This pipeline computes the correlation between significant arm-level copy number variations (cnvs) and subtypes.
Testing the association between copy number variation 51 arm-level results and 8 molecular subtypes across 220 patients, 49 significant findings detected with Q value < 0.25.
-
1p gain cnv correlated to 'MRNASEQ_CNMF'.
-
7p gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CNMF'.
-
7q gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CNMF'.
-
10p gain cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
19p gain cnv correlated to 'CN_CNMF'.
-
19q gain cnv correlated to 'CN_CNMF' and 'MRNASEQ_CNMF'.
-
20p gain cnv correlated to 'CN_CNMF' and 'MRNASEQ_CNMF'.
-
20q gain cnv correlated to 'CN_CNMF' and 'MRNASEQ_CNMF'.
-
1p loss cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
1q loss cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
4p loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CNMF'.
-
4q loss cnv correlated to 'METHLYATION_CNMF' and 'MIRSEQ_CNMF'.
-
10p loss cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CNMF'.
-
10q loss cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CNMF'.
-
11p loss cnv correlated to 'METHLYATION_CNMF'.
-
19q loss cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
22q loss cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 51 arm-level results and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 49 significant findings detected.
|
Molecular subtypes |
MRNA CNMF |
MRNA CHIERARCHICAL |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Chi-square test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| 1p loss | 67 (30%) | 153 |
0.00361 (0.974) |
0.00862 (1.00) |
4.56e-47 (1.57e-44) |
1.46e-42 (4.96e-40) |
1.61e-24 (5.41e-22) |
1.37e-30 (4.66e-28) |
1.41e-19 (4.72e-17) |
3.05e-05 (0.00951) |
| 19q loss | 76 (35%) | 144 |
0.247 (1.00) |
0.145 (1.00) |
1.4e-45 (4.79e-43) |
9.63e-33 (3.27e-30) |
9.57e-22 (3.22e-19) |
4.5e-27 (1.52e-24) |
3.01e-17 (9.94e-15) |
3.32e-05 (0.0103) |
| 7p gain | 37 (17%) | 183 |
0.175 (1.00) |
0.162 (1.00) |
1.86e-17 (6.17e-15) |
3.65e-07 (0.000118) |
8.17e-12 (2.65e-09) |
0.000204 (0.0615) |
0.000552 (0.164) |
0.0071 (1.00) |
| 4p loss | 15 (7%) | 205 |
0.415 (1.00) |
0.329 (1.00) |
0.000882 (0.259) |
0.000169 (0.0513) |
0.000108 (0.0328) |
0.000418 (0.125) |
4.98e-06 (0.00159) |
0.833 (1.00) |
| 7q gain | 51 (23%) | 169 |
0.282 (1.00) |
0.144 (1.00) |
6.78e-15 (2.21e-12) |
6.01e-06 (0.00192) |
6.56e-08 (2.13e-05) |
0.00139 (0.396) |
0.00423 (1.00) |
0.128 (1.00) |
| 1q loss | 9 (4%) | 211 |
0.0889 (1.00) |
0.0971 (1.00) |
8.38e-05 (0.0256) |
3.23e-05 (0.01) |
0.00603 (1.00) |
0.00013 (0.0397) |
0.0106 (1.00) |
0.715 (1.00) |
| 10p loss | 31 (14%) | 189 |
0.271 (1.00) |
0.241 (1.00) |
2.3e-17 (7.62e-15) |
7.92e-17 (2.61e-14) |
4.79e-19 (1.59e-16) |
0.0016 (0.454) |
0.0191 (1.00) |
0.0066 (1.00) |
| 10q loss | 32 (15%) | 188 |
0.271 (1.00) |
0.241 (1.00) |
3.46e-16 (1.14e-13) |
6.63e-16 (2.17e-13) |
3.81e-20 (1.28e-17) |
0.00109 (0.318) |
0.0128 (1.00) |
0.0178 (1.00) |
| 22q loss | 15 (7%) | 205 |
1 (1.00) |
0.763 (1.00) |
7.15e-06 (0.00227) |
0.00075 (0.221) |
1.92e-05 (0.00605) |
0.0451 (1.00) |
0.149 (1.00) |
0.149 (1.00) |
| 10p gain | 21 (10%) | 199 |
6.41e-07 (0.000206) |
2.89e-05 (0.00905) |
0.0175 (1.00) |
0.094 (1.00) |
0.54 (1.00) |
0.93 (1.00) |
||
| 19q gain | 9 (4%) | 211 |
0.00034 (0.102) |
