(primary solid tumor cohort)
This pipeline computes the correlation between cancer subtypes identified by different molecular patterns and selected clinical features.
Testing the association between subtypes identified by 8 different clustering approaches and 6 clinical features across 179 patients, 12 significant findings detected with P value < 0.05 and Q value < 0.25.
-
CNMF clustering analysis on array-based mRNA expression data identified 3 subtypes that do not correlate to any clinical features.
-
Consensus hierarchical clustering analysis on array-based mRNA expression data identified 3 subtypes that do not correlate to any clinical features.
-
3 subtypes identified in current cancer cohort by 'CN CNMF'. These subtypes correlate to 'HISTOLOGICAL.TYPE'.
-
4 subtypes identified in current cancer cohort by 'METHLYATION CNMF'. These subtypes correlate to 'Time to Death', 'AGE', 'HISTOLOGICAL.TYPE', and 'RADIATIONS.RADIATION.REGIMENINDICATION'.
-
CNMF clustering analysis on sequencing-based mRNA expression data identified 4 subtypes that correlate to 'Time to Death', 'HISTOLOGICAL.TYPE', and 'RADIATIONS.RADIATION.REGIMENINDICATION'.
-
Consensus hierarchical clustering analysis on sequencing-based mRNA expression data identified 4 subtypes that correlate to 'Time to Death', 'AGE', and 'HISTOLOGICAL.TYPE'.
-
CNMF clustering analysis on sequencing-based miR expression data identified 7 subtypes that correlate to 'HISTOLOGICAL.TYPE'.
-
Consensus hierarchical clustering analysis on sequencing-based miR expression data identified 3 subtypes that do not correlate to any clinical features.
Table 1. Get Full Table Overview of the association between subtypes identified by 8 different clustering approaches and 6 clinical features. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 12 significant findings detected.
|
Clinical Features |
Time to Death |
AGE | GENDER |
KARNOFSKY PERFORMANCE SCORE |
HISTOLOGICAL TYPE |
RADIATIONS RADIATION REGIMENINDICATION |
| Statistical Tests | logrank test | ANOVA | Fisher's exact test | ANOVA | Chi-square test | Fisher's exact test |
| mRNA CNMF subtypes |
0.247 (1.00) |
0.326 (1.00) |
0.101 (1.00) |
0.441 (1.00) |
0.0226 (0.699) |
0.384 (1.00) |
| mRNA cHierClus subtypes |
0.31 (1.00) |
0.467 (1.00) |
0.172 (1.00) |
0.441 (1.00) |
0.0136 (0.463) |
0.757 (1.00) |
| CN CNMF |
0.0131 (0.458) |
0.162 (1.00) |
0.0354 (1.00) |
0.0523 (1.00) |
0.000129 (0.00555) |
0.0365 (1.00) |
| METHLYATION CNMF |
1.19e-05 (0.00055) |
2.76e-05 (0.00121) |
0.0183 (0.584) |
0.872 (1.00) |
1.55e-11 (7.43e-10) |
0.000172 (0.00723) |
| RNAseq CNMF subtypes |
0.000486 (0.0194) |
0.0677 (1.00) |
0.294 (1.00) |
0.204 (1.00) |
4.12e-06 (0.000193) |
0.00271 (0.103) |
| RNAseq cHierClus subtypes |
0.000316 (0.013) |
0.0059 (0.218) |
0.127 (1.00) |
0.361 (1.00) |
1.22e-05 (0.00055) |
0.0151 (0.497) |
| MIRseq CNMF subtypes |
0.324 (1.00) |
0.00901 (0.324) |
0.0591 (1.00) |
0.294 (1.00) |
0.000491 (0.0194) |
0.13 (1.00) |
| MIRseq cHierClus subtypes |
0.0326 (0.978) |
