This pipeline computes the correlation between significantly recurrent gene mutations and molecular subtypes.
Testing the association between mutation status of 209 genes and 10 molecular subtypes across 224 patients, 38 significant findings detected with P value < 0.05 and Q value < 0.25.
-
FBXW7 mutation correlated to 'CN_CNMF'.
-
BRAF mutation correlated to 'MRNA_CNMF', 'MRNA_CHIERARCHICAL', 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
KRAS mutation correlated to 'MRNASEQ_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
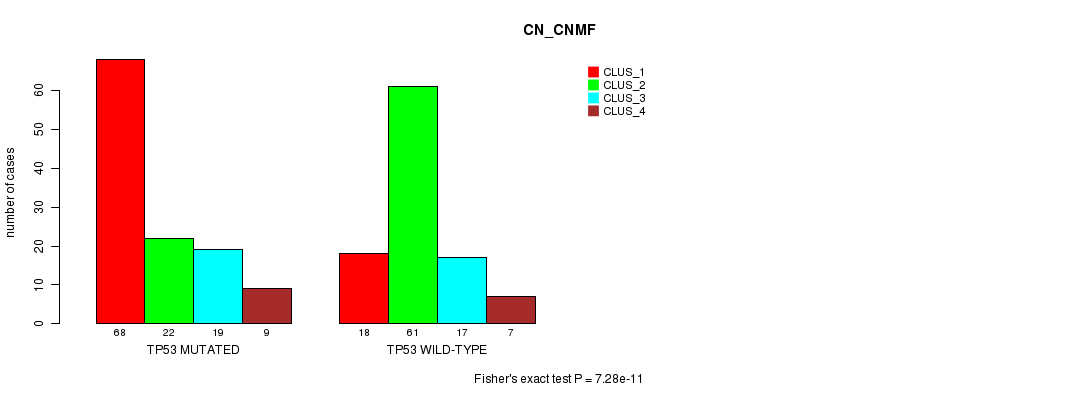
TP53 mutation correlated to 'CN_CNMF'.
-
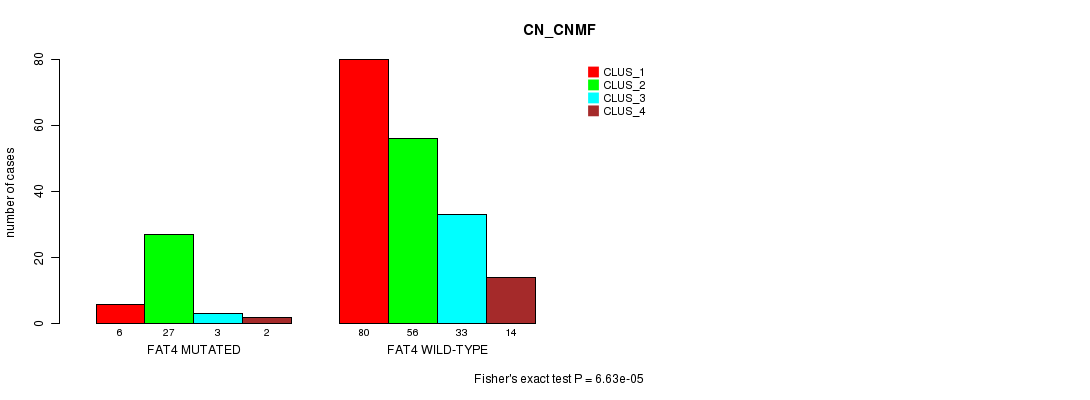
FAT4 mutation correlated to 'CN_CNMF'.
-
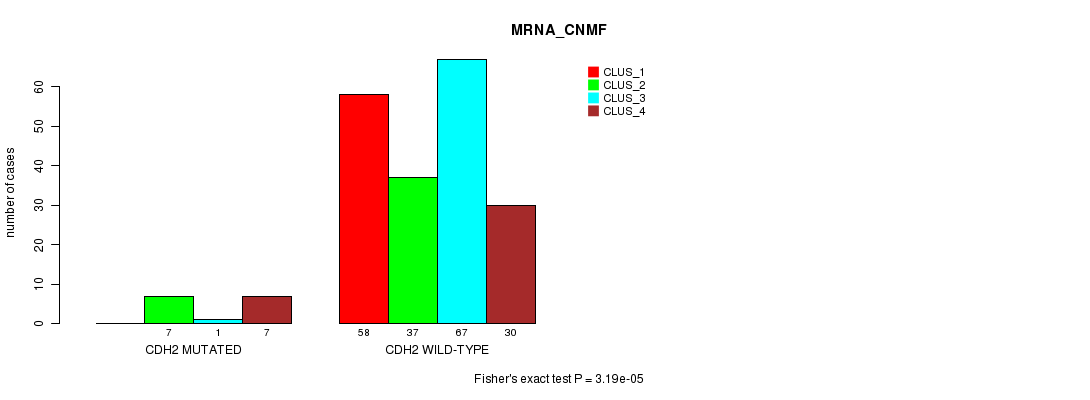
CDH2 mutation correlated to 'MRNA_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
FLG mutation correlated to 'MRNA_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
DNAH5 mutation correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
SDK1 mutation correlated to 'MRNASEQ_CHIERARCHICAL'.
-
TMEM132D mutation correlated to 'MRNA_CNMF'.
-
LRRN3 mutation correlated to 'MRNASEQ_CNMF'.
-
GUCY1A3 mutation correlated to 'MRNASEQ_CHIERARCHICAL'.
-
OR51V1 mutation correlated to 'MRNASEQ_CHIERARCHICAL'.
-
ZC3H13 mutation correlated to 'CN_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
CSMD3 mutation correlated to 'MRNASEQ_CHIERARCHICAL'.
-
DCAF4L2 mutation correlated to 'MRNASEQ_CHIERARCHICAL'.
-
DOCK2 mutation correlated to 'MRNA_CNMF', 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
RIMS1 mutation correlated to 'MRNASEQ_CHIERARCHICAL'.
-
CDH10 mutation correlated to 'MRNA_CNMF', 'CN_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
SYNE1 mutation correlated to 'MRNA_CNMF', 'MRNA_CHIERARCHICAL', and 'CN_CNMF'.
Table 1. Get Full Table Overview of the association between mutation status of 209 genes and 10 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 38 significant findings detected.
|
Clinical Features |
MRNA CNMF |
MRNA CHIERARCHICAL |
CN CNMF |
METHLYATION CNMF |
RPPA CNMF |
RPPA CHIERARCHICAL |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
||
| nMutated (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| BRAF | 22 (10%) | 202 |
1.13e-06 (0.00205) |
1.26e-07 (0.000228) |
4.17e-05 (0.0749) |
0.875 (1.00) |
0.227 (1.00) |
3.49e-08 (6.34e-05) |
1.92e-10 (3.49e-07) |
0.6 (1.00) |
1 (1.00) |
|
| DOCK2 | 21 (9%) | 203 |
0.000122 (0.217) |
0.00397 (1.00) |
0.000138 (0.245) |
0.0958 (1.00) |
0.779 (1.00) |
1.35e-05 (0.0244) |
7.91e-07 (0.00143) |
1 (1.00) |
0.101 (1.00) |
|
| CDH2 | 16 (7%) | 208 |
3.19e-05 (0.0573) |
0.00563 (1.00) |
0.00381 (1.00) |
0.0336 (1.00) |
0.817 (1.00) |
5.18e-05 (0.093) |
1.1e-05 (0.0198) |
0.41 (1.00) |
0.683 (1.00) |
|
| DNAH5 | 34 (15%) | 190 |
0.000387 (0.683) |
0.00585 (1.00) |
0.00013 (0.231) |
0.62 (1.00) |
0.301 (1.00) |
7.52e-05 (0.135) |
1.52e-06 (0.00275) |
1 (1.00) |
0.238 (1.00) |
|
| CDH10 | 14 (6%) | 210 |
4.46e-05 (0.0801) |
0.00333 (1.00) |
0.000131 (0.233) |
0.336 (1.00) |
0.605 (1.00) |
0.00456 (1.00) |
2.68e-05 (0.0483) |
1 (1.00) |
0.257 (1.00) |
|
| SYNE1 | 47 (21%) | 177 |
2.86e-05 (0.0514) |
2.56e-05 (0.046) |
1.63e-06 (0.00295) |
1 (1.00) |
0.641 (1.00) |
0.419 (1.00) |
0.0204 (1.00) |
0.000383 (0.677) |
0.367 (1.00) |
0.0326 (1.00) |
| KRAS | 96 (43%) | 128 |
0.000775 (1.00) |
0.00193 (1.00) |
0.485 (1.00) |
0.257 (1.00) |
0.742 (1.00) |
0.946 (1.00) |
1.94e-07 (0.000351) |
3.02e-07 (0.000547) |
0.0688 (1.00) |
0.735 (1.00) |
| FLG | 32 (14%) | 192 |
2.36e-06 (0.00426) |
0.00102 (1.00) |
0.00376 (1.00) |
0.0122 (1.00) |
0.813 (1.00) |
0.00123 (1.00) |
4.08e-05 (0.0734) |
1 (1.00) |
0.74 (1.00) |
|
| ZC3H13 | 21 (9%) | 203 |
0.000395 (0.697) |
0.000571 (1.00) |
8.41e-05 (0.15) |
0.345 (1.00) |
0.676 (1.00) |
0.000883 (1.00) |
6.7e-05 (0.12) |
1 (1.00) |
0.306 (1.00) |
|
| FBXW7 | 38 (17%) | 186 |
0.0186 (1.00) |
0.0302 (1.00) |
0.000122 (0.217) |
0.0191 (1.00) |
0.126 (1.00) |
0.000223 (0.397) |
0.000368 (0.651) |
0.356 (1.00) |
0.118 (1.00) |
|
| TP53 | 120 (54%) | 104 |
0.00037 (0.654) |
0.00159 (1.00) |
7.28e-11 (1.32e-07) |
0.887 (1.00) |
0.658 (1.00) |
0.00978 (1.00) |
0.00162 (1.00) |
0.0252 (1.00) |
0.134 (1.00) |
|
| FAT4 | 39 (17%) | 185 |
0.0134 (1.00) |
0.08 (1.00) |
6.63e-05 (0.119) |
0.597 (1.00) |
0.0868 (1.00) |
0.0448 (1.00) |
0.00381 (1.00) |
0.635 (1.00) |
0.683 (1.00) |
|
| SDK1 | 27 (12%) | 197 |
0.00909 (1.00) |
0.00366 (1.00) |
0.24 (1.00) |
0.764 (1.00) |
0.941 (1.00) |
0.00143 (1.00) |
0.000102 (0.182) |
0.6 (1.00) |
1 (1.00) |
|
| TMEM132D | 19 (8%) | 205 |
2.85e-05 (0.0512) |
0.00157 (1.00) |
0.00293 (1.00) |
0.0366 (1.00) |
0.913 (1.00) |
0.00222 (1.00) |
0.00162 (1.00) |
1 (1.00) |
1 (1.00) |
|
| LRRN3 | 10 (4%) | 214 |
0.00243 (1.00) |
0.123 (1.00) |
0.446 (1.00) |
0.387 (1.00) |
0.528 (1.00) |
9.57e-05 (0.171) |
0.0318 (1.00) |
0.274 (1.00) |
0.502 (1.00) |
|
| GUCY1A3 | 12 (5%) | 212 |
0.38 (1.00) |
0.172 (1.00) |
0.0311 (1.00) |
0.825 (1.00) |
1 (1.00) |
0.0145 (1.00) |
7.22e-05 (0.129) |
1 (1.00) |
0.584 (1.00) |
|
| OR51V1 | 9 (4%) | 215 |
0.00262 (1.00) |
0.00756 (1.00) |
0.0016 (1.00) |
0.799 (1.00) |
1 (1.00) |
0.000361 (0.639) |
2.77e-06 (0.005) |
1 (1.00) |
0.545 (1.00) |
|
| CSMD3 | 28 (12%) | 196 |
0.00625 (1.00) |
0.0145 (1.00) |
0.00457 (1.00) |
0.19 (1.00) |
0.0678 (1.00) |
0.0221 (1.00) |
0.000129 (0.23) |
0.6 (1.00) |
0.0587 (1.00) |
|
| DCAF4L2 | 11 (5%) | 213 |
0.00355 (1.00) |
0.0326 (1.00) |
0.027 (1.00) |
0.0276 (1.00) |
0.000116 (0.208) |
0.304 (1.00) |
0.545 (1.00) |
|||
| RIMS1 | 15 (7%) | 209 |
0.24 (1.00) |
0.423 (1.00) |
0.00135 (1.00) |
0.0193 (1.00) |
0.795 (1.00) |
0.0338 (1.00) |
0.000103 (0.183) |
0.458 (1.00) |
0.736 (1.00) |
|
| NRAS | 20 (9%) | 204 |
0.0358 (1.00) |
0.33 (1.00) |
0.274 (1.00) |
0.857 (1.00) |
0.676 (1.00) |
0.25 (1.00) |
0.636 (1.00) |
1 (1.00) |
1 (1.00) |
|
| PIK3CA | 33 (15%) | 191 |
0.194 (1.00) |
0.38 (1.00) |
0.00048 (0.844) |
0.0323 (1.00) |
0.0748 (1.00) |
0.0177 (1.00) |
0.000436 (0.768) |
0.0853 (1.00) |
1 (1.00) |
|
| APC | 160 (71%) | 64 |
0.0183 (1.00) |
0.00907 (1.00) |
0.221 (1.00) |
0.533 (1.00) |
0.374 (1.00) |
0.0164 (1.00) |
