This pipeline computes the correlation between significant arm-level copy number variations (cnvs) and subtypes.
Testing the association between copy number variation 59 arm-level results and 8 molecular subtypes across 127 patients, 40 significant findings detected with Q value < 0.25.
-
1q gain cnv correlated to 'METHLYATION_CNMF'.
-
3p gain cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
3q gain cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
5p gain cnv correlated to 'CN_CNMF'.
-
5q gain cnv correlated to 'CN_CNMF'.
-
7p gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
7q gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
12p gain cnv correlated to 'CN_CNMF'.
-
12q gain cnv correlated to 'CN_CNMF'.
-
16p gain cnv correlated to 'CN_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
16q gain cnv correlated to 'CN_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
17p gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
17q gain cnv correlated to 'CN_CNMF'.
-
4p loss cnv correlated to 'CN_CNMF'.
-
5q loss cnv correlated to 'CN_CNMF'.
-
6p loss cnv correlated to 'CN_CNMF'.
-
9p loss cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
9q loss cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
14q loss cnv correlated to 'CN_CNMF'.
-
22q loss cnv correlated to 'CN_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 59 arm-level results and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 40 significant findings detected.
|
Molecular subtypes |
MRNA CNMF |
MRNA CHIERARCHICAL |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Chi-square test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| 3p gain | 0 (0%) | 98 |
0.302 (1.00) |
0.0559 (1.00) |
7.99e-15 (3.22e-12) |
0.0132 (1.00) |
0.000268 (0.0993) |
3.64e-07 (0.000144) |
0.0375 (1.00) |
1.22e-05 (0.0047) |
| 3q gain | 0 (0%) | 94 |
0.302 (1.00) |
0.0559 (1.00) |
1.95e-12 (7.83e-10) |
0.00739 (1.00) |
3.65e-05 (0.0139) |
6.94e-06 (0.0027) |
0.00911 (1.00) |
0.000246 (0.0914) |
| 7p gain | 0 (0%) | 60 |
0.633 (1.00) |
0.432 (1.00) |
2.69e-07 (0.000107) |
2.62e-06 (0.00103) |
5.65e-05 (0.0214) |
2.01e-05 (0.00771) |
0.011 (1.00) |
0.00896 (1.00) |
| 7q gain | 0 (0%) | 59 |
0.633 (1.00) |
0.432 (1.00) |
5.79e-08 (2.31e-05) |
9.37e-06 (0.00363) |
0.000181 (0.068) |
3.74e-05 (0.0142) |
0.00419 (1.00) |
0.0113 (1.00) |
| 17p gain | 0 (0%) | 61 |
0.633 (1.00) |
0.432 (1.00) |
9.52e-08 (3.78e-05) |
1.3e-06 (0.000513) |
3.49e-05 (0.0133) |
2.21e-06 (0.000869) |
0.129 (1.00) |
0.00197 (0.692) |
| 16q gain | 0 (0%) | 69 |
0.302 (1.00) |
0.0559 (1.00) |
9.17e-06 (0.00356) |
0.0521 (1.00) |
0.00419 (1.00) |
1.46e-05 (0.00563) |
0.00179 (0.629) |
0.000223 (0.0835) |
| 16p gain | 0 (0%) | 67 |
0.126 (1.00) |
0.0699 (1.00) |
4.19e-06 (0.00164) |
0.221 (1.00) |
0.00874 (1.00) |
0.000182 (0.0684) |
0.00268 (0.934) |
0.000706 (0.256) |
| 9p loss | 0 (0%) | 112 |
1 (1.00) |
0.75 (1.00) |
0.000169 (0.0639) |
0.000318 (0.117) |
0.00782 (1.00) |
0.0678 (1.00) |
