This pipeline computes the correlation between significant copy number variation (cnv focal) genes and selected clinical features.
Testing the association between copy number variation 66 arm-level results and 10 clinical features across 885 patients, 26 significant findings detected with Q value < 0.25.
-
Amp Peak 5(3q26.32) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Amp Peak 6(4q13.3) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Amp Peak 8(6p23) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Amp Peak 12(8q24.21) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Amp Peak 13(10p15.1) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Amp Peak 15(11p13) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Amp Peak 23(15q26.3) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Amp Peak 25(17q23.1) cnv correlated to 'GENDER'.
-
Amp Peak 27(19q12) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Amp Peak 29(20q13.2) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 4(3p21.31) cnv correlated to 'AGE' and 'HISTOLOGICAL.TYPE'.
-
Del Peak 5(4p16.3) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 6(4q35.2) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 7(5q11.2) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 8(5q21.3) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 11(6q27) cnv correlated to 'AGE'.
-
Del Peak 14(8p23.2) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 16(9p21.3) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 26(13q14.2) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 27(14q24.3) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 28(15q13.1) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 29(16q24.3) cnv correlated to 'AGE' and 'HISTOLOGICAL.TYPE'.
-
Del Peak 33(19p13.3) cnv correlated to 'HISTOLOGICAL.TYPE'.
-
Del Peak 34(19q13.32) cnv correlated to 'HISTOLOGICAL.TYPE'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 66 arm-level results and 10 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 26 significant findings detected.
|
Clinical Features |
Time to Death |
AGE | GENDER |
HISTOLOGICAL TYPE |
RADIATIONS RADIATION REGIMENINDICATION |
DISTANT METASTASIS |
LYMPH NODE METASTASIS |
NUMBER OF LYMPH NODES |
TUMOR STAGECODE |
NEOPLASM DISEASESTAGE |
||
| nCNV (%) | nWild-Type | logrank test | t-test | Fisher's exact test | Chi-square test | Fisher's exact test | Fisher's exact test | Chi-square test | t-test | t-test | Chi-square test | |
| Del Peak 4(3p21 31) | 0 (0%) | 605 |
0.0077 (1.00) |
6.67e-05 (0.0384) |
0.475 (1.00) |
4.84e-05 (0.028) |
0.736 (1.00) |
0.461 (1.00) |
0.729 (1.00) |
0.859 (1.00) |
0.433 (1.00) |
|
| Del Peak 29(16q24 3) | 0 (0%) | 316 |
0.106 (1.00) |
0.00038 (0.217) |
1 (1.00) |
1.68e-15 (9.98e-13) |
0.622 (1.00) |
0.186 (1.00) |
0.402 (1.00) |
0.671 (1.00) |
0.256 (1.00) |
|
| Amp Peak 5(3q26 32) | 0 (0%) | 605 |
0.533 (1.00) |
0.271 (1.00) |
0.727 (1.00) |
1.16e-10 (6.84e-08) |
0.0752 (1.00) |
0.123 (1.00) |
0.822 (1.00) |
0.737 (1.00) |
0.607 (1.00) |
|
| Amp Peak 6(4q13 3) | 0 (0%) | 745 |
0.00151 (0.845) |
0.328 (1.00) |
1 (1.00) |
5.92e-05 (0.0342) |
0.747 (1.00) |
0.717 (1.00) |
0.296 (1.00) |
0.333 (1.00) |
0.292 (1.00) |
|
| Amp Peak 8(6p23) | 0 (0%) | 652 |