0.00922 (1.00) |
0.689 (1.00) |
0.464 (1.00) |
|
| SMAD4 | 26 (12%) | 198 |
0.023 (1.00) |
0.134 (1.00) |
0.0355 (1.00) |
0.388 (1.00) |
0.162 (1.00) |
0.00333 (1.00) |
0.00127 (1.00) |
1 (1.00) |
0.846 (1.00) |
|
| FAM123B | 25 (11%) | 199 |
0.0765 (1.00) |
0.0348 (1.00) |
0.00539 (1.00) |
0.141 (1.00) |
0.945 (1.00) |
0.0084 (1.00) |
0.00267 (1.00) |
1 (1.00) |
0.459 (1.00) |
|
| TTN | 85 (38%) | 139 |
0.00535 (1.00) |
0.105 (1.00) |
0.00133 (1.00) |
0.657 (1.00) |
0.0909 (1.00) |
0.849 (1.00) |
0.0392 (1.00) |
0.000551 (0.967) |
0.151 (1.00) |
0.927 (1.00) |
| SMAD2 | 15 (7%) | 209 |
0.114 (1.00) |
0.604 (1.00) |
0.0393 (1.00) |
0.286 (1.00) |
0.143 (1.00) |
0.265 (1.00) |
0.294 (1.00) |
1 (1.00) |
0.711 (1.00) |
|
| TCF7L2 | 18 (8%) | 206 |
1 (1.00) |
0.311 (1.00) |
0.417 (1.00) |
0.355 (1.00) |
0.372 (1.00) |
0.238 (1.00) |
0.792 (1.00) |
1 (1.00) |
0.23 (1.00) |
|
| KRTAP5-5 | 4 (2%) | 220 |
0.353 (1.00) |
1 (1.00) |
0.48 (1.00) |
0.172 (1.00) |
1 (1.00) |
0.579 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|
| ACVR1B | 14 (6%) | 210 |
0.109 (1.00) |
0.423 (1.00) |
0.0496 (1.00) |
0.463 (1.00) |
0.15 (1.00) |
0.0815 (1.00) |
0.0126 (1.00) |
0.435 (1.00) |
0.711 (1.00) |
|
| WBSCR17 | 19 (8%) | 205 |
0.0966 (1.00) |
0.31 (1.00) |
0.141 (1.00) |
0.262 (1.00) |
0.267 (1.00) |
0.654 (1.00) |
0.0146 (1.00) |
0.544 (1.00) |
0.618 (1.00) |
|
| PCDH9 | 22 (10%) | 202 |
0.727 (1.00) |
0.462 (1.00) |
0.0495 (1.00) |
0.875 (1.00) |
0.478 (1.00) |
0.489 (1.00) |
0.0447 (1.00) |
0.563 (1.00) |
1 (1.00) |
|
| LRP1B | 39 (17%) | 185 |
0.00288 (1.00) |
0.0211 (1.00) |
0.00332 (1.00) |
0.168 (1.00) |
1 (1.00) |
0.0113 (1.00) |
0.00162 (1.00) |
1 (1.00) |
0.087 (1.00) |
|
| TNFRSF10C | 6 (3%) | 218 |
0.376 (1.00) |
1 (1.00) |
0.675 (1.00) |
1 (1.00) |
0.236 (1.00) |
0.89 (1.00) |
0.622 (1.00) |
1 (1.00) |
1 (1.00) |
|
| MAP2K4 | 11 (5%) | 213 |
0.0756 (1.00) |
0.423 (1.00) |
0.123 (1.00) |
0.336 (1.00) |
1 (1.00) |
0.497 (1.00) |
0.134 (1.00) |
1 (1.00) |
0.545 (1.00) |
|
| KIAA1804 | 15 (7%) | 209 |
0.23 (1.00) |
0.411 (1.00) |
0.0659 (1.00) |
0.137 (1.00) |
0.614 (1.00) |
0.46 (1.00) |
0.575 (1.00) |
1 (1.00) |
0.736 (1.00) |
|
| NRXN1 | 20 (9%) | 204 |
0.432 (1.00) |
0.354 (1.00) |
0.0843 (1.00) |
0.876 (1.00) |
0.696 (1.00) |
0.193 (1.00) |
0.0376 (1.00) |
1 (1.00) |
0.13 (1.00) |
|
| KLHL4 | 16 (7%) | 208 |
0.0126 (1.00) |
0.0613 (1.00) |
0.00117 (1.00) |
0.387 (1.00) |
1 (1.00) |
0.0421 (1.00) |
0.0109 (1.00) |
0.41 (1.00) |
0.0126 (1.00) |
|
| SOX9 | 10 (4%) | 214 |
0.142 (1.00) |
0.352 (1.00) |
0.00451 (1.00) |
0.519 (1.00) |
1 (1.00) |
0.839 (1.00) |
0.659 (1.00) |
1 (1.00) |
0.584 (1.00) |
|
| PRDM9 | 16 (7%) | 208 |
0.00501 (1.00) |
1 (1.00) |
0.0701 (1.00) |
0.122 (1.00) |
0.86 (1.00) |
0.032 (1.00) |
0.22 (1.00) |
1 (1.00) |
0.255 (1.00) |
|
| PCBP1 | 6 (3%) | 218 |
0.175 (1.00) |
0.244 (1.00) |
0.396 (1.00) |
0.163 (1.00) |
0.203 (1.00) |
1 (1.00) |
1 (1.00) |
|||
| DMD | 33 (15%) | 191 |
0.0178 (1.00) |
0.0412 (1.00) |
0.0052 (1.00) |
0.624 (1.00) |
0.826 (1.00) |
0.0421 (1.00) |
0.00721 (1.00) |
1 (1.00) |
0.156 (1.00) |
|
| CSMD1 | 26 (12%) | 198 |
0.152 (1.00) |
0.122 (1.00) |
0.0523 (1.00) |
1 (1.00) |
0.137 (1.00) |
0.739 (1.00) |
0.0885 (1.00) |
0.0155 (1.00) |
0.0581 (1.00) |
0.142 (1.00) |
| LIFR | 18 (8%) | 206 |
0.116 (1.00) |
0.579 (1.00) |
0.219 (1.00) |
0.579 (1.00) |
0.676 (1.00) |
0.251 (1.00) |
0.00809 (1.00) |
1 (1.00) |
0.064 (1.00) |
|
| OR6N1 | 10 (4%) | 214 |
0.00197 (1.00) |
0.123 (1.00) |
0.013 (1.00) |
0.463 (1.00) |
1 (1.00) |
0.0167 (1.00) |
0.00422 (1.00) |
0.304 (1.00) |
1 (1.00) |
|
| CCDC160 | 7 (3%) | 217 |
0.0679 (1.00) |
0.742 (1.00) |
0.108 (1.00) |
0.215 (1.00) |
1 (1.00) |
0.258 (1.00) |
0.499 (1.00) |
1 (1.00) |
0.0575 (1.00) |
|
| CNTN6 | 17 (8%) | 207 |
0.153 (1.00) |
0.288 (1.00) |
0.0782 (1.00) |
0.266 (1.00) |
0.449 (1.00) |
0.333 (1.00) |
0.0051 (1.00) |
1 (1.00) |
0.322 (1.00) |
|
| CPXCR1 | 9 (4%) | 215 |
0.00223 (1.00) |
0.407 (1.00) |
0.191 (1.00) |
0.519 (1.00) |
1 (1.00) |
0.347 (1.00) |
0.0523 (1.00) |
0.304 (1.00) |
0.0285 (1.00) |
|
| SLITRK1 | 15 (7%) | 209 |
0.504 (1.00) |
0.881 (1.00) |
0.159 (1.00) |
0.206 (1.00) |
0.662 (1.00) |
0.963 (1.00) |
0.0496 (1.00) |
1 (1.00) |
0.181 (1.00) |
|
| CNBD1 | 7 (3%) | 217 |
0.767 (1.00) |
1 (1.00) |
0.293 (1.00) |
0.258 (1.00) |
0.0835 (1.00) |
1 (1.00) |
0.406 (1.00) |
|||
| CTNNB1 | 11 (5%) | 213 |
0.763 (1.00) |
0.832 (1.00) |
0.048 (1.00) |
0.648 (1.00) |
0.525 (1.00) |
0.332 (1.00) |
0.061 (1.00) |
1 (1.00) |
0.545 (1.00) |
|
| DKK2 | 6 (3%) | 218 |
0.0322 (1.00) |
0.0134 (1.00) |
0.628 (1.00) |
0.332 (1.00) |
0.871 (1.00) |
0.0371 (1.00) |
0.367 (1.00) |
1 (1.00) |
1 (1.00) |
|
| ERBB4 | 18 (8%) | 206 |
0.115 (1.00) |
0.67 (1.00) |
0.00631 (1.00) |
0.118 (1.00) |
1 (1.00) |
0.000461 (0.811) |
0.165 (1.00) |
0.481 (1.00) |
0.76 (1.00) |
|
| ARID1A | 21 (9%) | 203 |
0.0216 (1.00) |
0.0883 (1.00) |
0.0136 (1.00) |
0.699 (1.00) |
0.438 (1.00) |
0.0585 (1.00) |
0.0238 (1.00) |
1 (1.00) |
0.064 (1.00) |
|
| KIAA1486 | 13 (6%) | 211 |
0.337 (1.00) |
1 (1.00) |
0.306 (1.00) |
0.463 (1.00) |
1 (1.00) |
0.787 (1.00) |
0.481 (1.00) |
1 (1.00) |
0.0638 (1.00) |
|
| ROBO1 | 18 (8%) | 206 |
0.0046 (1.00) |
0.0617 (1.00) |
0.0221 (1.00) |
0.0201 (1.00) |
0.871 (1.00) |
0.0274 (1.00) |
0.00173 (1.00) |
0.502 (1.00) |
0.281 (1.00) |
|
| ZFHX4 | 24 (11%) | 200 |
0.703 (1.00) |
0.327 (1.00) |
0.0166 (1.00) |
0.699 (1.00) |
0.913 (1.00) |
0.594 (1.00) |
0.0452 (1.00) |
1 (1.00) |
0.072 (1.00) |
|
| EPHA3 | 16 (7%) | 208 |
0.000438 (0.771) |
0.0299 (1.00) |
0.214 (1.00) |
0.313 (1.00) |
0.905 (1.00) |
0.0513 (1.00) |
0.000308 (0.546) |
1 (1.00) |
0.23 (1.00) |
|
| PRIM2 | 8 (4%) | 216 |
0.285 (1.00) |
0.603 (1.00) |
0.889 (1.00) |
0.137 (1.00) |
0.636 (1.00) |
0.325 (1.00) |
0.611 (1.00) |
1 (1.00) |
0.406 (1.00) |
|
| UBE2NL | 7 (3%) | 217 |
0.102 (1.00) |
0.458 (1.00) |
0.402 (1.00) |
0.137 (1.00) |
1 (1.00) |
0.647 (1.00) |
0.15 (1.00) |
1 (1.00) |
0.351 (1.00) |
|
| ACVR2A | 9 (4%) | 215 |
0.00784 (1.00) |
0.452 (1.00) |
0.0325 (1.00) |
0.0487 (1.00) |
0.741 (1.00) |
0.0467 (1.00) |
0.0126 (1.00) |
1 (1.00) |
0.545 (1.00) |
|
| ADAM29 | 11 (5%) | 213 |
0.45 (1.00) |
