(primary solid tumor cohort)
This pipeline computes the correlation between significantly recurrent gene mutations and molecular subtypes.
Testing the association between mutation status of 94 genes and 8 molecular subtypes across 306 patients, 12 significant findings detected with P value < 0.05 and Q value < 0.25.
-
NOTCH1 mutation correlated to 'MRNASEQ_CHIERARCHICAL' and 'MIRSEQ_CNMF'.
-
HRAS mutation correlated to 'CN_CNMF'.
-
NFE2L2 mutation correlated to 'METHLYATION_CNMF'.
-
TP53 mutation correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
CASP8 mutation correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
NSD1 mutation correlated to 'METHLYATION_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
Table 1. Get Full Table Overview of the association between mutation status of 94 genes and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 12 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
RPPA CNMF |
RPPA CHIERARCHICAL |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
||
| nMutated (%) | nWild-Type | Chi-square test | Chi-square test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| CASP8 | 27 (9%) | 279 |
0.000326 (0.236) |
7.22e-06 (0.00525) |
0.699 (1.00) |
0.739 (1.00) |
3e-07 (0.000219) |
5.28e-07 (0.000385) |
0.329 (1.00) |
0.286 (1.00) |
| NOTCH1 | 57 (19%) | 249 |
0.488 (1.00) |
0.000746 (0.536) |
0.0187 (1.00) |
0.279 (1.00) |
0.000428 (0.308) |
2.54e-06 (0.00185) |
7.73e-05 (0.056) |
0.572 (1.00) |
| TP53 | 214 (70%) | 92 |
1.84e-16 (1.35e-13) |
3.14e-18 (2.3e-15) |
0.11 (1.00) |
0.776 (1.00) |
0.0113 (1.00) |
0.00143 (1.00) |
0.0426 (1.00) |
0.000589 (0.424) |
| NSD1 | 33 (11%) | 273 |
0.377 (1.00) |
3.04e-23 (2.23e-20) |
0.244 (1.00) |
0.73 (1.00) |
0.000411 (0.297) |
6.41e-06 (0.00466) |
0.0133 (1.00) |
0.0339 (1.00) |
| HRAS | 10 (3%) | 296 |
1.33e-05 (0.00968) |
0.0612 (1.00) |
0.168 (1.00) |
0.599 (1.00) |
0.00339 (1.00) |
0.0011 (0.785) |
0.109 (1.00) |
0.504 (1.00) |
| NFE2L2 | 17 (6%) | 289 |
0.909 (1.00) |
0.000297 (0.215) |
0.125 (1.00) |
0.765 (1.00) |
1 (1.00) |
0.464 (1.00) |
0.223 (1.00) |
0.11 (1.00) |
| CDKN2A | 65 (21%) | 241 |
0.0451 (1.00) |
0.00306 (1.00) |
0.0349 (1.00) |
0.583 (1.00) |
0.201 (1.00) |
0.206 (1.00) |
0.354 (1.00) |
0.115 (1.00) |
| PIK3CA | 64 (21%) | 242 |
0.0123 (1.00) |
0.0554 (1.00) |
0.0948 (1.00) |
0.928 (1.00) |
0.119 (1.00) |
0.398 (1.00) |
0.0234 (1.00) |
0.152 (1.00) |
| FAT1 | 72 (24%) | 234 |
0.308 (1.00) |
