This pipeline computes the correlation between significant arm-level copy number variations (cnvs) and molecular subtypes.
Testing the association between copy number variation 61 arm-level events and 8 molecular subtypes across 66 patients, 54 significant findings detected with P value < 0.05 and Q value < 0.25.
-
4p gain cnv correlated to 'MRNASEQ_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
4q gain cnv correlated to 'MRNASEQ_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
11p gain cnv correlated to 'MRNASEQ_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
11q gain cnv correlated to 'MRNASEQ_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
16p gain cnv correlated to 'MRNASEQ_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
16q gain cnv correlated to 'MRNASEQ_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
1p loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
1q loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
2p loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
2q loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
6p loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
6q loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
8p loss cnv correlated to 'CN_CNMF'.
-
8q loss cnv correlated to 'CN_CNMF'.
-
10p loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
10q loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
13q loss cnv correlated to 'METHLYATION_CNMF'.
-
17p loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
17q loss cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 61 arm-level events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 54 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Chi-square test | Fisher's exact test | Fisher's exact test | Chi-square test | Fisher's exact test | |
| 2p loss | 46 (70%) | 20 |
0.0875 (1.00) |
3.7e-07 (0.000178) |
8.54e-08 (4.16e-05) |
2.91e-07 (0.000141) |
0.0891 (1.00) |
0.000161 (0.072) |
0.0277 (1.00) |
2.02e-05 (0.00934) |
| 2q loss | 46 (70%) | 20 |
0.0875 (1.00) |
3.7e-07 (0.000178) |
8.54e-08 (4.16e-05) |
2.91e-07 (0.000141) |
0.0891 (1.00) |
0.000161 (0.072) |
0.0277 (1.00) |
2.02e-05 (0.00934) |
| 17p loss | 50 (76%) | 16 |
0.299 (1.00) |
4.75e-07 (0.000228) |
5.74e-07 (0.000273) |
5.55e-07 (0.000266) |
0.0506 (1.00) |
0.000396 (0.173) |
0.05 (1.00) |
0.000108 (0.0486) |
| 17q loss | 50 (76%) | 16 |
0.299 (1.00) |
4.75e-07 (0.000228) |
5.74e-07 (0.000273) |
5.55e-07 (0.000266) |
0.0506 (1.00) |
0.000396 (0.173) |
0.05 (1.00) |
0.000108 (0.0486) |
| 10p loss | 48 (73%) | 18 |
0.342 (1.00) |
0.00038 (0.166) |
8.47e-05 (0.0383) |
5.23e-06 (0.00245) |
0.287 (1.00) |
0.0097 (1.00) |
0.0145 (1.00) |
0.000336 (0.148) |
| 1p loss | 53 (80%) | 13 |
0.671 (1.00) |
2.22e-05 (0.0102) |
0.000362 (0.159) |
5.73e-05 (0.0259) |
0.16 (1.00) |
0.00115 (0.49) |
0.166 (1.00) |
0.00118 (0.5) |
| 1q loss | 52 (79%) | 14 |
0.475 (1.00) |
2.29e-06 (0.00108) |
5.05e-05 (0.0229) |
4.21e-06 (0.00198) |
0.24 (1.00) |
0.00192 (0.801) |
0.133 (1.00) |
0.00184 (0.768) |
| 6p loss | 51 (77%) | 15 |
0.33 (1.00) |
0.000107 (0.0482) |
4.76e-05 (0.0217) |
0.00033 (0.146) |
0.355 (1.00) |
0.00305 (1.00) |
0.306 (1.00) |
0.00201 (0.837) |
| 6q loss | 51 (77%) | 15 |
0.33 (1.00) |
0.000107 (0.0482) |
4.76e-05 (0.0217) |
0.00033 (0.146) |
0.355 (1.00) |
0.00305 (1.00) |
0.306 (1.00) |
0.00201 (0.837) |
| 10q loss | 49 (74%) | 17 |
0.248 (1.00) |
0.000262 (0.116) |
2.53e-05 (0.0116) |
2.88e-06 (0.00136) |
0.243 (1.00) |
0.00681 (1.00) |
0.0296 (1.00) |
0.00108 (0.458) |
| 4p gain | 24 (36%) | 42 |
0.0552 (1.00) |
1 (1.00) |
2.78e-05 (0.0127) |
0.000215 (0.0956) |
0.788 (1.00) |
0.139 (1.00) |
0.734 (1.00) |
0.0309 (1.00) |
| 4q gain | 24 (36%) | 42 |
0.0552 (1.00) |
1 (1.00) |
2.78e-05 (0.0127) |
0.000215 (0.0956) |
0.788 (1.00) |
0.139 (1.00) |
0.734 (1.00) |
