This pipeline computes the correlation between significantly recurrent gene mutations and molecular subtypes.
Testing the association between mutation status of 7 genes and 10 molecular subtypes across 401 patients, 30 significant findings detected with P value < 0.05 and Q value < 0.25.
-
NRAS mutation correlated to 'METHLYATION_CNMF', 'RPPA_CHIERARCHICAL', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
BRAF mutation correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'RPPA_CNMF', 'RPPA_CHIERARCHICAL', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
HRAS mutation correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'RPPA_CNMF', 'RPPA_CHIERARCHICAL', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
OTUD4 mutation correlated to 'MRNASEQ_CNMF'.
-
EIF1AX mutation correlated to 'MRNASEQ_CHIERARCHICAL'.
Table 1. Get Full Table Overview of the association between mutation status of 7 genes and 10 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 30 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
RPPA CNMF |
RPPA CHIERARCHICAL |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nMutated (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| BRAF | 240 (60%) | 161 |
0.000423 (0.0161) |
1.56e-41 (9.04e-40) |
1.51e-05 (0.000618) |
6.5e-10 (3.25e-08) |
8.11e-54 (5.19e-52) |
1.54e-51 (9.73e-50) |
9.16e-45 (5.5e-43) |
2.33e-49 (1.44e-47) |
6.03e-42 (3.56e-40) |
9.72e-48 (5.93e-46) |
| HRAS | 14 (3%) | 387 |
7.2e-06 (0.000302) |
1.13e-07 (5.32e-06) |
7.6e-05 (0.00304) |
0.00362 (0.134) |
9.11e-07 (3.92e-05) |
