This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 55 focal events and 8 molecular subtypes across 192 patients, 33 significant findings detected with P value < 0.05 and Q value < 0.25.
-
amp_2q33.1 cnv correlated to 'MRNASEQ_CNMF'.
-
amp_3q26.31 cnv correlated to 'CN_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_3q28 cnv correlated to 'CN_CNMF' and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_5p15.33 cnv correlated to 'CN_CNMF'.
-
amp_7p11.2 cnv correlated to 'CN_CNMF'.
-
amp_11q22.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
amp_17q25.1 cnv correlated to 'CN_CNMF'.
-
amp_19q13.2 cnv correlated to 'CN_CNMF'.
-
amp_20q11.21 cnv correlated to 'CN_CNMF'.
-
del_1p36.31 cnv correlated to 'MIRSEQ_CHIERARCHICAL'.
-
del_3p14.2 cnv correlated to 'CN_CNMF'.
-
del_4q21.3 cnv correlated to 'CN_CNMF'.
-
del_4q35.2 cnv correlated to 'CN_CNMF'.
-
del_5q13.3 cnv correlated to 'CN_CNMF'.
-
del_5q35.2 cnv correlated to 'CN_CNMF'.
-
del_11q25 cnv correlated to 'CN_CNMF'.
-
del_14q32.31 cnv correlated to 'MRNASEQ_CHIERARCHICAL' and 'MIRSEQ_MATURE_CNMF'.
-
del_17q25.3 cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MIRSEQ_CNMF', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_18q21.2 cnv correlated to 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
del_19p13.3 cnv correlated to 'MIRSEQ_CHIERARCHICAL'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 55 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 33 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Chi-square test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| del 17q25 3 | 32 (17%) | 160 |
0.395 (1.00) |
0.000214 (0.0896) |
1.93e-05 (0.0083) |
0.00265 (1.00) |
1.51e-05 (0.00656) |
0.0012 (0.472) |
2.93e-05 (0.0126) |
0.000237 (0.0987) |
| amp 11q22 1 | 33 (17%) | 159 |
5.94e-05 (0.0252) |
1.69e-05 (0.0073) |
0.000284 (0.117) |
0.000103 (0.0435) |
0.0165 (1.00) |
0.0166 (1.00) |
0.0132 (1.00) |
0.00797 (1.00) |
| amp 3q26 31 | 148 (77%) | 44 |
2.63e-09 (1.16e-06) |
0.000813 (0.329) |
0.017 (1.00) |
0.000584 (0.238) |
0.0355 (1.00) |
0.00464 (1.00) |
0.0515 (1.00) |
0.000225 (0.0938) |
| del 18q21 2 | 53 (28%) | 139 |
0.0158 (1.00) |
0.00141 (0.55) |
9.61e-05 (0.0407) |
0.000583 (0.238) |
0.0637 (1.00) |
0.000273 (0.113) |
0.00136 (0.533) |
0.00966 (1.00) |
| amp 3q28 | 148 (77%) | 44 |
2.63e-09 (1.16e-06) |
0.00305 (1.00) |
0.0446 (1.00) |
0.00084 (0.339) |
0.0225 (1.00) |
0.00251 (0.955) |
0.0388 (1.00) |
0.000133 (0.0562) |
| del 14q32 31 | 31 (16%) | 161 |
0.00574 (1.00) |
0.262 (1.00) |
0.00893 (1.00) |
0.000272 (0.112) |
0.0381 (1.00) |
0.0369 (1.00) |
0.00054 (0.221) |
0.0361 (1.00) |
| amp 2q33 1 | 30 (16%) | 162 |
0.00326 (1.00) |
0.0597 (1.00) |
0.000145 (0.0607) |
0.0584 (1.00) |
0.254 (1.00) |
0.00822 (1.00) |
0.0965 (1.00) |
0.155 (1.00) |