0.0103 (1.00) |
0.000336 (0.101) |
0.0217 (1.00) |
0.115 (1.00) |
0.162 (1.00) |
||
| 20p gain | 17 (8%) | 203 |
1.66e-05 (0.00528) |
0.00225 (0.627) |
1.7e-05 (0.00537) |
0.259 (1.00) |
0.0281 (1.00) |
0.267 (1.00) |
||
| 20q gain | 16 (7%) | 204 |
8.04e-05 (0.0247) |
0.00128 (0.365) |
7.35e-08 (2.37e-05) |
0.0252 (1.00) |
0.00356 (0.966) |
0.0462 (1.00) |
||
| 4q loss | 24 (11%) | 196 |
0.839 (1.00) |
0.828 (1.00) |
0.0049 (1.00) |
0.000613 (0.181) |
0.002 (0.56) |
0.00355 (0.966) |
6.65e-05 (0.0205) |
0.6 (1.00) |
| 1p gain | 6 (3%) | 214 |
0.0272 (1.00) |
0.115 (1.00) |
3.39e-05 (0.0105) |
0.103 (1.00) |
0.234 (1.00) |
0.208 (1.00) |
||
| 19p gain | 11 (5%) | 209 |
2.25e-05 (0.00708) |
0.00124 (0.356) |
0.00122 (0.353) |
0.0102 (1.00) |
0.0861 (1.00) |
0.0595 (1.00) |
||
| 11p loss | 20 (9%) | 200 |
0.815 (1.00) |
0.526 (1.00) |
0.00199 (0.56) |
0.000474 (0.141) |
0.0162 (1.00) |
0.00897 (1.00) |
0.0113 (1.00) |
0.0469 (1.00) |
| 1q gain | 8 (4%) | 212 |
0.00126 (0.362) |
0.0898 (1.00) |
0.0312 (1.00) |
0.618 (1.00) |
0.851 (1.00) |
0.569 (1.00) |
||
| 6p gain | 4 (2%) | 216 |
0.213 (1.00) |
0.426 (1.00) |
0.0517 (1.00) |
0.279 (1.00) |
0.0922 (1.00) |
0.717 (1.00) |
||
| 8p gain | 14 (6%) | 206 |
0.0337 (1.00) |
0.26 (1.00) |
0.606 (1.00) |
0.464 (1.00) |
0.205 (1.00) |
0.459 (1.00) |
||
| 8q gain | 17 (8%) | 203 |
0.00467 (1.00) |
0.0872 (1.00) |
0.377 (1.00) |
0.203 (1.00) |
0.125 (1.00) |
0.172 (1.00) |
||
| 9p gain | 4 (2%) | 216 |
0.213 (1.00) |
0.355 (1.00) |
0.0715 (1.00) |
1 (1.00) |
1 (1.00) |
0.159 (1.00) |
||
| 9q gain | 6 (3%) | 214 |
0.0272 (1.00) |
0.274 (1.00) |
0.00231 (0.642) |
0.519 (1.00) |
0.747 (1.00) |
0.37 (1.00) |
||
| 11p gain | 9 (4%) | 211 |
0.5 (1.00) |
0.192 (1.00) |
0.00395 (1.00) |
0.00261 (0.717) |
0.138 (1.00) |
0.426 (1.00) |
||
| 11q gain | 14 (6%) | 206 |
0.353 (1.00) |
0.301 (1.00) |
0.172 (1.00) |
0.107 (1.00) |
0.708 (1.00) |
0.521 (1.00) |
||
| 12p gain | 11 (5%) | 209 |
0.0368 (1.00) |
0.179 (1.00) |
0.286 (1.00) |
0.17 (1.00) |
0.0299 (1.00) |
0.05 (1.00) |
||
| 12q gain | 5 (2%) | 215 |
0.848 (1.00) |
0.885 (1.00) |
0.934 (1.00) |
0.346 (1.00) |
0.0605 (1.00) |
0.157 (1.00) |
||
| 18p gain | 4 (2%) | 216 |
0.213 (1.00) |
0.504 (1.00) |
0.569 (1.00) |
0.279 (1.00) |
0.37 (1.00) |
0.0958 (1.00) |
||
| 21q gain | 3 (1%) | 217 |
0.328 (1.00) |
0.709 (1.00) |
0.158 (1.00) |
0.0879 (1.00) |
1 (1.00) |
1 (1.00) |
||
| 2p loss | 4 (2%) | 216 |
0.825 (1.00) |
1 (1.00) |
0.464 (1.00) |
0.37 (1.00) |
0.37 (1.00) |
0.0958 (1.00) |
||
| 2q loss | 3 (1%) | 217 |
0.796 (1.00) |
1 (1.00) |
0.513 (1.00) |
0.259 (1.00) |
0.0543 (1.00) |
0.387 (1.00) |
||
| 3p loss | 4 (2%) | 216 |
0.553 (1.00) |
0.504 (1.00) |
0.356 (1.00) |
0.279 (1.00) |
0.191 (1.00) |
0.241 (1.00) |
||
| 3q loss | 9 (4%) | 211 |
0.175 (1.00) |
1 (1.00) |
0.128 (1.00) |
0.109 (1.00) |
0.547 (1.00) |
0.453 (1.00) |
0.96 (1.00) |
0.849 (1.00) |
| 5p loss | 11 (5%) | 209 |
1 (1.00) |
0.388 (1.00) |
0.0162 (1.00) |
0.0317 (1.00) |
0.847 (1.00) |
0.36 (1.00) |
0.161 (1.00) |
0.766 (1.00) |
| 5q loss | 9 (4%) | 211 |
0.615 (1.00) |
0.0971 (1.00) |
0.0453 (1.00) |
0.0734 (1.00) |
0.0439 (1.00) |
0.0217 (1.00) |
0.00506 (1.00) |
0.139 (1.00) |
| 6p loss | 7 (3%) | 213 |
0.12 (1.00) |
0.0431 (1.00) |
0.139 (1.00) |
1 (1.00) |
0.693 (1.00) |
0.157 (1.00) |
||
| 6q loss | 21 (10%) | 199 |
0.00431 (1.00) |
0.0237 (1.00) |