1.94e-07 (8.72e-06) |
1.79e-07 (8.22e-06) |
1.12e-08 (5.38e-07) |
4.91e-07 (2.16e-05) |
8.59e-09 (4.21e-07) |
| NRAS | 34 (8%) | 367 |
0.0106 (0.361) |
8.51e-19 (4.6e-17) |
1 (1.00) |
0.000377 (0.0147) |
1.44e-17 (7.65e-16) |
2.64e-19 (1.48e-17) |
1.04e-16 (5.41e-15) |
4.4e-19 (2.42e-17) |
3.09e-15 (1.57e-13) |
2.21e-19 (1.26e-17) |
| OTUD4 | 5 (1%) | 396 |
1 (1.00) |
0.32 (1.00) |
0.00555 (0.194) |
0.0182 (0.601) |
1 (1.00) |
0.0707 (1.00) |
0.168 (1.00) |
0.384 (1.00) |
||
| EIF1AX | 6 (1%) | 395 |
0.73 (1.00) |
0.13 (1.00) |
0.324 (1.00) |
0.13 (1.00) |
0.137 (1.00) |
0.00381 (0.137) |
0.331 (1.00) |
0.586 (1.00) |
0.38 (1.00) |
0.516 (1.00) |
| NUP93 | 4 (1%) | 397 |
1 (1.00) |
0.827 (1.00) |
0.504 (1.00) |
0.87 (1.00) |
1 (1.00) |
1 (1.00) |
0.841 (1.00) |
1 (1.00) |
||
| NLRP6 | 3 (1%) | 398 |
0.518 (1.00) |
0.295 (1.00) |
0.37 (1.00) |
0.686 (1.00) |
0.777 (1.00) |
0.0536 (1.00) |
0.533 (1.00) |
0.0614 (1.00) |
P value = 8.51e-19 (Fisher's exact test), Q value = 4.6e-17
Table S1. Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 70 | 204 |
| NRAS MUTATED | 34 | 0 | 0 |
| NRAS WILD-TYPE | 93 | 70 | 204 |
Figure S1. Get High-res Image Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'

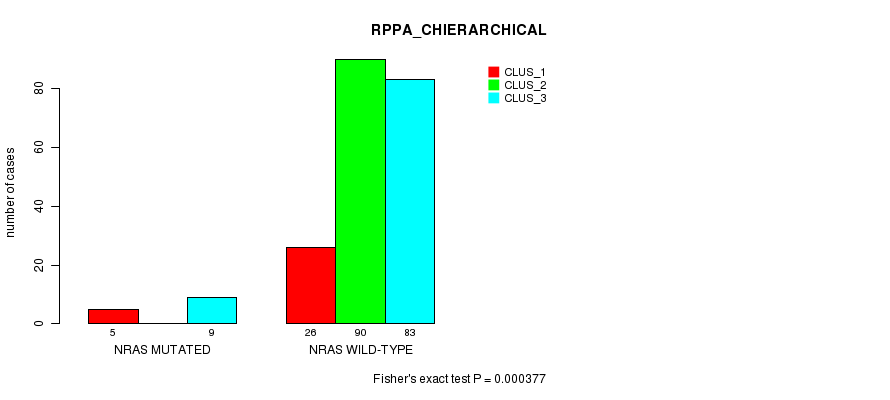
P value = 0.000377 (Fisher's exact test), Q value = 0.015
Table S2. Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 90 | 92 |
| NRAS MUTATED | 5 | 0 | 9 |
| NRAS WILD-TYPE | 26 | 90 | 83 |
Figure S2. Get High-res Image Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'