| amp 5p15 33 | 80 (42%) | 112 |
1.41e-07 (6.14e-05) |
0.0257 (1.00) |
0.114 (1.00) |
0.0164 (1.00) |
0.0164 (1.00) |
0.011 (1.00) |
0.0255 (1.00) |
0.0526 (1.00) |
| amp 7p11 2 | 34 (18%) | 158 |
3.82e-05 (0.0163) |
0.0206 (1.00) |
0.0256 (1.00) |
0.0492 (1.00) |
0.00893 (1.00) |
0.0157 (1.00) |
0.00747 (1.00) |
0.0398 (1.00) |
| amp 17q25 1 | 53 (28%) | 139 |
0.000173 (0.0725) |
0.0193 (1.00) |
0.0984 (1.00) |
0.177 (1.00) |
0.0511 (1.00) |
0.602 (1.00) |
0.0285 (1.00) |
0.65 (1.00) |
| amp 19q13 2 | 59 (31%) | 133 |
7.28e-07 (0.000316) |
0.0657 (1.00) |
0.402 (1.00) |
0.528 (1.00) |
0.447 (1.00) |
0.435 (1.00) |
0.195 (1.00) |
0.74 (1.00) |
| amp 20q11 21 | 87 (45%) | 105 |
0.00026 (0.108) |
0.15 (1.00) |
0.156 (1.00) |
0.242 (1.00) |
0.888 (1.00) |
0.319 (1.00) |
0.441 (1.00) |
0.273 (1.00) |
| del 1p36 31 | 26 (14%) | 166 |
0.0314 (1.00) |
0.106 (1.00) |
0.0425 (1.00) |
0.00109 (0.431) |
0.0552 (1.00) |
8.18e-05 (0.0347) |
0.0127 (1.00) |
0.00166 (0.641) |
| del 3p14 2 | 78 (41%) | 114 |
1.1e-05 (0.00479) |
0.000901 (0.36) |
0.02 (1.00) |
0.00211 (0.814) |
0.00526 (1.00) |
0.00643 (1.00) |
0.0115 (1.00) |
0.000854 (0.343) |
| del 4q21 3 | 58 (30%) | 134 |
7.73e-08 (3.38e-05) |
0.0115 (1.00) |
0.581 (1.00) |
0.00383 (1.00) |
0.317 (1.00) |
0.652 (1.00) |
0.762 (1.00) |
0.436 (1.00) |
| del 4q35 2 | 70 (36%) | 122 |
3.9e-08 (1.71e-05) |
0.0763 (1.00) |
0.0946 (1.00) |
0.00174 (0.673) |
0.353 (1.00) |
0.448 (1.00) |
0.97 (1.00) |
0.681 (1.00) |
| del 5q13 3 | 47 (24%) | 145 |
2.42e-05 (0.0104) |
0.0376 (1.00) |
0.324 (1.00) |
0.38 (1.00) |
0.0524 (1.00) |
0.476 (1.00) |
0.0897 (1.00) |
0.392 (1.00) |
| del 5q35 2 | 49 (26%) | 143 |
1.88e-05 (0.0081) |
0.0632 (1.00) |
0.654 (1.00) |
0.879 (1.00) |
0.0426 (1.00) |
0.289 (1.00) |
0.0829 (1.00) |
0.36 (1.00) |
| del 11q25 | 107 (56%) | 85 |
4.37e-05 (0.0186) |
0.18 (1.00) |
0.323 (1.00) |
0.0289 (1.00) |
0.0876 (1.00) |
0.0327 (1.00) |
0.0395 (1.00) |
0.00794 (1.00) |
| del 19p13 3 | 58 (30%) | 134 |
0.00412 (1.00) |
0.000646 (0.263) |
0.00286 (1.00) |
0.00651 (1.00) |
0.433 (1.00) |
0.000519 (0.213) |
0.25 (1.00) |
0.0194 (1.00) |
| amp 1p31 3 | 62 (32%) | 130 |
0.683 (1.00) |
0.197 (1.00) |
0.138 (1.00) |
0.0984 (1.00) |
0.971 (1.00) |
0.156 (1.00) |
0.943 (1.00) |
0.492 (1.00) |
| amp 1q21 3 | 98 (51%) | 94 |
0.00569 (1.00) |
0.309 (1.00) |
0.367 (1.00) |
0.276 (1.00) |
0.782 (1.00) |
0.475 (1.00) |
0.857 (1.00) |
0.346 (1.00) |
| amp 2p24 3 | 50 (26%) | 142 |
0.0183 (1.00) |
0.0173 (1.00) |
0.41 (1.00) |
0.0475 (1.00) |
0.17 (1.00) |
0.000899 (0.36) |
0.33 (1.00) |
0.013 (1.00) |
| amp 6p21 33 | 51 (27%) | 141 |
0.0502 (1.00) |
0.00436 (1.00) |
0.0503 (1.00) |
0.0117 (1.00) |
0.215 (1.00) |
0.0505 (1.00) |
0.416 (1.00) |
0.554 (1.00) |
| amp 8q24 21 | 79 (41%) | 113 |
0.254 (1.00) |
0.694 (1.00) |
0.868 (1.00) |