0.0309 (1.00) |
| 11p gain | 15 (23%) | 51 |
0.282 (1.00) |
0.679 (1.00) |
2e-05 (0.00928) |
4.44e-06 (0.00208) |
0.835 (1.00) |
0.106 (1.00) |
0.741 (1.00) |
0.071 (1.00) |
| 11q gain | 15 (23%) | 51 |
0.282 (1.00) |
0.679 (1.00) |
6.69e-07 (0.000316) |
1.11e-07 (5.41e-05) |
0.63 (1.00) |
0.106 (1.00) |
0.441 (1.00) |
0.0453 (1.00) |
| 16p gain | 21 (32%) | 45 |
0.106 (1.00) |
1 (1.00) |
6.22e-06 (0.00291) |
9.49e-06 (0.00441) |
0.158 (1.00) |
0.0481 (1.00) |
0.776 (1.00) |
0.0499 (1.00) |
| 16q gain | 21 (32%) | 45 |
0.106 (1.00) |
1 (1.00) |
6.22e-06 (0.00291) |
9.49e-06 (0.00441) |
0.158 (1.00) |
0.0481 (1.00) |
0.776 (1.00) |
0.0499 (1.00) |
| 8p loss | 9 (14%) | 57 |
8.11e-08 (3.96e-05) |
0.0857 (1.00) |
0.0531 (1.00) |
0.0036 (1.00) |
0.148 (1.00) |
0.341 (1.00) |
0.0757 (1.00) |
0.12 (1.00) |
| 8q loss | 8 (12%) | 58 |
5.86e-07 (0.000278) |
0.0437 (1.00) |
0.134 (1.00) |
0.00434 (1.00) |
0.291 (1.00) |
0.586 (1.00) |
0.165 (1.00) |
0.224 (1.00) |
| 13q loss | 43 (65%) | 23 |
0.27 (1.00) |
0.000529 (0.23) |
0.00364 (1.00) |
0.00343 (1.00) |
0.853 (1.00) |
0.0555 (1.00) |
0.93 (1.00) |
0.158 (1.00) |
| 3p gain | 8 (12%) | 58 |
0.27 (1.00) |
0.53 (1.00) |
0.952 (1.00) |
0.614 (1.00) |
0.515 (1.00) |
0.298 (1.00) |
0.0206 (1.00) |
0.823 (1.00) |
| 3q gain | 8 (12%) | 58 |
0.27 (1.00) |
0.53 (1.00) |
0.952 (1.00) |
0.614 (1.00) |
0.515 (1.00) |
0.298 (1.00) |
0.0206 (1.00) |
0.823 (1.00) |
| 5p gain | 8 (12%) | 58 |
0.424 (1.00) |
0.682 (1.00) |
0.448 (1.00) |
0.629 (1.00) |
0.363 (1.00) |
0.0706 (1.00) |
0.633 (1.00) |
0.166 (1.00) |
| 5q gain | 8 (12%) | 58 |
0.424 (1.00) |
0.682 (1.00) |
0.448 (1.00) |
0.629 (1.00) |
0.363 (1.00) |
0.0706 (1.00) |
0.633 (1.00) |
0.166 (1.00) |
| 7p gain | 24 (36%) | 42 |
0.151 (1.00) |
1 (1.00) |
0.0169 (1.00) |
0.0167 (1.00) |
0.844 (1.00) |
1 (1.00) |
0.741 (1.00) |
0.423 (1.00) |
| 7q gain | 24 (36%) | 42 |
0.151 (1.00) |
1 (1.00) |
0.0169 (1.00) |
0.0167 (1.00) |
0.844 (1.00) |
1 (1.00) |
0.741 (1.00) |
0.423 (1.00) |
| 8p gain | 17 (26%) | 49 |
0.411 (1.00) |
0.411 (1.00) |
0.00393 (1.00) |
0.00701 (1.00) |
0.442 (1.00) |
1 (1.00) |
0.712 (1.00) |
0.155 (1.00) |
| 8q gain | 18 (27%) | 48 |
0.342 (1.00) |
0.399 (1.00) |
0.00128 (0.54) |
0.00181 (0.76) |
0.679 (1.00) |
1 (1.00) |
0.926 (1.00) |
0.0785 (1.00) |
| 9p gain | 10 (15%) | 56 |
1 (1.00) |
1 (1.00) |
0.722 (1.00) |
0.644 (1.00) |
0.832 (1.00) |
0.616 (1.00) |
0.188 (1.00) |
0.47 (1.00) |
| 9q gain | 10 (15%) | 56 |
1 (1.00) |
1 (1.00) |
0.722 (1.00) |
0.644 (1.00) |
0.832 (1.00) |
0.616 (1.00) |
0.188 (1.00) |
0.47 (1.00) |
| 10p gain | 4 (6%) | 62 |
0.438 (1.00) |
0.162 (1.00) |
0.515 (1.00) |
0.336 (1.00) |
1 (1.00) |
0.452 (1.00) |
0.534 (1.00) |
0.0739 (1.00) |
| 12p gain | 19 (29%) | 47 |
0.482 (1.00) |
0.766 (1.00) |
0.0793 (1.00) |
0.13 (1.00) |
1 (1.00) |
0.709 (1.00) |
0.845 (1.00) |
0.0871 (1.00) |
| 12q gain | 20 (30%) | 46 |
0.445 (1.00) |
0.591 (1.00) |
0.14 (1.00) |
0.052 (1.00) |
0.917 (1.00) |
1 (1.00) |
0.921 (1.00) |
0.169 (1.00) |
| 14q gain | 21 (32%) | 45 |
0.106 (1.00) |
0.796 (1.00) |
0.000833 (0.36) |
0.00261 (1.00) |
0.423 (1.00) |
0.252 (1.00) |
0.878 (1.00) |
0.0499 (1.00) |
| 15q gain | 21 (32%) | 45 |
0.106 (1.00) |
0.796 (1.00) |
0.00209 (0.866) |
0.00789 (1.00) |
0.761 (1.00) |
0.252 (1.00) |
0.998 (1.00) |
0.312 (1.00) |
| 18p gain | 17 (26%) | 49 |
0.518 (1.00) |
0.806 (1.00) |
0.00215 (0.887) |
0.00497 (1.00) |
0.905 (1.00) |
0.684 (1.00) |
0.315 (1.00) |
0.033 (1.00) |
| 18q gain | 16 (24%) | 50 |
0.638 (1.00) |
0.928 (1.00) |
0.00105 (0.448) |
0.00379 (1.00) |
0.785 (1.00) |
1 (1.00) |
0.385 (1.00) |
0.0366 (1.00) |
| 19p gain | 19 (29%) | 47 |
0.482 (1.00) |
0.268 (1.00) |
0.0218 (1.00) |
0.0111 (1.00) |
0.642 (1.00) |
1 (1.00) |
0.929 (1.00) |
0.0518 (1.00) |
| 19q gain | 17 (26%) | 49 |
0.518 (1.00) |
0.305 (1.00) |
