This pipeline computes the correlation between significant arm-level copy number variations (cnvs) and selected clinical features.
Testing the association between copy number variation 78 arm-level events and 3 clinical features across 57 patients, 5 significant findings detected with Q value < 0.25.
-
7Q GAIN MUTATION ANALYSIS cnv correlated to 'AGE'.
-
8P GAIN MUTATION ANALYSIS cnv correlated to 'Time to Death'.
-
8Q GAIN MUTATION ANALYSIS cnv correlated to 'Time to Death'.
-
12P GAIN MUTATION ANALYSIS cnv correlated to 'Time to Death' and 'AGE'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 78 arm-level events and 3 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 5 significant findings detected.
|
Clinical Features |
Time to Death |
AGE | GENDER | ||
| nCNV (%) | nWild-Type | logrank test | t-test | Fisher's exact test | |
| 12P GAIN MUTATION ANALYSIS | 8 (14%) | 49 |
0.00024 (0.0554) |
6.58e-05 (0.0153) |
1 (1.00) |
| 7Q GAIN MUTATION ANALYSIS | 14 (25%) | 43 |
0.755 (1.00) |
0.000596 (0.137) |
0.23 (1.00) |
| 8P GAIN MUTATION ANALYSIS | 12 (21%) | 45 |
2.87e-05 (0.00671) |
0.13 (1.00) |
0.53 (1.00) |
| 8Q GAIN MUTATION ANALYSIS | 12 (21%) | 45 |
6.12e-05 (0.0143) |
0.252 (1.00) |
1 (1.00) |
| 1P GAIN MUTATION ANALYSIS | 11 (19%) | 46 |
0.0108 (1.00) |
0.0238 (1.00) |
0.747 (1.00) |
| 1Q GAIN MUTATION ANALYSIS | 9 (16%) | 48 |
0.796 (1.00) |
0.0322 (1.00) |
0.73 (1.00) |
| 2P GAIN MUTATION ANALYSIS | 9 (16%) | 48 |
0.273 (1.00) |
0.0451 (1.00) |
0.73 (1.00) |
| 2Q GAIN MUTATION ANALYSIS | 6 (11%) | 51 |
0.709 (1.00) |
0.204 (1.00) |
1 (1.00) |
| 3P GAIN MUTATION ANALYSIS | 6 (11%) | 51 |
0.551 (1.00) |
0.426 (1.00) |
1 (1.00) |
| 3Q GAIN MUTATION ANALYSIS | 7 (12%) | 50 |
0.14 (1.00) |
0.323 (1.00) |
0.706 (1.00) |
| 4P GAIN MUTATION ANALYSIS | 10 (18%) | 47 |
0.823 (1.00) |
0.199 (1.00) |
0.504 (1.00) |
| 4Q GAIN MUTATION ANALYSIS | 8 (14%) | 49 |
0.936 (1.00) |
0.337 (1.00) |
0.706 (1.00) |
| 5P GAIN MUTATION ANALYSIS | 13 (23%) | 44 |
0.642 (1.00) |
0.0377 (1.00) |
1 (1.00) |
| 5Q GAIN MUTATION ANALYSIS | 11 (19%) | 46 |
0.447 (1.00) |
0.161 (1.00) |