0.961 (1.00) |
0.0131 (1.00) |
0.302 (1.00) |
0.101 (1.00) |
0.239 (1.00) |
| amp 9p24 1 | 41 (21%) | 151 |
0.225 (1.00) |
0.0578 (1.00) |
0.647 (1.00) |
0.00884 (1.00) |
0.445 (1.00) |
0.185 (1.00) |
0.828 (1.00) |
0.182 (1.00) |
| amp 11p13 | 16 (8%) | 176 |
0.241 (1.00) |
0.587 (1.00) |
0.347 (1.00) |
0.92 (1.00) |
0.578 (1.00) |
0.463 (1.00) |
0.773 (1.00) |
0.477 (1.00) |
| amp 11q13 3 | 24 (12%) | 168 |
0.00314 (1.00) |
0.00403 (1.00) |
0.00728 (1.00) |
0.00715 (1.00) |
0.0178 (1.00) |
0.00536 (1.00) |
0.0162 (1.00) |
0.0248 (1.00) |
| amp 13q22 1 | 41 (21%) | 151 |
0.011 (1.00) |
0.14 (1.00) |
0.148 (1.00) |
0.934 (1.00) |
0.674 (1.00) |
0.524 (1.00) |
0.687 (1.00) |
0.306 (1.00) |
| amp 13q34 | 43 (22%) | 149 |
0.00295 (1.00) |
0.807 (1.00) |
0.934 (1.00) |
0.934 (1.00) |
0.724 (1.00) |
0.154 (1.00) |
0.61 (1.00) |
0.0815 (1.00) |
| amp 15q26 1 | 46 (24%) | 146 |
0.964 (1.00) |
0.433 (1.00) |
0.761 (1.00) |
0.391 (1.00) |
0.286 (1.00) |
1 (1.00) |
0.189 (1.00) |
0.787 (1.00) |
| amp 16p13 13 | 39 (20%) | 153 |
0.193 (1.00) |
0.481 (1.00) |
0.358 (1.00) |
0.314 (1.00) |
0.257 (1.00) |
0.427 (1.00) |
0.492 (1.00) |
0.31 (1.00) |
| amp 17q12 | 36 (19%) | 156 |
0.0317 (1.00) |
0.0459 (1.00) |
0.157 (1.00) |
0.136 (1.00) |
0.175 (1.00) |
0.0779 (1.00) |
0.163 (1.00) |
0.178 (1.00) |
| amp xq28 | 47 (24%) | 145 |
0.782 (1.00) |
0.992 (1.00) |
0.753 (1.00) |
0.581 (1.00) |
0.00479 (1.00) |
0.0368 (1.00) |
0.0308 (1.00) |
0.0812 (1.00) |
| del 2q22 1 | 37 (19%) | 155 |
0.792 (1.00) |
0.915 (1.00) |
0.383 (1.00) |
0.279 (1.00) |
0.73 (1.00) |
0.761 (1.00) |
0.862 (1.00) |
0.731 (1.00) |
| del 2q37 1 | 81 (42%) | 111 |
0.172 (1.00) |
0.00824 (1.00) |
0.00833 (1.00) |
0.000736 (0.299) |
0.0183 (1.00) |
0.00713 (1.00) |
0.0144 (1.00) |
0.00291 (1.00) |
| del 6p24 2 | 29 (15%) | 163 |
0.556 (1.00) |
0.0932 (1.00) |
0.244 (1.00) |
0.523 (1.00) |
0.196 (1.00) |
0.833 (1.00) |
0.865 (1.00) |
0.556 (1.00) |
| del 6q26 | 65 (34%) | 127 |
0.00982 (1.00) |
0.419 (1.00) |
0.746 (1.00) |
0.639 (1.00) |
0.602 (1.00) |
0.227 (1.00) |
0.705 (1.00) |
0.225 (1.00) |
| del 7q34 | 43 (22%) | 149 |
0.00505 (1.00) |
0.965 (1.00) |
0.683 (1.00) |
0.696 (1.00) |
0.453 (1.00) |
0.528 (1.00) |
0.603 (1.00) |
0.676 (1.00) |
| del 8p23 3 | 66 (34%) | 126 |
0.00344 (1.00) |
0.136 (1.00) |
0.0915 (1.00) |
0.851 (1.00) |
0.831 (1.00) |
0.157 (1.00) |
0.829 (1.00) |
0.273 (1.00) |
| del 10q23 31 | 55 (29%) | 137 |
0.0314 (1.00) |
0.197 (1.00) |
0.242 (1.00) |
0.65 (1.00) |
0.669 (1.00) |
1 (1.00) |
0.973 (1.00) |
1 (1.00) |
| del 11p15 1 | 62 (32%) | 130 |
0.0146 (1.00) |
0.046 (1.00) |
0.0596 (1.00) |
0.0372 (1.00) |
0.187 (1.00) |
0.0821 (1.00) |
0.168 (1.00) |
0.0615 (1.00) |
| del 13q12 12 | 68 (35%) | 124 |
0.000866 (0.347) |
0.0143 (1.00) |
0.167 (1.00) |
0.00308 (1.00) |