1 (1.00) |
0.447 (1.00) |
0.898 (1.00) |
0.318 (1.00) |
1 (1.00) |
0.545 (1.00) |
|||
| ATP10A | 19 (8%) | 205 |
0.0169 (1.00) |
0.0882 (1.00) |
0.141 (1.00) |
0.000382 (0.675) |
0.365 (1.00) |
0.0319 (1.00) |
0.11 (1.00) |
1 (1.00) |
0.386 (1.00) |
|
| GABRA5 | 9 (4%) | 215 |
0.473 (1.00) |
1 (1.00) |
0.756 (1.00) |
0.546 (1.00) |
0.0637 (1.00) |
0.244 (1.00) |
0.456 (1.00) |
|||
| NALCN | 20 (9%) | 204 |
0.0365 (1.00) |
0.234 (1.00) |
0.141 (1.00) |
0.962 (1.00) |
0.757 (1.00) |
0.14 (1.00) |
0.0117 (1.00) |
1 (1.00) |
0.053 (1.00) |
|
| OTOL1 | 8 (4%) | 216 |
0.131 (1.00) |
1 (1.00) |
0.943 (1.00) |
0.623 (1.00) |
0.145 (1.00) |
0.274 (1.00) |
0.502 (1.00) |
|||
| PTEN | 7 (3%) | 217 |
0.0188 (1.00) |
0.78 (1.00) |
0.0429 (1.00) |
0.312 (1.00) |
0.636 (1.00) |
0.0334 (1.00) |
0.002 (1.00) |
0.244 (1.00) |
0.0575 (1.00) |
|
| TLL1 | 15 (7%) | 209 |
0.0352 (1.00) |
0.596 (1.00) |
0.02 (1.00) |
0.0853 (1.00) |
0.236 (1.00) |
0.262 (1.00) |
0.0603 (1.00) |
0.359 (1.00) |
0.00952 (1.00) |
|
| TRPC4 | 17 (8%) | 207 |
0.0046 (1.00) |
0.0827 (1.00) |
0.0139 (1.00) |
0.612 (1.00) |
0.162 (1.00) |
0.0231 (1.00) |
0.00421 (1.00) |
0.481 (1.00) |
0.171 (1.00) |
|
| ATM | 25 (11%) | 199 |
0.00284 (1.00) |
0.202 (1.00) |
0.000415 (0.731) |
0.325 (1.00) |
0.395 (1.00) |
0.015 (1.00) |
0.000574 (1.00) |
0.6 (1.00) |
0.162 (1.00) |
|
| GPC6 | 10 (4%) | 214 |
0.568 (1.00) |
0.636 (1.00) |
0.509 (1.00) |
0.735 (1.00) |
1 (1.00) |
0.455 (1.00) |
0.598 (1.00) |
1 (1.00) |
1 (1.00) |
|
| CDH8 | 14 (6%) | 210 |
0.00207 (1.00) |
0.00139 (1.00) |
0.124 (1.00) |
0.803 (1.00) |
0.478 (1.00) |
0.0504 (1.00) |
0.00103 (1.00) |
1 (1.00) |
1 (1.00) |
|
| MGC26647 | 7 (3%) | 217 |
0.275 (1.00) |
0.78 (1.00) |
0.0498 (1.00) |
0.579 (1.00) |
0.741 (1.00) |
0.574 (1.00) |
0.15 (1.00) |
1 (1.00) |
0.351 (1.00) |
|
| DKK4 | 7 (3%) | 217 |
0.297 (1.00) |
0.165 (1.00) |
0.402 (1.00) |
1 (1.00) |
1 (1.00) |
0.647 (1.00) |
0.575 (1.00) |
1 (1.00) |
1 (1.00) |
|
| PTPRM | 20 (9%) | 204 |
0.0446 (1.00) |
0.0145 (1.00) |
0.00293 (1.00) |
0.327 (1.00) |
0.568 (1.00) |
0.0772 (1.00) |
0.000589 (1.00) |
0.523 (1.00) |
0.0381 (1.00) |
|
| AFF2 | 17 (8%) | 207 |
0.0472 (1.00) |
0.0612 (1.00) |
0.201 (1.00) |
0.528 (1.00) |
0.119 (1.00) |
0.0331 (1.00) |
0.0497 (1.00) |
0.481 (1.00) |
0.255 (1.00) |
|
| NCAM2 | 12 (5%) | 212 |
0.0109 (1.00) |
0.451 (1.00) |
0.0617 (1.00) |
1 (1.00) |
1 (1.00) |
0.323 (1.00) |
0.00407 (1.00) |
1 (1.00) |
0.0387 (1.00) |
|
| OR2M4 | 8 (4%) | 216 |
0.00906 (1.00) |
0.267 (1.00) |
0.0382 (1.00) |
0.463 (1.00) |
0.741 (1.00) |
0.0152 (1.00) |
0.242 (1.00) |
1 (1.00) |
0.502 (1.00) |
|
| RYR2 | 29 (13%) | 195 |
0.0366 (1.00) |
0.248 (1.00) |
0.0195 (1.00) |
0.0123 (1.00) |
0.871 (1.00) |
0.0534 (1.00) |
0.00107 (1.00) |
0.6 (1.00) |
0.0755 (1.00) |
|
| CDH9 | 15 (7%) | 209 |
0.153 (1.00) |
0.288 (1.00) |
0.0226 (1.00) |
0.0231 (1.00) |
0.449 (1.00) |
0.0952 (1.00) |
0.0294 (1.00) |
1 (1.00) |
0.158 (1.00) |
|
| GRIK3 | 17 (8%) | 207 |
0.00763 (1.00) |
0.00174 (1.00) |
0.196 (1.00) |
0.163 (1.00) |
0.497 (1.00) |
0.148 (1.00) |
0.137 (1.00) |
1 (1.00) |
0.711 (1.00) |
|
| ELF3 | 6 (3%) | 218 |
0.311 (1.00) |
1 (1.00) |
0.0514 (1.00) |
0.668 (1.00) |
0.318 (1.00) |
0.0509 (1.00) |
0.0382 (1.00) |
1 (1.00) |
0.406 (1.00) |
|
| TRPS1 | 17 (8%) | 207 |
0.00936 (1.00) |
0.071 (1.00) |
0.00133 (1.00) |
0.0459 (1.00) |
1 (1.00) |
0.0101 (1.00) |
0.00057 (0.999) |
0.458 (1.00) |
0.354 (1.00) |
|
| FAM133A | 7 (3%) | 217 |
0.319 (1.00) |
0.78 (1.00) |
0.0183 (1.00) |
0.215 (1.00) |
0.636 (1.00) |
0.0783 (1.00) |
1 (1.00) |
1 (1.00) |
0.456 (1.00) |
|
| IL1RAPL2 | 10 (4%) | 214 |
0.334 (1.00) |
0.636 (1.00) |
0.0473 (1.00) |
0.387 (1.00) |
0.817 (1.00) |
0.255 (1.00) |
0.398 (1.00) |
0.0412 (1.00) |
1 (1.00) |
|
| CCBP2 | 10 (4%) | 214 |
0.149 (1.00) |
0.267 (1.00) |
0.623 (1.00) |
0.134 (1.00) |
0.905 (1.00) |
0.635 (1.00) |
0.398 (1.00) |
1 (1.00) |
0.162 (1.00) |
|
| OPCML | 10 (4%) | 214 |
0.321 (1.00) |
0.267 (1.00) |
0.543 (1.00) |
0.387 (1.00) |
0.236 (1.00) |
0.352 (1.00) |
0.543 (1.00) |
1 (1.00) |
0.0935 (1.00) |
|
| CDH18 | 13 (6%) | 211 |
0.00173 (1.00) |
0.273 (1.00) |
0.0984 (1.00) |
0.0515 (1.00) |
0.905 (1.00) |
0.0996 (1.00) |
0.0493 (1.00) |
1 (1.00) |
0.62 (1.00) |
|
| OR2L13 | 10 (4%) | 214 |
0.237 (1.00) |
0.599 (1.00) |
0.0275 (1.00) |
0.134 (1.00) |
0.739 (1.00) |
0.123 (1.00) |
0.221 (1.00) |
1 (1.00) |
1 (1.00) |
|
| BAI3 | 18 (8%) | 206 |
0.00719 (1.00) |
0.47 (1.00) |
0.423 (1.00) |
0.592 (1.00) |
1 (1.00) |
0.304 (1.00) |
0.207 (1.00) |
1 (1.00) |
0.255 (1.00) |
|
| MGC42105 | 11 (5%) | 213 |
0.00139 (1.00) |
0.172 (1.00) |
0.147 (1.00) |
0.0753 (1.00) |
0.119 (1.00) |
0.0167 (1.00) |
0.00422 (1.00) |
1 (1.00) |
1 (1.00) |
|
| TXNDC3 | 11 (5%) | 213 |
0.00269 (1.00) |
0.293 (1.00) |
0.0125 (1.00) |
1 (1.00) |
0.0299 (1.00) |
0.0918 (1.00) |
0.00398 (1.00) |
1 (1.00) |
0.00311 (1.00) |
|
| LPPR4 | 13 (6%) | 211 |
0.074 (1.00) |
0.0357 (1.00) |
0.0342 (1.00) |
0.799 (1.00) |
0.525 (1.00) |
0.223 (1.00) |
0.0738 (1.00) |
1 (1.00) |
0.653 (1.00) |
|
| UNC13C | 17 (8%) | 207 |
0.00713 (1.00) |
0.217 (1.00) |
0.0257 (1.00) |
0.0567 (1.00) |
0.86 (1.00) |
0.0612 (1.00) |
0.0827 (1.00) |
1 (1.00) |
0.255 (1.00) |
|
| SCN7A | 15 (7%) | 209 |
0.000649 (1.00) |
0.0609 (1.00) |
0.0492 (1.00) |
0.299 (1.00) |
0.913 (1.00) |
0.0111 (1.00) |
0.00632 (1.00) |
1 (1.00) |
0.205 (1.00) |
|
| ACOT4 | 3 (1%) | 221 |
0.296 (1.00) |
0.302 (1.00) |
0.829 (1.00) |
1 (1.00) |
0.0713 (1.00) |
1 (1.00) |
1 (1.00) |
|||
| GGT1 | 3 (1%) | 221 |
0.254 (1.00) |
0.184 (1.00) |
0.377 (1.00) |
1 (1.00) |
1 (1.00) |
|||||
| GRIA1 | 15 (7%) | 209 |
0.0535 (1.00) |
0.32 (1.00) |
0.0104 (1.00) |
0.558 (1.00) |
1 (1.00) |
0.0272 (1.00) |
0.00324 (1.00) |
1 (1.00) |
0.0226 (1.00) |
|
| ING1 | 6 (3%) | 218 |
0.291 (1.00) |
0.709 (1.00) |
0.628 (1.00) |
0.458 (1.00) |
0.203 (1.00) |
1 (1.00) |
0.406 (1.00) |
|||
| PCDHA10 | 10 (4%) | 214 |
0.079 (1.00) |
0.355 (1.00) |
0.108 (1.00) |
0.62 (1.00) |
0.0318 (1.00) |
1 (1.00) |
0.545 (1.00) |
|||
| ZNF208 | 11 (5%) | 213 |
0.182 (1.00) |
1 (1.00) |
0.249 (1.00) |
0.558 (1.00) |
1 (1.00) |
0.399 (1.00) |
0.628 (1.00) |
1 (1.00) |
0.114 (1.00) |
|