P value = 1.44e-17 (Fisher's exact test), Q value = 7.7e-16
Table S3. Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 125 | 46 | 91 | 128 |
| NRAS MUTATED | 34 | 0 | 0 | 0 |
| NRAS WILD-TYPE | 91 | 46 | 91 | 128 |
Figure S3. Get High-res Image Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'

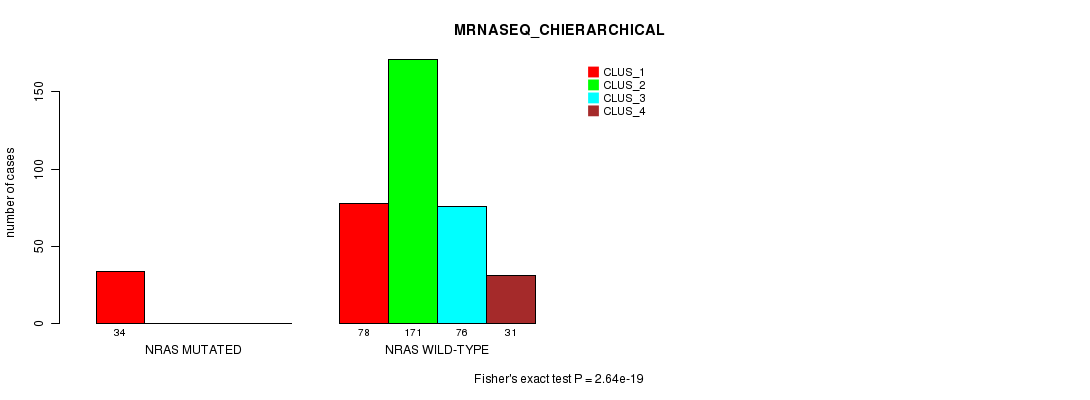
P value = 2.64e-19 (Fisher's exact test), Q value = 1.5e-17
Table S4. Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 112 | 171 | 76 | 31 |
| NRAS MUTATED | 34 | 0 | 0 | 0 |
| NRAS WILD-TYPE | 78 | 171 | 76 | 31 |
Figure S4. Get High-res Image Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
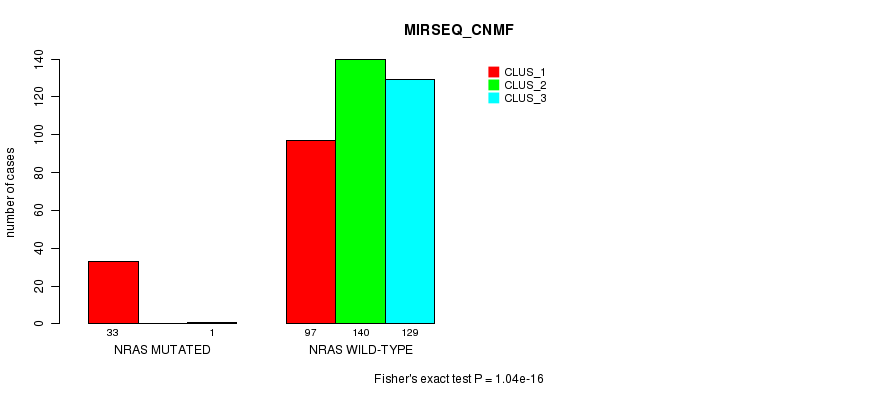
P value = 1.04e-16 (Fisher's exact test), Q value = 5.4e-15
Table S5. Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 130 | 140 | 130 |
| NRAS MUTATED | 33 | 0 | 1 |
| NRAS WILD-TYPE | 97 | 140 | 129 |
Figure S5. Get High-res Image Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
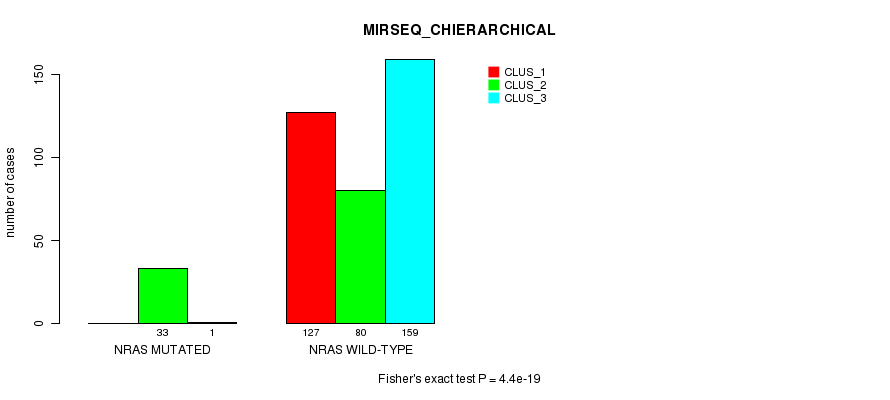
P value = 4.4e-19 (Fisher's exact test), Q value = 2.4e-17
Table S6. Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 113 | 160 |
| NRAS MUTATED | 0 | 33 | 1 |
| NRAS WILD-TYPE | 127 | 80 | 159 |
Figure S6. Get High-res Image Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
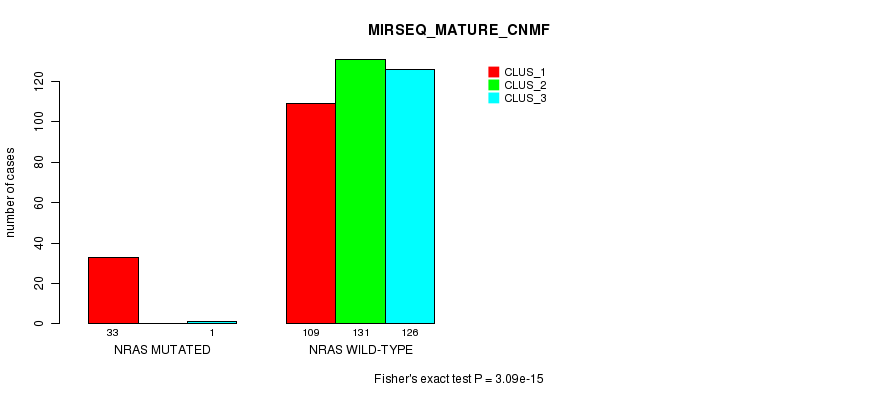
P value = 3.09e-15 (Fisher's exact test), Q value = 1.6e-13
Table S7. Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 142 | 131 | 127 |
| NRAS MUTATED | 33 | 0 | 1 |
| NRAS WILD-TYPE | 109 | 131 | 126 |