0.759 (1.00) |
0.156 (1.00) |
0.459 (1.00) |
9.93e-06 (0.00577) |
0.211 (1.00) |
0.597 (1.00) |
0.197 (1.00) |
0.242 (1.00) |
0.628 (1.00) |
|
| Amp Peak 12(8q24 21) | 0 (0%) | 316 |
0.0711 (1.00) |
0.0721 (1.00) |
0.503 (1.00) |
9.59e-13 (5.69e-10) |
0.367 (1.00) |
0.21 (1.00) |
0.0959 (1.00) |
0.191 (1.00) |
0.398 (1.00) |
|
| Amp Peak 13(10p15 1) | 0 (0%) | 632 |
0.171 (1.00) |
0.888 (1.00) |
0.0668 (1.00) |
4.05e-07 (0.000236) |
0.296 (1.00) |
0.926 (1.00) |
0.88 (1.00) |
0.88 (1.00) |
0.521 (1.00) |
|
| Amp Peak 15(11p13) | 0 (0%) | 700 |
0.554 (1.00) |
0.602 (1.00) |
1 (1.00) |
0.000433 (0.247) |
0.384 (1.00) |
0.457 (1.00) |
0.465 (1.00) |
0.792 (1.00) |
0.349 (1.00) |
|
| Amp Peak 23(15q26 3) | 0 (0%) | 723 |
0.25 (1.00) |
0.0196 (1.00) |
0.0639 (1.00) |
0.00041 (0.234) |
0.761 (1.00) |
0.104 (1.00) |
0.688 (1.00) |
0.559 (1.00) |
0.716 (1.00) |
|
| Amp Peak 25(17q23 1) | 0 (0%) | 517 |
0.0563 (1.00) |
0.48 (1.00) |
0.000351 (0.201) |
0.00387 (1.00) |
0.425 (1.00) |
0.0368 (1.00) |
0.535 (1.00) |
0.602 (1.00) |
0.0717 (1.00) |
|
| Amp Peak 27(19q12) | 0 (0%) | 663 |
0.332 (1.00) |
0.362 (1.00) |
0.699 (1.00) |
0.000181 (0.104) |
0.0842 (1.00) |
0.0487 (1.00) |
0.0993 (1.00) |
0.46 (1.00) |
0.554 (1.00) |
|
| Amp Peak 29(20q13 2) | 0 (0%) | 429 |
0.576 (1.00) |
0.176 (1.00) |
0.179 (1.00) |
1.68e-11 (9.93e-09) |
0.0334 (1.00) |
0.283 (1.00) |
0.277 (1.00) |
0.435 (1.00) |
0.37 (1.00) |
|
| Del Peak 5(4p16 3) | 0 (0%) | 611 |
0.733 (1.00) |
0.0648 (1.00) |
0.729 (1.00) |
1.42e-06 (0.00083) |
0.609 (1.00) |
0.834 (1.00) |
0.839 (1.00) |
0.848 (1.00) |
0.412 (1.00) |
|
| Del Peak 6(4q35 2) | 0 (0%) | 608 |
0.0847 (1.00) |
0.933 (1.00) |
0.287 (1.00) |
1.46e-05 (0.00844) |
0.932 (1.00) |
0.0796 (1.00) |
0.475 (1.00) |
0.869 (1.00) |
0.864 (1.00) |
|
| Del Peak 7(5q11 2) | 0 (0%) | 657 |
0.198 (1.00) |
0.0038 (1.00) |
0.46 (1.00) |
2.26e-10 (1.33e-07) |
0.53 (1.00) |
0.197 (1.00) |
0.607 (1.00) |
0.231 (1.00) |
0.476 (1.00) |
|
| Del Peak 8(5q21 3) | 0 (0%) | 667 |
0.31 (1.00) |
0.0108 (1.00) |
0.123 (1.00) |
2.71e-12 (1.6e-09) |
0.927 (1.00) |
0.447 (1.00) |
0.298 (1.00) |
0.000491 (0.279) |
0.267 (1.00) |
|
| Del Peak 11(6q27) | 0 (0%) | 583 |
0.113 (1.00) |
1.6e-05 (0.00925) |
0.178 (1.00) |
0.32 (1.00) |
0.561 (1.00) |
0.0802 (1.00) |
0.305 (1.00) |
0.349 (1.00) |
0.516 (1.00) |
|
| Del Peak 14(8p23 2) | 0 (0%) | 418 |
0.235 (1.00) |
0.613 (1.00) |
0.512 (1.00) |
7.95e-08 (4.65e-05) |
0.237 (1.00) |
0.693 (1.00) |
0.178 (1.00) |
0.761 (1.00) |
0.98 (1.00) |
|
| Del Peak 16(9p21 3) | 0 (0%) | 583 |
0.0222 (1.00) |
0.968 (1.00) |
0.287 (1.00) |
3.13e-08 (1.84e-05) |
0.245 (1.00) |
0.134 (1.00) |
0.000852 (0.479) |
0.144 (1.00) |
0.00584 (1.00) |
|
| Del Peak 26(13q14 2) | 0 (0%) | 477 |
0.0668 (1.00) |
0.674 (1.00) |
0.74 (1.00) |
0.000113 (0.065) |
0.752 (1.00) |
0.976 (1.00) |
0.489 (1.00) |
0.118 (1.00) |
0.963 (1.00) |
|
| Del Peak 27(14q24 3) | 0 (0%) | 604 |
0.151 (1.00) |
0.637 (1.00) |
1 (1.00) |
6.51e-09 (3.83e-06) |
0.352 (1.00) |
0.711 (1.00) |
0.124 (1.00) |
0.682 (1.00) |
0.846 (1.00) |
|
| Del Peak 28(15q13 1) | 0 (0%) | 578 |
0.353 (1.00) |
0.147 (1.00) |
1 (1.00) |
0.000267 (0.153) |
0.564 (1.00) |
0.204 (1.00) |
0.214 (1.00) |
0.842 (1.00) |
0.451 (1.00) |
|
