This pipeline computes the correlation between significantly recurrent gene mutations and molecular subtypes.
Testing the association between mutation status of 6 genes and 8 molecular subtypes across 66 patients, no significant finding detected with P value < 0.05 and Q value < 0.25.
-
No gene mutations related to molecuar subtypes.
Table 1. Get Full Table Overview of the association between mutation status of 6 genes and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, no significant finding detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nMutated (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Chi-square test | Fisher's exact test | Fisher's exact test | Chi-square test | Fisher's exact test | |
| MUC2 | 8 (12%) | 58 |
1 (1.00) |
0.276 (1.00) |
0.745 (1.00) |
0.71 (1.00) |
0.572 (1.00) |
0.586 (1.00) |
0.49 (1.00) |
0.481 (1.00) |
| MUC6 | 21 (32%) | 45 |
0.178 (1.00) |
0.634 (1.00) |
0.253 (1.00) |
0.174 (1.00) |
0.306 (1.00) |
0.252 (1.00) |
0.205 (1.00) |
0.267 (1.00) |
| TP53 | 22 (33%) | 44 |
0.615 (1.00) |
0.0856 (1.00) |
0.0665 (1.00) |
0.0593 (1.00) |
0.655 (1.00) |
0.706 (1.00) |
0.76 (1.00) |
0.86 (1.00) |
| PTEN | 6 (9%) | 60 |
0.0988 (1.00) |
0.321 (1.00) |
0.351 (1.00) |
0.13 (1.00) |
0.354 (1.00) |
0.585 (1.00) |
0.00617 (0.296) |
0.0812 (1.00) |
| PRSS3 | 7 (11%) | 59 |
0.386 (1.00) |
0.668 (1.00) |
0.0182 (0.855) |
0.252 (1.00) |
0.0815 (1.00) |
0.581 (1.00) |
0.61 (1.00) |
0.159 (1.00) |
| HLA-C | 7 (11%) | 59 |
0.49 (1.00) |
0.574 (1.00) |
0.714 (1.00) |
0.569 (1.00) |
0.158 (1.00) |
0.581 (1.00) |
0.401 (1.00) |
0.561 (1.00) |
P value = 1 (Fisher's exact test), Q value = 1
Table S1. Gene #1: 'MUC2 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 5 | 47 | 14 |
| MUC2 MUTATED | 0 | 6 | 2 |
| MUC2 WILD-TYPE | 5 | 41 | 12 |
P value = 0.276 (Fisher's exact test), Q value = 1
Table S2. Gene #1: 'MUC2 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| MUC2 MUTATED | 2 | 6 | 0 |
| MUC2 WILD-TYPE | 16 | 29 | 13 |
P value = 0.745 (Fisher's exact test), Q value = 1
Table S3. Gene #1: 'MUC2 MUTATION STATUS' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| MUC2 MUTATED | 3 | 3 | 2 | 0 |
| MUC2 WILD-TYPE | 16 | 19 | 13 | 10 |
P value = 0.71 (Chi-square test), Q value = 1
Table S4. Gene #1: 'MUC2 MUTATION STATUS' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| MUC2 MUTATED | 1 | 0 | 2 | 1 | 4 |
| MUC2 WILD-TYPE | 9 | 10 | 15 | 5 | 19 |
P value = 0.572 (Fisher's exact test), Q value = 1
Table S5. Gene #1: 'MUC2 MUTATION STATUS' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 20 | 24 | 22 |
| MUC2 MUTATED | 1 | 4 | 3 |
| MUC2 WILD-TYPE | 19 | 20 | 19 |
P value = 0.586 (Fisher's exact test), Q value = 1
Table S6. Gene #1: 'MUC2 MUTATION STATUS' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 9 | 57 |
| MUC2 MUTATED | 0 | 8 |
| MUC2 WILD-TYPE | 9 | 49 |
P value = 0.49 (Chi-square test), Q value = 1
Table S7. Gene #1: 'MUC2 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 16 | 16 | 10 | 14 |
| MUC2 MUTATED | 1 | 1 | 4 | 1 | 1 |
| MUC2 WILD-TYPE | 9 | 15 | 12 | 9 | 13 |