Figure S7. Get High-res Image Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #9: 'MIRSEQ_MATURE_CNMF'
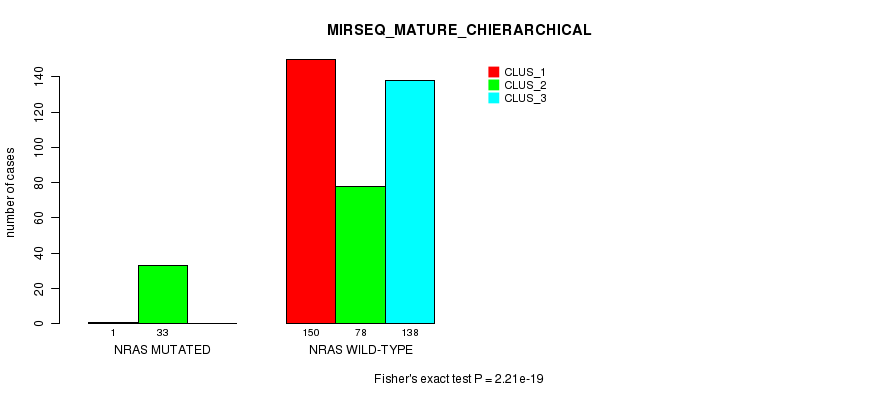
P value = 2.21e-19 (Fisher's exact test), Q value = 1.3e-17
Table S8. Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 151 | 111 | 138 |
| NRAS MUTATED | 1 | 33 | 0 |
| NRAS WILD-TYPE | 150 | 78 | 138 |
Figure S8. Get High-res Image Gene #1: 'NRAS MUTATION STATUS' versus Clinical Feature #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
P value = 0.000423 (Fisher's exact test), Q value = 0.016
Table S9. Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 26 | 313 | 60 |
| BRAF MUTATED | 8 | 203 | 29 |
| BRAF WILD-TYPE | 18 | 110 | 31 |
Figure S9. Get High-res Image Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'

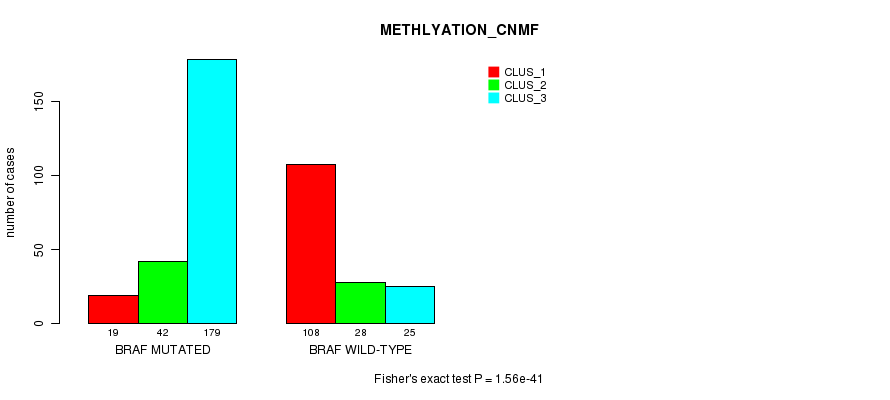
P value = 1.56e-41 (Fisher's exact test), Q value = 9e-40
Table S10. Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 70 | 204 |
| BRAF MUTATED | 19 | 42 | 179 |
| BRAF WILD-TYPE | 108 | 28 | 25 |
Figure S10. Get High-res Image Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
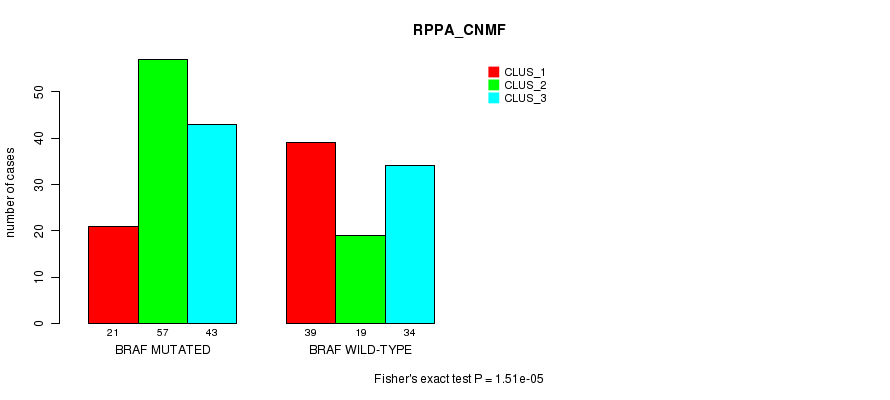
P value = 1.51e-05 (Fisher's exact test), Q value = 0.00062
Table S11. Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 76 | 77 |
| BRAF MUTATED | 21 | 57 | 43 |
| BRAF WILD-TYPE | 39 | 19 | 34 |
Figure S11. Get High-res Image Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
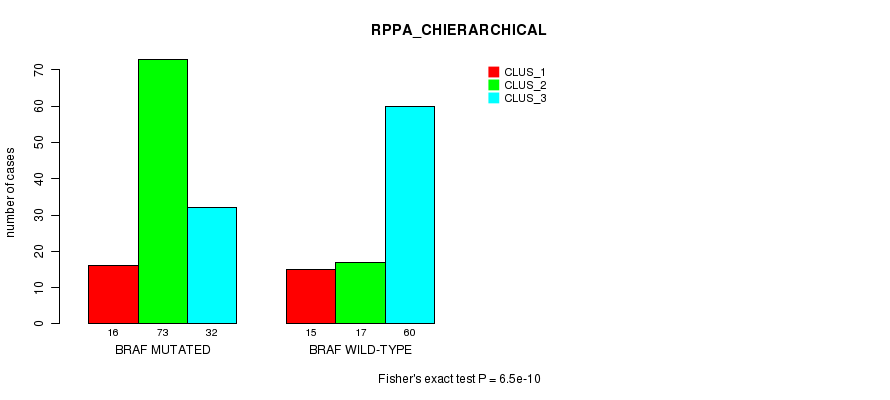
P value = 6.5e-10 (Fisher's exact test), Q value = 3.3e-08
Table S12. Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 90 | 92 |