0.232 (1.00) |
0.587 (1.00) |
| 9q loss | 0 (0%) | 110 |
1 (1.00) |
0.000445 (0.163) |
3.17e-05 (0.0121) |
0.0256 (1.00) |
0.131 (1.00) |
0.37 (1.00) |
0.254 (1.00) |
|
| 1q gain | 0 (0%) | 118 |
0.438 (1.00) |
0.00553 (1.00) |
0.000353 (0.13) |
0.00232 (0.813) |
0.00827 (1.00) |
0.433 (1.00) |
0.145 (1.00) |
|
| 5p gain | 0 (0%) | 113 |
0.175 (1.00) |
1.96e-08 (7.8e-06) |
0.0521 (1.00) |
0.156 (1.00) |
0.8 (1.00) |
0.697 (1.00) |
0.731 (1.00) |
|
| 5q gain | 0 (0%) | 113 |
0.175 (1.00) |
2.18e-10 (8.74e-08) |
0.0521 (1.00) |
0.579 (1.00) |
0.8 (1.00) |
0.919 (1.00) |
0.301 (1.00) |
|
| 12p gain | 0 (0%) | 87 |
1 (1.00) |
0.0909 (1.00) |
0.00055 (0.201) |
0.833 (1.00) |
0.161 (1.00) |
0.0648 (1.00) |
0.146 (1.00) |
0.0968 (1.00) |
| 12q gain | 0 (0%) | 87 |
1 (1.00) |
0.0909 (1.00) |
0.00055 (0.201) |
0.833 (1.00) |
0.161 (1.00) |
0.0648 (1.00) |
0.146 (1.00) |
0.0968 (1.00) |
| 17q gain | 0 (0%) | 50 |
0.308 (1.00) |
0.808 (1.00) |
5.4e-06 (0.00211) |
0.00104 (0.372) |
0.0144 (1.00) |
0.000926 (0.333) |
0.362 (1.00) |
0.156 (1.00) |
| 4p loss | 0 (0%) | 116 |
1 (1.00) |
0.000247 (0.0917) |
0.00449 (1.00) |
0.0129 (1.00) |
0.18 (1.00) |
0.535 (1.00) |
1 (1.00) |
|
| 5q loss | 0 (0%) | 121 |
1.96e-05 (0.00754) |
0.23 (1.00) |
0.41 (1.00) |
0.211 (1.00) |
0.236 (1.00) |
0.916 (1.00) |
||
| 6p loss | 0 (0%) | 118 |
1 (1.00) |
0.00043 (0.158) |
0.443 (1.00) |
0.468 (1.00) |
1 (1.00) |
0.331 (1.00) |
0.539 (1.00) |
|
| 14q loss | 0 (0%) | 104 |
1 (1.00) |
0.385 (1.00) |
1.54e-11 (6.18e-09) |
0.267 (1.00) |
0.135 (1.00) |
0.0977 (1.00) |
0.457 (1.00) |
0.968 (1.00) |
| 22q loss | 0 (0%) | 103 |
1 (1.00) |
0.000233 (0.087) |
0.00271 (0.944) |
0.673 (1.00) |
0.0477 (1.00) |
0.186 (1.00) |
0.433 (1.00) |
|
| 2p gain | 0 (0%) | 114 |
0.55 (1.00) |
0.75 (1.00) |
0.0607 (1.00) |
0.132 (1.00) |
0.585 (1.00) |
0.338 (1.00) |
0.869 (1.00) |
0.0669 (1.00) |
| 2q gain | 0 (0%) | 112 |
0.55 (1.00) |
0.75 (1.00) |
0.0309 (1.00) |
0.354 (1.00) |
0.329 (1.00) |
0.205 (1.00) |
0.856 (1.00) |
0.174 (1.00) |
| 4p gain | 0 (0%) | 123 |
0.175 (1.00) |
0.382 (1.00) |
0.457 (1.00) |
1 (1.00) |
0.683 (1.00) |
0.881 (1.00) |
||
| 4q gain | 0 (0%) | 124 |
0.438 (1.00) |
0.572 (1.00) |
0.645 (1.00) |
0.602 (1.00) |
0.767 (1.00) |
0.618 (1.00) |
||
| 6p gain | 0 (0%) | 123 |
0.438 (1.00) |
0.503 (1.00) |
0.44 (1.00) |
0.114 (1.00) |
0.677 (1.00) |
0.165 (1.00) |
0.374 (1.00) |
|
| 6q gain | 0 (0%) | 124 |
0.438 (1.00) |
0.213 (1.00) |
0.41 (1.00) |
1 (1.00) |
0.231 (1.00) |
0.397 (1.00) |
||
| 8p gain | 0 (0%) | 118 |
1 (1.00) |
0.318 (1.00) |
0.087 (1.00) |
0.732 (1.00) |
0.232 (1.00) |
0.617 (1.00) |
0.854 (1.00) |
|
| 8q gain | 0 (0%) | 116 |
1 (1.00) |
0.00446 (1.00) |
0.0131 (1.00) |
0.243 (1.00) |
0.0683 (1.00) |
0.187 (1.00) |
0.767 (1.00) |
|
| 10p gain | 0 (0%) | 123 |
1 (1.00) |
0.184 (1.00) |
0.645 (1.00) |
0.602 (1.00) |
0.189 (1.00) |
0.881 (1.00) |
||
| 10q gain | 0 (0%) | 124 |
1 (1.00) |
0.393 (1.00) |
0.454 (1.00) |
0.341 (1.00) |
||||
