This pipeline computes the correlation between significant arm-level copy number variations (cnvs) and selected clinical features.
Testing the association between copy number variation 57 arm-level events and 12 clinical features across 52 patients, 4 significant findings detected with Q value < 0.25.
-
8P GAIN MUTATION ANALYSIS cnv correlated to 'Time to Death'.
-
22Q GAIN MUTATION ANALYSIS cnv correlated to 'Time to Death'.
-
12P LOSS MUTATION ANALYSIS cnv correlated to 'Time to Death'.
-
20P LOSS MUTATION ANALYSIS cnv correlated to 'Time to Death'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 57 arm-level events and 12 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 4 significant findings detected.
|
Clinical Features |
Time to Death |
AGE |
NEOPLASM DISEASESTAGE |
PATHOLOGY T STAGE |
PATHOLOGY N STAGE |
PATHOLOGY M STAGE |
GENDER |
HISTOLOGICAL TYPE |
NUMBERPACKYEARSSMOKED | YEAROFTOBACCOSMOKINGONSET |
COMPLETENESS OF RESECTION |
NUMBER OF LYMPH NODES |
||
| nCNV (%) | nWild-Type | logrank test | t-test | Chi-square test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | t-test | t-test | Fisher's exact test | t-test | |
| 8P GAIN MUTATION ANALYSIS | 7 (13%) | 45 |
1.15e-05 (0.00684) |
0.869 (1.00) |
0.657 (1.00) |
1 (1.00) |
0.656 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.755 (1.00) |
0.752 (1.00) |
||
| 22Q GAIN MUTATION ANALYSIS | 3 (6%) | 49 |
0.000271 (0.16) |
0.9 (1.00) |
0.874 (1.00) |
1 (1.00) |
0.553 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.703 (1.00) |
|||
| 12P LOSS MUTATION ANALYSIS | 4 (8%) | 48 |
5.31e-05 (0.0316) |
0.674 (1.00) |
0.257 (1.00) |
1 (1.00) |
0.224 (1.00) |
0.0392 (1.00) |
1 (1.00) |
1 (1.00) |
0.404 (1.00) |
0.232 (1.00) |
||
| 20P LOSS MUTATION ANALYSIS | 4 (8%) | 48 |
7.24e-08 (4.31e-05) |
0.809 (1.00) |
0.902 (1.00) |
0.341 (1.00) |
0.562 (1.00) |
0.423 (1.00) |
1 (1.00) |
0.347 (1.00) |
0.0358 (1.00) |
0.837 (1.00) |
||
| 1P GAIN MUTATION ANALYSIS | 3 (6%) | 49 |
0.816 (1.00) |
0.947 (1.00) |
1 (1.00) |
1 (1.00) |
0.635 (1.00) |
0.59 (1.00) |
1 (1.00) |
1 (1.00) |
