(primary solid tumor cohort)
This pipeline computes the correlation between significantly recurrent gene mutations and selected clinical features.
Testing the association between mutation status of 69 genes and 4 clinical features across 248 patients, 10 significant findings detected with Q value < 0.25.
-
PTEN mutation correlated to 'HISTOLOGICAL.TYPE'.
-
TP53 mutation correlated to 'AGE' and 'HISTOLOGICAL.TYPE'.
-
KRAS mutation correlated to 'AGE'.
-
PIK3R1 mutation correlated to 'HISTOLOGICAL.TYPE'.
-
CTNNB1 mutation correlated to 'HISTOLOGICAL.TYPE'.
-
CYLC1 mutation correlated to 'HISTOLOGICAL.TYPE'.
-
STEAP4 mutation correlated to 'HISTOLOGICAL.TYPE'.
-
ANKRD31 mutation correlated to 'HISTOLOGICAL.TYPE'.
-
ZNF69 mutation correlated to 'HISTOLOGICAL.TYPE'.
Table 1. Get Full Table Overview of the association between mutation status of 69 genes and 4 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 10 significant findings detected.
|
Clinical Features |
Time to Death |
AGE |
HISTOLOGICAL TYPE |
RADIATIONS RADIATION REGIMENINDICATION |
||
| nMutated (%) | nWild-Type | logrank test | t-test | Chi-square test | Fisher's exact test | |
| TP53 | 69 (28%) | 179 |
0.325 (1.00) |
6.15e-06 (0.00168) |
3.98e-23 (1.1e-20) |
0.371 (1.00) |
| PTEN | 161 (65%) | 87 |
0.19 (1.00) |
0.0276 (1.00) |
5.44e-19 (1.5e-16) |
0.0503 (1.00) |
| KRAS | 53 (21%) | 195 |
0.072 (1.00) |
0.000274 (0.0739) |
0.00692 (1.00) |
0.253 (1.00) |
| PIK3R1 | 83 (33%) | 165 |
0.871 (1.00) |
0.806 (1.00) |
0.000827 (0.221) |
0.395 (1.00) |
| CTNNB1 | 74 (30%) | 174 |
0.878 (1.00) |
0.0017 (0.449) |
1.04e-05 (0.00284) |