P value = 0.481 (Fisher's exact test), Q value = 1
Table S8. Gene #1: 'MUC2 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| MUC2 MUTATED | 0 | 1 | 3 | 4 |
| MUC2 WILD-TYPE | 12 | 7 | 22 | 17 |
P value = 0.178 (Fisher's exact test), Q value = 1
Table S9. Gene #2: 'MUC6 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 5 | 47 | 14 |
| MUC6 MUTATED | 2 | 12 | 7 |
| MUC6 WILD-TYPE | 3 | 35 | 7 |
P value = 0.634 (Fisher's exact test), Q value = 1
Table S10. Gene #2: 'MUC6 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| MUC6 MUTATED | 7 | 11 | 3 |
| MUC6 WILD-TYPE | 11 | 24 | 10 |
P value = 0.253 (Fisher's exact test), Q value = 1
Table S11. Gene #2: 'MUC6 MUTATION STATUS' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| MUC6 MUTATED | 6 | 10 | 4 | 1 |
| MUC6 WILD-TYPE | 13 | 12 | 11 | 9 |
P value = 0.174 (Chi-square test), Q value = 1
Table S12. Gene #2: 'MUC6 MUTATION STATUS' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| MUC6 MUTATED | 2 | 1 | 9 | 2 | 7 |
| MUC6 WILD-TYPE | 8 | 9 | 8 | 4 | 16 |
P value = 0.306 (Fisher's exact test), Q value = 1
Table S13. Gene #2: 'MUC6 MUTATION STATUS' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 20 | 24 | 22 |
| MUC6 MUTATED | 4 | 10 | 7 |
| MUC6 WILD-TYPE | 16 | 14 | 15 |
P value = 0.252 (Fisher's exact test), Q value = 1
Table S14. Gene #2: 'MUC6 MUTATION STATUS' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 9 | 57 |
| MUC6 MUTATED | 1 | 20 |
| MUC6 WILD-TYPE | 8 | 37 |
P value = 0.205 (Chi-square test), Q value = 1
Table S15. Gene #2: 'MUC6 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 16 | 16 | 10 | 14 |
| MUC6 MUTATED | 6 | 5 | 3 | 4 | 3 |
| MUC6 WILD-TYPE | 4 | 11 | 13 | 6 | 11 |
P value = 0.267 (Fisher's exact test), Q value = 1
Table S16. Gene #2: 'MUC6 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| MUC6 MUTATED | 1 | 3 | 9 | 8 |
| MUC6 WILD-TYPE | 11 | 5 | 16 | 13 |
P value = 0.615 (Fisher's exact test), Q value = 1
Table S17. Gene #3: 'TP53 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 5 | 47 | 14 |
| TP53 MUTATED | 2 | 14 | 6 |
| TP53 WILD-TYPE | 3 | 33 | 8 |
P value = 0.0856 (Fisher's exact test), Q value = 1
Table S18. Gene #3: 'TP53 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| TP53 MUTATED | 3 | 16 | 3 |
| TP53 WILD-TYPE | 15 | 19 | 10 |
P value = 0.0665 (Fisher's exact test), Q value = 1
Table S19. Gene #3: 'TP53 MUTATION STATUS' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| TP53 MUTATED | 6 | 6 | 9 | 1 |
| TP53 WILD-TYPE | 13 | 16 | 6 | 9 |
P value = 0.0593 (Chi-square test), Q value = 1
Table S20. Gene #3: 'TP53 MUTATION STATUS' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| TP53 MUTATED | 7 | 1 | 6 | 2 | 6 |
| TP53 WILD-TYPE | 3 | 9 | 11 | 4 | 17 |
P value = 0.655 (Fisher's exact test), Q value = 1
Table S21. Gene #3: 'TP53 MUTATION STATUS' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 20 | 24 | 22 |
| TP53 MUTATED | 5 | 9 | 8 |
| TP53 WILD-TYPE | 15 | 15 | 14 |
P value = 0.706 (Fisher's exact test), Q value = 1
Table S22. Gene #3: 'TP53 MUTATION STATUS' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 9 | 57 |