| BRAF MUTATED | 16 | 73 | 32 |
| BRAF WILD-TYPE | 15 | 17 | 60 |
Figure S12. Get High-res Image Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'
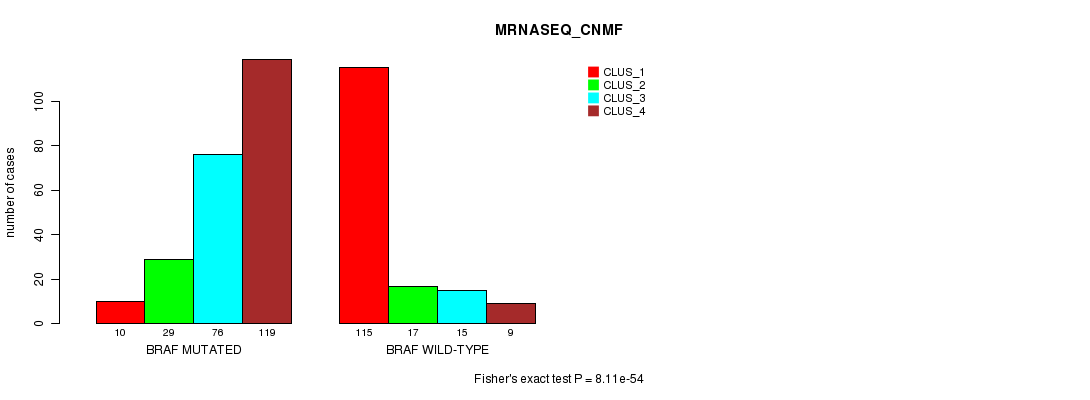
P value = 8.11e-54 (Fisher's exact test), Q value = 5.2e-52
Table S13. Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 125 | 46 | 91 | 128 |
| BRAF MUTATED | 10 | 29 | 76 | 119 |
| BRAF WILD-TYPE | 115 | 17 | 15 | 9 |
Figure S13. Get High-res Image Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
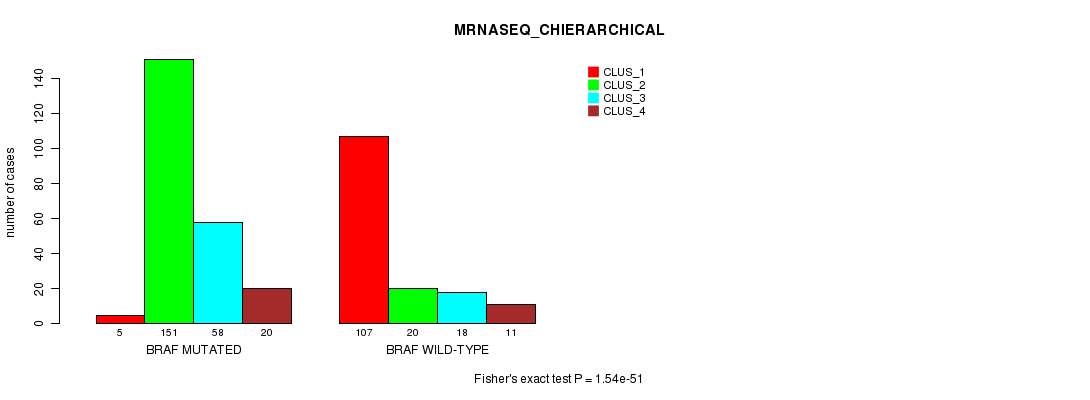
P value = 1.54e-51 (Fisher's exact test), Q value = 9.7e-50
Table S14. Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 112 | 171 | 76 | 31 |
| BRAF MUTATED | 5 | 151 | 58 | 20 |
| BRAF WILD-TYPE | 107 | 20 | 18 | 11 |
Figure S14. Get High-res Image Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
P value = 9.16e-45 (Fisher's exact test), Q value = 5.5e-43
Table S15. Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 130 | 140 | 130 |
| BRAF MUTATED | 15 | 122 | 102 |
| BRAF WILD-TYPE | 115 | 18 | 28 |
Figure S15. Get High-res Image Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'

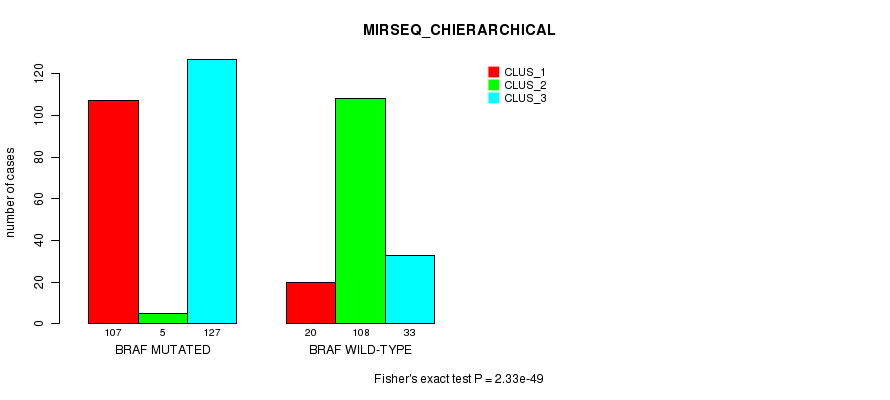
P value = 2.33e-49 (Fisher's exact test), Q value = 1.4e-47
Table S16. Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 113 | 160 |
| BRAF MUTATED | 107 | 5 | 127 |
| BRAF WILD-TYPE | 20 | 108 | 33 |
Figure S16. Get High-res Image Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
P value = 6.03e-42 (Fisher's exact test), Q value = 3.6e-40