0.381 (1.00) |
| CYLC1 | 18 (7%) | 230 |
0.198 (1.00) |
0.765 (1.00) |
0.000601 (0.162) |
0.313 (1.00) |
| STEAP4 | 12 (5%) | 236 |
0.286 (1.00) |
0.933 (1.00) |
0.000714 (0.191) |
0.756 (1.00) |
| ANKRD31 | 16 (6%) | 232 |
0.234 (1.00) |
0.883 (1.00) |
9.81e-05 (0.0267) |
0.588 (1.00) |
| ZNF69 | 7 (3%) | 241 |
0.514 (1.00) |
0.0163 (1.00) |
0.000147 (0.0397) |
1 (1.00) |
| ARID1A | 83 (33%) | 165 |
0.008 (1.00) |
0.00721 (1.00) |
0.00127 (0.337) |
0.673 (1.00) |
| FGFR2 | 31 (12%) | 217 |
0.9 (1.00) |
0.998 (1.00) |
0.719 (1.00) |
1 (1.00) |
| PIK3CA | 132 (53%) | 116 |
0.055 (1.00) |
0.195 (1.00) |
0.555 (1.00) |
0.593 (1.00) |
| CTCF | 45 (18%) | 203 |
0.405 (1.00) |
0.0198 (1.00) |
0.0021 (0.552) |
0.863 (1.00) |
| FBXW7 | 39 (16%) | 209 |
0.772 (1.00) |
0.904 (1.00) |
0.0473 (1.00) |
0.855 (1.00) |
| SPOP | 21 (8%) | 227 |
0.65 (1.00) |
0.527 (1.00) |
0.332 (1.00) |
0.811 (1.00) |
| PPP2R1A | 27 (11%) | 221 |
0.121 (1.00) |
0.0374 (1.00) |
0.00977 (1.00) |
0.52 (1.00) |
| CHD4 | 35 (14%) | 213 |
0.361 (1.00) |
0.418 (1.00) |
0.779 (1.00) |
0.565 (1.00) |
| P2RY11 | 9 (4%) | 239 |
0.395 (1.00) |
0.396 (1.00) |
0.987 (1.00) |
0.497 (1.00) |
| CCND1 | 15 (6%) | 233 |
0.729 (1.00) |
0.0556 (1.00) |
0.413 (1.00) |
0.4 (1.00) |
| NFE2L2 | 15 (6%) | 233 |
0.903 (1.00) |
0.191 (1.00) |
0.447 (1.00) |
0.589 (1.00) |
| FOXA2 | 12 (5%) | 236 |
0.799 (1.00) |
0.531 (1.00) |
0.972 (1.00) |
0.229 (1.00) |
| FAM9A | 14 (6%) | 234 |
0.378 (1.00) |
0.913 (1.00) |
0.89 (1.00) |
1 (1.00) |
| SOX17 | 7 (3%) | 241 |
0.669 (1.00) |
0.848 (1.00) |
0.00567 (1.00) |
0.428 (1.00) |
| MAX | 11 (4%) | 237 |
0.889 (1.00) |