0.362 (1.00) |
0.332 (1.00) |
0.0529 (1.00) |
0.32 (1.00) |
||
| 8p loss | 3 (1%) | 217 |
0.17 (1.00) |
0.571 (1.00) |
1 (1.00) |
0.125 (1.00) |
0.387 (1.00) |
|||
| 9p loss | 38 (17%) | 182 |
0.384 (1.00) |
0.828 (1.00) |
0.00289 (0.791) |
0.0555 (1.00) |
0.692 (1.00) |
0.245 (1.00) |
0.804 (1.00) |
0.915 (1.00) |
| 9q loss | 6 (3%) | 214 |
0.0702 (1.00) |
0.0542 (1.00) |
0.0111 (1.00) |
0.000882 (0.259) |
0.00806 (1.00) |
0.795 (1.00) |
||
| 11q loss | 4 (2%) | 216 |
0.00243 (0.669) |
0.238 (1.00) |
0.00187 (0.527) |
0.279 (1.00) |
0.697 (1.00) |
0.0958 (1.00) |
||
| 12q loss | 8 (4%) | 212 |
0.0444 (1.00) |
0.153 (1.00) |
0.686 (1.00) |
0.618 (1.00) |
0.182 (1.00) |
0.833 (1.00) |
||
| 13q loss | 28 (13%) | 192 |
1 (1.00) |
0.388 (1.00) |
0.635 (1.00) |
0.171 (1.00) |
0.86 (1.00) |
0.537 (1.00) |
0.914 (1.00) |
0.364 (1.00) |
| 14q loss | 24 (11%) | 196 |
0.00111 (0.322) |
0.0183 (1.00) |
0.00126 (0.362) |
0.452 (1.00) |
0.0127 (1.00) |
0.103 (1.00) |
||
| 15q loss | 12 (5%) | 208 |
0.00582 (1.00) |
0.0161 (1.00) |
0.000977 (0.285) |
0.00504 (1.00) |
0.248 (1.00) |
0.89 (1.00) |
||
| 16q loss | 6 (3%) | 214 |
0.662 (1.00) |
0.115 (1.00) |
0.302 (1.00) |
0.519 (1.00) |
0.549 (1.00) |
1 (1.00) |
||
| 18p loss | 15 (7%) | 205 |
0.344 (1.00) |
0.413 (1.00) |
0.144 (1.00) |
0.0484 (1.00) |
0.119 (1.00) |
0.117 (1.00) |
0.0755 (1.00) |
0.354 (1.00) |
| 18q loss | 16 (7%) | 204 |
0.0889 (1.00) |
0.0971 (1.00) |
0.00296 (0.807) |
0.00953 (1.00) |
0.0024 (0.664) |
0.0264 (1.00) |
0.0111 (1.00) |
0.51 (1.00) |
| 19p loss | 10 (5%) | 210 |
0.302 (1.00) |
0.0773 (1.00) |
0.0618 (1.00) |
0.0581 (1.00) |
0.0747 (1.00) |
0.0204 (1.00) |
0.0118 (1.00) |
0.744 (1.00) |
| 21q loss | 8 (4%) | 212 |
0.588 (1.00) |
0.423 (1.00) |
0.46 (1.00) |
0.618 (1.00) |
0.33 (1.00) |
0.197 (1.00) |
||
| Xq loss | 6 (3%) | 214 |
0.132 (1.00) |
0.203 (1.00) |
0.698 (1.00) |
0.103 (1.00) |
0.234 (1.00) |
0.208 (1.00) |
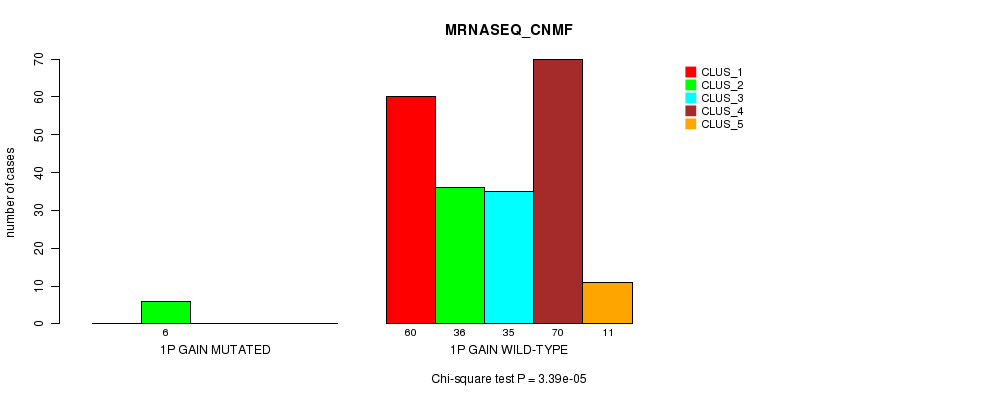
P value = 3.39e-05 (Chi-square test), Q value = 0.01
Table S1. Gene #1: '1p gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 1P GAIN MUTATED | 0 | 6 | 0 | 0 | 0 |
| 1P GAIN WILD-TYPE | 60 | 36 | 35 | 70 | 11 |
Figure S1. Get High-res Image Gene #1: '1p gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
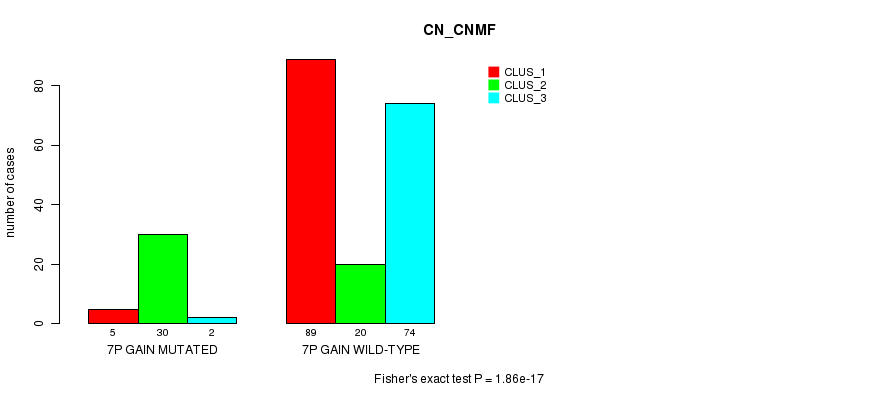
P value = 1.86e-17 (Fisher's exact test), Q value = 6.2e-15
Table S2. Gene #4: '7p gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 7P GAIN MUTATED | 5 | 30 | 2 |
| 7P GAIN WILD-TYPE | 89 | 20 | 74 |
Figure S2. Get High-res Image Gene #4: '7p gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
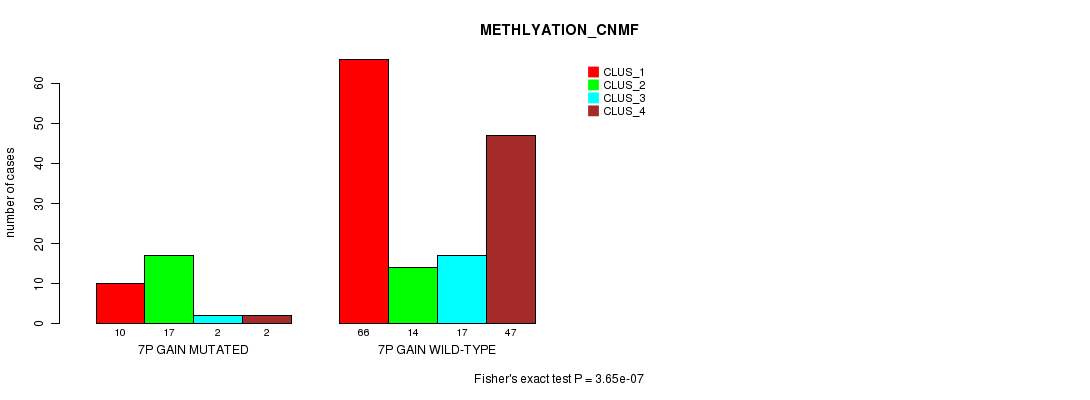
P value = 3.65e-07 (Fisher's exact test), Q value = 0.00012
Table S3. Gene #4: '7p gain mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 7P GAIN MUTATED | 10 | 17 | 2 | 2 |
| 7P GAIN WILD-TYPE | 66 | 14 | 17 | 47 |
Figure S3. Get High-res Image Gene #4: '7p gain mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
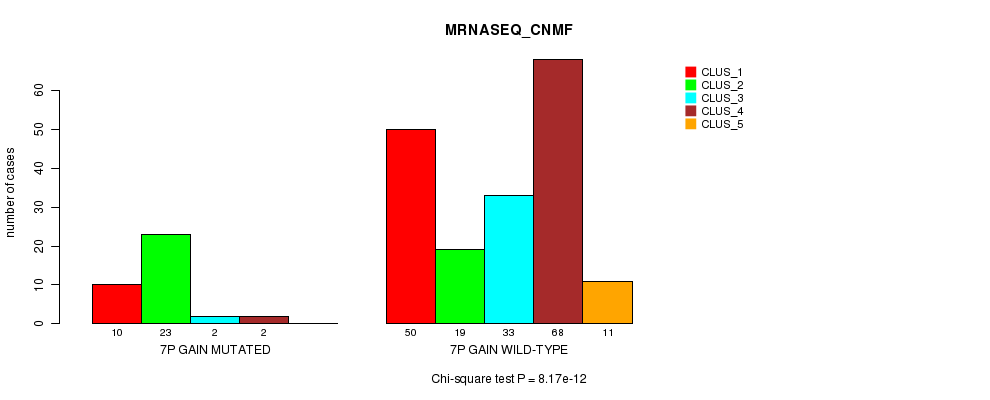
P value = 8.17e-12 (Chi-square test), Q value = 2.7e-09
Table S4. Gene #4: '7p gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 7P GAIN MUTATED | 10 | 23 | 2 | 2 | 0 |
| 7P GAIN WILD-TYPE | 50 | 19 | 33 | 68 | 11 |
Figure S4. Get High-res Image Gene #4: '7p gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
P value = 0.000204 (Fisher's exact test), Q value = 0.062