0.804 (1.00) |
0.693 (1.00) |
0.852 (1.00) |
0.0496 (1.00) |
0.0781 (1.00) |
Table S1. Get Full Table Description of clustering approach #1: 'mRNA CNMF subtypes'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 9 | 10 | 8 |
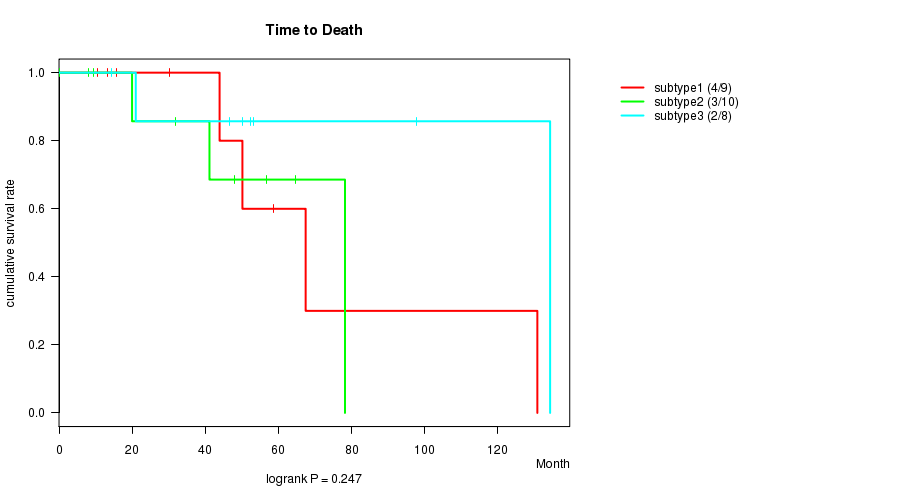
P value = 0.247 (logrank test), Q value = 1
Table S2. Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 27 | 9 | 0.1 - 134.3 (46.6) |
| subtype1 | 9 | 4 | 10.6 - 130.8 (43.9) |
| subtype2 | 10 | 3 | 0.1 - 78.2 (36.5) |
| subtype3 | 8 | 2 | 14.4 - 134.3 (51.3) |
Figure S1. Get High-res Image Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
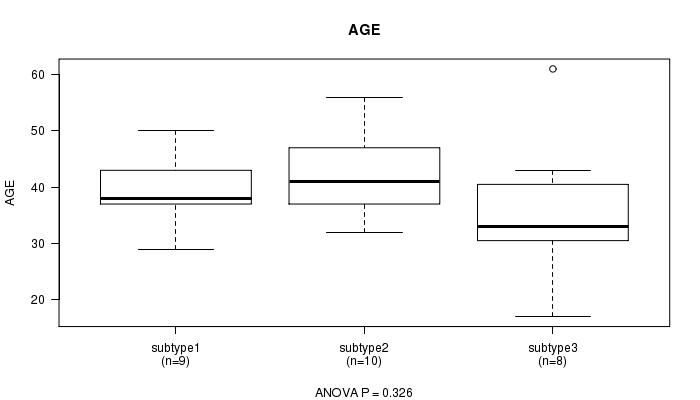
P value = 0.326 (ANOVA), Q value = 1
Table S3. Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 27 | 39.3 (9.1) |
| subtype1 | 9 | 39.2 (6.2) |
| subtype2 | 10 | 42.3 (7.6) |
| subtype3 | 8 | 35.8 (12.6) |
Figure S2. Get High-res Image Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #2: 'AGE'
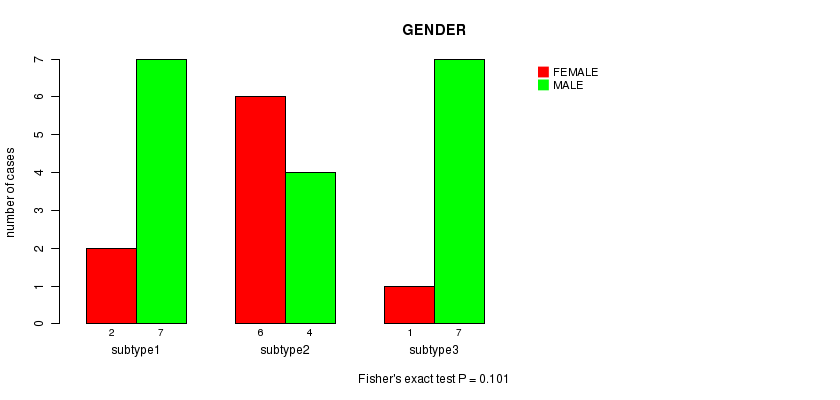
P value = 0.101 (Fisher's exact test), Q value = 1
Table S4. Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #3: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 9 | 18 |
| subtype1 | 2 | 7 |
| subtype2 | 6 | 4 |
| subtype3 | 1 | 7 |
Figure S3. Get High-res Image Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #3: 'GENDER'
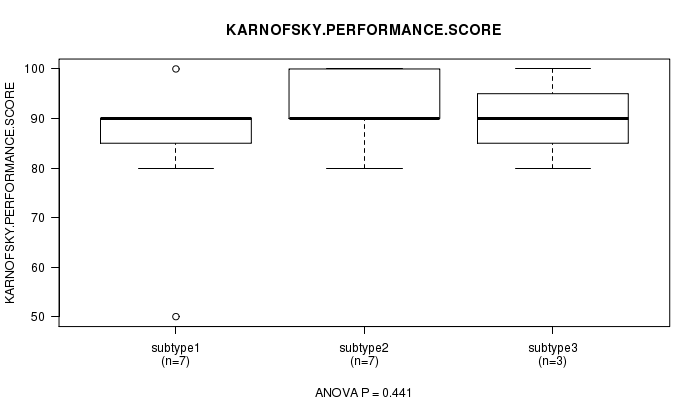
P value = 0.441 (ANOVA), Q value = 1
Table S5. Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 17 | 88.8 (12.2) |
| subtype1 | 7 | 84.3 (16.2) |
| subtype2 | 7 | 92.9 (7.6) |
| subtype3 | 3 | 90.0 (10.0) |
Figure S4. Get High-res Image Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
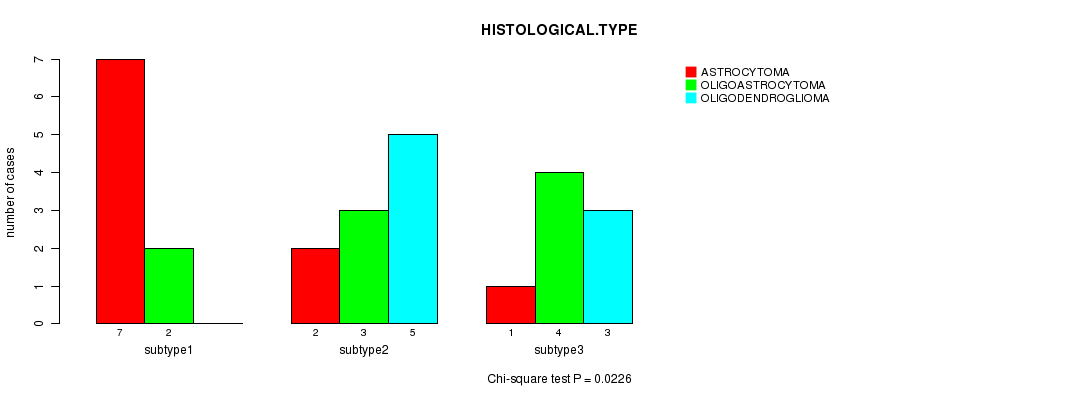
P value = 0.0226 (Chi-square test), Q value = 0.7
Table S6. Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 10 | 9 | 8 |
| subtype1 | 7 | 2 | 0 |
| subtype2 | 2 | 3 | 5 |
| subtype3 | 1 | 4 | 3 |
Figure S5. Get High-res Image Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
P value = 0.384 (Fisher's exact test), Q value = 1
Table S7. Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 19 | 8 |
| subtype1 | 7 | 2 |
| subtype2 | 8 | 2 |
| subtype3 | 4 | 4 |
Figure S6. Get High-res Image Clustering Approach #1: 'mRNA CNMF subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
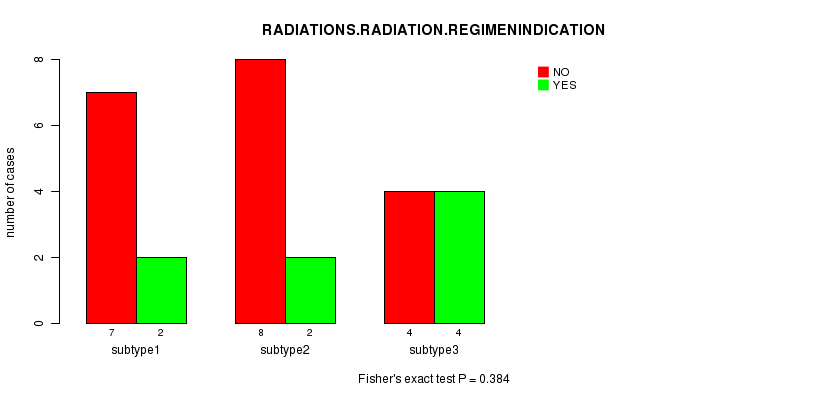
Table S8. Get Full Table Description of clustering approach #2: 'mRNA cHierClus subtypes'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 9 | 7 | 11 |
P value = 0.31 (logrank test), Q value = 1
Table S9. Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 27 | 9 | 0.1 - 134.3 (46.6) |
| subtype1 | 9 | 4 | 10.6 - 130.8 (43.9) |
| subtype2 | 7 | 2 | 14.4 - 134.3 (52.4) |
| subtype3 | 11 | 3 | 0.1 - 78.2 (41.1) |
Figure S7. Get High-res Image Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
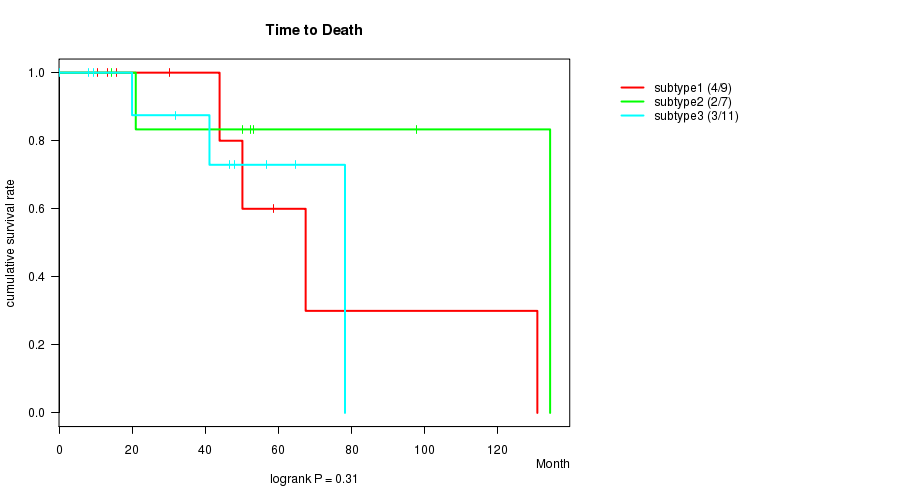
P value = 0.467 (ANOVA), Q value = 1
Table S10. Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 27 | 39.3 (9.1) |