0.926 (1.00) |
|||
| 1Q GAIN MUTATION ANALYSIS | 11 (21%) | 41 |
0.995 (1.00) |
0.831 (1.00) |
0.678 (1.00) |
0.571 (1.00) |
0.701 (1.00) |
0.198 (1.00) |
0.308 (1.00) |
0.658 (1.00) |
0.413 (1.00) |
0.0889 (1.00) |
0.844 (1.00) |
0.761 (1.00) |
| 2P GAIN MUTATION ANALYSIS | 4 (8%) | 48 |
0.874 (1.00) |
0.65 (1.00) |
0.93 (1.00) |
1 (1.00) |
1 (1.00) |
0.247 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.715 (1.00) |
||
| 2Q GAIN MUTATION ANALYSIS | 5 (10%) | 47 |
0.432 (1.00) |
0.381 (1.00) |
0.942 (1.00) |
1 (1.00) |
1 (1.00) |
0.0719 (1.00) |
0.652 (1.00) |
1 (1.00) |
1 (1.00) |
0.726 (1.00) |
||
| 3Q GAIN MUTATION ANALYSIS | 8 (15%) | 44 |
0.0792 (1.00) |
0.551 (1.00) |
0.472 (1.00) |
1 (1.00) |
0.366 (1.00) |
0.373 (1.00) |
0.447 (1.00) |
0.59 (1.00) |
0.473 (1.00) |
0.425 (1.00) |
||
| 4P GAIN MUTATION ANALYSIS | 3 (6%) | 49 |
0.852 (1.00) |
0.159 (1.00) |
1 (1.00) |
0.553 (1.00) |
0.0252 (1.00) |
0.24 (1.00) |
1 (1.00) |
1 (1.00) |
0.199 (1.00) |
|||
| 4Q GAIN MUTATION ANALYSIS | 3 (6%) | 49 |
0.852 (1.00) |
0.159 (1.00) |
1 (1.00) |
0.553 (1.00) |
0.0252 (1.00) |
0.24 (1.00) |
1 (1.00) |
1 (1.00) |
0.199 (1.00) |
|||
| 5P GAIN MUTATION ANALYSIS | 3 (6%) | 49 |
0.4 (1.00) |
0.947 (1.00) |
1 (1.00) |
1 (1.00) |
0.193 (1.00) |
1 (1.00) |
0.271 (1.00) |
0.0604 (1.00) |
0.19 (1.00) |
|||
| 6Q GAIN MUTATION ANALYSIS | 3 (6%) | 49 |
0.00457 (1.00) |
0.0643 (1.00) |
0.947 (1.00) |
1 (1.00) |
1 (1.00) |
0.635 (1.00) |
0.0916 (1.00) |
0.271 (1.00) |
0.614 (1.00) |
0.236 (1.00) |
||
| 7P GAIN MUTATION ANALYSIS | 11 (21%) | 41 |
0.928 (1.00) |
0.569 (1.00) |
0.727 (1.00) |
0.571 (1.00) |
1 (1.00) |
0.198 (1.00) |
0.735 (1.00) |
1 (1.00) |
0.0177 (1.00) |
0.832 (1.00) |
||
| 7Q GAIN MUTATION ANALYSIS | 11 (21%) | 41 |
0.928 (1.00) |
0.569 (1.00) |
0.727 (1.00) |
0.571 (1.00) |
1 (1.00) |
0.198 (1.00) |
0.735 (1.00) |
1 (1.00) |
0.0177 (1.00) |
0.832 (1.00) |
||
| 8Q GAIN MUTATION ANALYSIS | 14 (27%) | 38 |
0.304 (1.00) |
0.422 (1.00) |
0.748 (1.00) |
1 (1.00) |
0.712 (1.00) |
0.56 (1.00) |
0.366 (1.00) |
1 (1.00) |
0.452 (1.00) |
0.989 (1.00) |