| Del Peak 33(19p13 3) | 0 (0%) | 583 |
0.343 (1.00) |
0.0154 (1.00) |
0.726 (1.00) |
5.43e-08 (3.18e-05) |
0.868 (1.00) |
0.604 (1.00) |
0.762 (1.00) |
0.152 (1.00) |
0.937 (1.00) |
|
| Del Peak 34(19q13 32) | 0 (0%) | 708 |
0.745 (1.00) |
0.642 (1.00) |
0.696 (1.00) |
8.07e-06 (0.0047) |
0.491 (1.00) |
0.293 (1.00) |
0.216 (1.00) |
0.905 (1.00) |
0.392 (1.00) |
|
| Amp Peak 1(1p22 3) | 0 (0%) | 691 |
0.982 (1.00) |
0.806 (1.00) |
1 (1.00) |
0.341 (1.00) |
0.776 (1.00) |
0.519 (1.00) |
0.348 (1.00) |
0.119 (1.00) |
0.641 (1.00) |
|
| Amp Peak 2(1q21 3) | 0 (0%) | 252 |
0.713 (1.00) |
0.543 (1.00) |
0.284 (1.00) |
0.809 (1.00) |
0.163 (1.00) |
0.448 (1.00) |
0.567 (1.00) |
0.672 (1.00) |
0.362 (1.00) |
|
| Amp Peak 3(1q44) | 0 (0%) | 253 |
0.00866 (1.00) |
0.798 (1.00) |
0.72 (1.00) |
0.154 (1.00) |
0.339 (1.00) |
0.551 (1.00) |
0.308 (1.00) |
0.748 (1.00) |
0.522 (1.00) |
|
| Amp Peak 4(3p26 1) | 0 (0%) | 700 |
0.0038 (1.00) |
0.785 (1.00) |
0.217 (1.00) |
0.00449 (1.00) |
0.772 (1.00) |
0.0281 (1.00) |
0.121 (1.00) |
0.627 (1.00) |
0.128 (1.00) |
|
| Amp Peak 7(5p15 33) | 0 (0%) | 567 |
0.143 (1.00) |
0.237 (1.00) |
1 (1.00) |
0.0602 (1.00) |
0.163 (1.00) |
0.267 (1.00) |
0.0183 (1.00) |
0.574 (1.00) |
0.758 (1.00) |
|
| Amp Peak 9(6q21) | 0 (0%) | 721 |
0.0119 (1.00) |
0.421 (1.00) |
1 (1.00) |
0.000586 (0.332) |
0.612 (1.00) |
0.868 (1.00) |
0.726 (1.00) |
0.791 (1.00) |
0.251 (1.00) |
|
| Amp Peak 10(8p11 23) | 0 (0%) | 547 |
0.0163 (1.00) |
0.11 (1.00) |
0.165 (1.00) |
0.556 (1.00) |
0.419 (1.00) |
0.0789 (1.00) |
0.199 (1.00) |
0.0101 (1.00) |
0.0352 (1.00) |
|
| Amp Peak 11(8p11 21) | 0 (0%) | 543 |
0.696 (1.00) |
0.0617 (1.00) |
0.165 (1.00) |
0.071 (1.00) |
0.0238 (1.00) |
0.644 (1.00) |
0.634 (1.00) |
0.503 (1.00) |
0.468 (1.00) |
|
| Amp Peak 14(10q22 3) | 0 (0%) | 754 |
0.0996 (1.00) |
0.59 (1.00) |
1 (1.00) |
0.0113 (1.00) |
0.438 (1.00) |
0.104 (1.00) |
0.617 (1.00) |
0.666 (1.00) |
0.318 (1.00) |
|
| Amp Peak 16(11q13 3) | 0 (0%) | 549 |
0.248 (1.00) |
0.33 (1.00) |
0.312 (1.00) |
0.506 (1.00) |
0.571 (1.00) |
0.427 (1.00) |
0.625 (1.00) |
0.412 (1.00) |
0.378 (1.00) |
|
| Amp Peak 17(11q14 1) | 0 (0%) | 658 |
0.16 (1.00) |
0.33 (1.00) |
0.701 (1.00) |
0.0232 (1.00) |
0.857 (1.00) |
0.224 (1.00) |
0.72 (1.00) |
0.403 (1.00) |
0.246 (1.00) |
|
| Amp Peak 18(12p13 33) | 0 (0%) | 656 |
0.728 (1.00) |
0.42 (1.00) |
0.46 (1.00) |
0.00238 (1.00) |
0.0724 (1.00) |
0.635 (1.00) |
0.323 (1.00) |
0.904 (1.00) |
0.349 (1.00) |
|
| Amp Peak 19(12q15) | 0 (0%) | 655 |
0.0289 (1.00) |
0.396 (1.00) |
0.0118 (1.00) |
0.0471 (1.00) |
0.928 (1.00) |
0.266 (1.00) |
0.0769 (1.00) |
0.228 (1.00) |
0.215 (1.00) |
|
| Amp Peak 20(13q12 3) | 0 (0%) | 759 |
0.0281 (1.00) |
0.209 (1.00) |
0.624 (1.00) |
0.013 (1.00) |
0.43 (1.00) |
0.14 (1.00) |
0.13 (1.00) |
0.605 (1.00) |
0.592 (1.00) |
|
| Amp Peak 21(13q34) | 0 (0%) | 733 |
0.9 (1.00) |
0.0661 (1.00) |
0.371 (1.00) |
0.0209 (1.00) |
1 (1.00) |
0.277 (1.00) |
0.205 (1.00) |
0.784 (1.00) |
0.179 (1.00) |
|
| Amp Peak 22(14q21 1) | 0 (0%) | 704 |
0.357 (1.00) |
0.0239 (1.00) |
0.695 (1.00) |
0.165 (1.00) |
0.697 (1.00) |
0.539 (1.00) |
0.0158 (1.00) |
0.0546 (1.00) |
0.0415 (1.00) |
|
| Amp Peak 24(17q12) | 0 (0%) | 597 |
0.146 (1.00) |
0.997 (1.00) |
