This pipeline computes the correlation between significant copy number variation (cnv focal) genes and selected clinical features.
Testing the association between copy number variation 46 focal events and 6 clinical features across 160 patients, 4 significant findings detected with Q value < 0.25.
-
AMP PEAK 11(20Q13.33) MUTATION ANALYSIS cnv correlated to 'PATHOLOGY.T.STAGE'.
-
AMP PEAK 19(XQ25) MUTATION ANALYSIS cnv correlated to 'AGE'.
-
DEL PEAK 6(4Q28.2) MUTATION ANALYSIS cnv correlated to 'AGE'.
-
DEL PEAK 11(8P21.3) MUTATION ANALYSIS cnv correlated to 'PATHOLOGY.T.STAGE'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 46 focal events and 6 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 4 significant findings detected.
|
Clinical Features |
Time to Death |
AGE |
PATHOLOGY T STAGE |
PATHOLOGY N STAGE |
COMPLETENESS OF RESECTION |
NUMBER OF LYMPH NODES |
||
| nCNV (%) | nWild-Type | logrank test | t-test | Fisher's exact test | Fisher's exact test | Fisher's exact test | t-test | |
| AMP PEAK 11(20Q13 33) MUTATION ANALYSIS | 12 (8%) | 148 |
100 (1.00) |
0.00419 (1.00) |
0.000273 (0.0749) |
0.363 (1.00) |
0.0405 (1.00) |
0.0015 (0.404) |
| AMP PEAK 19(XQ25) MUTATION ANALYSIS | 3 (2%) | 157 |
100 (1.00) |
6.59e-09 (1.82e-06) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.00152 (0.405) |
| DEL PEAK 6(4Q28 2) MUTATION ANALYSIS | 14 (9%) | 146 |
100 (1.00) |
4.49e-05 (0.0123) |
0.0745 (1.00) |
1 (1.00) |
1 (1.00) |
0.159 (1.00) |
| DEL PEAK 11(8P21 3) MUTATION ANALYSIS | 97 (61%) | 63 |
100 (1.00) |
0.432 (1.00) |
0.000841 (0.23) |
0.0491 (1.00) |
1 (1.00) |
0.011 (1.00) |
| AMP PEAK 1(1Q21 3) MUTATION ANALYSIS | 13 (8%) | 147 |
100 (1.00) |
0.347 (1.00) |
0.311 (1.00) |
0.631 (1.00) |
0.795 (1.00) |
0.632 (1.00) |
| AMP PEAK 2(3Q26 2) MUTATION ANALYSIS | 25 (16%) | 135 |
100 (1.00) |
0.13 (1.00) |
0.0325 (1.00) |
0.287 (1.00) |
0.767 (1.00) |
0.274 (1.00) |
| AMP PEAK 3(7P15 3) MUTATION ANALYSIS | 32 (20%) | 128 |
100 (1.00) |
0.432 (1.00) |
0.0166 (1.00) |
0.511 (1.00) |
0.0239 (1.00) |