| CRTC1 | 6 (3%) | 218 |
0.122 (1.00) |
0.0889 (1.00) |
0.837 (1.00) |
0.312 (1.00) |
0.636 (1.00) |
0.228 (1.00) |
0.725 (1.00) |
1 (1.00) |
0.406 (1.00) |
|
| FAM5C | 13 (6%) | 211 |
0.0975 (1.00) |
0.601 (1.00) |
0.0186 (1.00) |
0.0231 (1.00) |
0.143 (1.00) |
0.466 (1.00) |
0.0353 (1.00) |
0.0711 (1.00) |
0.653 (1.00) |
|
| FAM22F | 9 (4%) | 215 |
0.69 (1.00) |
0.0958 (1.00) |
1 (1.00) |
0.915 (1.00) |
0.871 (1.00) |
0.749 (1.00) |
0.456 (1.00) |
1 (1.00) |
1 (1.00) |
|
| CHRDL1 | 11 (5%) | 213 |
0.166 (1.00) |
0.821 (1.00) |
0.472 (1.00) |
0.246 (1.00) |
1 (1.00) |
0.215 (1.00) |
0.916 (1.00) |
1 (1.00) |
0.584 (1.00) |
|
| EDNRB | 10 (4%) | 214 |
0.0629 (1.00) |
1 (1.00) |
0.757 (1.00) |
0.0128 (1.00) |
0.905 (1.00) |
0.839 (1.00) |
0.0982 (1.00) |
1 (1.00) |
0.545 (1.00) |
|
| IFT80 | 10 (4%) | 214 |
0.0879 (1.00) |
0.78 (1.00) |
0.254 (1.00) |
0.347 (1.00) |
0.0523 (1.00) |
0.304 (1.00) |
0.162 (1.00) |
|||
| ANK2 | 30 (13%) | 194 |
0.000689 (1.00) |
0.000889 (1.00) |
0.0015 (1.00) |
0.028 (1.00) |
0.614 (1.00) |
0.000963 (1.00) |
0.000734 (1.00) |
1 (1.00) |
0.0416 (1.00) |
|
| CASP14 | 8 (4%) | 216 |
1 (1.00) |
0.23 (1.00) |
0.889 (1.00) |
1 (1.00) |
1 (1.00) |
0.526 (1.00) |
0.464 (1.00) |
0.274 (1.00) |
1 (1.00) |
|
| LASS3 | 9 (4%) | 215 |
0.00355 (1.00) |
0.28 (1.00) |
0.104 (1.00) |
0.0946 (1.00) |
1 (1.00) |
0.0548 (1.00) |
0.0306 (1.00) |
1 (1.00) |
0.0747 (1.00) |
|
| P2RY10 | 7 (3%) | 217 |
0.66 (1.00) |
0.402 (1.00) |
0.402 (1.00) |
0.459 (1.00) |
1 (1.00) |
0.237 (1.00) |
0.499 (1.00) |
1 (1.00) |
1 (1.00) |
|
| CNTN1 | 14 (6%) | 210 |
0.0164 (1.00) |
0.193 (1.00) |
0.00405 (1.00) |
0.876 (1.00) |
0.696 (1.00) |
0.146 (1.00) |
0.00159 (1.00) |
1 (1.00) |
0.181 (1.00) |
|
| ISL1 | 8 (4%) | 216 |
0.302 (1.00) |
0.452 (1.00) |
0.435 (1.00) |
1 (1.00) |
1 (1.00) |
0.235 (1.00) |
0.242 (1.00) |
1 (1.00) |
0.502 (1.00) |
|
| ARID2 | 15 (7%) | 209 |
0.208 (1.00) |
0.221 (1.00) |
0.0254 (1.00) |
0.0459 (1.00) |
0.905 (1.00) |
0.231 (1.00) |
0.00632 (1.00) |
1 (1.00) |
0.0638 (1.00) |
|
| EIF4A2 | 5 (2%) | 219 |
0.647 (1.00) |
0.182 (1.00) |
0.73 (1.00) |
0.558 (1.00) |
0.636 (1.00) |
0.216 (1.00) |
0.288 (1.00) |
1 (1.00) |
0.141 (1.00) |
|
| CDH11 | 14 (6%) | 210 |
0.061 (1.00) |
0.273 (1.00) |
0.0175 (1.00) |
1 (1.00) |
1 (1.00) |
0.147 (1.00) |
0.0353 (1.00) |
0.385 (1.00) |
1 (1.00) |
|
| CHST9 | 8 (4%) | 216 |
0.0111 (1.00) |
0.8 (1.00) |
0.0706 (1.00) |
0.0144 (1.00) |
0.0988 (1.00) |
1 (1.00) |
0.456 (1.00) |
|||
| HTR3B | 6 (3%) | 218 |
0.13 (1.00) |
0.646 (1.00) |
0.628 (1.00) |
0.00853 (1.00) |
0.636 (1.00) |
0.587 (1.00) |
0.367 (1.00) |
1 (1.00) |
0.351 (1.00) |
|
| LRRIQ3 | 10 (4%) | 214 |
0.167 (1.00) |
0.697 (1.00) |
0.157 (1.00) |
0.232 (1.00) |
0.437 (1.00) |
0.467 (1.00) |
0.221 (1.00) |
1 (1.00) |
0.114 (1.00) |
|
| HCN1 | 10 (4%) | 214 |
0.344 (1.00) |
0.707 (1.00) |
0.0201 (1.00) |
0.735 (1.00) |
1 (1.00) |
0.24 (1.00) |
0.0423 (1.00) |
1 (1.00) |
1 (1.00) |
|
| CHRM2 | 10 (4%) | 214 |
0.338 (1.00) |
0.599 (1.00) |
0.115 (1.00) |
0.648 (1.00) |
1 (1.00) |
0.123 (1.00) |
0.00607 (1.00) |
1 (1.00) |
0.545 (1.00) |
|
| DACH2 | 12 (5%) | 212 |
0.123 (1.00) |
0.347 (1.00) |
0.256 (1.00) |
0.78 (1.00) |
0.0575 (1.00) |
1 (1.00) |
0.114 (1.00) |
|||
| PCDH11X | 14 (6%) | 210 |
0.0179 (1.00) |
0.521 (1.00) |
0.107 (1.00) |
0.681 (1.00) |
1 (1.00) |
0.0559 (1.00) |
0.289 (1.00) |
1 (1.00) |
0.181 (1.00) |
|
| KCNA4 | 12 (5%) | 212 |
0.101 (1.00) |
0.172 (1.00) |
0.00254 (1.00) |
0.0336 (1.00) |
0.528 (1.00) |
0.0486 (1.00) |
0.00407 (1.00) |
1 (1.00) |
0.584 (1.00) |
|
| DPP10 | 13 (6%) | 211 |
0.00128 (1.00) |
0.601 (1.00) |
0.0895 (1.00) |
0.699 (1.00) |
0.542 (1.00) |
0.276 (1.00) |
0.0493 (1.00) |
0.385 (1.00) |
0.158 (1.00) |
|
| EVC2 | 17 (8%) | 207 |
0.00338 (1.00) |
0.00644 (1.00) |
0.0383 (1.00) |
0.255 (1.00) |
0.144 (1.00) |
0.0125 (1.00) |
0.0597 (1.00) |
1 (1.00) |
0.711 (1.00) |
|
| C15ORF2 | 12 (5%) | 212 |
0.0174 (1.00) |
0.452 (1.00) |
0.166 (1.00) |
0.232 (1.00) |
0.152 (1.00) |
0.131 (1.00) |
0.00398 (1.00) |
0.304 (1.00) |
0.0492 (1.00) |
|
| NLRP5 | 12 (5%) | 212 |
0.159 (1.00) |
1 (1.00) |
0.0649 (1.00) |
0.025 (1.00) |
0.15 (1.00) |
0.00618 (1.00) |
0.0297 (1.00) |
1 (1.00) |
0.584 (1.00) |
|
| BTNL8 | 3 (1%) | 221 |
0.296 (1.00) |
0.302 (1.00) |
0.254 (1.00) |
0.215 (1.00) |
1 (1.00) |
0.184 (1.00) |
0.227 (1.00) |
1 (1.00) |
1 (1.00) |
|
| KLK2 | 3 (1%) | 221 |
0.468 (1.00) |
0.302 (1.00) |
0.829 (1.00) |
0.327 (1.00) |
0.227 (1.00) |
1 (1.00) |
||||
| LUZP2 | 6 (3%) | 218 |
0.0835 (1.00) |
0.202 (1.00) |
0.175 (1.00) |
0.126 (1.00) |
0.418 (1.00) |
0.182 (1.00) |
0.85 (1.00) |
1 (1.00) |
1 (1.00) |
|
| NXF5 | 7 (3%) | 217 |
0.243 (1.00) |
0.863 (1.00) |
0.425 (1.00) |
0.215 (1.00) |
1 (1.00) |
0.732 (1.00) |
0.575 (1.00) |
1 (1.00) |
0.0421 (1.00) |
|
| PCDHB5 | 14 (6%) | 210 |
0.148 (1.00) |
0.573 (1.00) |
0.0582 (1.00) |
0.492 (1.00) |
0.227 (1.00) |
0.211 (1.00) |
0.0185 (1.00) |
0.385 (1.00) |
0.653 (1.00) |
|
| LOXL2 | 9 (4%) | 215 |
0.653 (1.00) |
0.707 (1.00) |
0.191 (1.00) |
0.0316 (1.00) |
0.328 (1.00) |
0.0156 (1.00) |
0.363 (1.00) |
1 (1.00) |
0.32 (1.00) |
|
| UBQLNL | 5 (2%) | 219 |
0.12 (1.00) |
0.173 (1.00) |
1 (1.00) |
0.133 (1.00) |
0.164 (1.00) |
1 (1.00) |
1 (1.00) |
|||
| MYO1B | 13 (6%) | 211 |
0.00916 (1.00) |
0.0199 (1.00) |
0.00752 (1.00) |
0.0284 (1.00) |
1 (1.00) |
0.0119 (1.00) |
0.0108 (1.00) |
1 (1.00) |
0.653 (1.00) |
|
| SPATA17 | 9 (4%) | 215 |
0.102 (1.00) |
0.486 (1.00) |
0.0519 (1.00) |
1 (1.00) |
0.817 (1.00) |
0.288 (1.00) |
0.0637 (1.00) |
1 (1.00) |
1 (1.00) |
|
| COL12A1 | 21 (9%) | 203 |
0.01 (1.00) |
0.505 (1.00) |
0.0451 (1.00) |
0.148 (1.00) |
1 (1.00) |
0.0684 (1.00) |
0.0351 (1.00) |
0.523 (1.00) |
0.0753 (1.00) |
|
| JAKMIP2 | 13 (6%) | 211 |
0.0146 (1.00) |
0.273 (1.00) |
0.0855 (1.00) |
0.283 (1.00) |
0.173 (1.00) |
0.0258 (1.00) |
0.0261 (1.00) |
0.385 (1.00) |
0.158 (1.00) |
|
| EPHA5 | 13 (6%) | 211 |
0.0611 (1.00) |
0.117 (1.00) |
0.0666 (1.00) |
0.283 (1.00) |
0.452 (1.00) |
0.0612 (1.00) |
0.00538 (1.00) |
1 (1.00) |
0.653 (1.00) |
|
| ODZ1 | 22 (10%) | 202 |
0.376 (1.00) |
0.312 (1.00) |
0.0474 (1.00) |
0.918 (1.00) |
0.413 (1.00) |
0.00842 (1.00) |