0.227 (1.00) |
0.794 (1.00) |
0.487 (1.00) |
0.152 (1.00) |
0.288 (1.00) |
0.82 (1.00) |
0.252 (1.00) |
| JUB | 18 (6%) | 288 |
0.264 (1.00) |
0.0799 (1.00) |
0.0646 (1.00) |
0.0816 (1.00) |
0.305 (1.00) |
0.221 (1.00) |
0.38 (1.00) |
1 (1.00) |
| MLL2 | 56 (18%) | 250 |
0.118 (1.00) |
0.0471 (1.00) |
0.687 (1.00) |
0.521 (1.00) |
0.0532 (1.00) |
0.00795 (1.00) |
0.0707 (1.00) |
0.00651 (1.00) |
| FBXW7 | 15 (5%) | 291 |
0.21 (1.00) |
0.134 (1.00) |
0.092 (1.00) |
0.303 (1.00) |
0.881 (1.00) |
0.636 (1.00) |
0.945 (1.00) |
0.868 (1.00) |
| BAGE2 | 11 (4%) | 295 |
0.279 (1.00) |
0.0999 (1.00) |
0.414 (1.00) |
0.0857 (1.00) |
0.253 (1.00) |
0.291 (1.00) |
0.178 (1.00) |
0.101 (1.00) |
| POTEC | 15 (5%) | 291 |
0.0719 (1.00) |
0.0364 (1.00) |
0.847 (1.00) |
1 (1.00) |
0.17 (1.00) |
0.254 (1.00) |
0.535 (1.00) |
0.21 (1.00) |
| ZNF750 | 13 (4%) | 293 |
0.721 (1.00) |
0.159 (1.00) |
0.946 (1.00) |
0.353 (1.00) |
0.119 (1.00) |
0.128 (1.00) |
0.632 (1.00) |
1 (1.00) |
| EP300 | 25 (8%) | 281 |
0.0345 (1.00) |
0.151 (1.00) |
0.769 (1.00) |
0.264 (1.00) |
0.00955 (1.00) |
0.00467 (1.00) |
0.338 (1.00) |
0.526 (1.00) |
| POM121L12 | 14 (5%) | 292 |
0.0578 (1.00) |
0.304 (1.00) |
0.116 (1.00) |
0.151 (1.00) |
0.658 (1.00) |
0.657 (1.00) |
0.48 (1.00) |
0.454 (1.00) |
| RHOA | 4 (1%) | 302 |
0.0826 (1.00) |
0.54 (1.00) |
0.379 (1.00) |
0.444 (1.00) |
0.245 (1.00) |
0.141 (1.00) |
0.24 (1.00) |
0.254 (1.00) |
| B2M | 7 (2%) | 299 |
0.767 (1.00) |
0.0618 (1.00) |
0.515 (1.00) |
0.517 (1.00) |
0.141 (1.00) |
0.767 (1.00) |
||
| RAC1 | 9 (3%) | 297 |
0.374 (1.00) |
0.585 (1.00) |
1 (1.00) |
0.519 (1.00) |
0.195 (1.00) |
0.192 (1.00) |
0.268 (1.00) |
0.301 (1.00) |
| ZNF804B | 21 (7%) | 285 |
0.825 (1.00) |
0.487 (1.00) |
0.626 (1.00) |
0.514 (1.00) |
0.812 (1.00) |
0.746 (1.00) |
0.92 (1.00) |
0.904 (1.00) |
| KCNT2 | 17 (6%) | 289 |
0.197 (1.00) |
0.235 (1.00) |
0.702 (1.00) |
0.562 (1.00) |
0.05 (1.00) |
0.121 (1.00) |
0.052 (1.00) |
0.0388 (1.00) |
| RASA1 | 14 (5%) | 292 |
0.0617 (1.00) |
0.24 (1.00) |
1 (1.00) |
0.402 (1.00) |
0.357 (1.00) |
0.271 (1.00) |
0.313 (1.00) |
0.0301 (1.00) |
| PRIM2 | 9 (3%) | 297 |
0.0807 (1.00) |
0.305 (1.00) |
0.822 (1.00) |
0.404 (1.00) |
0.4 (1.00) |
0.395 (1.00) |
0.268 (1.00) |
0.301 (1.00) |
| CDH10 | 23 (8%) | 283 |
0.326 (1.00) |
0.531 (1.00) |
0.329 (1.00) |
0.695 (1.00) |
0.0743 (1.00) |
0.265 (1.00) |
0.207 (1.00) |
0.694 (1.00) |
| CNTNAP5 | 15 (5%) | 291 |