0.00143 (0.602) |
0.00123 (0.52) |
0.4 (1.00) |
0.427 (1.00) |
0.912 (1.00) |
0.00957 (1.00) |
| 20p gain | 20 (30%) | 46 |
0.216 (1.00) |
0.591 (1.00) |
0.0316 (1.00) |
0.00987 (1.00) |
0.681 (1.00) |
0.712 (1.00) |
0.637 (1.00) |
0.349 (1.00) |
| 20q gain | 21 (32%) | 45 |
0.227 (1.00) |
0.394 (1.00) |
0.067 (1.00) |
0.0217 (1.00) |
0.667 (1.00) |
1 (1.00) |
0.519 (1.00) |
0.368 (1.00) |
| 21q gain | 4 (6%) | 62 |
0.438 (1.00) |
0.162 (1.00) |
0.515 (1.00) |
0.336 (1.00) |
0.678 (1.00) |
0.452 (1.00) |
0.461 (1.00) |
0.361 (1.00) |
| 22q gain | 19 (29%) | 47 |
0.901 (1.00) |
0.698 (1.00) |
0.0721 (1.00) |
0.16 (1.00) |
0.505 (1.00) |
1 (1.00) |
0.996 (1.00) |
0.169 (1.00) |
| xq gain | 6 (9%) | 60 |
1 (1.00) |
0.722 (1.00) |
0.00689 (1.00) |
0.163 (1.00) |
0.264 (1.00) |
0.585 (1.00) |
0.52 (1.00) |
0.00993 (1.00) |
| 3p loss | 9 (14%) | 57 |
0.0367 (1.00) |
0.895 (1.00) |
0.0643 (1.00) |
0.00352 (1.00) |
0.904 (1.00) |
0.341 (1.00) |
0.0267 (1.00) |
0.0345 (1.00) |
| 3q loss | 8 (12%) | 58 |
0.0211 (1.00) |
0.367 (1.00) |
0.0251 (1.00) |
0.000866 (0.371) |
0.572 (1.00) |
0.586 (1.00) |
0.00763 (1.00) |
0.0089 (1.00) |
| 5p loss | 10 (15%) | 56 |
0.0987 (1.00) |
0.253 (1.00) |
0.000748 (0.324) |
0.000841 (0.361) |
0.427 (1.00) |
0.334 (1.00) |
0.619 (1.00) |
0.154 (1.00) |
| 5q loss | 10 (15%) | 56 |
0.0987 (1.00) |
0.253 (1.00) |
0.000748 (0.324) |
0.000841 (0.361) |
0.427 (1.00) |
0.334 (1.00) |
0.619 (1.00) |
0.154 (1.00) |
| 9p loss | 10 (15%) | 56 |
1 (1.00) |
0.902 (1.00) |
0.0649 (1.00) |
0.714 (1.00) |
0.912 (1.00) |
0.616 (1.00) |
0.896 (1.00) |
0.371 (1.00) |
| 9q loss | 10 (15%) | 56 |
1 (1.00) |
0.902 (1.00) |
0.0649 (1.00) |
0.714 (1.00) |
0.912 (1.00) |
0.616 (1.00) |
0.896 (1.00) |
0.371 (1.00) |
| 11p loss | 7 (11%) | 59 |
0.628 (1.00) |
0.574 (1.00) |
0.11 (1.00) |
0.497 (1.00) |
0.203 (1.00) |
1 (1.00) |
0.269 (1.00) |
0.0436 (1.00) |
| 11q loss | 7 (11%) | 59 |
0.628 (1.00) |
0.574 (1.00) |
0.11 (1.00) |
0.497 (1.00) |
0.203 (1.00) |
1 (1.00) |
0.269 (1.00) |
0.0436 (1.00) |
| 16p loss | 4 (6%) | 62 |
0.685 (1.00) |
0.517 (1.00) |
0.0232 (1.00) |
0.00987 (1.00) |
0.678 (1.00) |
0.452 (1.00) |
0.461 (1.00) |
1 (1.00) |
| 16q loss | 5 (8%) | 61 |
0.721 (1.00) |
0.385 (1.00) |
0.00592 (1.00) |
0.000834 (0.36) |
0.428 (1.00) |
0.134 (1.00) |
0.192 (1.00) |
0.616 (1.00) |
| 18p loss | 8 (12%) | 58 |
0.677 (1.00) |
0.0965 (1.00) |
0.315 (1.00) |
0.0503 (1.00) |
0.72 (1.00) |
1 (1.00) |
0.877 (1.00) |
1 (1.00) |
| 18q loss | 10 (15%) | 56 |
0.62 (1.00) |
0.0796 (1.00) |
0.443 (1.00) |
0.138 (1.00) |
0.575 (1.00) |
0.616 (1.00) |
0.49 (1.00) |
0.958 (1.00) |
| 19q loss | 3 (5%) | 63 |
0.646 (1.00) |
0.766 (1.00) |
0.767 (1.00) |
0.823 (1.00) |
0.515 (1.00) |
0.361 (1.00) |
0.69 (1.00) |
0.439 (1.00) |
| 20p loss | 4 (6%) | 62 |
0.301 (1.00) |
0.807 (1.00) |
0.209 (1.00) |
0.213 (1.00) |
0.45 (1.00) |
1 (1.00) |
0.37 (1.00) |
0.202 (1.00) |
| 20q loss | 3 (5%) | 63 |
1 (1.00) |
0.418 (1.00) |
0.226 (1.00) |
0.132 (1.00) |
0.294 (1.00) |
1 (1.00) |
0.219 (1.00) |
0.439 (1.00) |
| 21q loss | 35 (53%) | 31 |
0.742 (1.00) |
0.00886 (1.00) |
0.247 (1.00) |
0.0443 (1.00) |
1 (1.00) |
0.724 (1.00) |
0.754 (1.00) |
0.141 (1.00) |
| 22q loss | 8 (12%) | 58 |
0.83 (1.00) |
0.134 (1.00) |
0.247 (1.00) |
0.0645 (1.00) |
0.897 (1.00) |
0.586 (1.00) |
0.731 (1.00) |
0.267 (1.00) |
| xq loss | 39 (59%) | 27 |
0.532 (1.00) |
0.0787 (1.00) |
0.00683 (1.00) |
0.00129 (0.545) |
0.293 (1.00) |
0.026 (1.00) |
0.0691 (1.00) |
0.0281 (1.00) |
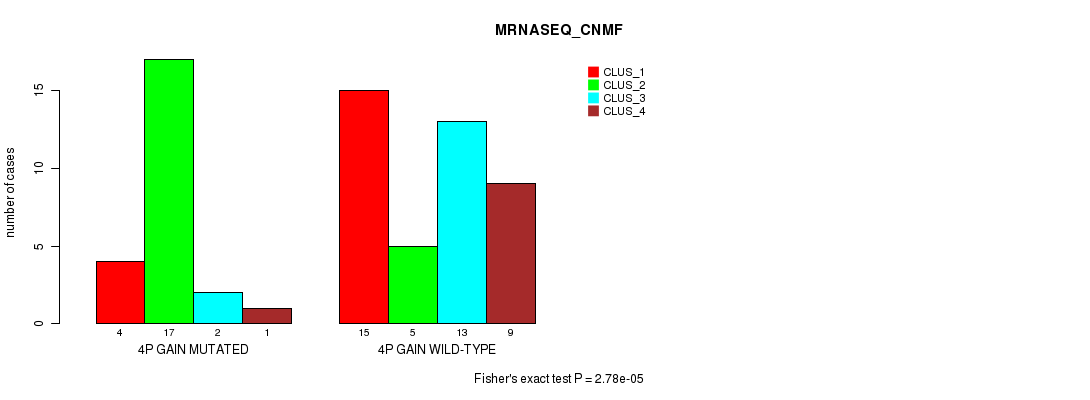
P value = 2.78e-05 (Fisher's exact test), Q value = 0.013
Table S1. Gene #3: '4p gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 4P GAIN MUTATED | 4 | 17 | 2 | 1 |
| 4P GAIN WILD-TYPE | 15 | 5 | 13 | 9 |
Figure S1. Get High-res Image Gene #3: '4p gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