0.336 (1.00) |
0.404 (1.00) |
0.753 (1.00) |
| ZNF267 | 16 (6%) | 232 |
0.565 (1.00) |
0.207 (1.00) |
0.0275 (1.00) |
0.79 (1.00) |
| RBMX | 13 (5%) | 235 |
0.336 (1.00) |
0.161 (1.00) |
0.4 (1.00) |
0.142 (1.00) |
| MORC4 | 20 (8%) | 228 |
0.152 (1.00) |
0.0204 (1.00) |
0.629 (1.00) |
1 (1.00) |
| OR4K2 | 11 (4%) | 237 |
0.304 (1.00) |
0.241 (1.00) |
0.00658 (1.00) |
1 (1.00) |
| ARID5B | 29 (12%) | 219 |
0.642 (1.00) |
0.649 (1.00) |
0.00865 (1.00) |
1 (1.00) |
| OR5D13 | 10 (4%) | 238 |
0.895 (1.00) |
0.741 (1.00) |
0.188 (1.00) |
0.318 (1.00) |
| OR5AS1 | 10 (4%) | 238 |
0.675 (1.00) |
0.503 (1.00) |
0.396 (1.00) |
0.501 (1.00) |
| OR4A15 | 13 (5%) | 235 |
0.213 (1.00) |
0.167 (1.00) |
0.00219 (0.573) |
1 (1.00) |
| CSDE1 | 21 (8%) | 227 |
0.802 (1.00) |
0.294 (1.00) |
0.0435 (1.00) |
1 (1.00) |
| CTXN3 | 6 (2%) | 242 |
0.512 (1.00) |
0.794 (1.00) |
0.548 (1.00) |
1 (1.00) |
| PAPD4 | 13 (5%) | 235 |
0.982 (1.00) |
0.181 (1.00) |
0.962 (1.00) |
0.378 (1.00) |
| RPL14 | 7 (3%) | 241 |
0.532 (1.00) |
0.677 (1.00) |
0.991 (1.00) |
0.428 (1.00) |
| DNER | 18 (7%) | 230 |
0.746 (1.00) |
0.0414 (1.00) |
0.0324 (1.00) |
0.44 (1.00) |
| RASA1 | 22 (9%) | 226 |
0.127 (1.00) |
0.466 (1.00) |
0.0211 (1.00) |
0.639 (1.00) |
| TAB3 | 18 (7%) | 230 |
0.22 (1.00) |
0.107 (1.00) |
0.00686 (1.00) |
0.616 (1.00) |
| UPF3B | 16 (6%) | 232 |
0.735 (1.00) |
0.81 (1.00) |
0.395 (1.00) |
0.588 (1.00) |
| SGK1 | 15 (6%) | 233 |
0.925 (1.00) |
0.427 (1.00) |
0.759 (1.00) |
0.158 (1.00) |
| C1ORF100 | 9 (4%) | 239 |
0.344 (1.00) |
0.246 (1.00) |
0.376 (1.00) |
0.497 (1.00) |
| ZNF334 | 17 (7%) | 231 |
0.95 (1.00) |