0.0108 (1.00) |
0.021 (1.00) |
0.0106 (1.00) |
0.0148 (1.00) |
| del 13q14 2 | 74 (39%) | 118 |
0.00231 (0.884) |
0.00853 (1.00) |
0.0679 (1.00) |
0.00147 (0.573) |
0.00354 (1.00) |
0.00276 (1.00) |
0.00286 (1.00) |
0.0067 (1.00) |
| del 15q21 1 | 38 (20%) | 154 |
0.0185 (1.00) |
0.0526 (1.00) |
0.414 (1.00) |
0.213 (1.00) |
0.27 (1.00) |
0.0232 (1.00) |
0.124 (1.00) |
0.113 (1.00) |
| del 16q12 1 | 39 (20%) | 153 |
0.575 (1.00) |
0.0175 (1.00) |
0.173 (1.00) |
0.347 (1.00) |
0.146 (1.00) |
0.0787 (1.00) |
0.0102 (1.00) |
0.154 (1.00) |
| del 16q23 1 | 39 (20%) | 153 |
0.511 (1.00) |
0.000931 (0.37) |
0.00398 (1.00) |
0.00388 (1.00) |
0.055 (1.00) |
0.00161 (0.623) |
0.00106 (0.421) |
0.00474 (1.00) |
| del 17p12 | 61 (32%) | 131 |
0.00332 (1.00) |
0.442 (1.00) |
0.217 (1.00) |
0.11 (1.00) |
0.0357 (1.00) |
0.375 (1.00) |
0.0637 (1.00) |
0.0959 (1.00) |
| del 19q13 33 | 22 (11%) | 170 |
0.244 (1.00) |
0.241 (1.00) |
0.102 (1.00) |
0.487 (1.00) |
0.467 (1.00) |
0.109 (1.00) |
0.488 (1.00) |
0.171 (1.00) |
| del 20p12 1 | 25 (13%) | 167 |
0.802 (1.00) |
0.446 (1.00) |
0.271 (1.00) |
0.75 (1.00) |
0.334 (1.00) |
0.16 (1.00) |
0.236 (1.00) |
0.732 (1.00) |
| del 21q21 1 | 35 (18%) | 157 |
0.0383 (1.00) |
0.387 (1.00) |
0.848 (1.00) |
0.00237 (0.904) |
0.756 (1.00) |
0.671 (1.00) |
0.929 (1.00) |
0.744 (1.00) |
| del 22q13 32 | 52 (27%) | 140 |
0.166 (1.00) |
0.13 (1.00) |
0.257 (1.00) |
0.385 (1.00) |
0.886 (1.00) |
0.203 (1.00) |
0.54 (1.00) |
0.365 (1.00) |
| del xp21 1 | 42 (22%) | 150 |
0.00104 (0.414) |
0.0907 (1.00) |
0.131 (1.00) |
0.286 (1.00) |
0.0124 (1.00) |
0.0171 (1.00) |
0.0104 (1.00) |
0.00277 (1.00) |
| del xp11 3 | 46 (24%) | 146 |
0.00141 (0.55) |
0.0688 (1.00) |
0.0865 (1.00) |
0.139 (1.00) |
0.00106 (0.42) |
0.0103 (1.00) |
0.000824 (0.333) |
0.00214 (0.823) |
| del xq21 33 | 39 (20%) | 153 |
0.063 (1.00) |
0.0253 (1.00) |
0.303 (1.00) |
0.0258 (1.00) |
0.0733 (1.00) |
0.0129 (1.00) |
0.15 (1.00) |
0.00715 (1.00) |
P value = 0.000145 (Fisher's exact test), Q value = 0.061
Table S1. Gene #4: 'amp_2q33.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 64 | 34 | 45 | 37 |
| AMP PEAK 4(2Q33.1) MUTATED | 1 | 11 | 7 | 7 |
| AMP PEAK 4(2Q33.1) WILD-TYPE | 63 | 23 | 38 | 30 |
Figure S1. Get High-res Image Gene #4: 'amp_2q33.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 2.63e-09 (Fisher's exact test), Q value = 1.2e-06
Table S2. Gene #5: 'amp_3q26.31' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| AMP PEAK 5(3Q26.31) MUTATED | 60 | 33 | 55 |
| AMP PEAK 5(3Q26.31) WILD-TYPE | 0 | 9 | 35 |
Figure S2. Get High-res Image Gene #5: 'amp_3q26.31' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000584 (Fisher's exact test), Q value = 0.24
Table S3. Gene #5: 'amp_3q26.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 19 | 112 | 49 |