0.000759 (0.428) |
0.0078 (1.00) |
1 (1.00) |
0.358 (1.00) |
0.0556 (1.00) |
0.44 (1.00) |
0.367 (1.00) |
|
| Amp Peak 26(19p13 12) | 0 (0%) | 701 |
0.704 (1.00) |
0.806 (1.00) |
0.0954 (1.00) |
0.119 (1.00) |
0.0254 (1.00) |
0.168 (1.00) |
0.351 (1.00) |
0.653 (1.00) |
0.788 (1.00) |
|
| Amp Peak 28(19q13 42) | 0 (0%) | 653 |
0.543 (1.00) |
0.148 (1.00) |
0.252 (1.00) |
0.131 (1.00) |
0.326 (1.00) |
0.0476 (1.00) |
0.585 (1.00) |
0.819 (1.00) |
0.981 (1.00) |
|
| Del Peak 1(1p36 13) | 0 (0%) | 512 |
0.179 (1.00) |
0.627 (1.00) |
0.178 (1.00) |
0.0268 (1.00) |
1 (1.00) |
0.277 (1.00) |
0.057 (1.00) |
0.0309 (1.00) |
0.0215 (1.00) |
|
| Del Peak 2(1p22 1) | 0 (0%) | 584 |
0.00252 (1.00) |
0.284 (1.00) |
0.499 (1.00) |
0.0809 (1.00) |
0.185 (1.00) |
0.592 (1.00) |
0.0103 (1.00) |
0.0251 (1.00) |
0.0195 (1.00) |
|
| Del Peak 3(2q37 3) | 0 (0%) | 627 |
0.954 (1.00) |
0.82 (1.00) |
0.46 (1.00) |
0.00211 (1.00) |
0.387 (1.00) |
0.569 (1.00) |
0.378 (1.00) |
0.601 (1.00) |
0.533 (1.00) |
|
| Del Peak 9(6p25 3) | 0 (0%) | 719 |
0.524 (1.00) |
0.0322 (1.00) |
0.679 (1.00) |
0.543 (1.00) |
0.158 (1.00) |
0.603 (1.00) |
0.544 (1.00) |
0.0286 (1.00) |
0.683 (1.00) |
|
| Del Peak 10(6q15) | 0 (0%) | 593 |
0.348 (1.00) |
0.057 (1.00) |
0.726 (1.00) |
0.611 (1.00) |
0.933 (1.00) |
0.551 (1.00) |
0.495 (1.00) |
0.115 (1.00) |
0.258 (1.00) |
|
| Del Peak 12(7p22 3) | 0 (0%) | 767 |
0.102 (1.00) |
0.238 (1.00) |
0.616 (1.00) |
0.000723 (0.408) |
0.728 (1.00) |
0.718 (1.00) |
0.797 (1.00) |
0.0255 (1.00) |
0.0715 (1.00) |
|
| Del Peak 13(7q36 1) | 0 (0%) | 710 |
0.0125 (1.00) |
0.932 (1.00) |
0.694 (1.00) |
0.0213 (1.00) |
0.374 (1.00) |
0.357 (1.00) |
0.831 (1.00) |
0.774 (1.00) |
0.908 (1.00) |
|
| Del Peak 15(8p11 21) | 0 (0%) | 714 |
0.0139 (1.00) |
0.307 (1.00) |
0.219 (1.00) |
0.0163 (1.00) |
0.765 (1.00) |
0.47 (1.00) |
0.0738 (1.00) |
0.264 (1.00) |
0.647 (1.00) |
|
| Del Peak 17(9q21 11) | 0 (0%) | 643 |
0.456 (1.00) |
0.991 (1.00) |
0.267 (1.00) |
0.000901 (0.506) |
0.724 (1.00) |
0.331 (1.00) |
0.296 (1.00) |
0.662 (1.00) |
0.355 (1.00) |
|
| Del Peak 18(10q23 31) | 0 (0%) | 631 |
0.808 (1.00) |
0.485 (1.00) |
0.0666 (1.00) |
0.00478 (1.00) |
0.487 (1.00) |
0.712 (1.00) |
0.399 (1.00) |
0.956 (1.00) |
0.722 (1.00) |
|
| Del Peak 19(10q26 3) | 0 (0%) | 627 |
0.0341 (1.00) |
0.814 (1.00) |
0.46 (1.00) |
0.0006 (0.34) |
0.729 (1.00) |
0.569 (1.00) |
0.237 (1.00) |
0.605 (1.00) |
0.559 (1.00) |
|
| Del Peak 20(11p15 5) | 0 (0%) | 628 |
0.316 (1.00) |
0.745 (1.00) |
0.293 (1.00) |
0.0224 (1.00) |
1 (1.00) |
0.968 (1.00) |
0.686 (1.00) |
0.143 (1.00) |
0.462 (1.00) |
|
| Del Peak 21(11q13 1) | 0 (0%) | 694 |
0.469 (1.00) |
0.96 (1.00) |
0.415 (1.00) |
0.674 (1.00) |
0.504 (1.00) |
0.659 (1.00) |
0.209 (1.00) |
0.746 (1.00) |
0.488 (1.00) |
|
| Del Peak 22(11q23 2) | 0 (0%) | 445 |
0.455 (1.00) |
0.31 (1.00) |
0.106 (1.00) |
0.171 (1.00) |
1 (1.00) |
0.564 (1.00) |
0.124 (1.00) |
0.61 (1.00) |
0.184 (1.00) |
|
| Del Peak 23(11q25) | 0 (0%) | 473 |
0.287 (1.00) |
0.0155 (1.00) |
0.0902 (1.00) |
0.0347 (1.00) |
0.528 (1.00) |
0.63 (1.00) |
0.385 (1.00) |
0.631 (1.00) |
0.09 (1.00) |
|
| Del Peak 24(12p13 1) | 0 (0%) | 731 |
0.641 (1.00) |
0.548 (1.00) |
0.66 (1.00) |
0.0031 (1.00) |
0.468 (1.00) |