Table S17. Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 142 | 131 | 127 |
| BRAF MUTATED | 23 | 117 | 99 |
| BRAF WILD-TYPE | 119 | 14 | 28 |
Figure S17. Get High-res Image Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #9: 'MIRSEQ_MATURE_CNMF'

P value = 9.72e-48 (Fisher's exact test), Q value = 5.9e-46
Table S18. Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 151 | 111 | 138 |
| BRAF MUTATED | 122 | 5 | 112 |
| BRAF WILD-TYPE | 29 | 106 | 26 |
Figure S18. Get High-res Image Gene #2: 'BRAF MUTATION STATUS' versus Clinical Feature #10: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 7.2e-06 (Fisher's exact test), Q value = 3e-04
Table S19. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 26 | 313 | 60 |
| HRAS MUTATED | 0 | 4 | 10 |
| HRAS WILD-TYPE | 26 | 309 | 50 |
Figure S19. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
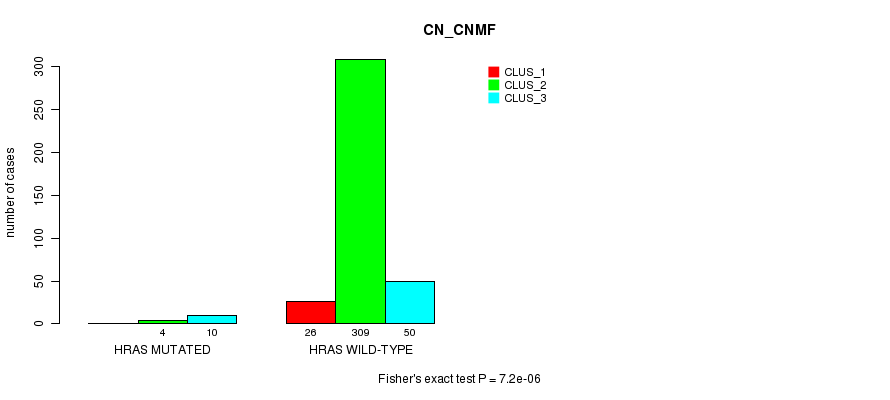
P value = 1.13e-07 (Fisher's exact test), Q value = 5.3e-06
Table S20. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 70 | 204 |
| HRAS MUTATED | 14 | 0 | 0 |
| HRAS WILD-TYPE | 113 | 70 | 204 |
Figure S20. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
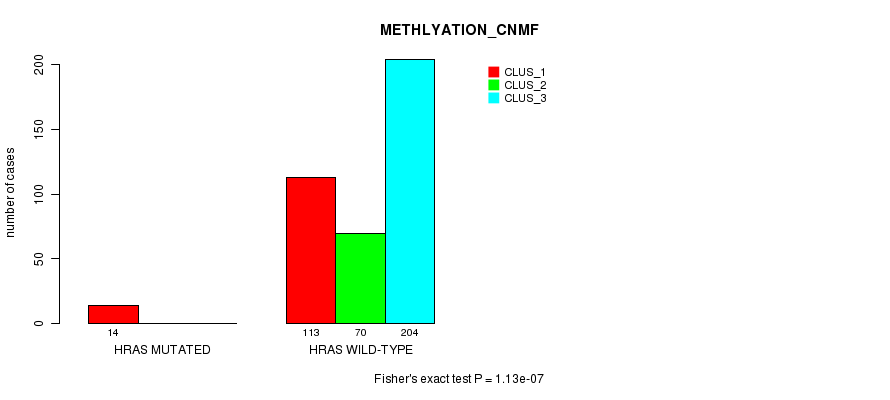
P value = 7.6e-05 (Fisher's exact test), Q value = 0.003
Table S21. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 76 | 77 |
| HRAS MUTATED | 9 | 1 | 0 |
| HRAS WILD-TYPE | 51 | 75 | 77 |
Figure S21. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
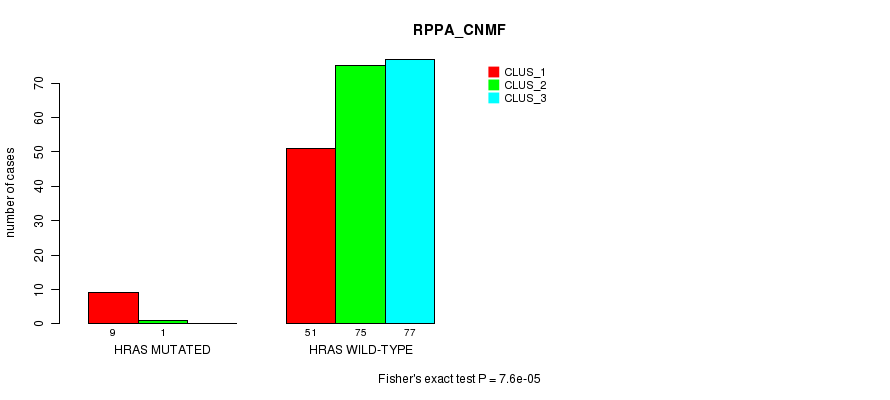
P value = 0.00362 (Fisher's exact test), Q value = 0.13
