6, MAP2K7, MAP3K1, MAP3K10, MAP3K12, MAP3K13, MAP3K14, MAP3K2, MAP3K3, MAP3K4, MAP3K5, MAP3K6, MAP3K7, MAP3K7IP1, MAP3K7IP2, MAP3K8, MAP4K1, MAP4K2, MAP4K3, MAP4K4, MAPK1, MAPK10, MAPK11, MAPK12, MAPK13, MAPK14, MAPK3, MAPK7, MAPK8, MAPK8IP1, MAPK8IP2, MAPK8IP3, MAPK9, MAPKAPK2, MAPKAPK3, MAPKAPK5, MAPT, MAX, MEF2C, MKNK1, MKNK2, MOS, MRAS, MYC, NF1, NFATC2, NFATC4, NFKB1, NFKB2, NGFB, NLK, NR4A1, NRAS, NTF3, NTF5, NTRK1, NTRK2, PAK1, PAK2, PDGFA, PDGFB, PDGFRA, PDGFRB, PLA2G10, PLA2G12A, PLA2G12B, PLA2G1B, PLA2G2A, PLA2G2D, PLA2G2E, PLA2G2F, PLA2G3, PLA2G4A, PLA2G5, PLA2G6, PPM1A, PPM1B, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PPP5C, PRKACA, PRKACB, PRKACG, PRKCA, PRKCB1, PRKCG, PRKX, PRKY, PTPN5, PTPN7, PTPRR, RAC1, RAC2, RAC3, RAF1, RAP1A, RAP1B, RAPGEF2, RASA1, RASA2, RASGRF1, RASGRF2, RASGRP1, RASGRP2, RASGRP3, RASGRP4, RPS6KA1, RPS6KA2, RPS6KA3, RPS6KA4, RPS6KA5, RPS6KA6, RRAS, RRAS2, SOS1, SOS2, SRF, STK3, STK4, STMN1, TAOK1, TAOK2, TAOK3, TGFB1, TGFB2, TGFB3, TGFBR1, TGFBR2, TNF, TNFRSF1A, TP53, TRAF2, TRAF6, ZAK | 231 | ACVR1B(4), ACVR1C(1), AKT1(12), AKT2(1), AKT3(2), ARRB1(1), ATF4(1), BDNF(1), BRAF(2), CACNA1A(5), CACNA1B(1), CACNA1C(4), CACNA1D(4), CACNA1E(8), CACNA1F(6), CACNA1G(5), CACNA1I(2), CACNA1S(1), CACNA2D1(4), CACNA2D2(2), CACNA2D3(5), CACNA2D4(1), CACNB1(1), CACNB2(2), CACNG2(1), CACNG3(2), CACNG4(1), CACNG6(1), CD14(1), CDC25B(3), CRKL(1), DUSP10(1), DUSP16(3), DUSP6(1), DUSP7(1), DUSP9(1), EGF(2), EGFR(4), ELK1(2), FAS(1), FASLG(1), FGF11(1), FGF13(2), FGF18(2), FGF23(1), FGF3(1), FGF5(1), FGF6(1), FGF7(2), FGFR2(2), FGFR4(3), FLNA(4), FLNB(7), FLNC(8), GRB2(1), IKBKB(2), IL1A(1), IL1R1(1), IL1R2(3), JUN(1), KRAS(1), MAP2K1(1), MAP2K3(1), MAP2K4(18), MAP2K5(1), MAP3K1(53), MAP3K10(4), MAP3K12(2), MAP3K13(3), MAP3K4(3), MAP3K5(1), MAP3K6(1), MAP3K8(2), MAP4K1(1), MAP4K4(1), MAPK1(1), MAPK10(1), MAPK14(1), MAPK3(1), MAPK8(2), MAPK8IP1(1), MAPKAPK3(1), MAPT(1), MAX(1), MEF2C(2), MOS(2), MYC(1), NF1(13), NFATC4(3), NFKB1(1), NFKB2(4), NLK(1), NR4A1(1), NTRK1(1), NTRK2(1), PAK1(2), PAK2(1), PDGFB(1), PDGFRA(3), PDGFRB(2), PLA2G2D(1), PLA2G2F(1), PLA2G3(3), PLA2G4A(6), PLA2G6(2), PPP3CA(3), PPP3CB(2), PRKACA(1), PRKCG(1), PTPN5(2), PTPN7(1), PTPRR(1), RAF1(1), RAPGEF2(1), RASA1(2), RASA2(1), RASGRF1(3), RASGRP2(2), RASGRP4(1), RPS6KA1(1), RPS6KA2(2), RPS6KA3(2), RPS6KA4(1), RPS6KA6(3), SOS1(3), SOS2(1), SRF(1), STK3(1), STK4(1), TAOK1(2), TAOK2(1), TGFB3(1), TGFBR1(2), TNFRSF1A(1), TP53(142), TRAF6(1), ZAK(1) | 202419750 | 491 | 304 | 428 | 74 | 74 | 134 | 70 | 28 | 152 | 33 | <1.00e-15 | <5.98e-15 |
| 4 | HSA04810_REGULATION_OF_ACTIN_CYTOSKELETON | Genes involved in regulation of actin cytoskeleton | ABI2, ACTN1, ACTN2, ACTN3, ACTN4, APC, APC2, ARAF, ARHGEF1, ARHGEF12, ARHGEF4, ARHGEF6, ARHGEF7, ARPC1A, ARPC1B, ARPC2, ARPC3, ARPC4, ARPC5, ARPC5L, BAIAP2, BCAR1, BDKRB1, BDKRB2, BRAF, C3orf10, CD14, CDC42, CFL1, CFL2, CHRM1, CHRM2, CHRM3, CHRM4, CHRM5, CRK, CRKL, CSK, CYFIP1, CYFIP2, DIAPH1, DIAPH2, DIAPH3, DOCK1, EGF, EGFR, EZR, F2, F2R, FGD1, FGD3, FGF1, FGF10, FGF11, FGF12, FGF13, FGF14, FGF16, FGF17, FGF18, FGF19, FGF2, FGF20, FGF21, FGF22, FGF23, FGF3, FGF4, FGF5, FGF6, FGF7, FGF8, FGF9, FGFR1, FGFR2, FGFR3, FGFR4, FN1, GIT1, GNA12, GNA13, GNG12, GRLF1, GSN, HRAS, INS, IQGAP1, IQGAP2, IQGAP3, ITGA1, ITGA10, ITGA11, ITGA2, ITGA2B, ITGA3, ITGA4, ITGA5, ITGA6, ITGA7, ITGA8, ITGA9, ITGAD, ITGAE, ITGAL, ITGAM, ITGAV, ITGAX, ITGB1, ITGB2, ITGB3, ITGB4, ITGB5, ITGB6, ITGB7, ITGB8, KRAS, LIMK1, LIMK2, LOC200025, LOC645126, LOC653888, MAP2K1, MAP2K2, MAPK1, MAPK3, MLCK, MOS, MRAS, MRCL3, MRLC2, MSN, MYH10, MYH14, MYH9, MYL2, MYL5, MYL7, MYL8P, MYL9, MYLC2PL, MYLK, MYLK2, MYLPF, NCKAP1, NCKAP1L, NRAS, PAK1, PAK2, PAK3, PAK4, PAK6, PAK7, PDGFA, PDGFB, PDGFRA, PDGFRB, PFN1, PFN2, PFN3, PFN4, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PIP4K2A, PIP4K2B, PIP4K2C, PIP5K1A, PIP5K1B, PIP5K1C, PIP5K3, PPP1CA, PPP1CB, PPP1CC, PPP1R12A, PPP1R12B, PTK2, PXN, RAC1, RAC2, RAC3, RAF1, RDX, RHOA, ROCK1, ROCK2, RRAS, RRAS2, SCIN, SLC9A1, SOS1, SOS2, SSH1, SSH2, SSH3, TIAM1, TIAM2, TMSB4X, TMSB4Y, TMSL3, VAV1, VAV2, VAV3, VCL, WAS, WASF1, WASF2, WASL | 193 | ACTN2(4), ACTN3(1), ACTN4(4), APC(3), ARAF(1), ARHGEF1(5), ARHGEF12(2), ARHGEF4(2), ARHGEF6(1), ARHGEF7(2), ARPC1A(1), ARPC1B(1), ARPC2(2), BAIAP2(1), BDKRB1(2), BRAF(2), C3orf10(1), CD14(1), CHRM1(2), CHRM2(2), CHRM3(1), CHRM5(2), CRKL(1), CYFIP2(2), DIAPH3(1), EGF(2), EGFR(4), EZR(2), F2(2), F2R(2), FGD3(2), FGF11(1), FGF13(2), FGF18(2), FGF23(1), FGF3(1), FGF5(1), FGF6(1), FGF7(2), FGFR2(2), FGFR4(3), FN1(3), GIT1(1), GRLF1(4), GSN(1), IQGAP1(1), IQGAP2(2), IQGAP3(3), ITGA1(3), ITGA10(1), ITGA11(4), ITGA2(2), ITGA2B(2), ITGA4(1), ITGA6(3), ITGA7(5), ITGA8(2), ITGAD(1), ITGAL(1), ITGAM(1), ITGAV(3), ITGAX(2), ITGB1(2), ITGB2(3), ITGB3(2), ITGB4(4), ITGB5(2), ITGB8(1), KRAS(1), LIMK1(3), LIMK2(1), MAP2K1(1), MAPK1(1), MAPK3(1), MOS(2), MSN(1), MYH10(4), MYH14(5), MYH9(5), MYL7(1), MYLK(5), NCKAP1(3), NCKAP1L(3), PAK1(2), PAK2(1), PAK3(1), PAK7(1), PDGFB(1), PDGFRA(3), PDGFRB(2), PIK3CA(168), PIK3CB(3), PIK3CG(2), PIK3R1(14), PIK3R3(1), PIP4K2A(1), PIP4K2C(5), PIP5K1A(1), PIP5K1C(3), PPP1R12A(1), PTK2(1), PXN(1), RAF1(1), RDX(1), RHOA(2), ROCK1(1), ROCK2(2), SCIN(2), SOS1(3), SOS2(1), SSH1(1), SSH2(1), SSH3(2), TIAM1(1), TIAM2(4), VAV1(1), VAV3(1), VCL(3), WAS(1), WASF1(1), WASL(1) | 197302092 | 416 | 288 | 275 | 90 | 103 | 88 | 140 | 24 | 61 | 0 | <1.00e-15 | <5.98e-15 |
| 5 | ARFPATHWAY | Cyclin-dependent kinase inhibitor 2A is a tumor suppressor that induces G1 arrest and can activate the p53 pathway, leading to G2/M arrest. | ABL1, CDKN2A, E2F1, MDM2, MYC, PIK3CA, PIK3R1, POLR1A, POLR1B, POLR1C, POLR1D, RAC1, RB1, TBX2, TP53, TWIST1 | 15 | ABL1(2), E2F1(2), MDM2(1), MYC(1), PIK3CA(168), PIK3R1(14), POLR1A(1), POLR1B(2), POLR1C(2), POLR1D(1), RB1(9), TP53(142) | 14867268 | 345 | 285 | 157 | 11 | 72 | 53 | 112 | 11 | 97 | 0 | <1.00e-15 | <5.98e-15 |
| 6 | HSA04910_INSULIN_SIGNALING_PATHWAY | Genes involved in insulin signaling pathway | ACACA, ACACB, AKT1, AKT2, AKT3, ARAF, BAD, BRAF, CALM1, CALM2, CALM3, CALML3, CALML6, CBL, CBLB, CBLC, CRK, CRKL, EIF4EBP1, ELK1, EXOC7, FASN, FBP1, FBP2, FLOT1, FLOT2, FOXO1, FRAP1, G6PC, G6PC2, GCK, GRB2, GSK3B, GYS1, GYS2, HRAS, IKBKB, INPP5D, INS, INSR, IRS1, IRS2, IRS4, KIAA1303, KRAS, LIPE, MAP2K1, MAP2K2, MAPK1, MAPK10, MAPK3, MAPK8, MAPK9, MKNK1, MKNK2, NRAS, PCK1, PCK2, PDE3A, PDE3B, PDPK1, PFKL, PFKM, PFKP, PHKA1, PHKA2, PHKB, PHKG1, PHKG2, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PKLR, PKM2, PPARGC1A, PPP1CA, PPP1CB, PPP1CC, PPP1R3A, PPP1R3B, PPP1R3C, PPP1R3D, PRKAA1, PRKAA2, PRKAB1, PRKAB2, PRKACA, PRKACB, PRKACG, PRKAG1, PRKAG2, PRKAG3, PRKAR1A, PRKAR1B, PRKAR2A, PRKAR2B, PRKCI, PRKCZ, PRKX, PRKY, PTPN1, PTPRF, PYGB, PYGL, PYGM, RAF1, RAPGEF1, RHEB, RHOQ, RPS6, RPS6KB1, RPS6KB2, SH2B2, SHC1, SHC2, SHC3, SHC4, SKIP, SLC2A4, SOCS1, SOCS2, SOCS3, SOCS4, SORBS1, SOS1, SOS2, SREBF1, TRIP10, TSC1, TSC2 | 123 | ACACA(1), ACACB(8), AKT1(12), AKT2(1), AKT3(2), ARAF(1), BAD(1), BRAF(2), CBLB(7), CBLC(3), CRKL(1), ELK1(2), G6PC2(1), GCK(2), GRB2(1), GYS1(5), GYS2(1), IKBKB(2), INPP5D(2), INSR(2), IRS1(1), IRS4(1), KRAS(1), LIPE(2), MAP2K1(1), MAPK1(1), MAPK10(1), MAPK3(1), MAPK8(2), PCK1(1), PCK2(1), PDE3A(6), PDE3B(1), PFKM(3), PFKP(2), PHKA2(4), PHKB(2), PHKG2(1), PIK3CA(168), PIK3CB(3), PIK3CG(2), PIK3R1(14), PIK3R3(1), PPARGC1A(1), PPP1R3A(6), PPP1R3B(1), PRKAA2(2), PRKAB1(1), PRKACA(1), PRKAG2(1), PRKAG3(2), PRKAR1A(1), PRKAR2A(1), PRKAR2B(1), PRKCI(1), PRKCZ(1), PTPRF(3), PYGB(3), PYGL(1), PYGM(2), RAF1(1), RAPGEF1(3), RPS6(1), RPS6KB2(3), SHC1(1), SHC3(1), SHC4(2), SLC2A4(1), SOCS4(1), SORBS1(4), SOS1(3), SOS2(1), TRIP10(3), TSC1(2), TSC2(2) | 107951961 | 335 | 261 | 185 | 44 | 86 | 81 | 112 | 14 | 42 | 0 | <1.00e-15 | <5.98e-15 |