0.0293 (1.00) |
0.403 (1.00) |
0.552 (1.00) |
0.282 (1.00) |
|
| Del Peak 25(12q24 21) | 0 (0%) | 718 |
0.58 (1.00) |
0.157 (1.00) |
1 (1.00) |
0.253 (1.00) |
0.615 (1.00) |
0.831 (1.00) |
0.165 (1.00) |
0.0754 (1.00) |
0.595 (1.00) |
|
| Del Peak 30(17p12) | 0 (0%) | 349 |
0.177 (1.00) |
0.00279 (1.00) |
0.495 (1.00) |
0.00972 (1.00) |
0.629 (1.00) |
0.95 (1.00) |
0.0287 (1.00) |
0.326 (1.00) |
0.738 (1.00) |
|
| Del Peak 31(17q21 31) | 0 (0%) | 577 |
0.483 (1.00) |
0.000759 (0.428) |
0.174 (1.00) |
0.00105 (0.586) |
0.934 (1.00) |
0.159 (1.00) |
0.0435 (1.00) |
0.137 (1.00) |
0.343 (1.00) |
|
| Del Peak 32(18q23) | 0 (0%) | 600 |
0.0143 (1.00) |
0.521 (1.00) |
1 (1.00) |
0.012 (1.00) |
0.206 (1.00) |
0.54 (1.00) |
0.0518 (1.00) |
0.179 (1.00) |
0.197 (1.00) |
|
| Del Peak 35(20p13) | 0 (0%) | 754 |
0.0111 (1.00) |
0.283 (1.00) |
0.629 (1.00) |
0.00277 (1.00) |
0.658 (1.00) |
0.549 (1.00) |
0.97 (1.00) |
0.828 (1.00) |
0.502 (1.00) |
|
| Del Peak 36(21q11 2) | 0 (0%) | 706 |
0.306 (1.00) |
0.427 (1.00) |
0.217 (1.00) |
0.013 (1.00) |
0.281 (1.00) |
0.552 (1.00) |
0.24 (1.00) |
0.811 (1.00) |
0.0188 (1.00) |
|
| Del Peak 37(22q11 1) | 0 (0%) | 515 |
0.311 (1.00) |
0.805 (1.00) |
0.318 (1.00) |
0.00696 (1.00) |
0.0253 (1.00) |
0.34 (1.00) |
0.347 (1.00) |
0.537 (1.00) |
0.841 (1.00) |
P value = 1.16e-10 (Chi-square test), Q value = 6.8e-08
Table S1. Gene #5: 'Amp Peak 5(3q26.32)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| AMP PEAK 5(3Q26.32) CNV | 251 | 9 | 4 | 5 | 1 | 10 |
| AMP PEAK 5(3Q26.32) WILD-TYPE | 431 | 112 | 0 | 21 | 10 | 31 |
Figure S1. Get High-res Image Gene #5: 'Amp Peak 5(3q26.32)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
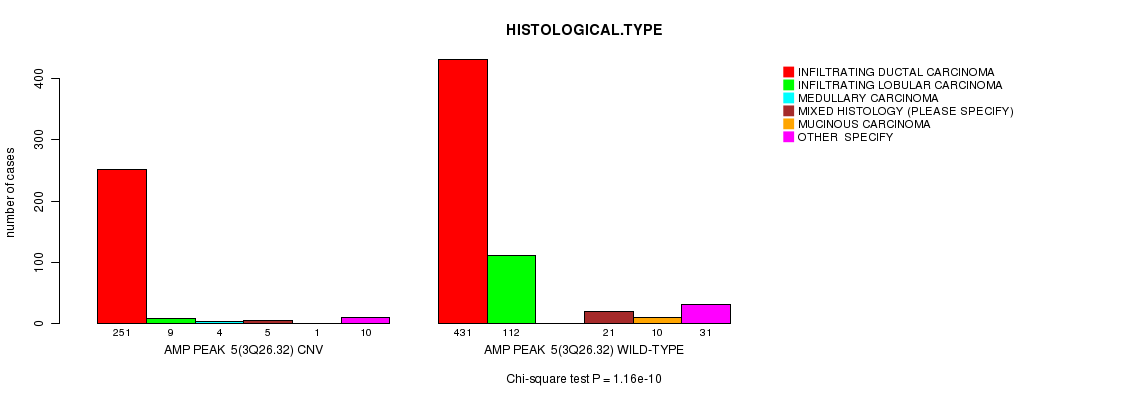
P value = 5.92e-05 (Chi-square test), Q value = 0.034
Table S2. Gene #6: 'Amp Peak 6(4q13.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| AMP PEAK 6(4Q13.3) CNV | 127 | 3 | 2 | 1 | 1 | 6 |
| AMP PEAK 6(4Q13.3) WILD-TYPE | 555 | 118 | 2 | 25 | 10 | 35 |
Figure S2. Get High-res Image Gene #6: 'Amp Peak 6(4q13.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
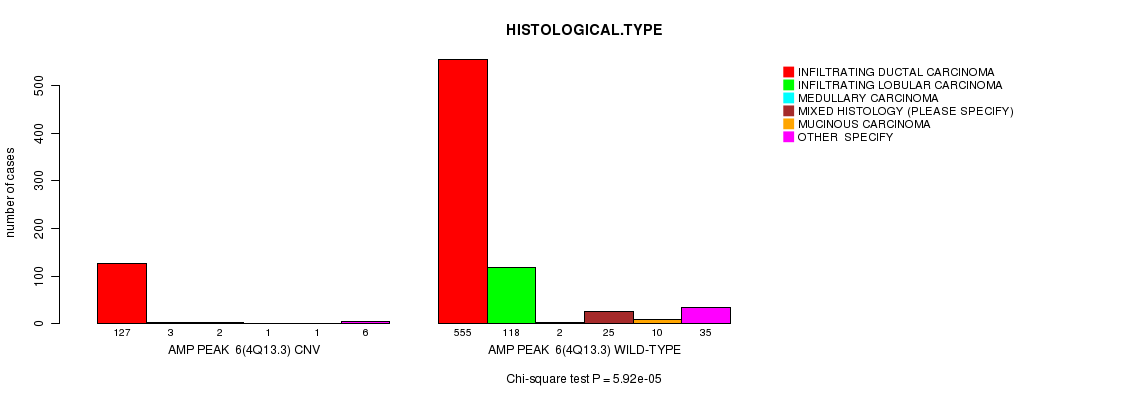
P value = 9.93e-06 (Chi-square test), Q value = 0.0058
Table S3. Gene #8: 'Amp Peak 8(6p23)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| AMP PEAK 8(6P23) CNV | 203 | 11 | 3 | 4 | 1 | 11 |
| AMP PEAK 8(6P23) WILD-TYPE | 479 | 110 | 1 | 22 | 10 | 30 |
Figure S3. Get High-res Image Gene #8: 'Amp Peak 8(6p23)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
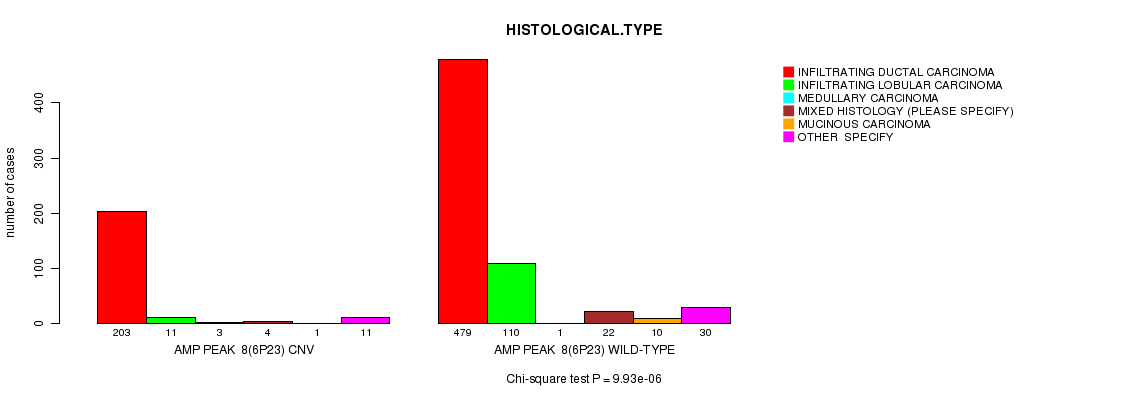
P value = 9.59e-13 (Chi-square test), Q value = 5.7e-10
Table S4. Gene #12: 'Amp Peak 12(8q24.21)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| AMP PEAK 12(8Q24.21) CNV | 481 | 42 | 4 | 15 | 4 | 23 |
| AMP PEAK 12(8Q24.21) WILD-TYPE | 201 | 79 | 0 | 11 | 7 | 18 |
Figure S4. Get High-res Image Gene #12: 'Amp Peak 12(8q24.21)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
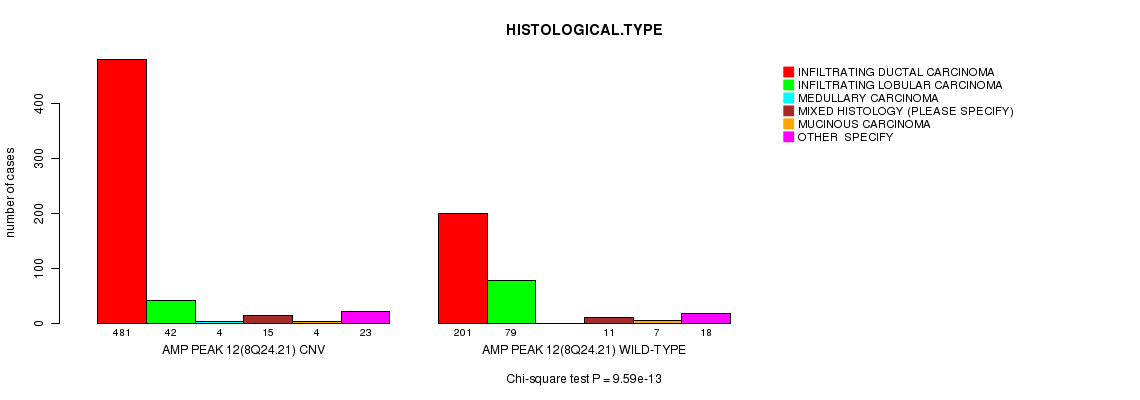
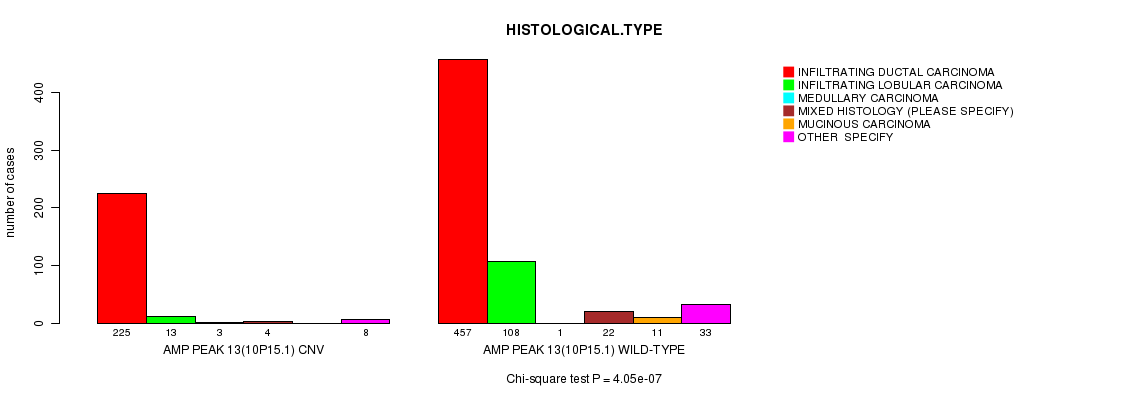
P value = 4.05e-07 (Chi-square test), Q value = 0.00024
Table S5. Gene #13: 'Amp Peak 13(10p15.1)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| AMP PEAK 13(10P15.1) CNV | 225 | 13 | 3 | 4 | 0 | 8 |