0.0232 (1.00) |
0.787 (1.00) |
1 (1.00) |
| METTL14 | 10 (4%) | 238 |
0.358 (1.00) |
0.833 (1.00) |
0.707 (1.00) |
1 (1.00) |
| NRAS | 9 (4%) | 239 |
0.354 (1.00) |
0.616 (1.00) |
0.0289 (1.00) |
0.722 (1.00) |
| HIST1H2BD | 6 (2%) | 242 |
0.0836 (1.00) |
0.474 (1.00) |
0.897 (1.00) |
0.415 (1.00) |
| CLIC2 | 10 (4%) | 238 |
0.294 (1.00) |
0.855 (1.00) |
0.00231 (0.601) |
1 (1.00) |
| CNPY1 | 7 (3%) | 241 |
0.449 (1.00) |
0.18 (1.00) |
0.294 (1.00) |
1 (1.00) |
| RAE1 | 11 (4%) | 237 |
0.282 (1.00) |
0.603 (1.00) |
0.0127 (1.00) |
0.34 (1.00) |
| IGFBP7 | 6 (2%) | 242 |
0.406 (1.00) |
0.659 (1.00) |
0.228 (1.00) |
0.667 (1.00) |
| ZNF300 | 15 (6%) | 233 |
0.222 (1.00) |
0.338 (1.00) |
0.135 (1.00) |
0.275 (1.00) |
| SNTG1 | 11 (4%) | 237 |
0.307 (1.00) |
0.506 (1.00) |
0.0644 (1.00) |
0.753 (1.00) |
| SSX5 | 11 (4%) | 237 |
0.328 (1.00) |
0.15 (1.00) |
0.00658 (1.00) |
1 (1.00) |
| FMR1 | 16 (6%) | 232 |
0.304 (1.00) |
0.643 (1.00) |
0.0688 (1.00) |
0.588 (1.00) |
| GYPB | 5 (2%) | 243 |
0.583 (1.00) |
0.843 (1.00) |
0.149 (1.00) |
0.168 (1.00) |
| LRRIQ3 | 12 (5%) | 236 |
0.372 (1.00) |
0.00386 (1) |
0.00221 (0.578) |
0.229 (1.00) |
| NDN | 6 (2%) | 242 |
0.535 (1.00) |
0.487 (1.00) |
0.699 (1.00) |
1 (1.00) |
| C14ORF118 | 15 (6%) | 233 |
0.268 (1.00) |
0.0138 (1.00) |
0.369 (1.00) |
0.589 (1.00) |
| COX19 | 4 (2%) | 244 |
0.652 (1.00) |
0.269 (1.00) |
0.998 (1.00) |
1 (1.00) |
| GNPDA2 | 8 (3%) | 240 |
0.789 (1.00) |
0.175 (1.00) |
0.518 (1.00) |
0.451 (1.00) |
| GPM6A | 11 (4%) | 237 |
0.306 (1.00) |
0.502 (1.00) |
0.545 (1.00) |
1 (1.00) |
| ZNF449 | 12 (5%) | 236 |
0.355 (1.00) |
0.95 (1.00) |