Table S22. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 90 | 92 |
| HRAS MUTATED | 1 | 0 | 9 |
| HRAS WILD-TYPE | 30 | 90 | 83 |
Figure S22. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'

P value = 9.11e-07 (Fisher's exact test), Q value = 3.9e-05
Table S23. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 125 | 46 | 91 | 128 |
| HRAS MUTATED | 14 | 0 | 0 | 0 |
| HRAS WILD-TYPE | 111 | 46 | 91 | 128 |
Figure S23. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'

P value = 1.94e-07 (Fisher's exact test), Q value = 8.7e-06
Table S24. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 112 | 171 | 76 | 31 |
| HRAS MUTATED | 14 | 0 | 0 | 0 |
| HRAS WILD-TYPE | 98 | 171 | 76 | 31 |
Figure S24. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
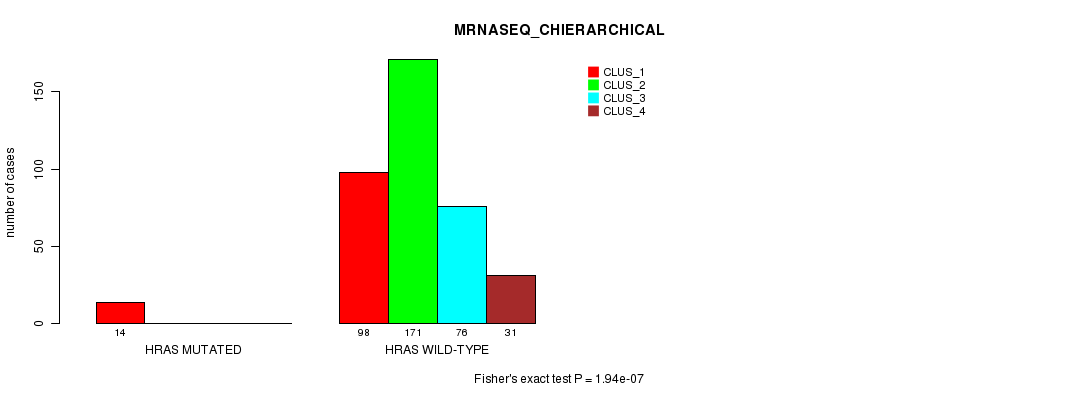
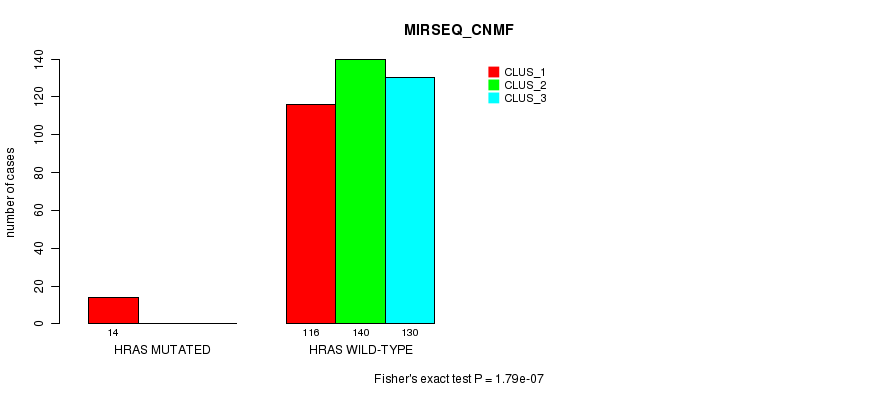
P value = 1.79e-07 (Fisher's exact test), Q value = 8.2e-06
Table S25. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 130 | 140 | 130 |
| HRAS MUTATED | 14 | 0 | 0 |
| HRAS WILD-TYPE | 116 | 140 | 130 |
Figure S25. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
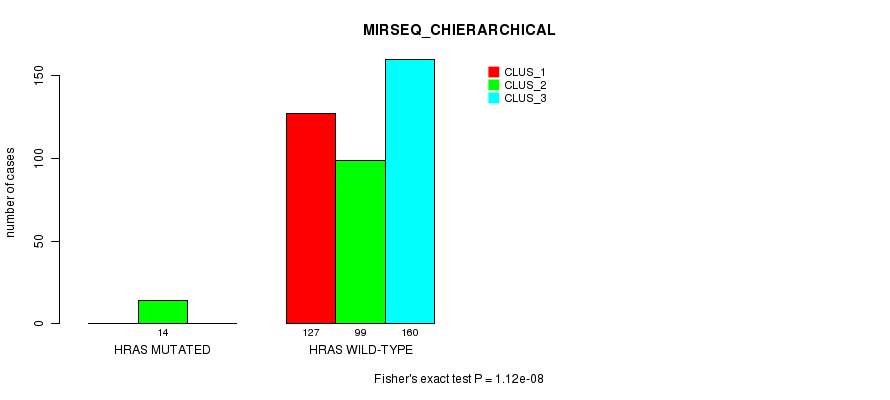
P value = 1.12e-08 (Fisher's exact test), Q value = 5.4e-07
Table S26. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 113 | 160 |
| HRAS MUTATED | 0 | 14 | 0 |
| HRAS WILD-TYPE | 127 | 99 | 160 |
Figure S26. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
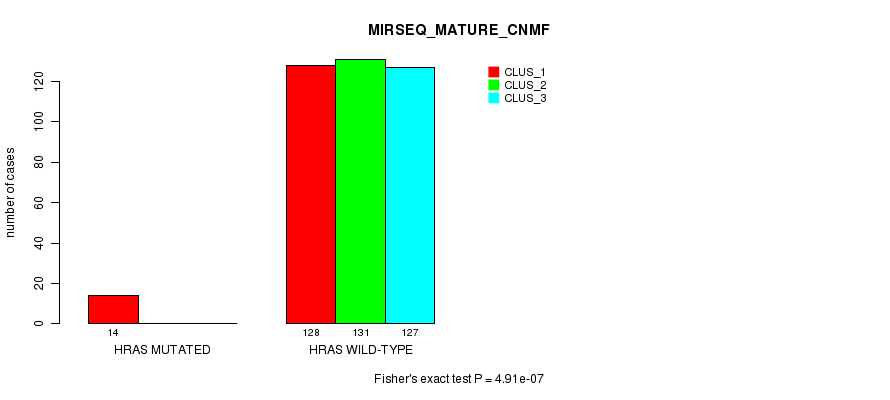
P value = 4.91e-07 (Fisher's exact test), Q value = 2.2e-05
