This pipeline computes the correlation between significantly recurrent gene mutations and selected clinical features.
Testing the association between mutation status of 20 genes and 11 clinical features across 220 patients, 5 significant findings detected with Q value < 0.25.
-
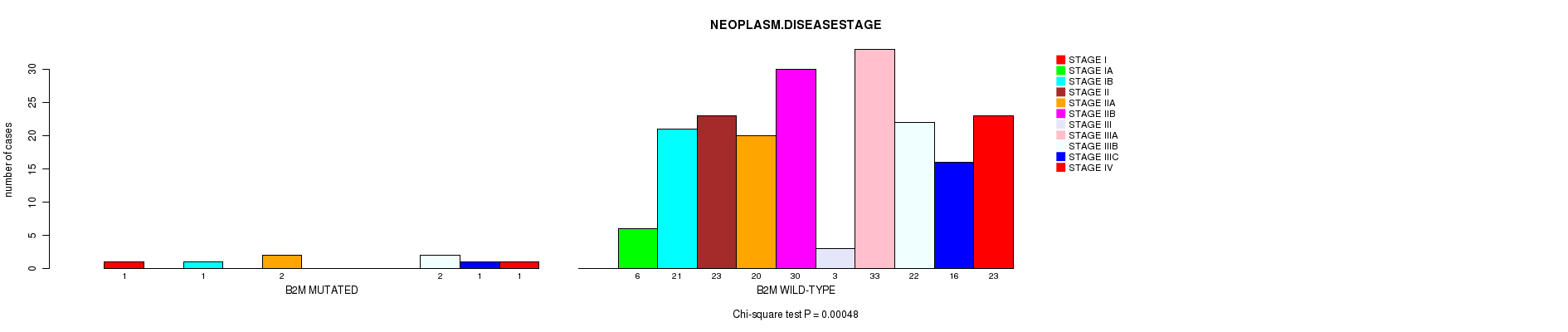
B2M mutation correlated to 'NEOPLASM.DISEASESTAGE'.
-
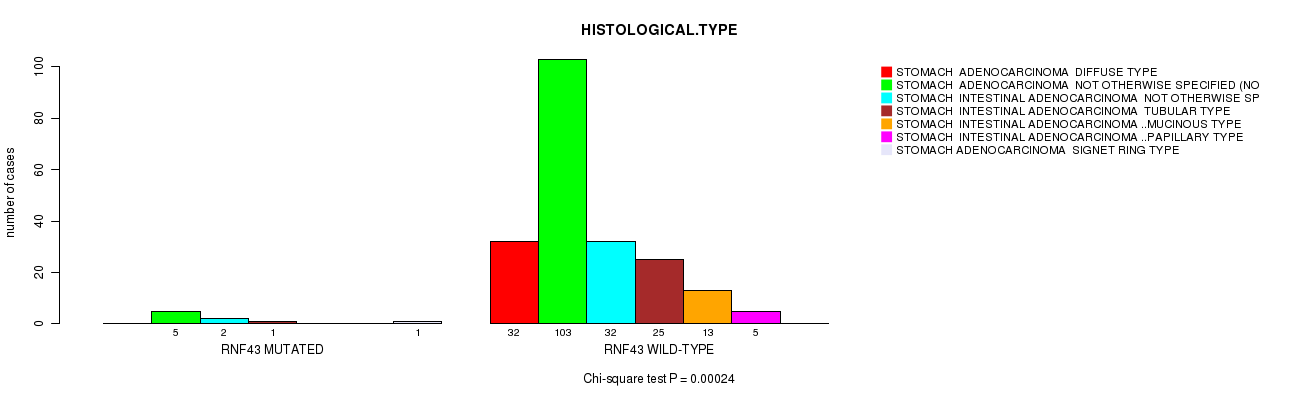
RNF43 mutation correlated to 'HISTOLOGICAL.TYPE'.