| AMP PEAK 5(3Q26.31) MUTATED | 10 | 96 | 32 |
| AMP PEAK 5(3Q26.31) WILD-TYPE | 9 | 16 | 17 |
Figure S3. Get High-res Image Gene #5: 'amp_3q26.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.000225 (Fisher's exact test), Q value = 0.094
Table S4. Gene #5: 'amp_3q26.31' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 70 | 55 |
| AMP PEAK 5(3Q26.31) MUTATED | 62 | 51 | 35 |
| AMP PEAK 5(3Q26.31) WILD-TYPE | 5 | 19 | 20 |
Figure S4. Get High-res Image Gene #5: 'amp_3q26.31' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 2.63e-09 (Fisher's exact test), Q value = 1.2e-06
Table S5. Gene #6: 'amp_3q28' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| AMP PEAK 6(3Q28) MUTATED | 60 | 33 | 55 |
| AMP PEAK 6(3Q28) WILD-TYPE | 0 | 9 | 35 |
Figure S5. Get High-res Image Gene #6: 'amp_3q28' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000133 (Fisher's exact test), Q value = 0.056
Table S6. Gene #6: 'amp_3q28' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 70 | 55 |
| AMP PEAK 6(3Q28) MUTATED | 62 | 52 | 34 |
| AMP PEAK 6(3Q28) WILD-TYPE | 5 | 18 | 21 |
Figure S6. Get High-res Image Gene #6: 'amp_3q28' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

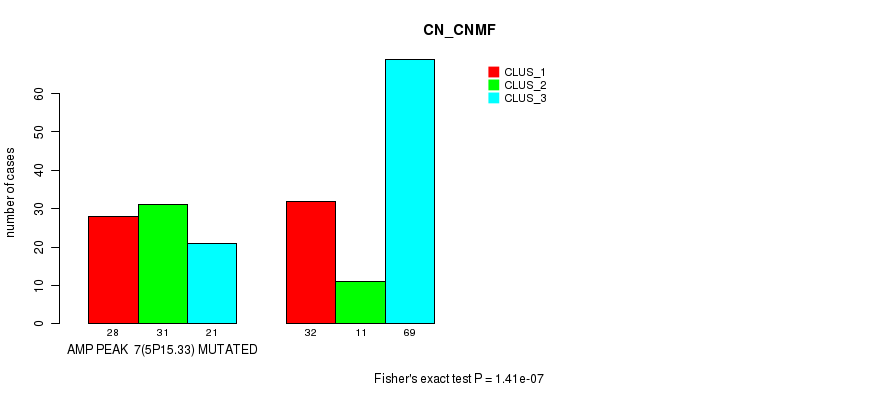
P value = 1.41e-07 (Fisher's exact test), Q value = 6.1e-05
Table S7. Gene #7: 'amp_5p15.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| AMP PEAK 7(5P15.33) MUTATED | 28 | 31 | 21 |
| AMP PEAK 7(5P15.33) WILD-TYPE | 32 | 11 | 69 |
Figure S7. Get High-res Image Gene #7: 'amp_5p15.33' versus Molecular Subtype #1: 'CN_CNMF'
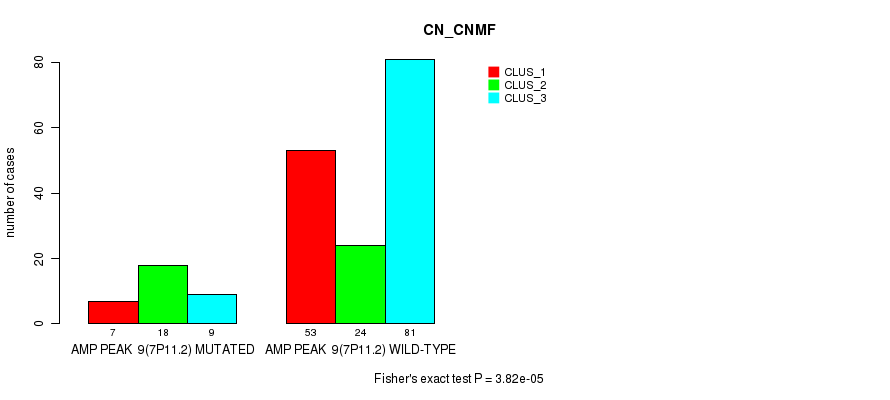
P value = 3.82e-05 (Fisher's exact test), Q value = 0.016
Table S8. Gene #9: 'amp_7p11.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| AMP PEAK 9(7P11.2) MUTATED | 7 | 18 | 9 |
| AMP PEAK 9(7P11.2) WILD-TYPE | 53 | 24 | 81 |
Figure S8. Get High-res Image Gene #9: 'amp_7p11.2' versus Molecular Subtype #1: 'CN_CNMF'
P value = 5.94e-05 (Fisher's exact test), Q value = 0.025
Table S9. Gene #14: 'amp_11q22.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| AMP PEAK 14(11Q22.1) MUTATED | 4 | 17 | 12 |
| AMP PEAK 14(11Q22.1) WILD-TYPE | 56 | 25 | 78 |
Figure S9. Get High-res Image Gene #14: 'amp_11q22.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.69e-05 (Chi-square test), Q value = 0.0073
Table S10. Gene #14: 'amp_11q22.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 32 | 27 | 21 | 32 | 31 | 34 | 15 |
| AMP PEAK 14(11Q22.1) MUTATED | 8 | 0 | 2 | 15 | 2 | 3 | 3 |
| AMP PEAK 14(11Q22.1) WILD-TYPE | 24 | 27 | 19 | 17 | 29 | 31 | 12 |
Figure S10. Get High-res Image Gene #14: 'amp_11q22.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'

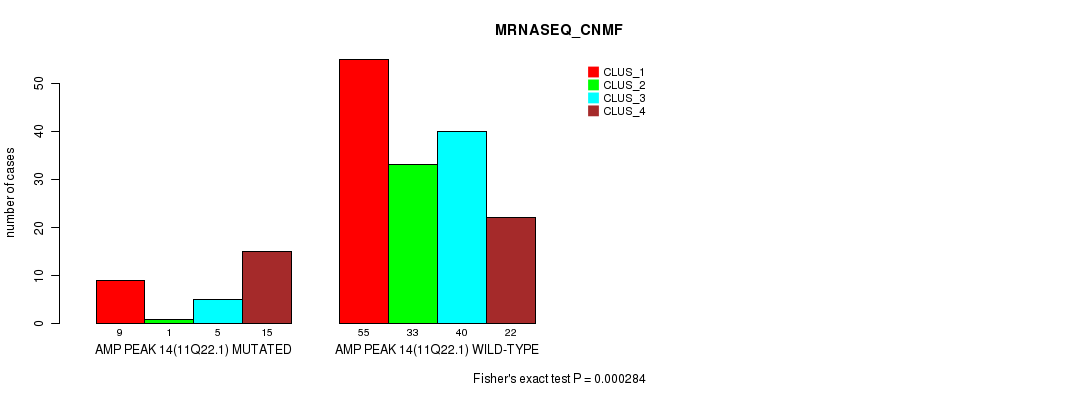
P value = 0.000284 (Fisher's exact test), Q value = 0.12