| subtype1 | 9 | 39.2 (6.2) |
| subtype2 | 7 | 36.0 (13.6) |
| subtype3 | 11 | 41.5 (7.6) |
Figure S8. Get High-res Image Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #2: 'AGE'
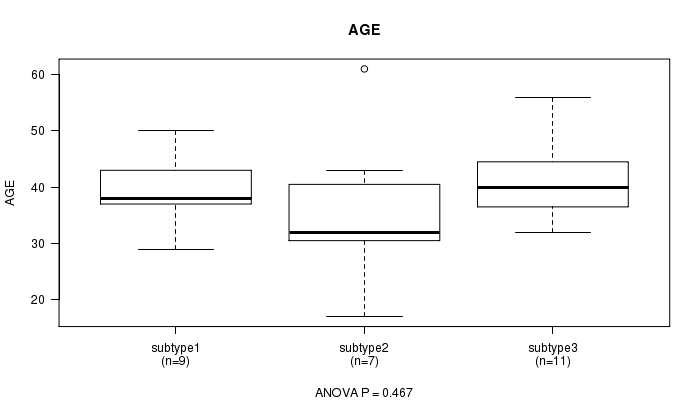
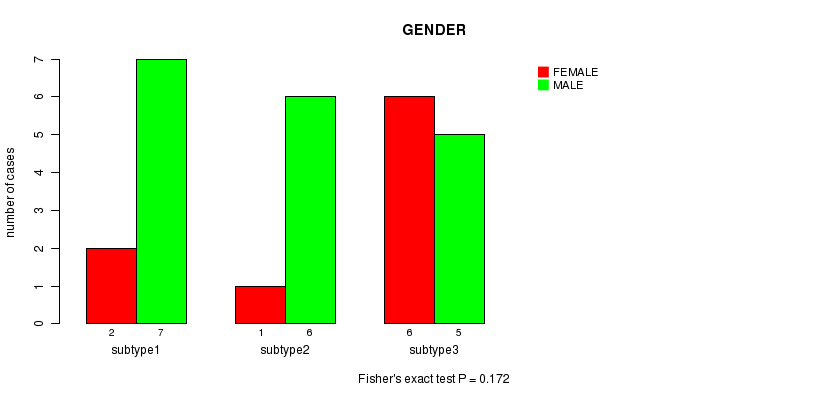
P value = 0.172 (Fisher's exact test), Q value = 1
Table S11. Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #3: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 9 | 18 |
| subtype1 | 2 | 7 |
| subtype2 | 1 | 6 |
| subtype3 | 6 | 5 |
Figure S9. Get High-res Image Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #3: 'GENDER'
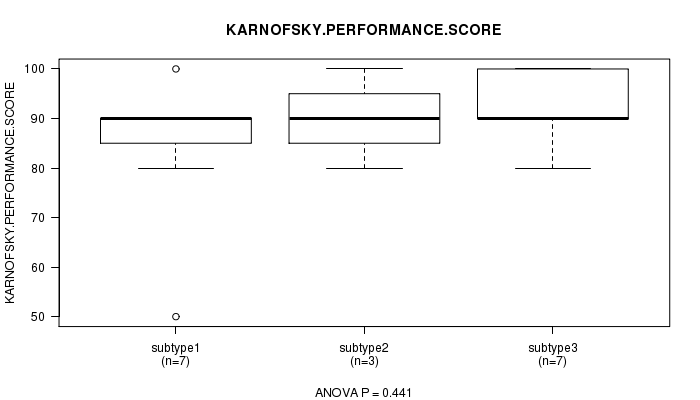
P value = 0.441 (ANOVA), Q value = 1
Table S12. Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 17 | 88.8 (12.2) |
| subtype1 | 7 | 84.3 (16.2) |
| subtype2 | 3 | 90.0 (10.0) |
| subtype3 | 7 | 92.9 (7.6) |
Figure S10. Get High-res Image Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
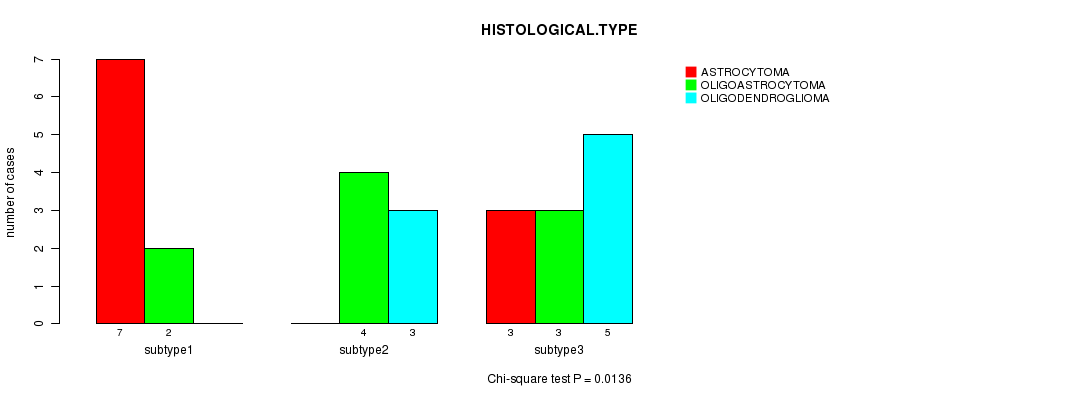
P value = 0.0136 (Chi-square test), Q value = 0.46
Table S13. Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 10 | 9 | 8 |
| subtype1 | 7 | 2 | 0 |
| subtype2 | 0 | 4 | 3 |
| subtype3 | 3 | 3 | 5 |
Figure S11. Get High-res Image Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
P value = 0.757 (Fisher's exact test), Q value = 1
Table S14. Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 19 | 8 |
| subtype1 | 7 | 2 |
| subtype2 | 4 | 3 |
| subtype3 | 8 | 3 |
Figure S12. Get High-res Image Clustering Approach #2: 'mRNA cHierClus subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
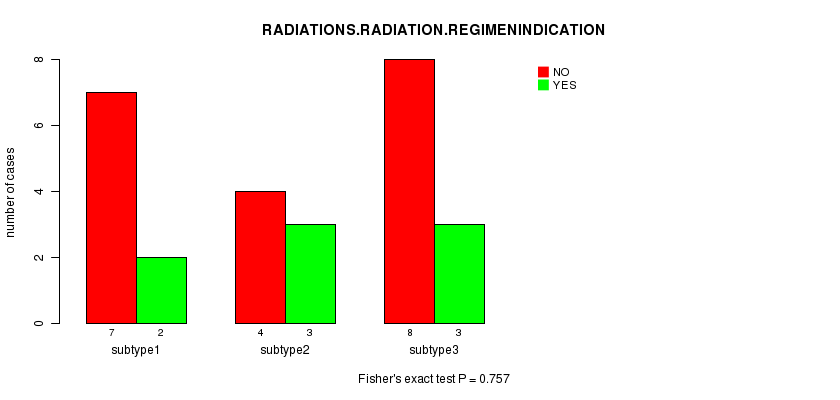
Table S15. Get Full Table Description of clustering approach #3: 'CN CNMF'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 59 | 50 | 69 |
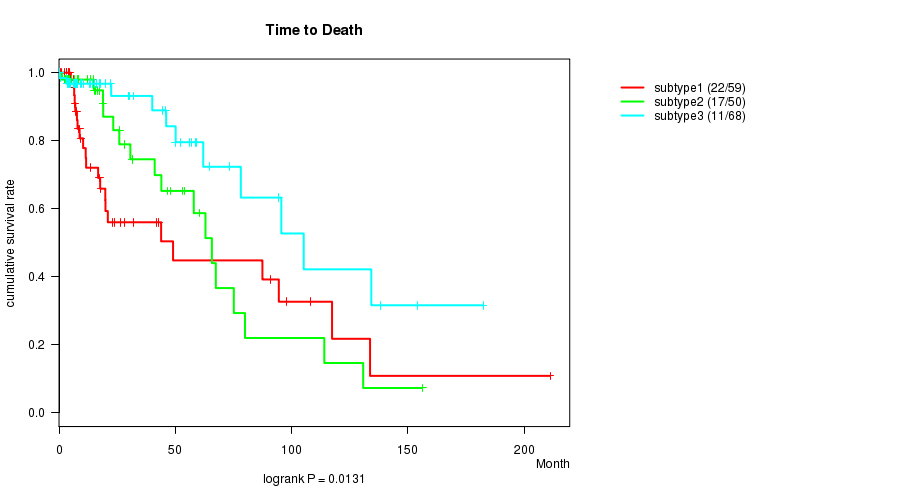
P value = 0.0131 (logrank test), Q value = 0.46
Table S16. Clustering Approach #3: 'CN CNMF' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 177 | 50 | 0.0 - 211.2 (14.7) |
| subtype1 | 59 | 22 | 0.1 - 211.2 (8.8) |
| subtype2 | 50 | 17 | 0.1 - 156.2 (18.2) |
| subtype3 | 68 | 11 | 0.0 - 182.3 (14.5) |
Figure S13. Get High-res Image Clustering Approach #3: 'CN CNMF' versus Clinical Feature #1: 'Time to Death'
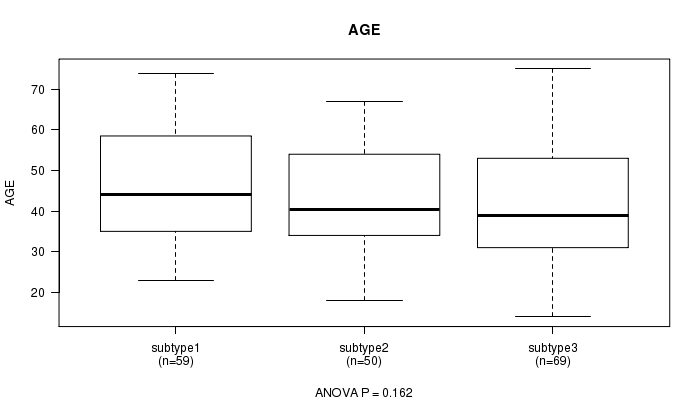
P value = 0.162 (ANOVA), Q value = 1
Table S17. Clustering Approach #3: 'CN CNMF' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 178 | 43.1 (13.4) |
| subtype1 | 59 | 45.8 (13.4) |
| subtype2 | 50 | 42.3 (12.5) |
| subtype3 | 69 | 41.5 (13.8) |
Figure S14. Get High-res Image Clustering Approach #3: 'CN CNMF' versus Clinical Feature #2: 'AGE'
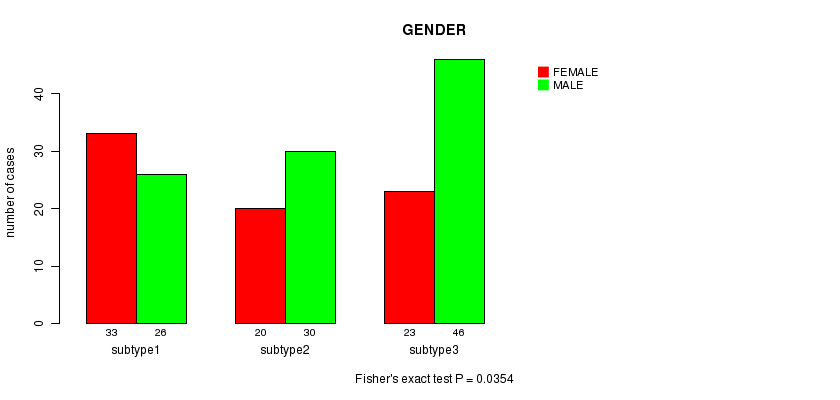
P value = 0.0354 (Fisher's exact test), Q value = 1
Table S18. Clustering Approach #3: 'CN CNMF' versus Clinical Feature #3: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 76 | 102 |
| subtype1 | 33 | 26 |
| subtype2 | 20 | 30 |
| subtype3 | 23 | 46 |
Figure S15. Get High-res Image Clustering Approach #3: 'CN CNMF' versus Clinical Feature #3: 'GENDER'