||
| 11P GAIN MUTATION ANALYSIS | 5 (10%) | 47 |
0.865 (1.00) |
0.517 (1.00) |
0.942 (1.00) |
1 (1.00) |
1 (1.00) |
0.476 (1.00) |
0.652 (1.00) |
1 (1.00) |
0.404 (1.00) |
0.374 (1.00) |
||
| 11Q GAIN MUTATION ANALYSIS | 5 (10%) | 47 |
0.192 (1.00) |
0.945 (1.00) |
0.942 (1.00) |
1 (1.00) |
1 (1.00) |
0.476 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.481 (1.00) |
||
| 12P GAIN MUTATION ANALYSIS | 5 (10%) | 47 |
0.153 (1.00) |
0.343 (1.00) |
0.408 (1.00) |
0.41 (1.00) |
1 (1.00) |
0.715 (1.00) |
0.169 (1.00) |
0.416 (1.00) |
0.31 (1.00) |
0.618 (1.00) |
||
| 12Q GAIN MUTATION ANALYSIS | 5 (10%) | 47 |
0.0423 (1.00) |
0.361 (1.00) |
0.844 (1.00) |
1 (1.00) |
0.578 (1.00) |
0.321 (1.00) |
0.169 (1.00) |
0.0691 (1.00) |
0.0695 (1.00) |
0.396 (1.00) |
||
| 13Q GAIN MUTATION ANALYSIS | 3 (6%) | 49 |
0.398 (1.00) |
0.895 (1.00) |
0.947 (1.00) |
1 (1.00) |
1 (1.00) |
0.635 (1.00) |
0.24 (1.00) |
0.271 (1.00) |
0.904 (1.00) |
|||
| 14Q GAIN MUTATION ANALYSIS | 7 (13%) | 45 |
0.333 (1.00) |
0.636 (1.00) |
0.502 (1.00) |
1 (1.00) |
1 (1.00) |
0.138 (1.00) |
1 (1.00) |
0.538 (1.00) |
0.396 (1.00) |
0.52 (1.00) |
||
| 15Q GAIN MUTATION ANALYSIS | 3 (6%) | 49 |
0.096 (1.00) |
0.305 (1.00) |
0.947 (1.00) |
1 (1.00) |
1 (1.00) |
0.635 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.366 (1.00) |
||
| 16P GAIN MUTATION ANALYSIS | 4 (8%) | 48 |
0.515 (1.00) |
0.903 (1.00) |
0.047 (1.00) |
0.341 (1.00) |
0.224 (1.00) |
0.182 (1.00) |
0.324 (1.00) |
0.347 (1.00) |
0.11 (1.00) |
0.0552 (1.00) |
||
| 16Q GAIN MUTATION ANALYSIS | 5 (10%) | 47 |
0.641 (1.00) |
0.851 (1.00) |
0.122 (1.00) |
0.41 (1.00) |
0.325 (1.00) |
0.16 (1.00) |
0.652 (1.00) |
0.416 (1.00) |
0.404 (1.00) |
0.246 (1.00) |
||
| 18P GAIN MUTATION ANALYSIS | 6 (12%) | 46 |
0.71 (1.00) |
0.341 (1.00) |
0.568 (1.00) |
1 (1.00) |
0.612 (1.00) |
0.284 (1.00) |
0.397 (1.00) |
0.48 (1.00) |
0.203 (1.00) |
0.021 (1.00) |
||
| 19Q GAIN MUTATION ANALYSIS | 6 (12%) | 46 |
0.654 (1.00) |
0.795 (1.00) |
0.472 (1.00) |
1 (1.00) |
1 (1.00) |
0.746 (1.00) |
0.397 (1.00) |
1 (1.00) |
0.0574 (1.00) |