Table S11. Gene #14: 'amp_11q22.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 64 | 34 | 45 | 37 |
| AMP PEAK 14(11Q22.1) MUTATED | 9 | 1 | 5 | 15 |
| AMP PEAK 14(11Q22.1) WILD-TYPE | 55 | 33 | 40 | 22 |
Figure S11. Get High-res Image Gene #14: 'amp_11q22.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
P value = 0.000103 (Fisher's exact test), Q value = 0.044
Table S12. Gene #14: 'amp_11q22.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 19 | 112 | 49 |
| AMP PEAK 14(11Q22.1) MUTATED | 10 | 17 | 3 |
| AMP PEAK 14(11Q22.1) WILD-TYPE | 9 | 95 | 46 |
Figure S12. Get High-res Image Gene #14: 'amp_11q22.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

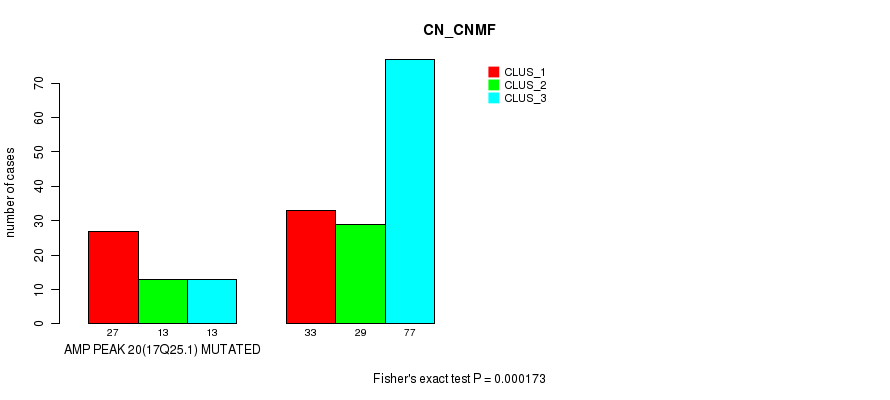
P value = 0.000173 (Fisher's exact test), Q value = 0.072
Table S13. Gene #20: 'amp_17q25.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| AMP PEAK 20(17Q25.1) MUTATED | 27 | 13 | 13 |
| AMP PEAK 20(17Q25.1) WILD-TYPE | 33 | 29 | 77 |
Figure S13. Get High-res Image Gene #20: 'amp_17q25.1' versus Molecular Subtype #1: 'CN_CNMF'
P value = 7.28e-07 (Fisher's exact test), Q value = 0.00032
Table S14. Gene #21: 'amp_19q13.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| AMP PEAK 21(19Q13.2) MUTATED | 16 | 27 | 16 |
| AMP PEAK 21(19Q13.2) WILD-TYPE | 44 | 15 | 74 |
Figure S14. Get High-res Image Gene #21: 'amp_19q13.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00026 (Fisher's exact test), Q value = 0.11
Table S15. Gene #22: 'amp_20q11.21' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| AMP PEAK 22(20Q11.21) MUTATED | 34 | 26 | 27 |
| AMP PEAK 22(20Q11.21) WILD-TYPE | 26 | 16 | 63 |
Figure S15. Get High-res Image Gene #22: 'amp_20q11.21' versus Molecular Subtype #1: 'CN_CNMF'

P value = 8.18e-05 (Fisher's exact test), Q value = 0.035
Table S16. Gene #24: 'del_1p36.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 71 | 61 | 60 |
| DEL PEAK 1(1P36.31) MUTATED | 1 | 10 | 15 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 70 | 51 | 45 |
Figure S16. Get High-res Image Gene #24: 'del_1p36.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 1.1e-05 (Fisher's exact test), Q value = 0.0048
Table S17. Gene #27: 'del_3p14.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| DEL PEAK 4(3P14.2) MUTATED | 26 | 29 | 23 |
| DEL PEAK 4(3P14.2) WILD-TYPE | 34 | 13 | 67 |
Figure S17. Get High-res Image Gene #27: 'del_3p14.2' versus Molecular Subtype #1: 'CN_CNMF'
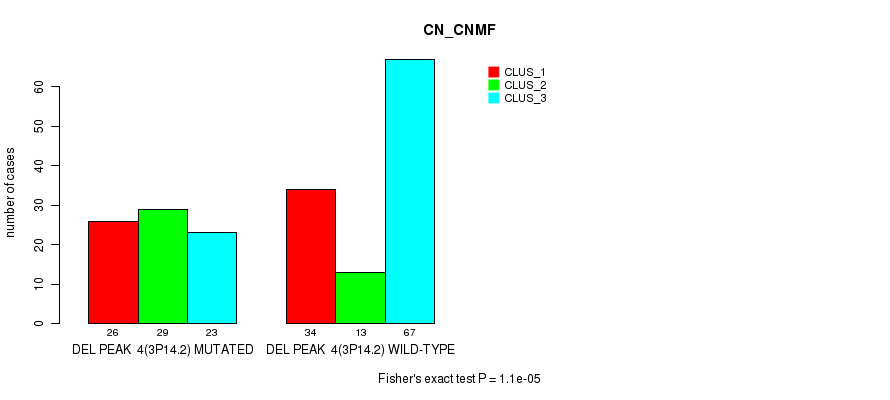
P value = 7.73e-08 (Fisher's exact test), Q value = 3.4e-05
Table S18. Gene #28: 'del_4q21.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| DEL PEAK 5(4Q21.3) MUTATED | 18 | 27 | 13 |
| DEL PEAK 5(4Q21.3) WILD-TYPE | 42 | 15 | 77 |
Figure S18. Get High-res Image Gene #28: 'del_4q21.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 3.9e-08 (Fisher's exact test), Q value = 1.7e-05
