This pipeline uses various statistical tests to identify genes whose promoter methylation levels correlated to selected clinical features.
Testing the association between 19985 genes and 10 clinical features across 665 samples, statistically thresholded by Q value < 0.05, 8 clinical features related to at least one genes.
-
440 genes correlated to 'AGE'.
-
KIAA1143 , KIF15 , C1ORF103 , LGALS8 , MEX3C , ...
-
41 genes correlated to 'NEOPLASM.DISEASESTAGE'.
-
WDR74 , RHBDL3 , FASTKD3 , MTRR__1 , SIL1 , ...
-
2 genes correlated to 'PATHOLOGY.N.STAGE'.
-
TCP11L1 , ADA
-
68 genes correlated to 'PATHOLOGY.M.STAGE'.
-
ABCE1 , ANAPC10 , RHBDL3 , FASTKD3 , MTRR__1 , ...
-
191 genes correlated to 'GENDER'.
-
ALDOC , CRIP1 , NMNAT3 , DNAJC15 , EML1 , ...
-
1201 genes correlated to 'HISTOLOGICAL.TYPE'.
-
CLUAP1 , RNF25 , STK36 , PRKAR2A , FAR1 , ...
-
116 genes correlated to 'RADIATIONS.RADIATION.REGIMENINDICATION'.
-
PRIM1 , METAP2 , MAEA , HMG20B , ZNF639 , ...
-
1 gene correlated to 'NUMBER.OF.LYMPH.NODES'.
-
TCP11L1
-
No genes correlated to 'Time to Death', and 'PATHOLOGY.T.STAGE'.
Complete statistical result table is provided in Supplement Table 1
Table 1. Get Full Table This table shows the clinical features, statistical methods used, and the number of genes that are significantly associated with each clinical feature at Q value < 0.05.
| Clinical feature | Statistical test | Significant genes | Associated with | Associated with | ||
|---|---|---|---|---|---|---|
| Time to Death | Cox regression test | N=0 | ||||
| AGE | Spearman correlation test | N=440 | older | N=343 | younger | N=97 |
| NEOPLASM DISEASESTAGE | ANOVA test | N=41 | ||||
| PATHOLOGY T STAGE | Spearman correlation test | N=0 | ||||
| PATHOLOGY N STAGE | Spearman correlation test | N=2 | higher stage | N=2 | lower stage | N=0 |
| PATHOLOGY M STAGE | ANOVA test | N=68 | ||||
| GENDER | t test | N=191 | male | N=54 | female | N=137 |
| HISTOLOGICAL TYPE | ANOVA test | N=1201 | ||||
| RADIATIONS RADIATION REGIMENINDICATION | t test | N=116 | yes | N=47 | no | N=69 |
| NUMBER OF LYMPH NODES | Spearman correlation test | N=1 | higher number.of.lymph.nodes | N=1 | lower number.of.lymph.nodes | N=0 |
Table S1. Basic characteristics of clinical feature: 'Time to Death'
| Time to Death | Duration (Months) | 0-234.3 (median=21.4) |
| censored | N = 585 | |
| death | N = 74 | |
| Significant markers | N = 0 |
Table S2. Basic characteristics of clinical feature: 'AGE'
| AGE | Mean (SD) | 58.06 (13) |
| Significant markers | N = 440 | |
| pos. correlated | 343 | |
| neg. correlated | 97 |
Table S3. Get Full Table List of top 10 genes significantly correlated to 'AGE' by Spearman correlation test
| SpearmanCorr | corrP | Q | |
|---|---|---|---|
| KIAA1143 | 0.3293 | 5.277e-18 | 1.05e-13 |
| KIF15 | 0.3293 | 5.277e-18 | 1.05e-13 |
| C1ORF103 | 0.3106 | 4.285e-16 | 8.56e-12 |
| LGALS8 | -0.3097 | 5.259e-16 | 1.05e-11 |
| MEX3C | 0.2998 | 4.805e-15 | 9.6e-11 |
| C20ORF199 | 0.2972 | 8.42e-15 | 1.68e-10 |
| SNORD12 | 0.2972 | 8.42e-15 | 1.68e-10 |
| C9ORF69 | 0.2856 | 9.625e-14 | 1.92e-09 |
| DNMT3A | 0.2854 | 1.001e-13 | 2e-09 |
| CACNA2D1 | 0.2839 | 1.362e-13 | 2.72e-09 |
Figure S1. Get High-res Image As an example, this figure shows the association of KIAA1143 to 'AGE'. P value = 5.28e-18 with Spearman correlation analysis. The straight line presents the best linear regression.