0.331 (1.00) |
| 6P GAIN MUTATION ANALYSIS | 10 (18%) | 47 |
0.945 (1.00) |
0.00694 (1.00) |
1 (1.00) |
| 6Q GAIN MUTATION ANALYSIS | 12 (21%) | 45 |
0.706 (1.00) |
0.0363 (1.00) |
0.747 (1.00) |
| 7P GAIN MUTATION ANALYSIS | 16 (28%) | 41 |
0.646 (1.00) |
0.00132 (0.302) |
0.0819 (1.00) |
| 9P GAIN MUTATION ANALYSIS | 9 (16%) | 48 |
0.621 (1.00) |
0.0144 (1.00) |
1 (1.00) |
| 9Q GAIN MUTATION ANALYSIS | 14 (25%) | 43 |
0.225 (1.00) |
0.0452 (1.00) |
0.76 (1.00) |
| 10P GAIN MUTATION ANALYSIS | 5 (9%) | 52 |
0.00429 (0.965) |
0.598 (1.00) |
1 (1.00) |
| 11P GAIN MUTATION ANALYSIS | 3 (5%) | 54 |
0.43 (1.00) |
0.0642 (1.00) |
0.112 (1.00) |
| 11Q GAIN MUTATION ANALYSIS | 3 (5%) | 54 |
0.454 (1.00) |
0.476 (1.00) |
0.112 (1.00) |
| 12Q GAIN MUTATION ANALYSIS | 4 (7%) | 53 |
0.176 (1.00) |
0.0115 (1.00) |
1 (1.00) |
| 14Q GAIN MUTATION ANALYSIS | 9 (16%) | 48 |
0.207 (1.00) |
0.636 (1.00) |
1 (1.00) |
| 15Q GAIN MUTATION ANALYSIS | 12 (21%) | 45 |
0.357 (1.00) |
0.0262 (1.00) |
1 (1.00) |
| 16P GAIN MUTATION ANALYSIS | 11 (19%) | 46 |
0.0463 (1.00) |
0.43 (1.00) |
0.747 (1.00) |
| 16Q GAIN MUTATION ANALYSIS | 4 (7%) | 53 |
0.803 (1.00) |
0.826 (1.00) |
1 (1.00) |
| 17P GAIN MUTATION ANALYSIS | 12 (21%) | 45 |
0.0334 (1.00) |
0.0258 (1.00) |
1 (1.00) |
| 17Q GAIN MUTATION ANALYSIS | 10 (18%) | 47 |
0.413 (1.00) |
0.204 (1.00) |
0.179 (1.00) |
| 18P GAIN MUTATION ANALYSIS | 10 (18%) | 47 |
0.31 (1.00) |
0.056 (1.00) |
0.179 (1.00) |
| 18Q GAIN MUTATION ANALYSIS | 10 (18%) | 47 |
0.957 (1.00) |
0.106 (1.00) |
0.0411 (1.00) |
| 19P GAIN MUTATION ANALYSIS | 19 (33%) | 38 |
0.0286 (1.00) |
0.00885 (1.00) |
0.576 (1.00) |
| 19Q GAIN MUTATION ANALYSIS | 13 (23%) | 44 |
0.517 (1.00) |
0.0259 (1.00) |
1 (1.00) |
| 20P GAIN MUTATION ANALYSIS | 17 (30%) | 40 |
0.0847 (1.00) |
0.0396 (1.00) |
0.777 (1.00) |
| 20Q GAIN MUTATION ANALYSIS | 20 (35%) | 37 |
0.221 (1.00) |
0.174 (1.00) |
1 (1.00) |
| 21Q GAIN MUTATION ANALYSIS | 11 (19%) | 46 |
0.0197 (1.00) |
0.118 (1.00) |
0.504 (1.00) |
| 22Q GAIN MUTATION ANALYSIS | 9 (16%) | 48 |
0.325 (1.00) |
0.105 (1.00) |
0.0787 (1.00) |