0.0449 (1.00) |
0.115 (1.00) |
0.553 (1.00) |
1 (1.00) |
0.0101 (1.00) |
0.0354 (1.00) |
0.0101 (1.00) |
0.3 (1.00) |
| POTEG | 10 (3%) | 296 |
0.00148 (1.00) |
0.0869 (1.00) |
0.229 (1.00) |
0.754 (1.00) |
0.16 (1.00) |
0.143 (1.00) |
0.728 (1.00) |
0.504 (1.00) |
| ZNF99 | 22 (7%) | 284 |
0.229 (1.00) |
0.126 (1.00) |
0.322 (1.00) |
0.402 (1.00) |
0.0343 (1.00) |
0.0116 (1.00) |
0.00606 (1.00) |
0.00653 (1.00) |
| OR2T12 | 11 (4%) | 295 |
0.0236 (1.00) |
0.12 (1.00) |
0.512 (1.00) |
0.599 (1.00) |
0.552 (1.00) |
0.502 (1.00) |
0.593 (1.00) |
0.672 (1.00) |
| REG1A | 8 (3%) | 298 |
0.168 (1.00) |
0.102 (1.00) |
0.496 (1.00) |
0.36 (1.00) |
0.058 (1.00) |
0.00131 (0.938) |
0.0569 (1.00) |
0.0617 (1.00) |
| PABPC5 | 9 (3%) | 297 |
0.207 (1.00) |
0.407 (1.00) |
0.692 (1.00) |
0.779 (1.00) |
0.089 (1.00) |
0.237 (1.00) |
0.268 (1.00) |
1 (1.00) |
| IL32 | 4 (1%) | 302 |
0.116 (1.00) |
0.00177 (1.00) |
0.561 (1.00) |
1 (1.00) |
0.563 (1.00) |
0.68 (1.00) |
||
| PRAMEF11 | 10 (3%) | 296 |
0.0932 (1.00) |
0.438 (1.00) |
0.963 (1.00) |
0.777 (1.00) |
0.669 (1.00) |
0.606 (1.00) |
0.551 (1.00) |
1 (1.00) |
| EPHA2 | 14 (5%) | 292 |
0.0127 (1.00) |
0.00256 (1.00) |
0.811 (1.00) |
0.906 (1.00) |
0.499 (1.00) |
0.57 (1.00) |
0.774 (1.00) |
0.071 (1.00) |
| FCRL4 | 13 (4%) | 293 |
0.12 (1.00) |
0.0179 (1.00) |
0.788 (1.00) |
1 (1.00) |
0.405 (1.00) |
0.106 (1.00) |
0.46 (1.00) |
0.847 (1.00) |
| LCP1 | 12 (4%) | 294 |
0.463 (1.00) |
0.000851 (0.61) |
0.667 (1.00) |
0.0436 (1.00) |
0.873 (1.00) |
0.758 (1.00) |
0.498 (1.00) |
0.842 (1.00) |
| LRFN5 | 13 (4%) | 293 |
0.00104 (0.748) |
0.07 (1.00) |
0.766 (1.00) |
0.865 (1.00) |
0.279 (1.00) |
0.0765 (1.00) |
0.0196 (1.00) |
0.424 (1.00) |
| MAPK1 | 4 (1%) | 302 |
0.725 (1.00) |
0.924 (1.00) |
0.569 (1.00) |
1 (1.00) |
0.691 (1.00) |
0.686 (1.00) |
0.691 (1.00) |
0.132 (1.00) |
| PEG3 | 23 (8%) | 283 |
0.172 (1.00) |
0.022 (1.00) |
0.0827 (1.00) |
0.103 (1.00) |
0.0217 (1.00) |
0.053 (1.00) |
0.253 (1.00) |
0.241 (1.00) |
| PRSS1 | 8 (3%) | 298 |
0.014 (1.00) |
0.478 (1.00) |
0.0589 (1.00) |
1 (1.00) |
1 (1.00) |
0.89 (1.00) |
0.82 (1.00) |
0.205 (1.00) |
| DOK6 | 8 (3%) | 298 |
0.375 (1.00) |
0.582 (1.00) |
0.705 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.666 (1.00) |
1 (1.00) |
| GABRB3 | 11 (4%) | 295 |
0.222 (1.00) |
0.0147 (1.00) |
0.148 (1.00) |
0.541 (1.00) |
0.253 (1.00) |
0.192 (1.00) |
0.593 (1.00) |
0.672 (1.00) |
| STEAP4 | 10 (3%) | 296 |
0.174 (1.00) |