| AMP PEAK 13(10P15.1) WILD-TYPE | 457 | 108 | 1 | 22 | 11 | 33 |
Figure S5. Get High-res Image Gene #13: 'Amp Peak 13(10p15.1)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
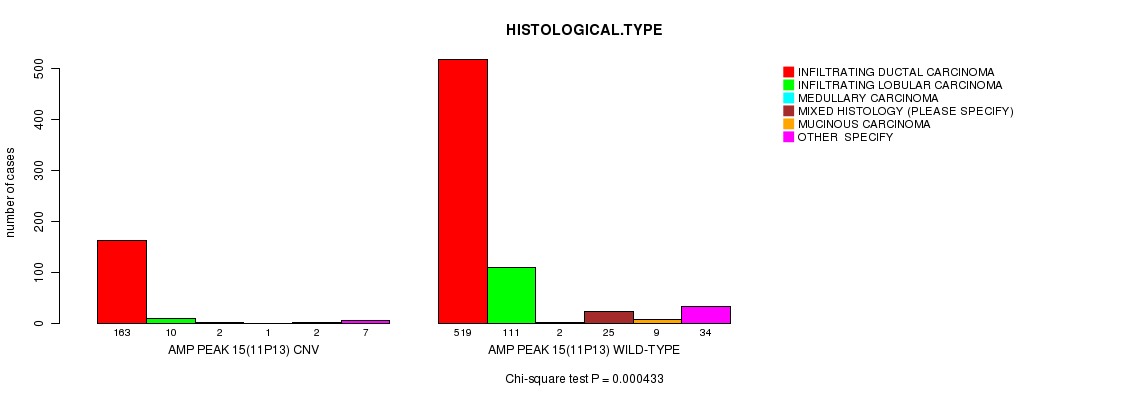
P value = 0.000433 (Chi-square test), Q value = 0.25
Table S6. Gene #15: 'Amp Peak 15(11p13)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| AMP PEAK 15(11P13) CNV | 163 | 10 | 2 | 1 | 2 | 7 |
| AMP PEAK 15(11P13) WILD-TYPE | 519 | 111 | 2 | 25 | 9 | 34 |
Figure S6. Get High-res Image Gene #15: 'Amp Peak 15(11p13)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
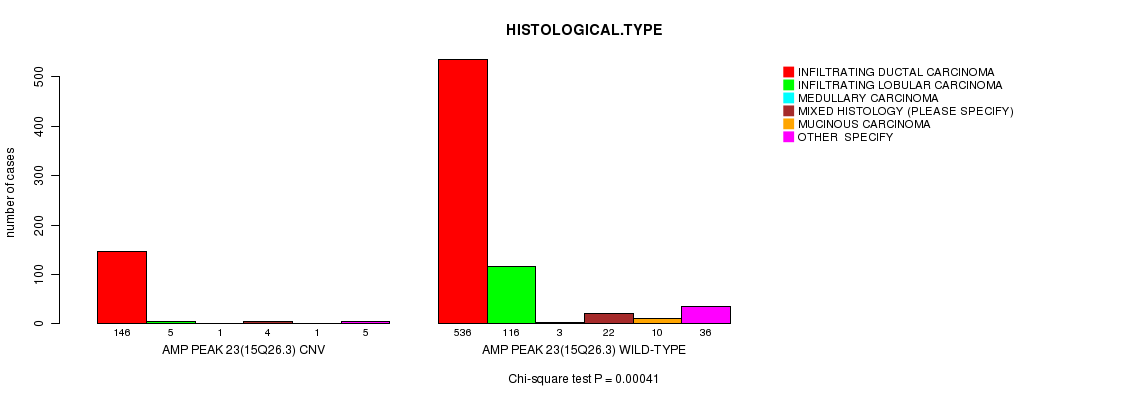
P value = 0.00041 (Chi-square test), Q value = 0.23
Table S7. Gene #23: 'Amp Peak 23(15q26.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| AMP PEAK 23(15Q26.3) CNV | 146 | 5 | 1 | 4 | 1 | 5 |
| AMP PEAK 23(15Q26.3) WILD-TYPE | 536 | 116 | 3 | 22 | 10 | 36 |
Figure S7. Get High-res Image Gene #23: 'Amp Peak 23(15q26.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
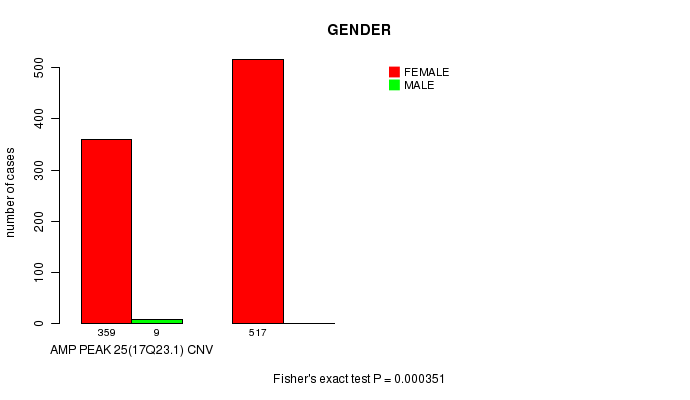
P value = 0.000351 (Fisher's exact test), Q value = 0.2
Table S8. Gene #25: 'Amp Peak 25(17q23.1)' versus Clinical Feature #3: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 876 | 9 |
| AMP PEAK 25(17Q23.1) CNV | 359 | 9 |
| AMP PEAK 25(17Q23.1) WILD-TYPE | 517 | 0 |
Figure S8. Get High-res Image Gene #25: 'Amp Peak 25(17q23.1)' versus Clinical Feature #3: 'GENDER'
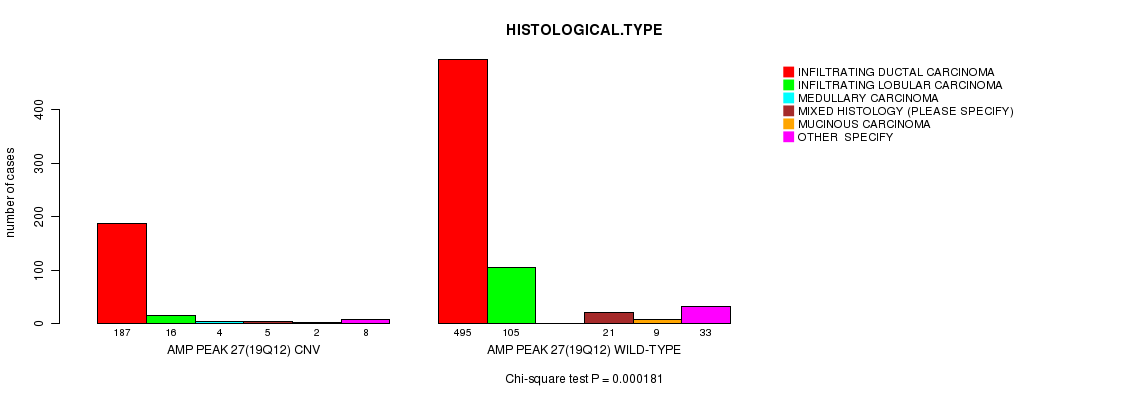
P value = 0.000181 (Chi-square test), Q value = 0.1
Table S9. Gene #27: 'Amp Peak 27(19q12)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| AMP PEAK 27(19Q12) CNV | 187 | 16 | 4 | 5 | 2 | 8 |
| AMP PEAK 27(19Q12) WILD-TYPE | 495 | 105 | 0 | 21 | 9 | 33 |
Figure S9. Get High-res Image Gene #27: 'Amp Peak 27(19q12)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
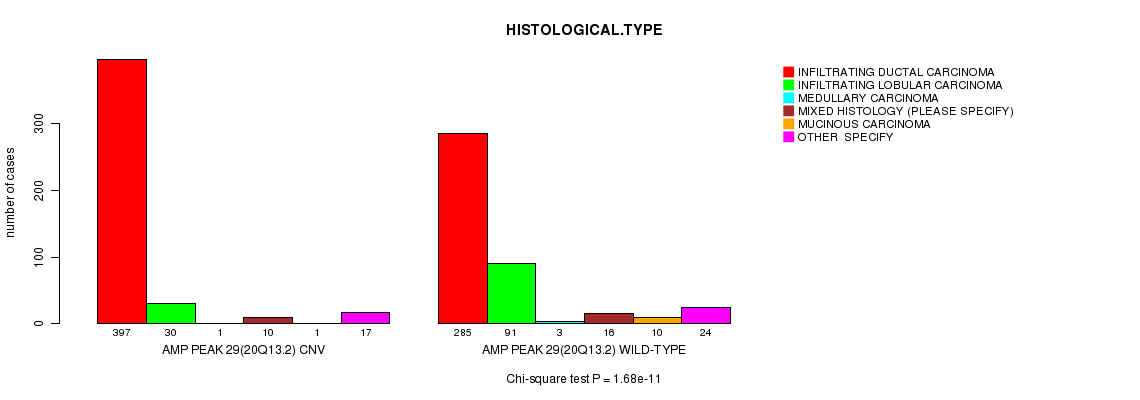
P value = 1.68e-11 (Chi-square test), Q value = 9.9e-09
Table S10. Gene #29: 'Amp Peak 29(20q13.2)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| AMP PEAK 29(20Q13.2) CNV | 397 | 30 | 1 | 10 | 1 | 17 |
| AMP PEAK 29(20Q13.2) WILD-TYPE | 285 | 91 | 3 | 16 | 10 | 24 |
Figure S10. Get High-res Image Gene #29: 'Amp Peak 29(20q13.2)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
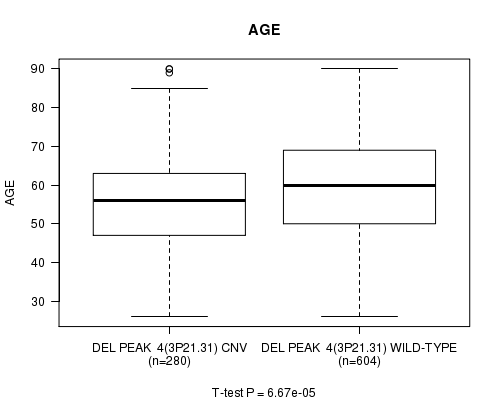
P value = 6.67e-05 (t-test), Q value = 0.038
Table S11. Gene #33: 'Del Peak 4(3p21.31)' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 884 | 58.6 (13.2) |
| DEL PEAK 4(3P21.31) CNV | 280 | 55.9 (13.3) |
| DEL PEAK 4(3P21.31) WILD-TYPE | 604 | 59.8 (13.0) |
Figure S11. Get High-res Image Gene #33: 'Del Peak 4(3p21.31)' versus Clinical Feature #2: 'AGE'
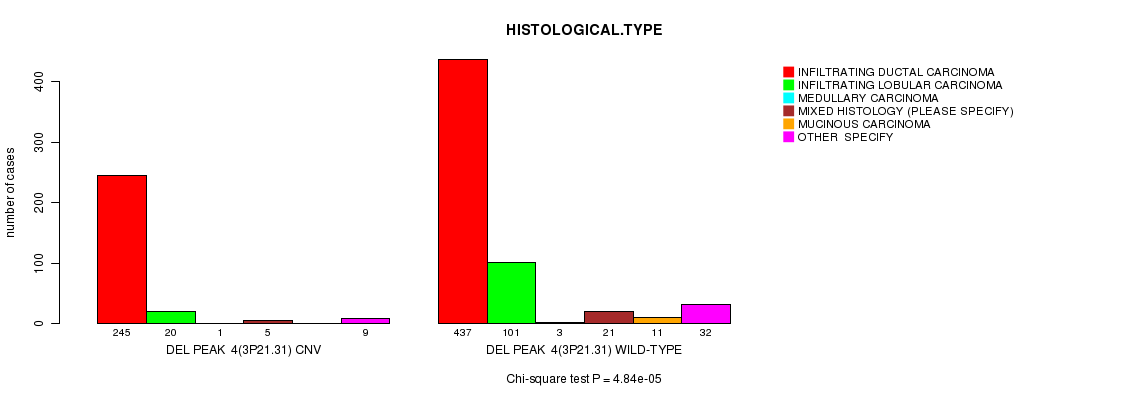
P value = 4.84e-05 (Chi-square test), Q value = 0.028
Table S12. Gene #33: 'Del Peak 4(3p21.31)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 4(3P21.31) CNV | 245 | 20 | 1 | 5 | 0 | 9 |
| DEL PEAK 4(3P21.31) WILD-TYPE | 437 | 101 | 3 | 21 | 11 | 32 |
Figure S12. Get High-res Image Gene #33: 'Del Peak 4(3p21.31)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
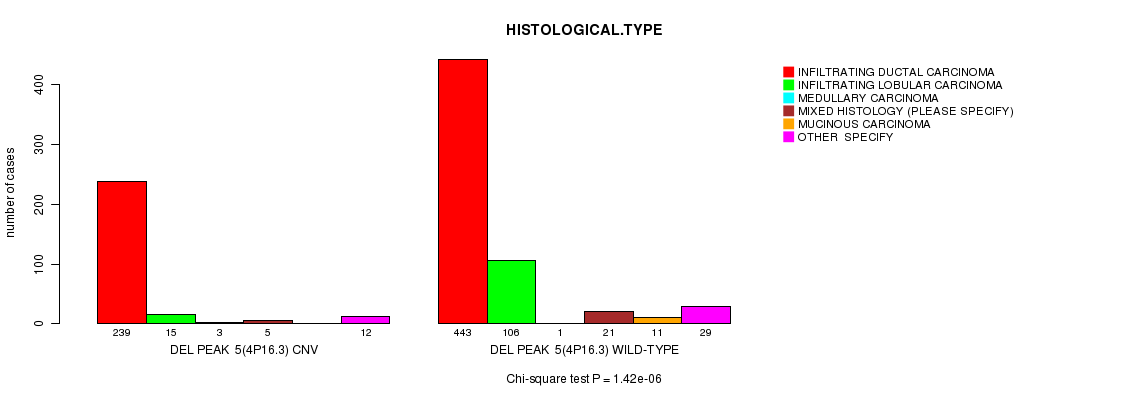
P value = 1.42e-06 (Chi-square test), Q value = 0.00083