Table S5. Gene #4: '7p gain mutation analysis' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 32 | 117 | 69 |
| 7P GAIN MUTATED | 2 | 31 | 4 |
| 7P GAIN WILD-TYPE | 30 | 86 | 65 |
Figure S5. Get High-res Image Gene #4: '7p gain mutation analysis' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
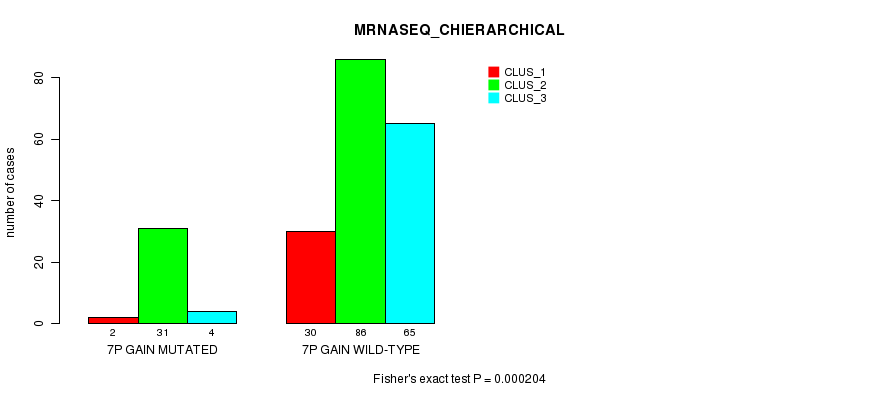
P value = 0.000552 (Fisher's exact test), Q value = 0.16
Table S6. Gene #4: '7p gain mutation analysis' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 50 | 50 | 32 | 87 |
| 7P GAIN MUTATED | 14 | 14 | 1 | 8 |
| 7P GAIN WILD-TYPE | 36 | 36 | 31 | 79 |
Figure S6. Get High-res Image Gene #4: '7p gain mutation analysis' versus Clinical Feature #7: 'MIRSEQ_CNMF'
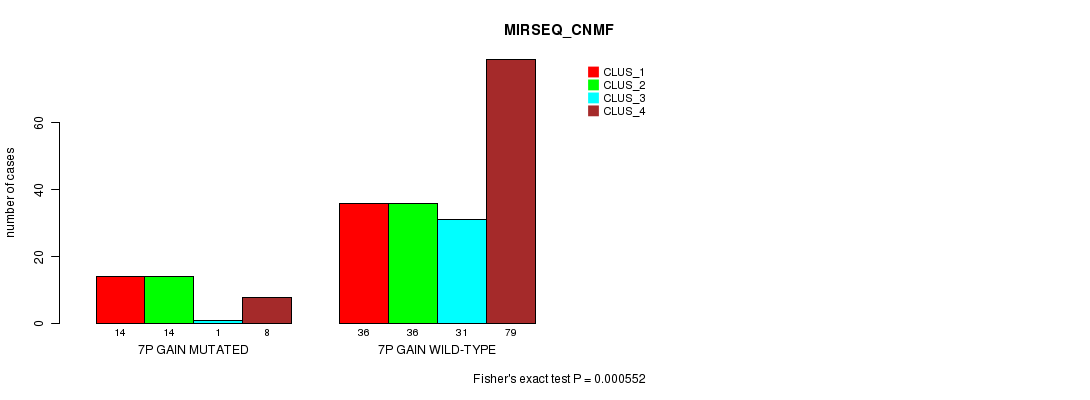
P value = 6.78e-15 (Fisher's exact test), Q value = 2.2e-12
Table S7. Gene #5: '7q gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 7Q GAIN MUTATED | 10 | 34 | 7 |
| 7Q GAIN WILD-TYPE | 84 | 16 | 69 |
Figure S7. Get High-res Image Gene #5: '7q gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'

P value = 6.01e-06 (Fisher's exact test), Q value = 0.0019
Table S8. Gene #5: '7q gain mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 7Q GAIN MUTATED | 16 | 19 | 3 | 5 |
| 7Q GAIN WILD-TYPE | 60 | 12 | 16 | 44 |
Figure S8. Get High-res Image Gene #5: '7q gain mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
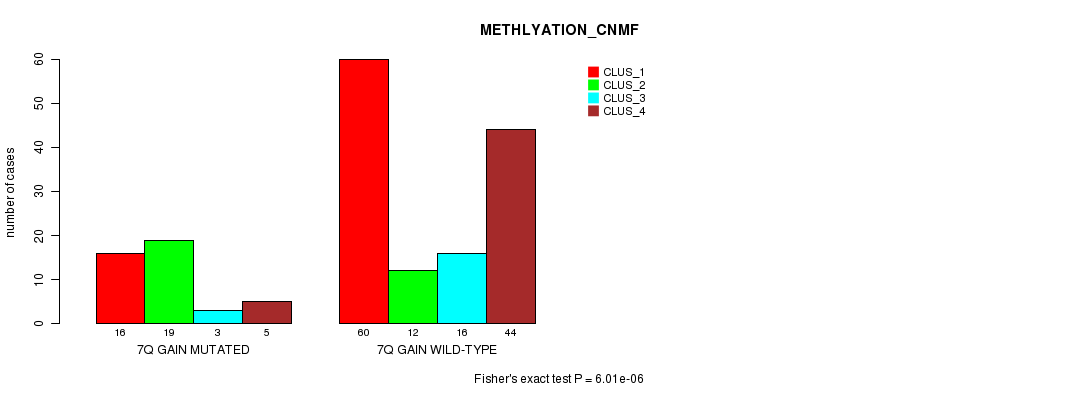
P value = 6.56e-08 (Chi-square test), Q value = 2.1e-05
Table S9. Gene #5: '7q gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 7Q GAIN MUTATED | 17 | 23 | 4 | 6 | 0 |
| 7Q GAIN WILD-TYPE | 43 | 19 | 31 | 64 | 11 |
Figure S9. Get High-res Image Gene #5: '7q gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
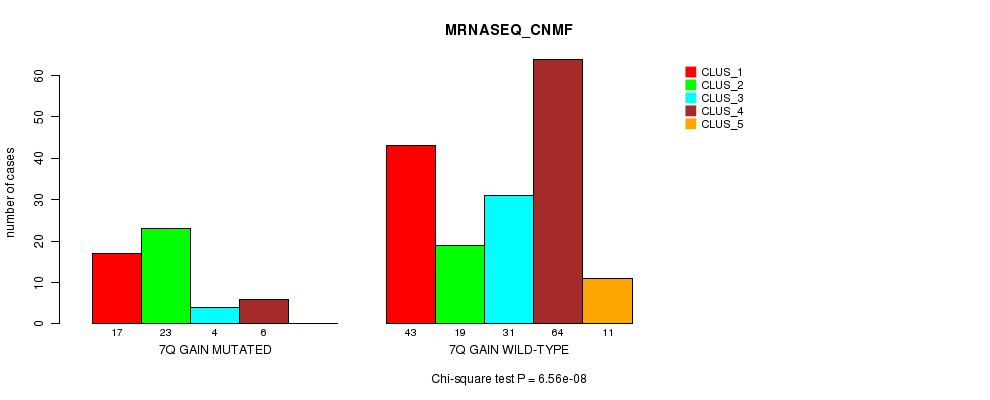
P value = 6.41e-07 (Fisher's exact test), Q value = 0.00021
Table S10. Gene #10: '10p gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 10P GAIN MUTATED | 20 | 0 | 1 |
| 10P GAIN WILD-TYPE | 74 | 50 | 75 |
Figure S10. Get High-res Image Gene #10: '10p gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
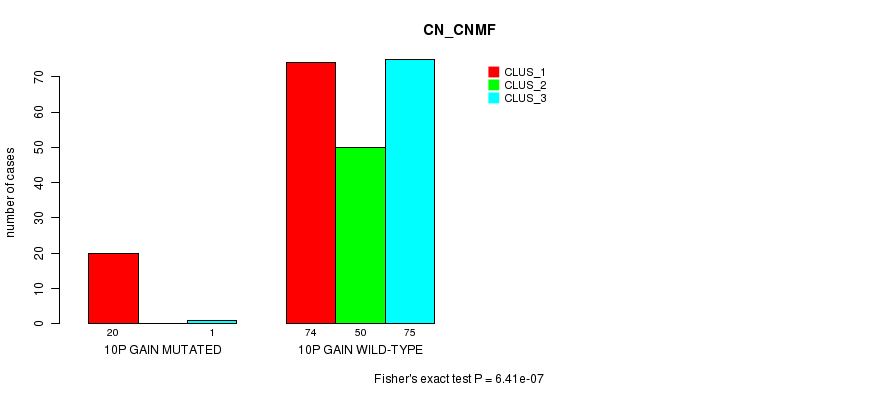
P value = 2.89e-05 (Fisher's exact test), Q value = 0.009
Table S11. Gene #10: '10p gain mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 10P GAIN MUTATED | 17 | 0 | 1 | 0 |
| 10P GAIN WILD-TYPE | 59 | 31 | 18 | 49 |
Figure S11. Get High-res Image Gene #10: '10p gain mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'

P value = 2.25e-05 (Fisher's exact test), Q value = 0.0071
Table S12. Gene #16: '19p gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 19P GAIN MUTATED | 2 | 9 | 0 |
| 19P GAIN WILD-TYPE | 92 | 41 | 76 |
Figure S12. Get High-res Image Gene #16: '19p gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
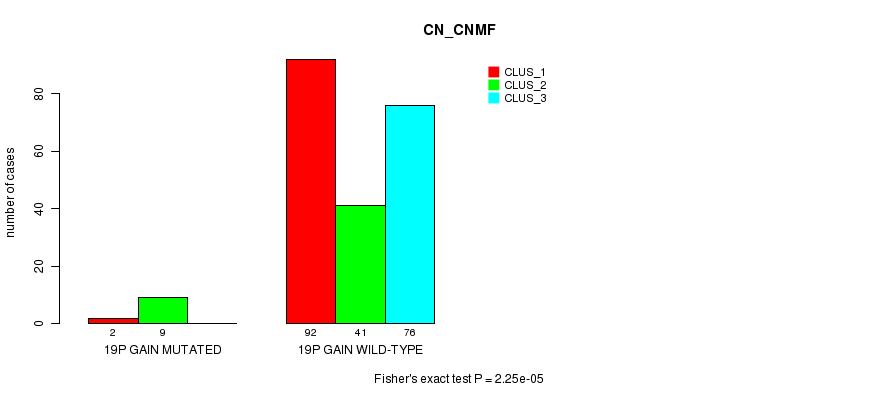
P value = 0.00034 (Fisher's exact test), Q value = 0.1
Table S13. Gene #17: '19q gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 19Q GAIN MUTATED | 2 | 7 | 0 |
| 19Q GAIN WILD-TYPE | 92 | 43 | 76 |
Figure S13. Get High-res Image Gene #17: '19q gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
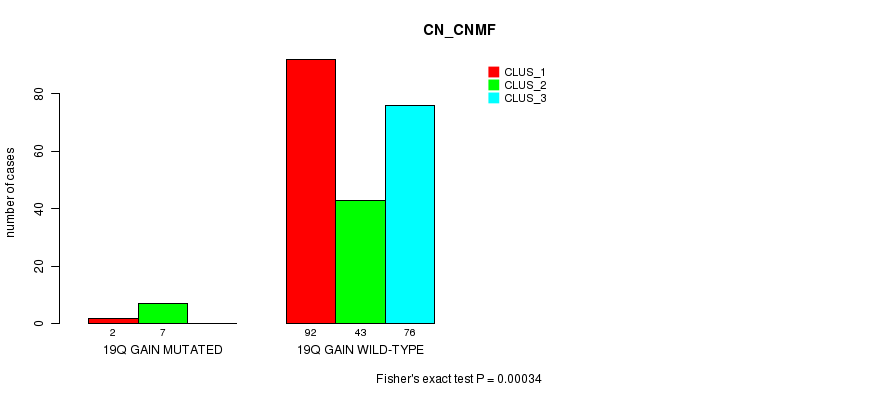
P value = 0.000336 (Chi-square test), Q value = 0.1
Table S14. Gene #17: '19q gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 19Q GAIN MUTATED | 1 | 7 | 0 | 1 | 0 |