0.636 (1.00) |
0.175 (1.00) |
0.656 (1.00) |
0.0813 (1.00) |
0.0997 (1.00) |
0.0893 (1.00) |
1 (1.00) |
| ASXL3 | 21 (7%) | 285 |
0.618 (1.00) |
0.632 (1.00) |
0.225 (1.00) |
0.356 (1.00) |
0.96 (1.00) |
0.714 (1.00) |
0.501 (1.00) |
0.383 (1.00) |
| NTM | 7 (2%) | 299 |
0.238 (1.00) |
0.24 (1.00) |
0.0949 (1.00) |
0.0238 (1.00) |
0.182 (1.00) |
0.0877 (1.00) |
0.104 (1.00) |
0.0893 (1.00) |
| FAM155A | 11 (4%) | 295 |
0.151 (1.00) |
0.0585 (1.00) |
0.0106 (1.00) |
0.0694 (1.00) |
0.233 (1.00) |
0.175 (1.00) |
0.119 (1.00) |
0.0667 (1.00) |
| HLA-A | 9 (3%) | 297 |
0.528 (1.00) |
0.335 (1.00) |
1 (1.00) |
1 (1.00) |
0.072 (1.00) |
0.0311 (1.00) |
0.633 (1.00) |
0.612 (1.00) |
| KRTAP1-5 | 3 (1%) | 303 |
0.826 (1.00) |
0.807 (1.00) |
0.78 (1.00) |
0.63 (1.00) |
0.0245 (1.00) |
1 (1.00) |
||
| CTCF | 11 (4%) | 295 |
0.551 (1.00) |
0.401 (1.00) |
0.393 (1.00) |
0.541 (1.00) |
0.636 (1.00) |
0.632 (1.00) |
0.86 (1.00) |
0.366 (1.00) |
| BRWD3 | 16 (5%) | 290 |
0.88 (1.00) |
0.532 (1.00) |
0.537 (1.00) |
1 (1.00) |
0.256 (1.00) |
0.136 (1.00) |
0.584 (1.00) |
0.234 (1.00) |
| CNPY3 | 3 (1%) | 303 |
0.598 (1.00) |
0.48 (1.00) |
0.5 (1.00) |
0.499 (1.00) |
0.495 (1.00) |
0.599 (1.00) |
||
| FAM101A | 4 (1%) | 302 |
0.581 (1.00) |
0.374 (1.00) |
0.691 (1.00) |
1 (1.00) |
0.0707 (1.00) |
1 (1.00) |
||
| HLA-B | 9 (3%) | 297 |
0.194 (1.00) |
0.0957 (1.00) |
0.952 (1.00) |
0.599 (1.00) |
0.00341 (1.00) |
0.00425 (1.00) |
0.361 (1.00) |
0.301 (1.00) |
| PSG8 | 9 (3%) | 297 |
0.221 (1.00) |
0.618 (1.00) |
1 (1.00) |
0.71 (1.00) |
0.483 (1.00) |
0.484 (1.00) |
0.918 (1.00) |
0.362 (1.00) |
| OR56A1 | 8 (3%) | 298 |
0.558 (1.00) |
0.0158 (1.00) |
0.894 (1.00) |
0.822 (1.00) |
0.357 (1.00) |
0.352 (1.00) |
0.201 (1.00) |
0.341 (1.00) |
| OR2M2 | 8 (3%) | 298 |
0.005 (1.00) |
0.126 (1.00) |
0.181 (1.00) |
0.0118 (1.00) |
0.605 (1.00) |
0.116 (1.00) |
||
| OR4M2 | 8 (3%) | 298 |
0.373 (1.00) |
0.88 (1.00) |
0.746 (1.00) |
0.519 (1.00) |
0.643 (1.00) |
0.639 (1.00) |
0.605 (1.00) |
0.776 (1.00) |
| CPXCR1 | 7 (2%) | 299 |
0.767 (1.00) |
0.536 (1.00) |
0.202 (1.00) |
0.209 (1.00) |
0.345 (1.00) |
0.498 (1.00) |
0.5 (1.00) |
0.399 (1.00) |
| EYA1 | 10 (3%) | 296 |
0.215 (1.00) |
0.636 (1.00) |
0.79 (1.00) |
0.843 (1.00) |
0.0274 (1.00) |
0.0263 (1.00) |
0.728 (1.00) |
0.215 (1.00) |
| LINGO2 | 10 (3%) | 296 |
0.316 (1.00) |
0.82 (1.00) |
0.284 (1.00) |
1 (1.00) |
0.015 (1.00) |
0.0144 (1.00) |
0.172 (1.00) |
0.138 (1.00) |