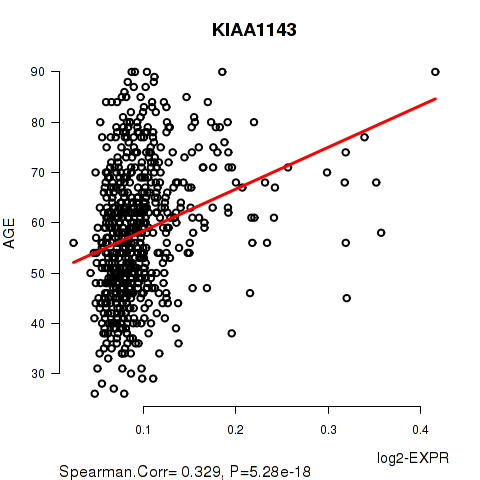
Table S4. Basic characteristics of clinical feature: 'NEOPLASM.DISEASESTAGE'
| NEOPLASM.DISEASESTAGE | Labels | N |
| STAGE I | 49 | |
| STAGE IA | 55 | |
| STAGE IB | 5 | |
| STAGE II | 8 | |
| STAGE IIA | 212 | |
| STAGE IIB | 155 | |
| STAGE III | 2 | |
| STAGE IIIA | 107 | |
| STAGE IIIB | 17 | |
| STAGE IIIC | 43 | |
| STAGE IV | 6 | |
| STAGE X | 5 | |
| Significant markers | N = 41 |
Table S5. Get Full Table List of top 10 genes differentially expressed by 'NEOPLASM.DISEASESTAGE'
| ANOVA_P | Q | |
|---|---|---|
| WDR74 | 1.747e-20 | 3.49e-16 |
| RHBDL3 | 2.37e-20 | 4.74e-16 |
| FASTKD3 | 7.679e-20 | 1.53e-15 |
| MTRR__1 | 7.679e-20 | 1.53e-15 |
| SIL1 | 7.436e-19 | 1.49e-14 |
| MMAB | 1.421e-17 | 2.84e-13 |
| MVK | 1.421e-17 | 2.84e-13 |
| ATP5J | 8.599e-17 | 1.72e-12 |
| GABPA | 8.599e-17 | 1.72e-12 |
| TTC32 | 4.644e-14 | 9.28e-10 |
Figure S2. Get High-res Image As an example, this figure shows the association of WDR74 to 'NEOPLASM.DISEASESTAGE'. P value = 1.75e-20 with ANOVA analysis.
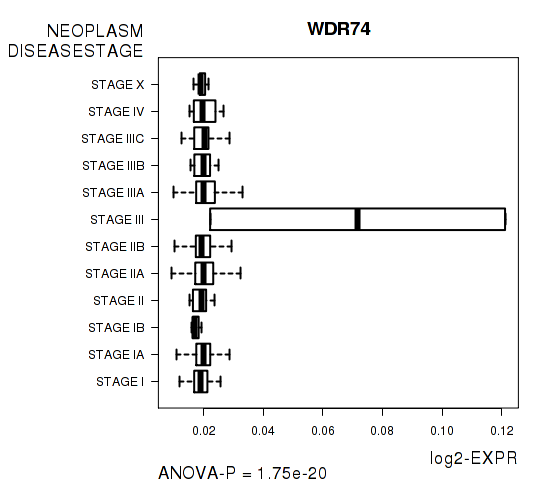
Table S6. Basic characteristics of clinical feature: 'PATHOLOGY.T.STAGE'
| PATHOLOGY.T.STAGE | Mean (SD) | 1.93 (0.71) |
| N | ||
| 1 | 174 | |
| 2 | 383 | |
| 3 | 86 | |
| 4 | 20 | |
| Significant markers | N = 0 |
Table S7. Basic characteristics of clinical feature: 'PATHOLOGY.N.STAGE'
| PATHOLOGY.N.STAGE | Mean (SD) | 0.82 (0.91) |
| N | ||
| 0 | 292 | |
| 1 | 235 | |
| 2 | 84 | |
| 3 | 46 | |
| Significant markers | N = 2 | |
| pos. correlated | 2 | |
| neg. correlated | 0 |
Table S8. Get Full Table List of 2 genes significantly correlated to 'PATHOLOGY.N.STAGE' by Spearman correlation test
| SpearmanCorr | corrP | Q | |
|---|---|---|---|
| TCP11L1 | 0.2024 | 1.677e-07 | 0.00335 |
| ADA | 0.1915 | 7.582e-07 | 0.0152 |
Figure S3. Get High-res Image As an example, this figure shows the association of TCP11L1 to 'PATHOLOGY.N.STAGE'. P value = 1.68e-07 with Spearman correlation analysis.