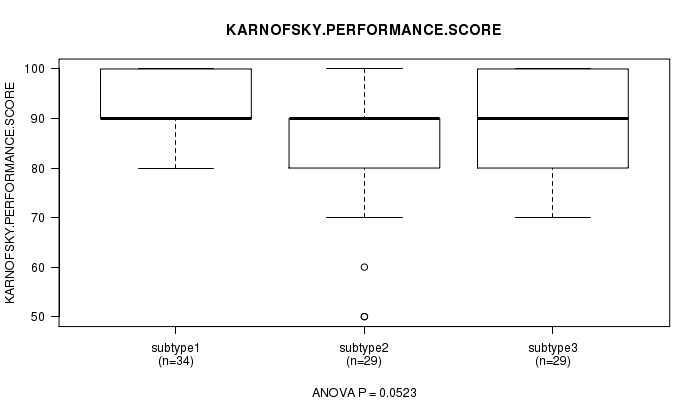
P value = 0.0523 (ANOVA), Q value = 1
Table S19. Clustering Approach #3: 'CN CNMF' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 92 | 88.5 (10.5) |
| subtype1 | 34 | 91.2 (6.9) |
| subtype2 | 29 | 84.8 (13.3) |
| subtype3 | 29 | 89.0 (10.1) |
Figure S16. Get High-res Image Clustering Approach #3: 'CN CNMF' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
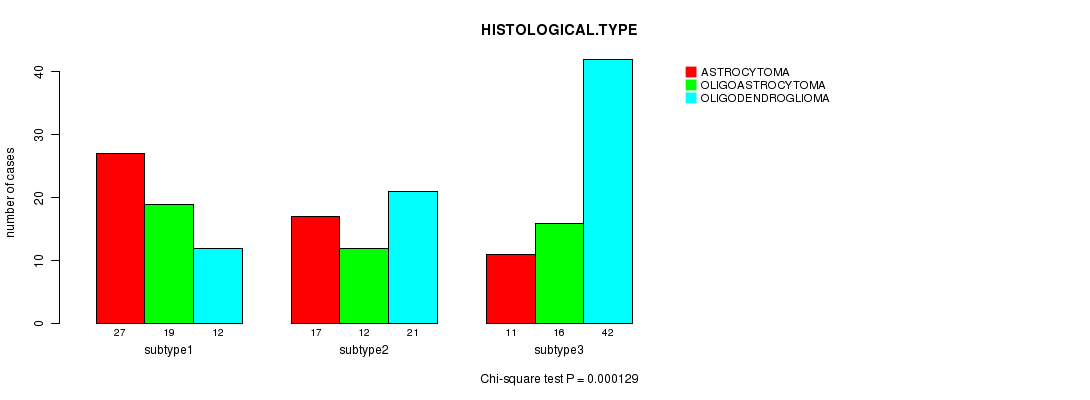
P value = 0.000129 (Chi-square test), Q value = 0.0055
Table S20. Clustering Approach #3: 'CN CNMF' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 55 | 47 | 75 |
| subtype1 | 27 | 19 | 12 |
| subtype2 | 17 | 12 | 21 |
| subtype3 | 11 | 16 | 42 |
Figure S17. Get High-res Image Clustering Approach #3: 'CN CNMF' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
P value = 0.0365 (Fisher's exact test), Q value = 1
Table S21. Clustering Approach #3: 'CN CNMF' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 96 | 82 |
| subtype1 | 35 | 24 |
| subtype2 | 32 | 18 |
| subtype3 | 29 | 40 |
Figure S18. Get High-res Image Clustering Approach #3: 'CN CNMF' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
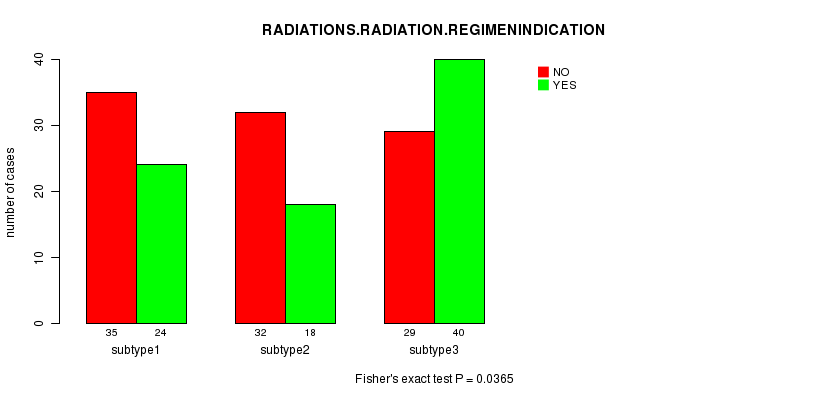
Table S22. Get Full Table Description of clustering approach #4: 'METHLYATION CNMF'
| Cluster Labels | 1 | 2 | 3 | 4 |
|---|---|---|---|---|
| Number of samples | 76 | 29 | 19 | 50 |
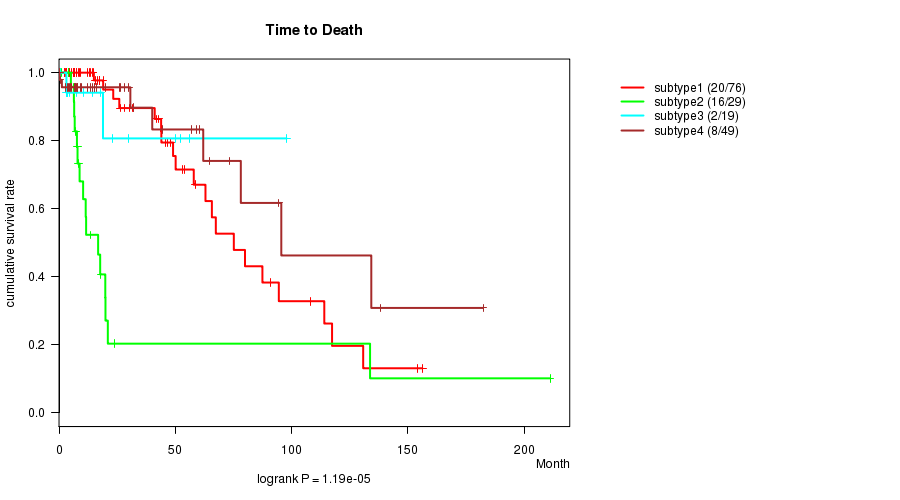
P value = 1.19e-05 (logrank test), Q value = 0.00055
Table S23. Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 173 | 46 | 0.0 - 211.2 (14.5) |
| subtype1 | 76 | 20 | 0.0 - 156.2 (17.5) |
| subtype2 | 29 | 16 | 0.1 - 211.2 (8.4) |
| subtype3 | 19 | 2 | 0.1 - 97.9 (10.6) |
| subtype4 | 49 | 8 | 0.1 - 182.3 (14.5) |
Figure S19. Get High-res Image Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #1: 'Time to Death'
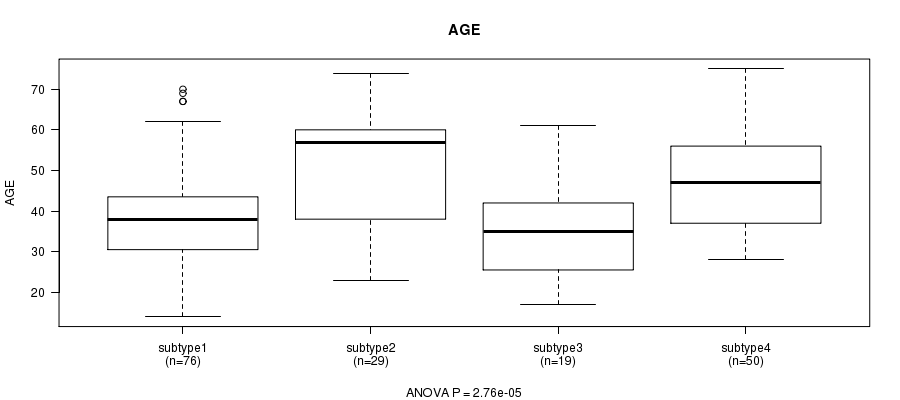
P value = 2.76e-05 (ANOVA), Q value = 0.0012
Table S24. Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 174 | 43.1 (13.4) |
| subtype1 | 76 | 39.8 (12.2) |
| subtype2 | 29 | 50.4 (14.1) |
| subtype3 | 19 | 35.6 (12.3) |
| subtype4 | 50 | 46.6 (12.3) |
Figure S20. Get High-res Image Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #2: 'AGE'
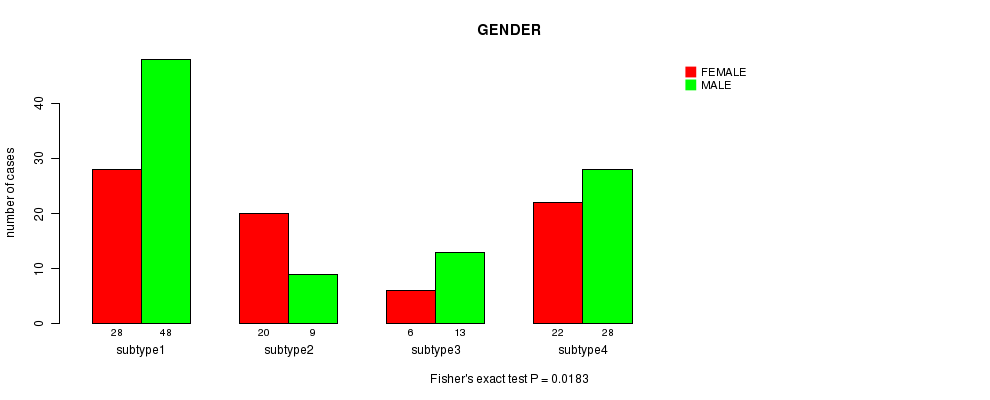
P value = 0.0183 (Fisher's exact test), Q value = 0.58
Table S25. Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #3: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 76 | 98 |
| subtype1 | 28 | 48 |
| subtype2 | 20 | 9 |
| subtype3 | 6 | 13 |
| subtype4 | 22 | 28 |
Figure S21. Get High-res Image Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #3: 'GENDER'
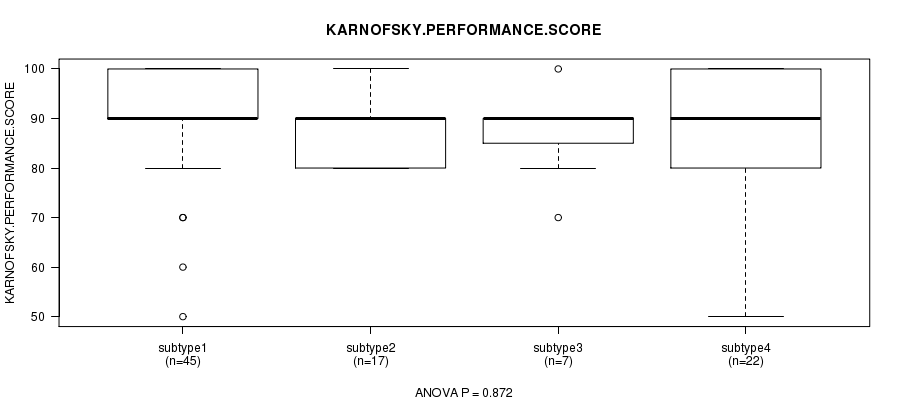
P value = 0.872 (ANOVA), Q value = 1
Table S26. Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 91 | 88.6 (10.5) |
| subtype1 | 45 | 89.3 (10.5) |
| subtype2 | 17 | 87.1 (5.9) |
| subtype3 | 7 | 87.1 (9.5) |
| subtype4 | 22 | 88.6 (13.6) |
Figure S22. Get High-res Image Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
