This pipeline computes the correlation between significantly recurrent gene mutations and selected clinical features.
Testing the association between mutation status of 89 genes and 8 clinical features across 293 patients, 3 significant findings detected with Q value < 0.25.
-
NUDT11 mutation correlated to 'AGE'.
-
FBXO43 mutation correlated to 'Time to Death'.
-
TMEM184C mutation correlated to 'AGE'.
Table 1. Get Full Table Overview of the association between mutation status of 89 genes and 8 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 3 significant findings detected.
|
Clinical Features |
Time to Death |
AGE | GENDER |
KARNOFSKY PERFORMANCE SCORE |
PATHOLOGY T |
PATHOLOGY N |
PATHOLOGICSPREAD(M) |
NEOADJUVANT THERAPY |
||
| nMutated (%) | nWild-Type | logrank test | t-test | Fisher's exact test | t-test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| NUDT11 | 3 (1%) | 290 |
0.6 (1.00) |
1.58e-24 (9.73e-22) |
0.554 (1.00) |
0.718 (1.00) |
1 (1.00) |
1 (1.00) |
||
| FBXO43 | 5 (2%) | 288 |
1.27e-13 (7.83e-11) |
0.621 (1.00) |
1 (1.00) |
0.446 (1.00) |
1 (1.00) |
0.513 (1.00) |
1 (1.00) |
|
| TMEM184C | 4 (1%) | 289 |
0.0172 (1.00) |
0.000391 (0.24) |
1 (1.00) |
0.796 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| VHL | 145 (49%) | 148 |
0.913 (1.00) |
0.0211 (1.00) |
0.113 (1.00) |
0.359 (1.00) |
0.498 (1.00) |
0.76 (1.00) |
0.732 (1.00) |
1 (1.00) |
| KDM5C | 18 (6%) | 275 |
0.0803 (1.00) |
0.206 (1.00) |
0.00486 (1.00) |
0.622 (1.00) |
0.516 (1.00) |
1 (1.00) |
1 (1.00) |
|
| PBRM1 | 110 (38%) | 183 |
0.439 (1.00) |
0.987 (1.00) |
0.129 (1.00) |
0.429 (1.00) |
0.613 (1.00) |
1 (1.00) |
0.478 (1.00) |
0.559 (1.00) |
| RRAD | 4 (1%) | 289 |
0.0836 (1.00) |
0.806 (1.00) |
0.123 (1.00) |
0.0613 (1.00) |
0.211 (1.00) |
0.437 (1.00) |
1 (1.00) |
|
| SV2C | 3 (1%) | 290 |
0.731 (1.00) |
0.0205 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| TOR1A | 3 (1%) | 290 |
0.0532 (1.00) |
0.704 (1.00) |
1 (1.00) |
1 (1.00) |
0.0753 (1.00) |
1 (1.00) |
1 (1.00) |
|
| SETD2 | 35 (12%) | 258 |
0.852 (1.00) |
0.123 (1.00) |
0.132 (1.00) |
0.101 (1.00) |
0.61 (1.00) |
0.0317 (1.00) |
0.318 (1.00) |
|
| PTEN | 15 (5%) | 278 |
0.87 (1.00) |
0.104 (1.00) |
1 (1.00) |
0.489 (1.00) |
0.0657 (1.00) |
1 (1.00) |
0.146 (1.00) |
|
| BAP1 | 28 (10%) | 265 |
0.0144 (1.00) |
0.695 (1.00) |
0.00345 (1.00) |
0.00203 (1.00) |
0.637 (1.00) |
0.00126 (0.771) |
0.261 (1.00) |
|
| PIK3CA | 14 (5%) | 279 |
0.797 (1.00) |
0.387 (1.00) |
0.57 (1.00) |
0.152 (1.00) |
0.555 (1.00) |
1 (1.00) |
1 (1.00) |
|
| MTOR | 24 (8%) | 269 |
0.123 (1.00) |
0.203 (1.00) |
0.119 (1.00) |
0.298 (1.00) |
0.0602 (1.00) |
1 (1.00) |
1 (1.00) |
|
| STAG3L2 | 6 (2%) | 287 |
0.252 (1.00) |
0.115 (1.00) |
1 (1.00) |
0.415 (1.00) |