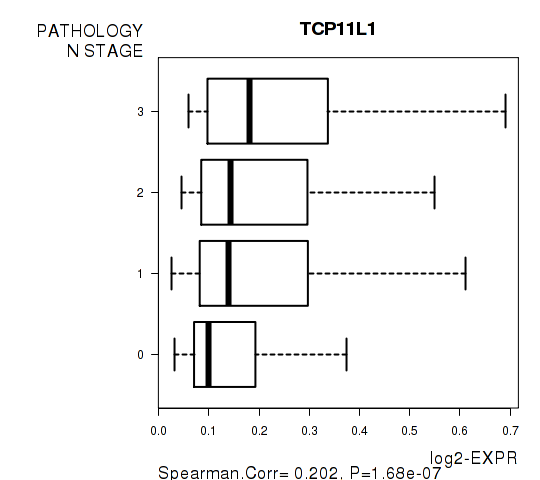
Table S9. Basic characteristics of clinical feature: 'PATHOLOGY.M.STAGE'
| PATHOLOGY.M.STAGE | Labels | N |
| CM0 (I+) | 1 | |
| M0 | 552 | |
| M1 | 6 | |
| MX | 106 | |
| Significant markers | N = 68 |
Table S10. Get Full Table List of top 10 genes differentially expressed by 'PATHOLOGY.M.STAGE'
| ANOVA_P | Q | |
|---|---|---|
| ABCE1 | 4.148e-108 | 8.29e-104 |
| ANAPC10 | 4.148e-108 | 8.29e-104 |
| RHBDL3 | 1.615e-25 | 3.23e-21 |
| FASTKD3 | 2.477e-25 | 4.95e-21 |
| MTRR__1 | 2.477e-25 | 4.95e-21 |
| MMAB | 1.28e-22 | 2.56e-18 |
| MVK | 1.28e-22 | 2.56e-18 |
| NHEDC1 | 1.348e-12 | 2.69e-08 |
| SAG | 2.353e-12 | 4.7e-08 |
| TSTD1 | 1.104e-10 | 2.2e-06 |
Figure S4. Get High-res Image As an example, this figure shows the association of ABCE1 to 'PATHOLOGY.M.STAGE'. P value = 4.15e-108 with ANOVA analysis.
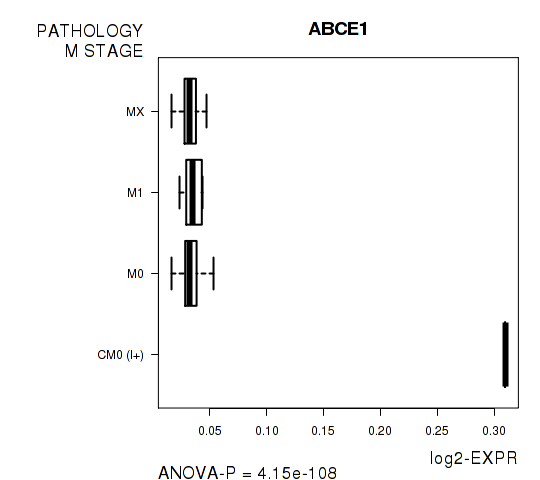
Table S11. Basic characteristics of clinical feature: 'GENDER'
| GENDER | Labels | N |
| FEMALE | 658 | |
| MALE | 7 | |
| Significant markers | N = 191 | |
| Higher in MALE | 54 | |
| Higher in FEMALE | 137 |
Table S12. Get Full Table List of top 10 genes differentially expressed by 'GENDER'
| T(pos if higher in 'MALE') | ttestP | Q | AUC | |
|---|---|---|---|---|
| ALDOC | -30.35 | 2.529e-72 | 5.05e-68 | 0.962 |
| CRIP1 | -20.11 | 1.688e-70 | 3.37e-66 | 0.8689 |
| NMNAT3 | -14.96 | 3.999e-43 | 7.99e-39 | 0.693 |
| DNAJC15 | -14.91 | 5.895e-42 | 1.18e-37 | 0.7093 |
| EML1 | -13.13 | 6.565e-35 | 1.31e-30 | 0.6235 |
| ADCY5 | 14.21 | 1.343e-31 | 2.68e-27 | 0.7312 |
| RND2 | -14.18 | 1.848e-31 | 3.69e-27 | 0.7579 |
| SLCO4C1 | -12.8 | 8.868e-30 | 1.77e-25 | 0.7178 |
| LOC400043 | -13.5 | 5.202e-29 | 1.04e-24 | 0.5597 |
| ACADS | -11.73 | 1.097e-28 | 2.19e-24 | 0.612 |
Figure S5. Get High-res Image As an example, this figure shows the association of ALDOC to 'GENDER'. P value = 2.53e-72 with T-test analysis.