-
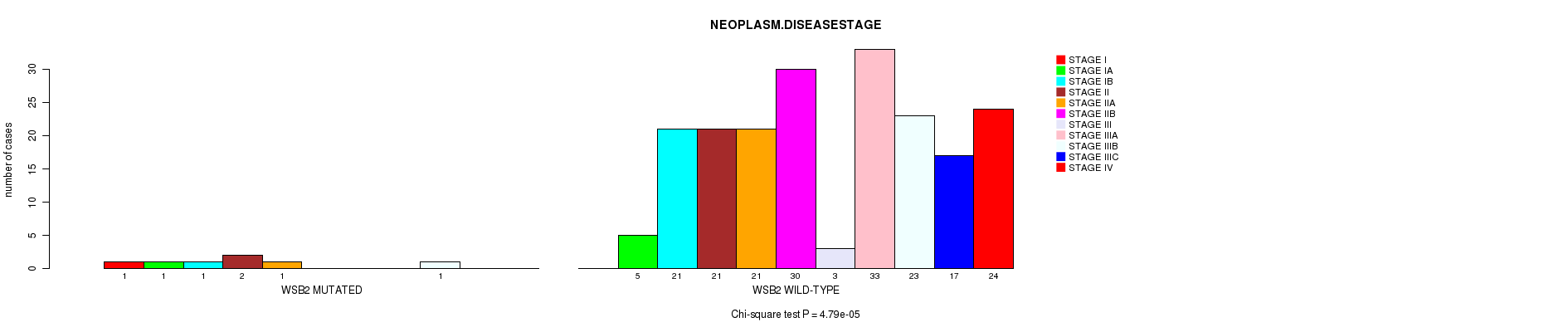
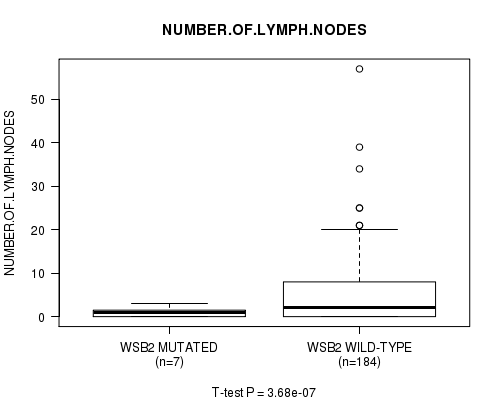
WSB2 mutation correlated to 'NEOPLASM.DISEASESTAGE' and 'NUMBER.OF.LYMPH.NODES'.
-
C13ORF33 mutation correlated to 'NEOPLASM.DISEASESTAGE'.
Table 1. Get Full Table Overview of the association between mutation status of 20 genes and 11 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 5 significant findings detected.
|
Clinical Features |
Time to Death |
AGE |
NEOPLASM DISEASESTAGE |
PATHOLOGY T STAGE |
PATHOLOGY N STAGE |
PATHOLOGY M STAGE |
GENDER |
HISTOLOGICAL TYPE |
RADIATIONS RADIATION REGIMENINDICATION |
COMPLETENESS OF RESECTION |
NUMBER OF LYMPH NODES |
||
| nMutated (%) | nWild-Type | logrank test | t-test | Chi-square test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Chi-square test | Fisher's exact test | Fisher's exact test | t-test | |
| WSB2 | 7 (3%) | 213 |
0.188 (1.00) |
0.0174 (1.00) |
4.79e-05 (0.0105) |
0.0133 (1.00) |
0.278 (1.00) |
0.347 (1.00) |
0.247 (1.00) |
0.251 (1.00) |
1 (1.00) |
0.772 (1.00) |
3.68e-07 (8.1e-05) |
| B2M | 8 (4%) | 212 |
0.317 (1.00) |
0.35 (1.00) |
0.00048 (0.104) |
0.282 (1.00) |
0.735 (1.00) |
0.395 (1.00) |
0.716 (1.00) |
0.491 (1.00) |
1 (1.00) |
0.772 (1.00) |
0.506 (1.00) |
| RNF43 | 9 (4%) | 211 |
0.166 (1.00) |
0.272 (1.00) |
0.346 (1.00) |
0.0917 (1.00) |
0.156 (1.00) |
0.441 (1.00) |
0.161 (1.00) |
0.00024 (0.0523) |
1 (1.00) |
0.199 (1.00) |
0.49 (1.00) |
| C13ORF33 | 6 (3%) | 214 |
0.815 (1.00) |
0.363 (1.00) |
0.000875 (0.189) |
0.133 (1.00) |
0.0478 (1.00) |
1 (1.00) |
0.685 (1.00) |
0.873 (1.00) |
0.155 (1.00) |
1 (1.00) |
0.198 (1.00) |
| PIK3CA | 48 (22%) | 172 |
0.724 (1.00) |
0.469 (1.00) |
0.252 (1.00) |
0.467 (1.00) |
0.9 (1.00) |
0.337 (1.00) |
0.868 (1.00) |
0.873 (1.00) |
1 (1.00) |
0.438 (1.00) |
0.226 (1.00) |
| PGM5 | 22 (10%) | 198 |
0.267 (1.00) |
0.0223 (1.00) |
0.0393 (1.00) |
0.0154 (1.00) |
0.884 (1.00) |
0.867 (1.00) |
0.00577 (1.00) |
0.924 (1.00) |
0.473 (1.00) |
0.667 (1.00) |
0.986 (1.00) |
| KRAS | 25 (11%) | 195 |
0.938 (1.00) |
0.458 (1.00) |