0.000214 (0.38) |
0.204 (1.00) |
0.669 (1.00) |
|
| POSTN | 11 (5%) | 213 |
0.0193 (1.00) |
0.108 (1.00) |
0.272 (1.00) |
0.681 (1.00) |
0.636 (1.00) |
0.407 (1.00) |
0.318 (1.00) |
1 (1.00) |
1 (1.00) |
|
| ZCWPW2 | 7 (3%) | 217 |
0.344 (1.00) |
0.517 (1.00) |
1 (1.00) |
0.903 (1.00) |
0.437 (1.00) |
0.844 (1.00) |
0.367 (1.00) |
1 (1.00) |
0.104 (1.00) |
|
| ZEB2 | 15 (7%) | 209 |
0.0402 (1.00) |
0.761 (1.00) |
0.164 (1.00) |
0.559 (1.00) |
1 (1.00) |
0.131 (1.00) |
0.0135 (1.00) |
0.385 (1.00) |
0.653 (1.00) |
|
| TUSC3 | 7 (3%) | 217 |
0.00548 (1.00) |
0.136 (1.00) |
0.04 (1.00) |
0.558 (1.00) |
1 (1.00) |
0.0194 (1.00) |
0.000529 (0.929) |
1 (1.00) |
0.00375 (1.00) |
|
| CACNG7 | 6 (3%) | 218 |
0.596 (1.00) |
0.322 (1.00) |
0.363 (1.00) |
0.126 (1.00) |
0.0541 (1.00) |
0.0605 (1.00) |
0.622 (1.00) |
1 (1.00) |
0.213 (1.00) |
|
| RWDD2B | 8 (4%) | 216 |
0.0518 (1.00) |
0.8 (1.00) |
0.182 (1.00) |
0.494 (1.00) |
0.242 (1.00) |
1 (1.00) |
0.00375 (1.00) |
|||
| SLC25A13 | 7 (3%) | 217 |
0.14 (1.00) |
0.349 (1.00) |
0.425 (1.00) |
0.232 (1.00) |
0.236 (1.00) |
0.381 (1.00) |
0.363 (1.00) |
1 (1.00) |
0.0421 (1.00) |
|
| DPF2 | 4 (2%) | 220 |
0.468 (1.00) |
0.302 (1.00) |
0.381 (1.00) |
0.215 (1.00) |
0.0777 (1.00) |
0.327 (1.00) |
1 (1.00) |
1 (1.00) |
0.141 (1.00) |
|
| PCDH10 | 17 (8%) | 207 |
0.0868 (1.00) |
0.113 (1.00) |
0.0139 (1.00) |
0.336 (1.00) |
0.696 (1.00) |
0.106 (1.00) |
0.000559 (0.98) |
1 (1.00) |
0.23 (1.00) |
|
| CDH12 | 13 (6%) | 211 |
0.00479 (1.00) |
0.32 (1.00) |
0.0874 (1.00) |
0.0946 (1.00) |
0.236 (1.00) |
0.0186 (1.00) |
0.00927 (1.00) |
1 (1.00) |
0.00946 (1.00) |
|
| MCF2 | 12 (5%) | 212 |
0.0808 (1.00) |
0.273 (1.00) |
0.263 (1.00) |
0.281 (1.00) |
0.817 (1.00) |
0.418 (1.00) |
0.121 (1.00) |
1 (1.00) |
0.135 (1.00) |
|
| NCKAP5 | 15 (7%) | 209 |
0.0233 (1.00) |
0.0161 (1.00) |
0.131 (1.00) |
0.00216 (1.00) |
0.905 (1.00) |
0.0606 (1.00) |
0.000936 (1.00) |
1 (1.00) |
0.711 (1.00) |
|
| RIMS2 | 13 (6%) | 211 |
0.00174 (1.00) |
0.32 (1.00) |
0.000306 (0.543) |
0.273 (1.00) |
1 (1.00) |
0.00301 (1.00) |
0.00538 (1.00) |
1 (1.00) |
0.653 (1.00) |
|
| ATP7A | 6 (3%) | 218 |
0.703 (1.00) |
1 (1.00) |
0.917 (1.00) |
0.205 (1.00) |
0.418 (1.00) |
0.844 (1.00) |
0.85 (1.00) |
1 (1.00) |
1 (1.00) |
|
| TPTE | 12 (5%) | 212 |
0.109 (1.00) |
0.461 (1.00) |
0.000369 (0.653) |
0.519 (1.00) |
1 (1.00) |
0.000818 (1.00) |
0.000394 (0.695) |
1 (1.00) |
0.62 (1.00) |
|
| OR8B4 | 6 (3%) | 218 |
0.0679 (1.00) |
0.407 (1.00) |
0.206 (1.00) |
0.458 (1.00) |
0.203 (1.00) |
1 (1.00) |
0.351 (1.00) |
|||
| SFMBT2 | 13 (6%) | 211 |
0.541 (1.00) |
0.273 (1.00) |
0.387 (1.00) |
0.767 (1.00) |
1 (1.00) |
0.264 (1.00) |
0.00927 (1.00) |
1 (1.00) |
0.683 (1.00) |
|
| FLRT2 | 13 (6%) | 211 |
0.138 (1.00) |
1 (1.00) |
0.019 (1.00) |
0.0552 (1.00) |
1 (1.00) |
0.369 (1.00) |
0.481 (1.00) |
1 (1.00) |
0.135 (1.00) |
|
| BCHE | 9 (4%) | 215 |
0.0539 (1.00) |
0.27 (1.00) |
0.314 (1.00) |
1 (1.00) |
1 (1.00) |
0.727 (1.00) |
0.0126 (1.00) |
1 (1.00) |
1 (1.00) |
|
| C8B | 10 (4%) | 214 |
0.0767 (1.00) |
0.27 (1.00) |
0.757 (1.00) |
0.668 (1.00) |
1 (1.00) |
0.635 (1.00) |
0.221 (1.00) |
1 (1.00) |
1 (1.00) |
|
| FREM2 | 23 (10%) | 201 |
0.217 (1.00) |
0.381 (1.00) |
0.299 (1.00) |
0.105 (1.00) |
0.542 (1.00) |
0.384 (1.00) |
0.716 (1.00) |
0.6 (1.00) |
0.409 (1.00) |
|
| ZNF167 | 7 (3%) | 217 |
0.275 (1.00) |
1 (1.00) |
0.0498 (1.00) |
0.825 (1.00) |
1 (1.00) |
0.237 (1.00) |
0.0163 (1.00) |
1 (1.00) |
0.406 (1.00) |
|
| GABRG1 | 8 (4%) | 216 |
0.561 (1.00) |
0.707 (1.00) |
0.459 (1.00) |
0.474 (1.00) |
0.817 (1.00) |
0.126 (1.00) |
0.242 (1.00) |
1 (1.00) |
0.0747 (1.00) |
|
| SAT1 | 4 (2%) | 220 |
0.00272 (1.00) |
0.0515 (1.00) |
0.0742 (1.00) |
0.825 (1.00) |
0.193 (1.00) |
0.0024 (1.00) |
0.0734 (1.00) |
1 (1.00) |
1 (1.00) |
|
| CHD4 | 17 (8%) | 207 |
0.00951 (1.00) |
0.171 (1.00) |
0.000862 (1.00) |
0.492 (1.00) |
0.913 (1.00) |
0.000324 (0.574) |
0.000812 (1.00) |
1 (1.00) |
0.0101 (1.00) |
|
| PCDHB2 | 14 (6%) | 210 |
0.0102 (1.00) |
0.0106 (1.00) |
0.0175 (1.00) |
0.648 (1.00) |
1 (1.00) |
0.0218 (1.00) |
0.00324 (1.00) |
1 (1.00) |
0.653 (1.00) |
|
| TMTC1 | 13 (6%) | 211 |
0.169 (1.00) |
0.273 (1.00) |
0.0147 (1.00) |
0.0504 (1.00) |
0.328 (1.00) |
0.0836 (1.00) |
0.00538 (1.00) |
0.385 (1.00) |
0.158 (1.00) |
|
| SH3GL3 | 8 (4%) | 216 |
0.102 (1.00) |
1 (1.00) |
0.596 (1.00) |
0.463 (1.00) |
1 (1.00) |
0.742 (1.00) |
0.464 (1.00) |
1 (1.00) |
0.133 (1.00) |
|
| ZFPM2 | 6 (3%) | 218 |
0.0235 (1.00) |
0.646 (1.00) |
0.541 (1.00) |
0.0487 (1.00) |
0.318 (1.00) |
0.338 (1.00) |
0.164 (1.00) |
1 (1.00) |
0.0549 (1.00) |
|
| CACNG3 | 8 (4%) | 216 |
0.0174 (1.00) |
0.27 (1.00) |
0.00923 (1.00) |
0.025 (1.00) |
0.318 (1.00) |
0.126 (1.00) |
0.0988 (1.00) |
0.274 (1.00) |
0.0747 (1.00) |
|
| POTEB | 8 (4%) | 216 |
0.307 (1.00) |
0.517 (1.00) |
0.287 (1.00) |
0.0388 (1.00) |
0.0125 (1.00) |
1 (1.00) |
1 (1.00) |
|||
| GRIN2A | 19 (8%) | 205 |
0.392 (1.00) |
0.369 (1.00) |
0.0439 (1.00) |
0.592 (1.00) |
0.86 (1.00) |
0.357 (1.00) |
0.0648 (1.00) |
0.481 (1.00) |
0.76 (1.00) |
|
| NRXN3 | 17 (8%) | 207 |
0.000398 (0.701) |
0.0827 (1.00) |
0.00293 (1.00) |
0.916 (1.00) |
0.179 (1.00) |
0.00121 (1.00) |
0.00057 (0.999) |
1 (1.00) |
0.736 (1.00) |
|
| OR7C1 | 6 (3%) | 218 |
0.13 (1.00) |
0.0134 (1.00) |
0.541 (1.00) |
0.128 (1.00) |
0.0683 (1.00) |
1 (1.00) |
0.406 (1.00) |
|||
| SPATA8 | 3 (1%) | 221 |
0.312 (1.00) |
0.439 (1.00) |
0.751 (1.00) |
1 (1.00) |
0.104 (1.00) |
|||||
| OR2M2 | 6 (3%) | 218 |
0.809 (1.00) |
0.562 (1.00) |
0.917 (1.00) |
0.528 (1.00) |
0.203 (1.00) |
1 (1.00) |
0.351 (1.00) |
|||
| SORCS1 | 15 (7%) | 209 |
0.236 (1.00) |
0.0518 (1.00) |
0.131 (1.00) |
0.936 (1.00) |
0.605 (1.00) |
0.149 (1.00) |
0.00632 (1.00) |
1 (1.00) |
0.205 (1.00) |
|
| IL18R1 | 10 (4%) | 214 |
0.0354 (1.00) |
0.486 (1.00) |
0.254 (1.00) |
0.463 (1.00) |
1 (1.00) |
0.186 (1.00) |
0.0724 (1.00) |
1 (1.00) |
0.114 (1.00) |
|
| DPY19L1 | 5 (2%) | 219 |
0.00985 (1.00) |
0.34 (1.00) |
0.194 (1.00) |
0.137 (1.00) |
1 (1.00) |
0.0911 (1.00) |
0.000861 (1.00) |
1 (1.00) |
0.292 (1.00) |
|
| C16ORF45 | 3 (1%) | 221 |
0.137 (1.00) |
0.594 (1.00) |
0.108 (1.00) |
0.145 (1.00) |