| TP53 MUTATED | 2 | 20 |
| TP53 WILD-TYPE | 7 | 37 |
P value = 0.76 (Chi-square test), Q value = 1
Table S23. Gene #3: 'TP53 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 16 | 16 | 10 | 14 |
| TP53 MUTATED | 5 | 5 | 4 | 3 | 5 |
| TP53 WILD-TYPE | 5 | 11 | 12 | 7 | 9 |
P value = 0.86 (Fisher's exact test), Q value = 1
Table S24. Gene #3: 'TP53 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| TP53 MUTATED | 3 | 3 | 8 | 8 |
| TP53 WILD-TYPE | 9 | 5 | 17 | 13 |
P value = 0.0988 (Fisher's exact test), Q value = 1
Table S25. Gene #4: 'PTEN MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 5 | 47 | 14 |
| PTEN MUTATED | 2 | 3 | 1 |
| PTEN WILD-TYPE | 3 | 44 | 13 |
P value = 0.321 (Fisher's exact test), Q value = 1
Table S26. Gene #4: 'PTEN MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| PTEN MUTATED | 3 | 3 | 0 |
| PTEN WILD-TYPE | 15 | 32 | 13 |
P value = 0.351 (Fisher's exact test), Q value = 1
Table S27. Gene #4: 'PTEN MUTATION STATUS' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| PTEN MUTATED | 2 | 1 | 3 | 0 |
| PTEN WILD-TYPE | 17 | 21 | 12 | 10 |
P value = 0.13 (Chi-square test), Q value = 1
Table S28. Gene #4: 'PTEN MUTATION STATUS' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| PTEN MUTATED | 3 | 0 | 1 | 0 | 2 |
| PTEN WILD-TYPE | 7 | 10 | 16 | 6 | 21 |
P value = 0.354 (Fisher's exact test), Q value = 1
Table S29. Gene #4: 'PTEN MUTATION STATUS' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 20 | 24 | 22 |
| PTEN MUTATED | 1 | 4 | 1 |
| PTEN WILD-TYPE | 19 | 20 | 21 |
P value = 0.585 (Fisher's exact test), Q value = 1
Table S30. Gene #4: 'PTEN MUTATION STATUS' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 9 | 57 |
| PTEN MUTATED | 0 | 6 |
| PTEN WILD-TYPE | 9 | 51 |
P value = 0.00617 (Chi-square test), Q value = 0.3
Table S31. Gene #4: 'PTEN MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 16 | 16 | 10 | 14 |
| PTEN MUTATED | 4 | 0 | 1 | 0 | 1 |
| PTEN WILD-TYPE | 6 | 16 | 15 | 10 | 13 |
Figure S1. Get High-res Image Gene #4: 'PTEN MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
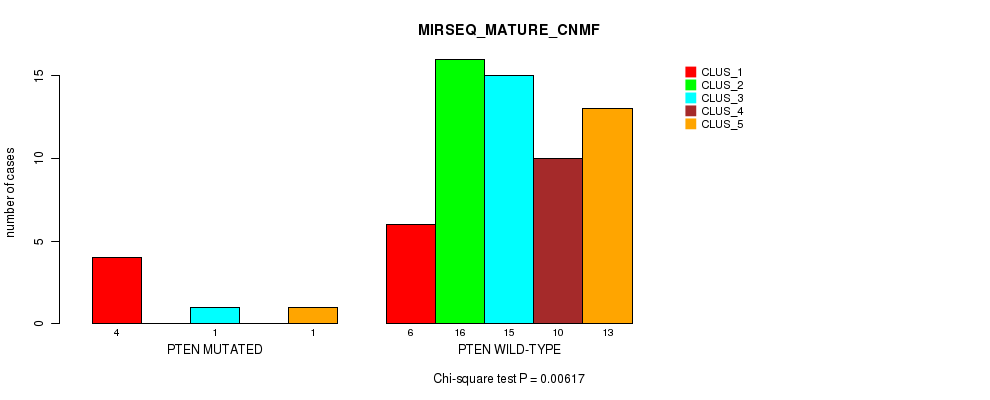
P value = 0.0812 (Fisher's exact test), Q value = 1
Table S32. Gene #4: 'PTEN MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| PTEN MUTATED | 0 | 0 | 1 | 5 |
| PTEN WILD-TYPE | 12 | 8 | 24 | 16 |
P value = 0.386 (Fisher's exact test), Q value = 1
Table S33. Gene #5: 'PRSS3 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 5 | 47 | 14 |
| PRSS3 MUTATED | 0 | 4 | 3 |
| PRSS3 WILD-TYPE | 5 | 43 | 11 |
P value = 0.668 (Fisher's exact test), Q value = 1
Table S34. Gene #5: 'PRSS3 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| PRSS3 MUTATED | 1 | 4 | 2 |
| PRSS3 WILD-TYPE | 17 | 31 | 11 |
P value = 0.0182 (Fisher's exact test), Q value = 0.86
Table S35. Gene #5: 'PRSS3 MUTATION STATUS' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| PRSS3 MUTATED | 0 | 6 | 1 | 0 |
| PRSS3 WILD-TYPE | 19 | 16 | 14 | 10 |
Figure S2. Get High-res Image Gene #5: 'PRSS3 MUTATION STATUS' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
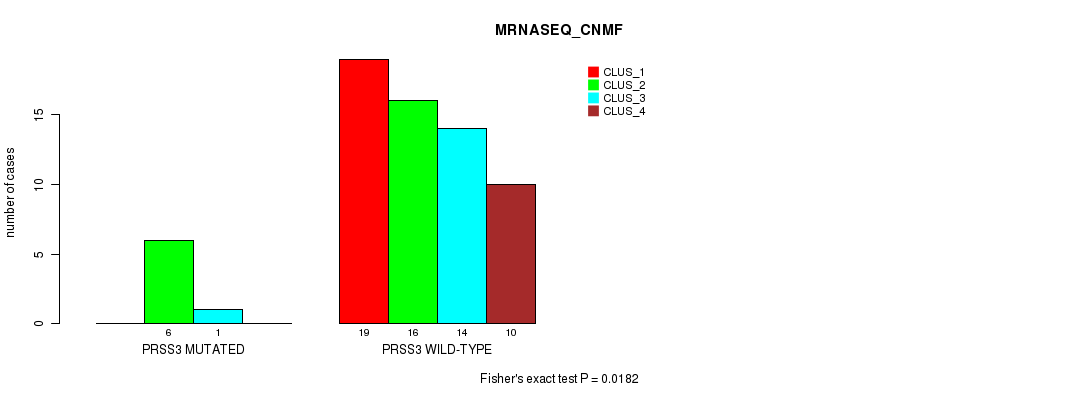