| XQ GAIN MUTATION ANALYSIS | 7 (12%) | 50 |
0.704 (1.00) |
0.677 (1.00) |
1 (1.00) |
| 1P LOSS MUTATION ANALYSIS | 10 (18%) | 47 |
0.112 (1.00) |
0.247 (1.00) |
0.0411 (1.00) |
| 1Q LOSS MUTATION ANALYSIS | 10 (18%) | 47 |
0.604 (1.00) |
0.353 (1.00) |
0.504 (1.00) |
| 2P LOSS MUTATION ANALYSIS | 10 (18%) | 47 |
0.902 (1.00) |
0.0102 (1.00) |
1 (1.00) |
| 2Q LOSS MUTATION ANALYSIS | 12 (21%) | 45 |
0.209 (1.00) |
0.00209 (0.475) |
1 (1.00) |
| 3P LOSS MUTATION ANALYSIS | 12 (21%) | 45 |
0.326 (1.00) |
0.0697 (1.00) |
1 (1.00) |
| 3Q LOSS MUTATION ANALYSIS | 12 (21%) | 45 |
0.855 (1.00) |
0.0803 (1.00) |
1 (1.00) |
| 4P LOSS MUTATION ANALYSIS | 11 (19%) | 46 |
0.522 (1.00) |
0.433 (1.00) |
0.331 (1.00) |
| 4Q LOSS MUTATION ANALYSIS | 14 (25%) | 43 |
0.506 (1.00) |
0.627 (1.00) |
0.07 (1.00) |
| 5P LOSS MUTATION ANALYSIS | 7 (12%) | 50 |
0.0773 (1.00) |
0.358 (1.00) |
1 (1.00) |
| 5Q LOSS MUTATION ANALYSIS | 10 (18%) | 47 |
0.219 (1.00) |
0.598 (1.00) |
1 (1.00) |
| 6P LOSS MUTATION ANALYSIS | 10 (18%) | 47 |
0.212 (1.00) |
0.796 (1.00) |
0.504 (1.00) |
| 6Q LOSS MUTATION ANALYSIS | 5 (9%) | 52 |
0.186 (1.00) |
0.961 (1.00) |
0.194 (1.00) |
| 7P LOSS MUTATION ANALYSIS | 4 (7%) | 53 |
0.234 (1.00) |
0.42 (1.00) |
1 (1.00) |
| 7Q LOSS MUTATION ANALYSIS | 4 (7%) | 53 |
0.494 (1.00) |
0.0585 (1.00) |
0.611 (1.00) |
| 8P LOSS MUTATION ANALYSIS | 13 (23%) | 44 |
0.482 (1.00) |
0.0459 (1.00) |
0.358 (1.00) |
| 8Q LOSS MUTATION ANALYSIS | 11 (19%) | 46 |
0.482 (1.00) |
0.0137 (1.00) |
0.331 (1.00) |
| 9P LOSS MUTATION ANALYSIS | 17 (30%) | 40 |
0.576 (1.00) |
0.00861 (1.00) |
0.395 (1.00) |
| 9Q LOSS MUTATION ANALYSIS | 11 (19%) | 46 |
0.455 (1.00) |
0.0278 (1.00) |
0.331 (1.00) |
| 10P LOSS MUTATION ANALYSIS | 23 (40%) | 34 |
0.712 (1.00) |
0.013 (1.00) |
0.00297 (0.675) |
| 10Q LOSS MUTATION ANALYSIS | 25 (44%) | 32 |
0.186 (1.00) |
0.23 (1.00) |
0.0033 (0.746) |
| 11P LOSS MUTATION ANALYSIS | 25 (44%) | 32 |
0.332 (1.00) |
0.0112 (1.00) |
0.596 (1.00) |
| 11Q LOSS MUTATION ANALYSIS | 19 (33%) | 38 |
0.186 (1.00) |
0.0091 (1.00) |
0.576 (1.00) |
| 12P LOSS MUTATION ANALYSIS | 12 (21%) | 45 |