0.227 (1.00) |
1 (1.00) |
0.491 (1.00) |
| 20P GAIN MUTATION ANALYSIS | 7 (13%) | 45 |
0.469 (1.00) |
0.348 (1.00) |
0.703 (1.00) |
1 (1.00) |
0.181 (1.00) |
0.768 (1.00) |
0.69 (1.00) |
0.538 (1.00) |
0.203 (1.00) |
0.371 (1.00) |
||
| 20Q GAIN MUTATION ANALYSIS | 10 (19%) | 42 |
0.191 (1.00) |
0.0636 (1.00) |
0.472 (1.00) |
1 (1.00) |
0.42 (1.00) |
0.828 (1.00) |
0.157 (1.00) |
1 (1.00) |
0.333 (1.00) |
0.205 (1.00) |
||
| 1P LOSS MUTATION ANALYSIS | 6 (12%) | 46 |
0.313 (1.00) |
0.103 (1.00) |
0.472 (1.00) |
0.473 (1.00) |
1 (1.00) |
0.375 (1.00) |
0.397 (1.00) |
1 (1.00) |
0.282 (1.00) |
0.155 (1.00) |
0.755 (1.00) |
0.539 (1.00) |
| 2P LOSS MUTATION ANALYSIS | 7 (13%) | 45 |
0.718 (1.00) |
0.858 (1.00) |
0.916 (1.00) |
1 (1.00) |
1 (1.00) |
0.574 (1.00) |
1 (1.00) |
0.538 (1.00) |
0.0943 (1.00) |
0.746 (1.00) |
||
| 3P LOSS MUTATION ANALYSIS | 11 (21%) | 41 |
0.427 (1.00) |
0.892 (1.00) |
0.727 (1.00) |
1 (1.00) |
1 (1.00) |
0.7 (1.00) |
0.0862 (1.00) |
1 (1.00) |
0.282 (1.00) |
0.69 (1.00) |
0.153 (1.00) |
|
| 3Q LOSS MUTATION ANALYSIS | 4 (8%) | 48 |
0.722 (1.00) |
0.77 (1.00) |
0.93 (1.00) |
1 (1.00) |
1 (1.00) |
0.672 (1.00) |
0.0393 (1.00) |
1 (1.00) |
1 (1.00) |
0.531 (1.00) |
||
| 4P LOSS MUTATION ANALYSIS | 5 (10%) | 47 |
0.0477 (1.00) |
0.166 (1.00) |
0.844 (1.00) |
1 (1.00) |
0.578 (1.00) |
1 (1.00) |
0.169 (1.00) |
1 (1.00) |
0.706 (1.00) |
0.306 (1.00) |
||
| 4Q LOSS MUTATION ANALYSIS | 4 (8%) | 48 |
0.718 (1.00) |
0.0463 (1.00) |
0.902 (1.00) |
1 (1.00) |
0.562 (1.00) |
0.423 (1.00) |
0.0393 (1.00) |
1 (1.00) |
0.404 (1.00) |
0.515 (1.00) |
||
| 5Q LOSS MUTATION ANALYSIS | 5 (10%) | 47 |
0.951 (1.00) |
0.395 (1.00) |
0.844 (1.00) |
1 (1.00) |
0.578 (1.00) |
0.715 (1.00) |
0.652 (1.00) |
0.416 (1.00) |
0.11 (1.00) |
0.431 (1.00) |
||
| 6P LOSS MUTATION ANALYSIS | 18 (35%) | 34 |
0.157 (1.00) |
0.48 (1.00) |
0.674 (1.00) |
0.648 (1.00) |
0.731 (1.00) |
0.0751 (1.00) |
0.149 (1.00) |
0.732 (1.00) |
0.953 (1.00) |
0.704 (1.00) |
0.512 (1.00) |
0.764 (1.00) |
| 6Q LOSS MUTATION ANALYSIS | 21 (40%) | 31 |
0.611 (1.00) |