| SEMA5A | 19 (6%) | 287 |
0.000737 (0.53) |
0.00318 (1.00) |
0.936 (1.00) |
1 (1.00) |
0.00661 (1.00) |
0.0234 (1.00) |
0.299 (1.00) |
0.0351 (1.00) |
| TGFBR2 | 10 (3%) | 296 |
0.46 (1.00) |
0.201 (1.00) |
0.469 (1.00) |
1 (1.00) |
0.313 (1.00) |
0.785 (1.00) |
0.79 (1.00) |
1 (1.00) |
| PLSCR1 | 5 (2%) | 301 |
0.141 (1.00) |
0.23 (1.00) |
1 (1.00) |
1 (1.00) |
0.524 (1.00) |
0.526 (1.00) |
0.155 (1.00) |
0.463 (1.00) |
| LIN28B | 6 (2%) | 300 |
0.634 (1.00) |
0.938 (1.00) |
0.513 (1.00) |
1 (1.00) |
0.737 (1.00) |
0.739 (1.00) |
0.515 (1.00) |
0.139 (1.00) |
| AGTR1 | 8 (3%) | 298 |
0.398 (1.00) |
0.401 (1.00) |
0.173 (1.00) |
0.108 (1.00) |
0.181 (1.00) |
0.2 (1.00) |
0.0569 (1.00) |
0.776 (1.00) |
| C6 | 12 (4%) | 294 |
0.58 (1.00) |
0.591 (1.00) |
0.062 (1.00) |
1 (1.00) |
0.0762 (1.00) |
0.222 (1.00) |
0.535 (1.00) |
1 (1.00) |
| MS4A14 | 9 (3%) | 297 |
0.238 (1.00) |
0.418 (1.00) |
0.588 (1.00) |
1 (1.00) |
0.4 (1.00) |
0.484 (1.00) |
0.486 (1.00) |
1 (1.00) |
| HIST1H4E | 5 (2%) | 301 |
0.658 (1.00) |
0.731 (1.00) |
0.255 (1.00) |
0.754 (1.00) |
0.866 (1.00) |
0.386 (1.00) |
0.333 (1.00) |
0.278 (1.00) |
| LBP | 7 (2%) | 299 |
0.426 (1.00) |
0.837 (1.00) |
0.775 (1.00) |
0.771 (1.00) |
1 (1.00) |
1 (1.00) |
||
| RGS17 | 5 (2%) | 301 |
0.706 (1.00) |
0.81 (1.00) |
0.157 (1.00) |
0.155 (1.00) |
0.155 (1.00) |
0.278 (1.00) |
||
| LILRB1 | 12 (4%) | 294 |
0.269 (1.00) |
0.166 (1.00) |
0.419 (1.00) |
0.822 (1.00) |
0.861 (1.00) |
0.929 (1.00) |
0.704 (1.00) |
0.462 (1.00) |
| MAGEL2 | 12 (4%) | 294 |
0.228 (1.00) |
0.00566 (1.00) |
0.17 (1.00) |
0.446 (1.00) |
0.538 (1.00) |
0.341 (1.00) |
0.397 (1.00) |
0.573 (1.00) |
| OR8D4 | 6 (2%) | 300 |
0.902 (1.00) |
0.119 (1.00) |
0.894 (1.00) |
0.438 (1.00) |
0.387 (1.00) |
1 (1.00) |
0.386 (1.00) |
0.732 (1.00) |
| HIST1H1B | 7 (2%) | 299 |
0.627 (1.00) |
0.356 (1.00) |
0.705 (1.00) |
0.822 (1.00) |
0.802 (1.00) |
0.717 (1.00) |
0.183 (1.00) |
0.125 (1.00) |
| C3ORF59 | 8 (3%) | 298 |
0.459 (1.00) |
0.476 (1.00) |
0.173 (1.00) |
0.00863 (1.00) |
1 (1.00) |
1 (1.00) |
0.908 (1.00) |
1 (1.00) |
| C8ORF34 | 7 (2%) | 299 |
0.931 (1.00) |
0.0892 (1.00) |
0.419 (1.00) |
0.0617 (1.00) |
0.802 (1.00) |
1 (1.00) |
0.345 (1.00) |
1 (1.00) |
| OR8J1 | 9 (3%) | 297 |
0.161 (1.00) |
0.319 (1.00) |
0.0346 (1.00) |
0.0238 (1.00) |
0.245 (1.00) |
0.265 (1.00) |
0.296 (1.00) |
0.612 (1.00) |
| RAB32 | 3 (1%) | 303 |
0.0793 (1.00) |
0.825 (1.00) |
0.856 (1.00) |