Table S19. Gene #29: 'del_4q35.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| DEL PEAK 6(4Q35.2) MUTATED | 20 | 31 | 19 |
| DEL PEAK 6(4Q35.2) WILD-TYPE | 40 | 11 | 71 |
Figure S19. Get High-res Image Gene #29: 'del_4q35.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 2.42e-05 (Fisher's exact test), Q value = 0.01
Table S20. Gene #30: 'del_5q13.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| DEL PEAK 7(5Q13.3) MUTATED | 15 | 21 | 11 |
| DEL PEAK 7(5Q13.3) WILD-TYPE | 45 | 21 | 79 |
Figure S20. Get High-res Image Gene #30: 'del_5q13.3' versus Molecular Subtype #1: 'CN_CNMF'
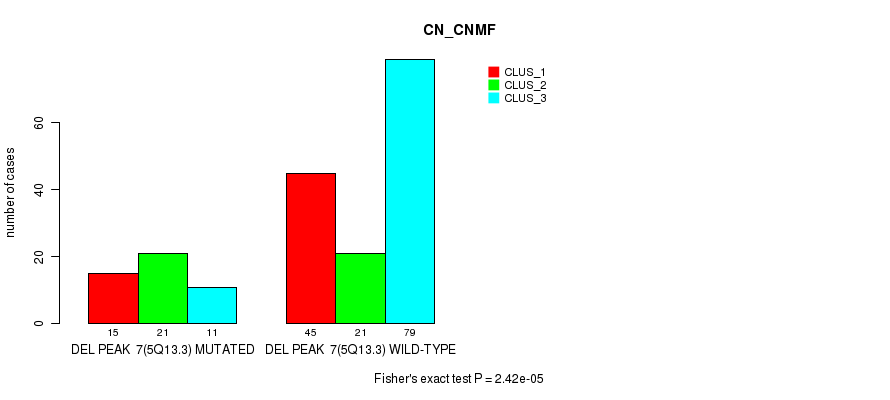
P value = 1.88e-05 (Fisher's exact test), Q value = 0.0081
Table S21. Gene #31: 'del_5q35.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| DEL PEAK 8(5Q35.2) MUTATED | 15 | 22 | 12 |
| DEL PEAK 8(5Q35.2) WILD-TYPE | 45 | 20 | 78 |
Figure S21. Get High-res Image Gene #31: 'del_5q35.2' versus Molecular Subtype #1: 'CN_CNMF'
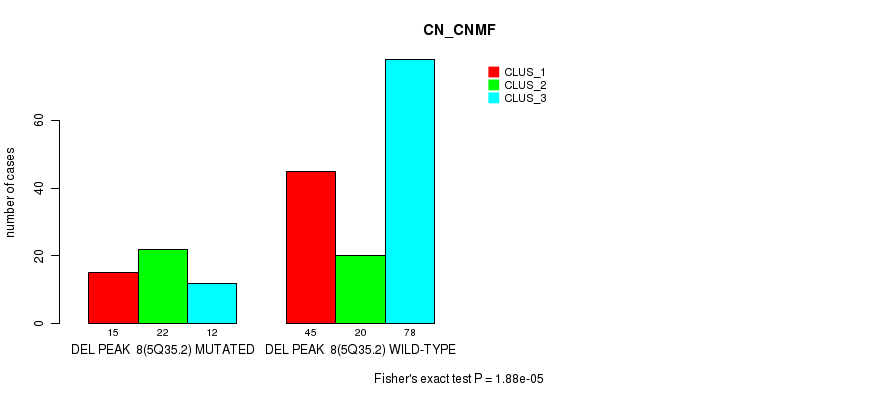
P value = 4.37e-05 (Fisher's exact test), Q value = 0.019
Table S22. Gene #38: 'del_11q25' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 60 | 42 | 90 |
| DEL PEAK 15(11Q25) MUTATED | 44 | 28 | 35 |
| DEL PEAK 15(11Q25) WILD-TYPE | 16 | 14 | 55 |
Figure S22. Get High-res Image Gene #38: 'del_11q25' versus Molecular Subtype #1: 'CN_CNMF'
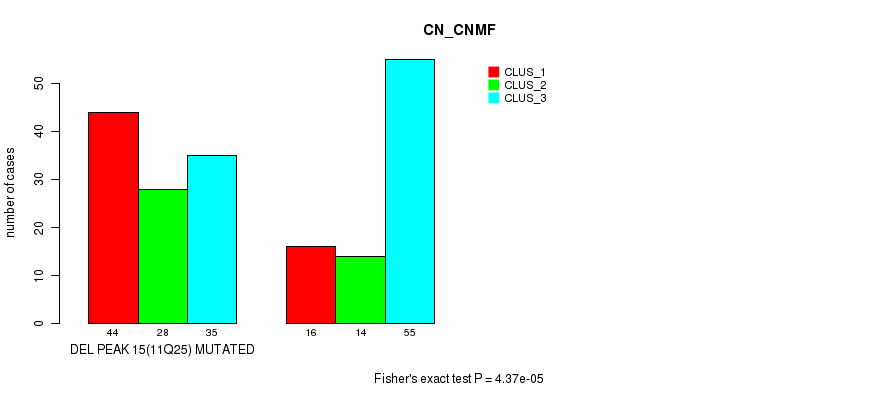
P value = 0.000272 (Fisher's exact test), Q value = 0.11
Table S23. Gene #41: 'del_14q32.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 19 | 112 | 49 |
| DEL PEAK 18(14Q32.31) MUTATED | 8 | 10 | 13 |
| DEL PEAK 18(14Q32.31) WILD-TYPE | 11 | 102 | 36 |
Figure S23. Get High-res Image Gene #41: 'del_14q32.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00054 (Fisher's exact test), Q value = 0.22
Table S24. Gene #41: 'del_14q32.31' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 72 | 52 | 68 |
| DEL PEAK 18(14Q32.31) MUTATED | 4 | 7 | 20 |
| DEL PEAK 18(14Q32.31) WILD-TYPE | 68 | 45 | 48 |
Figure S24. Get High-res Image Gene #41: 'del_14q32.31' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.000214 (Chi-square test), Q value = 0.09
Table S25. Gene #46: 'del_17q25.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 32 | 27 | 21 | 32 | 31 | 34 | 15 |
| DEL PEAK 23(17Q25.3) MUTATED | 5 | 2 | 2 | 10 | 12 | 0 | 1 |
| DEL PEAK 23(17Q25.3) WILD-TYPE | 27 | 25 | 19 | 22 | 19 | 34 | 14 |
Figure S25. Get High-res Image Gene #46: 'del_17q25.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1.93e-05 (Fisher's exact test), Q value = 0.0083
Table S26. Gene #46: 'del_17q25.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 64 | 34 | 45 | 37 |
| DEL PEAK 23(17Q25.3) MUTATED | 12 | 1 | 1 | 14 |
| DEL PEAK 23(17Q25.3) WILD-TYPE | 52 | 33 | 44 | 23 |