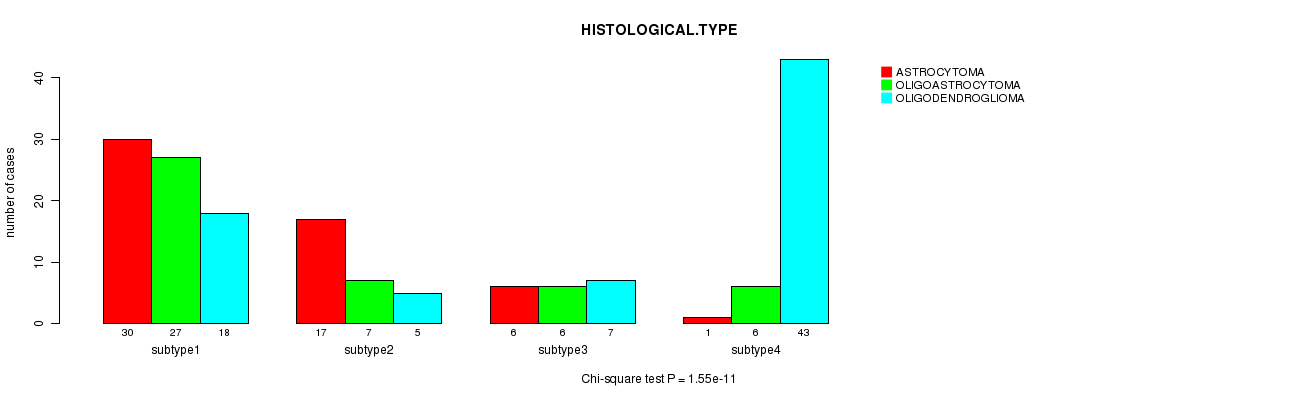
P value = 1.55e-11 (Chi-square test), Q value = 7.4e-10
Table S27. Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 54 | 46 | 73 |
| subtype1 | 30 | 27 | 18 |
| subtype2 | 17 | 7 | 5 |
| subtype3 | 6 | 6 | 7 |
| subtype4 | 1 | 6 | 43 |
Figure S23. Get High-res Image Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
P value = 0.000172 (Fisher's exact test), Q value = 0.0072
Table S28. Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 93 | 81 |
| subtype1 | 53 | 23 |
| subtype2 | 17 | 12 |
| subtype3 | 6 | 13 |
| subtype4 | 17 | 33 |
Figure S24. Get High-res Image Clustering Approach #4: 'METHLYATION CNMF' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
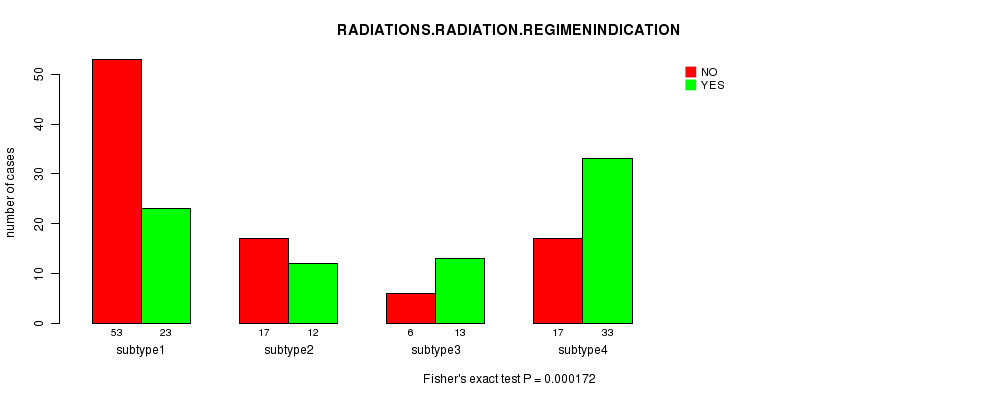
Table S29. Get Full Table Description of clustering approach #5: 'RNAseq CNMF subtypes'
| Cluster Labels | 1 | 2 | 3 | 4 |
|---|---|---|---|---|
| Number of samples | 23 | 29 | 24 | 34 |
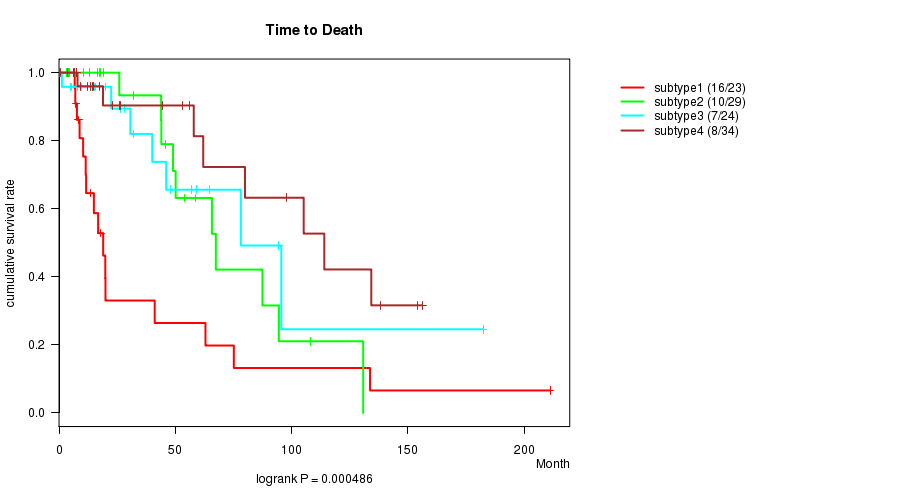
P value = 0.000486 (logrank test), Q value = 0.019
Table S30. Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 110 | 41 | 0.0 - 211.2 (19.0) |
| subtype1 | 23 | 16 | 4.1 - 211.2 (13.4) |
| subtype2 | 29 | 10 | 0.0 - 130.8 (25.9) |
| subtype3 | 24 | 7 | 1.2 - 182.3 (29.4) |
| subtype4 | 34 | 8 | 0.1 - 156.2 (18.2) |
Figure S25. Get High-res Image Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
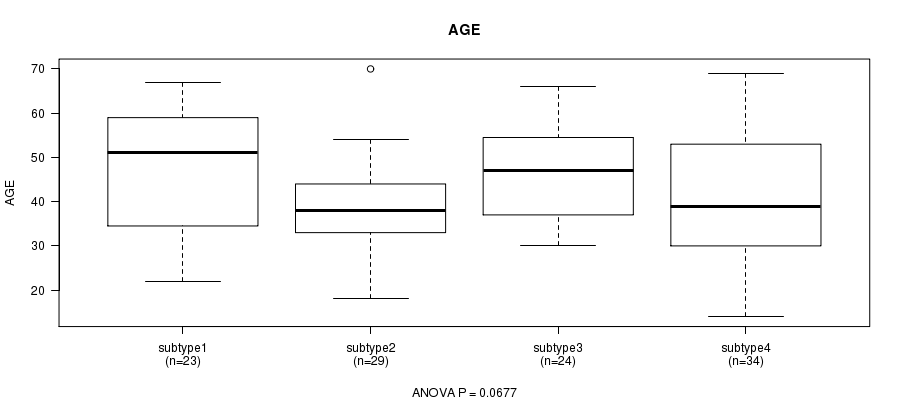
P value = 0.0677 (ANOVA), Q value = 1
Table S31. Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 110 | 42.9 (12.9) |
| subtype1 | 23 | 46.8 (14.3) |
| subtype2 | 29 | 39.5 (10.3) |
| subtype3 | 24 | 46.6 (10.8) |
| subtype4 | 34 | 40.7 (14.3) |
Figure S26. Get High-res Image Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #2: 'AGE'
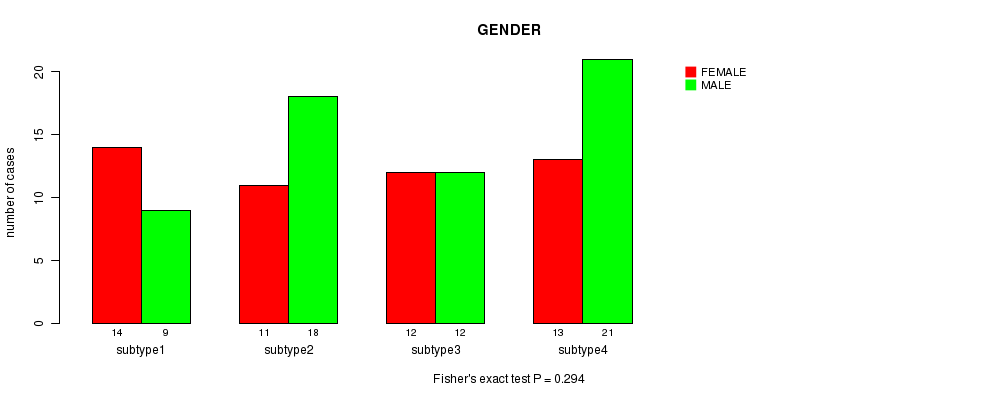
P value = 0.294 (Fisher's exact test), Q value = 1
Table S32. Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #3: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 50 | 60 |
| subtype1 | 14 | 9 |
| subtype2 | 11 | 18 |
| subtype3 | 12 | 12 |
| subtype4 | 13 | 21 |
Figure S27. Get High-res Image Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #3: 'GENDER'
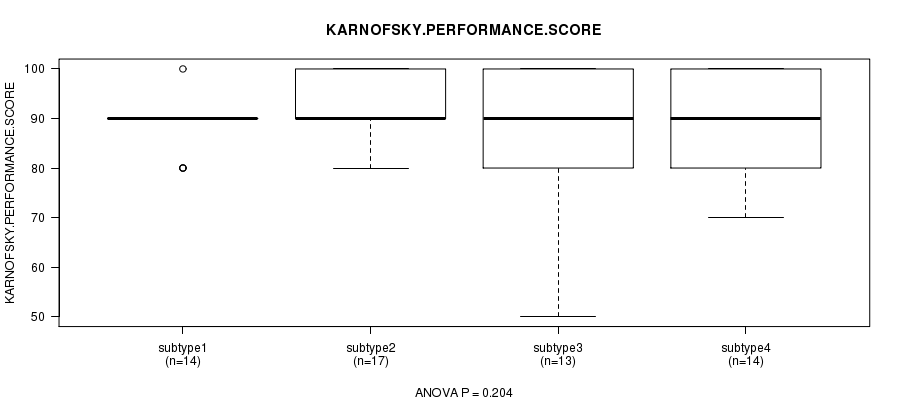
P value = 0.204 (ANOVA), Q value = 1
Table S33. Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 58 | 88.6 (11.0) |
| subtype1 | 14 | 88.6 (5.3) |
| subtype2 | 17 | 92.9 (5.9) |
| subtype3 | 13 | 84.6 (18.1) |
| subtype4 | 14 | 87.1 (10.7) |
Figure S28. Get High-res Image Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
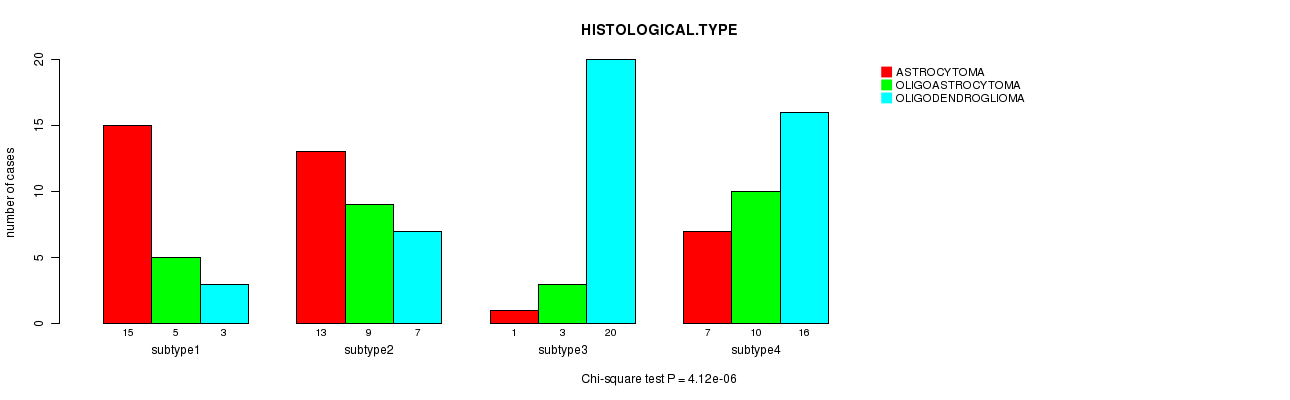
P value = 4.12e-06 (Chi-square test), Q value = 0.00019
Table S34. Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 36 | 27 | 46 |
| subtype1 | 15 | 5 | 3 |
| subtype2 | 13 | 9 | 7 |
| subtype3 | 1 | 3 | 20 |
| subtype4 | 7 | 10 | 16 |
Figure S29. Get High-res Image Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
P value = 0.00271 (Fisher's exact test), Q value = 0.1
Table S35. Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 66 | 44 |