1 (1.00) |
0.579 (1.00) |
1 (1.00) |
|
| EBPL | 6 (2%) | 287 |
0.48 (1.00) |
0.527 (1.00) |
0.00161 (0.987) |
0.117 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| KANK3 | 6 (2%) | 287 |
0.733 (1.00) |
0.513 (1.00) |
1 (1.00) |
0.51 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| ZCCHC3 | 3 (1%) | 290 |
0.392 (1.00) |
0.829 (1.00) |
0.554 (1.00) |
0.374 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| FAM174B | 3 (1%) | 290 |
0.476 (1.00) |
0.668 (1.00) |
1 (1.00) |
0.0917 (1.00) |
1 (1.00) |
1 (1.00) |
||
| ANKRD36 | 10 (3%) | 283 |
0.284 (1.00) |
0.339 (1.00) |
1 (1.00) |
0.391 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| TSPAN19 | 4 (1%) | 289 |
0.319 (1.00) |
0.467 (1.00) |
1 (1.00) |
0.458 (1.00) |
1 (1.00) |
0.437 (1.00) |
1 (1.00) |
|
| DACH2 | 7 (2%) | 286 |
0.972 (1.00) |
0.7 (1.00) |
0.242 (1.00) |
1 (1.00) |
0.328 (1.00) |
0.236 (1.00) |
0.0702 (1.00) |
|
| KRTAP1-1 | 3 (1%) | 290 |
0.296 (1.00) |
0.97 (1.00) |
0.0414 (1.00) |
0.0917 (1.00) |
0.35 (1.00) |
1 (1.00) |
||
| UQCRFS1 | 3 (1%) | 290 |
0.321 (1.00) |
0.518 (1.00) |
0.278 (1.00) |
0.0452 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| CR1 | 11 (4%) | 282 |
0.582 (1.00) |
0.936 (1.00) |
0.523 (1.00) |
0.831 (1.00) |
1 (1.00) |
0.645 (1.00) |
1 (1.00) |
|
| VCX2 | 3 (1%) | 290 |
0.293 (1.00) |
0.058 (1.00) |
1 (1.00) |
0.143 (1.00) |
1 (1.00) |
0.35 (1.00) |
1 (1.00) |
|
| KRT1 | 5 (2%) | 288 |
0.00506 (1.00) |
0.0322 (1.00) |
0.346 (1.00) |
0.172 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| ABCB1 | 8 (3%) | 285 |
0.116 (1.00) |
0.582 (1.00) |
0.718 (1.00) |
0.882 (1.00) |
1 (1.00) |
0.603 (1.00) |
1 (1.00) |
|
| MADCAM1 | 4 (1%) | 289 |
0.357 (1.00) |
0.751 (1.00) |
0.123 (1.00) |
0.0204 (1.00) |
1 (1.00) |
0.0871 (1.00) |
1 (1.00) |
|
| POLDIP2 | 4 (1%) | 289 |
0.458 (1.00) |
0.705 (1.00) |
1 (1.00) |
0.458 (1.00) |
1 (1.00) |
0.437 (1.00) |
1 (1.00) |
|
| PCDHGA8 | 4 (1%) | 289 |
0.251 (1.00) |
0.207 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| WDR52 | 9 (3%) | 284 |
0.967 (1.00) |
0.395 (1.00) |
0.724 (1.00) |
0.647 (1.00) |
1 (1.00) |
0.342 (1.00) |
1 (1.00) |
|
| DNAH9 | 18 (6%) | 275 |
0.857 (1.00) |
0.998 (1.00) |
0.445 (1.00) |
0.456 (1.00) |
0.792 (1.00) |
0.166 (1.00) |
0.484 (1.00) |
1 (1.00) |
| TPTE2 | 7 (2%) | 286 |
0.3 (1.00) |
0.309 (1.00) |
1 (1.00) |
0.48 (1.00) |
1 (1.00) |
0.599 (1.00) |
1 (1.00) |
|
| CCNB2 | 5 (2%) | 288 |
0.784 (1.00) |
0.459 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| OR5H1 | 4 (1%) | 289 |
0.919 (1.00) |
0.524 (1.00) |
0.612 (1.00) |
0.606 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| NFE2L2 | 5 (2%) | 288 |
0.163 (1.00) |
0.162 (1.00) |
1 (1.00) |
0.827 (1.00) |
1 (1.00) |
0.513 (1.00) |
1 (1.00) |