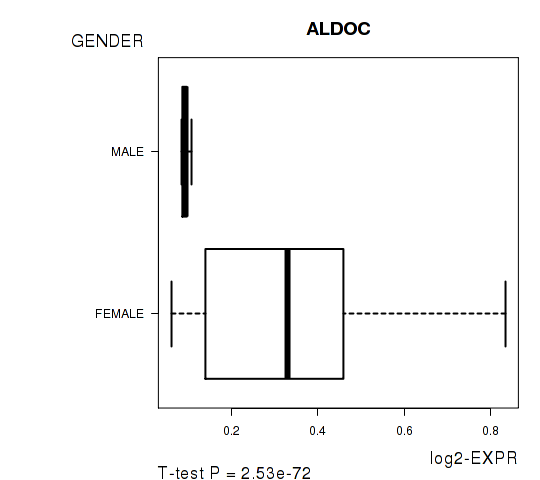
Table S13. Basic characteristics of clinical feature: 'HISTOLOGICAL.TYPE'
| HISTOLOGICAL.TYPE | Labels | N |
| INFILTRATING CARCINOMA NOS | 1 | |
| INFILTRATING DUCTAL CARCINOMA | 456 | |
| INFILTRATING LOBULAR CARCINOMA | 134 | |
| MEDULLARY CARCINOMA | 5 | |
| MIXED HISTOLOGY (PLEASE SPECIFY) | 24 | |
| MUCINOUS CARCINOMA | 12 | |
| OTHER SPECIFY | 32 | |
| Significant markers | N = 1201 |
Table S14. Get Full Table List of top 10 genes differentially expressed by 'HISTOLOGICAL.TYPE'
| ANOVA_P | Q | |
|---|---|---|
| CLUAP1 | 0 | 0 |
| RNF25 | 1.962e-271 | 3.92e-267 |
| STK36 | 1.962e-271 | 3.92e-267 |
| PRKAR2A | 3.391e-213 | 6.78e-209 |
| FAR1 | 5.401e-100 | 1.08e-95 |
| RNF26 | 2.007e-85 | 4.01e-81 |
| AP1M2 | 3.802e-81 | 7.6e-77 |
| CYB5D1__1 | 1.596e-73 | 3.19e-69 |
| LSMD1__1 | 1.596e-73 | 3.19e-69 |
| MPDU1 | 1.946e-58 | 3.89e-54 |
Figure S6. Get High-res Image As an example, this figure shows the association of CLUAP1 to 'HISTOLOGICAL.TYPE'. P value = 0 with ANOVA analysis.
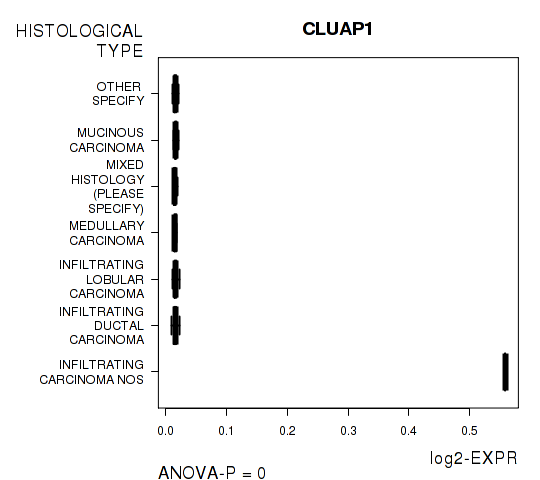
116 genes related to 'RADIATIONS.RADIATION.REGIMENINDICATION'.