0.358 (1.00) |
0.43 (1.00) |
0.637 (1.00) |
0.512 (1.00) |
0.0145 (1.00) |
1 (1.00) |
0.782 (1.00) |
0.913 (1.00) |
0.789 (1.00) |
0.785 (1.00) |
0.654 (1.00) |
| 8P LOSS MUTATION ANALYSIS | 8 (15%) | 44 |
0.635 (1.00) |
0.5 (1.00) |
0.357 (1.00) |
1 (1.00) |
1 (1.00) |
0.373 (1.00) |
0.447 (1.00) |
0.538 (1.00) |
0.222 (1.00) |
0.618 (1.00) |
||
| 9P LOSS MUTATION ANALYSIS | 21 (40%) | 31 |
0.822 (1.00) |
0.6 (1.00) |
0.102 (1.00) |
0.637 (1.00) |
0.0166 (1.00) |
0.423 (1.00) |
1 (1.00) |
1 (1.00) |
0.341 (1.00) |
0.206 (1.00) |
0.101 (1.00) |
0.0816 (1.00) |
| 9Q LOSS MUTATION ANALYSIS | 12 (23%) | 40 |
0.24 (1.00) |
0.0643 (1.00) |
0.657 (1.00) |
1 (1.00) |
0.253 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.592 (1.00) |
0.442 (1.00) |
0.349 (1.00) |
0.21 (1.00) |
| 10P LOSS MUTATION ANALYSIS | 9 (17%) | 43 |
0.288 (1.00) |
0.0978 (1.00) |
0.733 (1.00) |
1 (1.00) |
1 (1.00) |
0.26 (1.00) |
0.272 (1.00) |
0.638 (1.00) |
0.599 (1.00) |
0.67 (1.00) |
0.379 (1.00) |
0.757 (1.00) |
| 10Q LOSS MUTATION ANALYSIS | 10 (19%) | 42 |
0.253 (1.00) |
0.206 (1.00) |
0.738 (1.00) |
1 (1.00) |
1 (1.00) |
0.137 (1.00) |
0.483 (1.00) |
1 (1.00) |
0.152 (1.00) |
0.422 (1.00) |
0.348 (1.00) |
|
| 11P LOSS MUTATION ANALYSIS | 3 (6%) | 49 |
0.403 (1.00) |
0.588 (1.00) |
0.874 (1.00) |
1 (1.00) |
0.553 (1.00) |
0.326 (1.00) |
0.59 (1.00) |
1 (1.00) |
1 (1.00) |
0.761 (1.00) |
||
| 11Q LOSS MUTATION ANALYSIS | 3 (6%) | 49 |
0.487 (1.00) |
0.247 (1.00) |
0.874 (1.00) |
1 (1.00) |
0.553 (1.00) |
1 (1.00) |
0.0916 (1.00) |
0.271 (1.00) |
0.0604 (1.00) |
0.761 (1.00) |
||
| 12Q LOSS MUTATION ANALYSIS | 6 (12%) | 46 |
0.0374 (1.00) |
0.682 (1.00) |
0.223 (1.00) |
0.473 (1.00) |
0.127 (1.00) |
0.0275 (1.00) |
1 (1.00) |
1 (1.00) |
0.396 (1.00) |
0.0709 (1.00) |
||
| 13Q LOSS MUTATION ANALYSIS | 8 (15%) | 44 |
0.691 (1.00) |
0.296 (1.00) |
0.508 (1.00) |
1 (1.00) |
0.663 (1.00) |
0.222 (1.00) |
0.123 (1.00) |
0.254 (1.00) |
0.282 (1.00) |
0.773 (1.00) |
0.0635 (1.00) |
0.43 (1.00) |
| 15Q LOSS MUTATION ANALYSIS | 7 (13%) | 45 |
0.387 (1.00) |
0.998 (1.00) |