| subtype1 | 17 | 6 |
| subtype2 | 23 | 6 |
| subtype3 | 8 | 16 |
| subtype4 | 18 | 16 |
Figure S30. Get High-res Image Clustering Approach #5: 'RNAseq CNMF subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
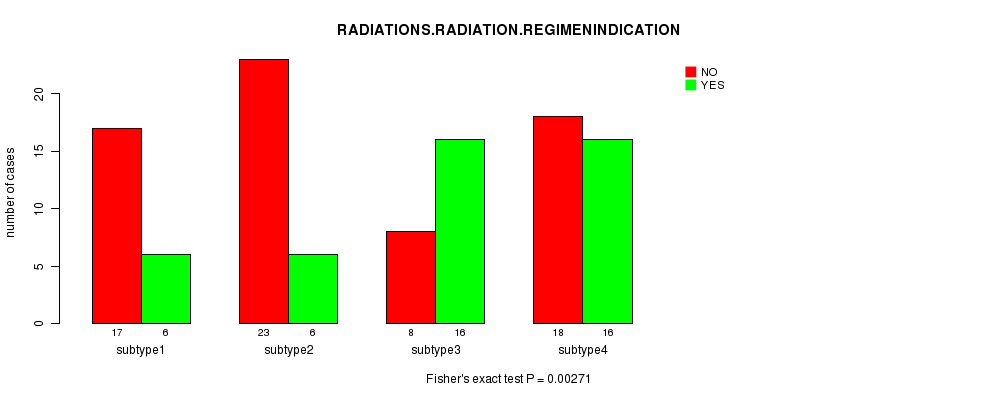
Table S36. Get Full Table Description of clustering approach #6: 'RNAseq cHierClus subtypes'
| Cluster Labels | 1 | 2 | 3 | 4 |
|---|---|---|---|---|
| Number of samples | 35 | 35 | 20 | 20 |
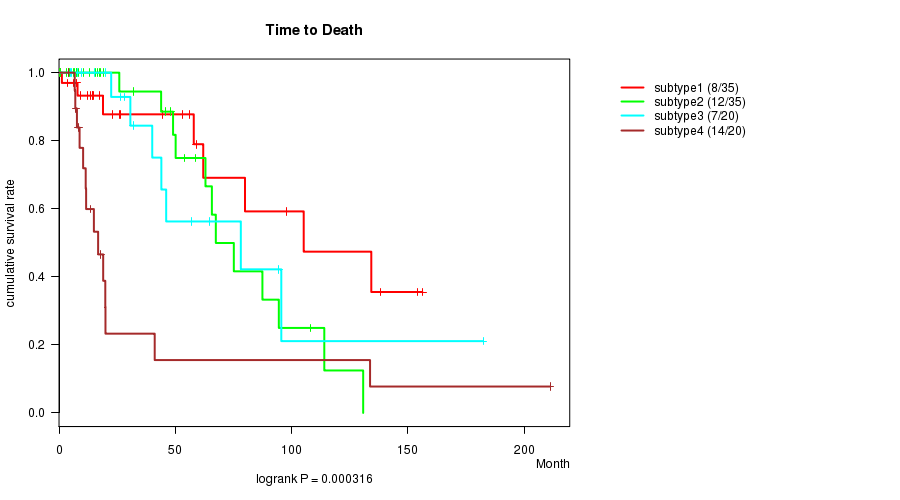
P value = 0.000316 (logrank test), Q value = 0.013
Table S37. Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 110 | 41 | 0.0 - 211.2 (19.0) |
| subtype1 | 35 | 8 | 0.1 - 156.2 (17.4) |
| subtype2 | 35 | 12 | 0.0 - 130.8 (25.9) |
| subtype3 | 20 | 7 | 4.8 - 182.3 (31.2) |
| subtype4 | 20 | 14 | 4.1 - 211.2 (12.5) |
Figure S31. Get High-res Image Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
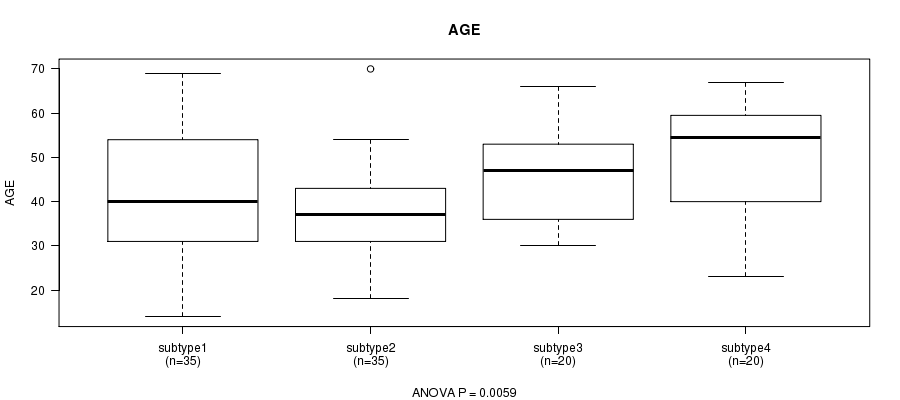
P value = 0.0059 (ANOVA), Q value = 0.22
Table S38. Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 110 | 42.9 (12.9) |
| subtype1 | 35 | 42.8 (14.3) |
| subtype2 | 35 | 37.7 (10.5) |
| subtype3 | 20 | 45.9 (10.6) |
| subtype4 | 20 | 49.5 (13.2) |
Figure S32. Get High-res Image Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #2: 'AGE'
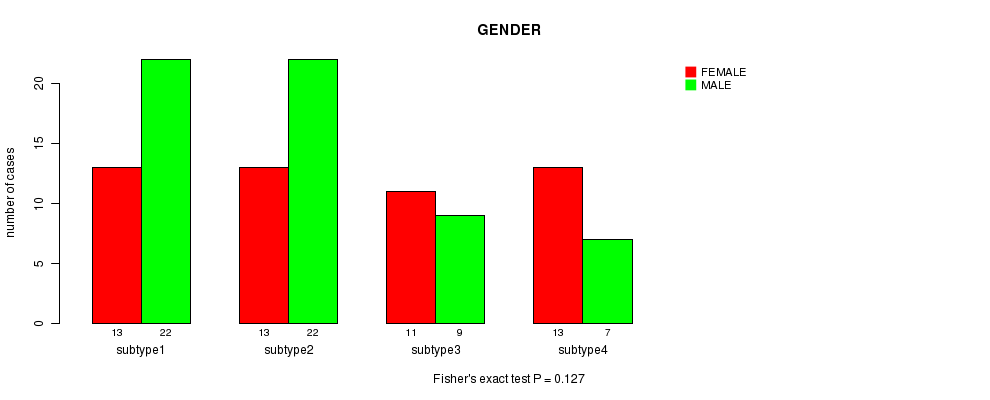
P value = 0.127 (Fisher's exact test), Q value = 1
Table S39. Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #3: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 50 | 60 |
| subtype1 | 13 | 22 |
| subtype2 | 13 | 22 |
| subtype3 | 11 | 9 |
| subtype4 | 13 | 7 |
Figure S33. Get High-res Image Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #3: 'GENDER'
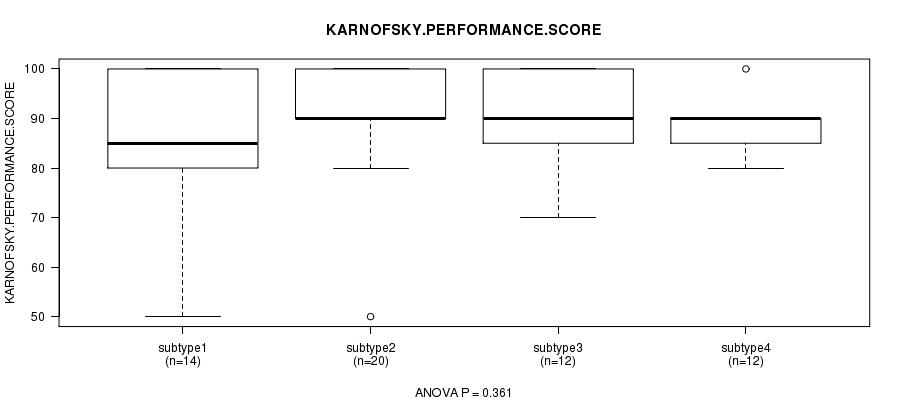
P value = 0.361 (ANOVA), Q value = 1
Table S40. Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 58 | 88.6 (11.0) |
| subtype1 | 14 | 84.3 (14.5) |
| subtype2 | 20 | 90.5 (11.0) |
| subtype3 | 12 | 90.8 (10.0) |
| subtype4 | 12 | 88.3 (5.8) |
Figure S34. Get High-res Image Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
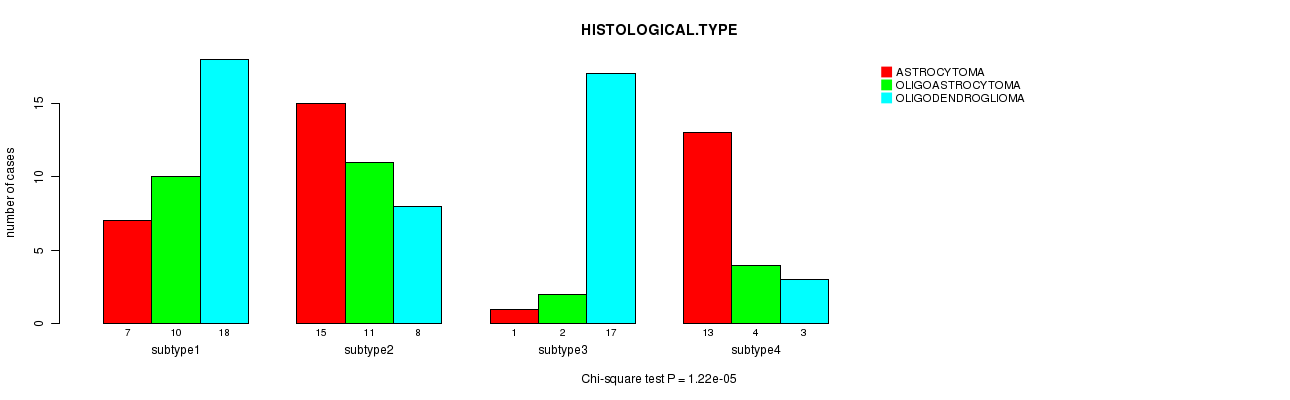
P value = 1.22e-05 (Chi-square test), Q value = 0.00055
Table S41. Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 36 | 27 | 46 |
| subtype1 | 7 | 10 | 18 |
| subtype2 | 15 | 11 | 8 |
| subtype3 | 1 | 2 | 17 |
| subtype4 | 13 | 4 | 3 |
Figure S35. Get High-res Image Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
P value = 0.0151 (Fisher's exact test), Q value = 0.5
Table S42. Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 66 | 44 |
| subtype1 | 17 | 18 |
| subtype2 | 25 | 10 |
| subtype3 | 8 | 12 |
| subtype4 | 16 | 4 |
Figure S36. Get High-res Image Clustering Approach #6: 'RNAseq cHierClus subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
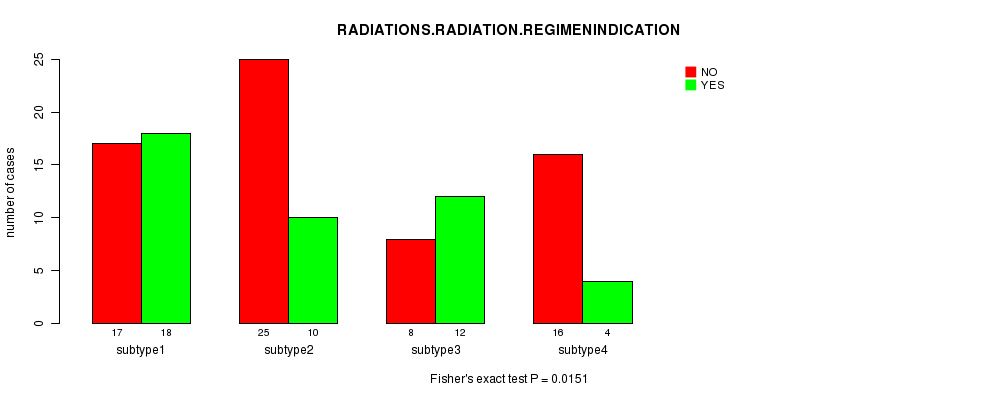
Table S43. Get Full Table Description of clustering approach #7: 'MIRseq CNMF subtypes'
| Cluster Labels | 1 | 2 | 3 | 4 | 5 | 6 | 7 |
|---|---|---|---|---|---|---|---|