| 13q gain | 0 (0%) | 113 |
1 (1.00) |
0.00662 (1.00) |
0.179 (1.00) |
0.324 (1.00) |
0.0445 (1.00) |
0.747 (1.00) |
0.0852 (1.00) |
|
| 18p gain | 0 (0%) | 121 |
0.55 (1.00) |
0.141 (1.00) |
0.16 (1.00) |
0.329 (1.00) |
0.863 (1.00) |
1 (1.00) |
0.845 (1.00) |
0.877 (1.00) |
| 18q gain | 0 (0%) | 123 |
0.438 (1.00) |
0.0227 (1.00) |
0.329 (1.00) |
0.77 (1.00) |
0.432 (1.00) |
0.91 (1.00) |
0.331 (1.00) |
|
| 20p gain | 0 (0%) | 88 |
0.126 (1.00) |
0.162 (1.00) |
0.0185 (1.00) |
0.0743 (1.00) |
0.000722 (0.261) |
0.0152 (1.00) |
0.305 (1.00) |
0.125 (1.00) |
| 20q gain | 0 (0%) | 87 |
0.315 (1.00) |
0.119 (1.00) |
0.0107 (1.00) |
0.142 (1.00) |
0.000813 (0.294) |
0.0381 (1.00) |
0.179 (1.00) |
0.371 (1.00) |
| 21q gain | 0 (0%) | 122 |
0.438 (1.00) |
0.637 (1.00) |
0.468 (1.00) |
0.11 (1.00) |
0.0492 (1.00) |
|||
| Xq gain | 0 (0%) | 122 |
1 (1.00) |
0.121 (1.00) |
0.137 (1.00) |
0.457 (1.00) |
1 (1.00) |
0.57 (1.00) |
0.881 (1.00) |
|
| 1p loss | 0 (0%) | 113 |
1 (1.00) |
0.0256 (1.00) |
0.328 (1.00) |
0.611 (1.00) |
0.589 (1.00) |
0.0114 (1.00) |
0.0873 (1.00) |
|
| 1q loss | 0 (0%) | 120 |
1 (1.00) |
0.0171 (1.00) |
0.193 (1.00) |
0.103 (1.00) |
0.432 (1.00) |
0.0359 (1.00) |
0.0124 (1.00) |
|
| 3p loss | 0 (0%) | 119 |
0.438 (1.00) |
0.00138 (0.493) |
0.00142 (0.504) |
0.058 (1.00) |
0.0149 (1.00) |
0.0591 (1.00) |
0.221 (1.00) |
|
| 3q loss | 0 (0%) | 124 |
0.248 (1.00) |
0.0335 (1.00) |
0.0288 (1.00) |
0.0762 (1.00) |
0.521 (1.00) |
0.72 (1.00) |
||
| 4q loss | 0 (0%) | 115 |
1 (1.00) |
0.00104 (0.372) |
0.00153 (0.543) |
0.0999 (1.00) |
0.0683 (1.00) |
0.853 (1.00) |
0.698 (1.00) |
|
| 5p loss | 0 (0%) | 122 |
0.00932 (1.00) |
0.246 (1.00) |
0.114 (1.00) |
0.0626 (1.00) |
0.236 (1.00) |
0.916 (1.00) |
||
| 6q loss | 0 (0%) | 115 |
1 (1.00) |
0.00107 (0.382) |
0.066 (1.00) |
0.374 (1.00) |
0.558 (1.00) |
0.783 (1.00) |
0.693 (1.00) |
|
| 8p loss | 0 (0%) | 124 |
1 (1.00) |
0.846 (1.00) |
1 (1.00) |
0.618 (1.00) |
||||
| 10p loss | 0 (0%) | 120 |
1 (1.00) |
0.0171 (1.00) |
0.056 (1.00) |
0.269 (1.00) |
0.18 (1.00) |
0.265 (1.00) |
0.675 (1.00) |
|
| 10q loss | 0 (0%) | 119 |
0.438 (1.00) |
0.00696 (1.00) |
0.0362 (1.00) |
0.269 (1.00) |
0.0871 (1.00) |
0.617 (1.00) |
0.221 (1.00) |
|
| 11p loss | 0 (0%) | 118 |
1 (1.00) |
0.167 (1.00) |
0.113 (1.00) |
0.269 (1.00) |
0.145 (1.00) |
1 (1.00) |
0.447 (1.00) |
|
| 11q loss | 0 (0%) | 116 |
0.00512 (1.00) |
0.00354 (1.00) |
0.296 (1.00) |
0.0563 (1.00) |
0.968 (1.00) |
0.9 (1.00) |
||
| 13q loss | 0 (0%) | 115 |
1 (1.00) |
0.019 (1.00) |
0.00153 (0.543) |
0.091 (1.00) |
0.403 (1.00) |
0.355 (1.00) |
0.234 (1.00) |
|
| 15q loss | 0 (0%) | 116 |
1 (1.00) |
0.00407 (1.00) |
0.119 (1.00) |
0.579 (1.00) |
0.487 (1.00) |
0.573 (1.00) |
0.816 (1.00) |
|
| 16q loss | 0 (0%) | 124 |
0.632 (1.00) |
0.139 (1.00) |
0.862 (1.00) |
0.286 (1.00) |
||||
| 17p loss | 0 (0%) | 122 |
0.438 (1.00) |
0.756 (1.00) |
0.0436 (1.00) |
0.0288 (1.00) |
0.0762 (1.00) |
0.219 (1.00) |
0.251 (1.00) |
|