| 19Q GAIN WILD-TYPE | 59 | 35 | 35 | 69 | 11 |
Figure S14. Get High-res Image Gene #17: '19q gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
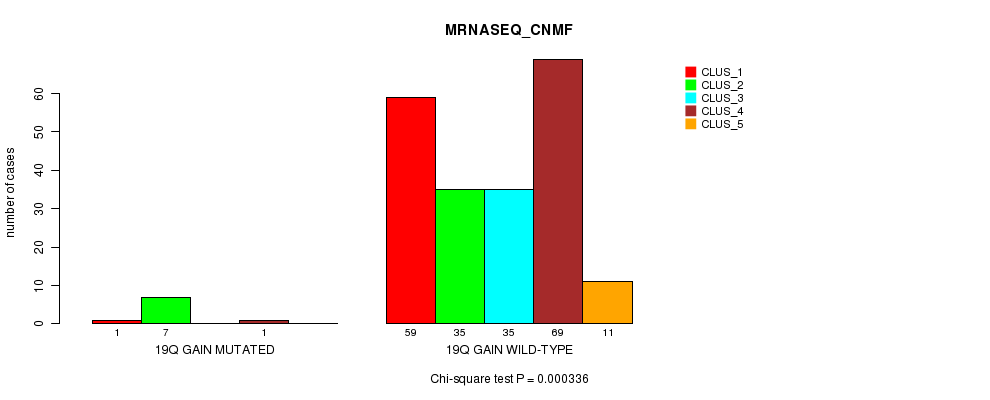
P value = 1.66e-05 (Fisher's exact test), Q value = 0.0053
Table S15. Gene #18: '20p gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 20P GAIN MUTATED | 4 | 12 | 1 |
| 20P GAIN WILD-TYPE | 90 | 38 | 75 |
Figure S15. Get High-res Image Gene #18: '20p gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
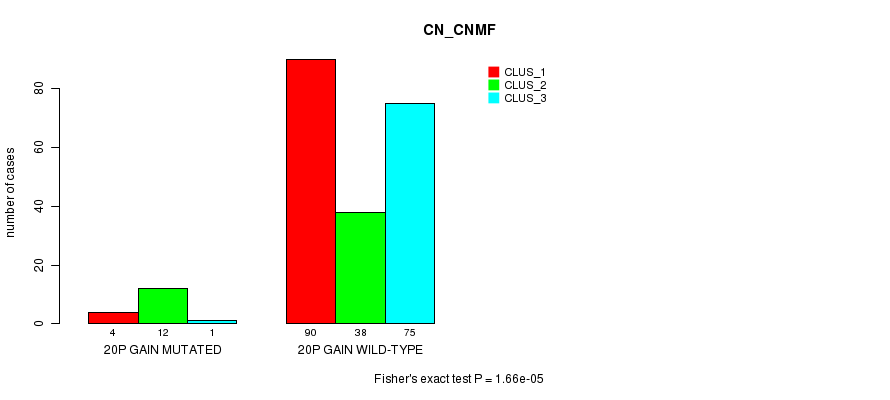
P value = 1.7e-05 (Chi-square test), Q value = 0.0054
Table S16. Gene #18: '20p gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 20P GAIN MUTATED | 2 | 11 | 1 | 2 | 0 |
| 20P GAIN WILD-TYPE | 58 | 31 | 34 | 68 | 11 |
Figure S16. Get High-res Image Gene #18: '20p gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
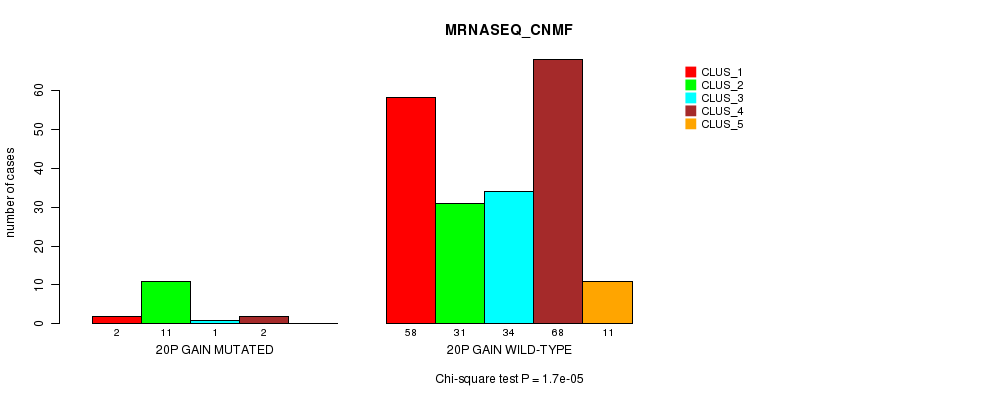
P value = 8.04e-05 (Fisher's exact test), Q value = 0.025
Table S17. Gene #19: '20q gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 20Q GAIN MUTATED | 4 | 11 | 1 |
| 20Q GAIN WILD-TYPE | 90 | 39 | 75 |
Figure S17. Get High-res Image Gene #19: '20q gain mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
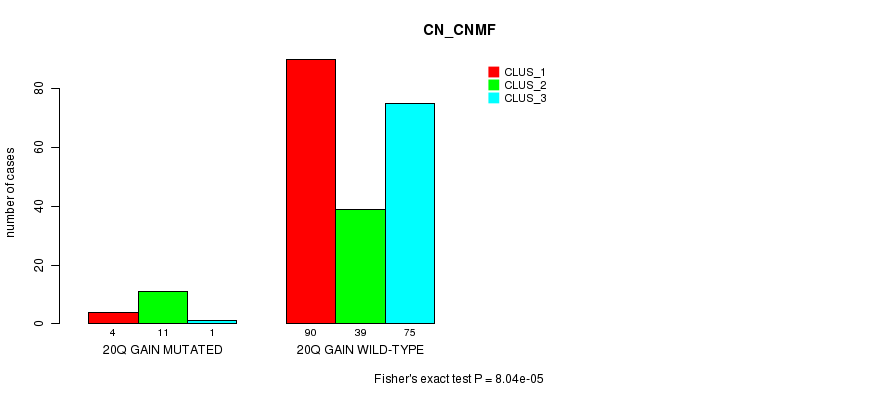
P value = 7.35e-08 (Chi-square test), Q value = 2.4e-05
Table S18. Gene #19: '20q gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 20Q GAIN MUTATED | 2 | 12 | 1 | 0 | 0 |
| 20Q GAIN WILD-TYPE | 58 | 30 | 34 | 70 | 11 |
Figure S18. Get High-res Image Gene #19: '20q gain mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
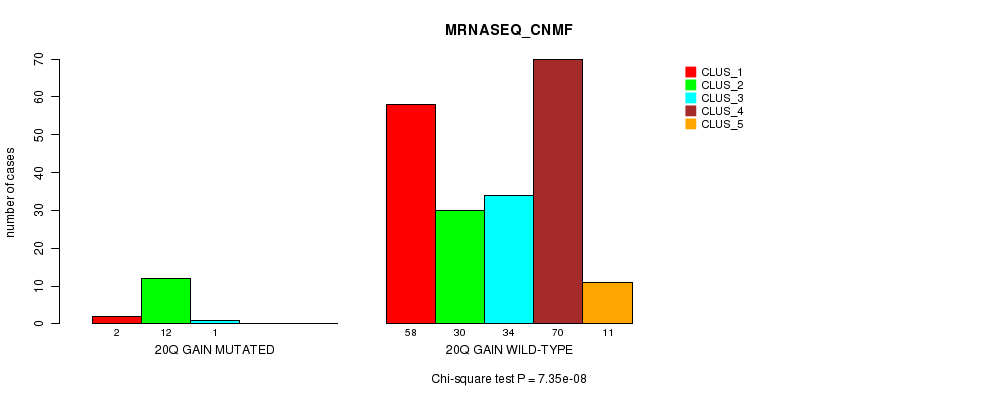
P value = 4.56e-47 (Fisher's exact test), Q value = 1.6e-44
Table S19. Gene #21: '1p loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 1P LOSS MUTATED | 0 | 0 | 67 |
| 1P LOSS WILD-TYPE | 94 | 50 | 9 |
Figure S19. Get High-res Image Gene #21: '1p loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
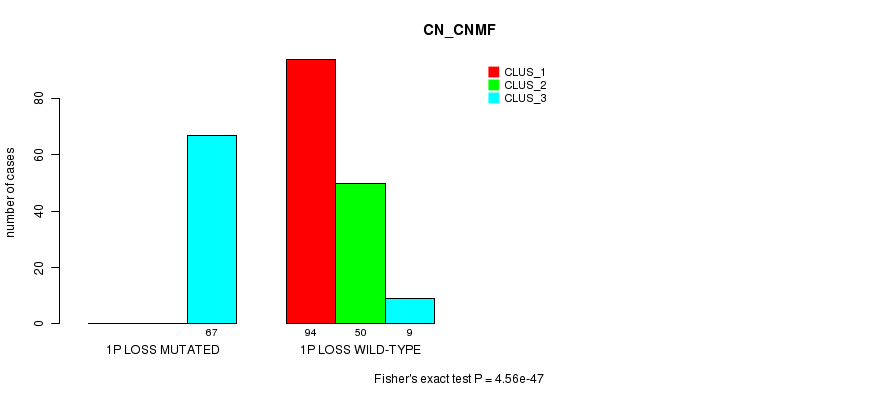
P value = 1.46e-42 (Fisher's exact test), Q value = 5e-40
Table S20. Gene #21: '1p loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 1P LOSS MUTATED | 0 | 0 | 4 | 49 |
| 1P LOSS WILD-TYPE | 76 | 31 | 15 | 0 |
Figure S20. Get High-res Image Gene #21: '1p loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'

P value = 1.61e-24 (Chi-square test), Q value = 5.4e-22
Table S21. Gene #21: '1p loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 1P LOSS MUTATED | 0 | 0 | 33 | 31 | 3 |
| 1P LOSS WILD-TYPE | 60 | 42 | 2 | 39 | 8 |
Figure S21. Get High-res Image Gene #21: '1p loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
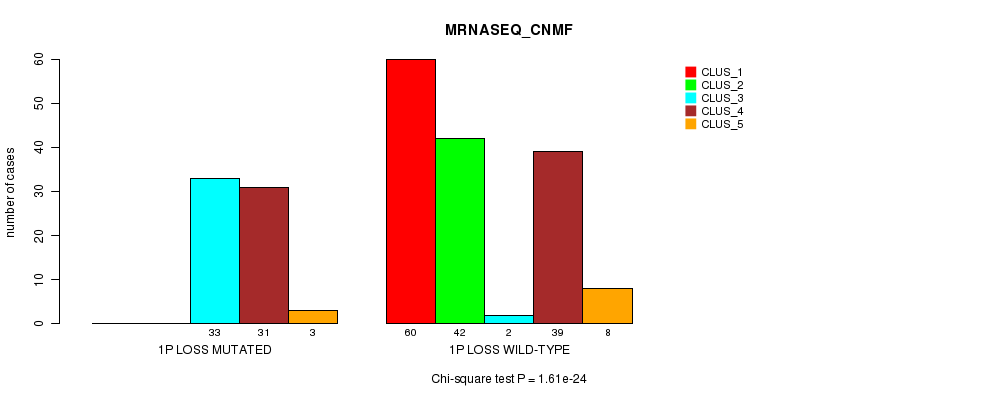
P value = 1.37e-30 (Fisher's exact test), Q value = 4.7e-28
Table S22. Gene #21: '1p loss mutation analysis' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 32 | 117 | 69 |
| 1P LOSS MUTATED | 32 | 4 | 31 |
| 1P LOSS WILD-TYPE | 0 | 113 | 38 |
Figure S22. Get High-res Image Gene #21: '1p loss mutation analysis' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
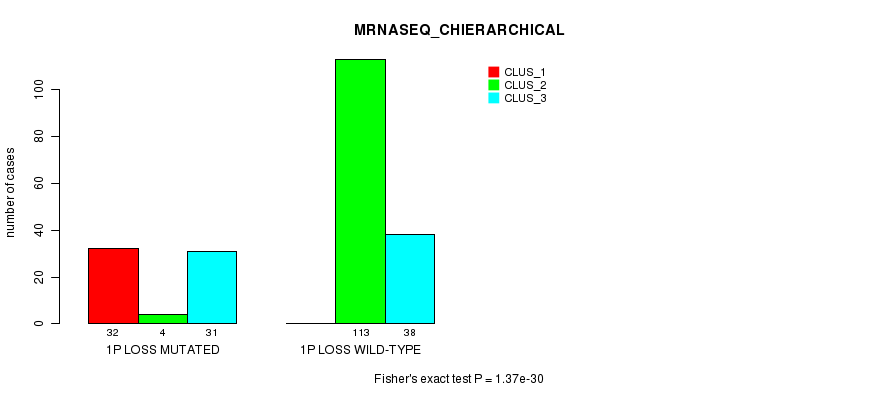
P value = 1.41e-19 (Fisher's exact test), Q value = 4.7e-17
Table S23. Gene #21: '1p loss mutation analysis' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 50 | 50 | 32 | 87 |
| 1P LOSS MUTATED | 3 | 0 | 25 | 39 |
| 1P LOSS WILD-TYPE | 47 | 50 | 7 | 48 |
Figure S23. Get High-res Image Gene #21: '1p loss mutation analysis' versus Clinical Feature #7: 'MIRSEQ_CNMF'