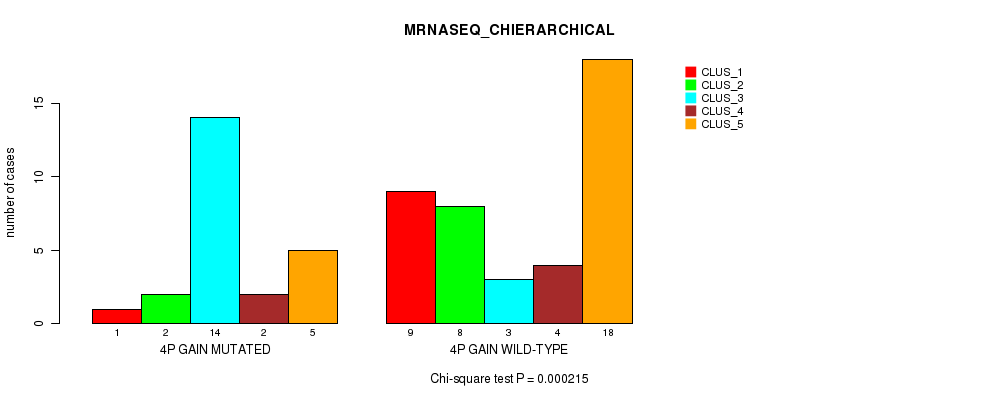
P value = 0.000215 (Chi-square test), Q value = 0.096
Table S2. Gene #3: '4p gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 4P GAIN MUTATED | 1 | 2 | 14 | 2 | 5 |
| 4P GAIN WILD-TYPE | 9 | 8 | 3 | 4 | 18 |
Figure S2. Get High-res Image Gene #3: '4p gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
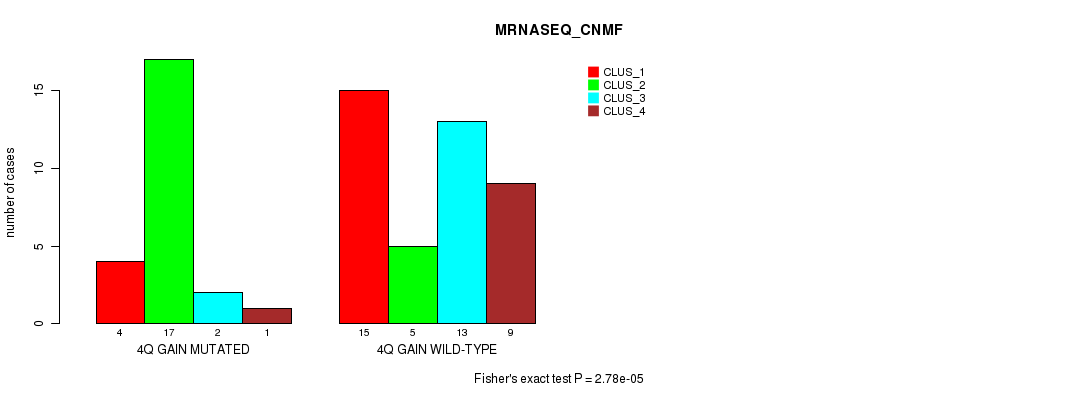
P value = 2.78e-05 (Fisher's exact test), Q value = 0.013
Table S3. Gene #4: '4q gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 4Q GAIN MUTATED | 4 | 17 | 2 | 1 |
| 4Q GAIN WILD-TYPE | 15 | 5 | 13 | 9 |
Figure S3. Get High-res Image Gene #4: '4q gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
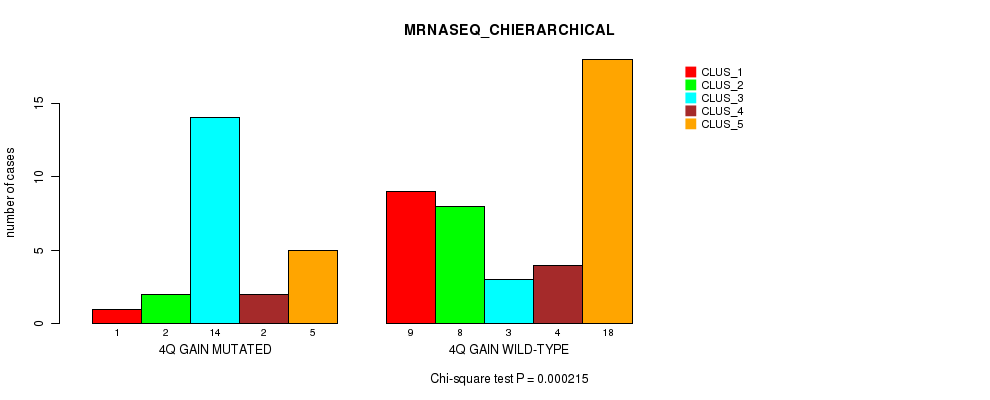
P value = 0.000215 (Chi-square test), Q value = 0.096
Table S4. Gene #4: '4q gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 4Q GAIN MUTATED | 1 | 2 | 14 | 2 | 5 |
| 4Q GAIN WILD-TYPE | 9 | 8 | 3 | 4 | 18 |
Figure S4. Get High-res Image Gene #4: '4q gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
P value = 2e-05 (Fisher's exact test), Q value = 0.0093
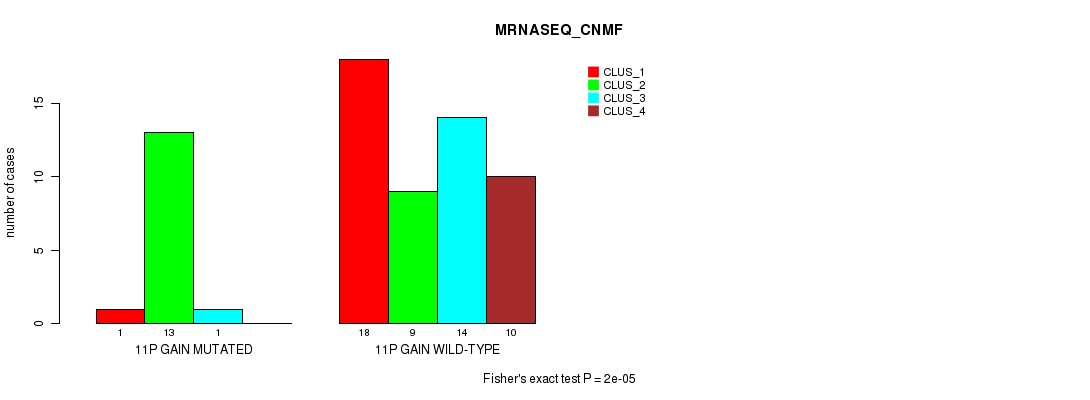
Table S5. Gene #14: '11p gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 11P GAIN MUTATED | 1 | 13 | 1 | 0 |
| 11P GAIN WILD-TYPE | 18 | 9 | 14 | 10 |
Figure S5. Get High-res Image Gene #14: '11p gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
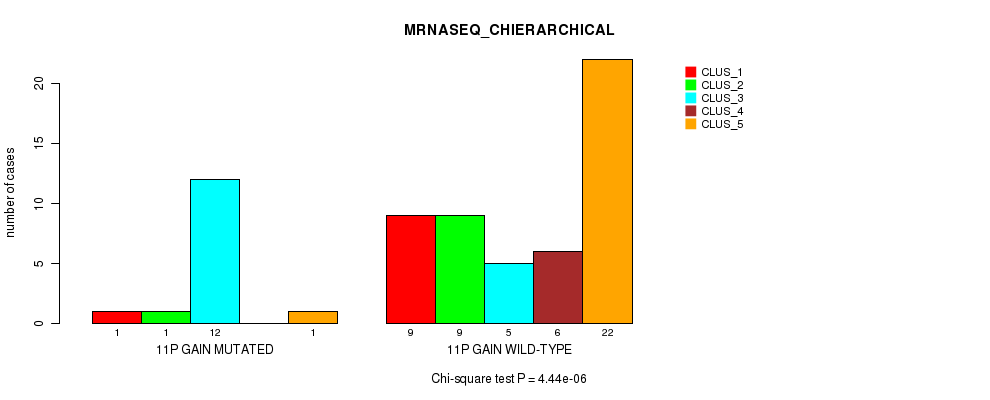
P value = 4.44e-06 (Chi-square test), Q value = 0.0021
Table S6. Gene #14: '11p gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 11P GAIN MUTATED | 1 | 1 | 12 | 0 | 1 |
| 11P GAIN WILD-TYPE | 9 | 9 | 5 | 6 | 22 |
Figure S6. Get High-res Image Gene #14: '11p gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
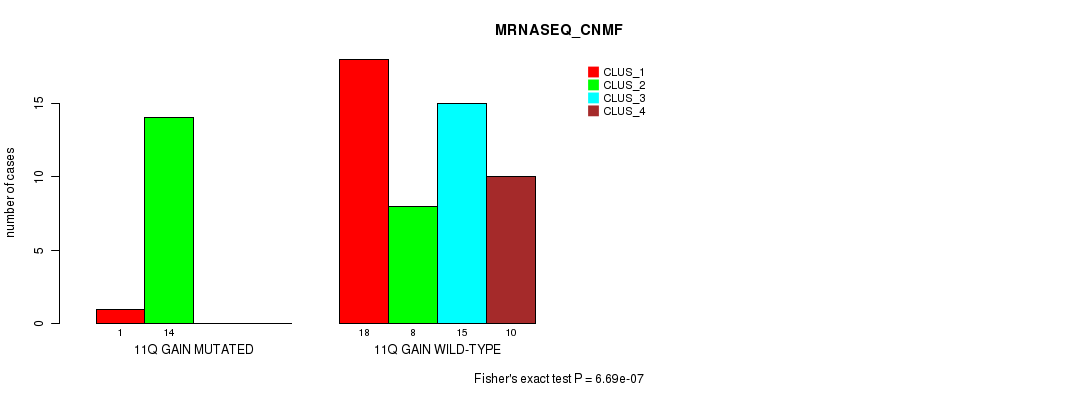
P value = 6.69e-07 (Fisher's exact test), Q value = 0.00032
Table S7. Gene #15: '11q gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 11Q GAIN MUTATED | 1 | 14 | 0 | 0 |
| 11Q GAIN WILD-TYPE | 18 | 8 | 15 | 10 |
Figure S7. Get High-res Image Gene #15: '11q gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
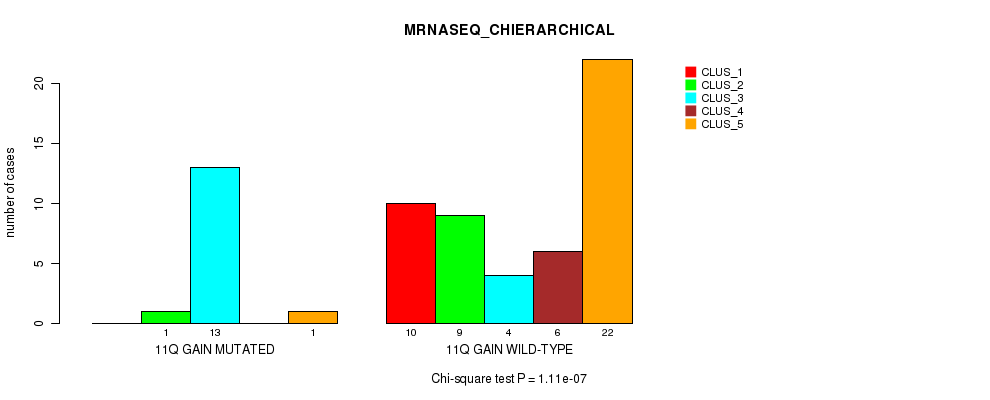
P value = 1.11e-07 (Chi-square test), Q value = 5.4e-05
Table S8. Gene #15: '11q gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 11Q GAIN MUTATED | 0 | 1 | 13 | 0 | 1 |
| 11Q GAIN WILD-TYPE | 10 | 9 | 4 | 6 | 22 |
Figure S8. Get High-res Image Gene #15: '11q gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
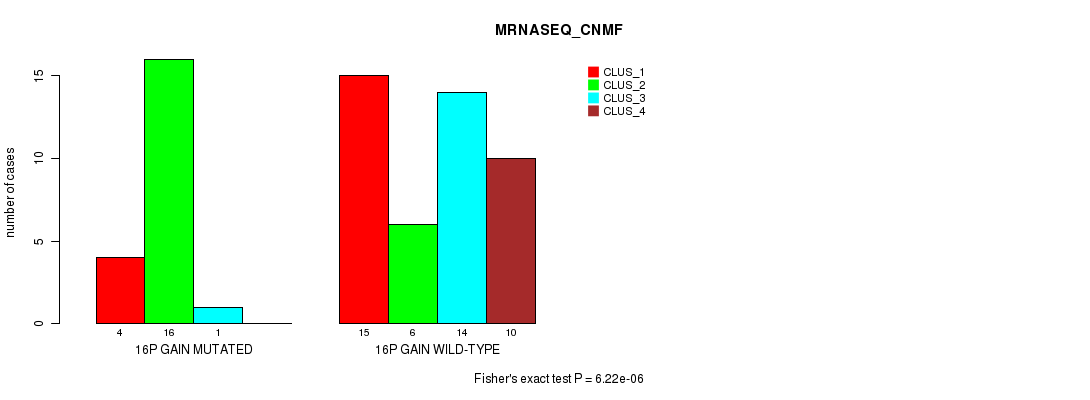
P value = 6.22e-06 (Fisher's exact test), Q value = 0.0029
Table S9. Gene #20: '16p gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 16P GAIN MUTATED | 4 | 16 | 1 | 0 |
| 16P GAIN WILD-TYPE | 15 | 6 | 14 | 10 |
Figure S9. Get High-res Image Gene #20: '16p gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
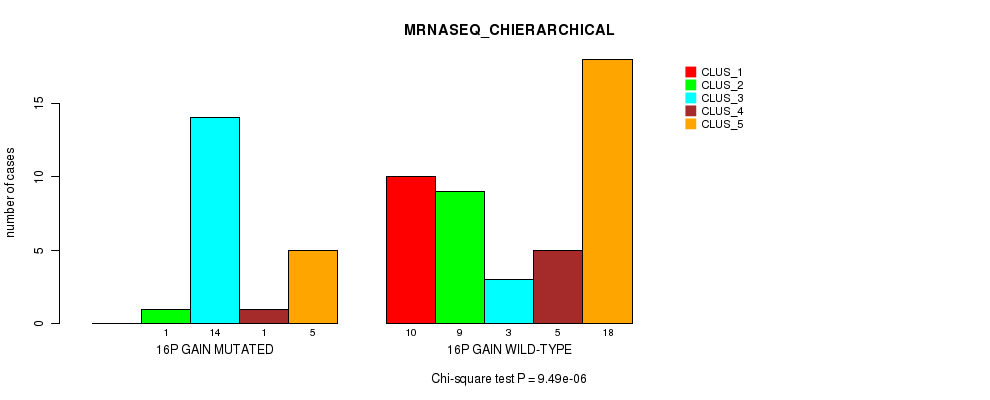
P value = 9.49e-06 (Chi-square test), Q value = 0.0044
Table S10. Gene #20: '16p gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 16P GAIN MUTATED | 0 | 1 | 14 | 1 | 5 |
| 16P GAIN WILD-TYPE | 10 | 9 | 3 | 5 | 18 |
Figure S10. Get High-res Image Gene #20: '16p gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
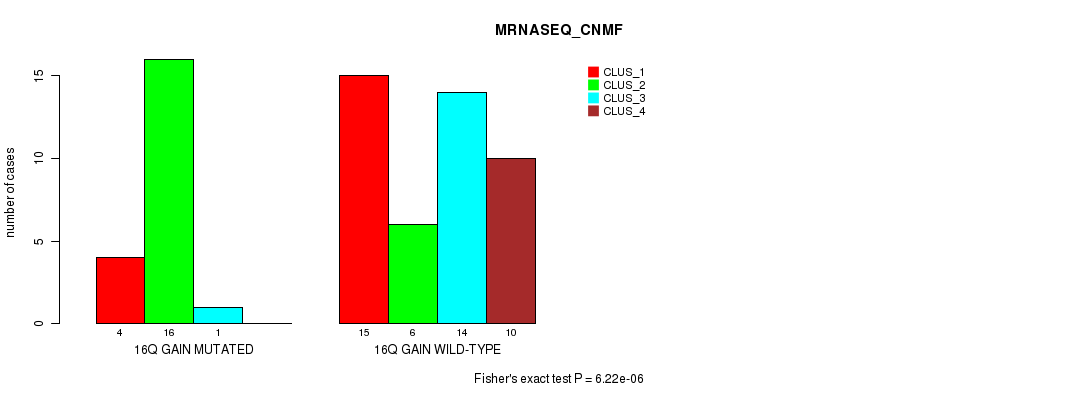
P value = 6.22e-06 (Fisher's exact test), Q value = 0.0029
Table S11. Gene #21: '16q gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 16Q GAIN MUTATED | 4 | 16 | 1 | 0 |
| 16Q GAIN WILD-TYPE | 15 | 6 | 14 | 10 |
Figure S11. Get High-res Image Gene #21: '16q gain' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
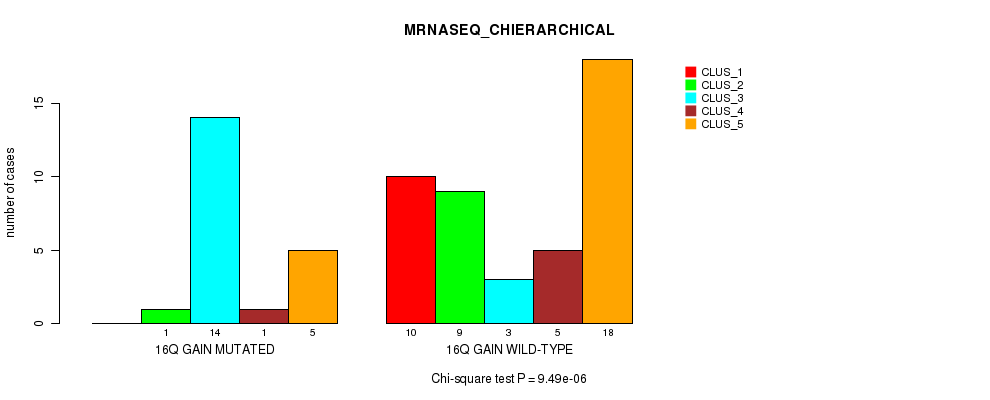
P value = 9.49e-06 (Chi-square test), Q value = 0.0044
Table S12. Gene #21: '16q gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 16Q GAIN MUTATED | 0 | 1 | 14 | 1 | 5 |