0.34 (1.00) |
| AMP PEAK 4(7Q34) MUTATION ANALYSIS | 29 (18%) | 131 |
100 (1.00) |
0.144 (1.00) |
0.252 (1.00) |
1 (1.00) |
0.193 (1.00) |
0.606 (1.00) |
| AMP PEAK 5(8P11 22) MUTATION ANALYSIS | 16 (10%) | 144 |
100 (1.00) |
0.449 (1.00) |
0.518 (1.00) |
0.0442 (1.00) |
0.66 (1.00) |
0.271 (1.00) |
| AMP PEAK 6(8Q24 21) MUTATION ANALYSIS | 42 (26%) | 118 |
100 (1.00) |
0.295 (1.00) |
0.109 (1.00) |
0.129 (1.00) |
1 (1.00) |
0.314 (1.00) |
| AMP PEAK 7(9Q34 3) MUTATION ANALYSIS | 15 (9%) | 145 |
100 (1.00) |
0.175 (1.00) |
0.0218 (1.00) |
0.000949 (0.258) |
0.377 (1.00) |
0.0733 (1.00) |
| AMP PEAK 8(11Q13 2) MUTATION ANALYSIS | 9 (6%) | 151 |
100 (1.00) |
0.524 (1.00) |
0.0517 (1.00) |
0.0557 (1.00) |
0.241 (1.00) |
0.294 (1.00) |
| AMP PEAK 9(12Q24 32) MUTATION ANALYSIS | 7 (4%) | 153 |
100 (1.00) |
0.959 (1.00) |
0.182 (1.00) |
0.162 (1.00) |
0.266 (1.00) |
0.398 (1.00) |
| AMP PEAK 10(14Q21 1) MUTATION ANALYSIS | 8 (5%) | 152 |
100 (1.00) |
0.00135 (0.366) |
0.459 (1.00) |
0.553 (1.00) |
0.458 (1.00) |
0.717 (1.00) |
| AMP PEAK 12(XP22 11) MUTATION ANALYSIS | 6 (4%) | 154 |
100 (1.00) |
0.529 (1.00) |
0.746 (1.00) |
0.497 (1.00) |
1 (1.00) |
0.694 (1.00) |
| AMP PEAK 13(XP22 11) MUTATION ANALYSIS | 6 (4%) | 154 |
100 (1.00) |
0.0772 (1.00) |
0.746 (1.00) |
0.497 (1.00) |
1 (1.00) |
0.694 (1.00) |
| AMP PEAK 14(XP21 1) MUTATION ANALYSIS | 5 (3%) | 155 |
100 (1.00) |
0.37 (1.00) |
0.473 (1.00) |
0.435 (1.00) |
0.344 (1.00) |
0.995 (1.00) |
| AMP PEAK 15(XQ12) MUTATION ANALYSIS | 3 (2%) | 157 |
100 (1.00) |
0.63 (1.00) |
0.322 (1.00) |
1 (1.00) |
0.532 (1.00) |
0.00152 (0.405) |
| AMP PEAK 16(XQ21 1) MUTATION ANALYSIS | 7 (4%) | 153 |
100 (1.00) |
0.691 (1.00) |
0.571 (1.00) |
0.553 (1.00) |
0.432 (1.00) |
0.764 (1.00) |
| AMP PEAK 17(XQ21 1) MUTATION ANALYSIS | 5 (3%) | 155 |
100 (1.00) |
0.316 (1.00) |
0.473 (1.00) |
0.0878 (1.00) |
0.624 (1.00) |
0.346 (1.00) |
| AMP PEAK 18(XQ21 31) MUTATION ANALYSIS | 5 (3%) | 155 |
100 (1.00) |
0.224 (1.00) |
1 (1.00) |
1 (1.00) |
0.624 (1.00) |
0.00152 (0.405) |