| 18p loss | 0 (0%) | 108 |
1 (1.00) |
0.0863 (1.00) |
0.0306 (1.00) |
0.0772 (1.00) |
0.0114 (1.00) |
0.158 (1.00) |
0.445 (1.00) |
|
| 18q loss | 0 (0%) | 107 |
1 (1.00) |
0.209 (1.00) |
0.0155 (1.00) |
0.0392 (1.00) |
0.00363 (1.00) |
0.322 (1.00) |
0.373 (1.00) |
|
| 19p loss | 0 (0%) | 123 |
0.0513 (1.00) |
1 (1.00) |
0.746 (1.00) |
0.331 (1.00) |
||||
| 19q loss | 0 (0%) | 123 |
0.0513 (1.00) |
1 (1.00) |
0.279 (1.00) |
0.618 (1.00) |
||||
| 21q loss | 0 (0%) | 112 |
0.55 (1.00) |
0.413 (1.00) |
0.504 (1.00) |
0.231 (1.00) |
0.246 (1.00) |
0.728 (1.00) |
0.219 (1.00) |
0.284 (1.00) |
| Xq loss | 0 (0%) | 124 |
1 (1.00) |
0.632 (1.00) |
0.767 (1.00) |
0.1 (1.00) |
P value = 0.000353 (Fisher's exact test), Q value = 0.13
Table S1. Gene #1: '1q gain' versus Molecular Subtype #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 53 | 31 |
| 1Q GAIN CNV | 1 | 0 | 7 |
| 1Q GAIN WILD-TYPE | 26 | 53 | 24 |
Figure S1. Get High-res Image Gene #1: '1q gain' versus Molecular Subtype #4: 'METHLYATION_CNMF'

P value = 7.99e-15 (Chi-square test), Q value = 3.2e-12
Table S2. Gene #4: '3p gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 3P GAIN CNV | 1 | 3 | 1 | 5 | 19 |
| 3P GAIN WILD-TYPE | 23 | 41 | 12 | 21 | 1 |
Figure S2. Get High-res Image Gene #4: '3p gain' versus Molecular Subtype #3: 'CN_CNMF'

P value = 0.000268 (Fisher's exact test), Q value = 0.099
Table S3. Gene #4: '3p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 24 | 25 |
| 3P GAIN CNV | 12 | 0 | 7 |
| 3P GAIN WILD-TYPE | 15 | 24 | 18 |
Figure S3. Get High-res Image Gene #4: '3p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
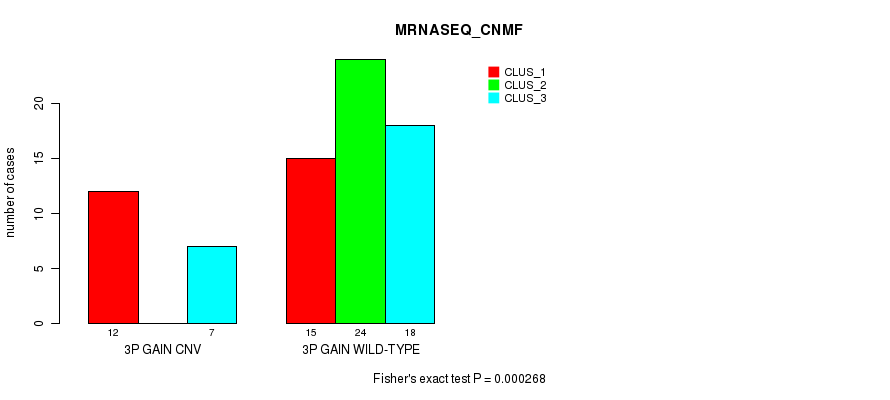
P value = 3.64e-07 (Fisher's exact test), Q value = 0.00014
Table S4. Gene #4: '3p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 37 | 21 |
| 3P GAIN CNV | 0 | 19 | 0 |
| 3P GAIN WILD-TYPE | 18 | 18 | 21 |
Figure S4. Get High-res Image Gene #4: '3p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
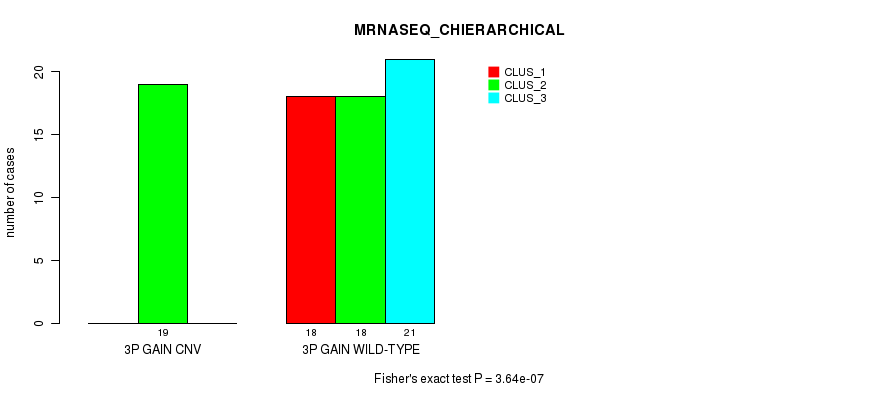
P value = 1.22e-05 (Fisher's exact test), Q value = 0.0047
Table S5. Gene #4: '3p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 27 | 45 | 12 | 33 |
| 3P GAIN CNV | 1 | 6 | 8 | 13 |
| 3P GAIN WILD-TYPE | 26 | 39 | 4 | 20 |
Figure S5. Get High-res Image Gene #4: '3p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
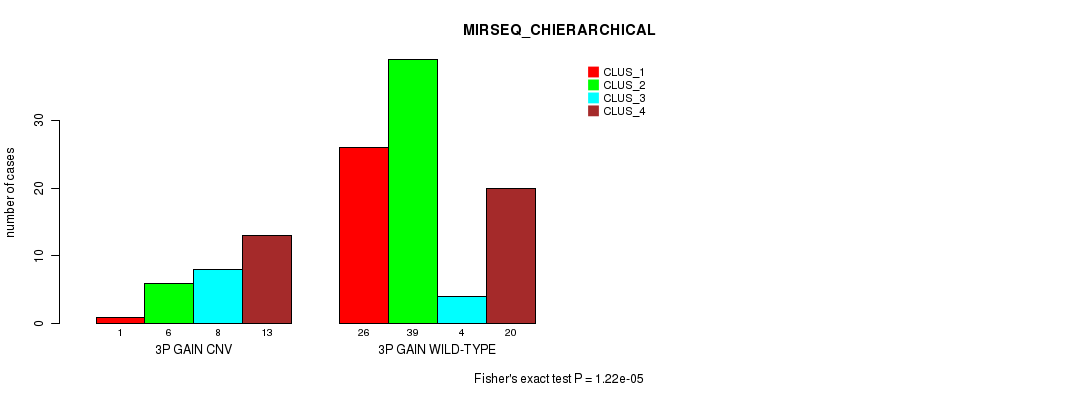
P value = 1.95e-12 (Chi-square test), Q value = 7.8e-10
Table S6. Gene #5: '3q gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 3Q GAIN CNV | 2 | 4 | 2 | 6 | 19 |
| 3Q GAIN WILD-TYPE | 22 | 40 | 11 | 20 | 1 |
Figure S6. Get High-res Image Gene #5: '3q gain' versus Molecular Subtype #3: 'CN_CNMF'

P value = 3.65e-05 (Fisher's exact test), Q value = 0.014
Table S7. Gene #5: '3q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 24 | 25 |
| 3Q GAIN CNV | 14 | 0 | 8 |
| 3Q GAIN WILD-TYPE | 13 | 24 | 17 |
Figure S7. Get High-res Image Gene #5: '3q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