| 16Q GAIN WILD-TYPE | 10 | 9 | 3 | 5 | 18 |
Figure S12. Get High-res Image Gene #21: '16q gain' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
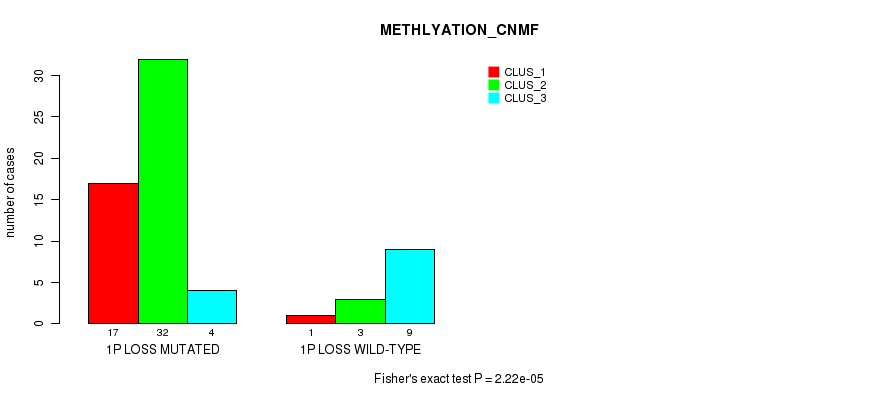
P value = 2.22e-05 (Fisher's exact test), Q value = 0.01
Table S13. Gene #31: '1p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 1P LOSS MUTATED | 17 | 32 | 4 |
| 1P LOSS WILD-TYPE | 1 | 3 | 9 |
Figure S13. Get High-res Image Gene #31: '1p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
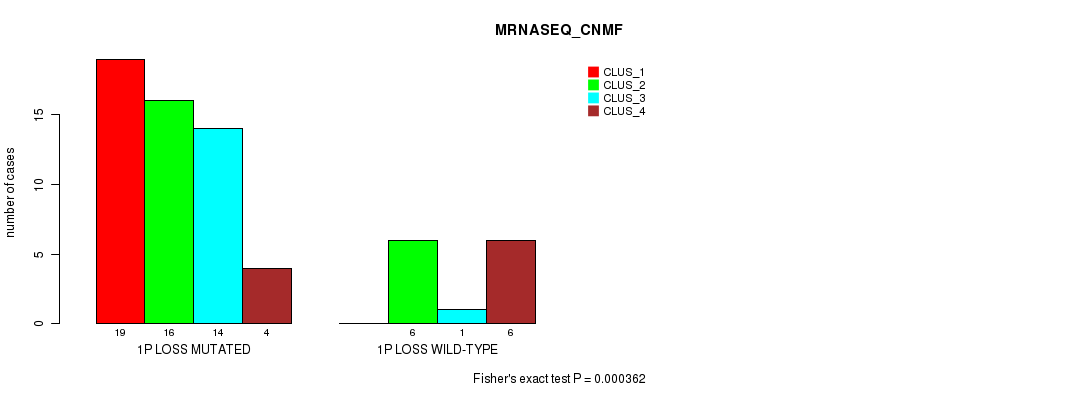
P value = 0.000362 (Fisher's exact test), Q value = 0.16
Table S14. Gene #31: '1p loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 1P LOSS MUTATED | 19 | 16 | 14 | 4 |
| 1P LOSS WILD-TYPE | 0 | 6 | 1 | 6 |
Figure S14. Get High-res Image Gene #31: '1p loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
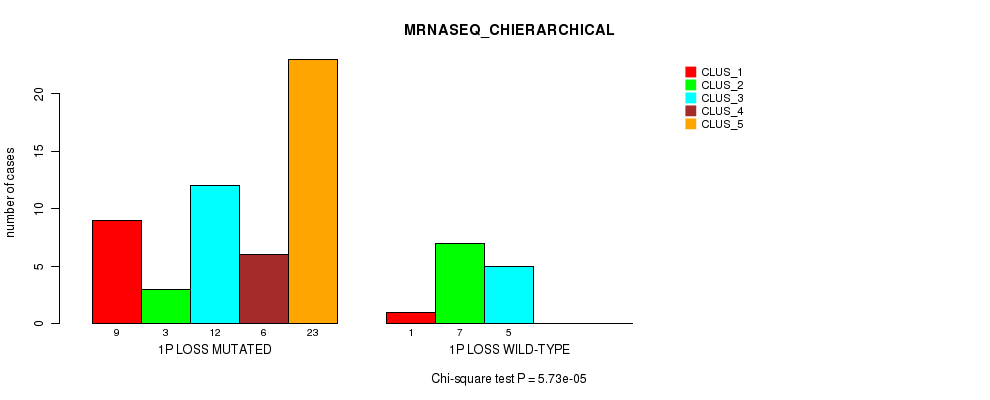
P value = 5.73e-05 (Chi-square test), Q value = 0.026
Table S15. Gene #31: '1p loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 1P LOSS MUTATED | 9 | 3 | 12 | 6 | 23 |
| 1P LOSS WILD-TYPE | 1 | 7 | 5 | 0 | 0 |
Figure S15. Get High-res Image Gene #31: '1p loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
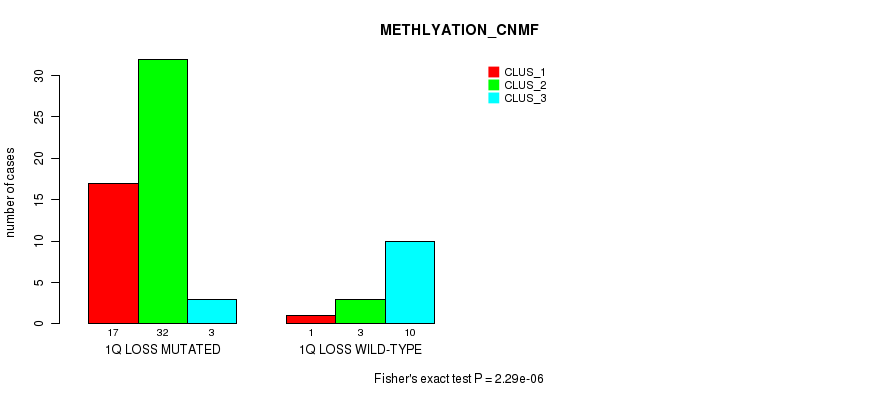
P value = 2.29e-06 (Fisher's exact test), Q value = 0.0011
Table S16. Gene #32: '1q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 1Q LOSS MUTATED | 17 | 32 | 3 |
| 1Q LOSS WILD-TYPE | 1 | 3 | 10 |
Figure S16. Get High-res Image Gene #32: '1q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
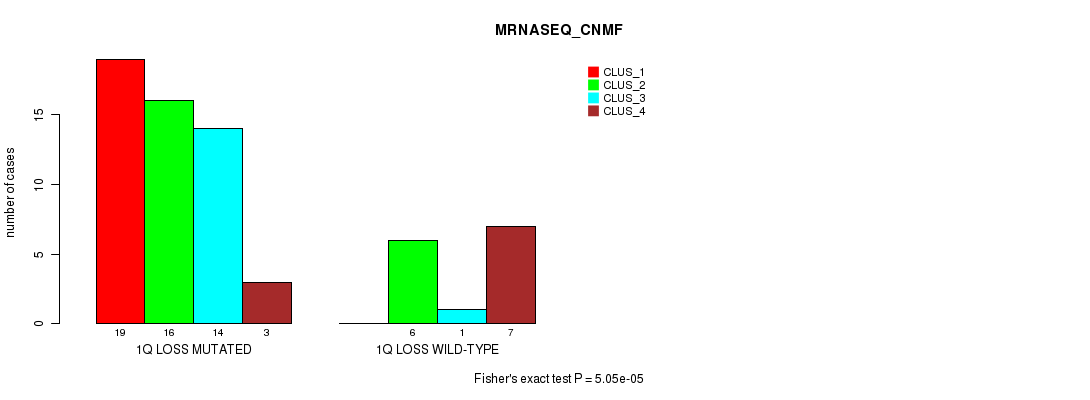
P value = 5.05e-05 (Fisher's exact test), Q value = 0.023
Table S17. Gene #32: '1q loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 1Q LOSS MUTATED | 19 | 16 | 14 | 3 |
| 1Q LOSS WILD-TYPE | 0 | 6 | 1 | 7 |
Figure S17. Get High-res Image Gene #32: '1q loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
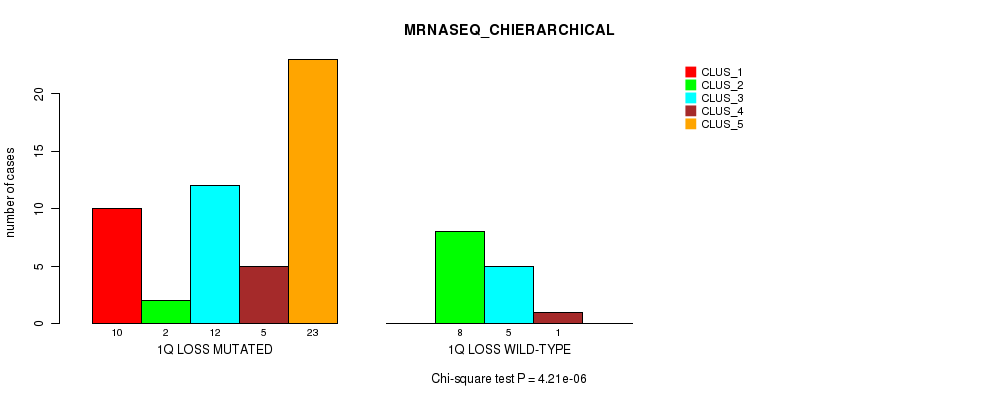
P value = 4.21e-06 (Chi-square test), Q value = 0.002
Table S18. Gene #32: '1q loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 1Q LOSS MUTATED | 10 | 2 | 12 | 5 | 23 |
| 1Q LOSS WILD-TYPE | 0 | 8 | 5 | 1 | 0 |
Figure S18. Get High-res Image Gene #32: '1q loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
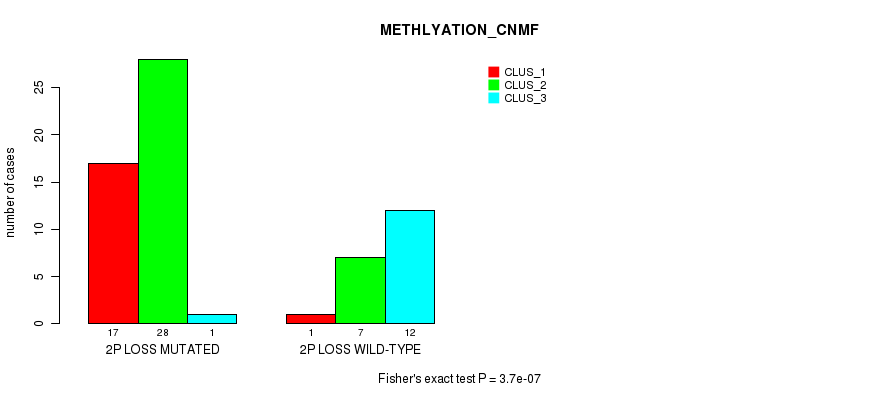
P value = 3.7e-07 (Fisher's exact test), Q value = 0.00018
Table S19. Gene #33: '2p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 2P LOSS MUTATED | 17 | 28 | 1 |
| 2P LOSS WILD-TYPE | 1 | 7 | 12 |
Figure S19. Get High-res Image Gene #33: '2p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
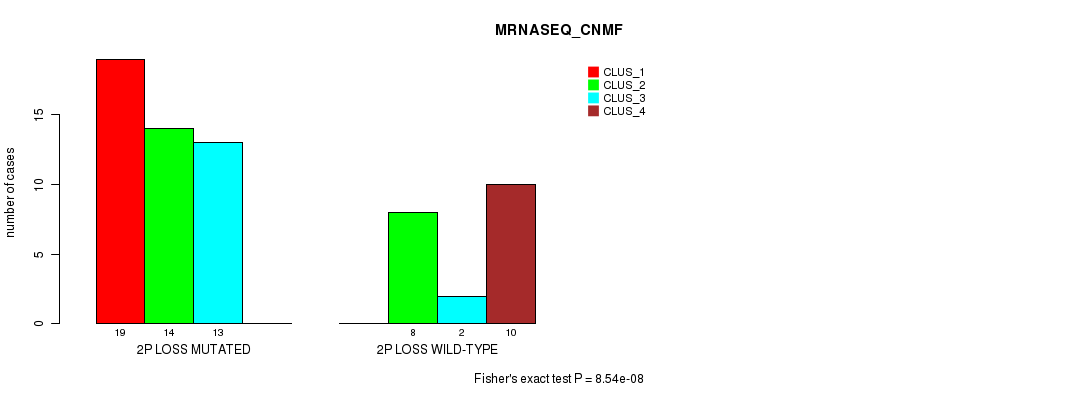
P value = 8.54e-08 (Fisher's exact test), Q value = 4.2e-05
Table S20. Gene #33: '2p loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 2P LOSS MUTATED | 19 | 14 | 13 | 0 |
| 2P LOSS WILD-TYPE | 0 | 8 | 2 | 10 |
Figure S20. Get High-res Image Gene #33: '2p loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
P value = 2.91e-07 (Chi-square test), Q value = 0.00014
Table S21. Gene #33: '2p loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 2P LOSS MUTATED | 10 | 0 | 11 | 3 | 22 |
| 2P LOSS WILD-TYPE | 0 | 10 | 6 | 3 | 1 |
Figure S21. Get High-res Image Gene #33: '2p loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
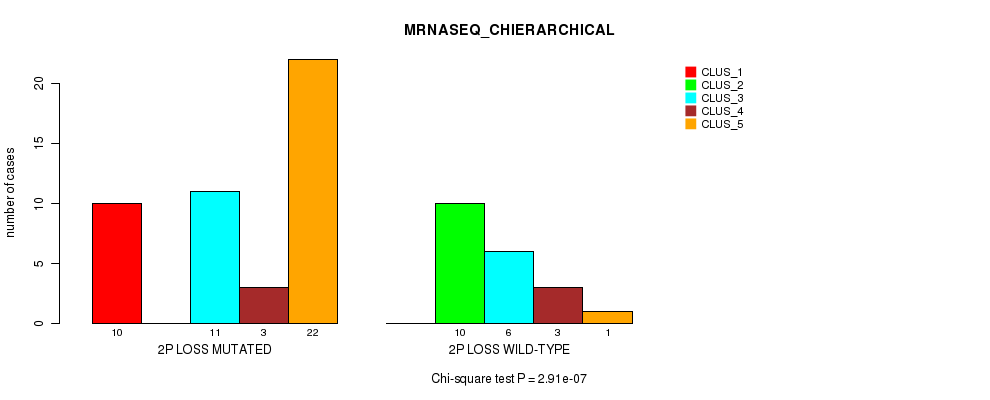
P value = 0.000161 (Fisher's exact test), Q value = 0.072
Table S22. Gene #33: '2p loss' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 9 | 57 |
| 2P LOSS MUTATED | 1 | 45 |
| 2P LOSS WILD-TYPE | 8 | 12 |
Figure S22. Get High-res Image Gene #33: '2p loss' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
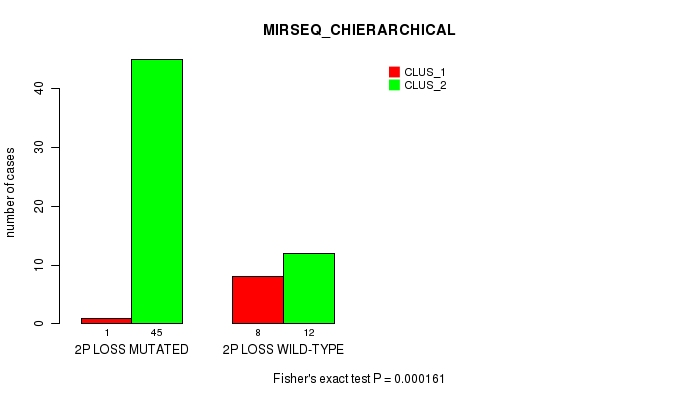
P value = 2.02e-05 (Fisher's exact test), Q value = 0.0093
Table S23. Gene #33: '2p loss' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| 2P LOSS MUTATED | 3 | 4 | 18 | 21 |