| AMP PEAK 20(XQ25) MUTATION ANALYSIS | 6 (4%) | 154 |
100 (1.00) |
0.753 (1.00) |
0.746 (1.00) |
0.497 (1.00) |
0.42 (1.00) |
0.857 (1.00) |
| AMP PEAK 21(XQ27 1) MUTATION ANALYSIS | 9 (6%) | 151 |
100 (1.00) |
0.282 (1.00) |
1 (1.00) |
0.6 (1.00) |
1 (1.00) |
0.00151 (0.405) |
| DEL PEAK 1(1P31 3) MUTATION ANALYSIS | 21 (13%) | 139 |
100 (1.00) |
0.784 (1.00) |
0.0816 (1.00) |
1 (1.00) |
0.606 (1.00) |
0.885 (1.00) |
| DEL PEAK 2(1Q42 13) MUTATION ANALYSIS | 15 (9%) | 145 |
100 (1.00) |
0.177 (1.00) |
0.262 (1.00) |
0.662 (1.00) |
1 (1.00) |
0.729 (1.00) |
| DEL PEAK 3(2Q22 1) MUTATION ANALYSIS | 20 (12%) | 140 |
100 (1.00) |
0.0556 (1.00) |
0.661 (1.00) |
1 (1.00) |
1 (1.00) |
0.549 (1.00) |
| DEL PEAK 4(3P13) MUTATION ANALYSIS | 27 (17%) | 133 |
100 (1.00) |
0.207 (1.00) |
0.206 (1.00) |
1 (1.00) |
0.519 (1.00) |
0.745 (1.00) |
| DEL PEAK 5(3Q29) MUTATION ANALYSIS | 10 (6%) | 150 |
100 (1.00) |
0.255 (1.00) |
0.65 (1.00) |
1 (1.00) |
1 (1.00) |
0.994 (1.00) |
| DEL PEAK 7(5Q11 2) MUTATION ANALYSIS | 24 (15%) | 136 |
100 (1.00) |
0.177 (1.00) |
0.0018 (0.477) |
0.00251 (0.66) |
1 (1.00) |
0.0767 (1.00) |
| DEL PEAK 8(5Q21 1) MUTATION ANALYSIS | 28 (18%) | 132 |
100 (1.00) |
0.0163 (1.00) |
0.866 (1.00) |
0.178 (1.00) |
0.398 (1.00) |
0.305 (1.00) |
| DEL PEAK 9(6Q15) MUTATION ANALYSIS | 51 (32%) | 109 |
100 (1.00) |
0.0544 (1.00) |
0.862 (1.00) |
0.0466 (1.00) |
0.444 (1.00) |
0.0268 (1.00) |
| DEL PEAK 10(7Q36 1) MUTATION ANALYSIS | 7 (4%) | 153 |
100 (1.00) |
0.31 (1.00) |
1 (1.00) |
0.124 (1.00) |
1 (1.00) |
0.404 (1.00) |
| DEL PEAK 12(8P11 21) MUTATION ANALYSIS | 55 (34%) | 105 |
100 (1.00) |
0.289 (1.00) |
0.333 (1.00) |
0.78 (1.00) |
0.0219 (1.00) |
0.65 (1.00) |
| DEL PEAK 13(9P22 3) MUTATION ANALYSIS | 14 (9%) | 146 |
100 (1.00) |
0.261 (1.00) |
0.404 (1.00) |
0.00779 (1.00) |
0.795 (1.00) |
0.0952 (1.00) |
| DEL PEAK 14(10Q23 31) MUTATION ANALYSIS | 59 (37%) | 101 |
100 (1.00) |
0.824 (1.00) |
0.0255 (1.00) |
0.0261 (1.00) |
0.691 (1.00) |
0.324 (1.00) |
| DEL PEAK 15(11Q23 2) MUTATION ANALYSIS | 17 (11%) | 143 |