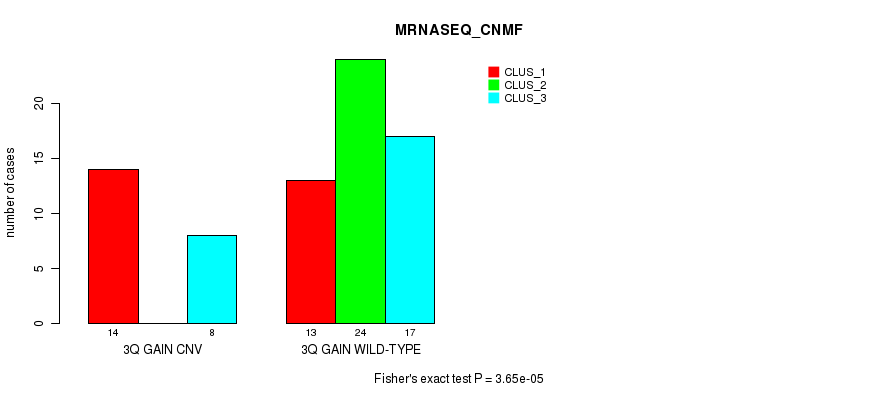
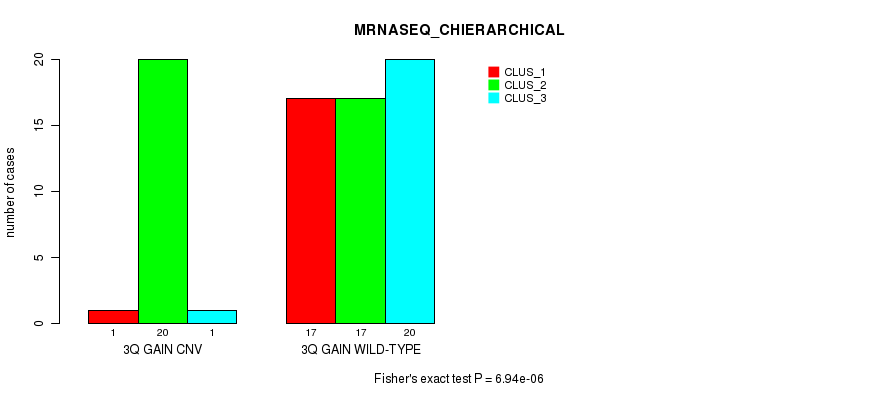
P value = 6.94e-06 (Fisher's exact test), Q value = 0.0027
Table S8. Gene #5: '3q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 37 | 21 |
| 3Q GAIN CNV | 1 | 20 | 1 |
| 3Q GAIN WILD-TYPE | 17 | 17 | 20 |
Figure S8. Get High-res Image Gene #5: '3q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
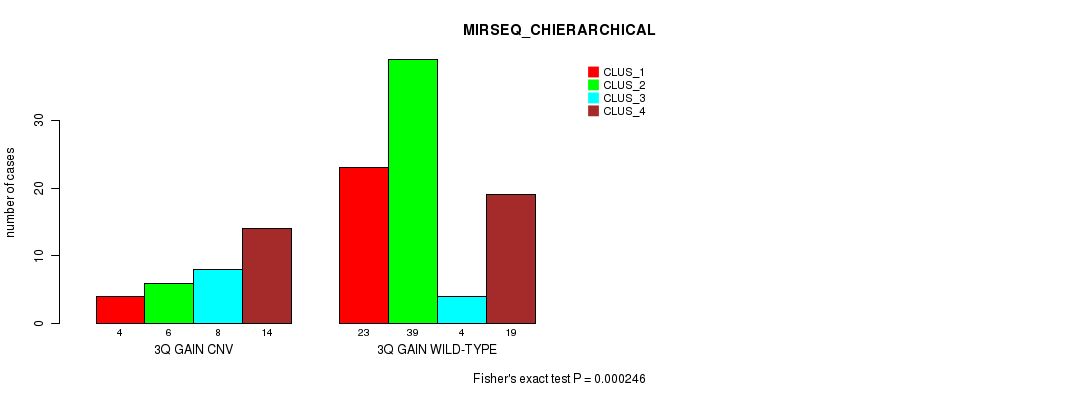
P value = 0.000246 (Fisher's exact test), Q value = 0.091
Table S9. Gene #5: '3q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 27 | 45 | 12 | 33 |
| 3Q GAIN CNV | 4 | 6 | 8 | 14 |
| 3Q GAIN WILD-TYPE | 23 | 39 | 4 | 19 |
Figure S9. Get High-res Image Gene #5: '3q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
P value = 1.96e-08 (Chi-square test), Q value = 7.8e-06
Table S10. Gene #8: '5p gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 5P GAIN CNV | 1 | 0 | 0 | 12 | 1 |
| 5P GAIN WILD-TYPE | 23 | 44 | 13 | 14 | 19 |
Figure S10. Get High-res Image Gene #8: '5p gain' versus Molecular Subtype #3: 'CN_CNMF'

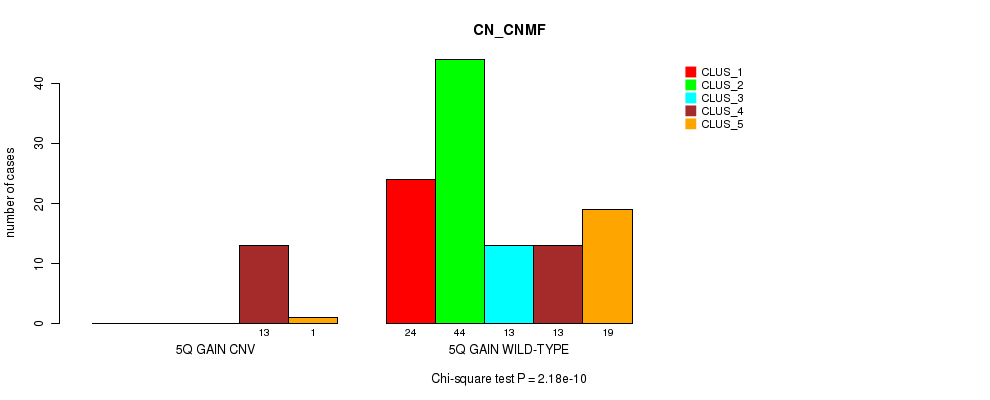
P value = 2.18e-10 (Chi-square test), Q value = 8.7e-08
Table S11. Gene #9: '5q gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 5Q GAIN CNV | 0 | 0 | 0 | 13 | 1 |
| 5Q GAIN WILD-TYPE | 24 | 44 | 13 | 13 | 19 |
Figure S11. Get High-res Image Gene #9: '5q gain' versus Molecular Subtype #3: 'CN_CNMF'
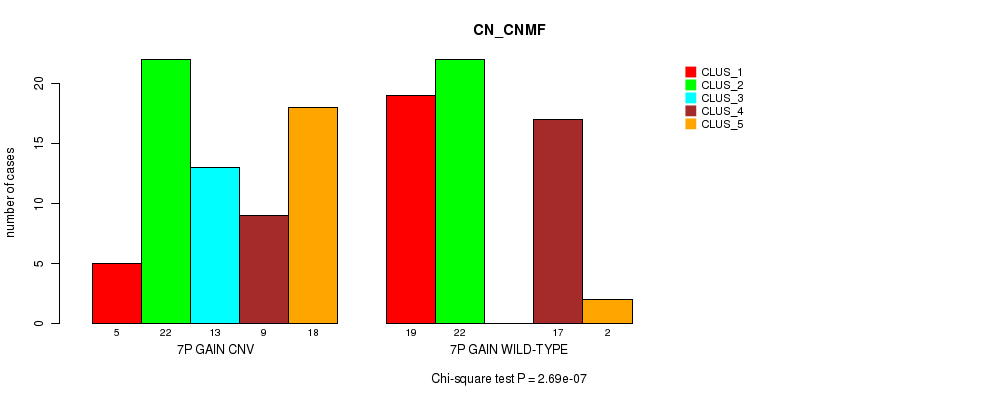
P value = 2.69e-07 (Chi-square test), Q value = 0.00011
Table S12. Gene #12: '7p gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 7P GAIN CNV | 5 | 22 | 13 | 9 | 18 |
| 7P GAIN WILD-TYPE | 19 | 22 | 0 | 17 | 2 |
Figure S12. Get High-res Image Gene #12: '7p gain' versus Molecular Subtype #3: 'CN_CNMF'
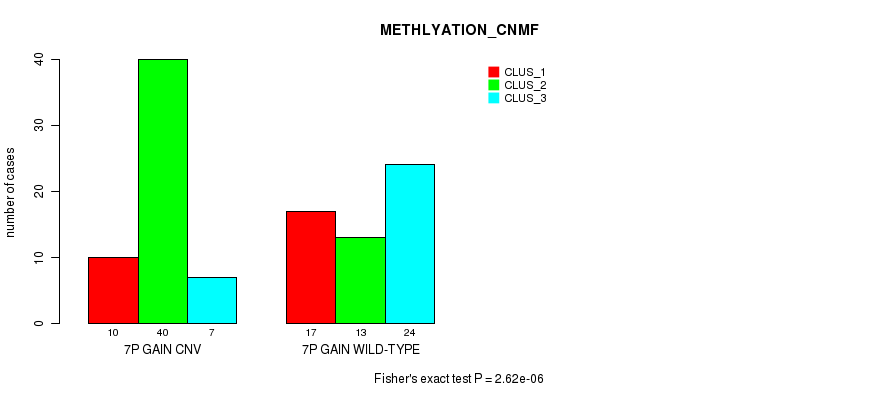
P value = 2.62e-06 (Fisher's exact test), Q value = 0.001
Table S13. Gene #12: '7p gain' versus Molecular Subtype #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 53 | 31 |
| 7P GAIN CNV | 10 | 40 | 7 |
| 7P GAIN WILD-TYPE | 17 | 13 | 24 |
Figure S13. Get High-res Image Gene #12: '7p gain' versus Molecular Subtype #4: 'METHLYATION_CNMF'
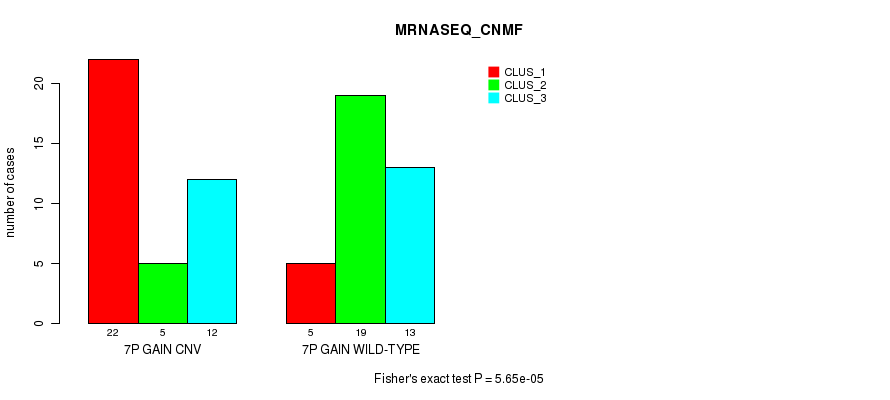
P value = 5.65e-05 (Fisher's exact test), Q value = 0.021
Table S14. Gene #12: '7p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 24 | 25 |
| 7P GAIN CNV | 22 | 5 | 12 |
| 7P GAIN WILD-TYPE | 5 | 19 | 13 |
Figure S14. Get High-res Image Gene #12: '7p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
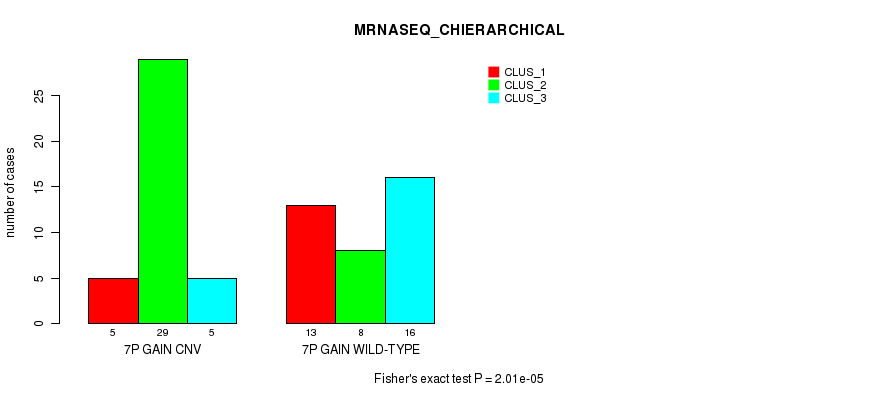
P value = 2.01e-05 (Fisher's exact test), Q value = 0.0077
Table S15. Gene #12: '7p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 37 | 21 |
| 7P GAIN CNV | 5 | 29 | 5 |
| 7P GAIN WILD-TYPE | 13 | 8 | 16 |
Figure S15. Get High-res Image Gene #12: '7p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
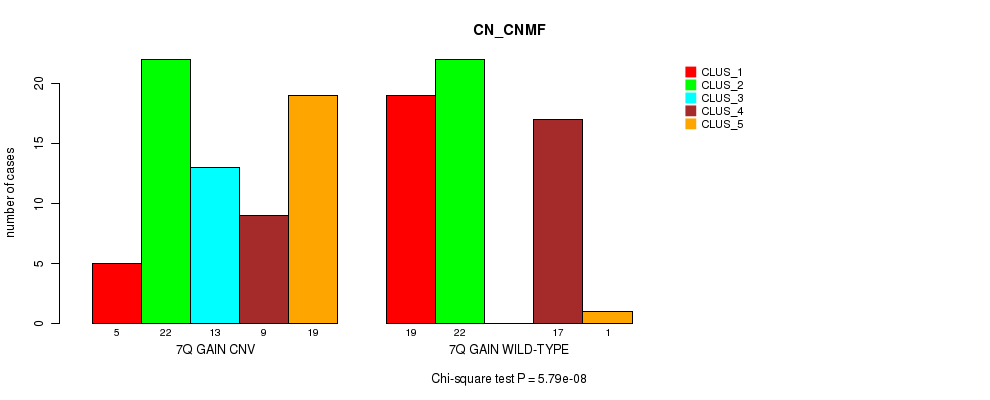
P value = 5.79e-08 (Chi-square test), Q value = 2.3e-05