0.227 (1.00) |
1 (1.00) |
1 (1.00) |
|||
| INHBA | 9 (4%) | 215 |
0.00486 (1.00) |
0.27 (1.00) |
0.0706 (1.00) |
0.0318 (1.00) |
0.00354 (1.00) |
0.025 (1.00) |
0.0575 (1.00) |
|||
| MIER3 | 11 (5%) | 213 |
0.000174 (0.309) |
0.0182 (1.00) |
0.00207 (1.00) |
0.799 (1.00) |
1 (1.00) |
0.00185 (1.00) |
0.00057 (0.999) |
1 (1.00) |
0.62 (1.00) |
|
| ZNF585A | 10 (4%) | 214 |
0.00256 (1.00) |
0.293 (1.00) |
0.00451 (1.00) |
0.463 (1.00) |
0.0666 (1.00) |
0.0025 (1.00) |
0.000975 (1.00) |
1 (1.00) |
0.0935 (1.00) |
|
| TMPRSS13 | 4 (2%) | 220 |
0.386 (1.00) |
1 (1.00) |
0.11 (1.00) |
0.21 (1.00) |
0.807 (1.00) |
1 (1.00) |
0.292 (1.00) |
|||
| FSTL5 | 11 (5%) | 213 |
0.0556 (1.00) |
0.172 (1.00) |
0.0316 (1.00) |
0.355 (1.00) |
0.542 (1.00) |
0.0399 (1.00) |
0.034 (1.00) |
1 (1.00) |
0.193 (1.00) |
|
| DOK5 | 6 (3%) | 218 |
0.893 (1.00) |
0.481 (1.00) |
0.628 (1.00) |
0.459 (1.00) |
1 (1.00) |
0.0605 (1.00) |
0.367 (1.00) |
1 (1.00) |
0.213 (1.00) |
|
| ZNF99 | 11 (5%) | 213 |
0.0467 (1.00) |
0.697 (1.00) |
0.0153 (1.00) |
0.137 (1.00) |
0.636 (1.00) |
0.00669 (1.00) |
0.00936 (1.00) |
1 (1.00) |
0.0387 (1.00) |
|
| GML | 6 (3%) | 218 |
0.412 (1.00) |
0.0684 (1.00) |
0.628 (1.00) |
0.458 (1.00) |
0.00787 (1.00) |
1 (1.00) |
1 (1.00) |
|||
| B2M | 5 (2%) | 219 |
0.0413 (1.00) |
0.661 (1.00) |
0.0452 (1.00) |
0.215 (1.00) |
1 (1.00) |
0.0911 (1.00) |
0.0143 (1.00) |
1 (1.00) |
0.0288 (1.00) |
|
| PPP2R2B | 3 (1%) | 221 |
0.468 (1.00) |
1 (1.00) |
0.445 (1.00) |
1 (1.00) |
0.377 (1.00) |
1 (1.00) |
0.228 (1.00) |
|||
| LRTM1 | 7 (3%) | 217 |
0.0879 (1.00) |
0.458 (1.00) |
0.402 (1.00) |
0.647 (1.00) |
0.0835 (1.00) |
1 (1.00) |
0.351 (1.00) |
|||
| OR10G9 | 6 (3%) | 218 |
0.238 (1.00) |
0.709 (1.00) |
0.206 (1.00) |
0.458 (1.00) |
0.0365 (1.00) |
1 (1.00) |
0.351 (1.00) |
|||
| ERBB3 | 14 (6%) | 210 |
0.14 (1.00) |
0.0872 (1.00) |
0.124 (1.00) |
0.947 (1.00) |
0.739 (1.00) |
0.167 (1.00) |
0.0185 (1.00) |
1 (1.00) |
0.158 (1.00) |
|
| PSG8 | 8 (4%) | 216 |
0.893 (1.00) |
0.0889 (1.00) |
0.107 (1.00) |
0.437 (1.00) |
0.411 (1.00) |
0.244 (1.00) |
0.249 (1.00) |
|||
| SYT16 | 9 (4%) | 215 |
0.071 (1.00) |
0.209 (1.00) |
0.182 (1.00) |
0.0336 (1.00) |
0.528 (1.00) |
0.115 (1.00) |
0.219 (1.00) |
0.304 (1.00) |
0.545 (1.00) |
|
| LPHN3 | 13 (6%) | 211 |
0.00292 (1.00) |
0.252 (1.00) |
0.387 (1.00) |
0.148 (1.00) |
1 (1.00) |
0.327 (1.00) |
0.00927 (1.00) |
1 (1.00) |
0.135 (1.00) |
|
| LRP2 | 35 (16%) | 189 |
0.00181 (1.00) |
0.0383 (1.00) |
0.0014 (1.00) |
0.523 (1.00) |
0.826 (1.00) |
0.0874 (1.00) |
0.0023 (1.00) |
0.342 (1.00) |
0.277 (1.00) |
|
| SOHLH2 | 9 (4%) | 215 |
0.859 (1.00) |
0.0418 (1.00) |
0.327 (1.00) |
0.0219 (1.00) |
0.297 (1.00) |
0.0708 (1.00) |
0.0908 (1.00) |
1 (1.00) |
0.502 (1.00) |
|
| ZIM3 | 9 (4%) | 215 |
0.0606 (1.00) |
0.9 (1.00) |
0.119 (1.00) |
0.00743 (1.00) |
0.817 (1.00) |
0.206 (1.00) |
0.57 (1.00) |
1 (1.00) |
0.0285 (1.00) |
|
| DSEL | 14 (6%) | 210 |
0.0128 (1.00) |
0.193 (1.00) |
0.00117 (1.00) |
0.336 (1.00) |
1 (1.00) |
0.0815 (1.00) |
0.0185 (1.00) |
1 (1.00) |
0.158 (1.00) |
|
| RNASE11 | 5 (2%) | 219 |
0.767 (1.00) |
0.709 (1.00) |
0.148 (1.00) |
0.81 (1.00) |
0.164 (1.00) |
1 (1.00) |
0.0177 (1.00) |
|||
| MDGA2 | 10 (4%) | 214 |
0.102 (1.00) |
0.124 (1.00) |
0.0201 (1.00) |
0.735 (1.00) |
0.817 (1.00) |
0.058 (1.00) |
0.00422 (1.00) |
0.304 (1.00) |
1 (1.00) |
|
| RBMS3 | 6 (3%) | 218 |
0.291 (1.00) |
0.842 (1.00) |
1 (1.00) |
0.844 (1.00) |
0.367 (1.00) |
1 (1.00) |
0.406 (1.00) |
|||
| COL6A6 | 16 (7%) | 208 |
0.0547 (1.00) |
0.573 (1.00) |
0.00426 (1.00) |
0.484 (1.00) |
0.452 (1.00) |
0.0375 (1.00) |
0.0294 (1.00) |
0.458 (1.00) |
0.736 (1.00) |
|
| TMBIM4 | 4 (2%) | 220 |
1 (1.00) |
0.322 (1.00) |
0.65 (1.00) |
0.528 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|||
| FAM135B | 15 (7%) | 209 |
0.0516 (1.00) |
0.273 (1.00) |
0.116 (1.00) |
0.772 (1.00) |
0.696 (1.00) |
0.00639 (1.00) |
0.00324 (1.00) |
0.107 (1.00) |
0.354 (1.00) |
|
| CYP7B1 | 7 (3%) | 217 |
0.00985 (1.00) |
0.0462 (1.00) |
0.108 (1.00) |
0.00513 (1.00) |
1 (1.00) |
0.105 (1.00) |
0.00142 (1.00) |
1 (1.00) |
0.351 (1.00) |
|
| SLC16A7 | 9 (4%) | 215 |
0.0275 (1.00) |
0.486 (1.00) |
0.119 (1.00) |
1 (1.00) |
0.297 (1.00) |
0.347 (1.00) |
0.0908 (1.00) |
1 (1.00) |
0.0747 (1.00) |
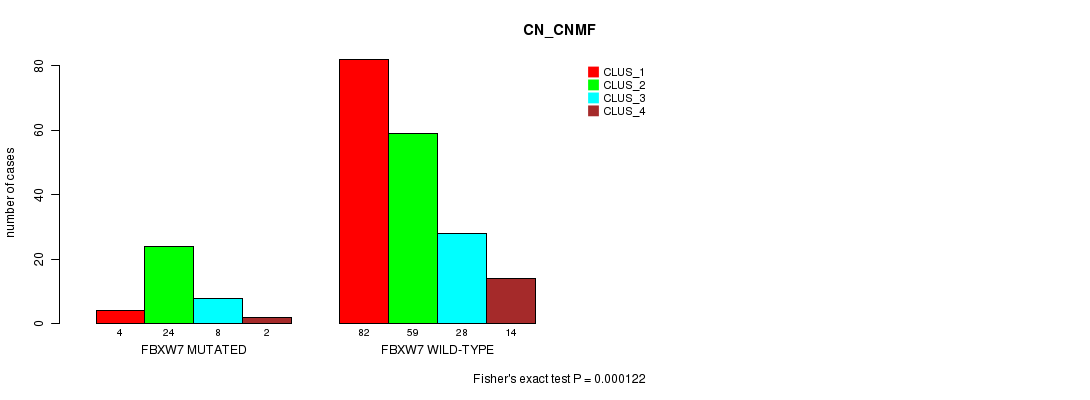
P value = 0.000122 (Fisher's exact test), Q value = 0.22
Table S1. Gene #1: 'FBXW7 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 86 | 83 | 36 | 16 |
| FBXW7 MUTATED | 4 | 24 | 8 | 2 |
| FBXW7 WILD-TYPE | 82 | 59 | 28 | 14 |
Figure S1. Get High-res Image Gene #1: 'FBXW7 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
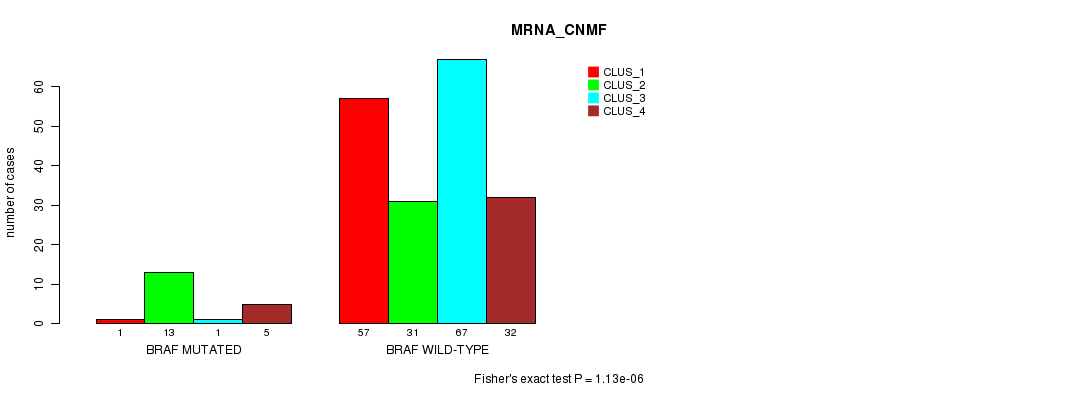
P value = 1.13e-06 (Fisher's exact test), Q value = 0.002
Table S2. Gene #5: 'BRAF MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 44 | 68 | 37 |
| BRAF MUTATED | 1 | 13 | 1 | 5 |
| BRAF WILD-TYPE | 57 | 31 | 67 | 32 |
Figure S2. Get High-res Image Gene #5: 'BRAF MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
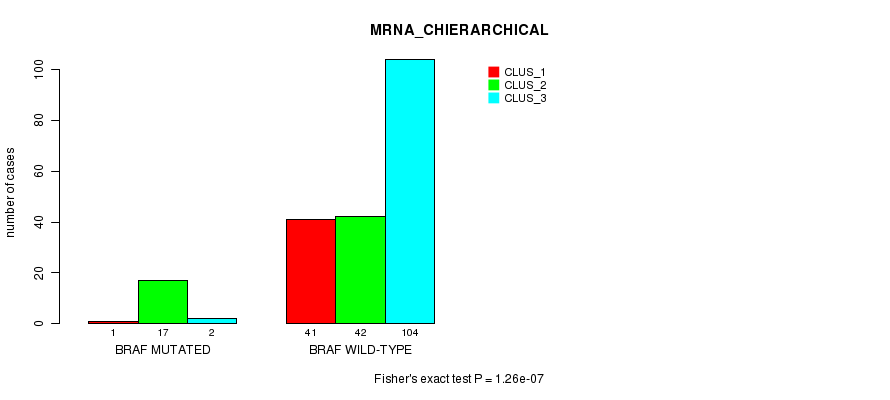
P value = 1.26e-07 (Fisher's exact test), Q value = 0.00023
Table S3. Gene #5: 'BRAF MUTATION STATUS' versus Clinical Feature #2: 'MRNA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 42 | 59 | 106 |
| BRAF MUTATED | 1 | 17 | 2 |
| BRAF WILD-TYPE | 41 | 42 | 104 |
Figure S3. Get High-res Image Gene #5: 'BRAF MUTATION STATUS' versus Clinical Feature #2: 'MRNA_CHIERARCHICAL'
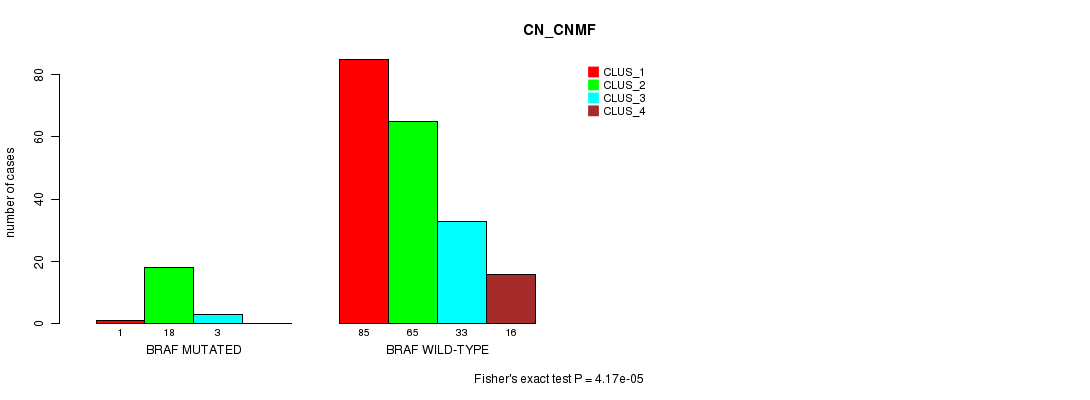
P value = 4.17e-05 (Fisher's exact test), Q value = 0.075
Table S4. Gene #5: 'BRAF MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 86 | 83 | 36 | 16 |
| BRAF MUTATED | 1 | 18 | 3 | 0 |
| BRAF WILD-TYPE | 85 | 65 | 33 | 16 |
Figure S4. Get High-res Image Gene #5: 'BRAF MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
P value = 3.49e-08 (Fisher's exact test), Q value = 6.3e-05
Table S5. Gene #5: 'BRAF MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 78 | 47 | 60 | 33 |
| BRAF MUTATED | 1 | 16 | 5 | 0 |
| BRAF WILD-TYPE | 77 | 31 | 55 | 33 |
Figure S5. Get High-res Image Gene #5: 'BRAF MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
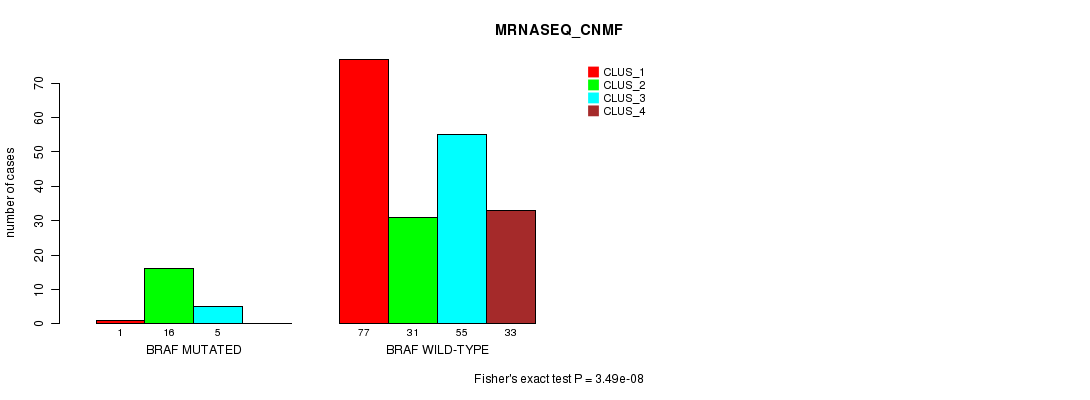