| 2P LOSS WILD-TYPE | 9 | 4 | 7 | 0 |
Figure S23. Get High-res Image Gene #33: '2p loss' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
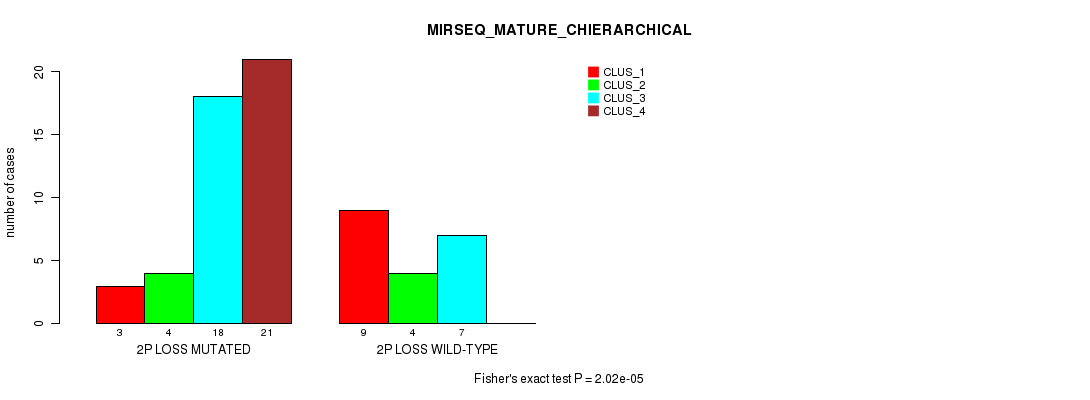
P value = 3.7e-07 (Fisher's exact test), Q value = 0.00018
Table S24. Gene #34: '2q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 2Q LOSS MUTATED | 17 | 28 | 1 |
| 2Q LOSS WILD-TYPE | 1 | 7 | 12 |
Figure S24. Get High-res Image Gene #34: '2q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
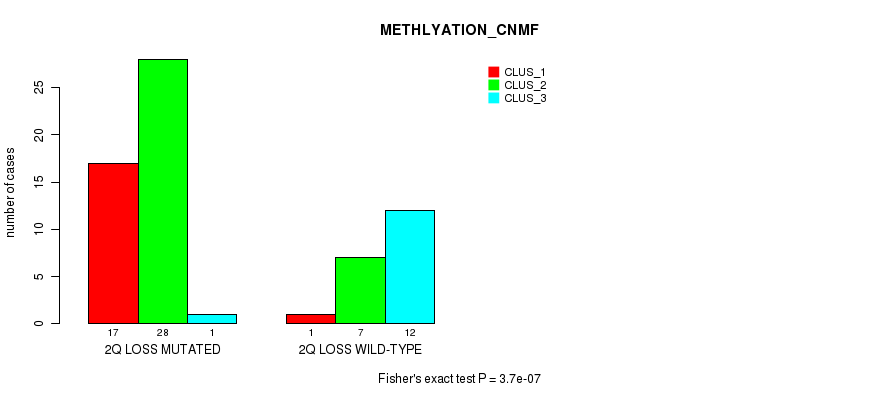
P value = 8.54e-08 (Fisher's exact test), Q value = 4.2e-05
Table S25. Gene #34: '2q loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 2Q LOSS MUTATED | 19 | 14 | 13 | 0 |
| 2Q LOSS WILD-TYPE | 0 | 8 | 2 | 10 |
Figure S25. Get High-res Image Gene #34: '2q loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
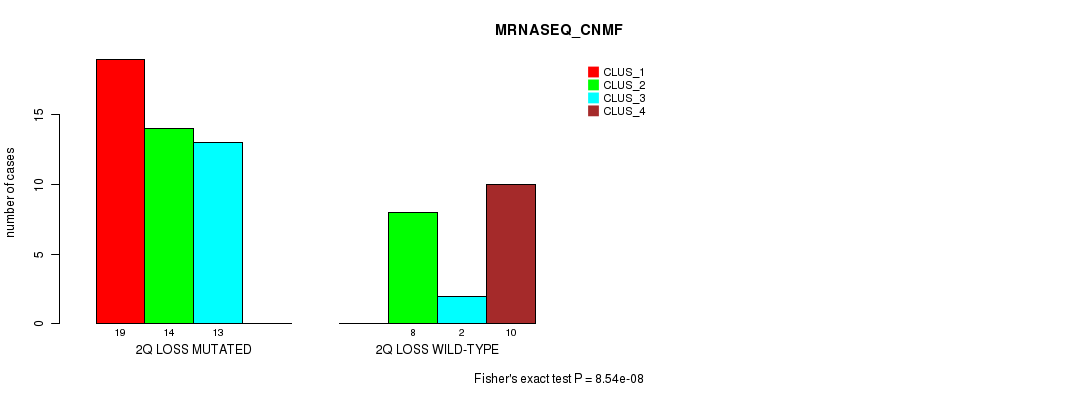
P value = 2.91e-07 (Chi-square test), Q value = 0.00014
Table S26. Gene #34: '2q loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 2Q LOSS MUTATED | 10 | 0 | 11 | 3 | 22 |
| 2Q LOSS WILD-TYPE | 0 | 10 | 6 | 3 | 1 |
Figure S26. Get High-res Image Gene #34: '2q loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
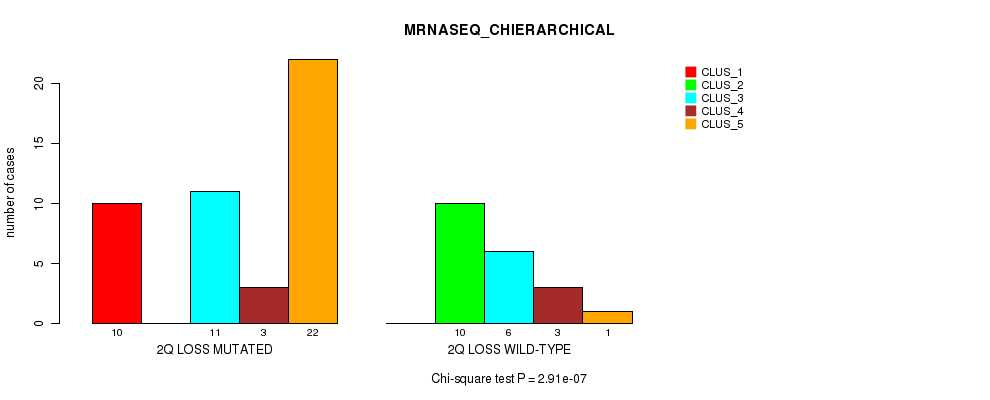
P value = 0.000161 (Fisher's exact test), Q value = 0.072
Table S27. Gene #34: '2q loss' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 9 | 57 |
| 2Q LOSS MUTATED | 1 | 45 |
| 2Q LOSS WILD-TYPE | 8 | 12 |
Figure S27. Get High-res Image Gene #34: '2q loss' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
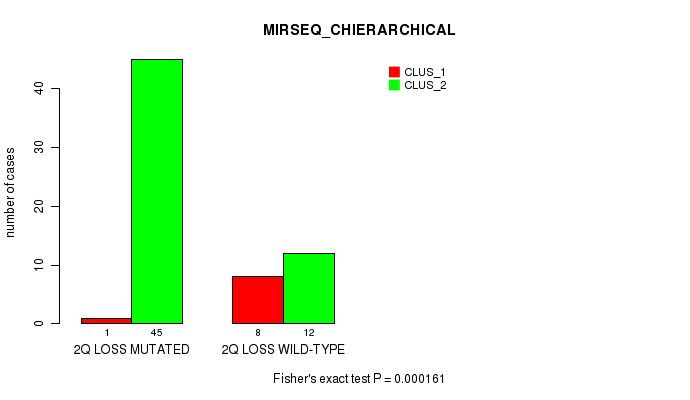
P value = 2.02e-05 (Fisher's exact test), Q value = 0.0093
Table S28. Gene #34: '2q loss' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| 2Q LOSS MUTATED | 3 | 4 | 18 | 21 |
| 2Q LOSS WILD-TYPE | 9 | 4 | 7 | 0 |
Figure S28. Get High-res Image Gene #34: '2q loss' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
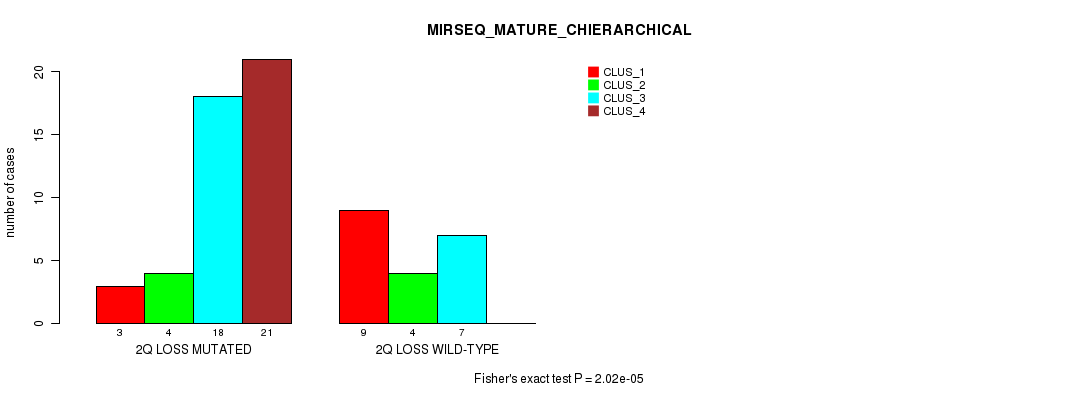
P value = 0.000107 (Fisher's exact test), Q value = 0.048
Table S29. Gene #39: '6p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 6P LOSS MUTATED | 17 | 30 | 4 |
| 6P LOSS WILD-TYPE | 1 | 5 | 9 |
Figure S29. Get High-res Image Gene #39: '6p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
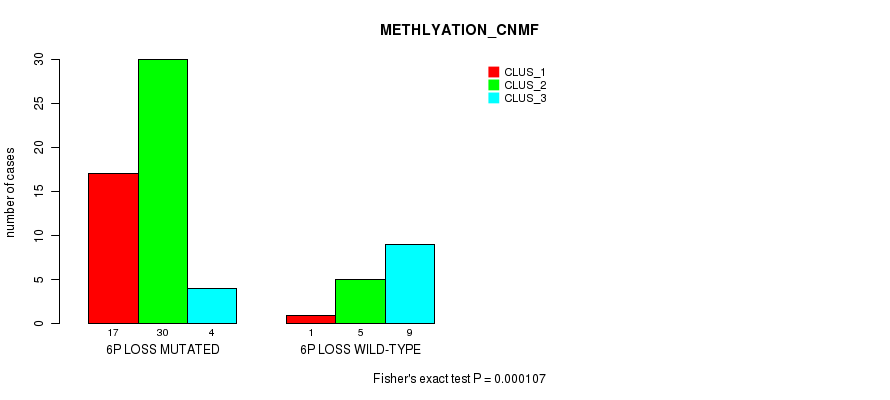
P value = 4.76e-05 (Fisher's exact test), Q value = 0.022
Table S30. Gene #39: '6p loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 6P LOSS MUTATED | 19 | 15 | 14 | 3 |
| 6P LOSS WILD-TYPE | 0 | 7 | 1 | 7 |
Figure S30. Get High-res Image Gene #39: '6p loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
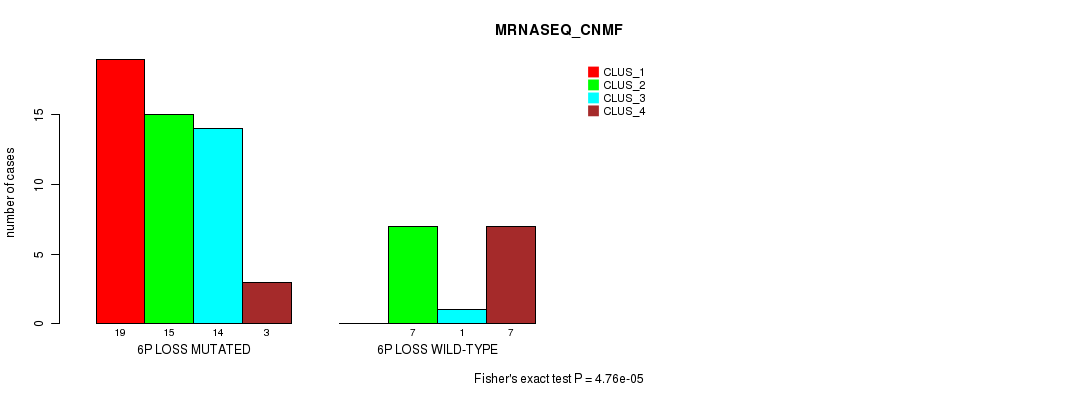
P value = 0.00033 (Chi-square test), Q value = 0.15
Table S31. Gene #39: '6p loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 6P LOSS MUTATED | 10 | 3 | 12 | 4 | 22 |
| 6P LOSS WILD-TYPE | 0 | 7 | 5 | 2 | 1 |
Figure S31. Get High-res Image Gene #39: '6p loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
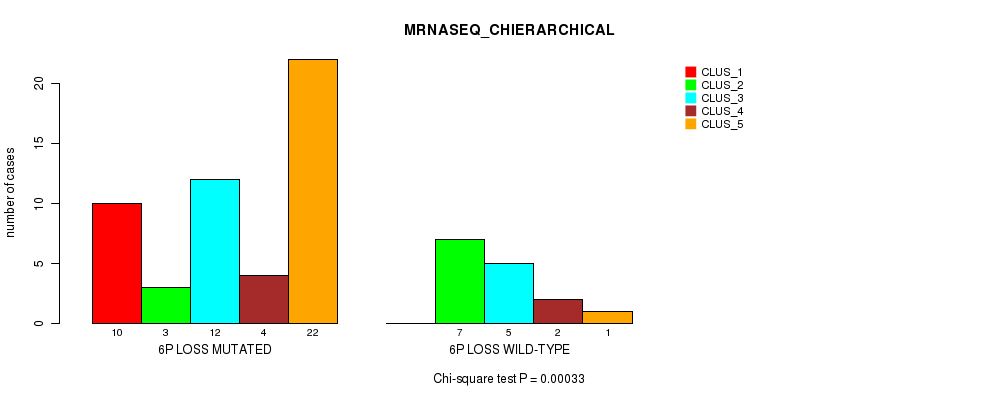
P value = 0.000107 (Fisher's exact test), Q value = 0.048
Table S32. Gene #40: '6q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 6Q LOSS MUTATED | 17 | 30 | 4 |
| 6Q LOSS WILD-TYPE | 1 | 5 | 9 |
Figure S32. Get High-res Image Gene #40: '6q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
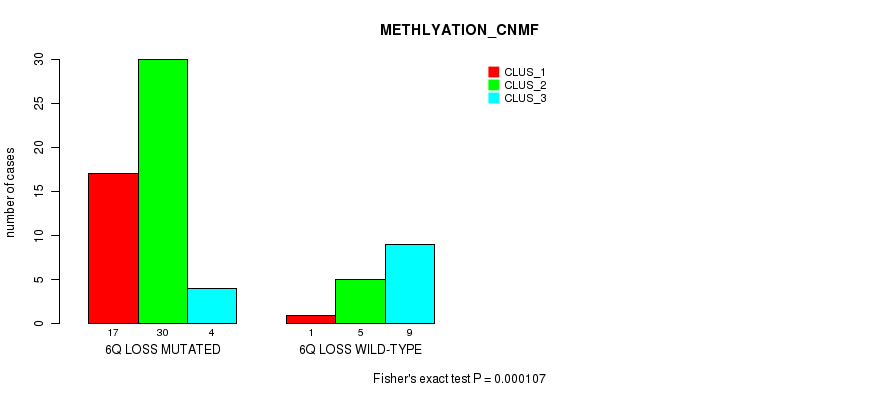
P value = 4.76e-05 (Fisher's exact test), Q value = 0.022
Table S33. Gene #40: '6q loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 6Q LOSS MUTATED | 19 | 15 | 14 | 3 |
| 6Q LOSS WILD-TYPE | 0 | 7 | 1 | 7 |
Figure S33. Get High-res Image Gene #40: '6q loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
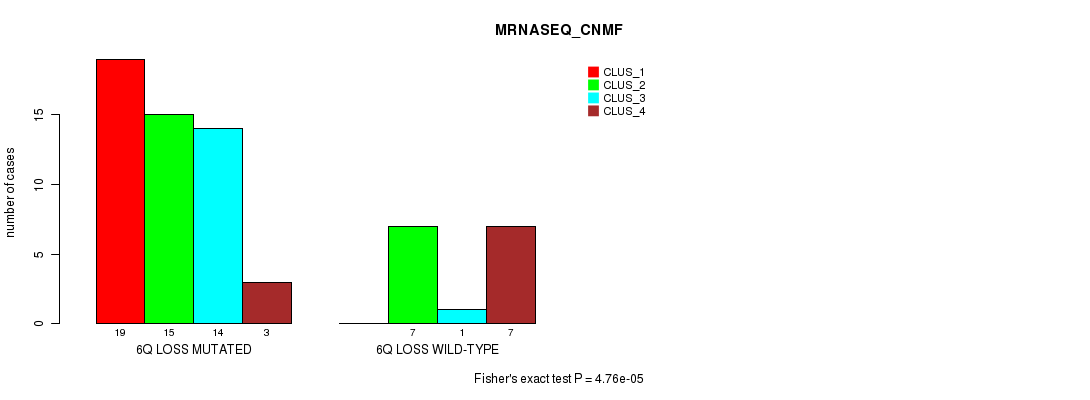
P value = 0.00033 (Chi-square test), Q value = 0.15