|
| GFRAL | 5 (2%) | 288 |
0.655 (1.00) |
0.0399 (1.00) |
1 (1.00) |
0.096 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| MSN | 4 (1%) | 289 |
0.63 (1.00) |
0.578 (1.00) |
1 (1.00) |
0.19 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| RECQL5 | 6 (2%) | 287 |
0.0507 (1.00) |
0.575 (1.00) |
0.188 (1.00) |
0.607 (1.00) |
0.328 (1.00) |
0.183 (1.00) |
1 (1.00) |
|
| NPNT | 6 (2%) | 287 |
0.0806 (1.00) |
0.222 (1.00) |
0.668 (1.00) |
0.342 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| SCAI | 7 (2%) | 286 |
0.461 (1.00) |
0.583 (1.00) |
0.428 (1.00) |
0.246 (1.00) |
1 (1.00) |
0.599 (1.00) |
1 (1.00) |
|
| BAGE2 | 4 (1%) | 289 |
0.961 (1.00) |
0.667 (1.00) |
0.612 (1.00) |
0.796 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| CNTNAP4 | 9 (3%) | 284 |
0.485 (1.00) |
0.5 (1.00) |
0.724 (1.00) |
0.51 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| ZNF800 | 6 (2%) | 287 |
0.166 (1.00) |
0.599 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.579 (1.00) |
1 (1.00) |
|
| SP8 | 3 (1%) | 290 |
0.252 (1.00) |
0.0518 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| UGT2A3 | 4 (1%) | 289 |
0.529 (1.00) |
0.354 (1.00) |
0.612 (1.00) |
1 (1.00) |
1 (1.00) |
0.0871 (1.00) |
1 (1.00) |
|
| LPAR4 | 4 (1%) | 289 |
0.534 (1.00) |
0.457 (1.00) |
0.612 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| SFRS15 | 9 (3%) | 284 |
0.564 (1.00) |
0.804 (1.00) |
0.724 (1.00) |
0.647 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| SPAM1 | 5 (2%) | 288 |
0.27 (1.00) |
0.192 (1.00) |
0.661 (1.00) |
0.096 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| MED12L | 8 (3%) | 285 |
0.018 (1.00) |
0.875 (1.00) |
1 (1.00) |
0.882 (1.00) |
1 (1.00) |
0.289 (1.00) |
1 (1.00) |
|
| PRB4 | 3 (1%) | 290 |
0.71 (1.00) |
0.0414 (1.00) |
0.519 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
||
| WASL | 5 (2%) | 288 |
0.612 (1.00) |
0.225 (1.00) |
0.346 (1.00) |
0.325 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| RNF115 | 4 (1%) | 289 |
0.611 (1.00) |
0.727 (1.00) |
0.612 (1.00) |
0.796 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| NF2 | 6 (2%) | 287 |
0.424 (1.00) |
0.853 (1.00) |
0.423 (1.00) |
0.177 (1.00) |
0.271 (1.00) |
0.579 (1.00) |
1 (1.00) |
|
| SERPINB5 | 3 (1%) | 290 |
0.764 (1.00) |
0.765 (1.00) |
0.554 (1.00) |
0.374 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| TP53 | 9 (3%) | 284 |
0.0454 (1.00) |
0.091 (1.00) |
0.285 (1.00) |
0.032 (1.00) |
0.328 (1.00) |
1 (1.00) |
1 (1.00) |
|
| PRB3 | 3 (1%) | 290 |
0.123 (1.00) |
0.554 (1.00) |
1 (1.00) |
0.0753 (1.00) |
1 (1.00) |
1 (1.00) |
||
| COL11A1 | 8 (3%) | 285 |
0.946 (1.00) |
0.546 (1.00) |
0.269 (1.00) |
0.783 (1.00) |
0.271 (1.00) |
0.603 (1.00) |
1 (1.00) |
|
| KCNH7 | 8 (3%) | 285 |