Table S13. Gene #34: 'Del Peak 5(4p16.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 5(4P16.3) CNV | 239 | 15 | 3 | 5 | 0 | 12 |
| DEL PEAK 5(4P16.3) WILD-TYPE | 443 | 106 | 1 | 21 | 11 | 29 |
Figure S13. Get High-res Image Gene #34: 'Del Peak 5(4p16.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
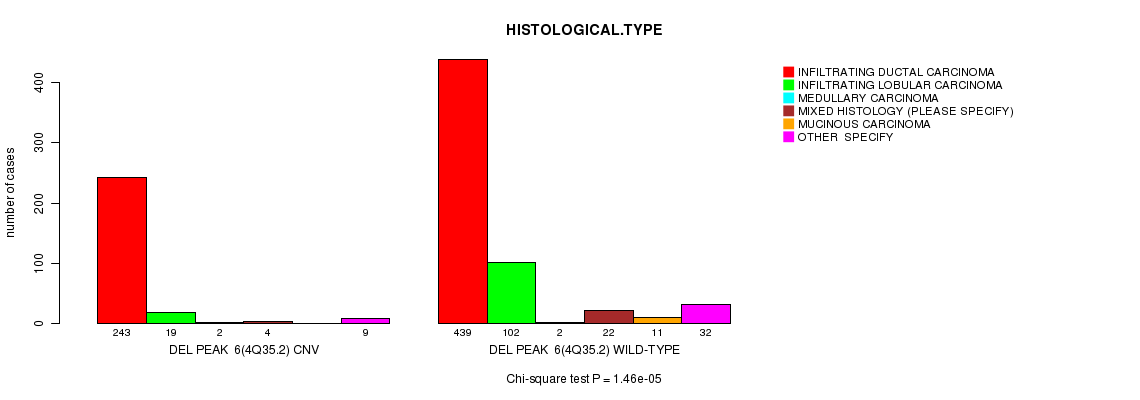
P value = 1.46e-05 (Chi-square test), Q value = 0.0084
Table S14. Gene #35: 'Del Peak 6(4q35.2)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 6(4Q35.2) CNV | 243 | 19 | 2 | 4 | 0 | 9 |
| DEL PEAK 6(4Q35.2) WILD-TYPE | 439 | 102 | 2 | 22 | 11 | 32 |
Figure S14. Get High-res Image Gene #35: 'Del Peak 6(4q35.2)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
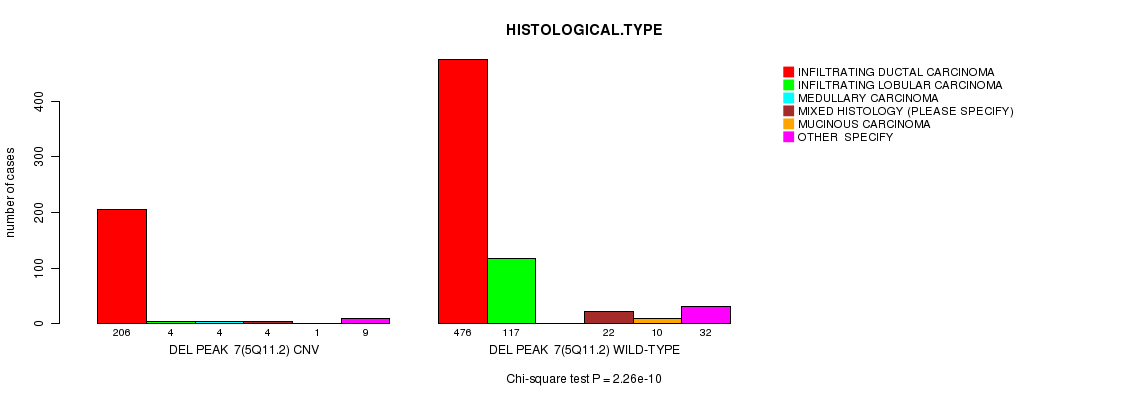
P value = 2.26e-10 (Chi-square test), Q value = 1.3e-07
Table S15. Gene #36: 'Del Peak 7(5q11.2)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 7(5Q11.2) CNV | 206 | 4 | 4 | 4 | 1 | 9 |
| DEL PEAK 7(5Q11.2) WILD-TYPE | 476 | 117 | 0 | 22 | 10 | 32 |
Figure S15. Get High-res Image Gene #36: 'Del Peak 7(5q11.2)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
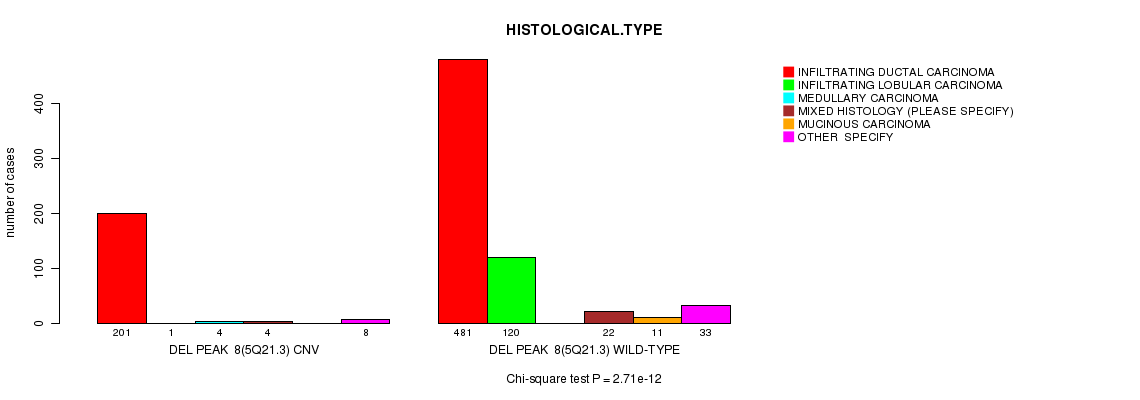
P value = 2.71e-12 (Chi-square test), Q value = 1.6e-09
Table S16. Gene #37: 'Del Peak 8(5q21.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 8(5Q21.3) CNV | 201 | 1 | 4 | 4 | 0 | 8 |
| DEL PEAK 8(5Q21.3) WILD-TYPE | 481 | 120 | 0 | 22 | 11 | 33 |
Figure S16. Get High-res Image Gene #37: 'Del Peak 8(5q21.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
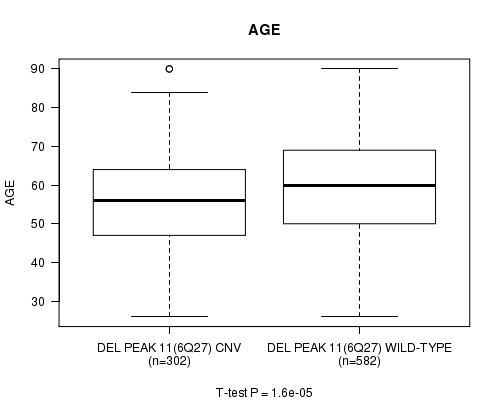
P value = 1.6e-05 (t-test), Q value = 0.0093
Table S17. Gene #40: 'Del Peak 11(6q27)' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 884 | 58.6 (13.2) |
| DEL PEAK 11(6Q27) CNV | 302 | 55.9 (12.7) |
| DEL PEAK 11(6Q27) WILD-TYPE | 582 | 59.9 (13.3) |
Figure S17. Get High-res Image Gene #40: 'Del Peak 11(6q27)' versus Clinical Feature #2: 'AGE'
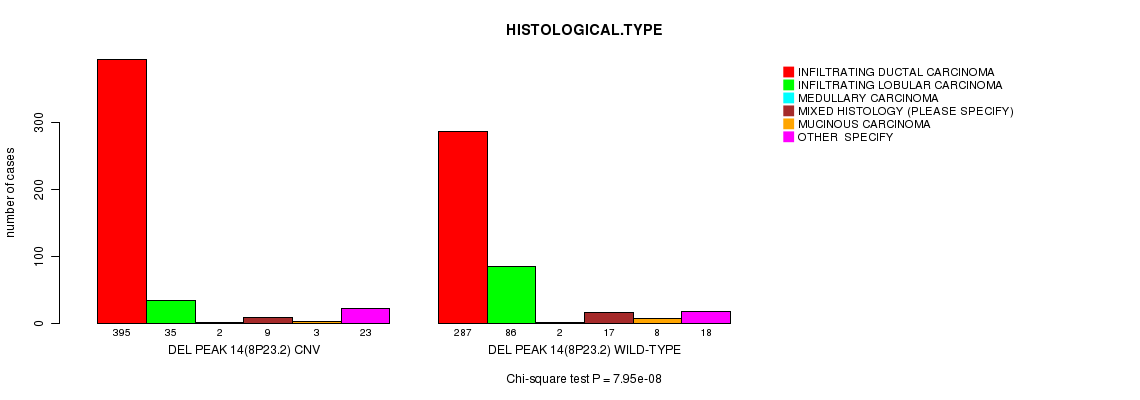
P value = 7.95e-08 (Chi-square test), Q value = 4.7e-05
Table S18. Gene #43: 'Del Peak 14(8p23.2)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 14(8P23.2) CNV | 395 | 35 | 2 | 9 | 3 | 23 |
| DEL PEAK 14(8P23.2) WILD-TYPE | 287 | 86 | 2 | 17 | 8 | 18 |
Figure S18. Get High-res Image Gene #43: 'Del Peak 14(8p23.2)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
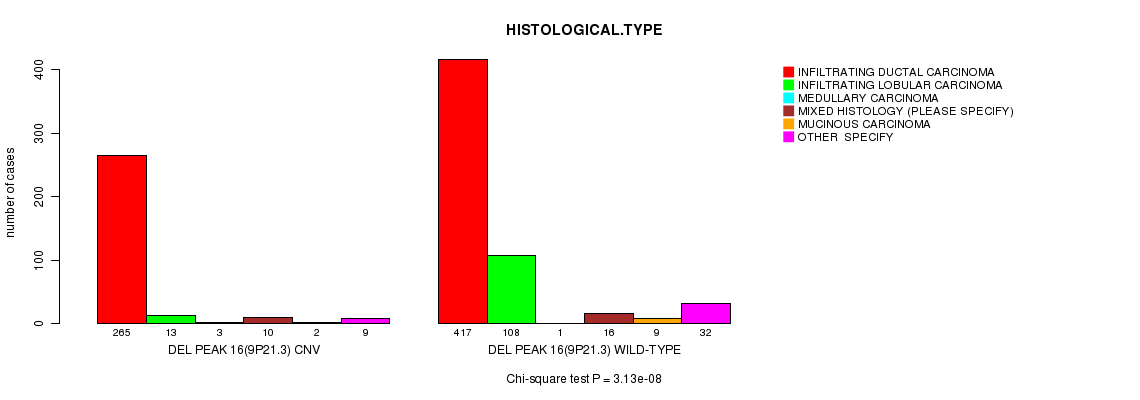
P value = 3.13e-08 (Chi-square test), Q value = 1.8e-05
Table S19. Gene #45: 'Del Peak 16(9p21.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 16(9P21.3) CNV | 265 | 13 | 3 | 10 | 2 | 9 |
| DEL PEAK 16(9P21.3) WILD-TYPE | 417 | 108 | 1 | 16 | 9 | 32 |
Figure S19. Get High-res Image Gene #45: 'Del Peak 16(9p21.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
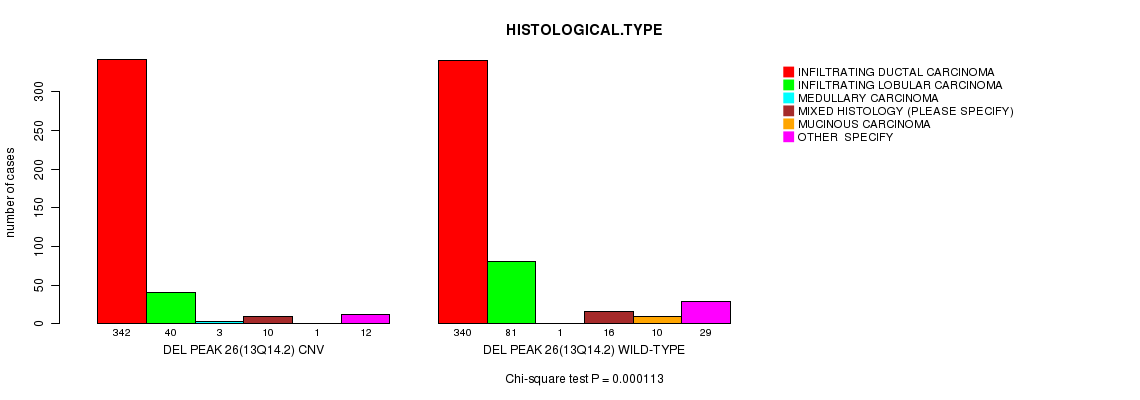
P value = 0.000113 (Chi-square test), Q value = 0.065
Table S20. Gene #55: 'Del Peak 26(13q14.2)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 26(13Q14.2) CNV | 342 | 40 | 3 | 10 | 1 | 12 |
| DEL PEAK 26(13Q14.2) WILD-TYPE | 340 | 81 | 1 | 16 | 10 | 29 |
Figure S20. Get High-res Image Gene #55: 'Del Peak 26(13q14.2)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
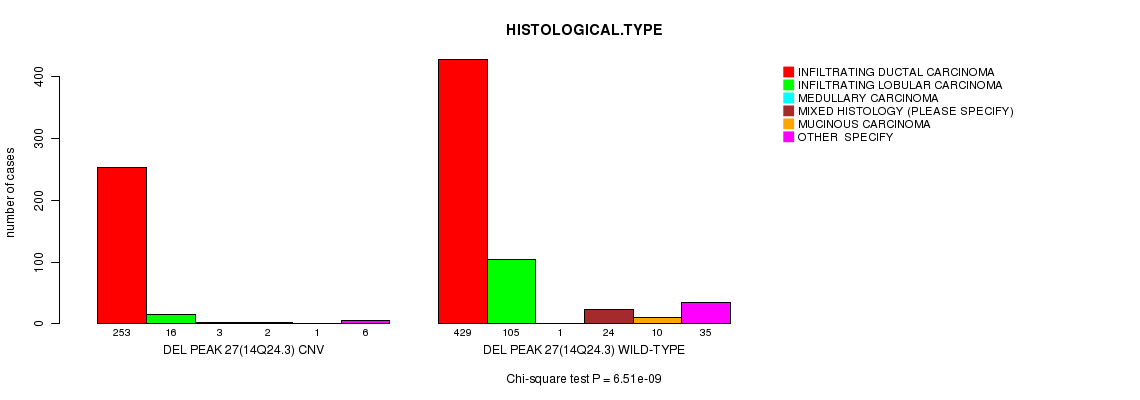
P value = 6.51e-09 (Chi-square test), Q value = 3.8e-06