0.672 (1.00) |
0.114 (1.00) |
0.116 (1.00) |
0.381 (1.00) |
0.35 (1.00) |
| TMEM195 | 8 (3%) | 298 |
0.508 (1.00) |
0.448 (1.00) |
0.79 (1.00) |
0.71 (1.00) |
0.821 (1.00) |
0.905 (1.00) |
0.82 (1.00) |
1 (1.00) |
| PCDH11X | 21 (7%) | 285 |
0.422 (1.00) |
0.71 (1.00) |
0.697 (1.00) |
0.0248 (1.00) |
0.346 (1.00) |
0.377 (1.00) |
0.78 (1.00) |
0.904 (1.00) |
| SLITRK4 | 12 (4%) | 294 |
0.147 (1.00) |
0.178 (1.00) |
0.465 (1.00) |
0.613 (1.00) |
0.092 (1.00) |
0.0896 (1.00) |
0.194 (1.00) |
0.283 (1.00) |
| FRG2B | 6 (2%) | 300 |
0.773 (1.00) |
0.692 (1.00) |
0.324 (1.00) |
0.672 (1.00) |
0.291 (1.00) |
0.289 (1.00) |
0.291 (1.00) |
0.732 (1.00) |
| OR2L13 | 7 (2%) | 299 |
0.239 (1.00) |
0.101 (1.00) |
0.00821 (1.00) |
1 (1.00) |
0.345 (1.00) |
0.498 (1.00) |
0.5 (1.00) |
0.00432 (1.00) |
| CUL3 | 10 (3%) | 296 |
0.0463 (1.00) |
0.124 (1.00) |
0.737 (1.00) |
0.636 (1.00) |
0.403 (1.00) |
0.852 (1.00) |
0.551 (1.00) |
0.403 (1.00) |
| FOSL2 | 7 (2%) | 299 |
0.395 (1.00) |
0.611 (1.00) |
0.644 (1.00) |
1 (1.00) |
0.345 (1.00) |
0.717 (1.00) |
0.345 (1.00) |
0.281 (1.00) |
| GPR158 | 12 (4%) | 294 |
0.454 (1.00) |
0.131 (1.00) |
1 (1.00) |
0.865 (1.00) |
0.538 (1.00) |
0.531 (1.00) |
0.934 (1.00) |
0.573 (1.00) |
| PTPRT | 19 (6%) | 287 |
0.743 (1.00) |
0.0925 (1.00) |
0.576 (1.00) |
0.752 (1.00) |
0.0996 (1.00) |
0.693 (1.00) |
0.477 (1.00) |
0.501 (1.00) |
| POTEH | 7 (2%) | 299 |
0.892 (1.00) |
0.55 (1.00) |
0.152 (1.00) |
0.155 (1.00) |
0.123 (1.00) |
0.103 (1.00) |
0.57 (1.00) |
0.767 (1.00) |
| OR5D13 | 7 (2%) | 299 |
0.756 (1.00) |
0.16 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.767 (1.00) |
| PRB1 | 7 (2%) | 299 |
0.5 (1.00) |
0.177 (1.00) |
0.757 (1.00) |
0.672 (1.00) |
0.89 (1.00) |
0.639 (1.00) |
0.104 (1.00) |
1 (1.00) |
| NEUROD6 | 7 (2%) | 299 |
0.892 (1.00) |
0.862 (1.00) |
0.502 (1.00) |
0.563 (1.00) |
1 (1.00) |
1 (1.00) |
||
| DPP10 | 16 (5%) | 290 |
0.0613 (1.00) |
0.12 (1.00) |
0.115 (1.00) |
0.26 (1.00) |
0.17 (1.00) |
0.0574 (1.00) |
0.0128 (1.00) |
0.0731 (1.00) |
| DPPA4 | 7 (2%) | 299 |
0.0625 (1.00) |
0.0035 (1.00) |
0.442 (1.00) |
0.0845 (1.00) |
0.123 (1.00) |
0.0461 (1.00) |
0.5 (1.00) |
0.767 (1.00) |
| EPDR1 | 6 (2%) | 300 |
0.773 (1.00) |
0.799 (1.00) |
0.561 (1.00) |
0.779 (1.00) |
0.327 (1.00) |
0.675 (1.00) |
0.881 (1.00) |
1 (1.00) |
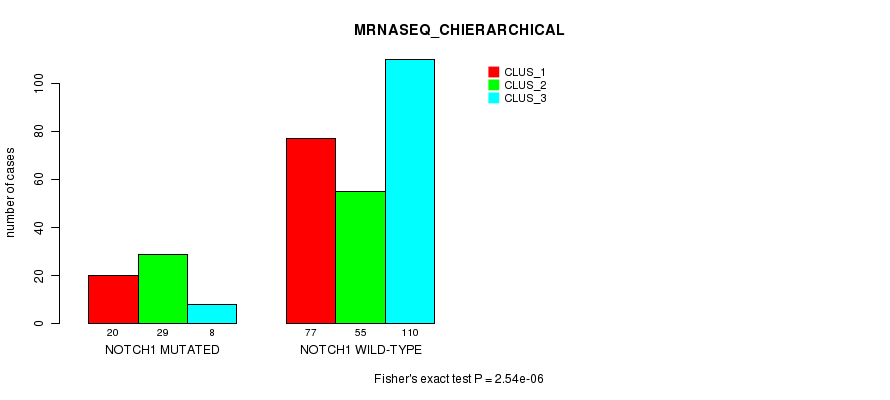
P value = 2.54e-06 (Fisher's exact test), Q value = 0.0019