0.655 (1.00) |
0.552 (1.00) |
| OR2M3 | 10 (4%) | 238 |
0.315 (1.00) |
0.822 (1.00) |
0.707 (1.00) |
0.739 (1.00) |
| XPA | 7 (3%) | 241 |
0.169 (1.00) |
0.9 (1.00) |
0.981 (1.00) |
0.236 (1.00) |
| BRS3 | 15 (6%) | 233 |
0.184 (1.00) |
0.133 (1.00) |
0.025 (1.00) |
0.4 (1.00) |
| GK2 | 17 (7%) | 231 |
0.202 (1.00) |
0.334 (1.00) |
0.0119 (1.00) |
0.432 (1.00) |
| TPTE | 15 (6%) | 233 |
0.273 (1.00) |
0.167 (1.00) |
0.00648 (1.00) |
1 (1.00) |
| IL24 | 7 (3%) | 241 |
0.426 (1.00) |
0.988 (1.00) |
0.00148 (0.392) |
0.428 (1.00) |
| CCDC160 | 11 (4%) | 237 |
0.271 (1.00) |
0.34 (1.00) |
0.00658 (1.00) |
1 (1.00) |
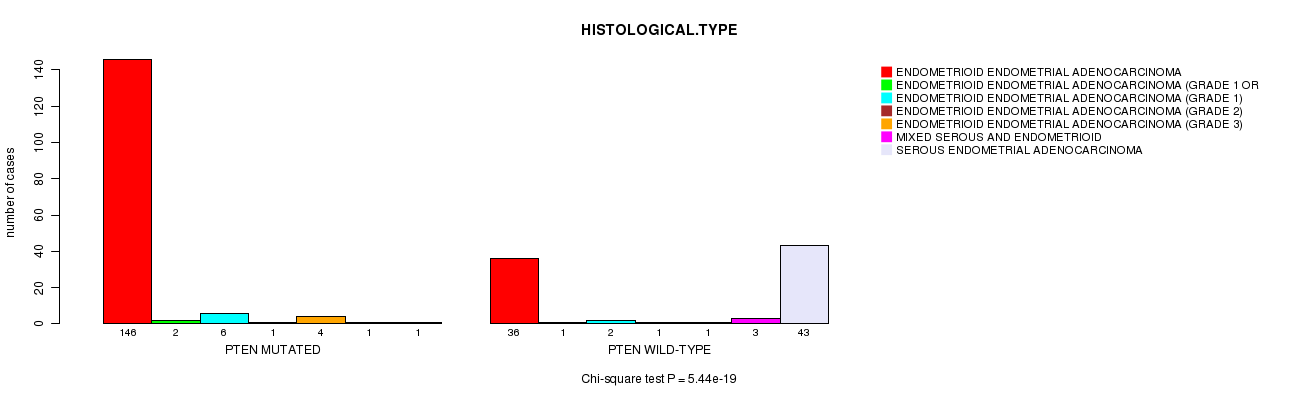
P value = 5.44e-19 (Chi-square test), Q value = 1.5e-16
Table S1. Gene #4: 'PTEN MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1 OR 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 3) | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|---|---|---|---|
| ALL | 182 | 3 | 8 | 2 | 5 | 4 | 44 |
| PTEN MUTATED | 146 | 2 | 6 | 1 | 4 | 1 | 1 |
| PTEN WILD-TYPE | 36 | 1 | 2 | 1 | 1 | 3 | 43 |
Figure S1. Get High-res Image Gene #4: 'PTEN MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
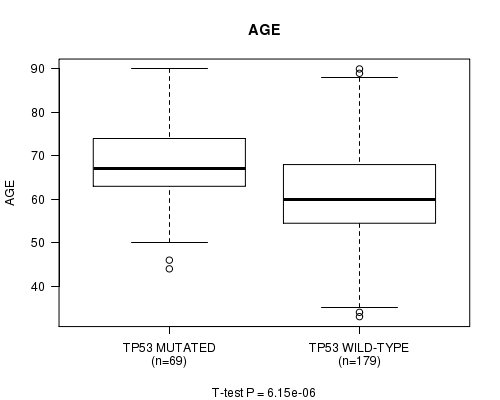
P value = 6.15e-06 (t-test), Q value = 0.0017
Table S2. Gene #5: 'TP53 MUTATION STATUS' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 248 | 63.1 (11.1) |
| TP53 MUTATED | 69 | 67.9 (9.4) |
| TP53 WILD-TYPE | 179 | 61.3 (11.2) |
Figure S2. Get High-res Image Gene #5: 'TP53 MUTATION STATUS' versus Clinical Feature #2: 'AGE'
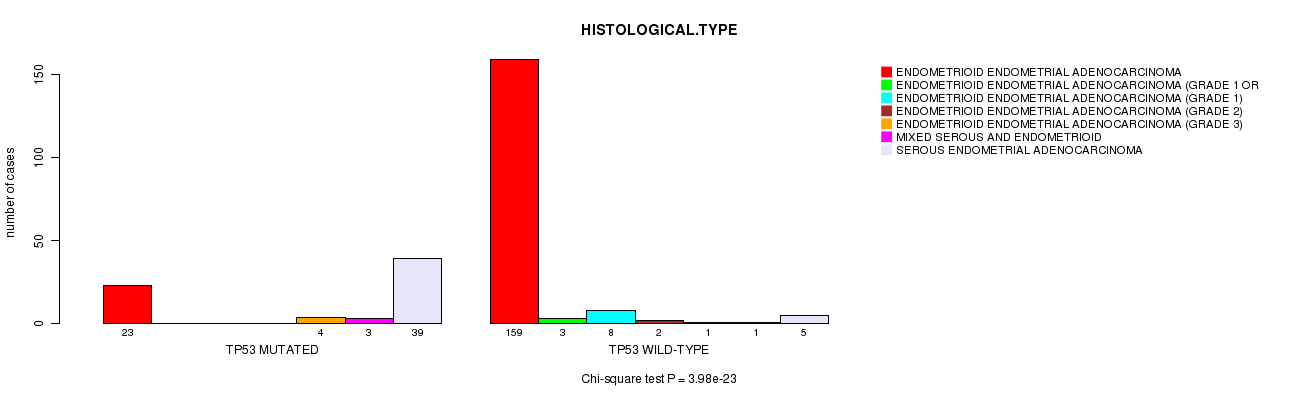
P value = 3.98e-23 (Chi-square test), Q value = 1.1e-20