Table S21. Gene #56: 'Del Peak 27(14q24.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 27(14Q24.3) CNV | 253 | 16 | 3 | 2 | 1 | 6 |
| DEL PEAK 27(14Q24.3) WILD-TYPE | 429 | 105 | 1 | 24 | 10 | 35 |
Figure S21. Get High-res Image Gene #56: 'Del Peak 27(14q24.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
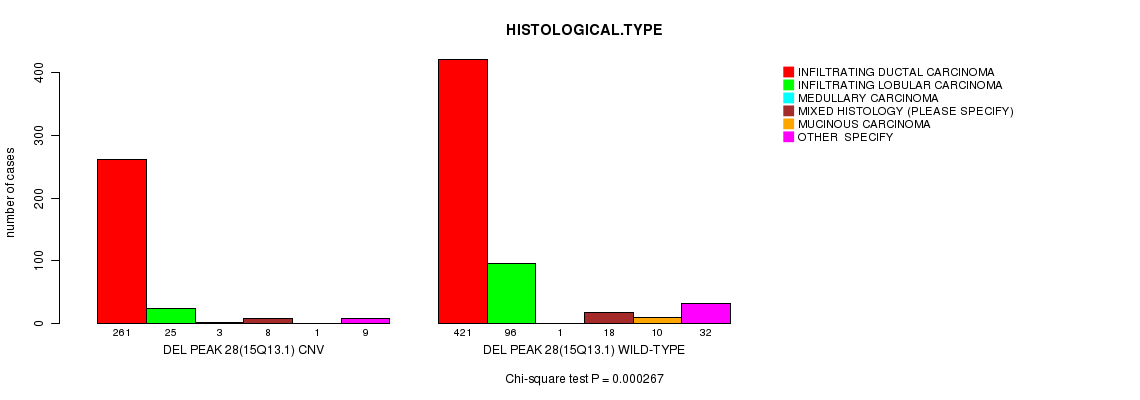
P value = 0.000267 (Chi-square test), Q value = 0.15
Table S22. Gene #57: 'Del Peak 28(15q13.1)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 28(15Q13.1) CNV | 261 | 25 | 3 | 8 | 1 | 9 |
| DEL PEAK 28(15Q13.1) WILD-TYPE | 421 | 96 | 1 | 18 | 10 | 32 |
Figure S22. Get High-res Image Gene #57: 'Del Peak 28(15q13.1)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
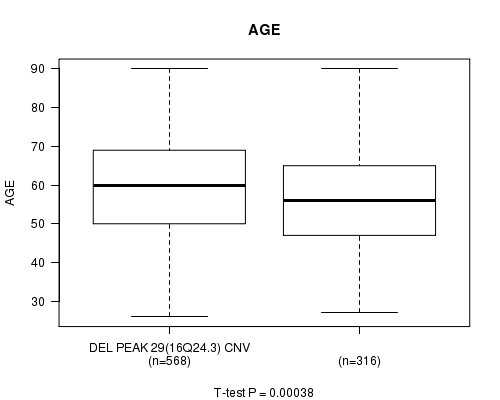
P value = 0.00038 (t-test), Q value = 0.22
Table S23. Gene #58: 'Del Peak 29(16q24.3)' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 884 | 58.6 (13.2) |
| DEL PEAK 29(16Q24.3) CNV | 568 | 59.7 (13.1) |
| DEL PEAK 29(16Q24.3) WILD-TYPE | 316 | 56.4 (13.2) |
Figure S23. Get High-res Image Gene #58: 'Del Peak 29(16q24.3)' versus Clinical Feature #2: 'AGE'
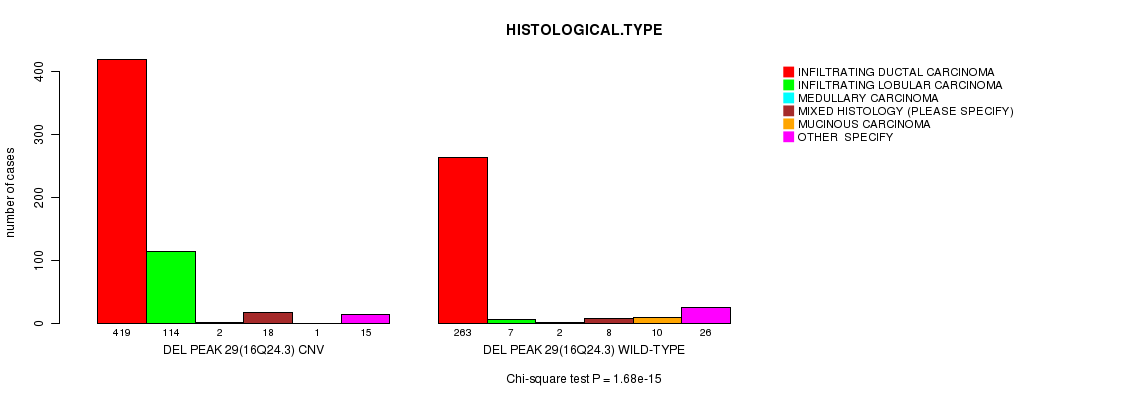
P value = 1.68e-15 (Chi-square test), Q value = 1e-12
Table S24. Gene #58: 'Del Peak 29(16q24.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 29(16Q24.3) CNV | 419 | 114 | 2 | 18 | 1 | 15 |
| DEL PEAK 29(16Q24.3) WILD-TYPE | 263 | 7 | 2 | 8 | 10 | 26 |
Figure S24. Get High-res Image Gene #58: 'Del Peak 29(16q24.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
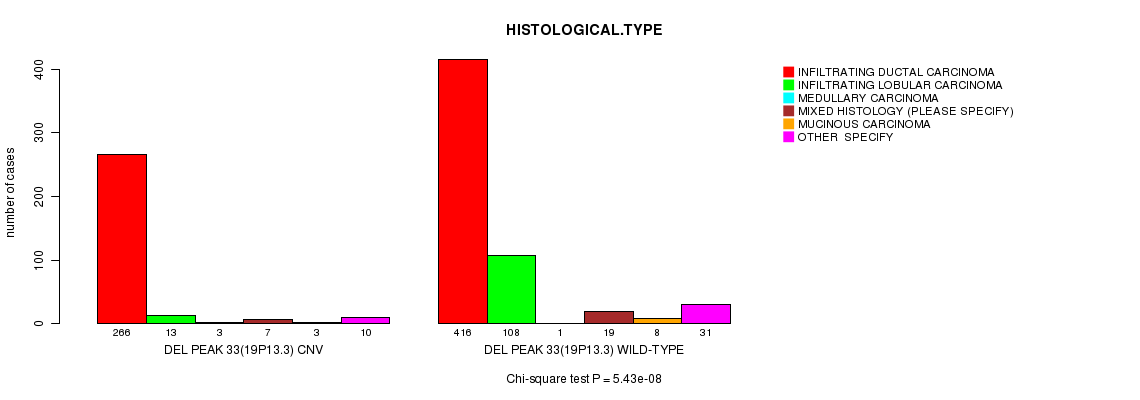
P value = 5.43e-08 (Chi-square test), Q value = 3.2e-05
Table S25. Gene #62: 'Del Peak 33(19p13.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 33(19P13.3) CNV | 266 | 13 | 3 | 7 | 3 | 10 |
| DEL PEAK 33(19P13.3) WILD-TYPE | 416 | 108 | 1 | 19 | 8 | 31 |
Figure S25. Get High-res Image Gene #62: 'Del Peak 33(19p13.3)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
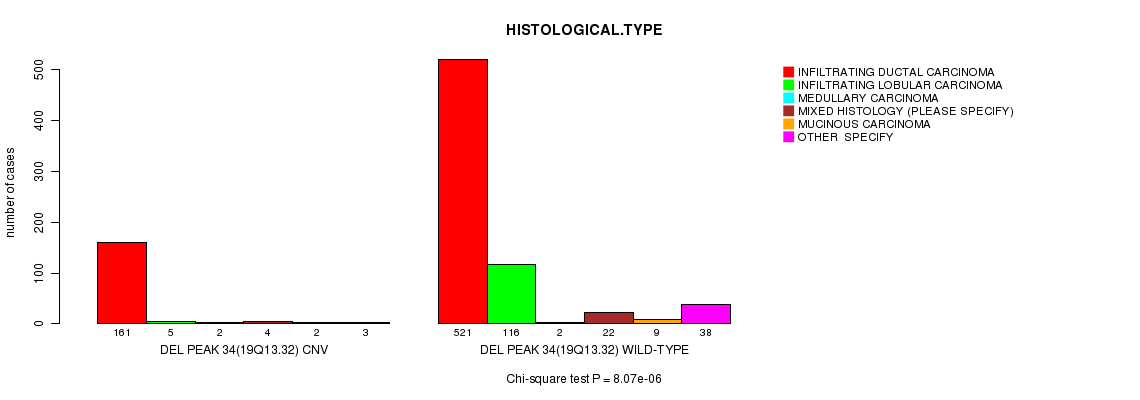
P value = 8.07e-06 (Chi-square test), Q value = 0.0047
Table S26. Gene #63: 'Del Peak 34(19q13.32)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
| nPatients | INFILTRATING DUCTAL CARCINOMA | INFILTRATING LOBULAR CARCINOMA | MEDULLARY CARCINOMA | MIXED HISTOLOGY (PLEASE SPECIFY) | MUCINOUS CARCINOMA | OTHER SPECIFY |
|---|---|---|---|---|---|---|
| ALL | 682 | 121 | 4 | 26 | 11 | 41 |
| DEL PEAK 34(19Q13.32) CNV | 161 | 5 | 2 | 4 | 2 | 3 |
| DEL PEAK 34(19Q13.32) WILD-TYPE | 521 | 116 | 2 | 22 | 9 | 38 |
Figure S26. Get High-res Image Gene #63: 'Del Peak 34(19q13.32)' versus Clinical Feature #4: 'HISTOLOGICAL.TYPE'
-
Mutation data file = all_lesions.conf_99.cnv.cluster.txt
-
Clinical data file = BRCA-TP.clin.merged.picked.txt
-
Number of patients = 885
-
Number of significantly arm-level cnvs = 66
-
Number of selected clinical features = 10
-
Exclude genes that fewer than K tumors have mutations, K = 3
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For continuous numerical clinical features, two-tailed Student's t test with unequal variance (Lehmann and Romano 2005) was applied to compare the clinical values between tumors with and without gene mutations using 't.test' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.