Table S15. Basic characteristics of clinical feature: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| RADIATIONS.RADIATION.REGIMENINDICATION | Labels | N |
| NO | 215 | |
| YES | 450 | |
| Significant markers | N = 116 | |
| Higher in YES | 47 | |
| Higher in NO | 69 |
Table S16. Get Full Table List of top 10 genes differentially expressed by 'RADIATIONS.RADIATION.REGIMENINDICATION'
| T(pos if higher in 'YES') | ttestP | Q | AUC | |
|---|---|---|---|---|
| PRIM1 | 7.8 | 2.499e-14 | 4.99e-10 | 0.6547 |
| METAP2 | 7.36 | 6.498e-13 | 1.3e-08 | 0.6625 |
| MAEA | -7.18 | 2.192e-12 | 4.38e-08 | 0.6474 |
| HMG20B | -7.13 | 3.126e-12 | 6.25e-08 | 0.648 |
| ZNF639 | 6.96 | 8.941e-12 | 1.79e-07 | 0.6478 |
| UHRF1BP1L | 6.92 | 1.261e-11 | 2.52e-07 | 0.6578 |
| C16ORF58 | -6.9 | 1.807e-11 | 3.61e-07 | 0.6577 |
| TSTD1 | -6.52 | 1.689e-10 | 3.37e-06 | 0.6318 |
| USF1 | -6.52 | 1.689e-10 | 3.37e-06 | 0.6318 |
| SRRM5 | -6.48 | 2.099e-10 | 4.19e-06 | 0.6352 |
Figure S7. Get High-res Image As an example, this figure shows the association of PRIM1 to 'RADIATIONS.RADIATION.REGIMENINDICATION'. P value = 2.5e-14 with T-test analysis.
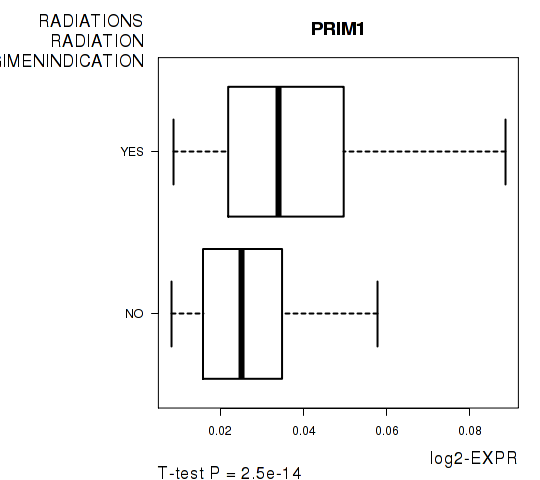
Table S17. Basic characteristics of clinical feature: 'NUMBER.OF.LYMPH.NODES'
| NUMBER.OF.LYMPH.NODES | Mean (SD) | 2.58 (4.8) |
| Significant markers | N = 1 | |
| pos. correlated | 1 | |
| neg. correlated | 0 |
Table S18. Get Full Table List of one gene significantly correlated to 'NUMBER.OF.LYMPH.NODES' by Spearman correlation test
| SpearmanCorr | corrP | Q | |
|---|---|---|---|
| TCP11L1 | 0.2267 | 1.392e-08 | 0.000278 |
Figure S8. Get High-res Image As an example, this figure shows the association of TCP11L1 to 'NUMBER.OF.LYMPH.NODES'. P value = 1.39e-08 with Spearman correlation analysis. The straight line presents the best linear regression.
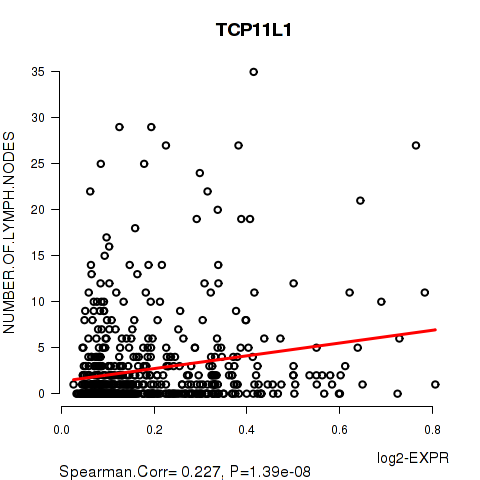
-
Expresson data file = BRCA-TP.meth.by_min_clin_corr.data.txt
-
Clinical data file = BRCA-TP.merged_data.txt
-
Number of patients = 665
-
Number of genes = 19985
-
Number of clinical features = 10
For survival clinical features, Wald's test in univariate Cox regression analysis with proportional hazards model (Andersen and Gill 1982) was used to estimate the P values using the 'coxph' function in R. Kaplan-Meier survival curves were plot using the four quartile subgroups of patients based on expression levels
For continuous numerical clinical features, Spearman's rank correlation coefficients (Spearman 1904) and two-tailed P values were estimated using 'cor.test' function in R
For multi-class clinical features (ordinal or nominal), one-way analysis of variance (Howell 2002) was applied to compare the log2-expression levels between different clinical classes using 'anova' function in R
For two-class clinical features, two-tailed Student's t test with unequal variance (Lehmann and Romano 2005) was applied to compare the log2-expression levels between the two clinical classes using 't.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.