100 (1.00) |
0.722 (1.00) |
0.0157 (1.00) |
1 (1.00) |
0.644 (1.00) |
0.103 (1.00) |
| DEL PEAK 16(12P13 2) MUTATION ANALYSIS | 34 (21%) | 126 |
100 (1.00) |
0.702 (1.00) |
0.185 (1.00) |
0.197 (1.00) |
0.737 (1.00) |
0.522 (1.00) |
| DEL PEAK 17(13Q14 13) MUTATION ANALYSIS | 74 (46%) | 86 |
100 (1.00) |
0.0276 (1.00) |
0.606 (1.00) |
0.412 (1.00) |
0.76 (1.00) |
0.275 (1.00) |
| DEL PEAK 18(16Q22 3) MUTATION ANALYSIS | 53 (33%) | 107 |
100 (1.00) |
0.15 (1.00) |
0.0736 (1.00) |
0.391 (1.00) |
0.924 (1.00) |
0.16 (1.00) |
| DEL PEAK 19(16Q24 1) MUTATION ANALYSIS | 68 (42%) | 92 |
100 (1.00) |
0.354 (1.00) |
0.0136 (1.00) |
0.278 (1.00) |
0.931 (1.00) |
0.17 (1.00) |
| DEL PEAK 20(17P13 1) MUTATION ANALYSIS | 42 (26%) | 118 |
100 (1.00) |
0.581 (1.00) |
0.0627 (1.00) |
0.123 (1.00) |
0.708 (1.00) |
0.291 (1.00) |
| DEL PEAK 21(17Q21 31) MUTATION ANALYSIS | 30 (19%) | 130 |
100 (1.00) |
0.755 (1.00) |
0.439 (1.00) |
0.738 (1.00) |
0.273 (1.00) |
0.364 (1.00) |
| DEL PEAK 22(18Q23) MUTATION ANALYSIS | 49 (31%) | 111 |
100 (1.00) |
0.151 (1.00) |
0.267 (1.00) |
0.0709 (1.00) |
0.435 (1.00) |
0.191 (1.00) |
| DEL PEAK 23(19Q13 2) MUTATION ANALYSIS | 13 (8%) | 147 |
100 (1.00) |
0.0597 (1.00) |
0.311 (1.00) |
0.145 (1.00) |
0.595 (1.00) |
0.425 (1.00) |
| DEL PEAK 24(21Q22 2) MUTATION ANALYSIS | 17 (11%) | 143 |
100 (1.00) |
0.504 (1.00) |
0.0393 (1.00) |
0.000949 (0.258) |
0.66 (1.00) |
0.0509 (1.00) |
| DEL PEAK 25(21Q22 3) MUTATION ANALYSIS | 52 (32%) | 108 |
100 (1.00) |
0.818 (1.00) |
0.0202 (1.00) |
0.0375 (1.00) |
0.388 (1.00) |
0.117 (1.00) |
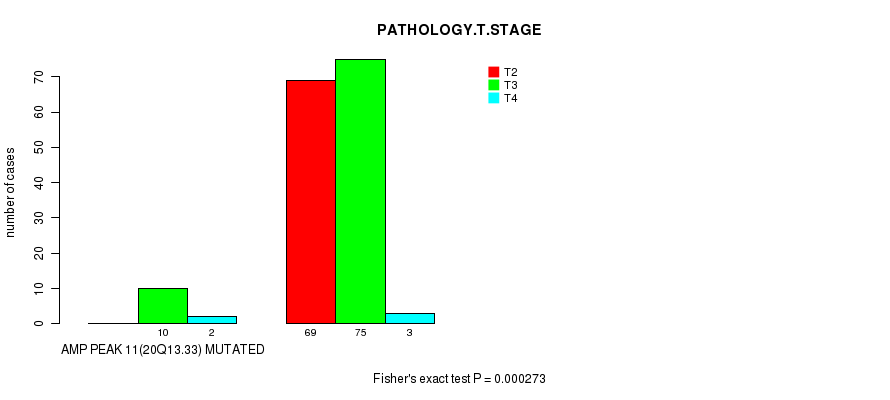
P value = 0.000273 (Fisher's exact test), Q value = 0.075
Table S1. Gene #11: 'AMP PEAK 11(20Q13.33) MUTATION STATUS' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
| nPatients | T2 | T3 | T4 |
|---|---|---|---|
| ALL | 69 | 85 | 5 |
| AMP PEAK 11(20Q13.33) MUTATED | 0 | 10 | 2 |
| AMP PEAK 11(20Q13.33) WILD-TYPE | 69 | 75 | 3 |