0.0636 (1.00) |
0.748 (1.00) |
0.361 (1.00) |
1 (1.00) |
0.0491 (1.00) |
0.517 (1.00) |
1 (1.00) |
0.0307 (1.00) |
0.0772 (1.00) |
| CBWD1 | 28 (13%) | 192 |
0.2 (1.00) |
0.0241 (1.00) |
0.414 (1.00) |
0.236 (1.00) |
0.582 (1.00) |
0.886 (1.00) |
0.0225 (1.00) |
0.0606 (1.00) |
0.563 (1.00) |
0.844 (1.00) |
0.066 (1.00) |
| TP53 | 99 (45%) | 121 |
0.513 (1.00) |
0.69 (1.00) |
0.83 (1.00) |
0.483 (1.00) |
0.637 (1.00) |
0.0269 (1.00) |
0.68 (1.00) |
0.197 (1.00) |
0.412 (1.00) |
0.247 (1.00) |
0.639 (1.00) |
| ARID1A | 41 (19%) | 179 |
0.411 (1.00) |
0.299 (1.00) |
0.207 (1.00) |
0.114 (1.00) |
0.323 (1.00) |
0.0398 (1.00) |
0.381 (1.00) |
0.244 (1.00) |
0.596 (1.00) |
0.115 (1.00) |
0.263 (1.00) |
| SMAD4 | 19 (9%) | 201 |
0.351 (1.00) |
0.385 (1.00) |
0.654 (1.00) |
0.958 (1.00) |
0.738 (1.00) |
0.606 (1.00) |
0.475 (1.00) |
0.825 (1.00) |
1 (1.00) |
0.472 (1.00) |
0.438 (1.00) |
| RHOA | 13 (6%) | 207 |
0.737 (1.00) |
0.297 (1.00) |
0.783 (1.00) |
0.869 (1.00) |
0.48 (1.00) |
1 (1.00) |
0.57 (1.00) |
0.422 (1.00) |
1 (1.00) |
0.783 (1.00) |
0.268 (1.00) |
| IRF2 | 14 (6%) | 206 |
0.404 (1.00) |
0.591 (1.00) |
0.215 (1.00) |
0.179 (1.00) |
0.156 (1.00) |
0.637 (1.00) |
1 (1.00) |
0.793 (1.00) |
1 (1.00) |
0.022 (1.00) |
0.00134 (0.288) |
| CDH1 | 18 (8%) | 202 |
0.128 (1.00) |
0.115 (1.00) |
0.5 (1.00) |
0.418 (1.00) |
0.548 (1.00) |
0.7 (1.00) |
0.623 (1.00) |
0.134 (1.00) |
1 (1.00) |
0.212 (1.00) |
0.386 (1.00) |
| PTEN | 14 (6%) | 206 |
0.154 (1.00) |
0.0483 (1.00) |
0.338 (1.00) |
0.869 (1.00) |
0.436 (1.00) |
0.494 (1.00) |
0.0213 (1.00) |
0.724 (1.00) |
1 (1.00) |
0.0629 (1.00) |
0.249 (1.00) |
| FBXW7 | 19 (9%) | 201 |
0.0889 (1.00) |
0.304 (1.00) |
0.734 (1.00) |
0.104 (1.00) |
0.965 (1.00) |
0.856 (1.00) |
1 (1.00) |
0.56 (1.00) |
0.422 (1.00) |
0.391 (1.00) |
0.0394 (1.00) |
| FAM46D | 6 (3%) | 214 |
0.111 (1.00) |
0.562 (1.00) |
0.804 (1.00) |
0.353 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.995 (1.00) |
1 (1.00) |
1 (1.00) |
0.645 (1.00) |
| APC | 33 (15%) | 187 |
0.195 (1.00) |
0.0486 (1.00) |
0.184 (1.00) |