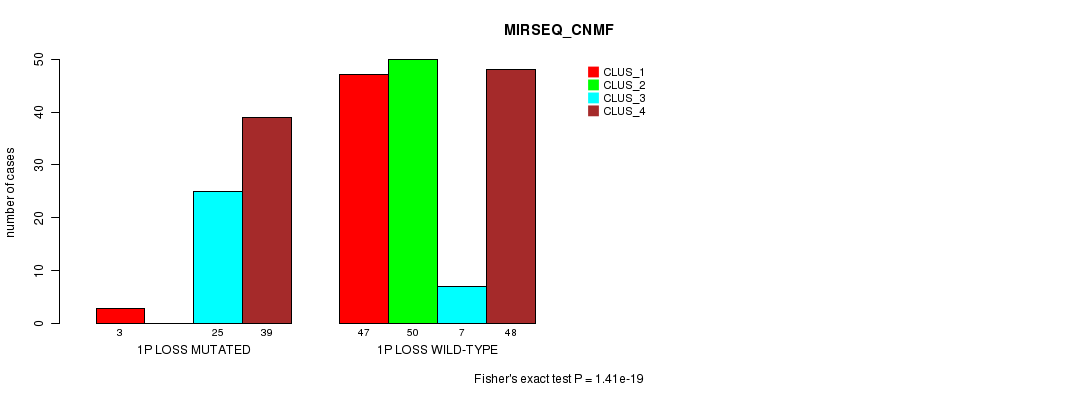
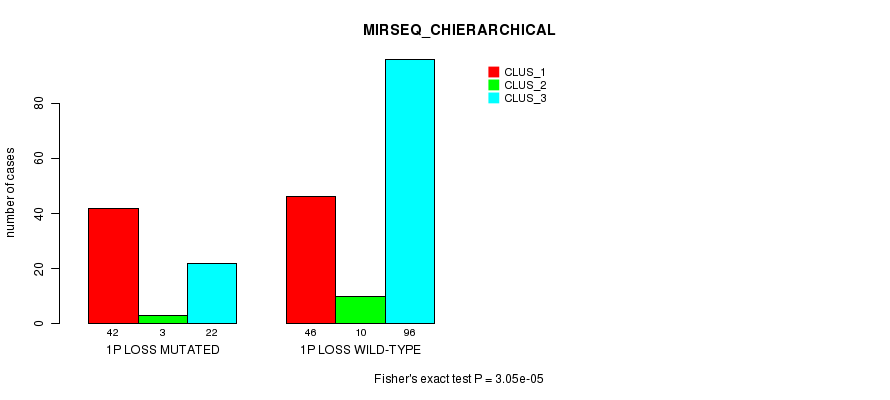
P value = 3.05e-05 (Fisher's exact test), Q value = 0.0095
Table S24. Gene #21: '1p loss mutation analysis' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 88 | 13 | 118 |
| 1P LOSS MUTATED | 42 | 3 | 22 |
| 1P LOSS WILD-TYPE | 46 | 10 | 96 |
Figure S24. Get High-res Image Gene #21: '1p loss mutation analysis' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
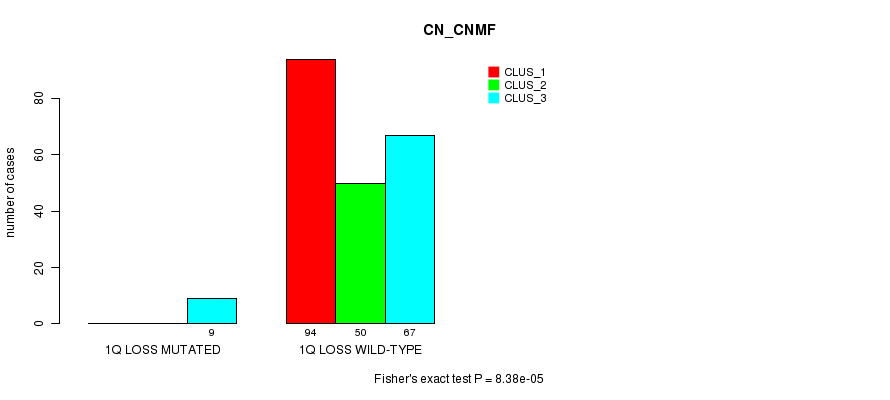
P value = 8.38e-05 (Fisher's exact test), Q value = 0.026
Table S25. Gene #22: '1q loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 1Q LOSS MUTATED | 0 | 0 | 9 |
| 1Q LOSS WILD-TYPE | 94 | 50 | 67 |
Figure S25. Get High-res Image Gene #22: '1q loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
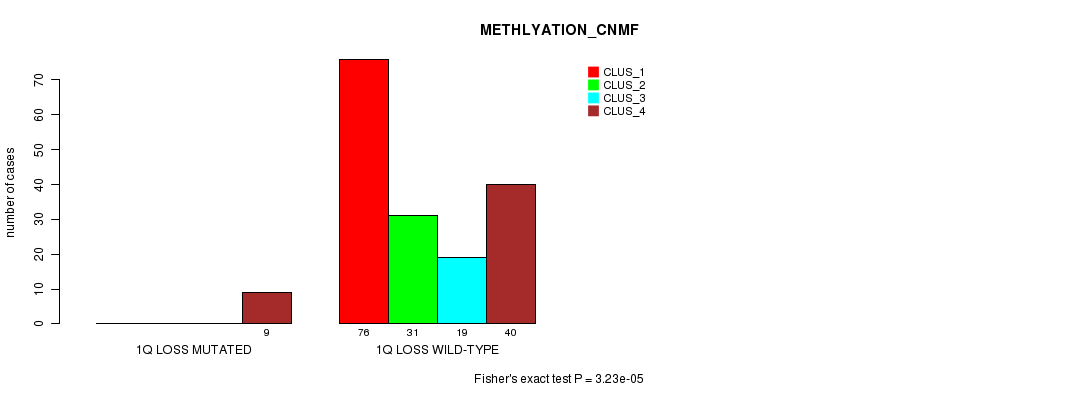
P value = 3.23e-05 (Fisher's exact test), Q value = 0.01
Table S26. Gene #22: '1q loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 1Q LOSS MUTATED | 0 | 0 | 0 | 9 |
| 1Q LOSS WILD-TYPE | 76 | 31 | 19 | 40 |
Figure S26. Get High-res Image Gene #22: '1q loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
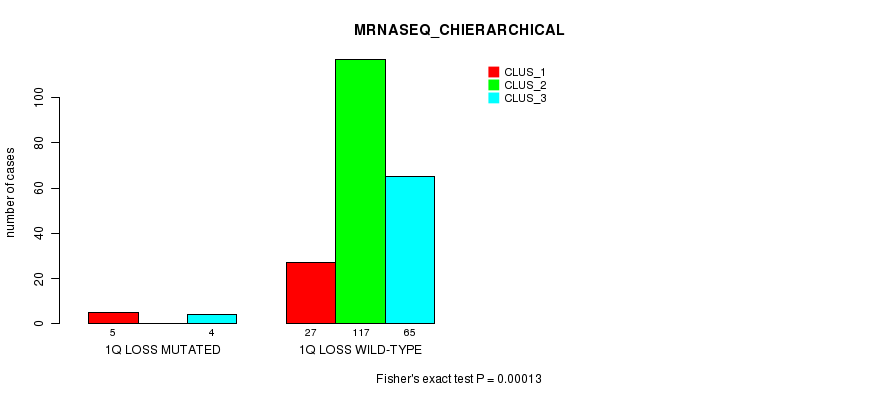
P value = 0.00013 (Fisher's exact test), Q value = 0.04
Table S27. Gene #22: '1q loss mutation analysis' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 32 | 117 | 69 |
| 1Q LOSS MUTATED | 5 | 0 | 4 |
| 1Q LOSS WILD-TYPE | 27 | 117 | 65 |
Figure S27. Get High-res Image Gene #22: '1q loss mutation analysis' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
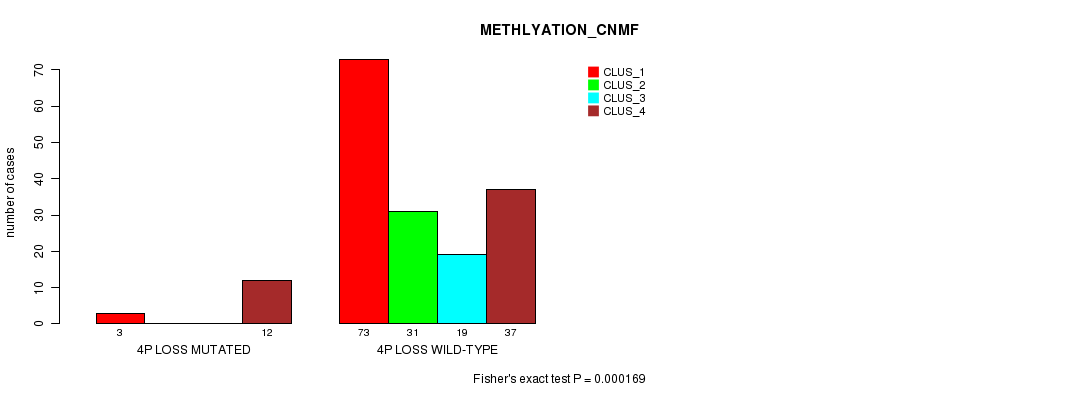
P value = 0.000169 (Fisher's exact test), Q value = 0.051
Table S28. Gene #27: '4p loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 4P LOSS MUTATED | 3 | 0 | 0 | 12 |
| 4P LOSS WILD-TYPE | 73 | 31 | 19 | 37 |
Figure S28. Get High-res Image Gene #27: '4p loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
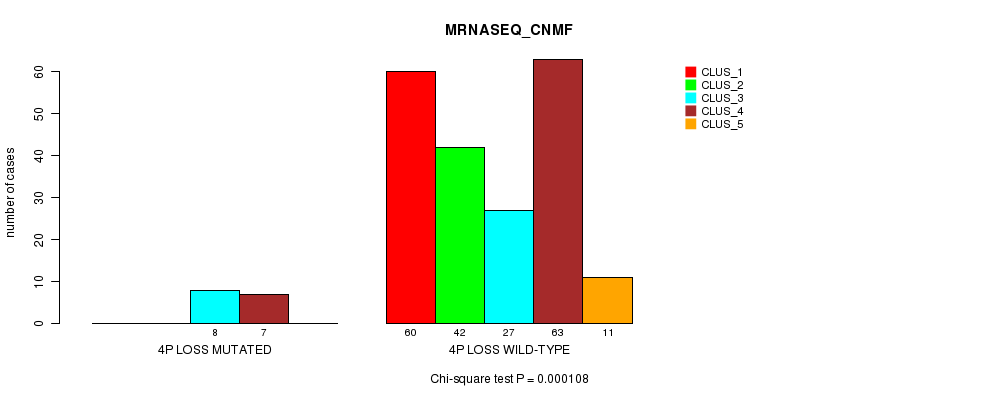
P value = 0.000108 (Chi-square test), Q value = 0.033
Table S29. Gene #27: '4p loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 4P LOSS MUTATED | 0 | 0 | 8 | 7 | 0 |
| 4P LOSS WILD-TYPE | 60 | 42 | 27 | 63 | 11 |
Figure S29. Get High-res Image Gene #27: '4p loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
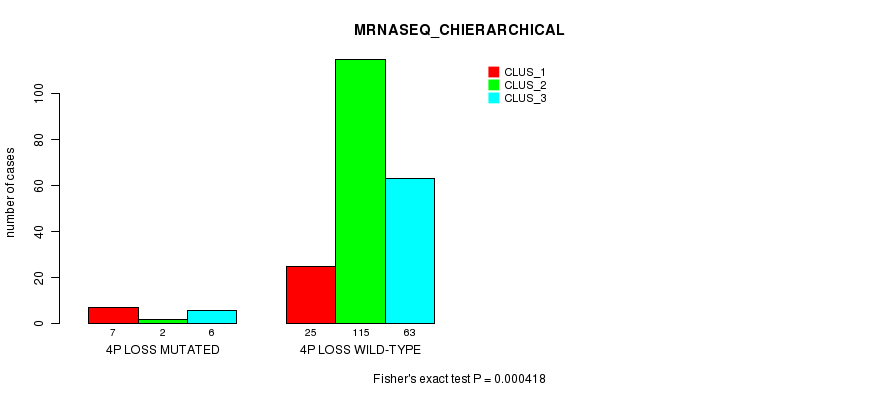
P value = 0.000418 (Fisher's exact test), Q value = 0.13
Table S30. Gene #27: '4p loss mutation analysis' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 32 | 117 | 69 |
| 4P LOSS MUTATED | 7 | 2 | 6 |
| 4P LOSS WILD-TYPE | 25 | 115 | 63 |
Figure S30. Get High-res Image Gene #27: '4p loss mutation analysis' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
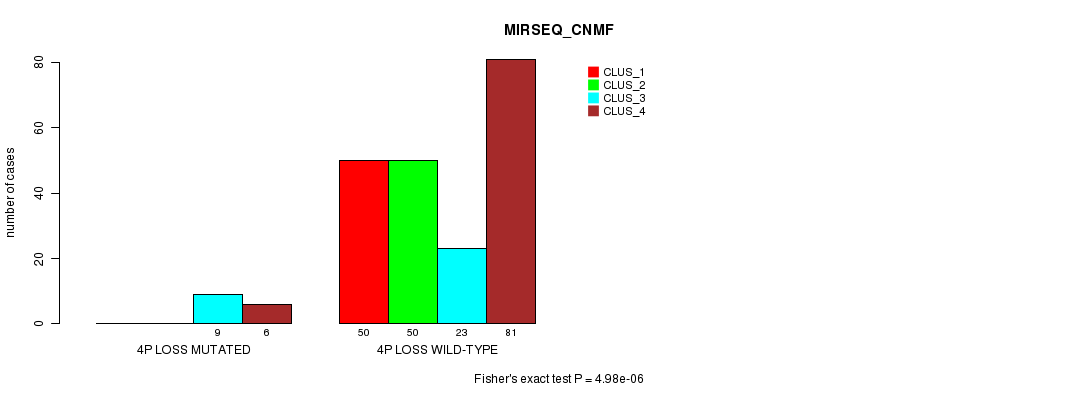
P value = 4.98e-06 (Fisher's exact test), Q value = 0.0016
Table S31. Gene #27: '4p loss mutation analysis' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 50 | 50 | 32 | 87 |
| 4P LOSS MUTATED | 0 | 0 | 9 | 6 |
| 4P LOSS WILD-TYPE | 50 | 50 | 23 | 81 |
Figure S31. Get High-res Image Gene #27: '4p loss mutation analysis' versus Clinical Feature #7: 'MIRSEQ_CNMF'
P value = 0.000613 (Fisher's exact test), Q value = 0.18
Table S32. Gene #28: '4q loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 4Q LOSS MUTATED | 8 | 1 | 0 | 15 |
| 4Q LOSS WILD-TYPE | 68 | 30 | 19 | 34 |