Table S16. Gene #13: '7q gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 7Q GAIN CNV | 5 | 22 | 13 | 9 | 19 |
| 7Q GAIN WILD-TYPE | 19 | 22 | 0 | 17 | 1 |
Figure S16. Get High-res Image Gene #13: '7q gain' versus Molecular Subtype #3: 'CN_CNMF'
P value = 9.37e-06 (Fisher's exact test), Q value = 0.0036
Table S17. Gene #13: '7q gain' versus Molecular Subtype #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 53 | 31 |
| 7Q GAIN CNV | 10 | 40 | 8 |
| 7Q GAIN WILD-TYPE | 17 | 13 | 23 |
Figure S17. Get High-res Image Gene #13: '7q gain' versus Molecular Subtype #4: 'METHLYATION_CNMF'
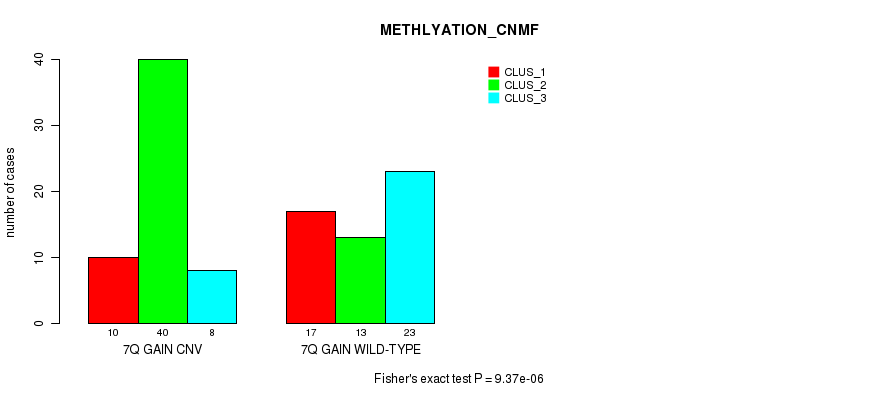
P value = 0.000181 (Fisher's exact test), Q value = 0.068
Table S18. Gene #13: '7q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 24 | 25 |
| 7Q GAIN CNV | 22 | 6 | 12 |
| 7Q GAIN WILD-TYPE | 5 | 18 | 13 |
Figure S18. Get High-res Image Gene #13: '7q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
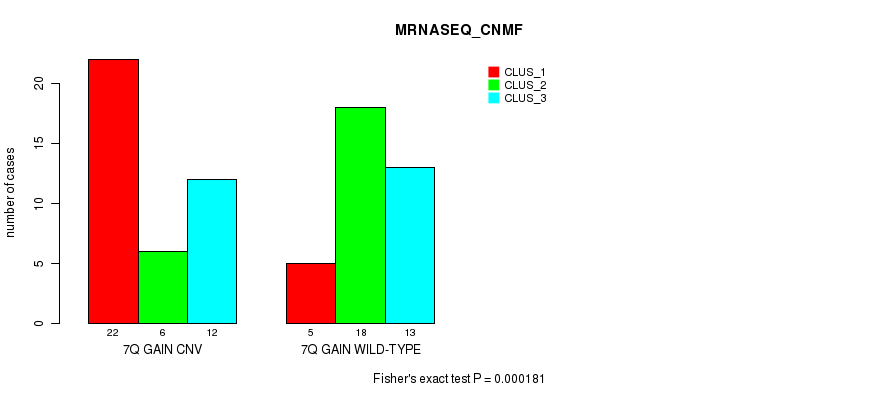
P value = 3.74e-05 (Fisher's exact test), Q value = 0.014
Table S19. Gene #13: '7q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 37 | 21 |
| 7Q GAIN CNV | 6 | 29 | 5 |
| 7Q GAIN WILD-TYPE | 12 | 8 | 16 |
Figure S19. Get High-res Image Gene #13: '7q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
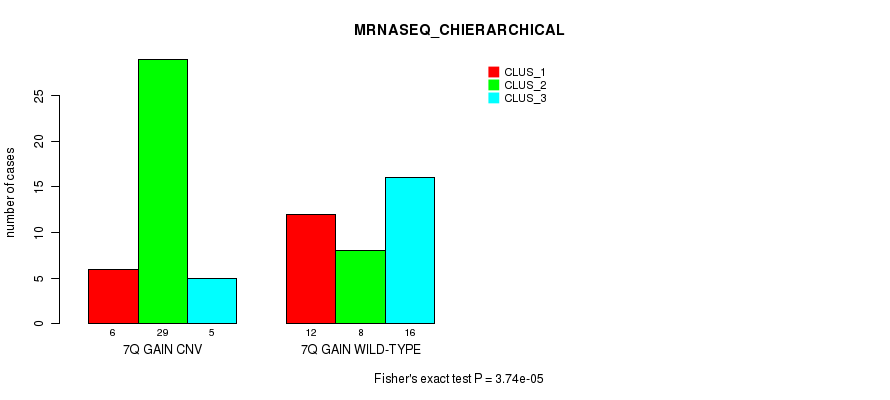
P value = 0.00055 (Chi-square test), Q value = 0.2
Table S20. Gene #18: '12p gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 12P GAIN CNV | 7 | 6 | 10 | 9 | 8 |
| 12P GAIN WILD-TYPE | 17 | 38 | 3 | 17 | 12 |
Figure S20. Get High-res Image Gene #18: '12p gain' versus Molecular Subtype #3: 'CN_CNMF'
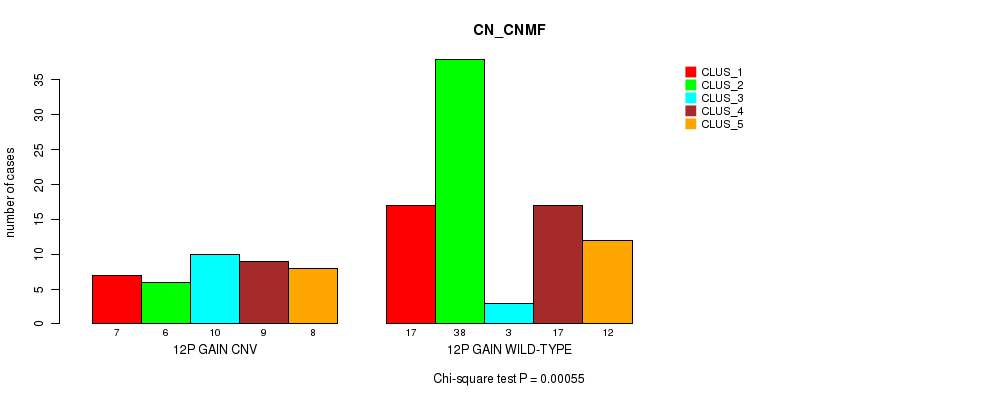
P value = 0.00055 (Chi-square test), Q value = 0.2
Table S21. Gene #19: '12q gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 12Q GAIN CNV | 7 | 6 | 10 | 9 | 8 |
| 12Q GAIN WILD-TYPE | 17 | 38 | 3 | 17 | 12 |
Figure S21. Get High-res Image Gene #19: '12q gain' versus Molecular Subtype #3: 'CN_CNMF'
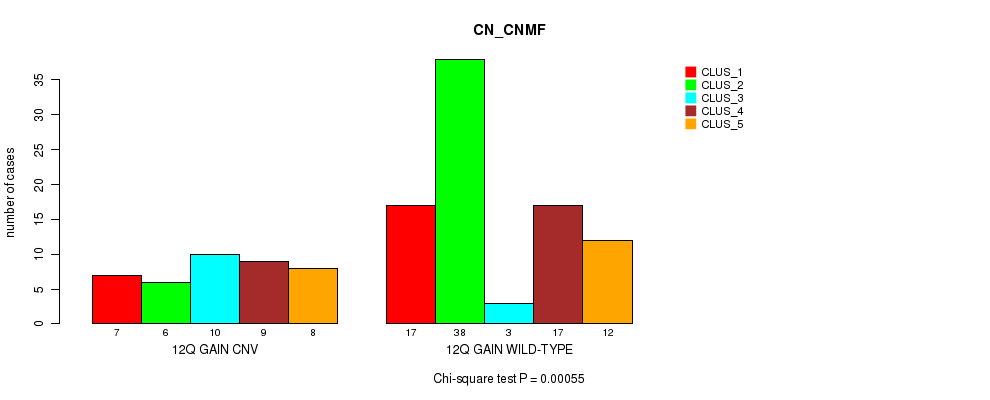
P value = 4.19e-06 (Chi-square test), Q value = 0.0016
Table S22. Gene #21: '16p gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 16P GAIN CNV | 6 | 16 | 3 | 16 | 19 |
| 16P GAIN WILD-TYPE | 18 | 28 | 10 | 10 | 1 |
Figure S22. Get High-res Image Gene #21: '16p gain' versus Molecular Subtype #3: 'CN_CNMF'

P value = 0.000182 (Fisher's exact test), Q value = 0.068
Table S23. Gene #21: '16p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 37 | 21 |
| 16P GAIN CNV | 2 | 24 | 6 |
| 16P GAIN WILD-TYPE | 16 | 13 | 15 |
Figure S23. Get High-res Image Gene #21: '16p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 9.17e-06 (Chi-square test), Q value = 0.0036
Table S24. Gene #22: '16q gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 16Q GAIN CNV | 6 | 16 | 3 | 14 | 19 |
| 16Q GAIN WILD-TYPE | 18 | 28 | 10 | 12 | 1 |
Figure S24. Get High-res Image Gene #22: '16q gain' versus Molecular Subtype #3: 'CN_CNMF'
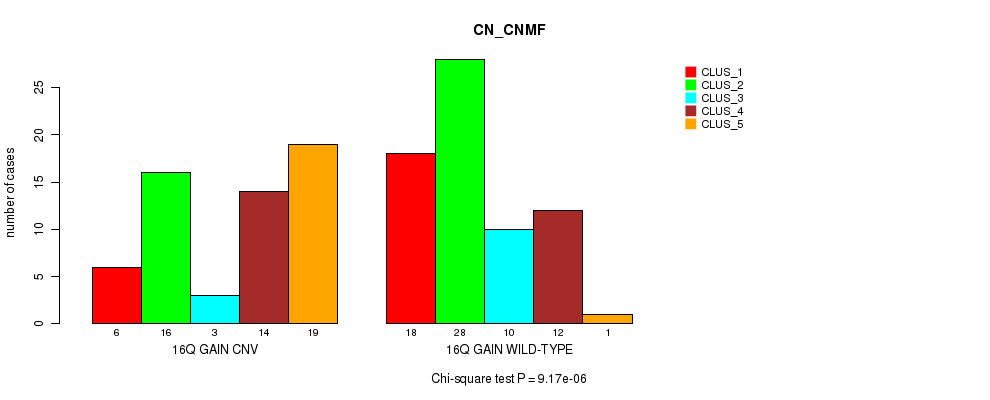