| 7 | HSA04070_PHOSPHATIDYLINOSITOL_SIGNALING_SYSTEM | Genes involved in phosphatidylinositol signaling system | CALM1, CALM2, CALM3, CALML3, CALML6, CARKL, CDIPT, CDS1, CDS2, DGKA, DGKB, DGKD, DGKE, DGKG, DGKH, DGKI, DGKQ, DGKZ, FN3K, IMPA1, IMPA2, INPP1, INPP4A, INPP4B, INPP5A, INPP5B, INPP5D, INPP5E, INPPL1, ITGB1BP3, ITPK1, ITPKA, ITPKB, ITPR1, ITPR2, ITPR3, OCRL, PI4KA, PI4KB, PIB5PA, PIK3C2A, PIK3C2B, PIK3C2G, PIK3C3, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PIP4K2A, PIP4K2B, PIP4K2C, PIP5K1A, PIP5K1B, PIP5K1C, PIP5K3, PLCB1, PLCB2, PLCB3, PLCB4, PLCD1, PLCD3, PLCD4, PLCE1, PLCG1, PLCG2, PLCZ1, PRKCA, PRKCB1, PRKCG, PTEN, PTPMT1, SKIP, SYNJ1, SYNJ2 | 70 | CDS1(2), DGKA(3), DGKD(1), DGKE(1), DGKG(4), DGKH(1), DGKI(5), IMPA1(1), INPP4A(2), INPP4B(3), INPP5B(4), INPP5D(2), INPPL1(1), ITGB1BP3(1), ITPK1(1), ITPKB(7), ITPR1(7), ITPR3(4), OCRL(3), PI4KA(3), PI4KB(1), PIK3C2A(2), PIK3C2B(2), PIK3C2G(1), PIK3C3(2), PIK3CA(168), PIK3CB(3), PIK3CG(2), PIK3R1(14), PIK3R3(1), PIP4K2A(1), PIP4K2C(5), PIP5K1A(1), PIP5K1C(3), PLCB1(3), PLCB2(2), PLCB3(1), PLCB4(6), PLCD1(1), PLCD3(2), PLCE1(5), PLCG1(4), PLCG2(2), PLCZ1(1), PRKCG(1), PTEN(15), SYNJ2(1) | 89527581 | 306 | 254 | 165 | 39 | 77 | 50 | 113 | 14 | 50 | 2 | <1.00e-15 | <5.98e-15 |
| 8 | HSA04630_JAK_STAT_SIGNALING_PATHWAY | Genes involved in Jak-STAT signaling pathway | AKT1, AKT2, AKT3, BCL2L1, CBL, CBLB, CBLC, CCND1, CCND2, CCND3, CISH, CLCF1, CNTF, CNTFR, CREBBP, CRLF2, CSF2, CSF2RA, CSF2RB, CSF3, CSF3R, CTF1, EP300, EPO, EPOR, GH1, GH2, GHR, GRB2, IFNA1, IFNA10, IFNA13, IFNA14, IFNA16, IFNA17, IFNA2, IFNA21, IFNA4, IFNA5, IFNA6, IFNA7, IFNA8, IFNAR1, IFNAR2, IFNB1, IFNE1, IFNG, IFNGR1, IFNGR2, IFNK, IFNW1, IL10, IL10RA, IL10RB, IL11, IL11RA, IL12A, IL12B, IL12RB1, IL12RB2, IL13, IL13RA1, IL13RA2, IL15, IL15RA, IL19, IL2, IL20, IL20RA, IL21, IL21R, IL22, IL22RA1, IL22RA2, IL23A, IL23R, IL24, IL26, IL28A, IL28B, IL28RA, IL29, IL2RA, IL2RB, IL2RG, IL3, IL3RA, IL4, IL4R, IL5, IL5RA, IL6, IL6R, IL6ST, IL7, IL7R, IL9, IL9R, IRF9, JAK1, JAK2, JAK3, LEP, LEPR, LIF, LIFR, MPL, MYC, OSM, OSMR, PIAS1, PIAS2, PIAS3, PIAS4, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PIM1, PRL, PRLR, PTPN11, PTPN6, SOCS1, SOCS2, SOCS3, SOCS4, SOCS5, SOCS7, SOS1, SOS2, SPRED1, SPRED2, SPRY1, SPRY2, SPRY3, SPRY4, STAM, STAM2, STAT1, STAT2, STAT3, STAT4, STAT5A, STAT5B, STAT6, TPO, TSLP, TYK2 | 148 | AKT1(12), AKT2(1), AKT3(2), CBLB(7), CBLC(3), CCND1(1), CCND3(5), CNTFR(1), CREBBP(2), CSF2RA(1), CSF2RB(2), CSF3R(2), EP300(3), EPOR(1), GH2(1), GHR(2), GRB2(1), IFNA10(1), IFNA14(1), IFNA2(2), IFNA4(1), IFNAR2(1), IFNB1(1), IFNK(1), IL10RA(1), IL10RB(3), IL11RA(1), IL12A(1), IL12B(1), IL12RB1(2), IL13RA1(1), IL13RA2(2), IL15(1), IL2(1), IL21(1), IL21R(1), IL22(1), IL22RA1(1), IL22RA2(1), IL23R(3), IL28B(3), IL28RA(1), IL2RG(1), IL3RA(1), IL5RA(1), IL6(2), IL6R(1), IL6ST(3), IRF9(2), JAK1(1), JAK2(2), JAK3(1), LEPR(3), LIF(3), LIFR(4), MYC(1), OSMR(3), PIAS2(1), PIAS4(2), PIK3CA(168), PIK3CB(3), PIK3CG(2), PIK3R1(14), PIK3R3(1), PRLR(1), SOCS4(1), SOCS5(1), SOS1(3), SOS2(1), SPRED1(2), SPRED2(1), STAM(1), STAM2(1), STAT1(1), STAT2(1), STAT3(1), STAT4(5), STAT5B(4), TPO(6), TSLP(1), TYK2(3) | 106057302 | 335 | 251 | 183 | 40 | 87 | 64 | 116 | 18 | 47 | 3 | <1.00e-15 | <5.98e-15 |
| 9 | HSA04012_ERBB_SIGNALING_PATHWAY | Genes involved in ErbB signaling pathway | ABL1, ABL2, AKT1, AKT2, AKT3, ARAF, AREG, BAD, BRAF, BTC, CAMK2A, CAMK2B, CAMK2D, CAMK2G, CBL, CBLB, CBLC, CDKN1A, CDKN1B, CRK, CRKL, EGF, EGFR, EIF4EBP1, ELK1, ERBB2, ERBB3, ERBB4, EREG, FRAP1, GAB1, GRB2, GSK3B, HBEGF, HRAS, JUN, KRAS, MAP2K1, MAP2K2, MAP2K4, MAP2K7, MAPK1, MAPK10, MAPK3, MAPK8, MAPK9, MYC, NCK1, NCK2, NRAS, NRG1, NRG2, NRG3, NRG4, PAK1, PAK2, PAK3, PAK4, PAK6, PAK7, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PLCG1, PLCG2, PRKCA, PRKCB1, PRKCG, PTK2, RAF1, RPS6KB1, RPS6KB2, SHC1, SHC2, SHC3, SHC4, SOS1, SOS2, SRC, STAT5A, STAT5B, TGFA | 82 | ABL1(2), ABL2(2), AKT1(12), AKT2(1), AKT3(2), ARAF(1), BAD(1), BRAF(2), BTC(1), CAMK2A(1), CAMK2D(3), CBLB(7), CBLC(3), CDKN1B(5), CRKL(1), EGF(2), EGFR(4), ELK1(2), ERBB2(4), ERBB3(8), ERBB4(4), GAB1(1), GRB2(1), JUN(1), KRAS(1), MAP2K1(1), MAP2K4(18), MAPK1(1), MAPK10(1), MAPK3(1), MAPK8(2), MYC(1), NRG3(1), PAK1(2), PAK2(1), PAK3(1), PAK7(1), PIK3CA(168), PIK3CB(3), PIK3CG(2), PIK3R1(14), PIK3R3(1), PLCG1(4), PLCG2(2), PRKCG(1), PTK2(1), RAF1(1), RPS6KB2(3), SHC1(1), SHC3(1), SHC4(2), SOS1(3), SOS2(1), STAT5B(4), TGFA(1) | 72661719 | 316 | 246 | 164 | 29 | 72 | 58 | 116 | 19 | 51 | 0 | <1.00e-15 | <5.98e-15 |