Table S1. Gene #1: 'NOTCH1 MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 97 | 84 | 118 |
| NOTCH1 MUTATED | 20 | 29 | 8 |
| NOTCH1 WILD-TYPE | 77 | 55 | 110 |
Figure S1. Get High-res Image Gene #1: 'NOTCH1 MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
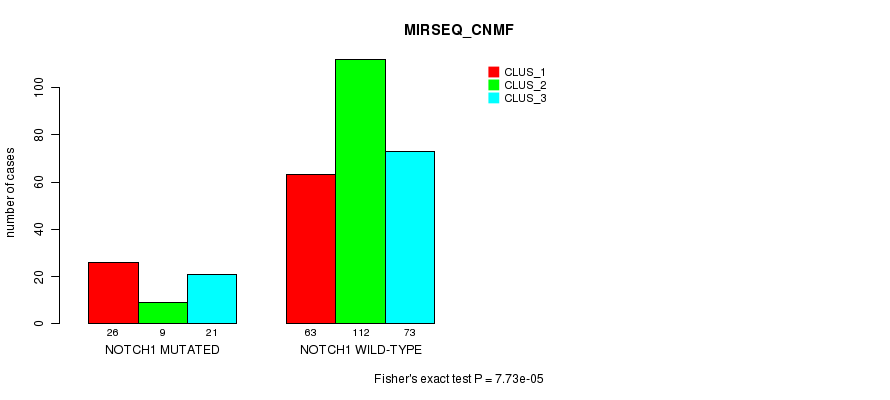
P value = 7.73e-05 (Fisher's exact test), Q value = 0.056
Table S2. Gene #1: 'NOTCH1 MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 89 | 121 | 94 |
| NOTCH1 MUTATED | 26 | 9 | 21 |
| NOTCH1 WILD-TYPE | 63 | 112 | 73 |
Figure S2. Get High-res Image Gene #1: 'NOTCH1 MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
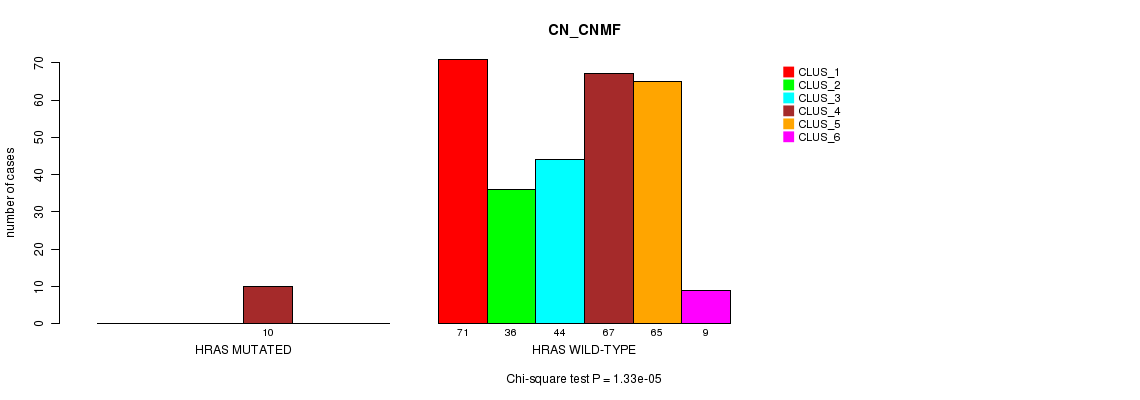
P value = 1.33e-05 (Chi-square test), Q value = 0.0097
Table S3. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 71 | 36 | 44 | 77 | 65 | 9 |
| HRAS MUTATED | 0 | 0 | 0 | 10 | 0 | 0 |
| HRAS WILD-TYPE | 71 | 36 | 44 | 67 | 65 | 9 |
Figure S3. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
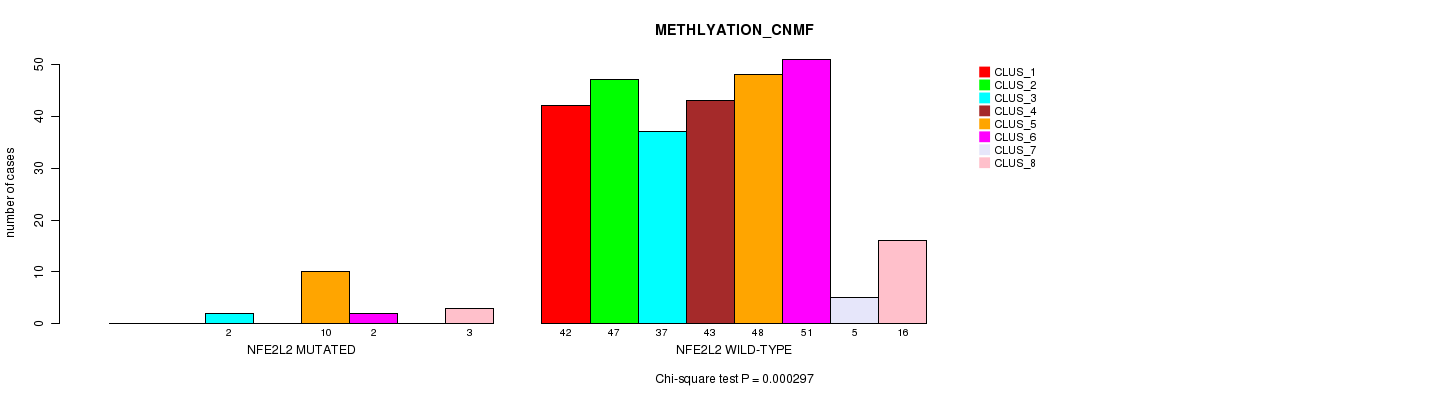
P value = 0.000297 (Chi-square test), Q value = 0.21
Table S4. Gene #4: 'NFE2L2 MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 42 | 47 | 39 | 43 | 58 | 53 | 5 | 19 |
| NFE2L2 MUTATED | 0 | 0 | 2 | 0 | 10 | 2 | 0 | 3 |
| NFE2L2 WILD-TYPE | 42 | 47 | 37 | 43 | 48 | 51 | 5 | 16 |
Figure S4. Get High-res Image Gene #4: 'NFE2L2 MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
P value = 1.84e-16 (Chi-square test), Q value = 1.3e-13
Table S5. Gene #6: 'TP53 MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 71 | 36 | 44 | 77 | 65 | 9 |
| TP53 MUTATED | 60 | 33 | 38 | 23 | 50 | 8 |
| TP53 WILD-TYPE | 11 | 3 | 6 | 54 | 15 | 1 |
Figure S5. Get High-res Image Gene #6: 'TP53 MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
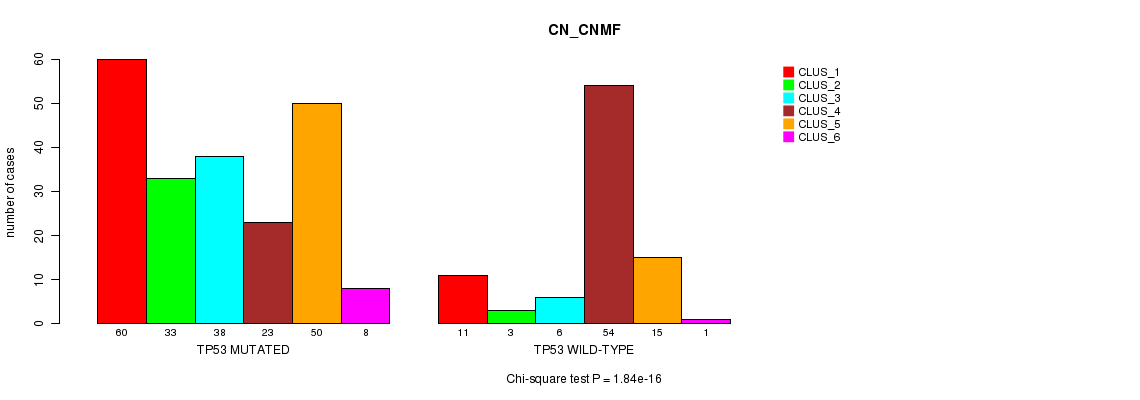
P value = 3.14e-18 (Chi-square test), Q value = 2.3e-15
Table S6. Gene #6: 'TP53 MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 42 | 47 | 39 | 43 | 58 | 53 | 5 | 19 |
| TP53 MUTATED | 34 | 43 | 29 | 4 | 51 | 36 | 3 | 14 |
| TP53 WILD-TYPE | 8 | 4 | 10 | 39 | 7 | 17 | 2 | 5 |
Figure S6. Get High-res Image Gene #6: 'TP53 MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
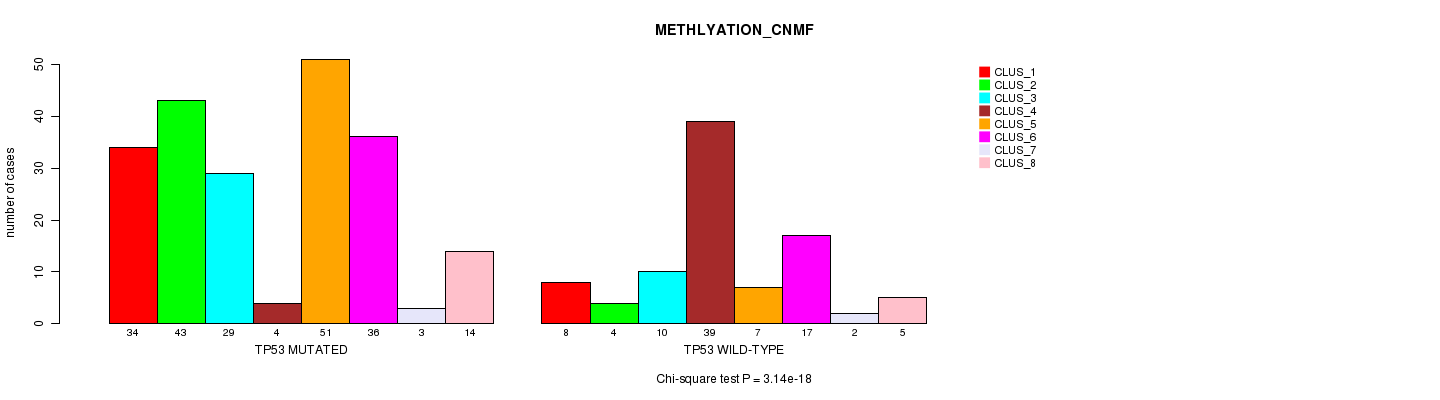
P value = 0.000326 (Chi-square test), Q value = 0.24
Table S7. Gene #7: 'CASP8 MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 71 | 36 | 44 | 77 | 65 | 9 |
| CASP8 MUTATED | 1 | 1 | 1 | 16 | 6 | 1 |
| CASP8 WILD-TYPE | 70 | 35 | 43 | 61 | 59 | 8 |