0.398 (1.00) |
0.348 (1.00) |
0.664 (1.00) |
0.703 (1.00) |
0.507 (1.00) |
0.595 (1.00) |
0.326 (1.00) |
0.722 (1.00) |
| MAP2K7 | 14 (6%) | 206 |
0.836 (1.00) |
0.116 (1.00) |
0.826 (1.00) |
0.142 (1.00) |
0.319 (1.00) |
1 (1.00) |
0.574 (1.00) |
0.816 (1.00) |
1 (1.00) |
0.695 (1.00) |
0.758 (1.00) |
| TRPS1 | 30 (14%) | 190 |
0.38 (1.00) |
0.37 (1.00) |
0.163 (1.00) |
0.955 (1.00) |
0.678 (1.00) |
0.642 (1.00) |
0.43 (1.00) |
0.835 (1.00) |
1 (1.00) |
0.445 (1.00) |
0.924 (1.00) |
P value = 0.00048 (Chi-square test), Q value = 0.1
Table S1. Gene #13: 'B2M MUTATION STATUS' versus Clinical Feature #3: 'NEOPLASM.DISEASESTAGE'
| nPatients | STAGE I | STAGE IA | STAGE IB | STAGE II | STAGE IIA | STAGE IIB | STAGE III | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 1 | 6 | 22 | 23 | 22 | 30 | 3 | 33 | 24 | 17 | 24 |
| B2M MUTATED | 1 | 0 | 1 | 0 | 2 | 0 | 0 | 0 | 2 | 1 | 1 |
| B2M WILD-TYPE | 0 | 6 | 21 | 23 | 20 | 30 | 3 | 33 | 22 | 16 | 23 |
Figure S1. Get High-res Image Gene #13: 'B2M MUTATION STATUS' versus Clinical Feature #3: 'NEOPLASM.DISEASESTAGE'
P value = 0.00024 (Chi-square test), Q value = 0.052
Table S2. Gene #15: 'RNF43 MUTATION STATUS' versus Clinical Feature #8: 'HISTOLOGICAL.TYPE'
| nPatients | STOMACH ADENOCARCINOMA DIFFUSE TYPE | STOMACH ADENOCARCINOMA NOT OTHERWISE SPECIFIED (NOS) | STOMACH INTESTINAL ADENOCARCINOMA NOT OTHERWISE SPECIFIED (NOS) | STOMACH INTESTINAL ADENOCARCINOMA TUBULAR TYPE | STOMACH INTESTINAL ADENOCARCINOMA MUCINOUS TYPE | STOMACH INTESTINAL ADENOCARCINOMA PAPILLARY TYPE | STOMACH ADENOCARCINOMA SIGNET RING TYPE |
|---|---|---|---|---|---|---|---|
| ALL | 32 | 108 | 34 | 26 | 13 | 5 | 1 |
| RNF43 MUTATED | 0 | 5 | 2 | 1 | 0 | 0 | 1 |
| RNF43 WILD-TYPE | 32 | 103 | 32 | 25 | 13 | 5 | 0 |
Figure S2. Get High-res Image Gene #15: 'RNF43 MUTATION STATUS' versus Clinical Feature #8: 'HISTOLOGICAL.TYPE'
P value = 4.79e-05 (Chi-square test), Q value = 0.01
Table S3. Gene #17: 'WSB2 MUTATION STATUS' versus Clinical Feature #3: 'NEOPLASM.DISEASESTAGE'