| 10 | ST_INTEGRIN_SIGNALING_PATHWAY | Integrins are transmembrane receptors that mediate cell growth, survival, and migration by binding to ligands in the extracellular matrix. | ABL1, ACK1, ACTN1, ACTR2, ACTR3, AKT1, AKT2, AKT3, ANGPTL2, ARHGEF6, ARHGEF7, BCAR1, BRAF, CAV1, CDC42, CDKN2A, CRK, CSE1L, DDEF1, DOCK1, EPHB2, FYN, GRAF, GRB2, GRB7, GRF2, GRLF1, ILK, ITGA1, ITGA10, ITGA11, ITGA2, ITGA3, ITGA4, ITGA5, ITGA6, ITGA7, ITGA8, ITGA9, ITGB3BP, MAP2K4, MAP2K7, MAP3K11, MAPK1, MAPK10, MAPK8, MAPK8IP1, MAPK8IP2, MAPK8IP3, MAPK9, MRAS, MYLK, MYLK2, P4HB, PAK1, PAK2, PAK3, PAK4, PAK6, PAK7, PIK3CA, PIK3CB, PKLR, PLCG1, PLCG2, PTEN, PTK2, RAF1, RALA, RHO, ROCK1, ROCK2, SHC1, SOS1, SOS2, SRC, TERF2IP, TLN1, TLN2, VASP, WAS, ZYX | 74 | ABL1(2), AKT1(12), AKT2(1), AKT3(2), ARHGEF6(1), ARHGEF7(2), BRAF(2), CAV1(1), CSE1L(2), EPHB2(1), GRB2(1), GRB7(3), GRLF1(4), ILK(2), ITGA1(3), ITGA10(1), ITGA11(4), ITGA2(2), ITGA4(1), ITGA6(3), ITGA7(5), ITGA8(2), ITGB3BP(1), MAP2K4(18), MAP3K11(1), MAPK1(1), MAPK10(1), MAPK8(2), MAPK8IP1(1), MYLK(5), PAK1(2), PAK2(1), PAK3(1), PAK7(1), PIK3CA(168), PIK3CB(3), PLCG1(4), PLCG2(2), PTEN(15), PTK2(1), RAF1(1), RHO(2), ROCK1(1), ROCK2(2), SHC1(1), SOS1(3), SOS2(1), TERF2IP(2), TLN1(7), TLN2(4), WAS(1) | 87730266 | 310 | 241 | 160 | 37 | 74 | 59 | 111 | 10 | 54 | 2 | <1.00e-15 | <5.98e-15 |
Table 7. Get Full Table A Ranked List of Significantly Mutated Genesets (Excluding Significantly Mutated Genes). Number of significant genesets found: 16. Number of genesets displayed: 10
| rank | geneset | description | genes | N_genes | mut_tally | N | n | npat | nsite | nsil | n1 | n2 | n3 | n4 | n5 | n6 | p | q |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HSA04510_FOCAL_ADHESION | Genes involved in focal adhesion | ACTB, ACTG1, ACTN1, ACTN2, ACTN3, ACTN4, AKT1, AKT2, AKT3, ARHGAP5, BAD, BCAR1, BCL2, BIRC2, BIRC3, BIRC4, BRAF, CAPN2, CAV1, CAV2, CAV3, CCND1, CCND2, CCND3, CDC42, CHAD, COL11A1, COL11A2, COL1A1, COL1A2, COL2A1, COL3A1, COL4A1, COL4A2, COL4A4, COL4A6, COL5A1, COL5A2, COL5A3, COL6A1, COL6A2, COL6A3, COL6A6, COMP, CRK, CRKL, CTNNB1, DIAPH1, DOCK1, EGF, EGFR, ELK1, ERBB2, FARP2, FIGF, FLNA, FLNB, FLNC, FLT1, FN1, FYN, GRB2, GRLF1, GSK3B, HGF, HRAS, IBSP, IGF1, IGF1R, ILK, ITGA1, ITGA10, ITGA11, ITGA2, ITGA2B, ITGA3, ITGA4, ITGA5, ITGA6, ITGA7, ITGA8, ITGA9, ITGAV, ITGB1, ITGB3, ITGB4, ITGB5, ITGB6, ITGB7, ITGB8, JUN, KDR, LAMA1, LAMA2, LAMA3, LAMA4, LAMA5, LAMB1, LAMB2, LAMB3, LAMB4, LAMC1, LAMC2, LAMC3, LOC653852, MAP2K1, MAPK1, MAPK10, MAPK3, MAPK8, MAPK9, MET, MLCK, MRCL3, MRLC2, MYL2, MYL5, MYL7, MYL8P, MYL9, MYLC2PL, MYLK, MYLK2, MYLPF, PAK1, PAK2, PAK3, PAK4, PAK6, PAK7, PARVA, PARVB, PARVG, PDGFA, PDGFB, PDGFC, PDGFD, PDGFRA, PDGFRB, PDPK1, PGF, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PIP5K1C, PPP1CA, PPP1CB, PPP1CC, PPP1R12A, PRKCA, PRKCB1, PRKCG, PTEN, PTK2, PXN, RAC1, RAC2, RAC3, RAF1, RAP1A, RAP1B, RAPGEF1, RELN, RHOA, ROCK1, ROCK2, SHC1, SHC2, SHC3, SHC4, SOS1, SOS2, SPP1, SRC, THBS1, THBS2, THBS3, THBS4, TLN1, TLN2, TNC, TNN, TNR, TNXB, VASP, VAV1, VAV2, VAV3, VCL, VEGFA, VEGFB, VEGFC, VTN, VWF, ZYX | 181 | ACTB(2), ACTG1(1), ACTN2(4), ACTN3(1), ACTN4(4), AKT2(1), AKT3(2), ARHGAP5(3), BAD(1), BRAF(2), CAPN2(2), CAV1(1), CCND1(1), CCND3(5), COL11A1(2), COL11A2(1), COL1A1(6), COL1A2(3), COL2A1(3), COL3A1(1), COL4A1(4), COL4A2(2), COL4A4(1), COL4A6(3), COL5A1(2), COL5A2(4), COL5A3(4), COL6A3(5), COL6A6(4), COMP(3), CRKL(1), EGF(2), EGFR(4), ELK1(2), ERBB2(4), FARP2(3), FIGF(1), FLNA(4), FLNB(7), FLNC(8), FLT1(2), FN1(3), GRB2(1), GRLF1(4), HGF(3), IGF1(1), IGF1R(1), ILK(2), ITGA1(3), ITGA10(1), ITGA11(4), ITGA2(2), ITGA2B(2), ITGA4(1), ITGA6(3), ITGA7(5), ITGA8(2), ITGAV(3), ITGB1(2), ITGB3(2), ITGB4(4), ITGB5(2), ITGB8(1), JUN(1), KDR(2), LAMA1(6), LAMA2(7), LAMA3(6), LAMA4(4), LAMB1(3), LAMB2(3), LAMB3(4), LAMB4(6), LAMC1(2), LAMC2(1), LAMC3(1), MAP2K1(1), MAPK1(1), MAPK10(1), MAPK3(1), MAPK8(2), MET(4), MYL7(1), MYLK(5), PAK1(2), PAK2(1), PAK3(1), PAK7(1), PARVA(1), PDGFB(1), PDGFRA(3), PDGFRB(2), PIK3CB(3), PIK3CG(2), PIK3R3(1), PIP5K1C(3), PPP1R12A(1), PRKCG(1), PTK2(1), PXN(1), RAF1(1), RAPGEF1(3), RELN(10), RHOA(2), ROCK1(1), ROCK2(2), SHC1(1), SHC3(1), SHC4(2), SOS1(3), SOS2(1), SPP1(1), THBS1(3), THBS3(1), THBS4(1), TLN1(7), TLN2(4), TNC(3), TNN(3), TNR(1), TNXB(3), VAV1(1), VAV3(1), VCL(3), VWF(6) | 259172823 | 320 | 212 | 319 | 111 | 57 | 101 | 70 | 27 | 59 | 6 | <1.00e-15 | <6.16e-13 |