Figure S32. Get High-res Image Gene #28: '4q loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
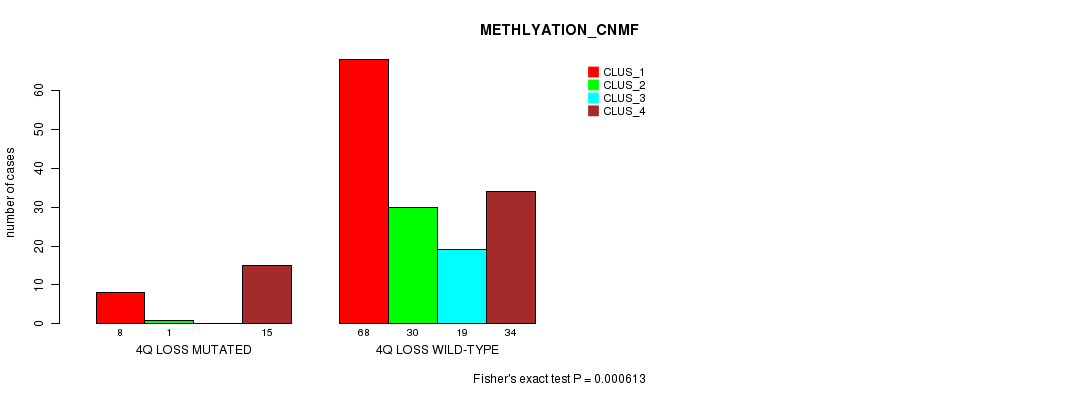
P value = 6.65e-05 (Fisher's exact test), Q value = 0.02
Table S33. Gene #28: '4q loss mutation analysis' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 50 | 50 | 32 | 87 |
| 4Q LOSS MUTATED | 1 | 2 | 11 | 10 |
| 4Q LOSS WILD-TYPE | 49 | 48 | 21 | 77 |
Figure S33. Get High-res Image Gene #28: '4q loss mutation analysis' versus Clinical Feature #7: 'MIRSEQ_CNMF'
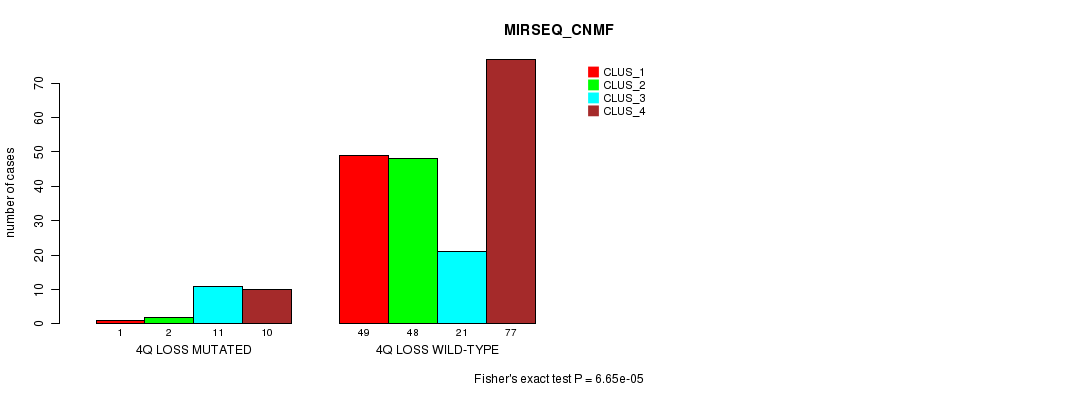
P value = 2.3e-17 (Fisher's exact test), Q value = 7.6e-15
Table S34. Gene #36: '10p loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 10P LOSS MUTATED | 1 | 27 | 3 |
| 10P LOSS WILD-TYPE | 93 | 23 | 73 |
Figure S34. Get High-res Image Gene #36: '10p loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
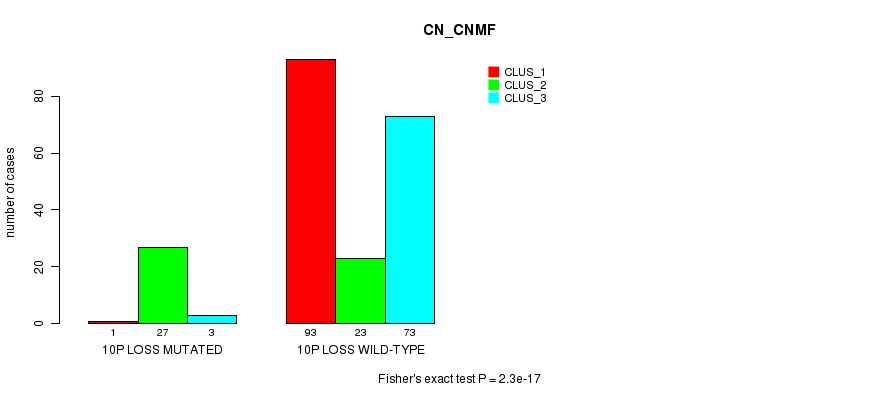
P value = 7.92e-17 (Fisher's exact test), Q value = 2.6e-14
Table S35. Gene #36: '10p loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 10P LOSS MUTATED | 1 | 22 | 0 | 3 |
| 10P LOSS WILD-TYPE | 75 | 9 | 19 | 46 |
Figure S35. Get High-res Image Gene #36: '10p loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
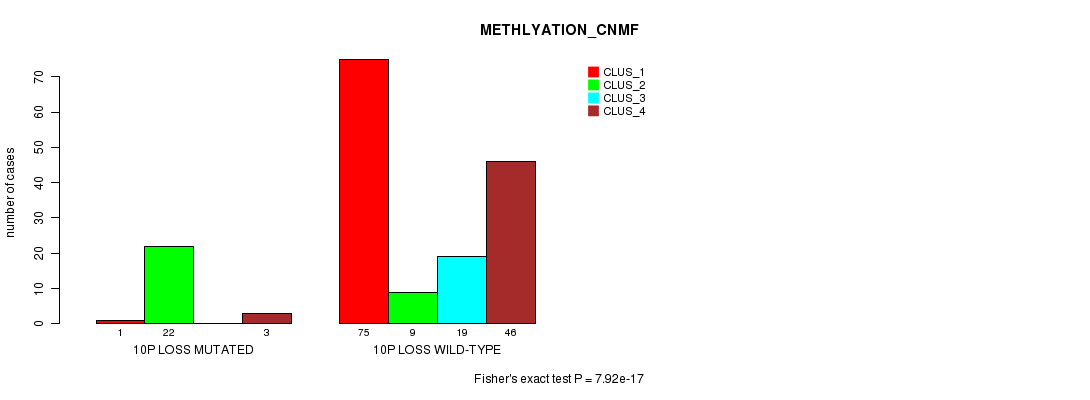
P value = 4.79e-19 (Chi-square test), Q value = 1.6e-16
Table S36. Gene #36: '10p loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 10P LOSS MUTATED | 1 | 25 | 1 | 3 | 0 |
| 10P LOSS WILD-TYPE | 59 | 17 | 34 | 67 | 11 |
Figure S36. Get High-res Image Gene #36: '10p loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
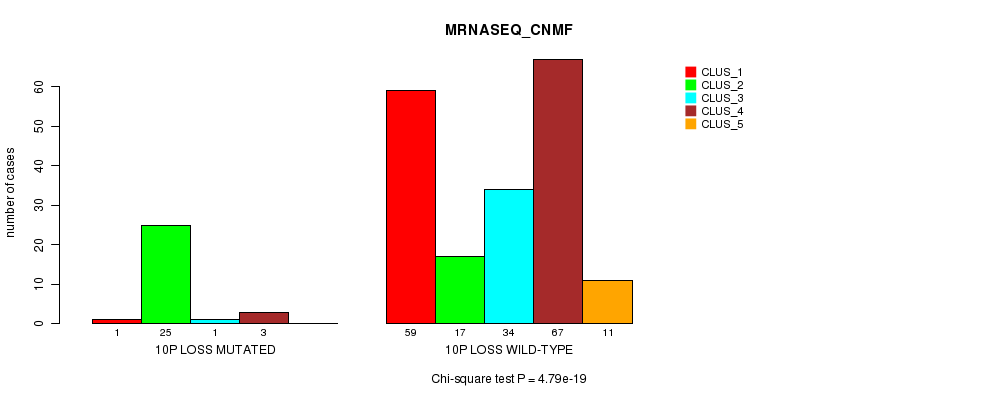
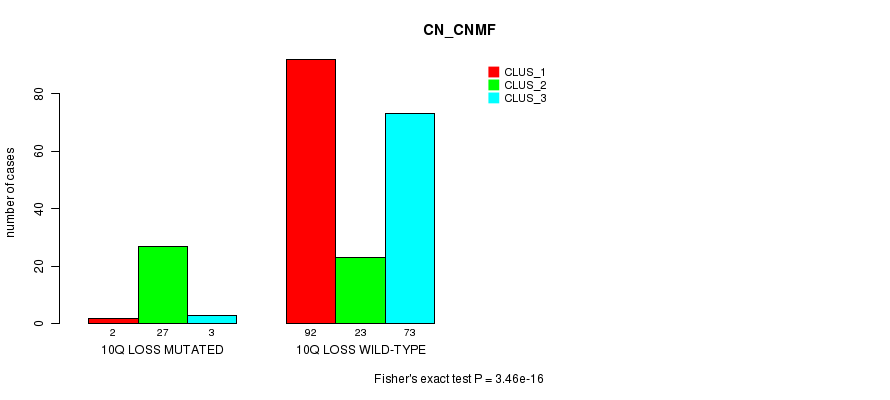
P value = 3.46e-16 (Fisher's exact test), Q value = 1.1e-13
Table S37. Gene #37: '10q loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 10Q LOSS MUTATED | 2 | 27 | 3 |
| 10Q LOSS WILD-TYPE | 92 | 23 | 73 |
Figure S37. Get High-res Image Gene #37: '10q loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
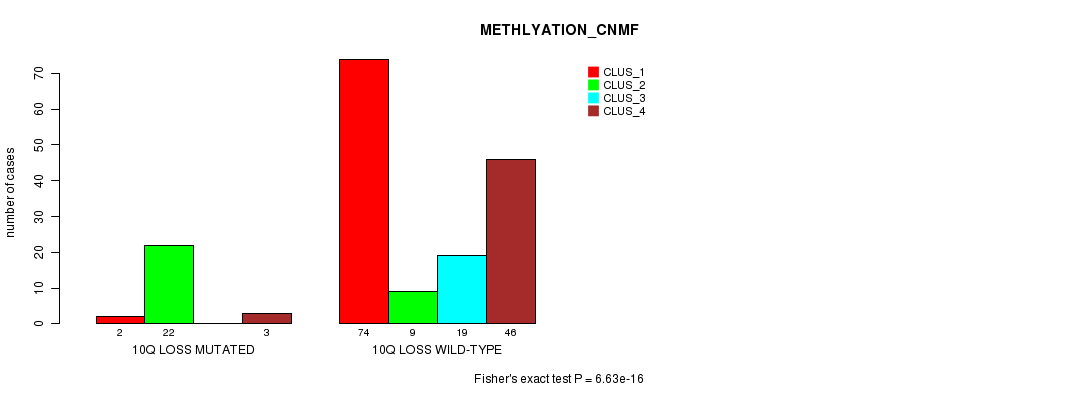
P value = 6.63e-16 (Fisher's exact test), Q value = 2.2e-13
Table S38. Gene #37: '10q loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 10Q LOSS MUTATED | 2 | 22 | 0 | 3 |
| 10Q LOSS WILD-TYPE | 74 | 9 | 19 | 46 |
Figure S38. Get High-res Image Gene #37: '10q loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
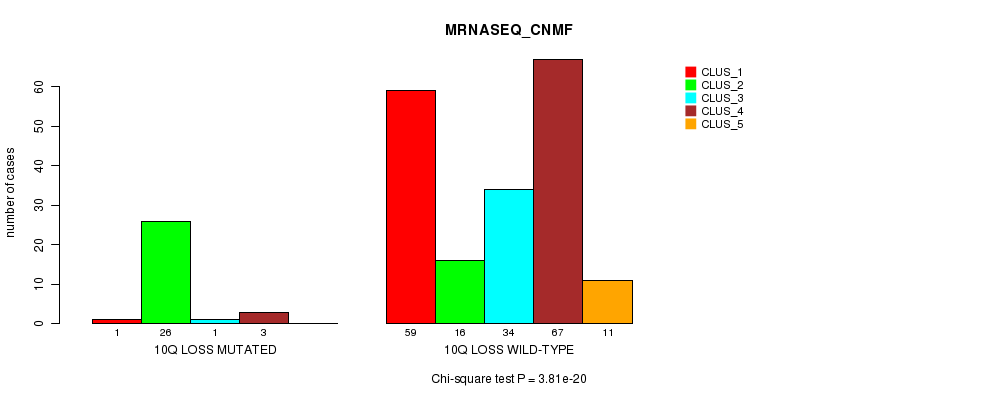
P value = 3.81e-20 (Chi-square test), Q value = 1.3e-17
Table S39. Gene #37: '10q loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 10Q LOSS MUTATED | 1 | 26 | 1 | 3 | 0 |
| 10Q LOSS WILD-TYPE | 59 | 16 | 34 | 67 | 11 |
Figure S39. Get High-res Image Gene #37: '10q loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
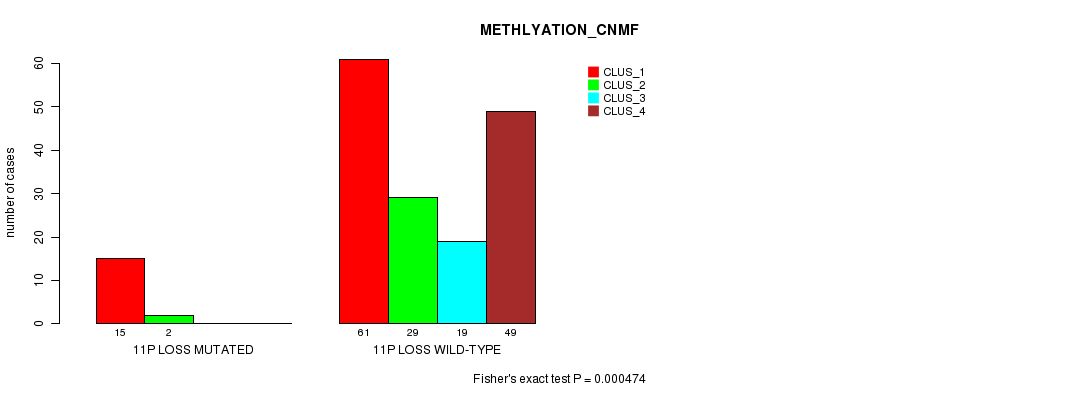
P value = 0.000474 (Fisher's exact test), Q value = 0.14
Table S40. Gene #38: '11p loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 11P LOSS MUTATED | 15 | 2 | 0 | 0 |
| 11P LOSS WILD-TYPE | 61 | 29 | 19 | 49 |
Figure S40. Get High-res Image Gene #38: '11p loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
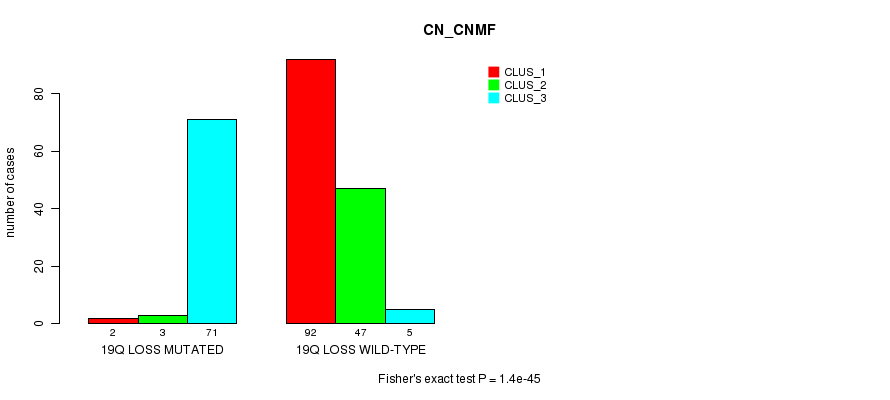
P value = 1.4e-45 (Fisher's exact test), Q value = 4.8e-43
Table S41. Gene #48: '19q loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 19Q LOSS MUTATED | 2 | 3 | 71 |
| 19Q LOSS WILD-TYPE | 92 | 47 | 5 |