Table S3. Gene #5: 'TP53 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1 OR 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 3) | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|---|---|---|---|
| ALL | 182 | 3 | 8 | 2 | 5 | 4 | 44 |
| TP53 MUTATED | 23 | 0 | 0 | 0 | 4 | 3 | 39 |
| TP53 WILD-TYPE | 159 | 3 | 8 | 2 | 1 | 1 | 5 |
Figure S3. Get High-res Image Gene #5: 'TP53 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
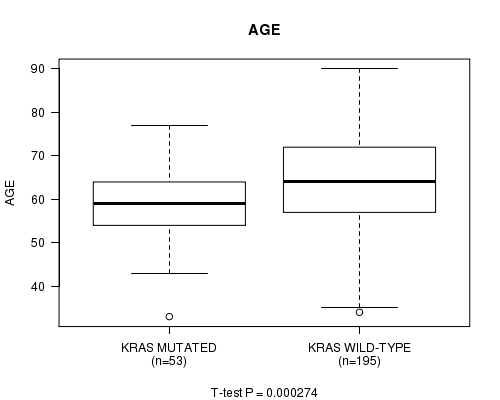
P value = 0.000274 (t-test), Q value = 0.074
Table S4. Gene #7: 'KRAS MUTATION STATUS' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 248 | 63.1 (11.1) |
| KRAS MUTATED | 53 | 58.8 (8.7) |
| KRAS WILD-TYPE | 195 | 64.3 (11.4) |
Figure S4. Get High-res Image Gene #7: 'KRAS MUTATION STATUS' versus Clinical Feature #2: 'AGE'
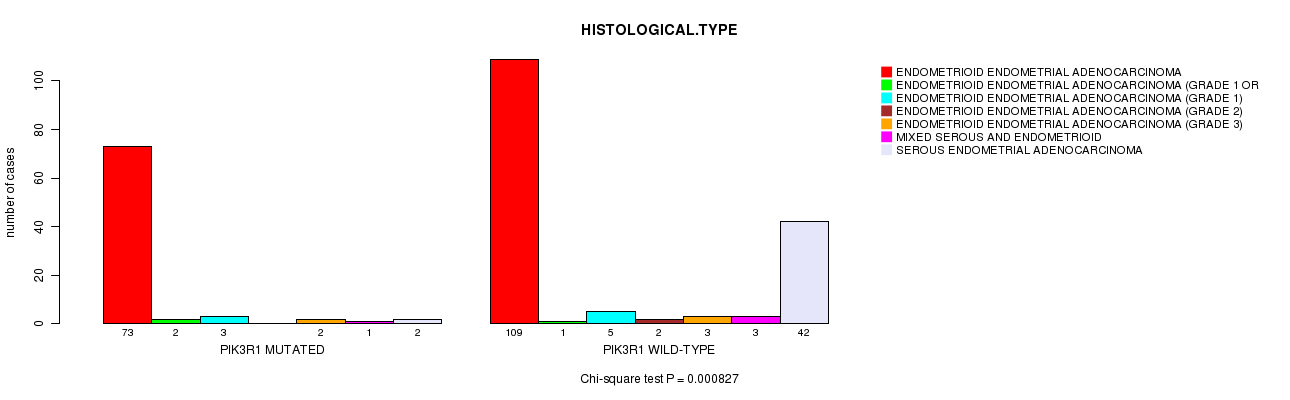
P value = 0.000827 (Chi-square test), Q value = 0.22
Table S5. Gene #8: 'PIK3R1 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1 OR 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 3) | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|---|---|---|---|
| ALL | 182 | 3 | 8 | 2 | 5 | 4 | 44 |
| PIK3R1 MUTATED | 73 | 2 | 3 | 0 | 2 | 1 | 2 |
| PIK3R1 WILD-TYPE | 109 | 1 | 5 | 2 | 3 | 3 | 42 |
Figure S5. Get High-res Image Gene #8: 'PIK3R1 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
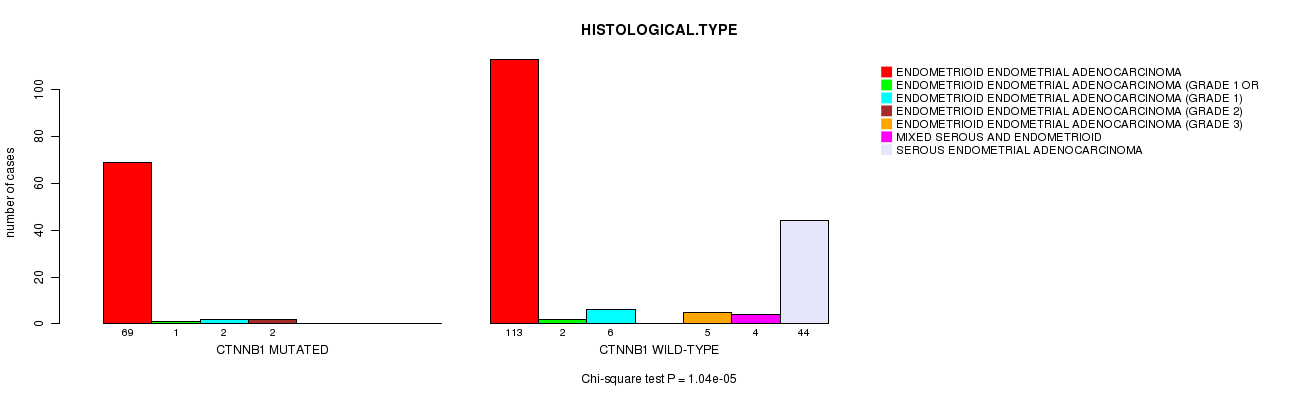
P value = 1.04e-05 (Chi-square test), Q value = 0.0028
Table S6. Gene #9: 'CTNNB1 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1 OR 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 3) | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|---|---|---|---|