P value = 1.92e-10 (Fisher's exact test), Q value = 3.5e-07
Table S6. Gene #5: 'BRAF MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| BRAF MUTATED | 19 | 2 | 1 |
| BRAF WILD-TYPE | 34 | 125 | 37 |
Figure S6. Get High-res Image Gene #5: 'BRAF MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
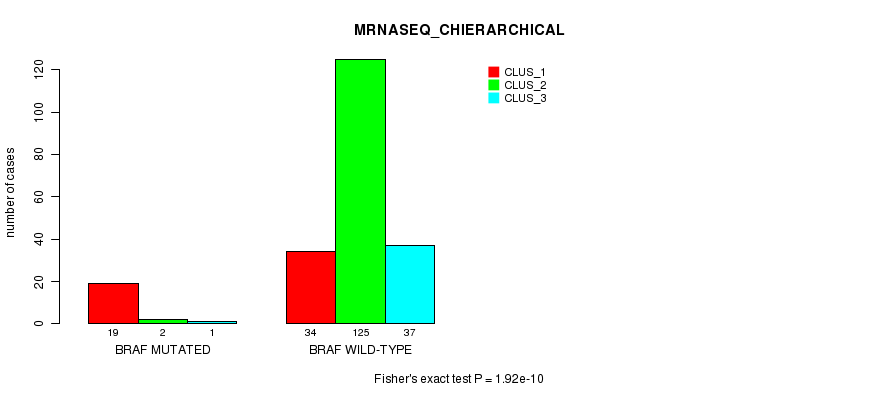
P value = 1.94e-07 (Fisher's exact test), Q value = 0.00035
Table S7. Gene #6: 'KRAS MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 78 | 47 | 60 | 33 |
| KRAS MUTATED | 21 | 15 | 44 | 12 |
| KRAS WILD-TYPE | 57 | 32 | 16 | 21 |
Figure S7. Get High-res Image Gene #6: 'KRAS MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
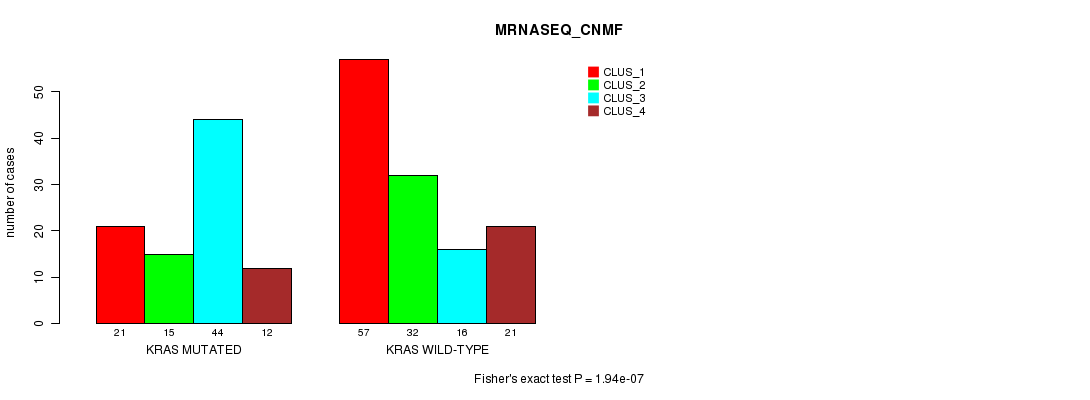
P value = 3.02e-07 (Fisher's exact test), Q value = 0.00055
Table S8. Gene #6: 'KRAS MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| KRAS MUTATED | 20 | 41 | 31 |
| KRAS WILD-TYPE | 33 | 86 | 7 |
Figure S8. Get High-res Image Gene #6: 'KRAS MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
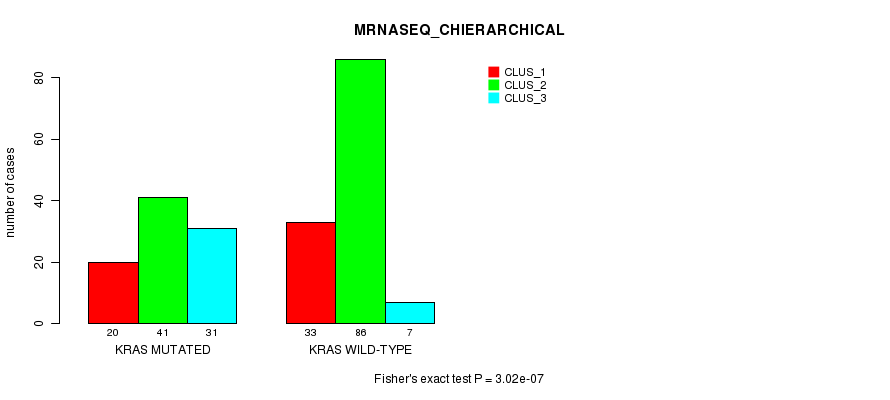
P value = 7.28e-11 (Fisher's exact test), Q value = 1.3e-07
Table S9. Gene #8: 'TP53 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 86 | 83 | 36 | 16 |
| TP53 MUTATED | 68 | 22 | 19 | 9 |
| TP53 WILD-TYPE | 18 | 61 | 17 | 7 |
Figure S9. Get High-res Image Gene #8: 'TP53 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
P value = 6.63e-05 (Fisher's exact test), Q value = 0.12
Table S10. Gene #13: 'FAT4 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 86 | 83 | 36 | 16 |
| FAT4 MUTATED | 6 | 27 | 3 | 2 |
| FAT4 WILD-TYPE | 80 | 56 | 33 | 14 |
Figure S10. Get High-res Image Gene #13: 'FAT4 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
P value = 3.19e-05 (Fisher's exact test), Q value = 0.057
Table S11. Gene #35: 'CDH2 MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 44 | 68 | 37 |
| CDH2 MUTATED | 0 | 7 | 1 | 7 |
| CDH2 WILD-TYPE | 58 | 37 | 67 | 30 |
Figure S11. Get High-res Image Gene #35: 'CDH2 MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
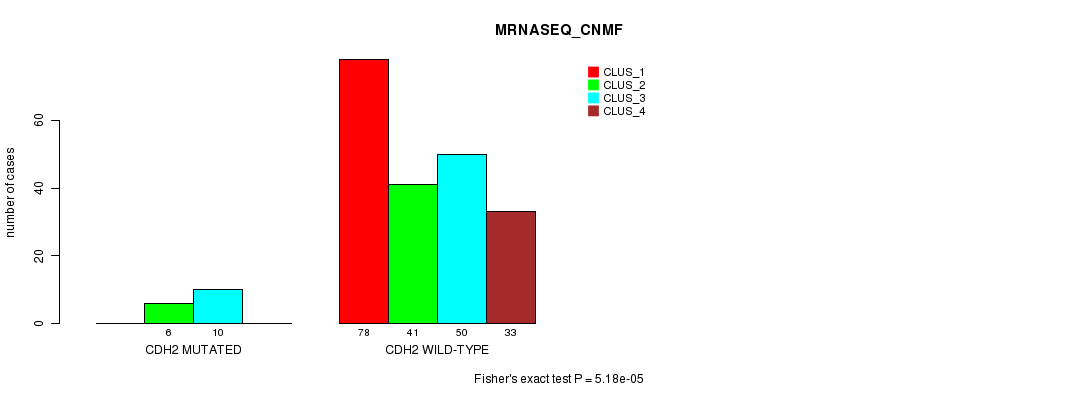
P value = 5.18e-05 (Fisher's exact test), Q value = 0.093
Table S12. Gene #35: 'CDH2 MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 78 | 47 | 60 | 33 |
| CDH2 MUTATED | 0 | 6 | 10 | 0 |
| CDH2 WILD-TYPE | 78 | 41 | 50 | 33 |
Figure S12. Get High-res Image Gene #35: 'CDH2 MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
P value = 1.1e-05 (Fisher's exact test), Q value = 0.02
Table S13. Gene #35: 'CDH2 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| CDH2 MUTATED | 10 | 1 | 5 |
| CDH2 WILD-TYPE | 43 | 126 | 33 |
Figure S13. Get High-res Image Gene #35: 'CDH2 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
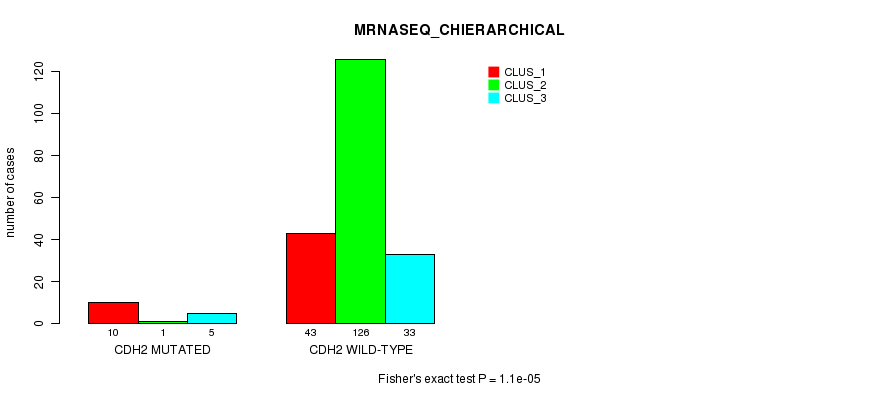
P value = 2.36e-06 (Fisher's exact test), Q value = 0.0043
Table S14. Gene #40: 'FLG MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 44 | 68 | 37 |
| FLG MUTATED | 0 | 14 | 6 | 8 |
| FLG WILD-TYPE | 58 | 30 | 62 | 29 |
Figure S14. Get High-res Image Gene #40: 'FLG MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
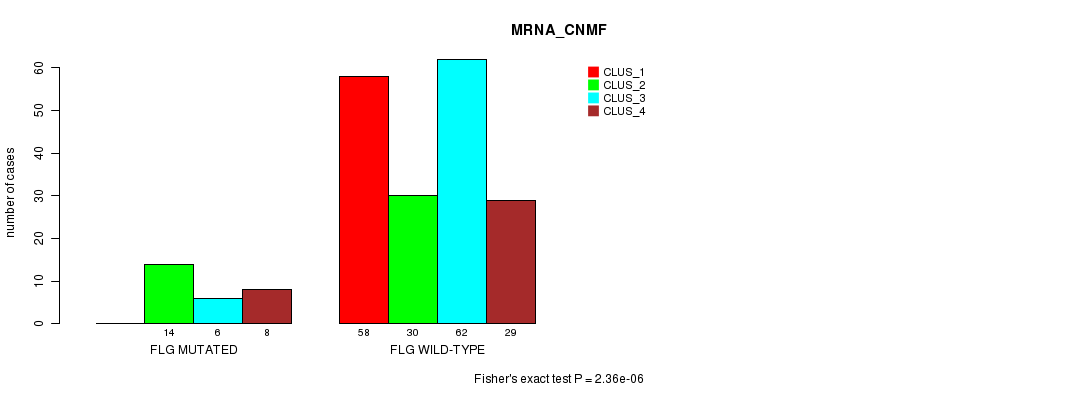
P value = 4.08e-05 (Fisher's exact test), Q value = 0.073
Table S15. Gene #40: 'FLG MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| FLG MUTATED | 18 | 9 | 5 |
| FLG WILD-TYPE | 35 | 118 | 33 |
Figure S15. Get High-res Image Gene #40: 'FLG MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
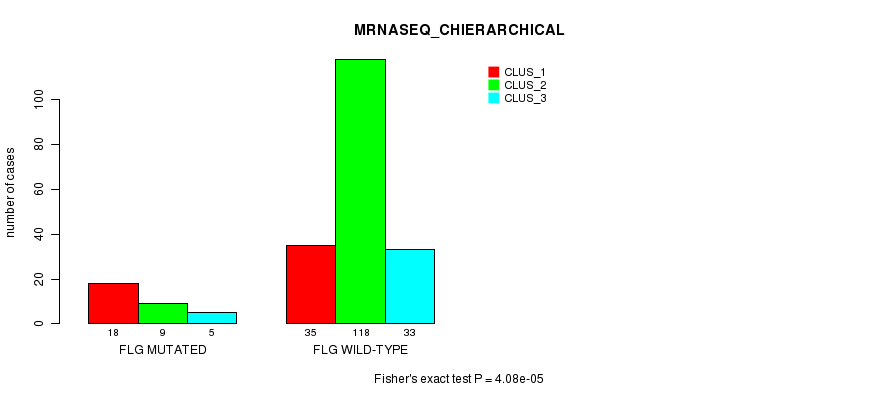
P value = 0.00013 (Fisher's exact test), Q value = 0.23
Table S16. Gene #51: 'DNAH5 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 86 | 83 | 36 | 16 |
| DNAH5 MUTATED | 4 | 24 | 4 | 2 |
| DNAH5 WILD-TYPE | 82 | 59 | 32 | 14 |
Figure S16. Get High-res Image Gene #51: 'DNAH5 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
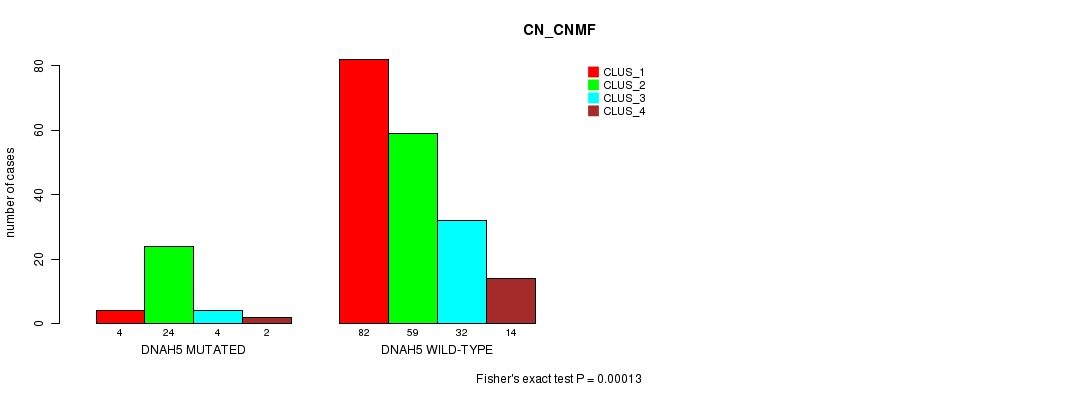
P value = 7.52e-05 (Fisher's exact test), Q value = 0.13
Table S17. Gene #51: 'DNAH5 MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 78 | 47 | 60 | 33 |
| DNAH5 MUTATED | 3 | 15 | 13 | 3 |
| DNAH5 WILD-TYPE | 75 | 32 | 47 | 30 |
Figure S17. Get High-res Image Gene #51: 'DNAH5 MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
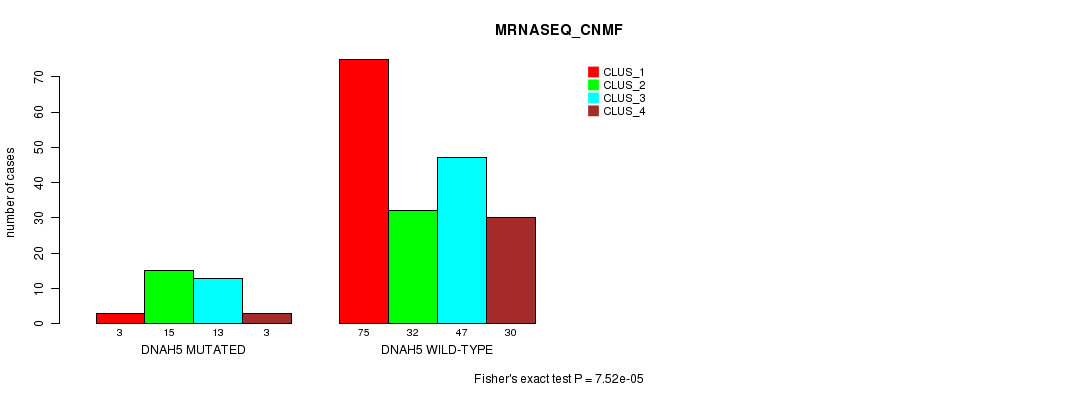
P value = 1.52e-06 (Fisher's exact test), Q value = 0.0027
Table S18. Gene #51: 'DNAH5 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| DNAH5 MUTATED | 20 | 8 | 6 |
| DNAH5 WILD-TYPE | 33 | 119 | 32 |
Figure S18. Get High-res Image Gene #51: 'DNAH5 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
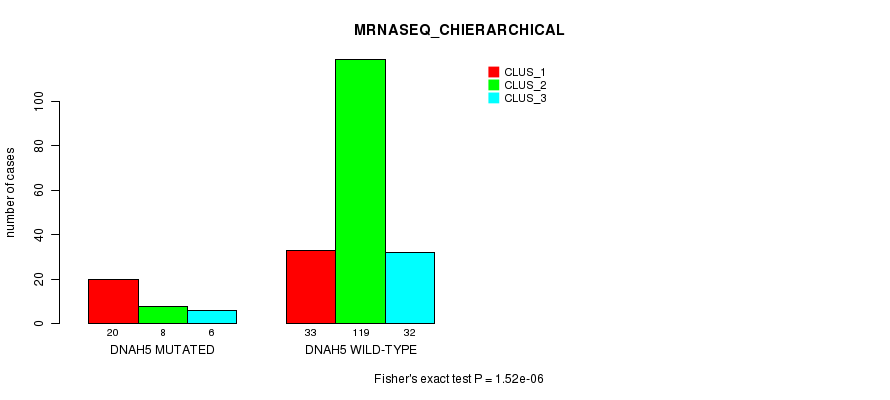
P value = 0.000102 (Fisher's exact test), Q value = 0.18
Table S19. Gene #56: 'SDK1 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| SDK1 MUTATED | 16 | 8 | 3 |
| SDK1 WILD-TYPE | 37 | 119 | 35 |
Figure S19. Get High-res Image Gene #56: 'SDK1 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
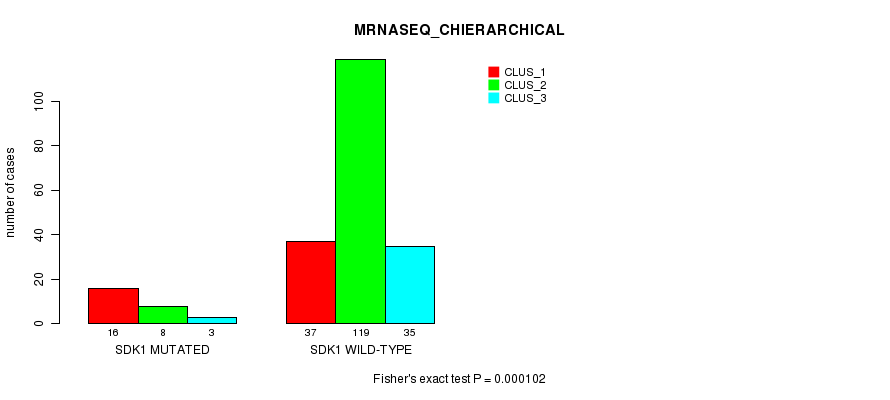
P value = 2.85e-05 (Fisher's exact test), Q value = 0.051
Table S20. Gene #58: 'TMEM132D MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 44 | 68 | 37 |
| TMEM132D MUTATED | 1 | 11 | 1 | 3 |
| TMEM132D WILD-TYPE | 57 | 33 | 67 | 34 |
Figure S20. Get High-res Image Gene #58: 'TMEM132D MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
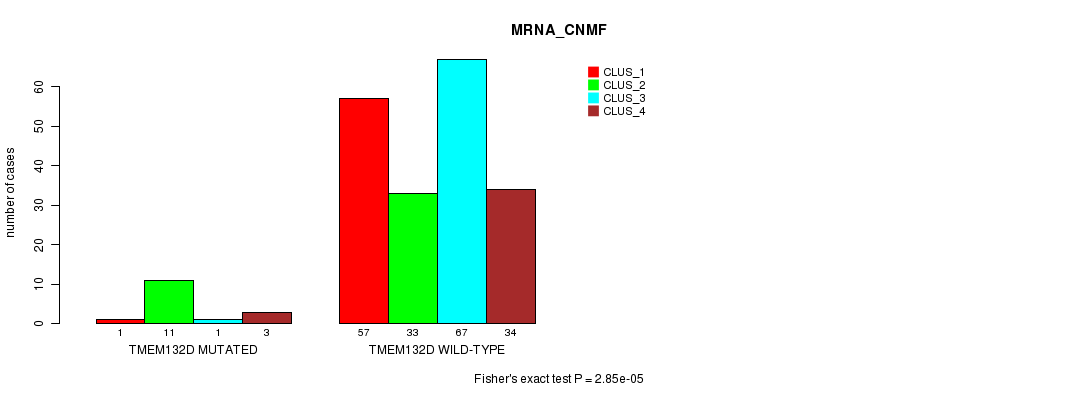
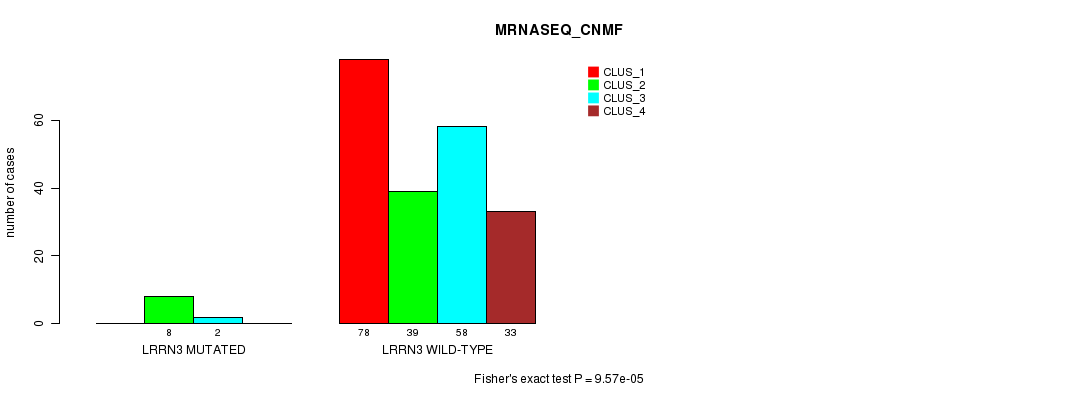
P value = 9.57e-05 (Fisher's exact test), Q value = 0.17
Table S21. Gene #80: 'LRRN3 MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 78 | 47 | 60 | 33 |
| LRRN3 MUTATED | 0 | 8 | 2 | 0 |
| LRRN3 WILD-TYPE | 78 | 39 | 58 | 33 |
Figure S21. Get High-res Image Gene #80: 'LRRN3 MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
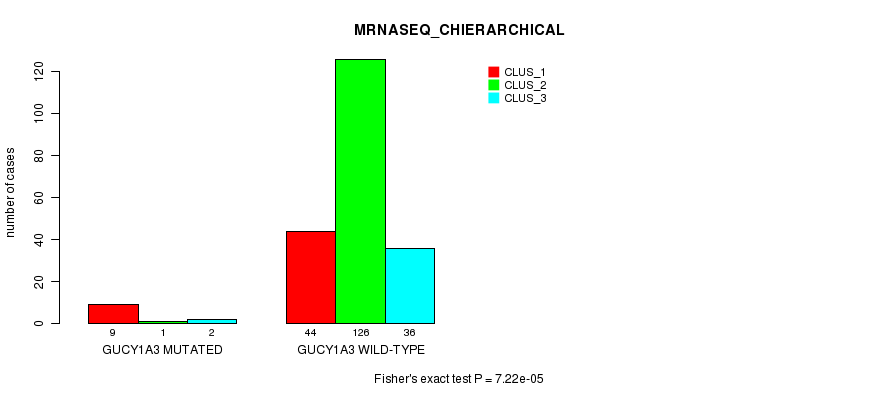
P value = 7.22e-05 (Fisher's exact test), Q value = 0.13
Table S22. Gene #95: 'GUCY1A3 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| GUCY1A3 MUTATED | 9 | 1 | 2 |
| GUCY1A3 WILD-TYPE | 44 | 126 | 36 |
Figure S22. Get High-res Image Gene #95: 'GUCY1A3 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
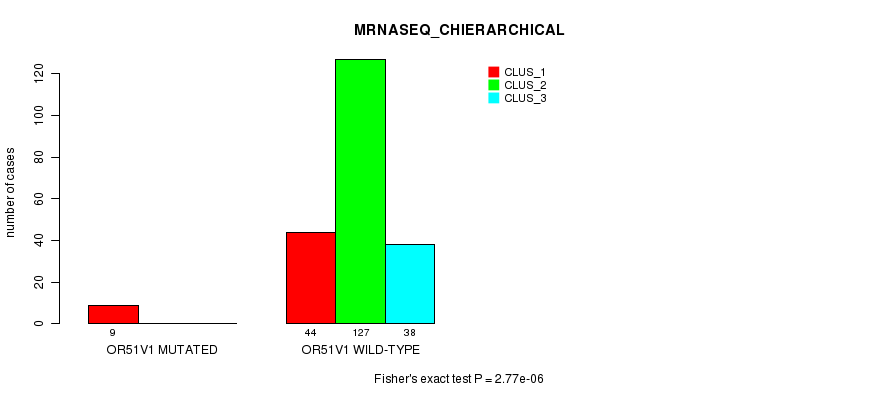
P value = 2.77e-06 (Fisher's exact test), Q value = 0.005