| Number of samples | 23 | 29 | 19 | 20 | 47 | 31 | 9 |
P value = 0.324 (logrank test), Q value = 1
Table S44. Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 177 | 49 | 0.0 - 211.2 (14.7) |
| subtype1 | 23 | 4 | 0.0 - 107.9 (6.0) |
| subtype2 | 29 | 12 | 0.1 - 134.3 (28.2) |
| subtype3 | 19 | 8 | 0.8 - 117.4 (16.5) |
| subtype4 | 19 | 4 | 0.1 - 94.3 (17.3) |
| subtype5 | 47 | 8 | 0.1 - 156.2 (8.8) |
| subtype6 | 31 | 12 | 1.2 - 211.2 (17.9) |
| subtype7 | 9 | 1 | 3.1 - 95.6 (15.3) |
Figure S37. Get High-res Image Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
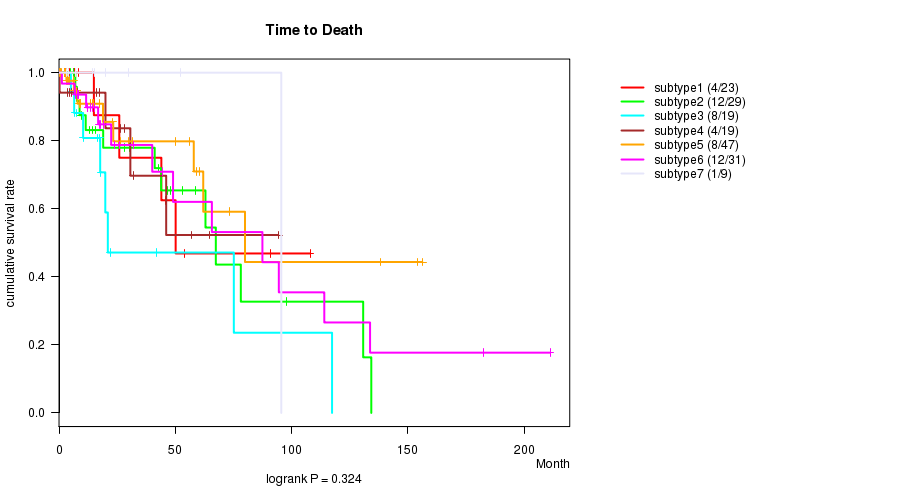
P value = 0.00901 (ANOVA), Q value = 0.32
Table S45. Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 178 | 43.3 (13.3) |
| subtype1 | 23 | 37.2 (10.9) |
| subtype2 | 29 | 42.0 (11.3) |
| subtype3 | 19 | 48.5 (12.9) |
| subtype4 | 20 | 49.4 (13.3) |
| subtype5 | 47 | 41.9 (14.0) |
| subtype6 | 31 | 46.1 (14.1) |
| subtype7 | 9 | 36.1 (11.2) |
Figure S38. Get High-res Image Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #2: 'AGE'
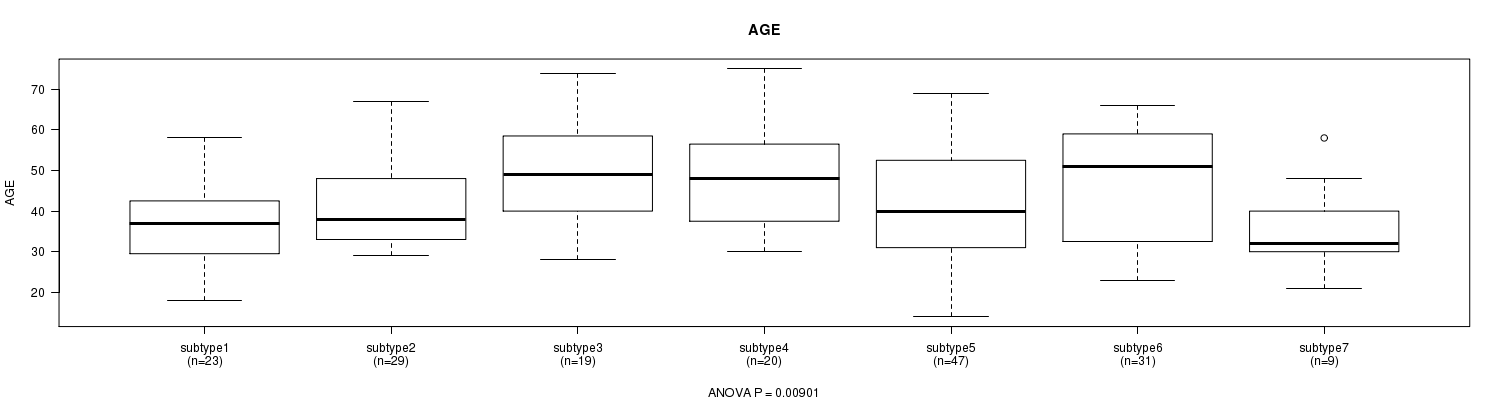
P value = 0.0591 (Chi-square test), Q value = 1
Table S46. Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #3: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 76 | 102 |
| subtype1 | 5 | 18 |
| subtype2 | 8 | 21 |
| subtype3 | 11 | 8 |
| subtype4 | 11 | 9 |
| subtype5 | 21 | 26 |
| subtype6 | 17 | 14 |
| subtype7 | 3 | 6 |
Figure S39. Get High-res Image Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #3: 'GENDER'
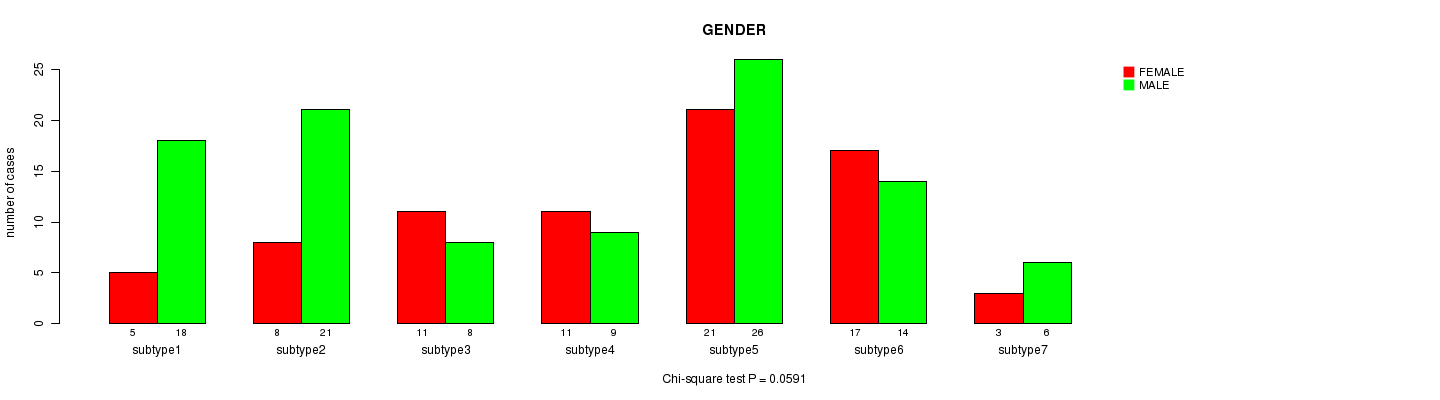
P value = 0.294 (ANOVA), Q value = 1
Table S47. Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 91 | 88.6 (10.5) |
| subtype1 | 13 | 93.8 (5.1) |
| subtype2 | 16 | 86.9 (12.0) |
| subtype3 | 11 | 91.8 (6.0) |
| subtype4 | 9 | 90.0 (10.0) |
| subtype5 | 21 | 85.7 (10.3) |
| subtype6 | 18 | 87.8 (12.2) |
| subtype7 | 3 | 83.3 (20.8) |
Figure S40. Get High-res Image Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
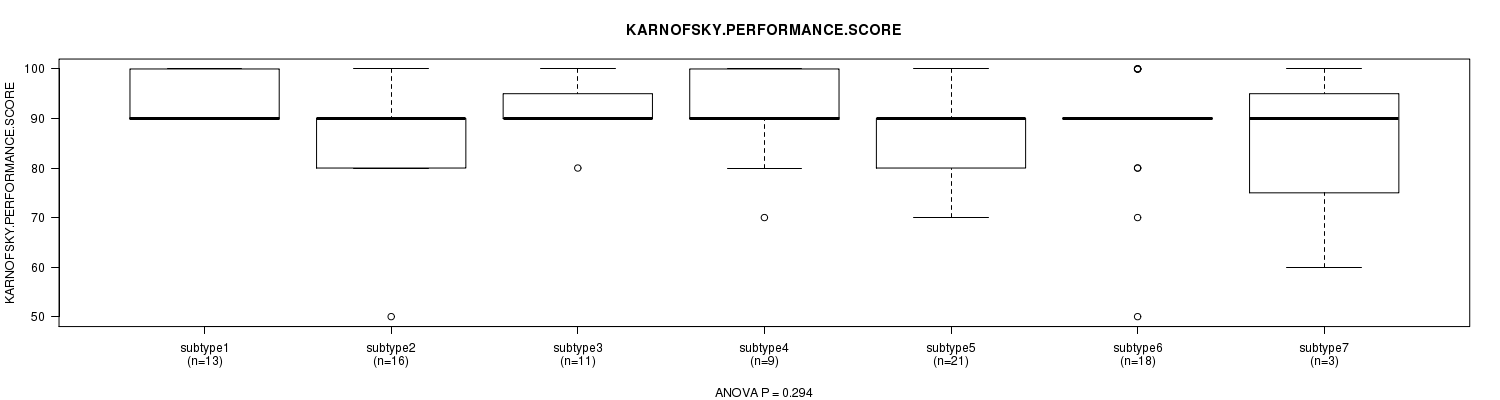
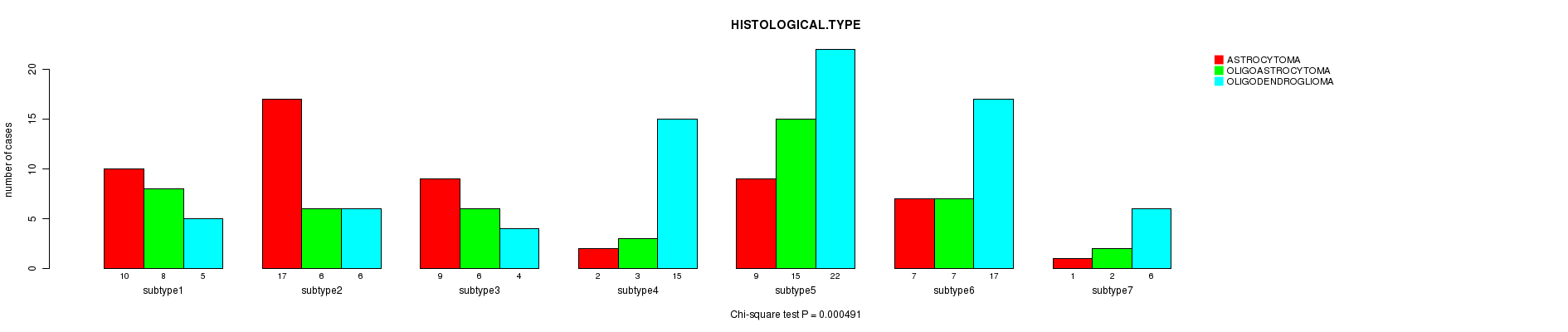
P value = 0.000491 (Chi-square test), Q value = 0.019
Table S48. Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 55 | 47 | 75 |
| subtype1 | 10 | 8 | 5 |
| subtype2 | 17 | 6 | 6 |
| subtype3 | 9 | 6 | 4 |
| subtype4 | 2 | 3 | 15 |
| subtype5 | 9 | 15 | 22 |
| subtype6 | 7 | 7 | 17 |
| subtype7 | 1 | 2 | 6 |
Figure S41. Get High-res Image Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
P value = 0.13 (Chi-square test), Q value = 1
Table S49. Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 95 | 83 |
| subtype1 | 9 | 14 |
| subtype2 | 19 | 10 |
| subtype3 | 11 | 8 |
| subtype4 | 9 | 11 |
| subtype5 | 21 | 26 |
| subtype6 | 22 | 9 |
| subtype7 | 4 | 5 |
Figure S42. Get High-res Image Clustering Approach #7: 'MIRseq CNMF subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
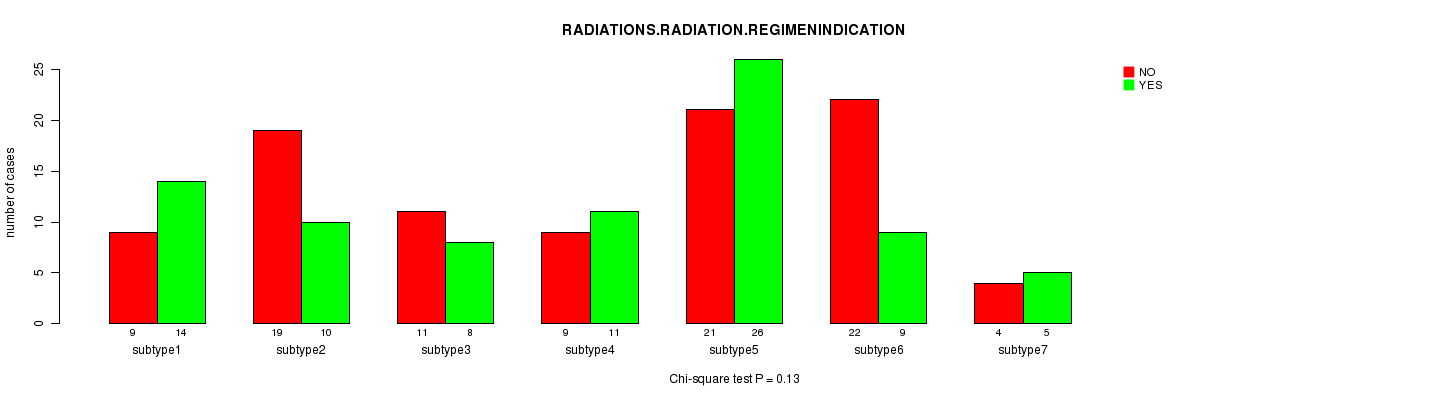
Table S50. Get Full Table Description of clustering approach #8: 'MIRseq cHierClus subtypes'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 7 | 94 | 77 |