| ALL | 182 | 3 | 8 | 2 | 5 | 4 | 44 |
| CTNNB1 MUTATED | 69 | 1 | 2 | 2 | 0 | 0 | 0 |
| CTNNB1 WILD-TYPE | 113 | 2 | 6 | 0 | 5 | 4 | 44 |
Figure S6. Get High-res Image Gene #9: 'CTNNB1 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
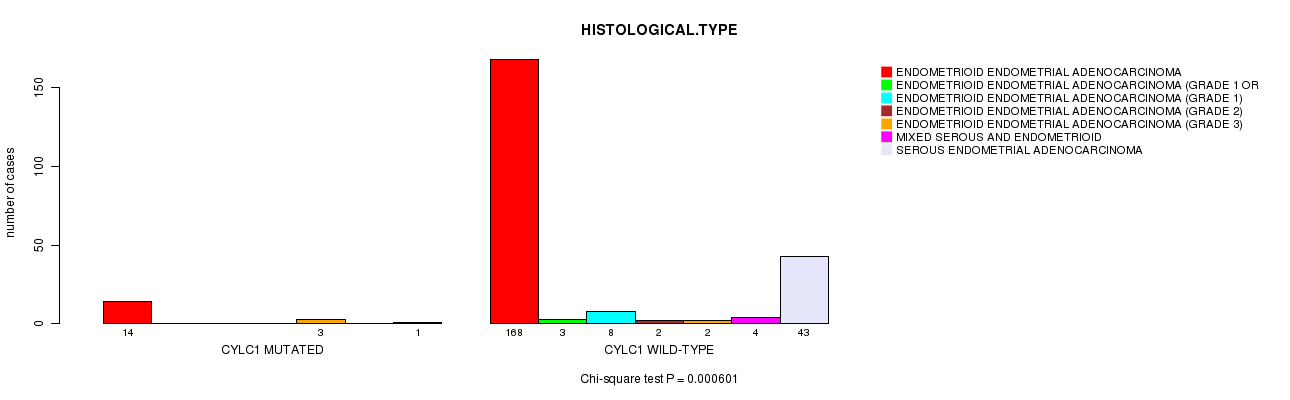
P value = 0.000601 (Chi-square test), Q value = 0.16
Table S7. Gene #22: 'CYLC1 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1 OR 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 3) | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|---|---|---|---|
| ALL | 182 | 3 | 8 | 2 | 5 | 4 | 44 |
| CYLC1 MUTATED | 14 | 0 | 0 | 0 | 3 | 0 | 1 |
| CYLC1 WILD-TYPE | 168 | 3 | 8 | 2 | 2 | 4 | 43 |
Figure S7. Get High-res Image Gene #22: 'CYLC1 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
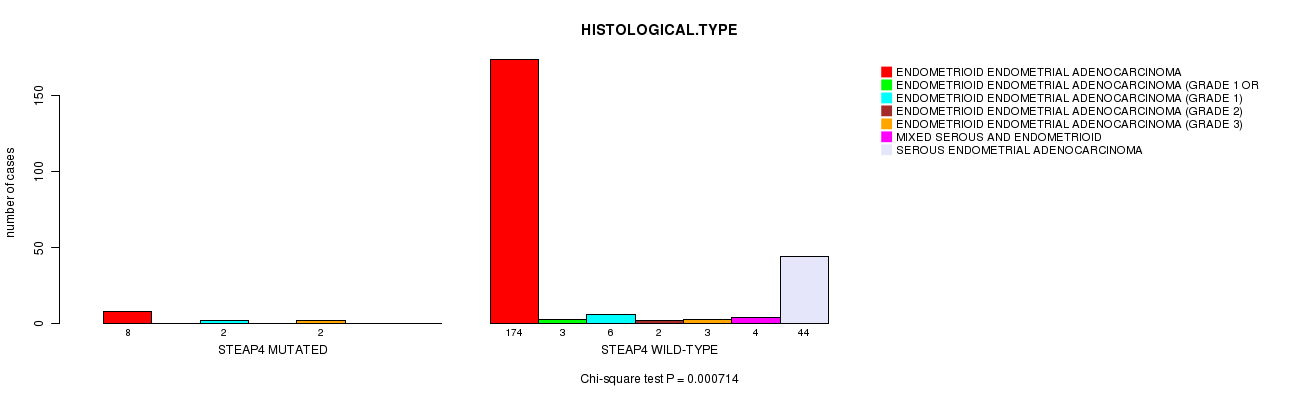
P value = 0.000714 (Chi-square test), Q value = 0.19
Table S8. Gene #44: 'STEAP4 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1 OR 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 3) | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|---|---|---|---|
| ALL | 182 | 3 | 8 | 2 | 5 | 4 | 44 |
| STEAP4 MUTATED | 8 | 0 | 2 | 0 | 2 | 0 | 0 |
| STEAP4 WILD-TYPE | 174 | 3 | 6 | 2 | 3 | 4 | 44 |
Figure S8. Get High-res Image Gene #44: 'STEAP4 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
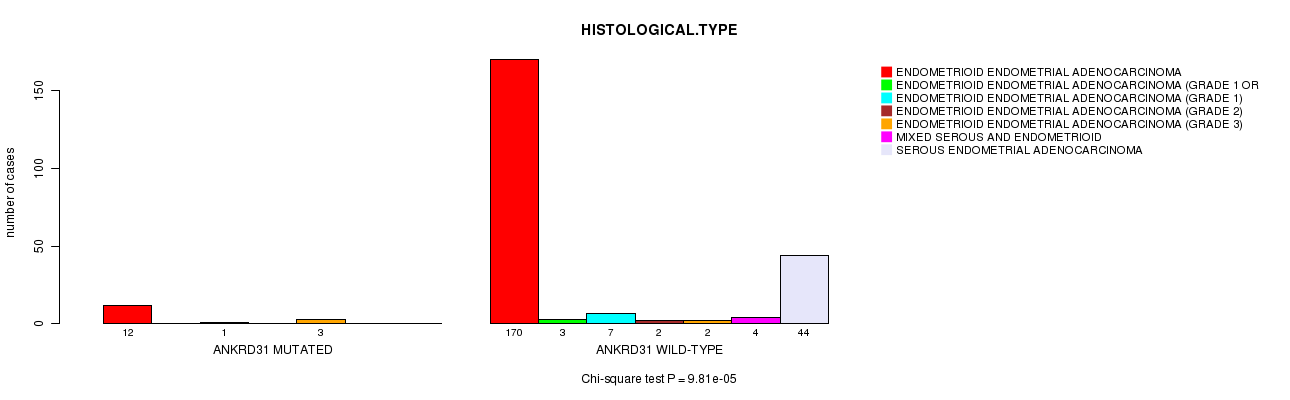
P value = 9.81e-05 (Chi-square test), Q value = 0.027
Table S9. Gene #46: 'ANKRD31 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1 OR 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 3) | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|---|---|---|---|