Table S34. Gene #40: '6q loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 6Q LOSS MUTATED | 10 | 3 | 12 | 4 | 22 |
| 6Q LOSS WILD-TYPE | 0 | 7 | 5 | 2 | 1 |
Figure S34. Get High-res Image Gene #40: '6q loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
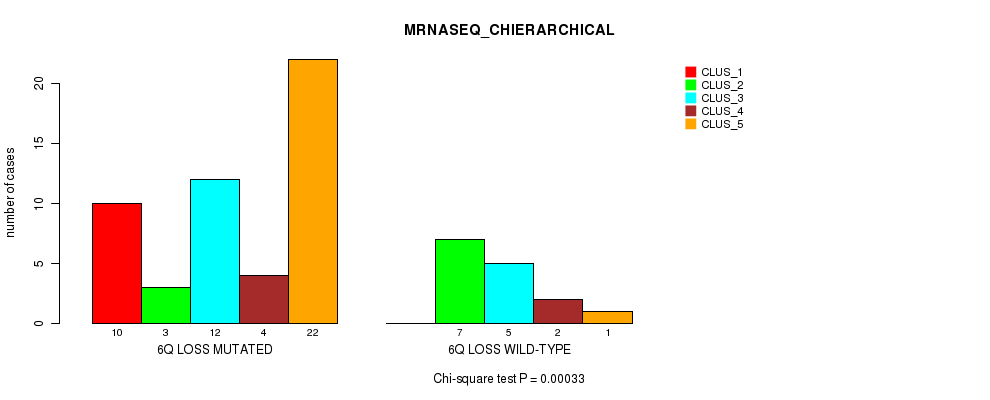
P value = 8.11e-08 (Fisher's exact test), Q value = 4e-05
Table S35. Gene #41: '8p loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 5 | 47 | 14 |
| 8P LOSS MUTATED | 0 | 0 | 9 |
| 8P LOSS WILD-TYPE | 5 | 47 | 5 |
Figure S35. Get High-res Image Gene #41: '8p loss' versus Molecular Subtype #1: 'CN_CNMF'
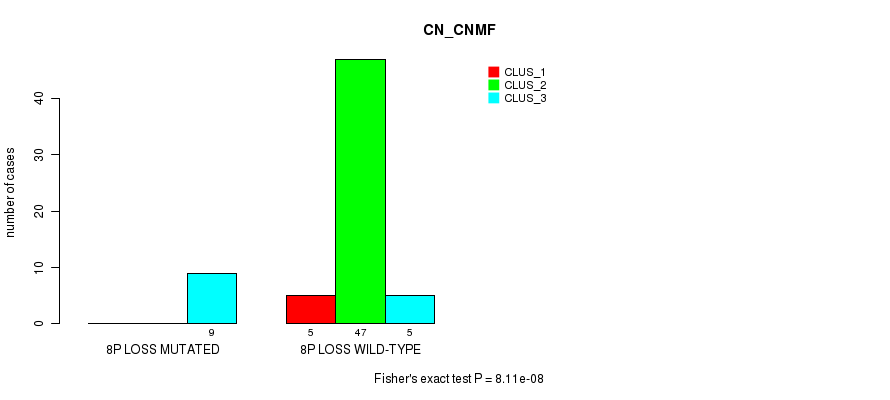
P value = 5.86e-07 (Fisher's exact test), Q value = 0.00028
Table S36. Gene #42: '8q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 5 | 47 | 14 |
| 8Q LOSS MUTATED | 0 | 0 | 8 |
| 8Q LOSS WILD-TYPE | 5 | 47 | 6 |
Figure S36. Get High-res Image Gene #42: '8q loss' versus Molecular Subtype #1: 'CN_CNMF'
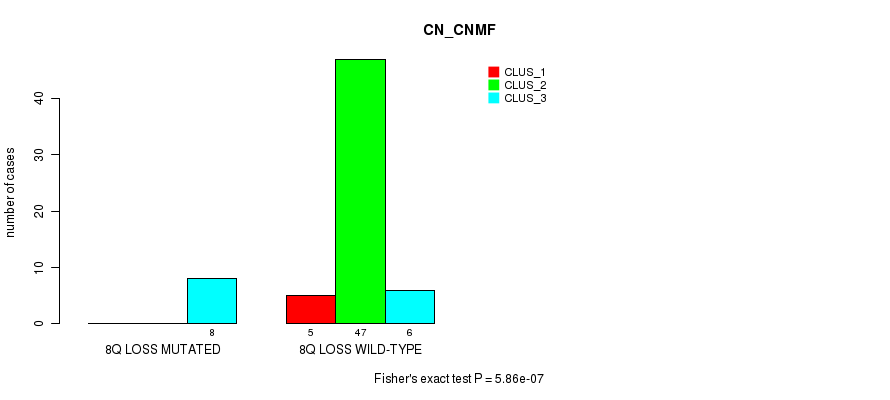
P value = 0.00038 (Fisher's exact test), Q value = 0.17
Table S37. Gene #45: '10p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 10P LOSS MUTATED | 17 | 27 | 4 |
| 10P LOSS WILD-TYPE | 1 | 8 | 9 |
Figure S37. Get High-res Image Gene #45: '10p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
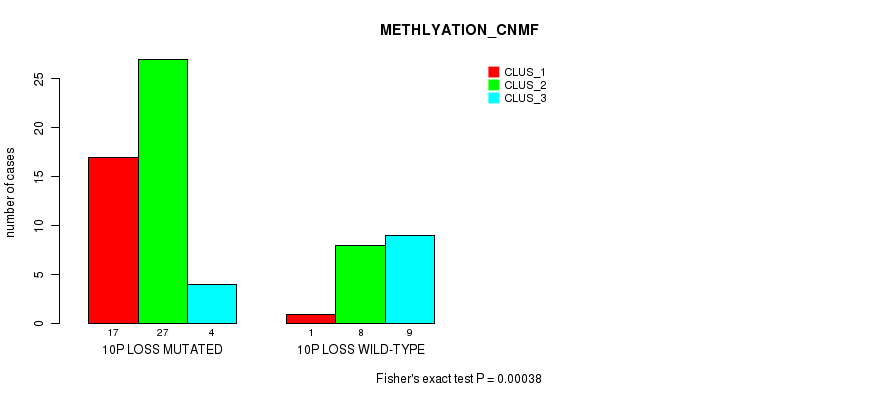
P value = 8.47e-05 (Fisher's exact test), Q value = 0.038
Table S38. Gene #45: '10p loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 10P LOSS MUTATED | 19 | 13 | 13 | 3 |
| 10P LOSS WILD-TYPE | 0 | 9 | 2 | 7 |
Figure S38. Get High-res Image Gene #45: '10p loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
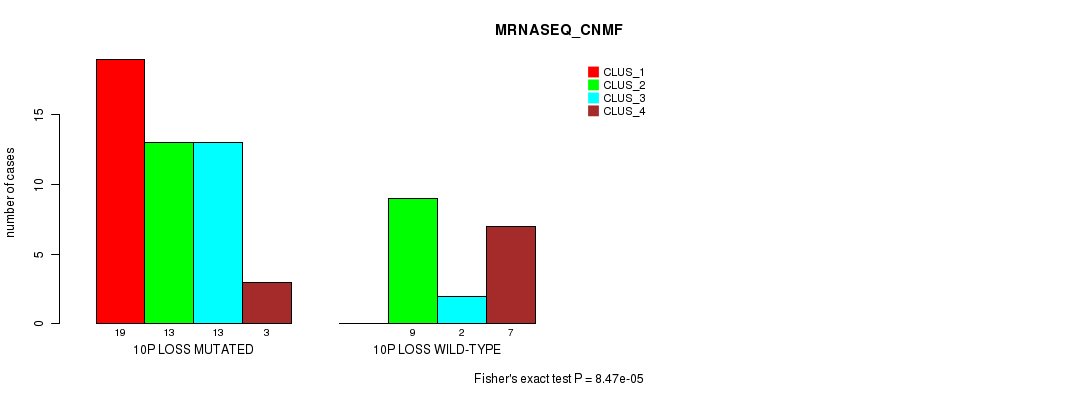
P value = 5.23e-06 (Chi-square test), Q value = 0.0024
Table S39. Gene #45: '10p loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 10P LOSS MUTATED | 10 | 2 | 9 | 4 | 23 |
| 10P LOSS WILD-TYPE | 0 | 8 | 8 | 2 | 0 |
Figure S39. Get High-res Image Gene #45: '10p loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
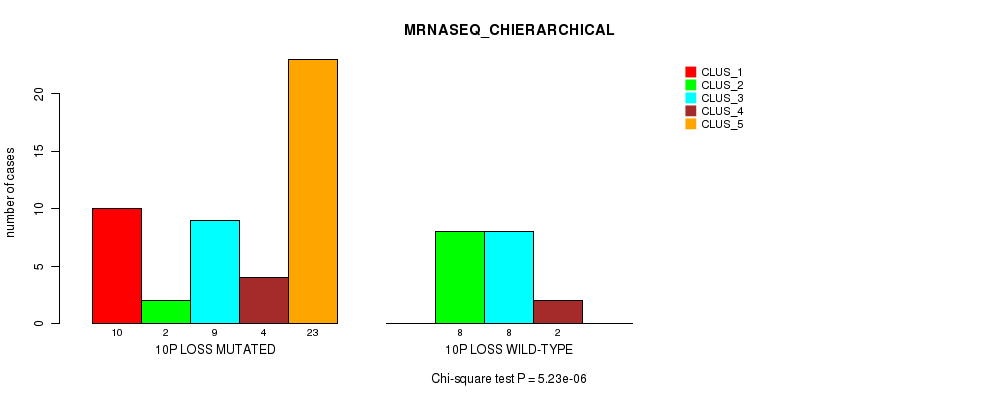
P value = 0.000336 (Fisher's exact test), Q value = 0.15
Table S40. Gene #45: '10p loss' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| 10P LOSS MUTATED | 6 | 3 | 18 | 21 |
| 10P LOSS WILD-TYPE | 6 | 5 | 7 | 0 |
Figure S40. Get High-res Image Gene #45: '10p loss' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
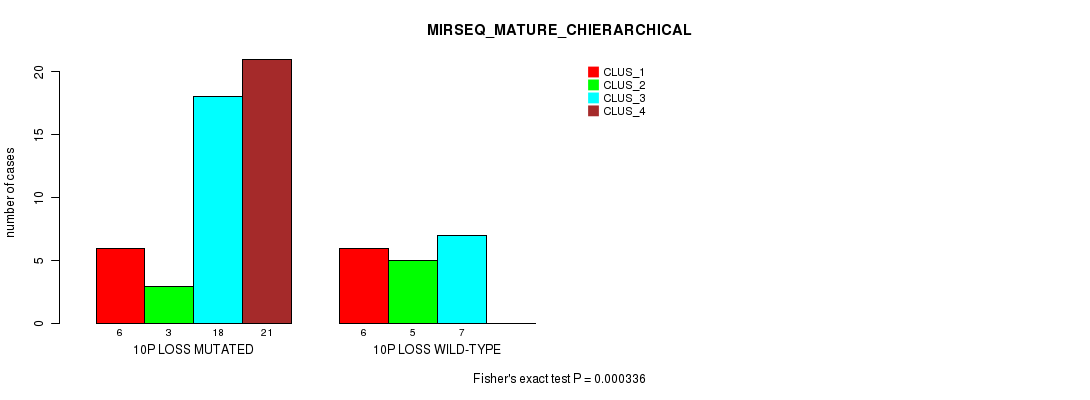
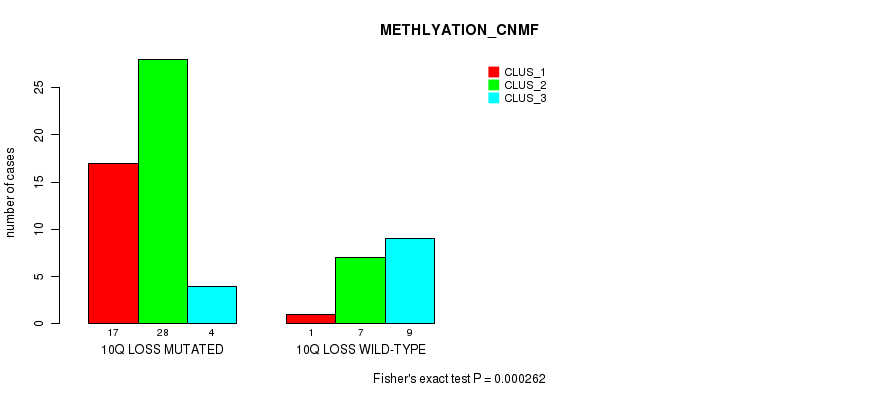
P value = 0.000262 (Fisher's exact test), Q value = 0.12
Table S41. Gene #46: '10q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 10Q LOSS MUTATED | 17 | 28 | 4 |
| 10Q LOSS WILD-TYPE | 1 | 7 | 9 |
Figure S41. Get High-res Image Gene #46: '10q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
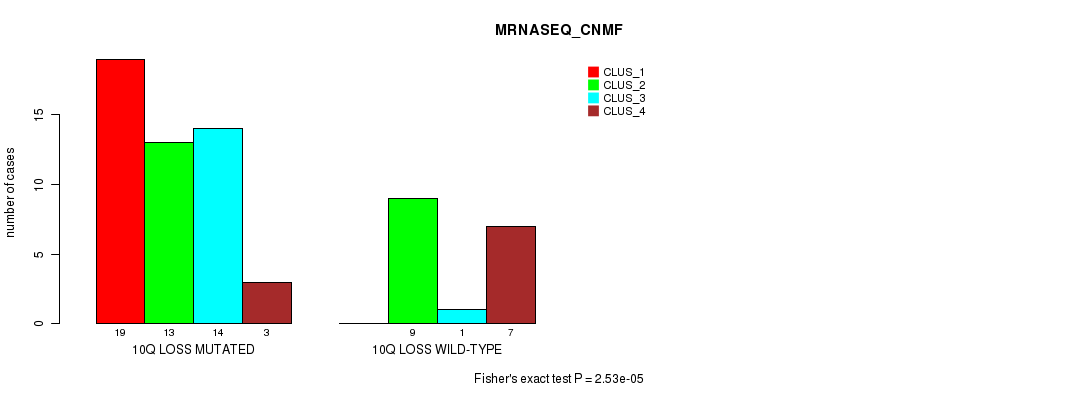
P value = 2.53e-05 (Fisher's exact test), Q value = 0.012
Table S42. Gene #46: '10q loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 10Q LOSS MUTATED | 19 | 13 | 14 | 3 |
| 10Q LOSS WILD-TYPE | 0 | 9 | 1 | 7 |
Figure S42. Get High-res Image Gene #46: '10q loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
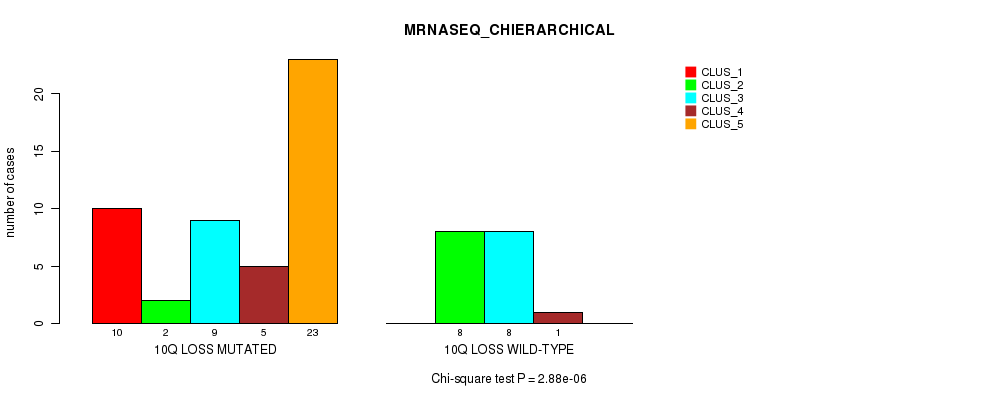
P value = 2.88e-06 (Chi-square test), Q value = 0.0014
Table S43. Gene #46: '10q loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 10Q LOSS MUTATED | 10 | 2 | 9 | 5 | 23 |
| 10Q LOSS WILD-TYPE | 0 | 8 | 8 | 1 | 0 |
Figure S43. Get High-res Image Gene #46: '10q loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
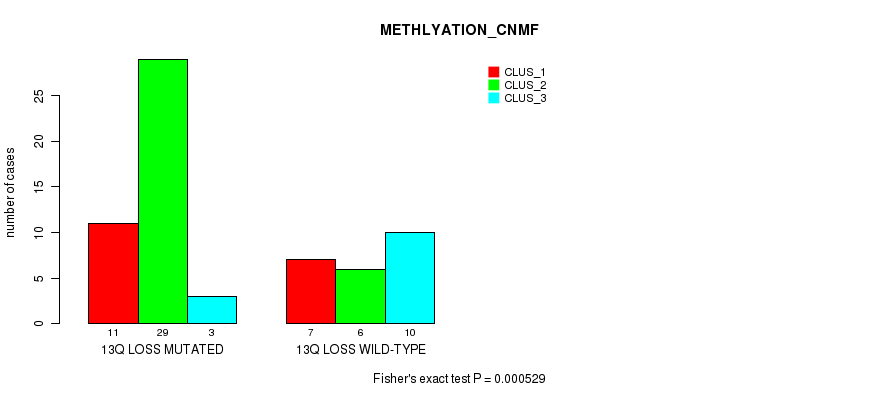
P value = 0.000529 (Fisher's exact test), Q value = 0.23
Table S44. Gene #49: '13q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 13Q LOSS MUTATED | 11 | 29 | 3 |
| 13Q LOSS WILD-TYPE | 7 | 6 | 10 |
Figure S44. Get High-res Image Gene #49: '13q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