Figure S1. Get High-res Image Gene #11: 'AMP PEAK 11(20Q13.33) MUTATION STATUS' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
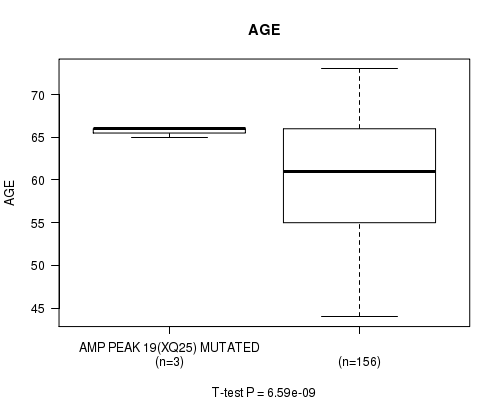
P value = 6.59e-09 (t-test), Q value = 1.8e-06
Table S2. Gene #19: 'AMP PEAK 19(XQ25) MUTATION STATUS' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 159 | 60.3 (6.9) |
| AMP PEAK 19(XQ25) MUTATED | 3 | 65.7 (0.6) |
| AMP PEAK 19(XQ25) WILD-TYPE | 156 | 60.2 (6.9) |
Figure S2. Get High-res Image Gene #19: 'AMP PEAK 19(XQ25) MUTATION STATUS' versus Clinical Feature #2: 'AGE'
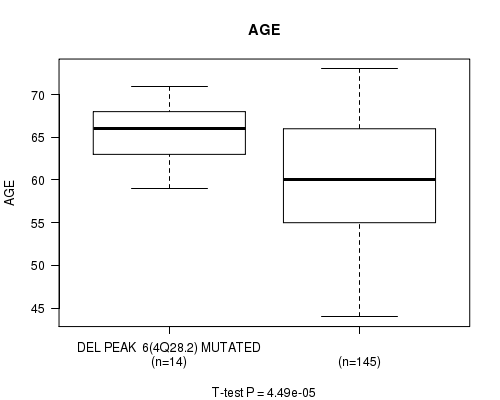
P value = 4.49e-05 (t-test), Q value = 0.012
Table S3. Gene #27: 'DEL PEAK 6(4Q28.2) MUTATION STATUS' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 159 | 60.3 (6.9) |
| DEL PEAK 6(4Q28.2) MUTATED | 14 | 65.2 (3.5) |
| DEL PEAK 6(4Q28.2) WILD-TYPE | 145 | 59.8 (6.9) |
Figure S3. Get High-res Image Gene #27: 'DEL PEAK 6(4Q28.2) MUTATION STATUS' versus Clinical Feature #2: 'AGE'
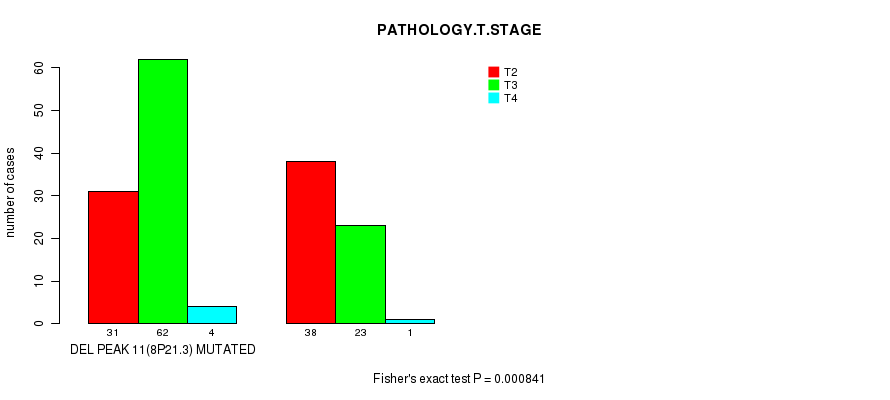
P value = 0.000841 (Fisher's exact test), Q value = 0.23
Table S4. Gene #32: 'DEL PEAK 11(8P21.3) MUTATION STATUS' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
| nPatients | T2 | T3 | T4 |
|---|---|---|---|
| ALL | 69 | 85 | 5 |
| DEL PEAK 11(8P21.3) MUTATED | 31 | 62 | 4 |
| DEL PEAK 11(8P21.3) WILD-TYPE | 38 | 23 | 1 |
Figure S4. Get High-res Image Gene #32: 'DEL PEAK 11(8P21.3) MUTATION STATUS' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
-
Copy number data file = transformed.cor.cli.txt
-
Clinical data file = PRAD-TP.clin.merged.picked.txt
-
Number of patients = 160
-
Number of significantly focal cnvs = 46
-
Number of selected clinical features = 6
-
Exclude genes that fewer than K tumors have mutations, K = 3
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For continuous numerical clinical features, two-tailed Student's t test with unequal variance (Lehmann and Romano 2005) was applied to compare the clinical values between tumors with and without gene mutations using 't.test' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.