| ALL | 182 | 3 | 8 | 2 | 5 | 4 | 44 |
| ANKRD31 MUTATED | 12 | 0 | 1 | 0 | 3 | 0 | 0 |
| ANKRD31 WILD-TYPE | 170 | 3 | 7 | 2 | 2 | 4 | 44 |
Figure S9. Get High-res Image Gene #46: 'ANKRD31 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
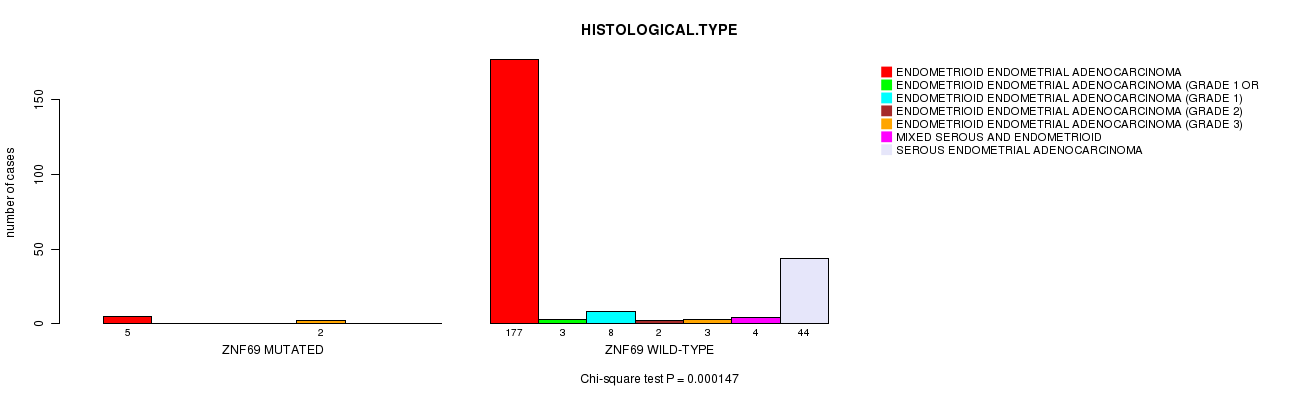
P value = 0.000147 (Chi-square test), Q value = 0.04
Table S10. Gene #64: 'ZNF69 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1 OR 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 1) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 2) | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA (GRADE 3) | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|---|---|---|---|
| ALL | 182 | 3 | 8 | 2 | 5 | 4 | 44 |
| ZNF69 MUTATED | 5 | 0 | 0 | 0 | 2 | 0 | 0 |
| ZNF69 WILD-TYPE | 177 | 3 | 8 | 2 | 3 | 4 | 44 |
Figure S10. Get High-res Image Gene #64: 'ZNF69 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL.TYPE'
-
Mutation data file = UCEC-TP.mutsig.cluster.txt
-
Clinical data file = UCEC-TP.clin.merged.picked.txt
-
Number of patients = 248
-
Number of significantly mutated genes = 69
-
Number of selected clinical features = 4
-
Exclude genes that fewer than K tumors have mutations, K = 3
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For continuous numerical clinical features, two-tailed Student's t test with unequal variance (Lehmann and Romano 2005) was applied to compare the clinical values between tumors with and without gene mutations using 't.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.