| 2 | HSA01430_CELL_COMMUNICATION | Genes involved in cell communication | ACTB, ACTG1, CHAD, COL11A1, COL11A2, COL17A1, COL1A1, COL1A2, COL2A1, COL3A1, COL4A1, COL4A2, COL4A4, COL4A6, COL5A1, COL5A2, COL5A3, COL6A1, COL6A2, COL6A3, COL6A6, COMP, DES, DSC1, DSC2, DSC3, DSG1, DSG2, DSG3, DSG4, FN1, GJA1, GJA10, GJA3, GJA4, GJA5, GJA8, GJA9, GJB1, GJB2, GJB3, GJB4, GJB5, GJB6, GJB7, GJC1, GJC2, GJC3, GJD2, GJD3, GJD4, IBSP, INA, ITGA6, ITGB4, KRT1, KRT10, KRT12, KRT13, KRT14, KRT15, KRT16, KRT17, KRT18, KRT19, KRT2, KRT20, KRT23, KRT24, KRT25, KRT27, KRT28, KRT3, KRT31, KRT32, KRT33A, KRT33B, KRT34, KRT35, KRT36, KRT37, KRT38, KRT39, KRT4, KRT40, KRT5, KRT6A, KRT6B, KRT6C, KRT7, KRT71, KRT72, KRT73, KRT74, KRT75, KRT76, KRT77, KRT78, KRT79, KRT8, KRT81, KRT82, KRT83, KRT84, KRT85, KRT86, KRT9, LAMA1, LAMA2, LAMA3, LAMA4, LAMA5, LAMB1, LAMB2, LAMB3, LAMB4, LAMC1, LAMC2, LAMC3, LMNA, LMNB1, LMNB2, LOC728760, NES, PRPH, RELN, SPP1, THBS1, THBS2, THBS3, THBS4, TNC, TNN, TNR, TNXB, VIM, VTN, VWF | 126 | ACTB(2), ACTG1(1), COL11A1(2), COL11A2(1), COL17A1(3), COL1A1(6), COL1A2(3), COL2A1(3), COL3A1(1), COL4A1(4), COL4A2(2), COL4A4(1), COL4A6(3), COL5A1(2), COL5A2(4), COL5A3(4), COL6A3(5), COL6A6(4), COMP(3), DSC2(3), DSC3(1), DSG1(2), DSG4(2), FN1(3), GJA1(2), GJA10(1), GJA4(1), GJA9(1), GJB2(1), GJB3(2), GJB4(1), GJB5(1), GJC1(1), INA(1), ITGA6(3), ITGB4(4), KRT1(3), KRT14(1), KRT15(1), KRT17(4), KRT18(1), KRT19(2), KRT2(2), KRT25(2), KRT28(1), KRT3(1), KRT33A(1), KRT34(1), KRT35(1), KRT36(2), KRT37(1), KRT4(2), KRT6A(1), KRT6C(1), KRT71(1), KRT72(1), KRT81(1), KRT82(2), KRT83(1), KRT84(1), KRT85(1), KRT86(1), KRT9(2), LAMA1(6), LAMA2(7), LAMA3(6), LAMA4(4), LAMB1(3), LAMB2(3), LAMB3(4), LAMB4(6), LAMC1(2), LAMC2(1), LAMC3(1), LMNB1(1), NES(3), RELN(10), SPP1(1), THBS1(3), THBS3(1), THBS4(1), TNC(3), TNN(3), TNR(1), TNXB(3), VWF(6) | 166605270 | 202 | 161 | 201 | 72 | 41 | 74 | 42 | 10 | 35 | 0 | 5.43e-08 | 0.000017 |
| 3 | HSA04350_TGF_BETA_SIGNALING_PATHWAY | Genes involved in TGF-beta signaling pathway | ACVR1, ACVR1B, ACVR1C, ACVR2A, ACVR2B, ACVRL1, AMH, AMHR2, BMP2, BMP4, BMP5, BMP6, BMP7, BMP8A, BMP8B, BMPR1A, BMPR1B, BMPR2, CDKN2B, CHRD, COMP, CREBBP, CUL1, DCN, E2F4, E2F5, EP300, FST, GDF5, GDF6, GDF7, hCG_1982709, ID1, ID2, ID3, ID4, IFNG, INHBA, INHBB, INHBC, INHBE, LEFTY1, LEFTY2, LTBP1, MAPK1, MAPK3, MYC, NODAL, NOG, PITX2, PPP2CA, PPP2CB, PPP2R1A, PPP2R1B, PPP2R2A, PPP2R2B, PPP2R2C, RBL1, RBL2, RBX1, RHOA, ROCK1, ROCK2, RPS6KB1, RPS6KB2, SKP1, SMAD1, SMAD2, SMAD3, SMAD4, SMAD5, SMAD6, SMAD7, SMAD9, SMURF1, SMURF2, SP1, TFDP1, TGFB1, TGFB2, TGFB3, TGFBR1, TGFBR2, THBS1, THBS2, THBS3, THBS4, TNF, ZFYVE16, ZFYVE9 | 84 | ACVR1(1), ACVR1B(4), ACVR1C(1), ACVR2A(3), AMHR2(2), BMP2(2), BMP4(2), BMP5(1), BMP6(1), BMP8B(1), BMPR1B(2), BMPR2(2), CHRD(2), COMP(3), CREBBP(2), CUL1(1), E2F4(2), EP300(3), FST(1), GDF6(2), ID2(1), INHBA(1), INHBC(1), INHBE(1), LEFTY2(1), LTBP1(2), MAPK1(1), MAPK3(1), MYC(1), NODAL(1), PITX2(1), PPP2CB(2), RBL1(1), RBL2(1), RHOA(2), ROCK1(1), ROCK2(2), RPS6KB2(3), SMAD2(3), SMAD3(2), SMAD4(1), SMAD9(2), SMURF2(1), SP1(1), TGFB3(1), TGFBR1(2), THBS1(3), THBS3(1), THBS4(1), ZFYVE16(3), ZFYVE9(3) | 70610904 | 86 | 77 | 84 | 29 | 13 | 23 | 18 | 7 | 25 | 0 | 2.99e-06 | 0.00061 |
| 4 | HSA04010_MAPK_SIGNALING_PATHWAY | Genes involved in MAPK signaling pathway | ACVR1B, ACVR1C, AKT1, AKT2, AKT3, ARRB1, ARRB2, ATF2, ATF4, BDNF, BRAF, CACNA1A, CACNA1B, CACNA1C, CACNA1D, CACNA1E, CACNA1F, CACNA1G, CACNA1H, CACNA1I, CACNA1S, CACNA2D1, CACNA2D2, CACNA2D3, CACNA2D4, CACNB1, CACNB2, CACNB3, CACNB4, CACNG1, CACNG2, CACNG3, CACNG4, CACNG5, CACNG6, CACNG7, CACNG8, CASP3, CD14, CDC25B, CDC42, CHP, CHUK, CRK, CRKL, DAXX, DDIT3, DUSP1, DUSP10, DUSP14, DUSP16, DUSP2, DUSP3, DUSP4, DUSP5, DUSP6, DUSP7, DUSP8, DUSP9, ECSIT, EGF, EGFR, ELK1, ELK4, EVI1, FAS, FASLG, FGF1, FGF10, FGF11, FGF12, FGF13, FGF14, FGF16, FGF17, FGF18, FGF19, FGF2, FGF20, FGF21, FGF22, FGF23, FGF3, FGF4, FGF5, FGF6, FGF7, FGF8, FGF9, FGFR1, FGFR2, FGFR3, FGFR4, FLNA, FLNB, FLNC, FOS, GADD45A, GADD45B, GADD45G, GNA12, GNG12, GRB2, HRAS, IKBKB, IKBKG, IL1A, IL1B, IL1R1, IL1R2, JUN, JUND, KRAS, LOC653852, MAP2K1, MAP2K1IP1, MAP2K2, MAP2K3, MAP2K4