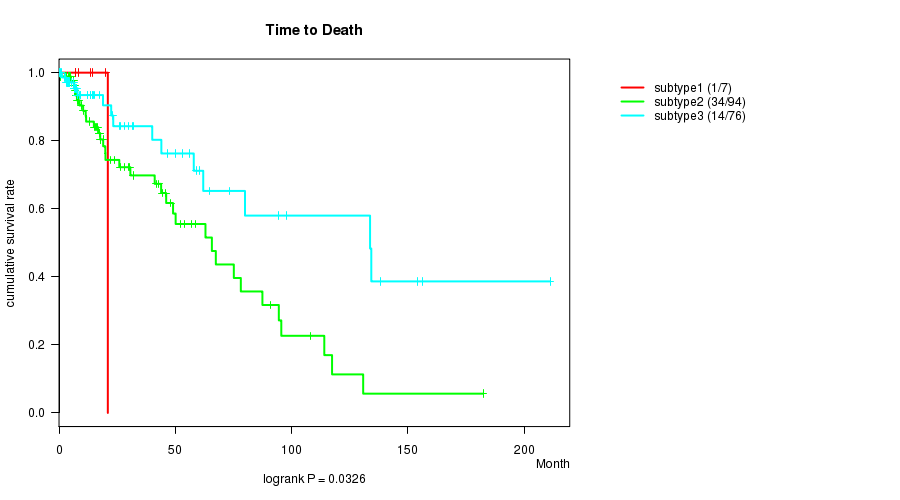
P value = 0.0326 (logrank test), Q value = 0.98
Table S51. Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 177 | 49 | 0.0 - 211.2 (14.7) |
| subtype1 | 7 | 1 | 7.1 - 21.0 (13.4) |
| subtype2 | 94 | 34 | 0.0 - 182.3 (17.0) |
| subtype3 | 76 | 14 | 0.1 - 211.2 (13.4) |
Figure S43. Get High-res Image Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
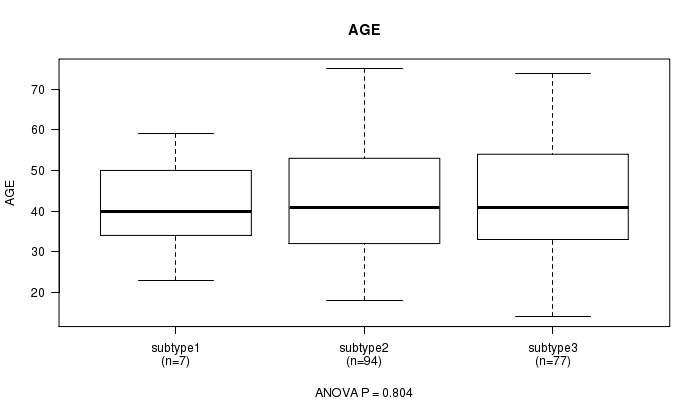
P value = 0.804 (ANOVA), Q value = 1
Table S52. Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 178 | 43.3 (13.3) |
| subtype1 | 7 | 41.4 (13.1) |
| subtype2 | 94 | 42.9 (13.1) |
| subtype3 | 77 | 44.0 (13.8) |
Figure S44. Get High-res Image Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #2: 'AGE'
P value = 0.693 (Fisher's exact test), Q value = 1
Table S53. Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #3: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 76 | 102 |
| subtype1 | 2 | 5 |
| subtype2 | 39 | 55 |
| subtype3 | 35 | 42 |
Figure S45. Get High-res Image Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #3: 'GENDER'
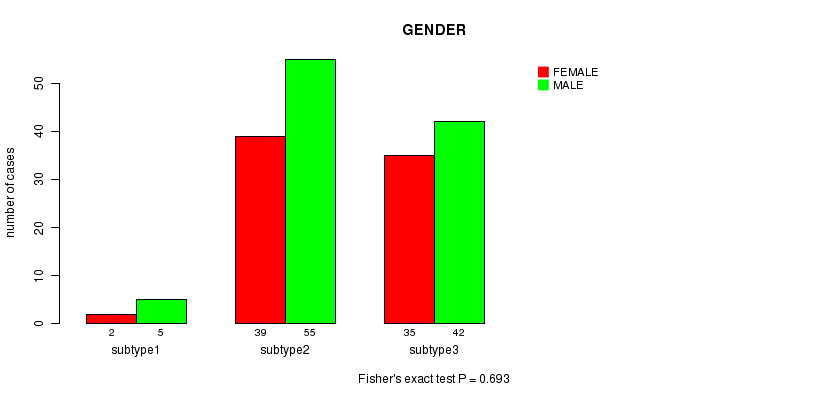
P value = 0.852 (ANOVA), Q value = 1
Table S54. Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 91 | 88.6 (10.5) |
| subtype1 | 5 | 88.0 (4.5) |
| subtype2 | 48 | 89.2 (9.9) |
| subtype3 | 38 | 87.9 (11.9) |
Figure S46. Get High-res Image Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #4: 'KARNOFSKY.PERFORMANCE.SCORE'
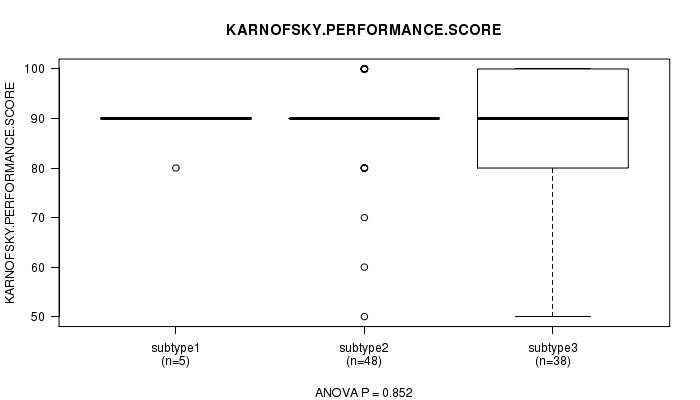
P value = 0.0496 (Chi-square test), Q value = 1
Table S55. Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 55 | 47 | 75 |
| subtype1 | 3 | 1 | 3 |
| subtype2 | 37 | 24 | 32 |
| subtype3 | 15 | 22 | 40 |
Figure S47. Get High-res Image Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #5: 'HISTOLOGICAL.TYPE'
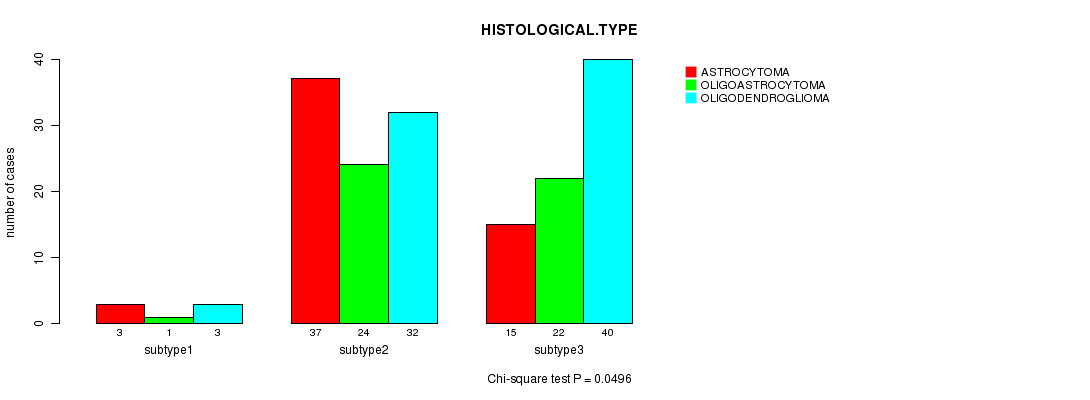
P value = 0.0781 (Fisher's exact test), Q value = 1
Table S56. Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 95 | 83 |
| subtype1 | 5 | 2 |
| subtype2 | 56 | 38 |
| subtype3 | 34 | 43 |
Figure S48. Get High-res Image Clustering Approach #8: 'MIRseq cHierClus subtypes' versus Clinical Feature #6: 'RADIATIONS.RADIATION.REGIMENINDICATION'
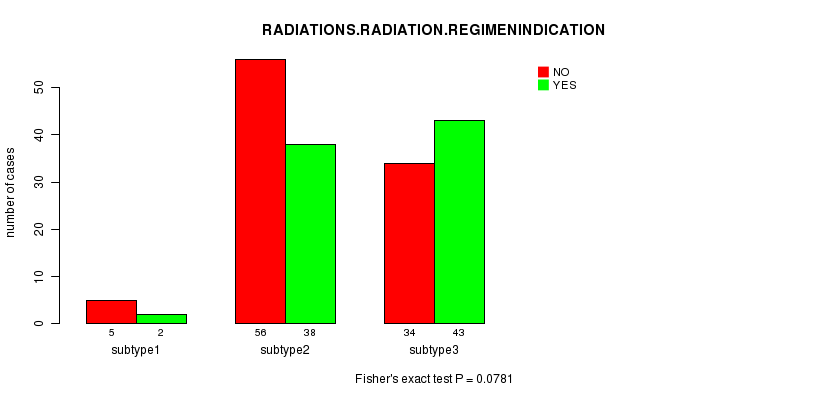
-
Cluster data file = LGG-TP.mergedcluster.txt
-
Clinical data file = LGG-TP.clin.merged.picked.txt
-
Number of patients = 179
-
Number of clustering approaches = 8
-
Number of selected clinical features = 6
-
Exclude small clusters that include fewer than K patients, K = 3
consensus non-negative matrix factorization clustering approach (Brunet et al. 2004)
Resampling-based clustering method (Monti et al. 2003)
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For continuous numerical clinical features, one-way analysis of variance (Howell 2002) was applied to compare the clinical values between tumor subtypes using 'anova' function in R
For binary clinical features, two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.