P value = 1.46e-05 (Fisher's exact test), Q value = 0.0056
Table S25. Gene #22: '16q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 37 | 21 |
| 16Q GAIN CNV | 2 | 24 | 3 |
| 16Q GAIN WILD-TYPE | 16 | 13 | 18 |
Figure S25. Get High-res Image Gene #22: '16q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
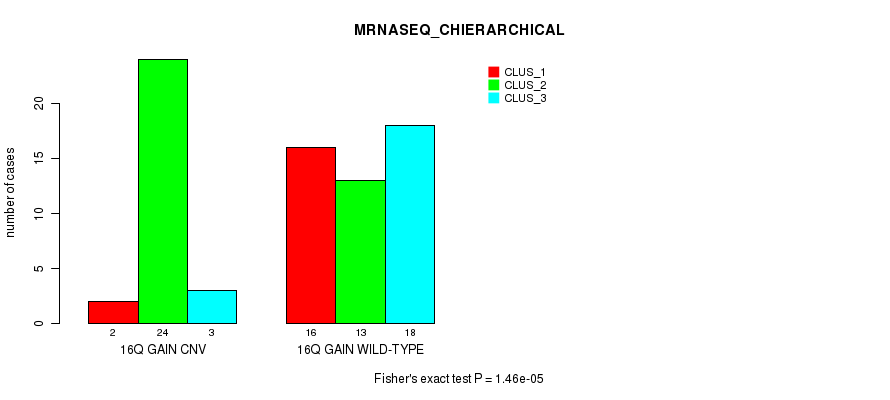
P value = 0.000223 (Fisher's exact test), Q value = 0.083
Table S26. Gene #22: '16q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 27 | 45 | 12 | 33 |
| 16Q GAIN CNV | 8 | 12 | 8 | 23 |
| 16Q GAIN WILD-TYPE | 19 | 33 | 4 | 10 |
Figure S26. Get High-res Image Gene #22: '16q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
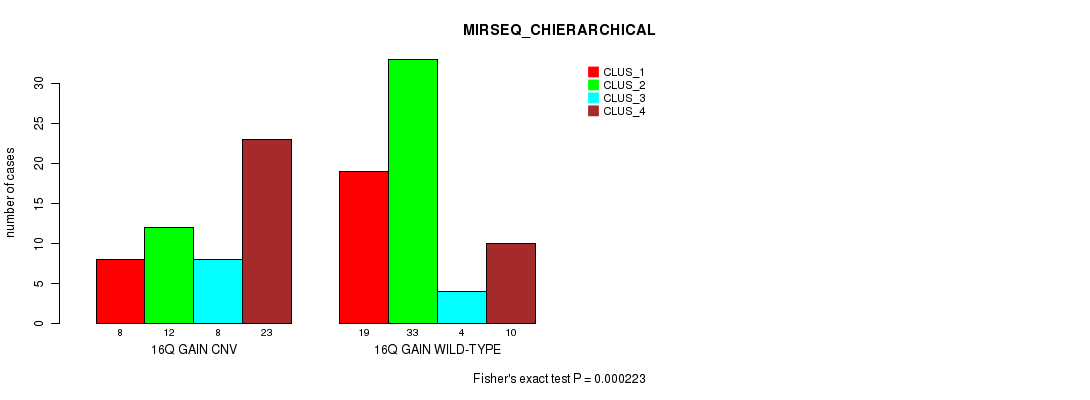
P value = 9.52e-08 (Chi-square test), Q value = 3.8e-05
Table S27. Gene #23: '17p gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 17P GAIN CNV | 6 | 19 | 13 | 9 | 19 |
| 17P GAIN WILD-TYPE | 18 | 25 | 0 | 17 | 1 |
Figure S27. Get High-res Image Gene #23: '17p gain' versus Molecular Subtype #3: 'CN_CNMF'

P value = 1.3e-06 (Fisher's exact test), Q value = 0.00051
Table S28. Gene #23: '17p gain' versus Molecular Subtype #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 53 | 31 |
| 17P GAIN CNV | 9 | 40 | 7 |
| 17P GAIN WILD-TYPE | 18 | 13 | 24 |
Figure S28. Get High-res Image Gene #23: '17p gain' versus Molecular Subtype #4: 'METHLYATION_CNMF'
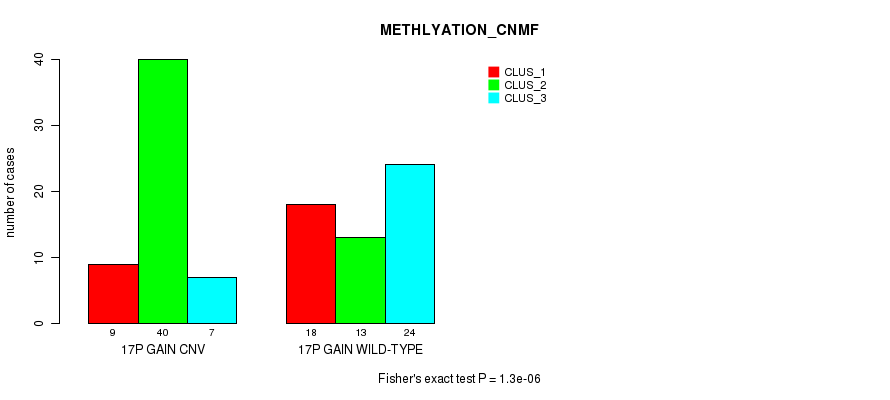
P value = 3.49e-05 (Fisher's exact test), Q value = 0.013
Table S29. Gene #23: '17p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 24 | 25 |
| 17P GAIN CNV | 20 | 3 | 10 |
| 17P GAIN WILD-TYPE | 7 | 21 | 15 |
Figure S29. Get High-res Image Gene #23: '17p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
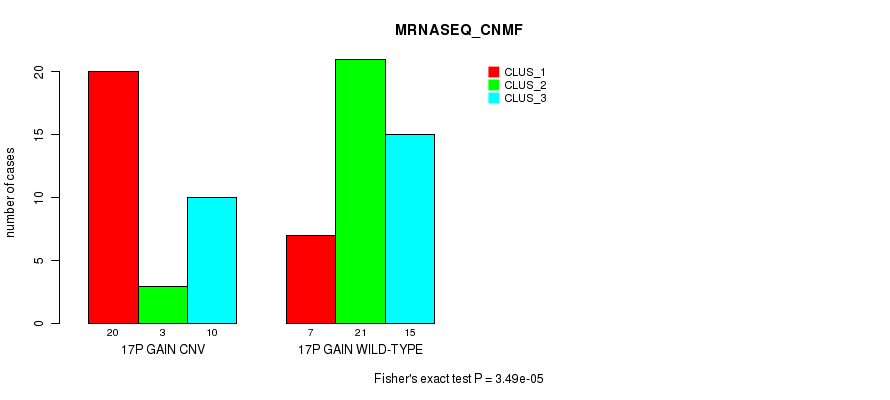
P value = 2.21e-06 (Fisher's exact test), Q value = 0.00087
Table S30. Gene #23: '17p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 37 | 21 |
| 17P GAIN CNV | 3 | 27 | 3 |
| 17P GAIN WILD-TYPE | 15 | 10 | 18 |
Figure S30. Get High-res Image Gene #23: '17p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
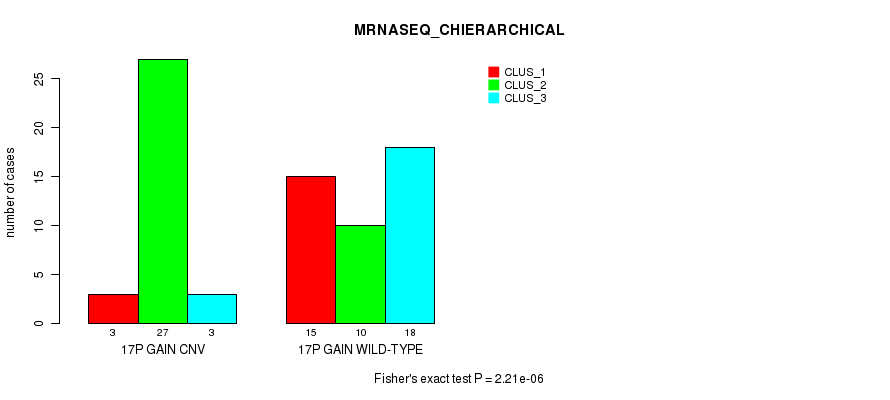
P value = 5.4e-06 (Chi-square test), Q value = 0.0021
Table S31. Gene #24: '17q gain' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 17Q GAIN CNV | 7 | 25 | 13 | 13 | 19 |
| 17Q GAIN WILD-TYPE | 17 | 19 | 0 | 13 | 1 |
Figure S31. Get High-res Image Gene #24: '17q gain' versus Molecular Subtype #3: 'CN_CNMF'
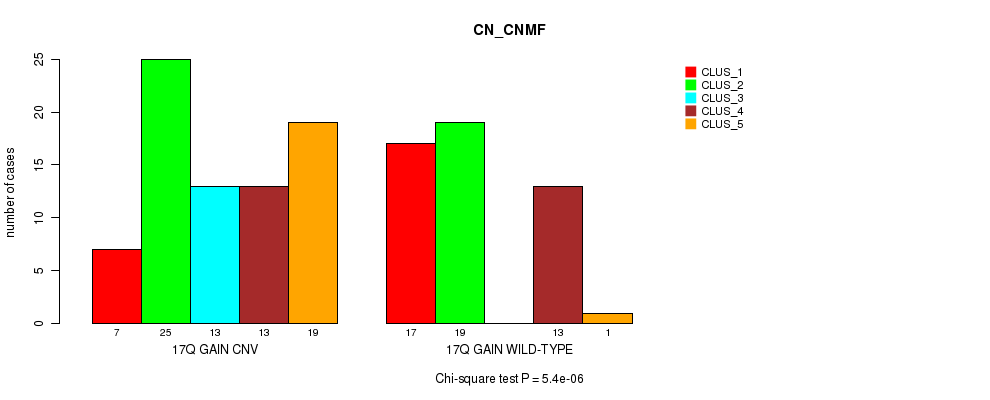
P value = 0.000247 (Chi-square test), Q value = 0.092
Table S32. Gene #35: '4p loss' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 4P LOSS CNV | 7 | 0 | 0 | 4 | 0 |
| 4P LOSS WILD-TYPE | 17 | 44 | 13 | 22 | 20 |
Figure S32. Get High-res Image Gene #35: '4p loss' versus Molecular Subtype #3: 'CN_CNMF'
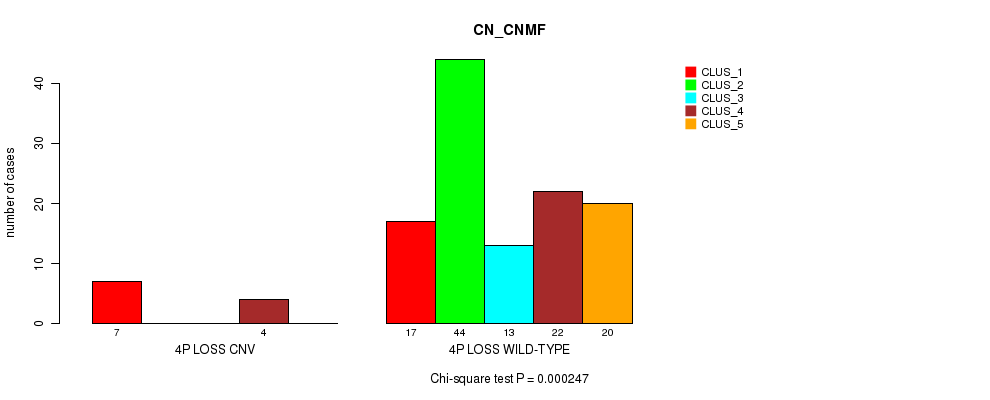
P value = 1.96e-05 (Chi-square test), Q value = 0.0075
Table S33. Gene #38: '5q loss' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 5Q LOSS CNV | 6 | 0 | 0 | 0 | 0 |