Figure S7. Get High-res Image Gene #7: 'CASP8 MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
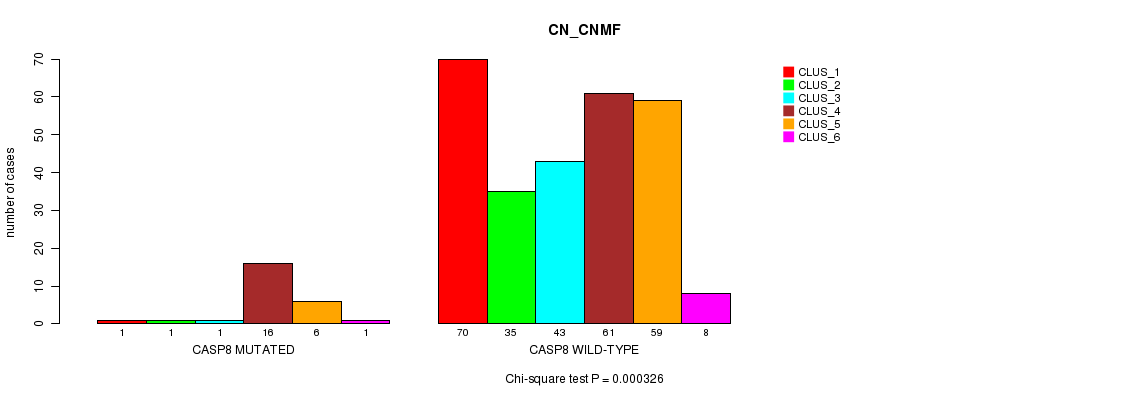
P value = 7.22e-06 (Chi-square test), Q value = 0.0052
Table S8. Gene #7: 'CASP8 MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 42 | 47 | 39 | 43 | 58 | 53 | 5 | 19 |
| CASP8 MUTATED | 1 | 3 | 8 | 0 | 0 | 9 | 0 | 6 |
| CASP8 WILD-TYPE | 41 | 44 | 31 | 43 | 58 | 44 | 5 | 13 |
Figure S8. Get High-res Image Gene #7: 'CASP8 MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
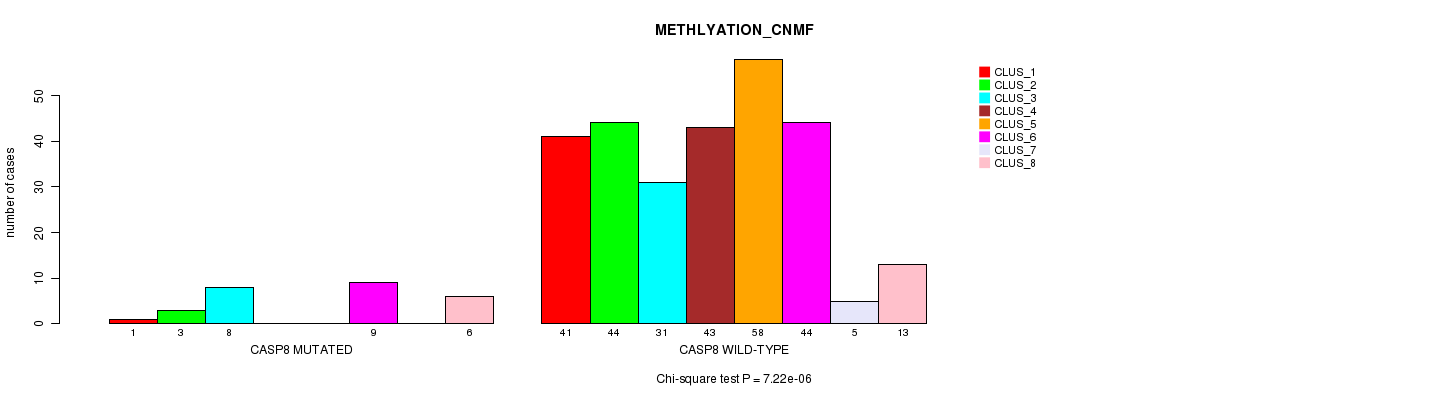
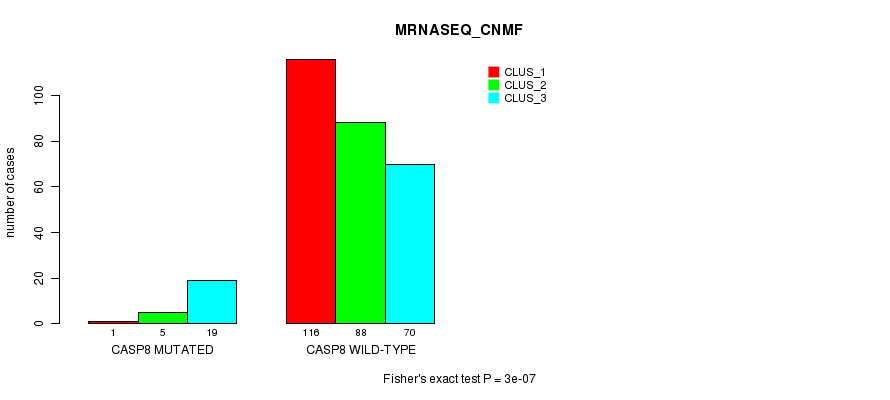
P value = 3e-07 (Fisher's exact test), Q value = 0.00022
Table S9. Gene #7: 'CASP8 MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 117 | 93 | 89 |
| CASP8 MUTATED | 1 | 5 | 19 |
| CASP8 WILD-TYPE | 116 | 88 | 70 |
Figure S9. Get High-res Image Gene #7: 'CASP8 MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
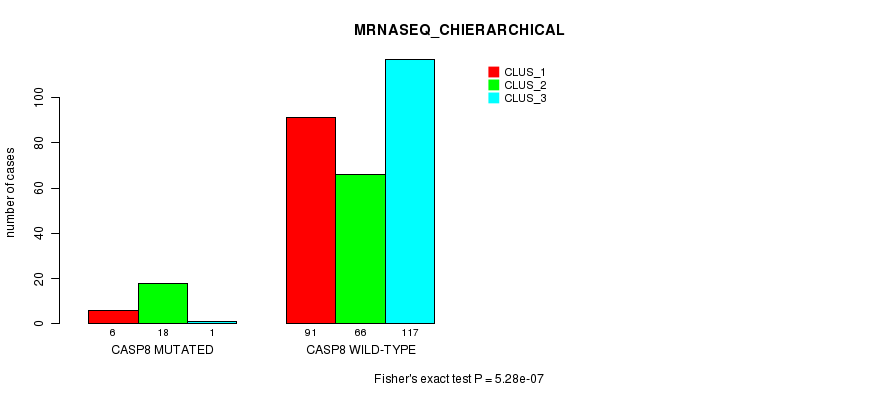
P value = 5.28e-07 (Fisher's exact test), Q value = 0.00039
Table S10. Gene #7: 'CASP8 MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 97 | 84 | 118 |
| CASP8 MUTATED | 6 | 18 | 1 |
| CASP8 WILD-TYPE | 91 | 66 | 117 |
Figure S10. Get High-res Image Gene #7: 'CASP8 MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
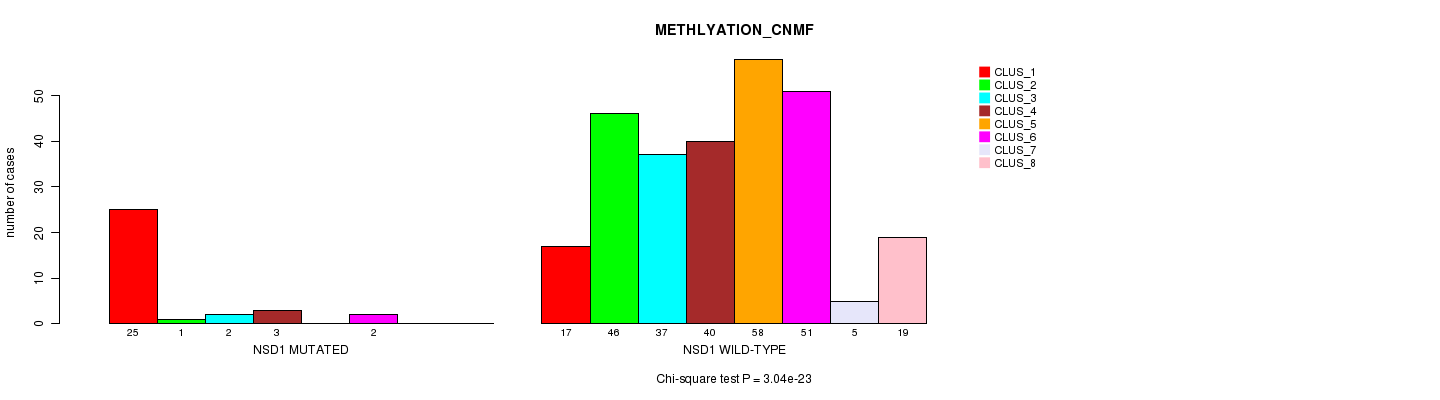
P value = 3.04e-23 (Chi-square test), Q value = 2.2e-20
Table S11. Gene #12: 'NSD1 MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 42 | 47 | 39 | 43 | 58 | 53 | 5 | 19 |
| NSD1 MUTATED | 25 | 1 | 2 | 3 | 0 | 2 | 0 | 0 |
| NSD1 WILD-TYPE | 17 | 46 | 37 | 40 | 58 | 51 | 5 | 19 |
Figure S11. Get High-res Image Gene #12: 'NSD1 MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
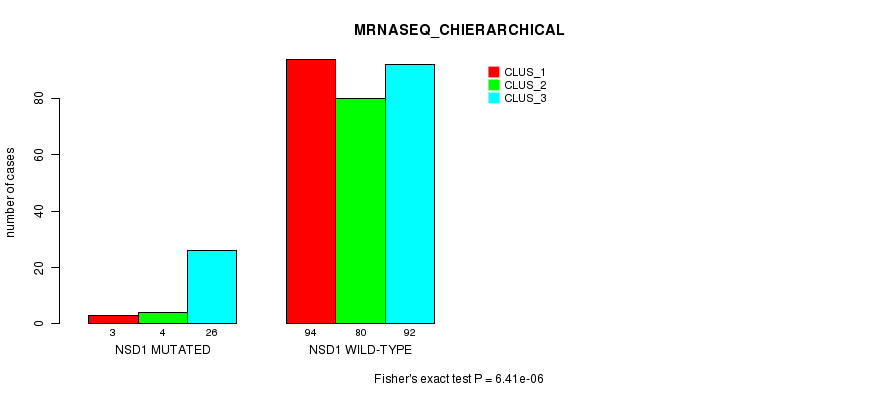
P value = 6.41e-06 (Fisher's exact test), Q value = 0.0047
Table S12. Gene #12: 'NSD1 MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 97 | 84 | 118 |
| NSD1 MUTATED | 3 | 4 | 26 |
| NSD1 WILD-TYPE | 94 | 80 | 92 |
Figure S12. Get High-res Image Gene #12: 'NSD1 MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
-
Mutation data file = HNSC-TP.mutsig.cluster.txt
-
Molecular subtypes file = HNSC-TP.transferedmergedcluster.txt
-
Number of patients = 306
-
Number of significantly mutated genes = 94
-
Number of Molecular subtypes = 8
-
Exclude genes that fewer than K tumors have mutations, K = 3
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.