, MAP2K5, MAP2K6, MAP2K7, MAP3K1, MAP3K10, MAP3K12, MAP3K13, MAP3K14, MAP3K2, MAP3K3, MAP3K4, MAP3K5, MAP3K6, MAP3K7, MAP3K7IP1, MAP3K7IP2, MAP3K8, MAP4K1, MAP4K2, MAP4K3, MAP4K4, MAPK1, MAPK10, MAPK11, MAPK12, MAPK13, MAPK14, MAPK3, MAPK7, MAPK8, MAPK8IP1, MAPK8IP2, MAPK8IP3, MAPK9, MAPKAPK2, MAPKAPK3, MAPKAPK5, MAPT, MAX, MEF2C, MKNK1, MKNK2, MOS, MRAS, MYC, NF1, NFATC2, NFATC4, NFKB1, NFKB2, NGFB, NLK, NR4A1, NRAS, NTF3, NTF5, NTRK1, NTRK2, PAK1, PAK2, PDGFA, PDGFB, PDGFRA, PDGFRB, PLA2G10, PLA2G12A, PLA2G12B, PLA2G1B, PLA2G2A, PLA2G2D, PLA2G2E, PLA2G2F, PLA2G3, PLA2G4A, PLA2G5, PLA2G6, PPM1A, PPM1B, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PPP5C, PRKACA, PRKACB, PRKACG, PRKCA, PRKCB1, PRKCG, PRKX, PRKY, PTPN5, PTPN7, PTPRR, RAC1, RAC2, RAC3, RAF1, RAP1A, RAP1B, RAPGEF2, RASA1, RASA2, RASGRF1, RASGRF2, RASGRP1, RASGRP2, RASGRP3, RASGRP4, RPS6KA1, RPS6KA2, RPS6KA3, RPS6KA4, RPS6KA5, RPS6KA6, RRAS, RRAS2, SOS1, SOS2, SRF, STK3, STK4, STMN1, TAOK1, TAOK2, TAOK3, TGFB1, TGFB2, TGFB3, TGFBR1, TGFBR2, TNF, TNFRSF1A, TP53, TRAF2, TRAF6, ZAK | 226 | ACVR1B(4), ACVR1C(1), AKT2(1), AKT3(2), ARRB1(1), ATF4(1), BDNF(1), BRAF(2), CACNA1A(5), CACNA1B(1), CACNA1C(4), CACNA1D(4), CACNA1E(8), CACNA1F(6), CACNA1G(5), CACNA1I(2), CACNA1S(1), CACNA2D1(4), CACNA2D2(2), CACNA2D3(5), CACNA2D4(1), CACNB1(1), CACNB2(2), CACNG2(1), CACNG3(2), CACNG4(1), CACNG6(1), CD14(1), CDC25B(3), CRKL(1), DUSP10(1), DUSP16(3), DUSP6(1), DUSP7(1), DUSP9(1), EGF(2), EGFR(4), ELK1(2), FAS(1), FASLG(1), FGF11(1), FGF13(2), FGF18(2), FGF23(1), FGF3(1), FGF5(1), FGF6(1), FGF7(2), FGFR2(2), FGFR4(3), FLNA(4), FLNB(7), FLNC(8), GRB2(1), IKBKB(2), IL1A(1), IL1R1(1), IL1R2(3), JUN(1), KRAS(1), MAP2K1(1), MAP2K3(1), MAP2K5(1), MAP3K10(4), MAP3K12(2), MAP3K13(3), MAP3K4(3), MAP3K5(1), MAP3K6(1), MAP3K8(2), MAP4K1(1), MAP4K4(1), MAPK1(1), MAPK10(1), MAPK14(1), MAPK3(1), MAPK8(2), MAPK8IP1(1), MAPKAPK3(1), MAPT(1), MAX(1), MEF2C(2), MOS(2), MYC(1), NFATC4(3), NFKB1(1), NFKB2(4), NLK(1), NR4A1(1), NTRK1(1), NTRK2(1), PAK1(2), PAK2(1), PDGFB(1), PDGFRA(3), PDGFRB(2), PLA2G2D(1), PLA2G2F(1), PLA2G3(3), PLA2G4A(6), PLA2G6(2), PPP3CA(3), PPP3CB(2), PRKACA(1), PRKCG(1), PTPN5(2), PTPN7(1), PTPRR(1), RAF1(1), RAPGEF2(1), RASA1(2), RASA2(1), RASGRF1(3), RASGRP2(2), RASGRP4(1), RPS6KA1(1), RPS6KA2(2), RPS6KA3(2), RPS6KA4(1), RPS6KA6(3), SOS1(3), SOS2(1), SRF(1), STK3(1), STK4(1), TAOK1(2), TAOK2(1), TGFB3(1), TGFBR1(2), TNFRSF1A(1), TRAF6(1), ZAK(1) | 193925472 | 253 | 175 | 250 | 69 | 54 | 88 | 46 | 18 | 44 | 3 | 9.28e-06 | 0.0014 |
| 5 | HSA04012_ERBB_SIGNALING_PATHWAY | Genes involved in ErbB signaling pathway | ABL1, ABL2, AKT1, AKT2, AKT3, ARAF, AREG, BAD, BRAF, BTC, CAMK2A, CAMK2B, CAMK2D, CAMK2G, CBL, CBLB, CBLC, CDKN1A, CDKN1B, CRK, CRKL, EGF, EGFR, EIF4EBP1, ELK1, ERBB2, ERBB3, ERBB4, EREG, FRAP1, GAB1, GRB2, GSK3B, HBEGF, HRAS, JUN, KRAS, MAP2K1, MAP2K2, MAP2K4, MAP2K7, MAPK1, MAPK10, MAPK3, MAPK8, MAPK9, MYC, NCK1, NCK2, NRAS, NRG1, NRG2, NRG3, NRG4, PAK1, PAK2, PAK3, PAK4, PAK6, PAK7, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PLCG1, PLCG2, PRKCA, PRKCB1, PRKCG, PTK2, RAF1, RPS6KB1, RPS6KB2, SHC1, SHC2, SHC3, SHC4, SOS1, SOS2, SRC, STAT5A, STAT5B, TGFA | 77 | ABL1(2), ABL2(2), AKT2(1), AKT3(2), ARAF(1), BAD(1), BRAF(2), BTC(1), CAMK2A(1), CAMK2D(3), CBLB(7), CBLC(3), CRKL(1), EGF(2), EGFR(4), ELK1(2), ERBB2(4), ERBB3(8), ERBB4(4), GAB1(1), GRB2(1), JUN(1), KRAS(1), MAP2K1(1), MAPK1(1), MAPK10(1), MAPK3(1), MAPK8(2), MYC(1), NRG3(1), PAK1(2), PAK2(1), PAK3(1), PAK7(1), PIK3CB(3), PIK3CG(2), PIK3R3(1), PLCG1(4), PLCG2(2), PRKCG(1), PTK2(1), RAF1(1), RPS6KB2(3), SHC1(1), SHC3(1), SHC4(2), SOS1(3), SOS2(1), STAT5B(4), TGFA(1) | 68217357 | 99 | 86 | 99 | 24 | 13 | 28 | 24 | 17 | 17 | 0 | 0.000018 | 0.0022 |