0.354 (1.00) |
0.53 (1.00) |
0.656 (1.00) |
0.0134 (1.00) |
0.69 (1.00) |
1 (1.00) |
0.0715 (1.00) |
0.746 (1.00) |
0.755 (1.00) |
0.581 (1.00) |
| 17P LOSS MUTATION ANALYSIS | 18 (35%) | 34 |
0.274 (1.00) |
0.484 (1.00) |
0.218 (1.00) |
0.15 (1.00) |
0.507 (1.00) |
0.183 (1.00) |
0.774 (1.00) |
0.732 (1.00) |
0.0459 (1.00) |
0.208 (1.00) |
0.00145 (0.861) |
0.544 (1.00) |
| 17Q LOSS MUTATION ANALYSIS | 9 (17%) | 43 |
0.164 (1.00) |
0.94 (1.00) |
0.733 (1.00) |
0.573 (1.00) |
1 (1.00) |
0.503 (1.00) |
0.272 (1.00) |
0.254 (1.00) |
0.146 (1.00) |
0.109 (1.00) |
||
| 18P LOSS MUTATION ANALYSIS | 17 (33%) | 35 |
0.868 (1.00) |
0.082 (1.00) |
0.318 (1.00) |
1 (1.00) |
1 (1.00) |
0.791 (1.00) |
0.769 (1.00) |
0.732 (1.00) |
0.295 (1.00) |
0.704 (1.00) |
0.178 (1.00) |
0.971 (1.00) |
| 18Q LOSS MUTATION ANALYSIS | 26 (50%) | 26 |
0.322 (1.00) |
0.728 (1.00) |
0.247 (1.00) |
1 (1.00) |
0.743 (1.00) |
0.417 (1.00) |
0.404 (1.00) |
1 (1.00) |
0.645 (1.00) |
0.071 (1.00) |
0.0286 (1.00) |
0.572 (1.00) |
| 19P LOSS MUTATION ANALYSIS | 5 (10%) | 47 |
0.492 (1.00) |
0.397 (1.00) |
0.408 (1.00) |
1 (1.00) |
1 (1.00) |
0.16 (1.00) |
0.358 (1.00) |
1 (1.00) |
1 (1.00) |
0.963 (1.00) |
||
| 19Q LOSS MUTATION ANALYSIS | 4 (8%) | 48 |
0.492 (1.00) |
0.56 (1.00) |
0.302 (1.00) |
1 (1.00) |
1 (1.00) |
0.0392 (1.00) |
0.615 (1.00) |
1 (1.00) |
1 (1.00) |
0.941 (1.00) |
||
| 21Q LOSS MUTATION ANALYSIS | 16 (31%) | 36 |
0.291 (1.00) |
0.418 (1.00) |
0.0625 (1.00) |
0.308 (1.00) |
0.0776 (1.00) |
0.0377 (1.00) |
0.229 (1.00) |
0.514 (1.00) |
0.609 (1.00) |
0.81 (1.00) |
0.189 (1.00) |
0.0824 (1.00) |
| 22Q LOSS MUTATION ANALYSIS | 8 (15%) | 44 |
0.163 (1.00) |
0.933 (1.00) |
0.709 (1.00) |
1 (1.00) |
1 (1.00) |
0.0619 (1.00) |
0.123 (1.00) |
1 (1.00) |
0.0344 (1.00) |
0.51 (1.00) |
0.0908 (1.00) |
0.844 (1.00) |
| XQ LOSS MUTATION ANALYSIS | 4 (8%) | 48 |
0.996 (1.00) |
0.796 (1.00) |
0.902 (1.00) |
1 (1.00) |
0.562 (1.00) |
0.247 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.488 (1.00) |
P value = 1.15e-05 (logrank test), Q value = 0.0068