0.539 (1.00) |
0.391 (1.00) |
0.747 (1.00) |
| 12Q LOSS MUTATION ANALYSIS | 14 (25%) | 43 |
0.362 (1.00) |
0.745 (1.00) |
1 (1.00) |
| 13Q LOSS MUTATION ANALYSIS | 25 (44%) | 32 |
0.458 (1.00) |
0.0351 (1.00) |
0.186 (1.00) |
| 14Q LOSS MUTATION ANALYSIS | 21 (37%) | 36 |
0.192 (1.00) |
0.974 (1.00) |
0.787 (1.00) |
| 15Q LOSS MUTATION ANALYSIS | 11 (19%) | 46 |
0.16 (1.00) |
0.0651 (1.00) |
0.331 (1.00) |
| 16P LOSS MUTATION ANALYSIS | 15 (26%) | 42 |
0.491 (1.00) |
0.208 (1.00) |
0.379 (1.00) |
| 16Q LOSS MUTATION ANALYSIS | 27 (47%) | 30 |
0.103 (1.00) |
0.161 (1.00) |
0.189 (1.00) |
| 17P LOSS MUTATION ANALYSIS | 11 (19%) | 46 |
0.357 (1.00) |
0.464 (1.00) |
0.747 (1.00) |
| 17Q LOSS MUTATION ANALYSIS | 7 (12%) | 50 |
0.495 (1.00) |
0.217 (1.00) |
0.706 (1.00) |
| 18P LOSS MUTATION ANALYSIS | 14 (25%) | 43 |
0.728 (1.00) |
0.697 (1.00) |
0.07 (1.00) |
| 18Q LOSS MUTATION ANALYSIS | 15 (26%) | 42 |
0.319 (1.00) |
0.0162 (1.00) |
0.379 (1.00) |
| 19P LOSS MUTATION ANALYSIS | 4 (7%) | 53 |
0.297 (1.00) |
0.165 (1.00) |
0.352 (1.00) |
| 19Q LOSS MUTATION ANALYSIS | 10 (18%) | 47 |
0.882 (1.00) |
0.138 (1.00) |
0.73 (1.00) |
| 20P LOSS MUTATION ANALYSIS | 7 (12%) | 50 |
0.518 (1.00) |
0.26 (1.00) |
1 (1.00) |
| 20Q LOSS MUTATION ANALYSIS | 5 (9%) | 52 |
0.827 (1.00) |
0.207 (1.00) |
1 (1.00) |
| 21Q LOSS MUTATION ANALYSIS | 13 (23%) | 44 |
0.147 (1.00) |
0.207 (1.00) |
0.76 (1.00) |
| 22Q LOSS MUTATION ANALYSIS | 21 (37%) | 36 |
0.839 (1.00) |
0.567 (1.00) |
0.787 (1.00) |
| XQ LOSS MUTATION ANALYSIS | 19 (33%) | 38 |
0.82 (1.00) |
0.956 (1.00) |
0.783 (1.00) |
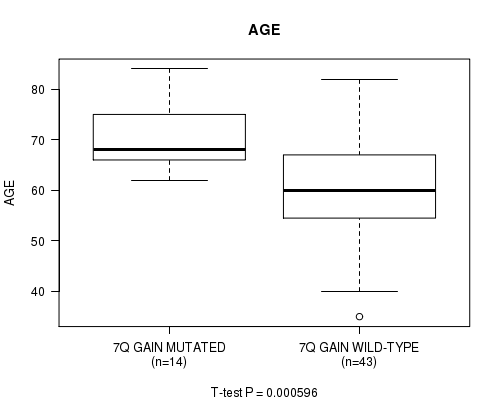
P value = 0.000596 (t-test), Q value = 0.14
Table S1. Gene #14: '7Q GAIN MUTATION STATUS' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 57 | 63.4 (11.5) |
| 7Q GAIN MUTATED | 14 | 70.5 (6.7) |
| 7Q GAIN WILD-TYPE | 43 | 61.0 (11.9) |
Figure S1. Get High-res Image Gene #14: '7Q GAIN MUTATION STATUS' versus Clinical Feature #2: 'AGE'
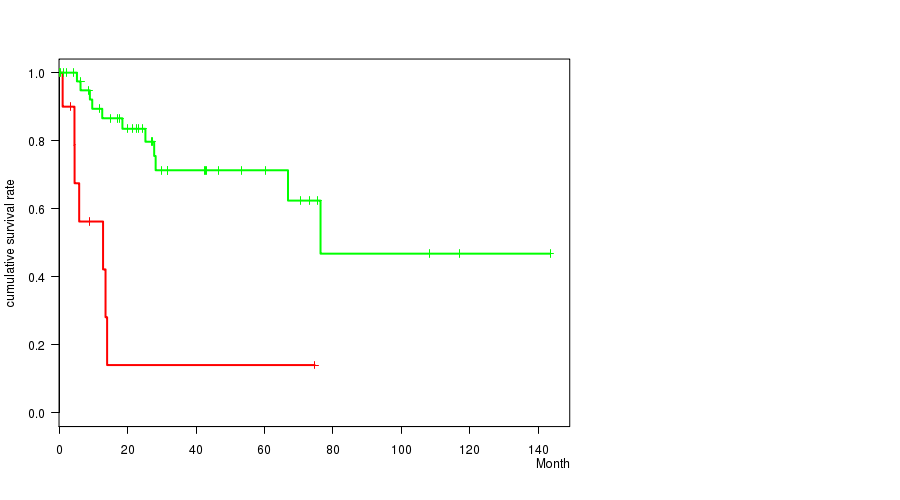
P value = 2.87e-05 (logrank test), Q value = 0.0067