0.787 (1.00) |
0.95 (1.00) |
1 (1.00) |
0.529 (1.00) |
0.211 (1.00) |
0.603 (1.00) |
1 (1.00) |
|
| LMO4 | 3 (1%) | 290 |
0.902 (1.00) |
0.0752 (1.00) |
1 (1.00) |
0.519 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| PCLO | 20 (7%) | 273 |
0.537 (1.00) |
0.775 (1.00) |
0.223 (1.00) |
0.763 (1.00) |
1 (1.00) |
0.737 (1.00) |
1 (1.00) |
|
| PRIM2 | 5 (2%) | 288 |
0.105 (1.00) |
0.796 (1.00) |
0.346 (1.00) |
0.096 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| PTPN18 | 4 (1%) | 289 |
0.416 (1.00) |
0.349 (1.00) |
1 (1.00) |
0.258 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| RPUSD2 | 3 (1%) | 290 |
0.789 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
||
| SLC2A14 | 3 (1%) | 290 |
0.713 (1.00) |
0.897 (1.00) |
1 (1.00) |
0.519 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| ZKSCAN1 | 4 (1%) | 289 |
0.558 (1.00) |
0.675 (1.00) |
0.612 (1.00) |
0.19 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| ZYG11B | 6 (2%) | 287 |
0.317 (1.00) |
0.00367 (1.00) |
1 (1.00) |
0.863 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| NBPF10 | 20 (7%) | 273 |
0.237 (1.00) |
0.321 (1.00) |
0.151 (1.00) |
0.202 (1.00) |
0.139 (1.00) |
0.0353 (1.00) |
1 (1.00) |
|
| ADCY8 | 5 (2%) | 288 |
0.706 (1.00) |
0.582 (1.00) |
0.167 (1.00) |
0.262 (1.00) |
0.0753 (1.00) |
0.513 (1.00) |
1 (1.00) |
|
| C5ORF13 | 3 (1%) | 290 |
0.32 (1.00) |
0.465 (1.00) |
0.278 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| OR10G7 | 5 (2%) | 288 |
0.0498 (1.00) |
0.301 (1.00) |
1 (1.00) |
0.568 (1.00) |
1 (1.00) |
0.513 (1.00) |
1 (1.00) |
|
| ADAMTS20 | 7 (2%) | 286 |
0.163 (1.00) |
0.243 (1.00) |
0.698 (1.00) |
0.878 (1.00) |
1 (1.00) |
0.0523 (1.00) |
1 (1.00) |
|
| DPP10 | 6 (2%) | 287 |
0.369 (1.00) |
0.902 (1.00) |
1 (1.00) |
0.225 (1.00) |
1 (1.00) |
0.183 (1.00) |
1 (1.00) |
|
| KRTAP4-1 | 3 (1%) | 290 |
0.927 (1.00) |
0.173 (1.00) |
0.0414 (1.00) |
0.718 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| MYO6 | 3 (1%) | 290 |
0.196 (1.00) |
0.554 (1.00) |
0.374 (1.00) |
1 (1.00) |
1 (1.00) |
|||
| POTEC | 10 (3%) | 283 |
0.386 (1.00) |
0.0352 (1.00) |
1 (1.00) |
0.81 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| GRIN2B | 10 (3%) | 283 |
0.873 (1.00) |
0.48 (1.00) |
0.743 (1.00) |
0.902 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| MAGEC1 | 7 (2%) | 286 |
0.586 (1.00) |
0.194 (1.00) |
0.242 (1.00) |
0.878 (1.00) |
0.328 (1.00) |
0.599 (1.00) |
1 (1.00) |
|
| LILRB3 | 3 (1%) | 290 |
0.803 (1.00) |
0.92 (1.00) |
0.278 (1.00) |
0.244 (1.00) |
0.35 (1.00) |
1 (1.00) |
||
| C15ORF26 | 4 (1%) | 289 |
0.221 (1.00) |
0.0652 (1.00) |
0.302 (1.00) |
0.458 (1.00) |
0.0871 (1.00) |
1 (1.00) |
||
| MARCKS | 3 (1%) | 290 |
0.298 (1.00) |
0.113 (1.00) |
0.278 (1.00) |