| 6 | HSA04910_INSULIN_SIGNALING_PATHWAY | Genes involved in insulin signaling pathway | ACACA, ACACB, AKT1, AKT2, AKT3, ARAF, BAD, BRAF, CALM1, CALM2, CALM3, CALML3, CALML6, CBL, CBLB, CBLC, CRK, CRKL, EIF4EBP1, ELK1, EXOC7, FASN, FBP1, FBP2, FLOT1, FLOT2, FOXO1, FRAP1, G6PC, G6PC2, GCK, GRB2, GSK3B, GYS1, GYS2, HRAS, IKBKB, INPP5D, INS, INSR, IRS1, IRS2, IRS4, KIAA1303, KRAS, LIPE, MAP2K1, MAP2K2, MAPK1, MAPK10, MAPK3, MAPK8, MAPK9, MKNK1, MKNK2, NRAS, PCK1, PCK2, PDE3A, PDE3B, PDPK1, PFKL, PFKM, PFKP, PHKA1, PHKA2, PHKB, PHKG1, PHKG2, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PKLR, PKM2, PPARGC1A, PPP1CA, PPP1CB, PPP1CC, PPP1R3A, PPP1R3B, PPP1R3C, PPP1R3D, PRKAA1, PRKAA2, PRKAB1, PRKAB2, PRKACA, PRKACB, PRKACG, PRKAG1, PRKAG2, PRKAG3, PRKAR1A, PRKAR1B, PRKAR2A, PRKAR2B, PRKCI, PRKCZ, PRKX, PRKY, PTPN1, PTPRF, PYGB, PYGL, PYGM, RAF1, RAPGEF1, RHEB, RHOQ, RPS6, RPS6KB1, RPS6KB2, SH2B2, SHC1, SHC2, SHC3, SHC4, SKIP, SLC2A4, SOCS1, SOCS2, SOCS3, SOCS4, SORBS1, SOS1, SOS2, SREBF1, TRIP10, TSC1, TSC2 | 120 | ACACA(1), ACACB(8), AKT2(1), AKT3(2), ARAF(1), BAD(1), BRAF(2), CBLB(7), CBLC(3), CRKL(1), ELK1(2), G6PC2(1), GCK(2), GRB2(1), GYS1(5), GYS2(1), IKBKB(2), INPP5D(2), INSR(2), IRS1(1), IRS4(1), KRAS(1), LIPE(2), MAP2K1(1), MAPK1(1), MAPK10(1), MAPK3(1), MAPK8(2), PCK1(1), PCK2(1), PDE3A(6), PDE3B(1), PFKM(3), PFKP(2), PHKA2(4), PHKB(2), PHKG2(1), PIK3CB(3), PIK3CG(2), PIK3R3(1), PPARGC1A(1), PPP1R3A(6), PPP1R3B(1), PRKAA2(2), PRKAB1(1), PRKACA(1), PRKAG2(1), PRKAG3(2), PRKAR1A(1), PRKAR2A(1), PRKAR2B(1), PRKCI(1), PRKCZ(1), PTPRF(3), PYGB(3), PYGL(1), PYGM(2), RAF1(1), RAPGEF1(3), RPS6(1), RPS6KB2(3), SHC1(1), SHC3(1), SHC4(2), SLC2A4(1), SOCS4(1), SORBS1(4), SOS1(3), SOS2(1), TRIP10(3), TSC1(2), TSC2(2) | 104354289 | 141 | 118 | 141 | 39 | 31 | 51 | 22 | 12 | 25 | 0 | 0.000051 | 0.0053 |
| 7 | HSA04630_JAK_STAT_SIGNALING_PATHWAY | Genes involved in Jak-STAT signaling pathway | AKT1, AKT2, AKT3, BCL2L1, CBL, CBLB, CBLC, CCND1, CCND2, CCND3, CISH, CLCF1, CNTF, CNTFR, CREBBP, CRLF2, CSF2, CSF2RA, CSF2RB, CSF3, CSF3R, CTF1, EP300, EPO, EPOR, GH1, GH2, GHR, GRB2, IFNA1, IFNA10, IFNA13, IFNA14, IFNA16, IFNA17, IFNA2, IFNA21, IFNA4, IFNA5, IFNA6, IFNA7, IFNA8, IFNAR1, IFNAR2, IFNB1, IFNE1, IFNG, IFNGR1, IFNGR2, IFNK, IFNW1, IL10, IL10RA, IL10RB, IL11, IL11RA, IL12A, IL12B, IL12RB1, IL12RB2, IL13, IL13RA1, IL13RA2, IL15, IL15RA, IL19, IL2, IL20, IL20RA, IL21, IL21R, IL22, IL22RA1, IL22RA2, IL23A, IL23R, IL24, IL26, IL28A, IL28B, IL28RA, IL29, IL2RA, IL2RB, IL2RG, IL3, IL3RA, IL4, IL4R, IL5, IL5RA, IL6, IL6R, IL6ST, IL7, IL7R, IL9, IL9R, IRF9, JAK1, JAK2, JAK3, LEP, LEPR, LIF, LIFR, MPL, MYC, OSM, OSMR, PIAS1, PIAS2, PIAS3, PIAS4, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PIM1, PRL, PRLR, PTPN11, PTPN6, SOCS1, SOCS2, SOCS3, SOCS4, SOCS5, SOCS7, SOS1, SOS2, SPRED1, SPRED2, SPRY1, SPRY2, SPRY3, SPRY4, STAM, STAM2, STAT1, STAT2, STAT3, STAT4, STAT5A, STAT5B, STAT6, TPO, TSLP, TYK2 | 145 | AKT2(1), AKT3(2), CBLB(7), CBLC(3), CCND1(1), CCND3(5), CNTFR(1), CREBBP(2), CSF2RA(1), CSF2RB(2), CSF3R(2), EP300(3), EPOR(1), GH2(1), GHR(2), GRB2(1), IFNA10(1), IFNA14(1), IFNA2(2), IFNA4(1), IFNAR2(1), IFNB1(1), IFNK(1), IL10RA(1), IL10RB(3), IL11RA(1), IL12A(1), IL12B(1), IL12RB1(2), IL13RA1(1), IL13RA2(2), IL15(1), IL2(1), IL21(1), IL21R(1), IL22(1), IL22RA1(1), IL22RA2(1), IL23R(3), IL28B(3), IL28RA(1), IL2RG(1), IL3RA(1), IL5RA(1), IL6(2), IL6R(1), IL6ST(3), IRF9(2), JAK1(1), JAK2(2), JAK3(1), LEPR(3), LIF(3), LIFR(4), MYC(1), OSMR(3), PIAS2(1), PIAS4(2), PIK3CB(3), PIK3CG(2), PIK3R3(1), PRLR(1), SOCS4(1), SOCS5(1), SOS1(3), SOS2(1), SPRED1(2), SPRED2(1), STAM(1), STAM2(1), STAT1(1), STAT2(1), STAT3(1), STAT4(5), STAT5B(4), TPO(6), TSLP(1), TYK2(3) | 102459630 | 141 | 113 | 139 | 35 | 32 | 34 | 26 | 16 | 30 | 3 | 0.000063 | 0.0055 |
| 8 | HSA04320_DORSO_VENTRAL_AXIS_FORMATION | Genes involved in dorso-ventral axis formation | BRAF, CPEB1, EGFR, ERBB2, ERBB4, ETS1, ETS2, ETV6, ETV7, FMN2, GRB2, KRAS, MAP2K1, MAPK1, MAPK3, NOTCH1, NOTCH2, NOTCH3, NOTCH4, PIWIL1, PIWIL2, PIWIL3, PIWIL4, RAF1, SOS1, SOS2, SPIRE1, SPIRE2 | 25 | BRAF(2), CPEB1(2), EGFR(4), ERBB2(4), ERBB4(4), ETS1(2), ETS2(3), ETV6(1), FMN2(7), GRB2(1), KRAS(1), MAP2K1(1), MAPK1(1), MAPK3(1), NOTCH2(4), NOTCH3(3), NOTCH4(6), PIWIL2(4), PIWIL4(1), RAF1(1), SOS1(3), SOS2(1), SPIRE1(1) | 33021417 | 58 | 50 | 57 | 12 | 8 | 19 | 13 | 7 | 11 | 0 | 0.000083 | 0.0064 |