| 5Q LOSS WILD-TYPE | 18 | 44 | 13 | 26 | 20 |
Figure S33. Get High-res Image Gene #38: '5q loss' versus Molecular Subtype #3: 'CN_CNMF'
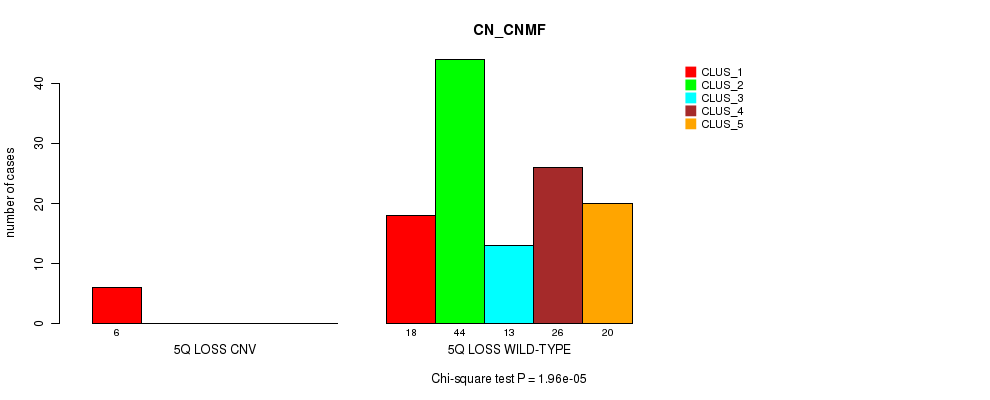
P value = 0.00043 (Chi-square test), Q value = 0.16
Table S34. Gene #39: '6p loss' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 6P LOSS CNV | 1 | 0 | 0 | 7 | 1 |
| 6P LOSS WILD-TYPE | 23 | 44 | 13 | 19 | 19 |
Figure S34. Get High-res Image Gene #39: '6p loss' versus Molecular Subtype #3: 'CN_CNMF'
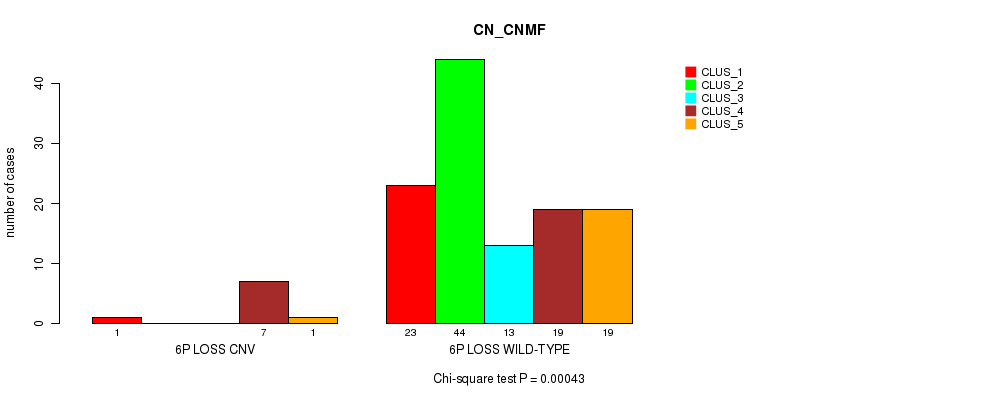
P value = 0.000169 (Chi-square test), Q value = 0.064
Table S35. Gene #42: '9p loss' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 9P LOSS CNV | 8 | 0 | 0 | 6 | 1 |
| 9P LOSS WILD-TYPE | 16 | 44 | 13 | 20 | 19 |
Figure S35. Get High-res Image Gene #42: '9p loss' versus Molecular Subtype #3: 'CN_CNMF'
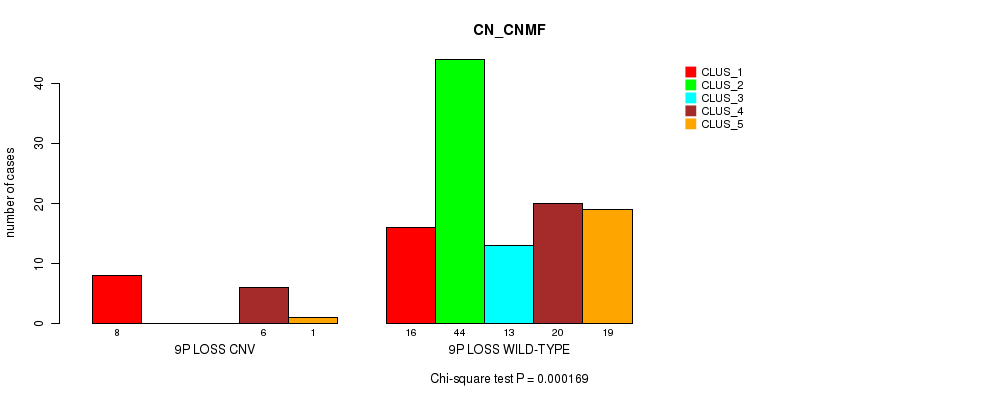
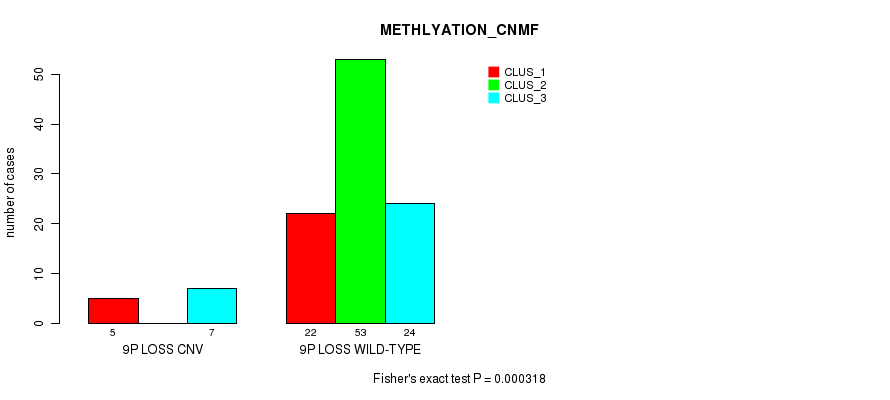
P value = 0.000318 (Fisher's exact test), Q value = 0.12
Table S36. Gene #42: '9p loss' versus Molecular Subtype #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 53 | 31 |
| 9P LOSS CNV | 5 | 0 | 7 |
| 9P LOSS WILD-TYPE | 22 | 53 | 24 |
Figure S36. Get High-res Image Gene #42: '9p loss' versus Molecular Subtype #4: 'METHLYATION_CNMF'
P value = 0.000445 (Chi-square test), Q value = 0.16
Table S37. Gene #43: '9q loss' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 9Q LOSS CNV | 8 | 1 | 0 | 7 | 1 |
| 9Q LOSS WILD-TYPE | 16 | 43 | 13 | 19 | 19 |
Figure S37. Get High-res Image Gene #43: '9q loss' versus Molecular Subtype #3: 'CN_CNMF'

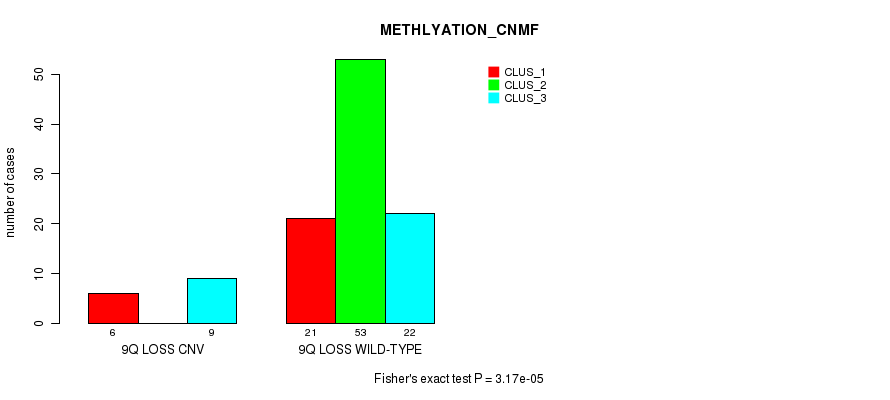
P value = 3.17e-05 (Fisher's exact test), Q value = 0.012
Table S38. Gene #43: '9q loss' versus Molecular Subtype #4: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 53 | 31 |
| 9Q LOSS CNV | 6 | 0 | 9 |
| 9Q LOSS WILD-TYPE | 21 | 53 | 22 |
Figure S38. Get High-res Image Gene #43: '9q loss' versus Molecular Subtype #4: 'METHLYATION_CNMF'
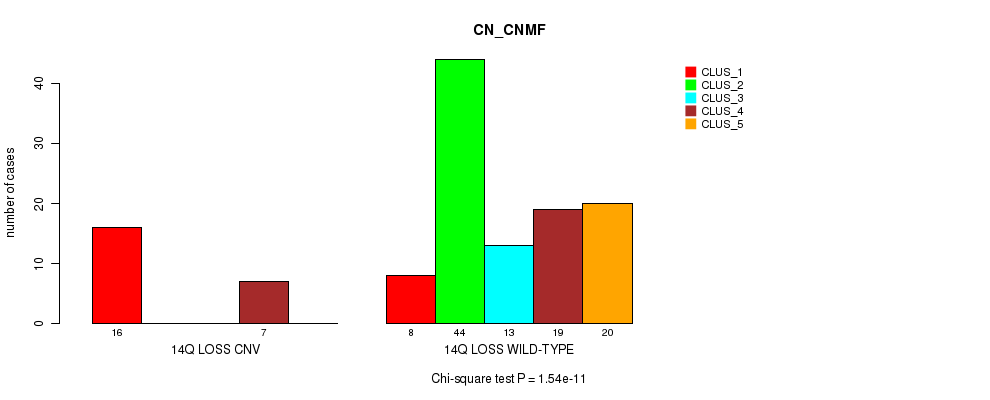
P value = 1.54e-11 (Chi-square test), Q value = 6.2e-09
Table S39. Gene #49: '14q loss' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 14Q LOSS CNV | 16 | 0 | 0 | 7 | 0 |
| 14Q LOSS WILD-TYPE | 8 | 44 | 13 | 19 | 20 |
Figure S39. Get High-res Image Gene #49: '14q loss' versus Molecular Subtype #3: 'CN_CNMF'
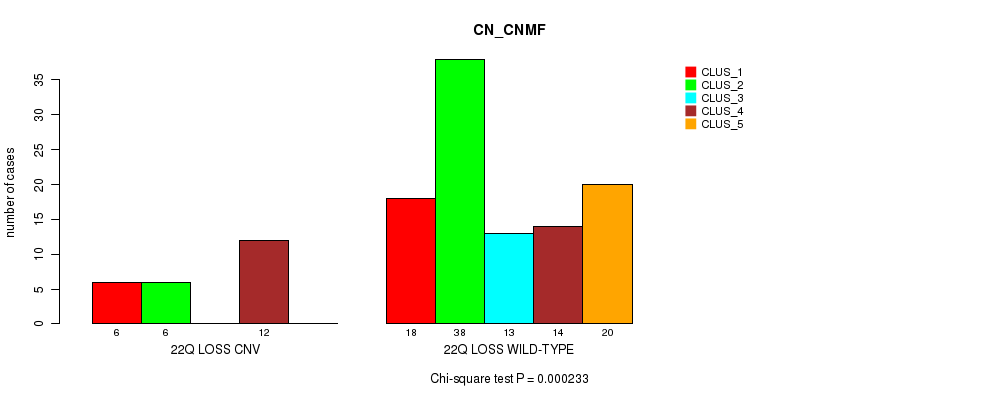
P value = 0.000233 (Chi-square test), Q value = 0.087
Table S40. Gene #58: '22q loss' versus Molecular Subtype #3: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 24 | 44 | 13 | 26 | 20 |
| 22Q LOSS CNV | 6 | 6 | 0 | 12 | 0 |
| 22Q LOSS WILD-TYPE | 18 | 38 | 13 | 14 | 20 |
Figure S40. Get High-res Image Gene #58: '22q loss' versus Molecular Subtype #3: 'CN_CNMF'
-
Mutation data file = broad_values_by_arm.mutsig.cluster.txt
-
Molecular subtypes file = KIRP-TP.transferedmergedcluster.txt
-
Number of patients = 127
-
Number of significantly arm-level cnvs = 59
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.