Figure S41. Get High-res Image Gene #48: '19q loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
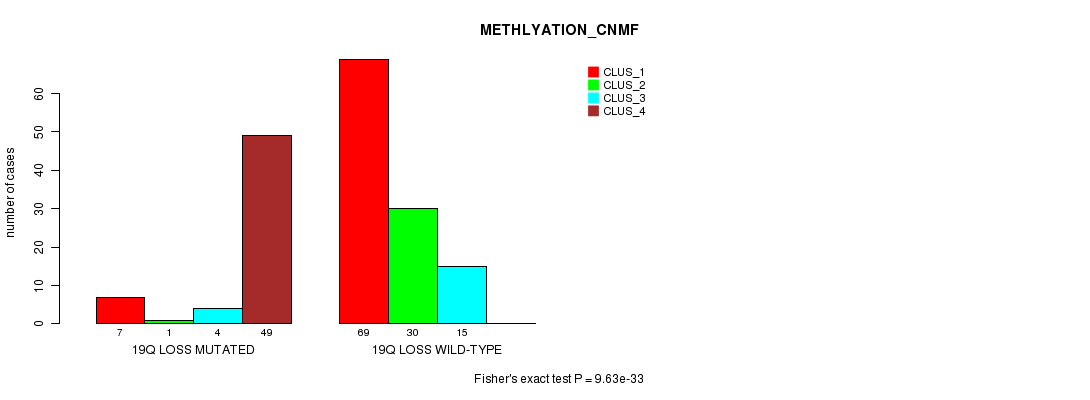
P value = 9.63e-33 (Fisher's exact test), Q value = 3.3e-30
Table S42. Gene #48: '19q loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 19Q LOSS MUTATED | 7 | 1 | 4 | 49 |
| 19Q LOSS WILD-TYPE | 69 | 30 | 15 | 0 |
Figure S42. Get High-res Image Gene #48: '19q loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
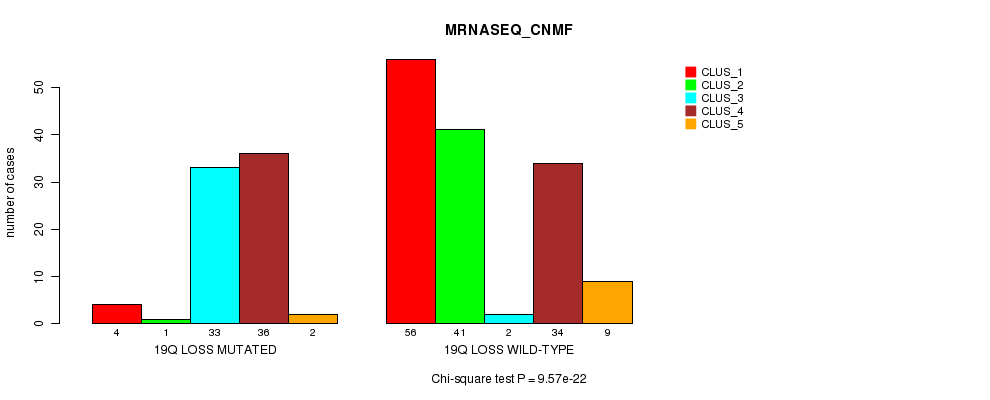
P value = 9.57e-22 (Chi-square test), Q value = 3.2e-19
Table S43. Gene #48: '19q loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 19Q LOSS MUTATED | 4 | 1 | 33 | 36 | 2 |
| 19Q LOSS WILD-TYPE | 56 | 41 | 2 | 34 | 9 |
Figure S43. Get High-res Image Gene #48: '19q loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
P value = 4.5e-27 (Fisher's exact test), Q value = 1.5e-24
Table S44. Gene #48: '19q loss mutation analysis' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 32 | 117 | 69 |
| 19Q LOSS MUTATED | 32 | 9 | 35 |
| 19Q LOSS WILD-TYPE | 0 | 108 | 34 |
Figure S44. Get High-res Image Gene #48: '19q loss mutation analysis' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'

P value = 3.01e-17 (Fisher's exact test), Q value = 9.9e-15
Table S45. Gene #48: '19q loss mutation analysis' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 50 | 50 | 32 | 87 |
| 19Q LOSS MUTATED | 4 | 3 | 26 | 43 |
| 19Q LOSS WILD-TYPE | 46 | 47 | 6 | 44 |
Figure S45. Get High-res Image Gene #48: '19q loss mutation analysis' versus Clinical Feature #7: 'MIRSEQ_CNMF'

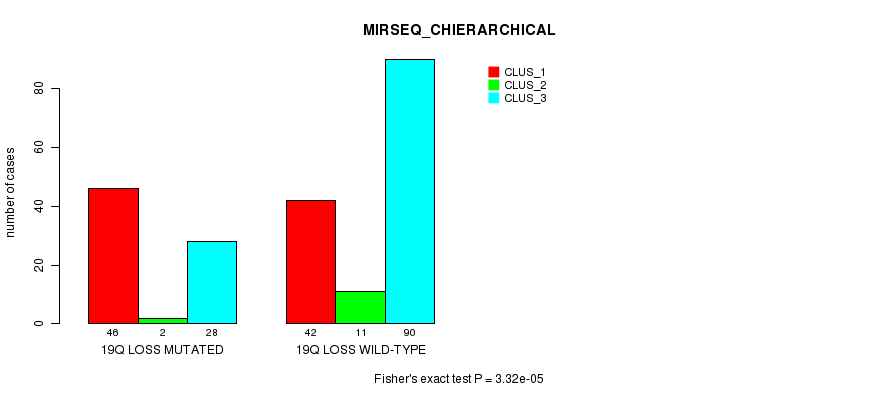
P value = 3.32e-05 (Fisher's exact test), Q value = 0.01
Table S46. Gene #48: '19q loss mutation analysis' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 88 | 13 | 118 |
| 19Q LOSS MUTATED | 46 | 2 | 28 |
| 19Q LOSS WILD-TYPE | 42 | 11 | 90 |
Figure S46. Get High-res Image Gene #48: '19q loss mutation analysis' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
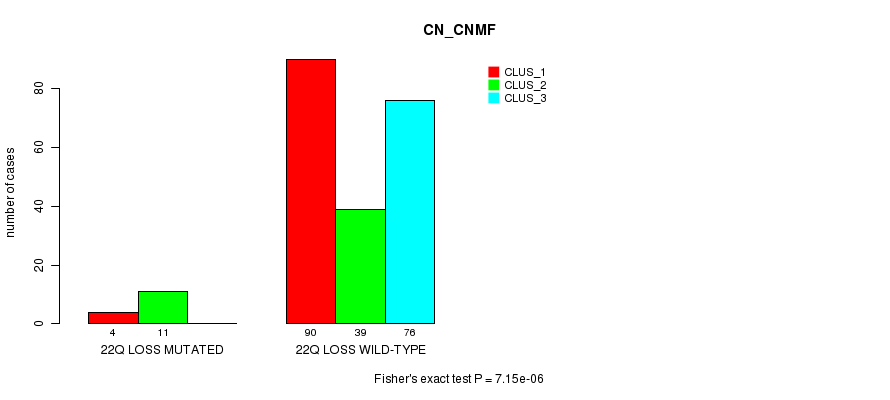
P value = 7.15e-06 (Fisher's exact test), Q value = 0.0023
Table S47. Gene #50: '22q loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 50 | 76 |
| 22Q LOSS MUTATED | 4 | 11 | 0 |
| 22Q LOSS WILD-TYPE | 90 | 39 | 76 |
Figure S47. Get High-res Image Gene #50: '22q loss mutation analysis' versus Clinical Feature #3: 'CN_CNMF'
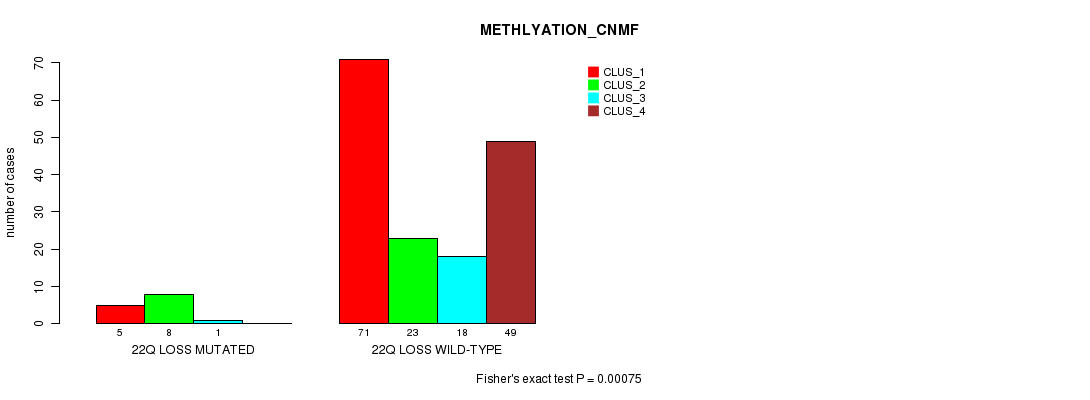
P value = 0.00075 (Fisher's exact test), Q value = 0.22
Table S48. Gene #50: '22q loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 76 | 31 | 19 | 49 |
| 22Q LOSS MUTATED | 5 | 8 | 1 | 0 |
| 22Q LOSS WILD-TYPE | 71 | 23 | 18 | 49 |
Figure S48. Get High-res Image Gene #50: '22q loss mutation analysis' versus Clinical Feature #4: 'METHLYATION_CNMF'
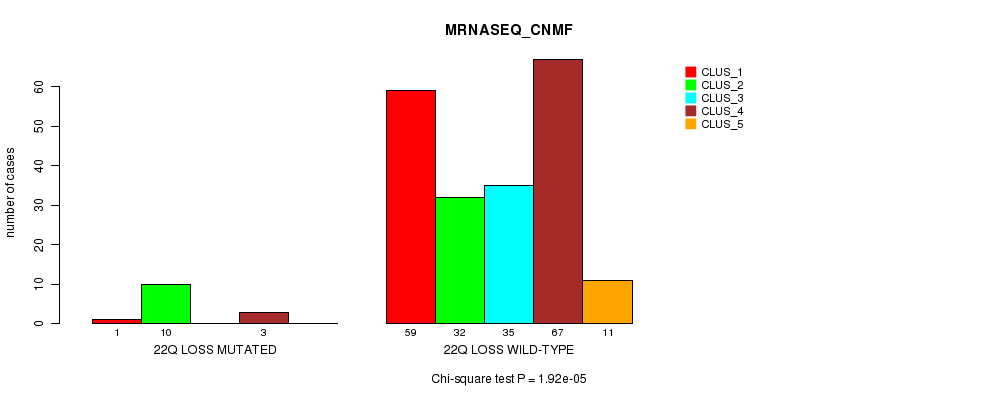
P value = 1.92e-05 (Chi-square test), Q value = 0.006
Table S49. Gene #50: '22q loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 60 | 42 | 35 | 70 | 11 |
| 22Q LOSS MUTATED | 1 | 10 | 0 | 3 | 0 |
| 22Q LOSS WILD-TYPE | 59 | 32 | 35 | 67 | 11 |
Figure S49. Get High-res Image Gene #50: '22q loss mutation analysis' versus Clinical Feature #5: 'MRNASEQ_CNMF'
-
Mutation data file = broad_values_by_arm.mutsig.cluster.txt
-
Molecular subtypes file = LGG-TP.transferedmergedcluster.txt
-
Number of patients = 220
-
Number of significantly arm-level cnvs = 51
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.