Table S23. Gene #108: 'OR51V1 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| OR51V1 MUTATED | 9 | 0 | 0 |
| OR51V1 WILD-TYPE | 44 | 127 | 38 |
Figure S23. Get High-res Image Gene #108: 'OR51V1 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
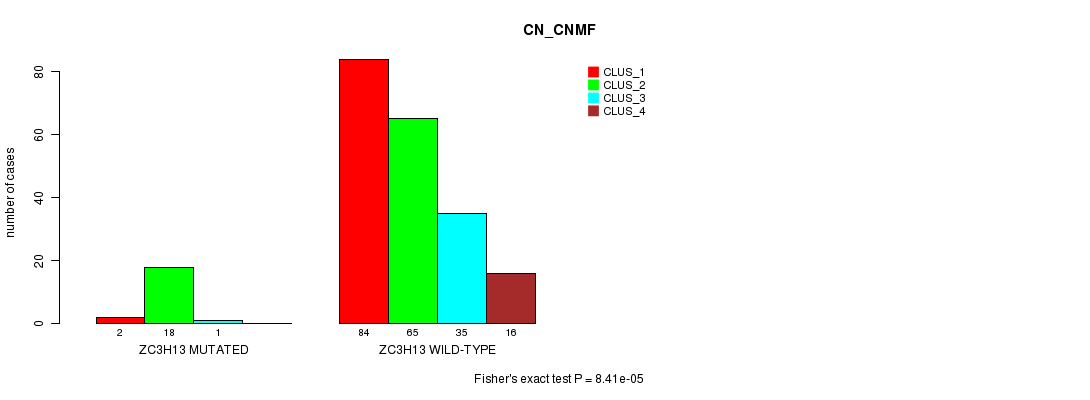
P value = 8.41e-05 (Fisher's exact test), Q value = 0.15
Table S24. Gene #109: 'ZC3H13 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 86 | 83 | 36 | 16 |
| ZC3H13 MUTATED | 2 | 18 | 1 | 0 |
| ZC3H13 WILD-TYPE | 84 | 65 | 35 | 16 |
Figure S24. Get High-res Image Gene #109: 'ZC3H13 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
P value = 6.7e-05 (Fisher's exact test), Q value = 0.12
Table S25. Gene #109: 'ZC3H13 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| ZC3H13 MUTATED | 13 | 4 | 4 |
| ZC3H13 WILD-TYPE | 40 | 123 | 34 |
Figure S25. Get High-res Image Gene #109: 'ZC3H13 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
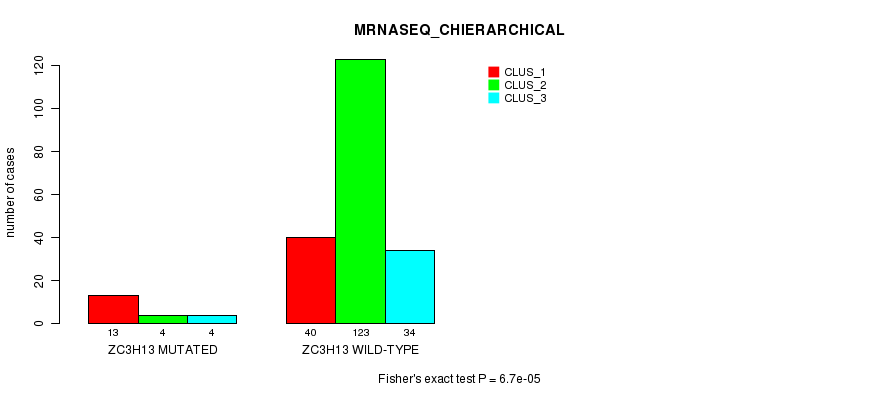
P value = 0.000129 (Fisher's exact test), Q value = 0.23
Table S26. Gene #116: 'CSMD3 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| CSMD3 MUTATED | 16 | 8 | 4 |
| CSMD3 WILD-TYPE | 37 | 119 | 34 |
Figure S26. Get High-res Image Gene #116: 'CSMD3 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
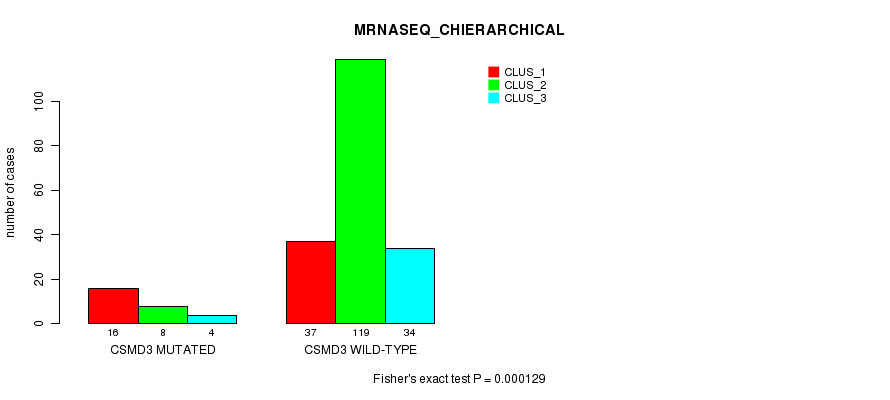
P value = 0.000116 (Fisher's exact test), Q value = 0.21
Table S27. Gene #117: 'DCAF4L2 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| DCAF4L2 MUTATED | 9 | 1 | 1 |
| DCAF4L2 WILD-TYPE | 44 | 126 | 37 |
Figure S27. Get High-res Image Gene #117: 'DCAF4L2 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
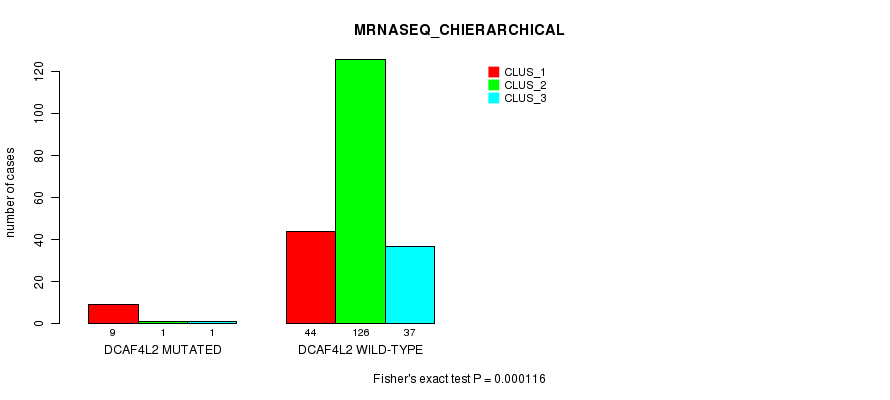
P value = 0.000122 (Fisher's exact test), Q value = 0.22
Table S28. Gene #119: 'DOCK2 MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 44 | 68 | 37 |
| DOCK2 MUTATED | 1 | 8 | 1 | 7 |
| DOCK2 WILD-TYPE | 57 | 36 | 67 | 30 |
Figure S28. Get High-res Image Gene #119: 'DOCK2 MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
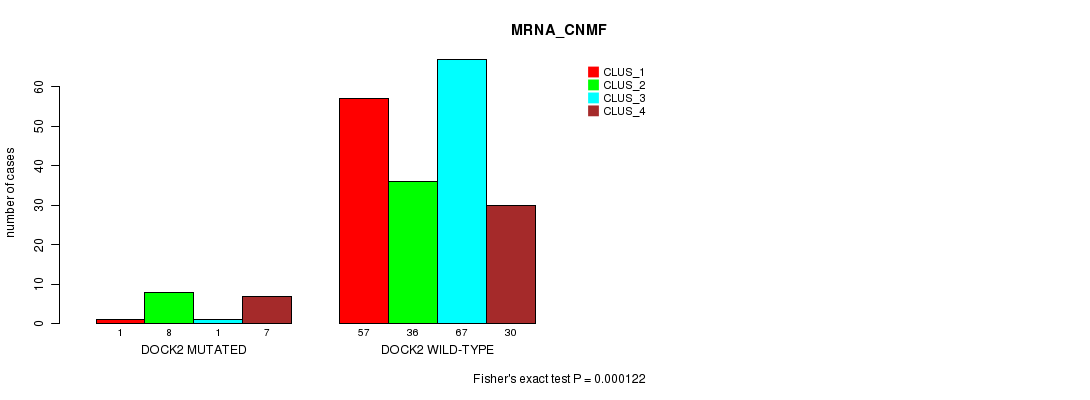
P value = 0.000138 (Fisher's exact test), Q value = 0.25
Table S29. Gene #119: 'DOCK2 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 86 | 83 | 36 | 16 |
| DOCK2 MUTATED | 1 | 17 | 2 | 1 |
| DOCK2 WILD-TYPE | 85 | 66 | 34 | 15 |
Figure S29. Get High-res Image Gene #119: 'DOCK2 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
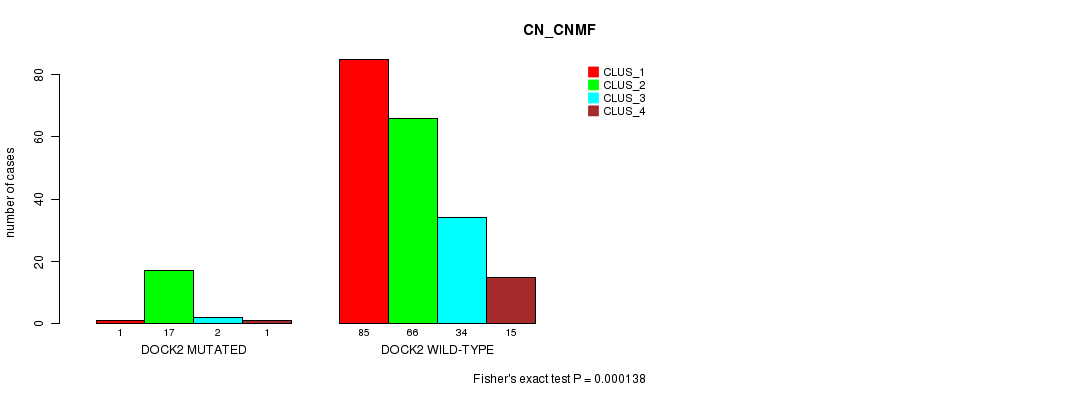
P value = 1.35e-05 (Fisher's exact test), Q value = 0.024
Table S30. Gene #119: 'DOCK2 MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 78 | 47 | 60 | 33 |
| DOCK2 MUTATED | 0 | 8 | 12 | 1 |
| DOCK2 WILD-TYPE | 78 | 39 | 48 | 32 |
Figure S30. Get High-res Image Gene #119: 'DOCK2 MUTATION STATUS' versus Clinical Feature #7: 'MRNASEQ_CNMF'
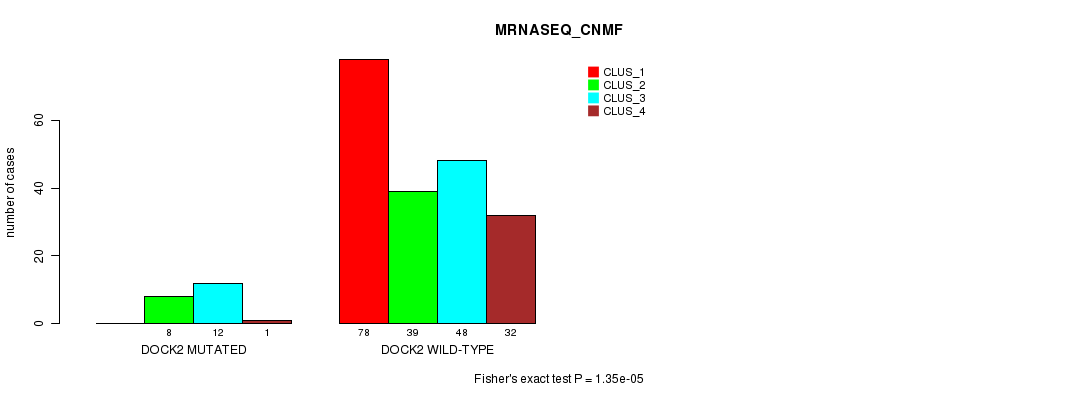
P value = 7.91e-07 (Fisher's exact test), Q value = 0.0014
Table S31. Gene #119: 'DOCK2 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| DOCK2 MUTATED | 14 | 2 | 5 |
| DOCK2 WILD-TYPE | 39 | 125 | 33 |
Figure S31. Get High-res Image Gene #119: 'DOCK2 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
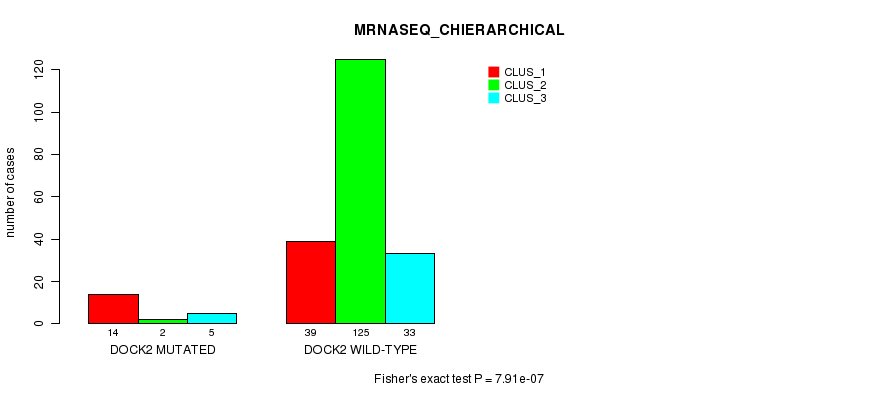
P value = 0.000103 (Fisher's exact test), Q value = 0.18
Table S32. Gene #135: 'RIMS1 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| RIMS1 MUTATED | 11 | 3 | 1 |
| RIMS1 WILD-TYPE | 42 | 124 | 37 |
Figure S32. Get High-res Image Gene #135: 'RIMS1 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
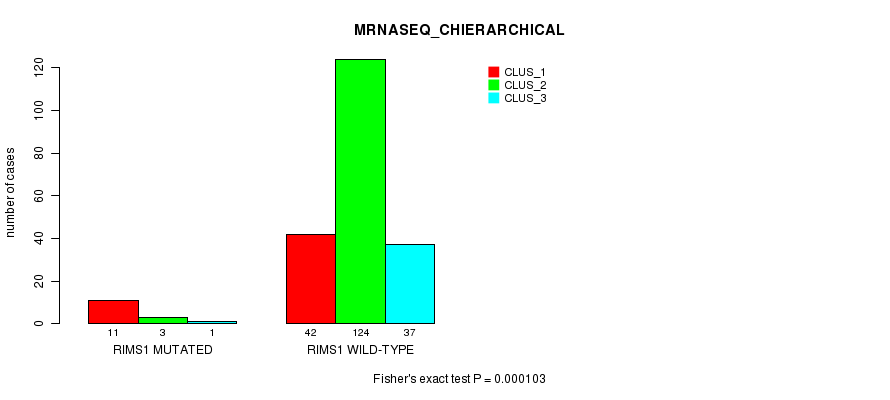
P value = 4.46e-05 (Fisher's exact test), Q value = 0.08
Table S33. Gene #138: 'CDH10 MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 44 | 68 | 37 |
| CDH10 MUTATED | 0 | 4 | 1 | 8 |
| CDH10 WILD-TYPE | 58 | 40 | 67 | 29 |
Figure S33. Get High-res Image Gene #138: 'CDH10 MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
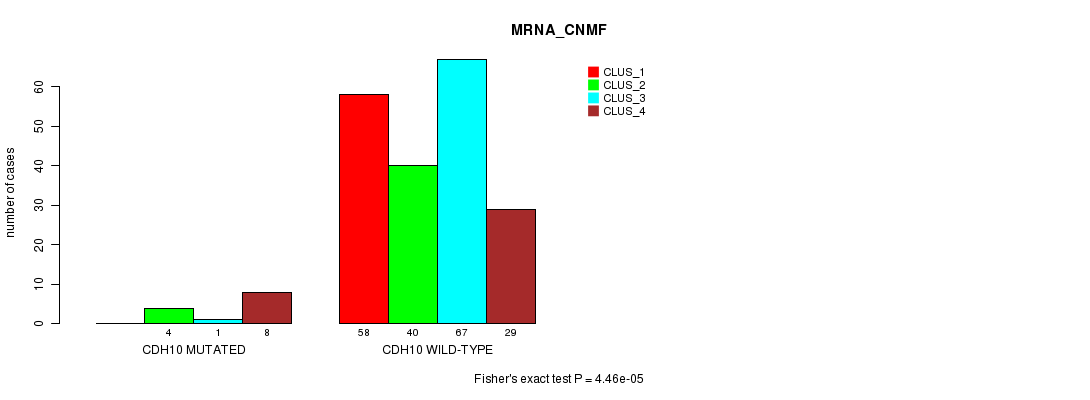
P value = 0.000131 (Fisher's exact test), Q value = 0.23
Table S34. Gene #138: 'CDH10 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 86 | 83 | 36 | 16 |
| CDH10 MUTATED | 0 | 13 | 1 | 0 |
| CDH10 WILD-TYPE | 86 | 70 | 35 | 16 |
Figure S34. Get High-res Image Gene #138: 'CDH10 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
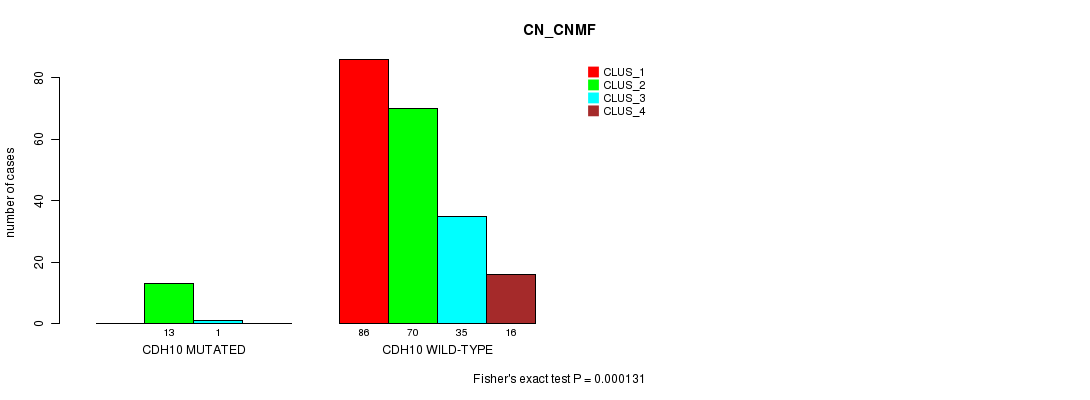
P value = 2.68e-05 (Fisher's exact test), Q value = 0.048
Table S35. Gene #138: 'CDH10 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 53 | 127 | 38 |
| CDH10 MUTATED | 11 | 2 | 1 |
| CDH10 WILD-TYPE | 42 | 125 | 37 |
Figure S35. Get High-res Image Gene #138: 'CDH10 MUTATION STATUS' versus Clinical Feature #8: 'MRNASEQ_CHIERARCHICAL'
P value = 2.86e-05 (Fisher's exact test), Q value = 0.051
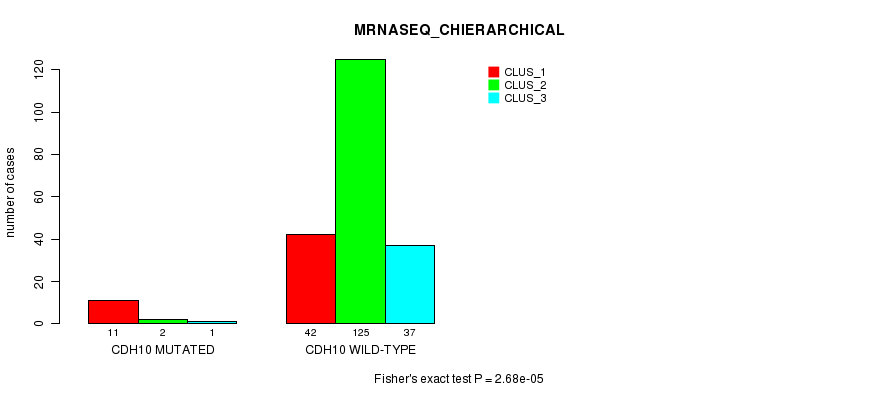
Table S36. Gene #142: 'SYNE1 MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 44 | 68 | 37 |
| SYNE1 MUTATED | 8 | 14 | 6 | 17 |
| SYNE1 WILD-TYPE | 50 | 30 | 62 | 20 |
Figure S36. Get High-res Image Gene #142: 'SYNE1 MUTATION STATUS' versus Clinical Feature #1: 'MRNA_CNMF'
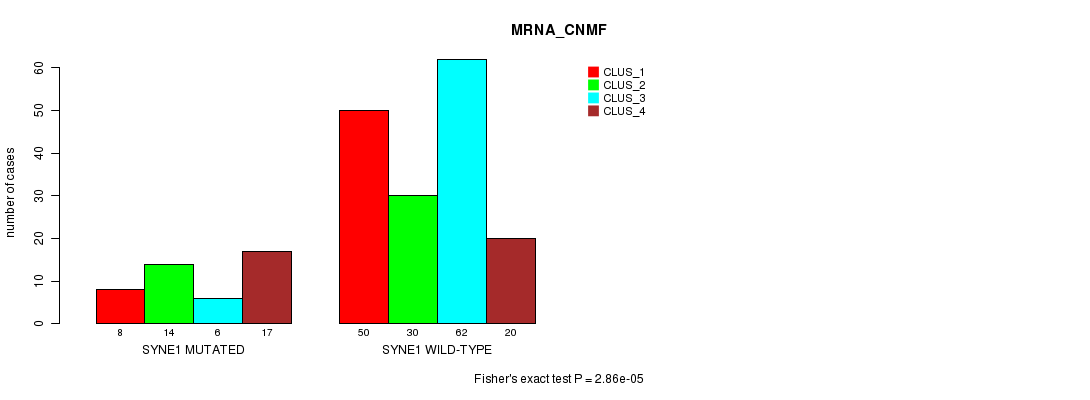
P value = 2.56e-05 (Fisher's exact test), Q value = 0.046
Table S37. Gene #142: 'SYNE1 MUTATION STATUS' versus Clinical Feature #2: 'MRNA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 42 | 59 | 106 |
| SYNE1 MUTATED | 3 | 25 | 17 |
| SYNE1 WILD-TYPE | 39 | 34 | 89 |
Figure S37. Get High-res Image Gene #142: 'SYNE1 MUTATION STATUS' versus Clinical Feature #2: 'MRNA_CHIERARCHICAL'
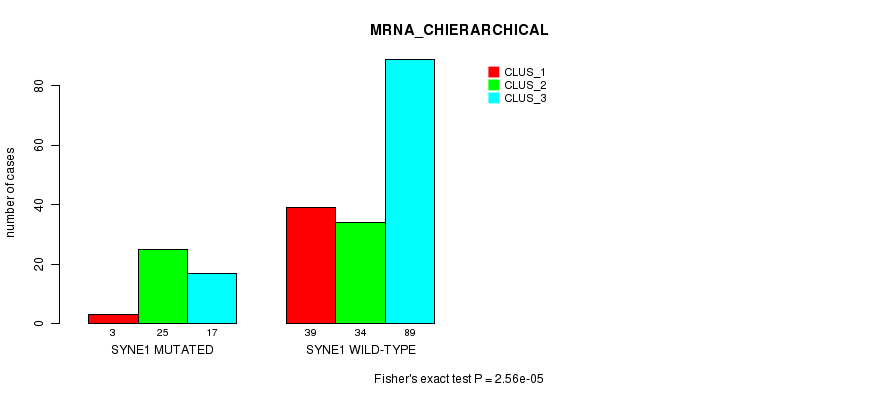
P value = 1.63e-06 (Fisher's exact test), Q value = 0.0029
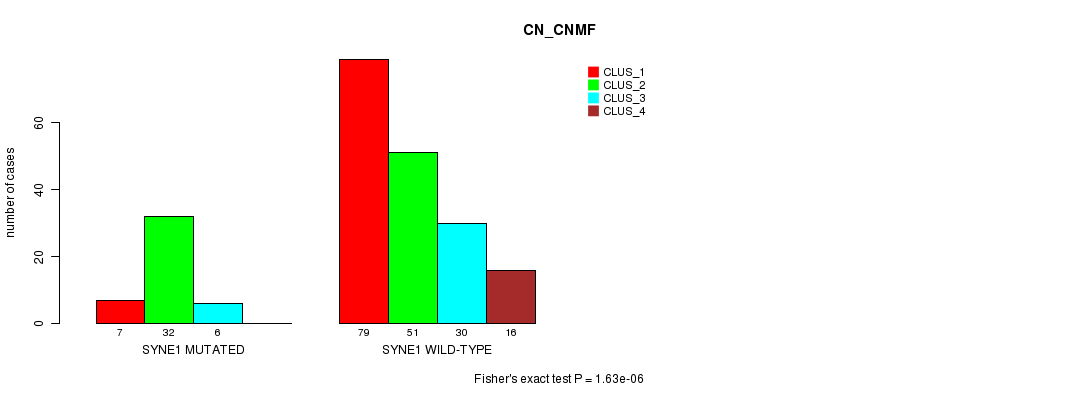
Table S38. Gene #142: 'SYNE1 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 86 | 83 | 36 | 16 |
| SYNE1 MUTATED | 7 | 32 | 6 | 0 |
| SYNE1 WILD-TYPE | 79 | 51 | 30 | 16 |
Figure S38. Get High-res Image Gene #142: 'SYNE1 MUTATION STATUS' versus Clinical Feature #3: 'CN_CNMF'
-
Mutation data file = COADREAD-TP.mutsig.cluster.txt
-
Molecular subtypes file = COADREAD-TP.transferedmergedcluster.txt
-
Number of patients = 224
-
Number of significantly mutated genes = 209
-
Number of Molecular subtypes = 10
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.