P value = 4.75e-07 (Fisher's exact test), Q value = 0.00023
Table S45. Gene #52: '17p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 17P LOSS MUTATED | 17 | 31 | 2 |
| 17P LOSS WILD-TYPE | 1 | 4 | 11 |
Figure S45. Get High-res Image Gene #52: '17p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
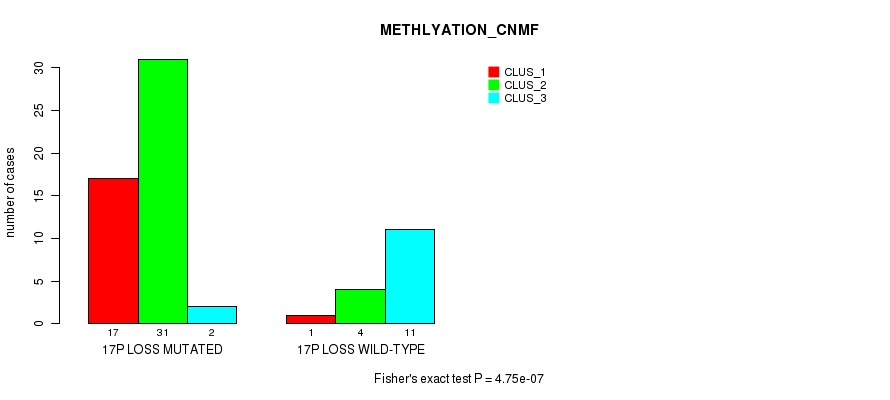
P value = 5.74e-07 (Fisher's exact test), Q value = 0.00027
Table S46. Gene #52: '17p loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 17P LOSS MUTATED | 19 | 16 | 14 | 1 |
| 17P LOSS WILD-TYPE | 0 | 6 | 1 | 9 |
Figure S46. Get High-res Image Gene #52: '17p loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
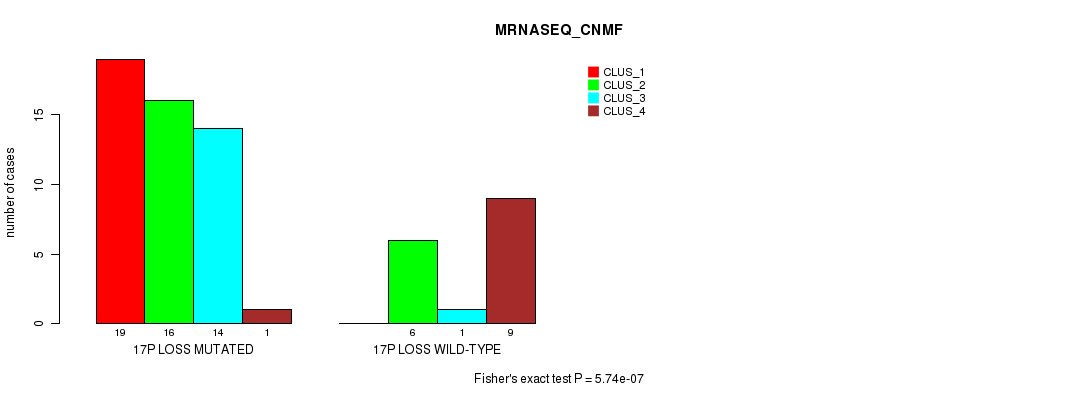
P value = 5.55e-07 (Chi-square test), Q value = 0.00027
Table S47. Gene #52: '17p loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 17P LOSS MUTATED | 10 | 1 | 12 | 4 | 23 |
| 17P LOSS WILD-TYPE | 0 | 9 | 5 | 2 | 0 |
Figure S47. Get High-res Image Gene #52: '17p loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
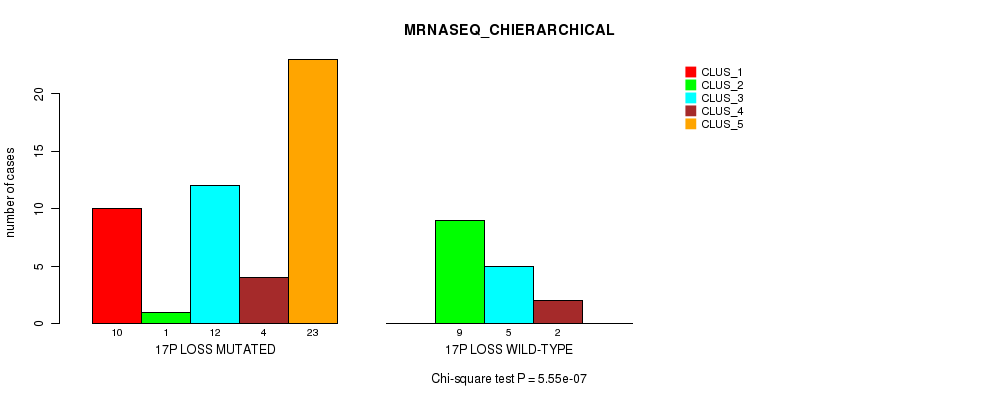
P value = 0.000396 (Fisher's exact test), Q value = 0.17
Table S48. Gene #52: '17p loss' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 9 | 57 |
| 17P LOSS MUTATED | 2 | 48 |
| 17P LOSS WILD-TYPE | 7 | 9 |
Figure S48. Get High-res Image Gene #52: '17p loss' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
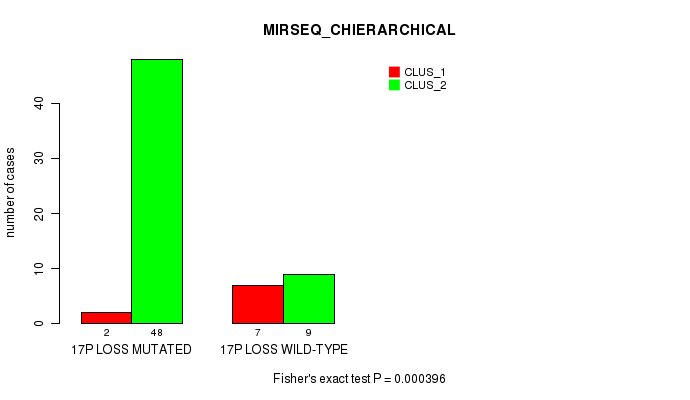
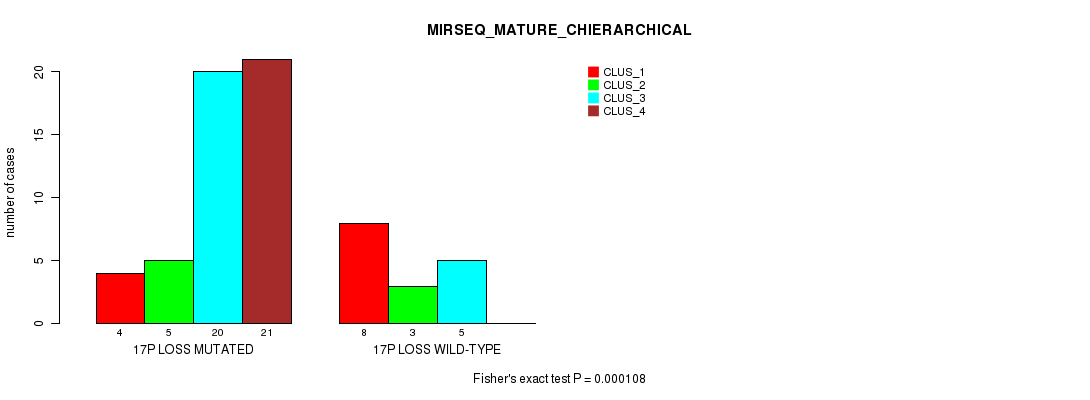
P value = 0.000108 (Fisher's exact test), Q value = 0.049
Table S49. Gene #52: '17p loss' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| 17P LOSS MUTATED | 4 | 5 | 20 | 21 |
| 17P LOSS WILD-TYPE | 8 | 3 | 5 | 0 |
Figure S49. Get High-res Image Gene #52: '17p loss' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
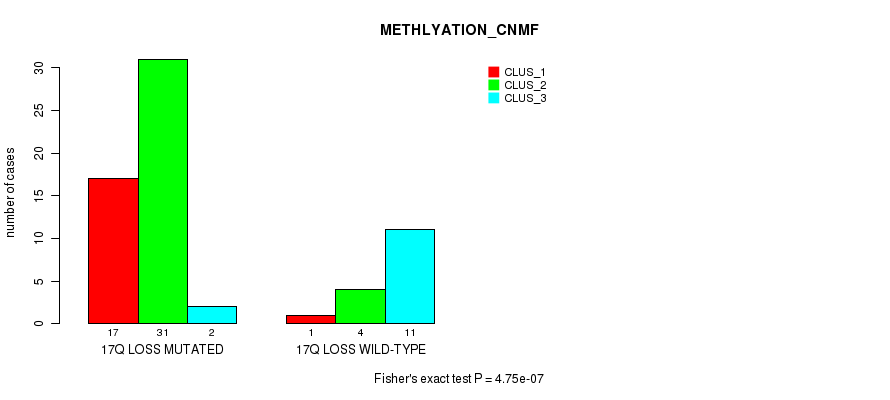
P value = 4.75e-07 (Fisher's exact test), Q value = 0.00023
Table S50. Gene #53: '17q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 35 | 13 |
| 17Q LOSS MUTATED | 17 | 31 | 2 |
| 17Q LOSS WILD-TYPE | 1 | 4 | 11 |
Figure S50. Get High-res Image Gene #53: '17q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
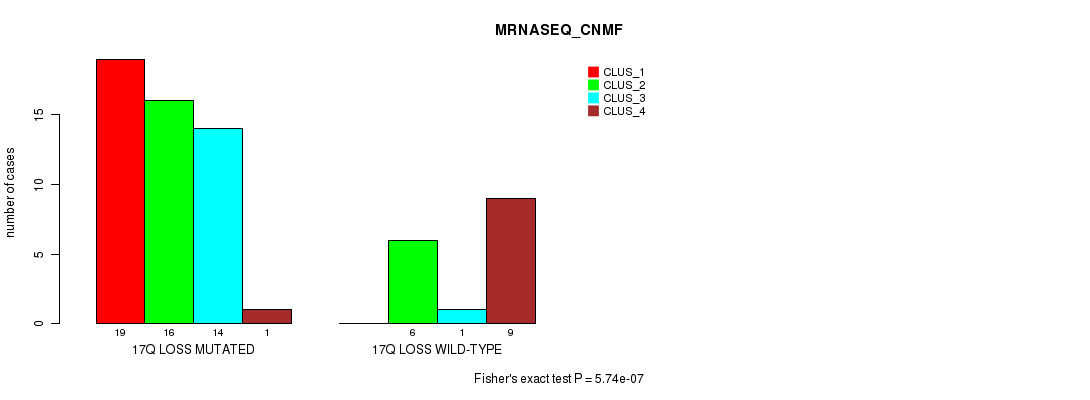
P value = 5.74e-07 (Fisher's exact test), Q value = 0.00027
Table S51. Gene #53: '17q loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 19 | 22 | 15 | 10 |
| 17Q LOSS MUTATED | 19 | 16 | 14 | 1 |
| 17Q LOSS WILD-TYPE | 0 | 6 | 1 | 9 |
Figure S51. Get High-res Image Gene #53: '17q loss' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
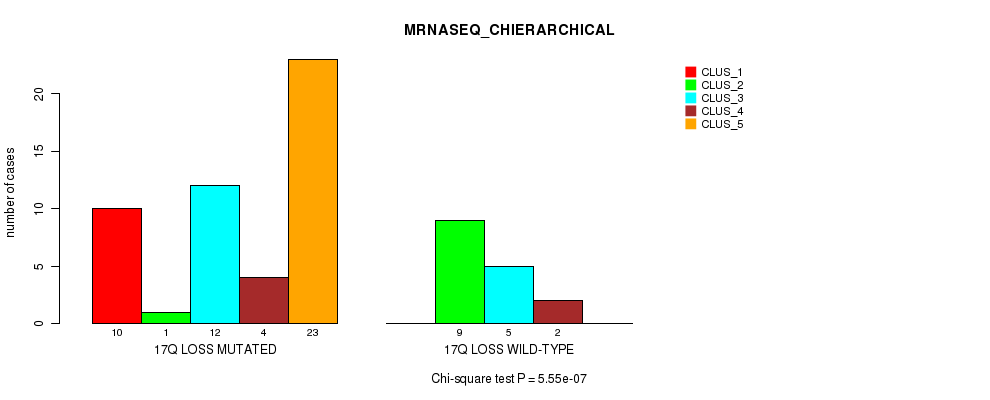
P value = 5.55e-07 (Chi-square test), Q value = 0.00027
Table S52. Gene #53: '17q loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 10 | 10 | 17 | 6 | 23 |
| 17Q LOSS MUTATED | 10 | 1 | 12 | 4 | 23 |
| 17Q LOSS WILD-TYPE | 0 | 9 | 5 | 2 | 0 |
Figure S52. Get High-res Image Gene #53: '17q loss' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
P value = 0.000396 (Fisher's exact test), Q value = 0.17
Table S53. Gene #53: '17q loss' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 9 | 57 |
| 17Q LOSS MUTATED | 2 | 48 |
| 17Q LOSS WILD-TYPE | 7 | 9 |
Figure S53. Get High-res Image Gene #53: '17q loss' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
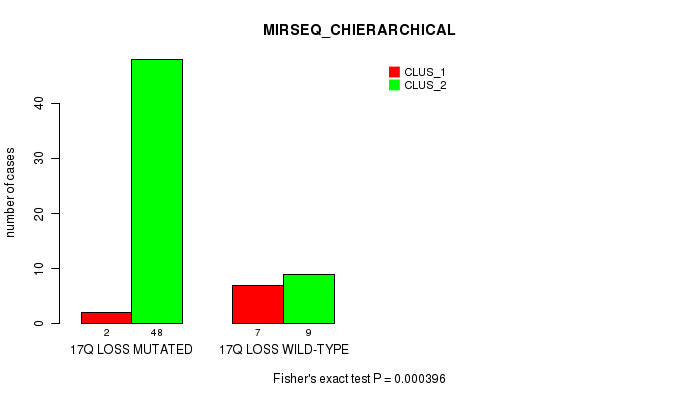
P value = 0.000108 (Fisher's exact test), Q value = 0.049
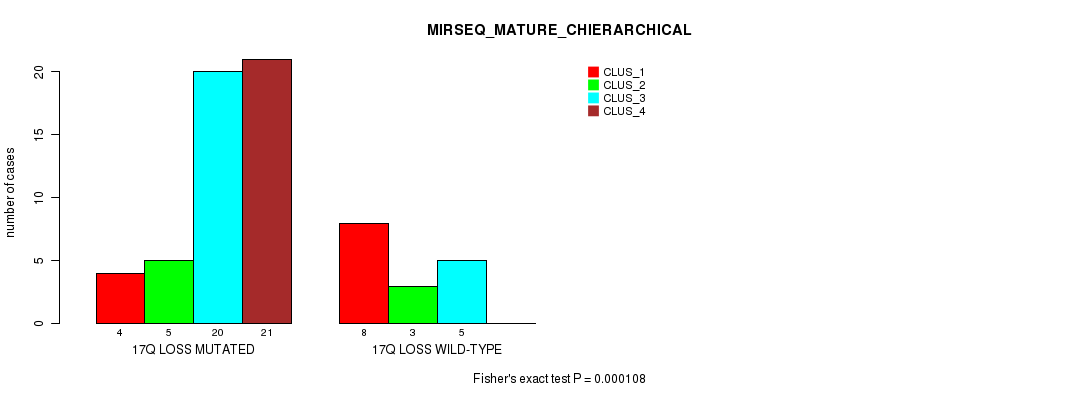
Table S54. Gene #53: '17q loss' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 8 | 25 | 21 |
| 17Q LOSS MUTATED | 4 | 5 | 20 | 21 |
| 17Q LOSS WILD-TYPE | 8 | 3 | 5 | 0 |
Figure S54. Get High-res Image Gene #53: '17q loss' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtypes file = KICH-TP.transferedmergedcluster.txt
-
Number of patients = 66
-
Number of significantly arm-level cnvs = 61
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.