Table S1. Gene #12: '8P GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 49 | 17 | 0.0 - 49.4 (5.0) |
| 8P GAIN MUTATED | 6 | 3 | 0.0 - 4.8 (2.4) |
| 8P GAIN WILD-TYPE | 43 | 14 | 0.1 - 49.4 (7.1) |
Figure S1. Get High-res Image Gene #12: '8P GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'

P value = 0.000271 (logrank test), Q value = 0.16
Table S2. Gene #27: '22Q GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 49 | 17 | 0.0 - 49.4 (5.0) |
| 22Q GAIN MUTATED | 3 | 1 | 0.0 - 3.9 (0.1) |
| 22Q GAIN WILD-TYPE | 46 | 16 | 0.1 - 49.4 (5.6) |
Figure S2. Get High-res Image Gene #27: '22Q GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
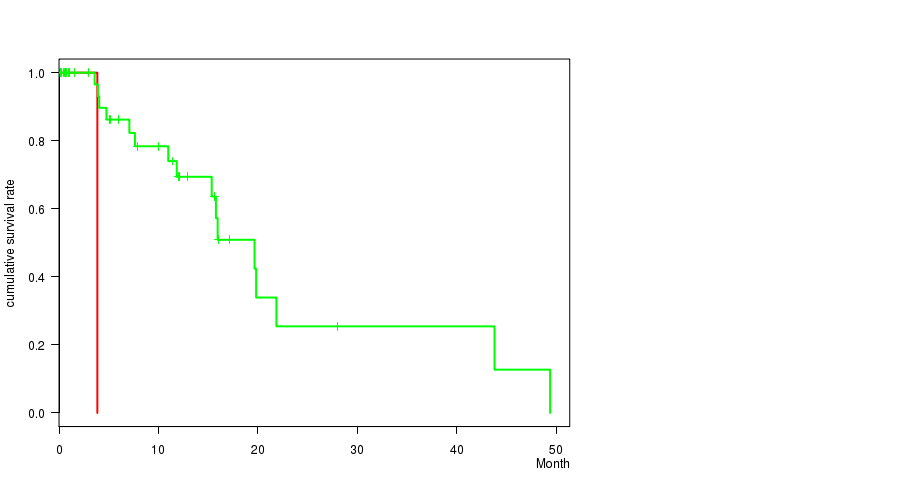
P value = 5.31e-05 (logrank test), Q value = 0.032
Table S3. Gene #44: '12P LOSS MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 49 | 17 | 0.0 - 49.4 (5.0) |
| 12P LOSS MUTATED | 4 | 2 | 0.2 - 4.0 (2.3) |
| 12P LOSS WILD-TYPE | 45 | 15 | 0.0 - 49.4 (6.0) |
Figure S3. Get High-res Image Gene #44: '12P LOSS MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'

P value = 7.24e-08 (logrank test), Q value = 4.3e-05
Table S4. Gene #54: '20P LOSS MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 49 | 17 | 0.0 - 49.4 (5.0) |
| 20P LOSS MUTATED | 3 | 1 | 0.8 - 3.6 (1.5) |
| 20P LOSS WILD-TYPE | 46 | 16 | 0.0 - 49.4 (5.6) |
Figure S4. Get High-res Image Gene #54: '20P LOSS MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
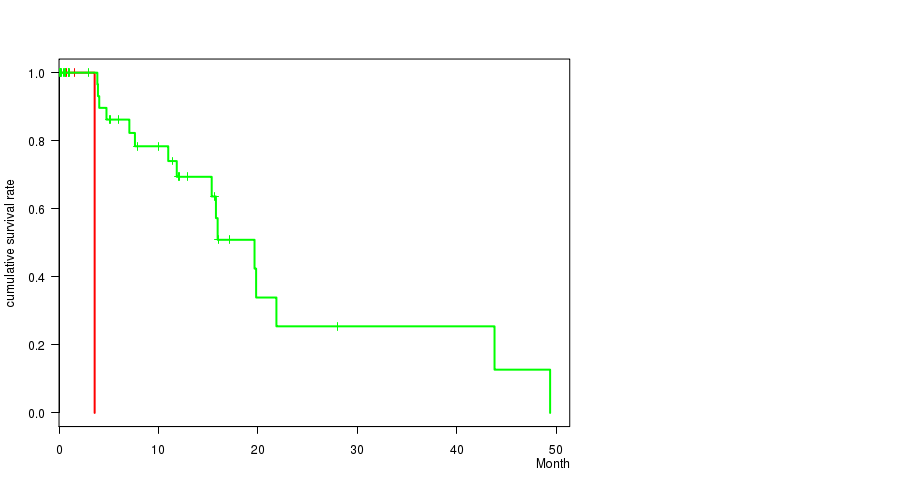
-
Copy number data file = transformed.cor.cli.txt
-
Clinical data file = PAAD-TP.clin.merged.picked.txt
-
Number of patients = 52
-
Number of significantly arm-level cnvs = 57
-
Number of selected clinical features = 12
-
Exclude regions that fewer than K tumors have mutations, K = 3
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For continuous numerical clinical features, two-tailed Student's t test with unequal variance (Lehmann and Romano 2005) was applied to compare the clinical values between tumors with and without gene mutations using 't.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.