0.0917 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| MSR1 | 5 (2%) | 288 |
0.156 (1.00) |
0.353 (1.00) |
1 (1.00) |
0.325 (1.00) |
0.0286 (1.00) |
0.513 (1.00) |
1 (1.00) |
|
| RALGAPA1 | 7 (2%) | 286 |
0.909 (1.00) |
0.71 (1.00) |
0.428 (1.00) |
0.0809 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| CLYBL | 4 (1%) | 289 |
0.375 (1.00) |
0.646 (1.00) |
0.302 (1.00) |
0.124 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| SDR16C5 | 4 (1%) | 289 |
0.825 (1.00) |
0.323 (1.00) |
0.302 (1.00) |
1 (1.00) |
1 (1.00) |
0.0871 (1.00) |
1 (1.00) |
|
| SPRYD5 | 4 (1%) | 289 |
0.516 (1.00) |
0.454 (1.00) |
1 (1.00) |
0.606 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| RARA | 4 (1%) | 289 |
0.462 (1.00) |
0.554 (1.00) |
1 (1.00) |
0.606 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| STAG2 | 10 (3%) | 283 |
0.951 (1.00) |
0.00867 (1.00) |
1 (1.00) |
0.0727 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
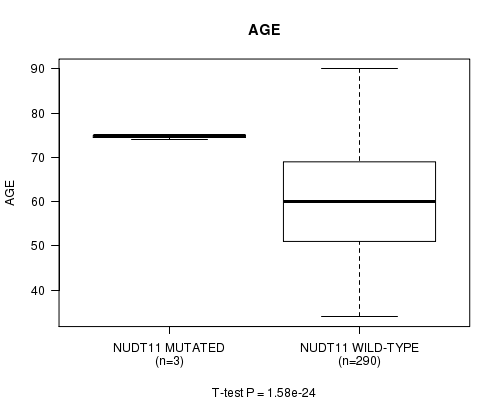
P value = 1.58e-24 (t-test), Q value = 9.7e-22
Table S1. Gene #15: 'NUDT11 MUTATION STATUS' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 293 | 60.3 (11.9) |
| NUDT11 MUTATED | 3 | 74.7 (0.6) |
| NUDT11 WILD-TYPE | 290 | 60.2 (11.9) |
Figure S1. Get High-res Image Gene #15: 'NUDT11 MUTATION STATUS' versus Clinical Feature #2: 'AGE'
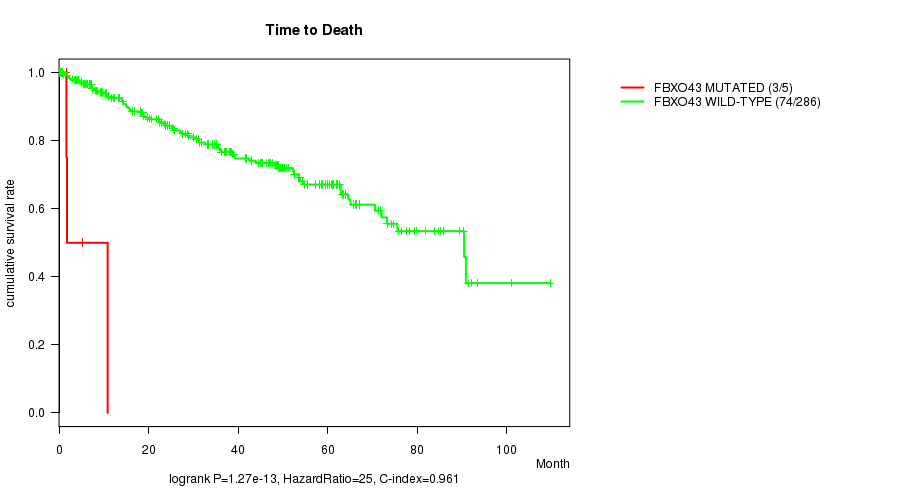
P value = 1.27e-13 (logrank test), Q value = 7.8e-11
Table S2. Gene #69: 'FBXO43 MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 291 | 77 | 0.1 - 109.6 (34.3) |
| FBXO43 MUTATED | 5 | 3 | 1.6 - 10.8 (1.8) |
| FBXO43 WILD-TYPE | 286 | 74 | 0.1 - 109.6 (34.9) |
Figure S2. Get High-res Image Gene #69: 'FBXO43 MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
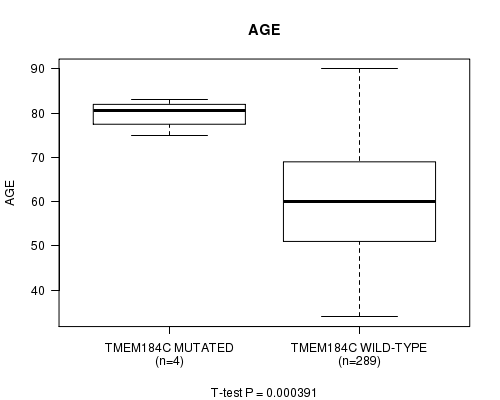
P value = 0.000391 (t-test), Q value = 0.24
Table S3. Gene #81: 'TMEM184C MUTATION STATUS' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 293 | 60.3 (11.9) |
| TMEM184C MUTATED | 4 | 79.8 (3.4) |
| TMEM184C WILD-TYPE | 289 | 60.1 (11.8) |
Figure S3. Get High-res Image Gene #81: 'TMEM184C MUTATION STATUS' versus Clinical Feature #2: 'AGE'
-
Mutation data file = KIRC.mutsig.cluster.txt
-
Clinical data file = KIRC.clin.merged.picked.txt
-
Number of patients = 293
-
Number of significantly mutated genes = 89
-
Number of selected clinical features = 8
-
Exclude genes that fewer than K tumors have mutations, K = 3
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For continuous numerical clinical features, two-tailed Student's t test with unequal variance (Lehmann and Romano 2005) was applied to compare the clinical values between tumors with and without gene mutations using 't.test' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.