Figure S26. Get High-res Image Gene #46: 'del_17q25.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 1.51e-05 (Fisher's exact test), Q value = 0.0066
Table S27. Gene #46: 'del_17q25.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 73 | 52 | 67 |
| DEL PEAK 23(17Q25.3) MUTATED | 23 | 7 | 2 |
| DEL PEAK 23(17Q25.3) WILD-TYPE | 50 | 45 | 65 |
Figure S27. Get High-res Image Gene #46: 'del_17q25.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 2.93e-05 (Fisher's exact test), Q value = 0.013
Table S28. Gene #46: 'del_17q25.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 72 | 52 | 68 |
| DEL PEAK 23(17Q25.3) MUTATED | 22 | 8 | 2 |
| DEL PEAK 23(17Q25.3) WILD-TYPE | 50 | 44 | 66 |
Figure S28. Get High-res Image Gene #46: 'del_17q25.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.000237 (Fisher's exact test), Q value = 0.099
Table S29. Gene #46: 'del_17q25.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 70 | 55 |
| DEL PEAK 23(17Q25.3) MUTATED | 9 | 21 | 2 |
| DEL PEAK 23(17Q25.3) WILD-TYPE | 58 | 49 | 53 |
Figure S29. Get High-res Image Gene #46: 'del_17q25.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 9.61e-05 (Fisher's exact test), Q value = 0.041
Table S30. Gene #47: 'del_18q21.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 64 | 34 | 45 | 37 |
| DEL PEAK 24(18Q21.2) MUTATED | 7 | 7 | 20 | 16 |
| DEL PEAK 24(18Q21.2) WILD-TYPE | 57 | 27 | 25 | 21 |
Figure S30. Get High-res Image Gene #47: 'del_18q21.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.000583 (Fisher's exact test), Q value = 0.24
Table S31. Gene #47: 'del_18q21.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 19 | 112 | 49 |
| DEL PEAK 24(18Q21.2) MUTATED | 9 | 20 | 21 |
| DEL PEAK 24(18Q21.2) WILD-TYPE | 10 | 92 | 28 |
Figure S31. Get High-res Image Gene #47: 'del_18q21.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

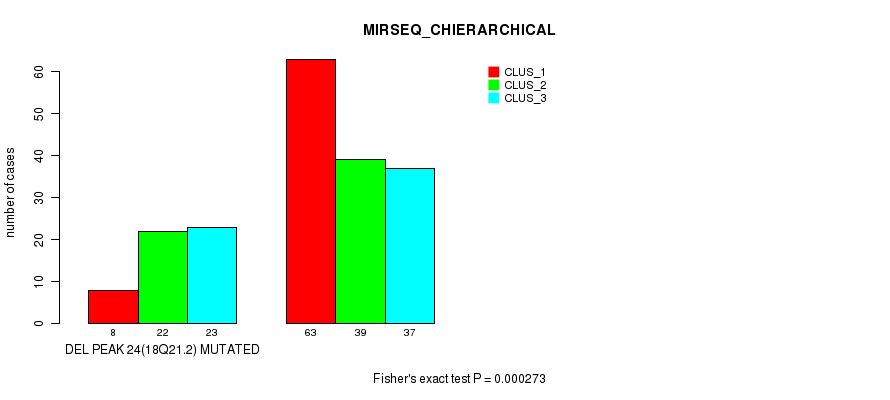
P value = 0.000273 (Fisher's exact test), Q value = 0.11
Table S32. Gene #47: 'del_18q21.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 71 | 61 | 60 |
| DEL PEAK 24(18Q21.2) MUTATED | 8 | 22 | 23 |
| DEL PEAK 24(18Q21.2) WILD-TYPE | 63 | 39 | 37 |
Figure S32. Get High-res Image Gene #47: 'del_18q21.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
P value = 0.000519 (Fisher's exact test), Q value = 0.21
Table S33. Gene #48: 'del_19p13.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 71 | 61 | 60 |
| DEL PEAK 25(19P13.3) MUTATED | 10 | 26 | 22 |
| DEL PEAK 25(19P13.3) WILD-TYPE | 61 | 35 | 38 |
Figure S33. Get High-res Image Gene #48: 'del_19p13.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtype file = CESC-TP.transferedmergedcluster.txt
-
Number of patients = 192
-
Number of significantly focal cnvs = 55
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.