Table S2. Gene #15: '8P GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 57 | 18 | 0.1 - 143.4 (18.5) |
| 8P GAIN MUTATED | 12 | 7 | 0.1 - 74.7 (5.2) |
| 8P GAIN WILD-TYPE | 45 | 11 | 0.1 - 143.4 (24.5) |
Figure S2. Get High-res Image Gene #15: '8P GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
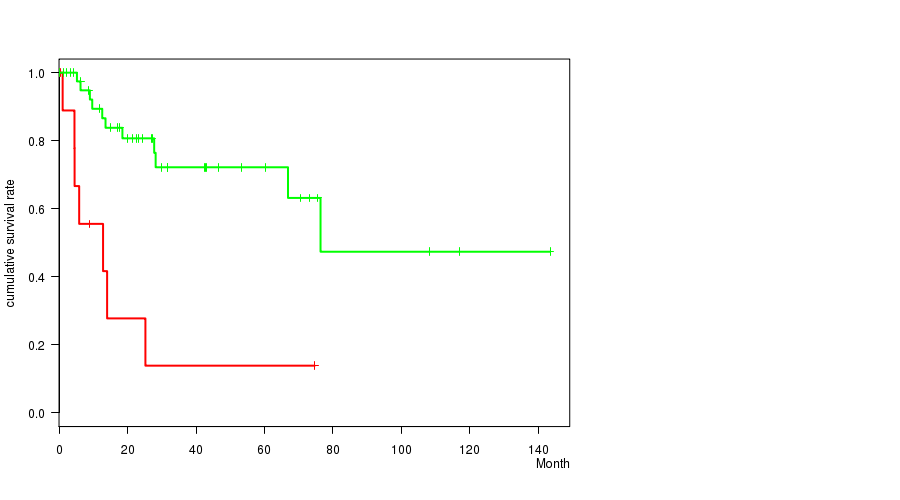
P value = 6.12e-05 (logrank test), Q value = 0.014
Table S3. Gene #16: '8Q GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 57 | 18 | 0.1 - 143.4 (18.5) |
| 8Q GAIN MUTATED | 12 | 7 | 0.1 - 74.7 (5.2) |
| 8Q GAIN WILD-TYPE | 45 | 11 | 0.1 - 143.4 (23.1) |
Figure S3. Get High-res Image Gene #16: '8Q GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
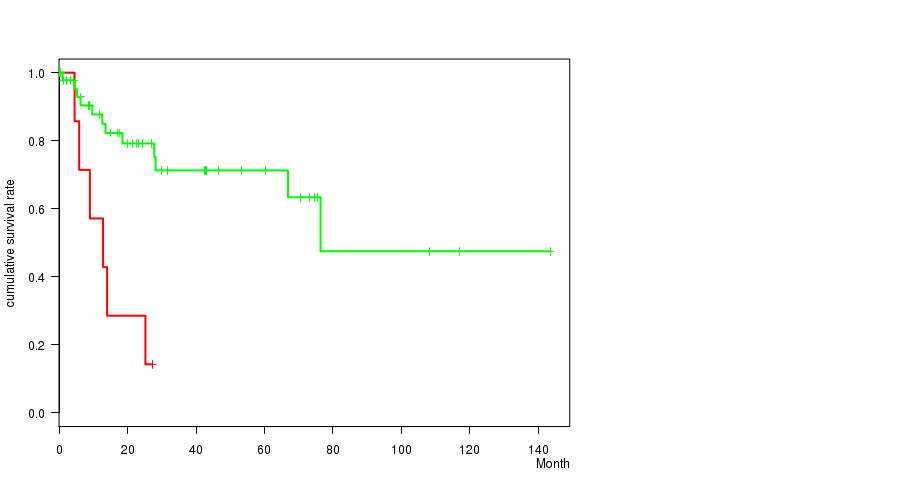
P value = 0.00024 (logrank test), Q value = 0.055
Table S4. Gene #22: '12P GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 57 | 18 | 0.1 - 143.4 (18.5) |
| 12P GAIN MUTATED | 8 | 6 | 0.1 - 27.2 (10.9) |
| 12P GAIN WILD-TYPE | 49 | 12 | 0.1 - 143.4 (21.4) |
Figure S4. Get High-res Image Gene #22: '12P GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
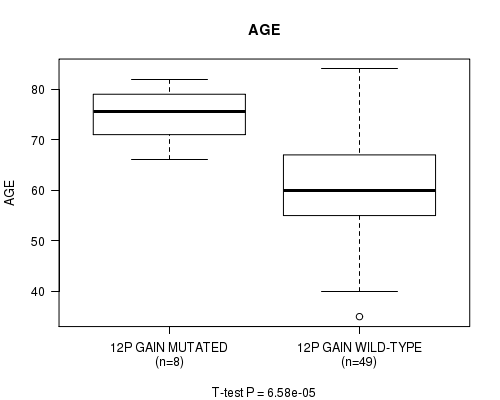
P value = 6.58e-05 (t-test), Q value = 0.015
Table S5. Gene #22: '12P GAIN MUTATION STATUS' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 57 | 63.4 (11.5) |
| 12P GAIN MUTATED | 8 | 74.9 (5.7) |
| 12P GAIN WILD-TYPE | 49 | 61.5 (11.2) |
Figure S5. Get High-res Image Gene #22: '12P GAIN MUTATION STATUS' versus Clinical Feature #2: 'AGE'
-
Copy number data file = transformed.cor.cli.txt
-
Clinical data file = SARC-TP.clin.merged.picked.txt
-
Number of patients = 57
-
Number of significantly arm-level cnvs = 78
-
Number of selected clinical features = 3
-
Exclude regions that fewer than K tumors have mutations, K = 3
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For continuous numerical clinical features, two-tailed Student's t test with unequal variance (Lehmann and Romano 2005) was applied to compare the clinical values between tumors with and without gene mutations using 't.test' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.