| nPatients | STAGE I | STAGE IA | STAGE IB | STAGE II | STAGE IIA | STAGE IIB | STAGE III | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 1 | 6 | 22 | 23 | 22 | 30 | 3 | 33 | 24 | 17 | 24 |
| WSB2 MUTATED | 1 | 1 | 1 | 2 | 1 | 0 | 0 | 0 | 1 | 0 | 0 |
| WSB2 WILD-TYPE | 0 | 5 | 21 | 21 | 21 | 30 | 3 | 33 | 23 | 17 | 24 |
Figure S3. Get High-res Image Gene #17: 'WSB2 MUTATION STATUS' versus Clinical Feature #3: 'NEOPLASM.DISEASESTAGE'
P value = 3.68e-07 (t-test), Q value = 8.1e-05
Table S4. Gene #17: 'WSB2 MUTATION STATUS' versus Clinical Feature #11: 'NUMBER.OF.LYMPH.NODES'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 191 | 5.2 (7.5) |
| WSB2 MUTATED | 7 | 1.0 (1.2) |
| WSB2 WILD-TYPE | 184 | 5.3 (7.6) |
Figure S4. Get High-res Image Gene #17: 'WSB2 MUTATION STATUS' versus Clinical Feature #11: 'NUMBER.OF.LYMPH.NODES'
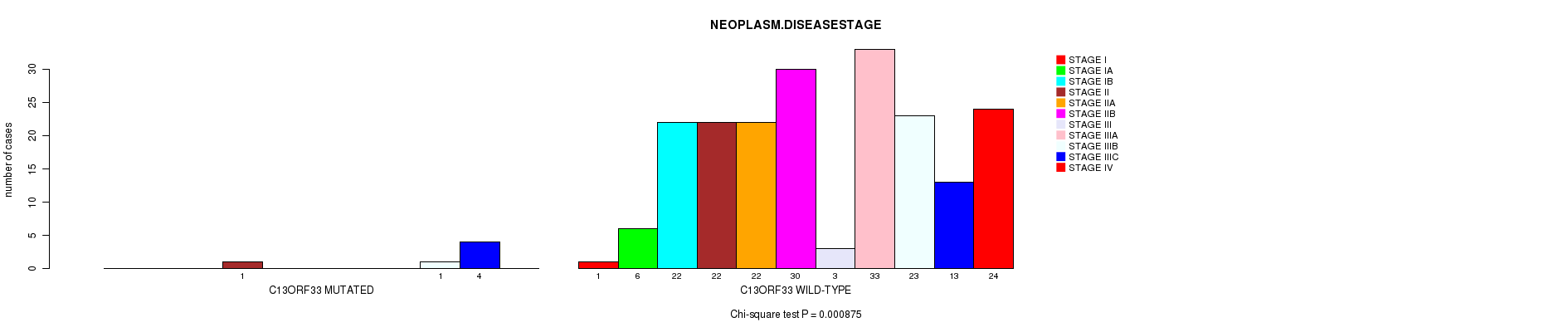
P value = 0.000875 (Chi-square test), Q value = 0.19
Table S5. Gene #20: 'C13ORF33 MUTATION STATUS' versus Clinical Feature #3: 'NEOPLASM.DISEASESTAGE'
| nPatients | STAGE I | STAGE IA | STAGE IB | STAGE II | STAGE IIA | STAGE IIB | STAGE III | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 1 | 6 | 22 | 23 | 22 | 30 | 3 | 33 | 24 | 17 | 24 |
| C13ORF33 MUTATED | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 4 | 0 |
| C13ORF33 WILD-TYPE | 1 | 6 | 22 | 22 | 22 | 30 | 3 | 33 | 23 | 13 | 24 |
Figure S5. Get High-res Image Gene #20: 'C13ORF33 MUTATION STATUS' versus Clinical Feature #3: 'NEOPLASM.DISEASESTAGE'
-
Mutation data file = transformed.cor.cli.txt
-
Clinical data file = STAD-TP.merged_data.txt
-
Number of patients = 220
-
Number of significantly mutated genes = 20
-
Number of selected clinical features = 11
-
Exclude genes that fewer than K tumors have mutations, K = 3
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For continuous numerical clinical features, two-tailed Student's t test with unequal variance (Lehmann and Romano 2005) was applied to compare the clinical values between tumors with and without gene mutations using 't.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.