| 9 | HSA04512_ECM_RECEPTOR_INTERACTION | Genes involved in ECM-receptor interaction | AGRN, CD36, CD44, CD47, CHAD, COL11A1, COL11A2, COL1A1, COL1A2, COL2A1, COL3A1, COL4A1, COL4A2, COL4A4, COL4A6, COL5A1, COL5A2, COL5A3, COL6A1, COL6A2, COL6A3, COL6A6, DAG1, FN1, FNDC1, FNDC3A, FNDC4, FNDC5, GP1BA, GP1BB, GP5, GP6, GP9, HMMR, HSPG2, IBSP, ITGA1, ITGA10, ITGA11, ITGA2, ITGA2B, ITGA3, ITGA4, ITGA5, ITGA6, ITGA7, ITGA8, ITGA9, ITGAV, ITGB1, ITGB3, ITGB4, ITGB5, ITGB6, ITGB7, ITGB8, LAMA1, LAMA2, LAMA3, LAMA4, LAMA5, LAMB1, LAMB2, LAMB3, LAMB4, LAMC1, LAMC2, LAMC3, RELN, SDC1, SDC2, SDC3, SDC4, SPP1, SV2A, SV2B, SV2C, THBS1, THBS2, THBS3, THBS4, TNC, TNN, TNR, TNXB, VTN, VWF | 82 | CD36(1), COL11A1(2), COL11A2(1), COL1A1(6), COL1A2(3), COL2A1(3), COL3A1(1), COL4A1(4), COL4A2(2), COL4A4(1), COL4A6(3), COL5A1(2), COL5A2(4), COL5A3(4), COL6A3(5), COL6A6(4), FN1(3), FNDC1(4), FNDC3A(1), GP1BA(3), GP6(1), HSPG2(5), ITGA1(3), ITGA10(1), ITGA11(4), ITGA2(2), ITGA2B(2), ITGA4(1), ITGA6(3), ITGA7(5), ITGA8(2), ITGAV(3), ITGB1(2), ITGB3(2), ITGB4(4), ITGB5(2), ITGB8(1), LAMA1(6), LAMA2(7), LAMA3(6), LAMA4(4), LAMB1(3), LAMB2(3), LAMB3(4), LAMB4(6), LAMC1(2), LAMC2(1), LAMC3(1), RELN(10), SDC4(1), SPP1(1), SV2A(1), SV2B(1), THBS1(3), THBS3(1), THBS4(1), TNC(3), TNN(3), TNR(1), TNXB(3), VWF(6) | 157698294 | 178 | 142 | 177 | 70 | 34 | 61 | 44 | 8 | 31 | 0 | 0.00014 | 0.0096 |
| 10 | HSA04020_CALCIUM_SIGNALING_PATHWAY | Genes involved in calcium signaling pathway | ADCY1, ADCY2, ADCY3, ADCY4, ADCY7, ADCY8, ADCY9, ADORA2A, ADORA2B, ADRA1A, ADRA1B, ADRA1D, ADRB1, ADRB2, ADRB3, AGTR1, ATP2A1, ATP2A2, ATP2A3, ATP2B1, ATP2B2, ATP2B3, ATP2B4, AVPR1A, AVPR1B, BDKRB1, BDKRB2, BST1, CACNA1A, CACNA1B, CACNA1C, CACNA1D, CACNA1E, CACNA1F, CACNA1G, CACNA1H, CACNA1I, CACNA1S, CALM1, CALM2, CALM3, CALML3, CALML6, CAMK2A, CAMK2B, CAMK2D, CAMK2G, CAMK4, CCKAR, CCKBR, CD38, CHP, CHRM1, CHRM2, CHRM3, CHRM5, CHRNA7, CYSLTR1, CYSLTR2, DRD1, EDNRA, EDNRB, EGFR, ERBB2, ERBB3, ERBB4, F2R, GNA11, GNA14, GNA15, GNAL, GNAQ, GNAS, GRIN1, GRIN2A, GRIN2C, GRIN2D, GRM1, GRM5, GRPR, HRH1, HRH2, HTR2A, HTR2B, HTR2C, HTR4, HTR5A, HTR6, HTR7, ITPKA, ITPKB, ITPR1, ITPR2, ITPR3, LHCGR, LTB4R2, MLCK, MYLK, MYLK2, NOS1, NOS2A, NOS3, NTSR1, OXTR, P2RX1, P2RX2, P2RX3, P2RX4, P2RX5, P2RX7, P2RXL1, PDE1A, PDE1B, PDE1C, PDGFRA, PDGFRB, PHKA1, PHKA2, PHKB, PHKG1, PHKG2, PLCB1, PLCB2, PLCB3, PLCB4, PLCD1, PLCD3, PLCD4, PLCE1, PLCG1, PLCG2, PLCZ1, PLN, PPID, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PRKACA, PRKACB, PRKACG, PRKCA, PRKCB1, PRKCG, PRKX, PRKY, PTAFR, PTGER1, PTGER3, PTGFR, PTK2B, RYR1, RYR2, RYR3, SLC25A4, SLC25A5, SLC25A6, SLC8A1, SLC8A2, SLC8A3, SPHK1, SPHK2, TACR1, TACR2, TACR3, TBXA2R, TNNC1, TNNC2, TRHR, TRPC1, VDAC1, VDAC2, VDAC3 | 161 | ADCY1(3), ADCY3(2), ADCY4(1), ADCY7(1), ADCY8(2), ADCY9(5), ADRA1A(1), ADRA1B(1), ATP2A1(2), ATP2B1(4), ATP2B2(4), ATP2B3(4), ATP2B4(1), AVPR1B(1), BDKRB1(2), CACNA1A(5), CACNA1B(1), CACNA1C(4), CACNA1D(4), CACNA1E(8), CACNA1F(6), CACNA1G(5), CACNA1I(2), CACNA1S(1), CAMK2A(1), CAMK2D(3), CD38(2), CHRM1(2), CHRM2(2), CHRM3(1), CHRM5(2), CYSLTR1(1), CYSLTR2(1), EDNRB(1), EGFR(4), ERBB2(4), ERBB3(8), ERBB4(4), F2R(2), GNAQ(1), GNAS(4), GRIN1(1), GRIN2C(4), GRIN2D(3), GRM1(2), GRM5(1), GRPR(1), HRH2(2), HTR2A(3), HTR2B(1), HTR2C(1), HTR5A(3), ITPR1(7), ITPR3(4), LHCGR(4), LTB4R2(1), MYLK(5), NOS1(3), NOS3(2), P2RX3(1), P2RX4(1), P2RX5(1), P2RX7(1), PDE1A(2), PDE1B(1), PDGFRA(3), PDGFRB(2), PHKA2(4), PHKB(2), PHKG2(1), PLCB1(3), PLCB2(2), PLCB3(1), PLCB4(6), PLCD1(1), PLCD3(2), PLCE1(5), PLCG1(4), PLCG2(2), PLCZ1(1), PPP3CA(3), PPP3CB(2), PRKACA(1), PRKCG(1), PTGER3(2), PTGFR(1), PTK2B(3), RYR1(5), RYR2(13), RYR3(11), SPHK2(2), TACR2(1), TNNC2(1), TRPC1(2), VDAC2(1) | 191542572 | 256 | 175 | 255 | 108 | 73 | 79 | 54 | 19 | 27 | 4 | 0.00023 | 0.014 |
In brief, we tabulate the number of mutations and the number of covered bases for each gene. The counts are broken down by mutation context category: four context categories that are discovered by MutSig, and one for indel and 'null' mutations, which include indels, nonsense mutations, splice-site mutations, and non-stop (read-through) mutations. For each gene, we calculate the probability of seeing the observed constellation of mutations, i.e. the product P1 x P2 x ... x Pm, or a more extreme one, given the background mutation rates calculated across the dataset.[1]
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.