Table S27. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 142 | 131 | 127 |
| HRAS MUTATED | 14 | 0 | 0 |
| HRAS WILD-TYPE | 128 | 131 | 127 |
Figure S27. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #9: 'MIRSEQ_MATURE_CNMF'
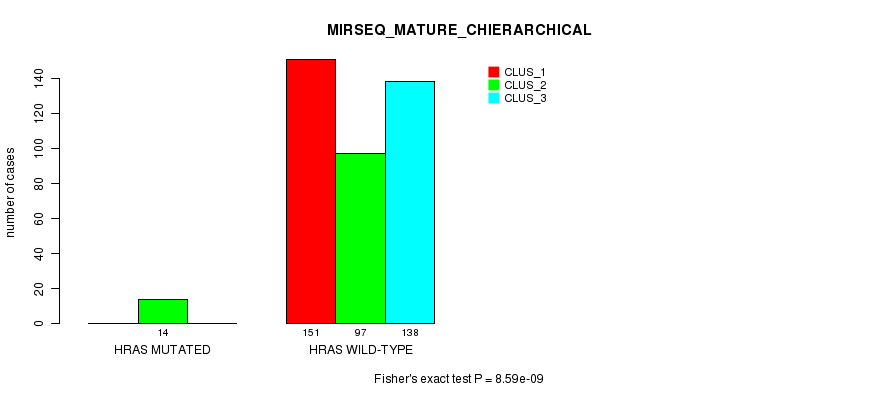
P value = 8.59e-09 (Fisher's exact test), Q value = 4.2e-07
Table S28. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 151 | 111 | 138 |
| HRAS MUTATED | 0 | 14 | 0 |
| HRAS WILD-TYPE | 151 | 97 | 138 |
Figure S28. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
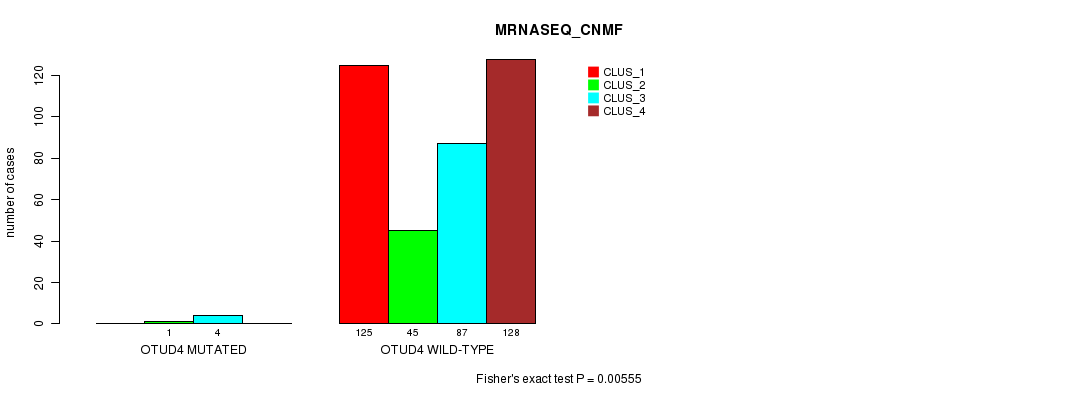
P value = 0.00555 (Fisher's exact test), Q value = 0.19
Table S29. Gene #4: 'OTUD4 MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 125 | 46 | 91 | 128 |
| OTUD4 MUTATED | 0 | 1 | 4 | 0 |
| OTUD4 WILD-TYPE | 125 | 45 | 87 | 128 |
Figure S29. Get High-res Image Gene #4: 'OTUD4 MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
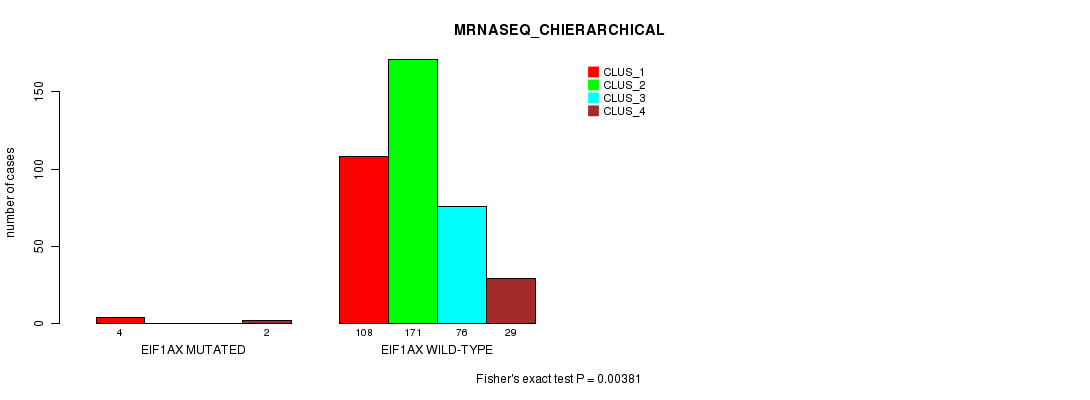
P value = 0.00381 (Fisher's exact test), Q value = 0.14
Table S30. Gene #5: 'EIF1AX MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 112 | 171 | 76 | 31 |
| EIF1AX MUTATED | 4 | 0 | 0 | 2 |
| EIF1AX WILD-TYPE | 108 | 171 | 76 | 29 |
Figure S30. Get High-res Image Gene #5: 'EIF1AX MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
-
Mutation data file = THCA-TP.mutsig.cluster.txt
-
Molecular subtypes file = THCA-TP.transferedmergedcluster.txt
-
Number of patients = 401
-
Number of significantly mutated genes = 7
-
Number of Molecular subtypes = 10
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.