P value = 0.252 (Chi-square test), Q value = 1
Table S36. Gene #5: 'PRSS3 MUTATION STATUS' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| PRSS3 MUTATED | 1 | 0 | 4 | 1 | 1 |
| PRSS3 WILD-TYPE | 9 | 10 | 13 | 5 | 22 |
P value = 0.0815 (Fisher's exact test), Q value = 1
Table S37. Gene #5: 'PRSS3 MUTATION STATUS' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 20 | 24 | 22 |
| PRSS3 MUTATED | 3 | 0 | 4 |
| PRSS3 WILD-TYPE | 17 | 24 | 18 |
P value = 0.581 (Fisher's exact test), Q value = 1
Table S38. Gene #5: 'PRSS3 MUTATION STATUS' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 9 | 57 |
| PRSS3 MUTATED | 0 | 7 |
| PRSS3 WILD-TYPE | 9 | 50 |
P value = 0.61 (Chi-square test), Q value = 1
Table S39. Gene #5: 'PRSS3 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 16 | 16 | 10 | 14 |
| PRSS3 MUTATED | 0 | 2 | 1 | 2 | 2 |
| PRSS3 WILD-TYPE | 10 | 14 | 15 | 8 | 12 |
P value = 0.159 (Fisher's exact test), Q value = 1
Table S40. Gene #5: 'PRSS3 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| PRSS3 MUTATED | 2 | 1 | 4 | 0 |
| PRSS3 WILD-TYPE | 10 | 7 | 21 | 21 |
P value = 0.49 (Fisher's exact test), Q value = 1
Table S41. Gene #6: 'HLA-C MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 5 | 47 | 14 |
| HLA-C MUTATED | 1 | 4 | 2 |
| HLA-C WILD-TYPE | 4 | 43 | 12 |
P value = 0.574 (Fisher's exact test), Q value = 1
Table S42. Gene #6: 'HLA-C MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| HLA-C MUTATED | 3 | 3 | 1 |
| HLA-C WILD-TYPE | 15 | 32 | 12 |
P value = 0.714 (Fisher's exact test), Q value = 1
Table S43. Gene #6: 'HLA-C MUTATION STATUS' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| HLA-C MUTATED | 3 | 3 | 1 | 0 |
| HLA-C WILD-TYPE | 16 | 19 | 14 | 10 |
P value = 0.569 (Chi-square test), Q value = 1
Table S44. Gene #6: 'HLA-C MUTATION STATUS' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| HLA-C MUTATED | 1 | 0 | 3 | 0 | 3 |
| HLA-C WILD-TYPE | 9 | 10 | 14 | 6 | 20 |
P value = 0.158 (Fisher's exact test), Q value = 1
Table S45. Gene #6: 'HLA-C MUTATION STATUS' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 20 | 24 | 22 |
| HLA-C MUTATED | 0 | 3 | 4 |
| HLA-C WILD-TYPE | 20 | 21 | 18 |
P value = 0.581 (Fisher's exact test), Q value = 1
Table S46. Gene #6: 'HLA-C MUTATION STATUS' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 9 | 57 |
| HLA-C MUTATED | 0 | 7 |
| HLA-C WILD-TYPE | 9 | 50 |
P value = 0.401 (Chi-square test), Q value = 1
Table S47. Gene #6: 'HLA-C MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 16 | 16 | 10 | 14 |
| HLA-C MUTATED | 1 | 1 | 3 | 2 | 0 |
| HLA-C WILD-TYPE | 9 | 15 | 13 | 8 | 14 |
P value = 0.561 (Fisher's exact test), Q value = 1
Table S48. Gene #6: 'HLA-C MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| HLA-C MUTATED | 0 | 1 | 4 | 2 |
| HLA-C WILD-TYPE | 12 | 7 | 21 | 19 |
-
Mutation data file = transformed.cor.cli.txt
-
Molecular subtypes file = KICH-TP.transferedmergedcluster.txt
-
Number of patients = 66
-
Number of significantly mutated genes = 6
-
Number of Molecular subtypes = 8
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.