This pipeline computes the correlation between significantly recurrent gene mutations and selected clinical features.
Testing the association between mutation status of 34 genes and 13 clinical features across 247 patients, 2 significant findings detected with Q value < 0.25.
-
TP53 mutation correlated to 'GLEASON_SCORE_COMBINED' and 'GLEASON_SCORE'.
Table 1. Get Full Table Overview of the association between mutation status of 34 genes and 13 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 2 significant findings detected.
|
Clinical Features |
AGE |
PATHOLOGY T STAGE |
PATHOLOGY N STAGE |
HISTOLOGICAL TYPE |
COMPLETENESS OF RESECTION |
NUMBER OF LYMPH NODES |
GLEASON SCORE COMBINED |
GLEASON SCORE PRIMARY |
GLEASON SCORE SECONDARY |
GLEASON SCORE |
PSA RESULT PREOP |
PSA VALUE |
RACE | ||
| nMutated (%) | nWild-Type | Wilcoxon-test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Wilcoxon-test | Wilcoxon-test | Wilcoxon-test | Wilcoxon-test | Wilcoxon-test | Wilcoxon-test | Wilcoxon-test | Fisher's exact test | |
| TP53 | 22 (9%) | 225 |
0.711 (1.00) |
0.29 (1.00) |
0.713 (1.00) |
1 (1.00) |
0.819 (1.00) |
0.674 (1.00) |
9.8e-05 (0.0412) |
0.00336 (1.00) |
0.041 (1.00) |
3.61e-05 (0.0152) |
0.328 (1.00) |
0.0882 (1.00) |
1 (1.00) |
| SPOP | 25 (10%) | 222 |
0.147 (1.00) |
0.81 (1.00) |
1 (1.00) |
0.0814 (1.00) |
0.621 (1.00) |
0.87 (1.00) |
0.833 (1.00) |
0.408 (1.00) |
0.568 (1.00) |
0.943 (1.00) |
0.902 (1.00) |
0.736 (1.00) |
0.548 (1.00) |
| PTEN | 11 (4%) | 236 |
0.627 (1.00) |
0.808 (1.00) |
1 (1.00) |
0.205 (1.00) |
1 (1.00) |
0.902 (1.00) |
0.53 (1.00) |
0.892 (1.00) |
0.603 (1.00) |
0.669 (1.00) |
0.603 (1.00) |
0.00795 (1.00) |
1 (1.00) |
| CTNNB1 | 9 (4%) | 238 |
0.508 (1.00) |
0.428 (1.00) |
1 (1.00) |
0.171 (1.00) |
0.765 (1.00) |
0.319 (1.00) |
0.439 (1.00) |
0.854 (1.00) |
0.358 (1.00) |
0.276 (1.00) |
0.0875 (1.00) |
0.441 (1.00) |
1 (1.00) |
| TCEB3 | 3 (1%) | 244 |
0.695 (1.00) |
1 (1.00) |
0.29 (1.00) |
1 (1.00) |
0.584 (1.00) |
0.24 (1.00) |
0.535 (1.00) |
0.251 (1.00) |
0.697 (1.00) |
0.601 (1.00) |
0.337 (1.00) |
0.875 (1.00) |
|
| FOXA1 | 11 (4%) | 236 |
0.181 (1.00) |
0.488 (1.00) |
1 (1.00) |
1 (1.00) |
0.412 (1.00) |
0.319 (1.00) |
0.863 (1.00) |
0.889 (1.00) |
0.603 (1.00) |
0.704 (1.00) |
0.981 (1.00) |
0.327 (1.00) |
0.0538 (1.00) |
| BRAF | 5 (2%) | 242 |
0.815 (1.00) |
0.44 (1.00) |
1 (1.00) |
0.098 (1.00) |
1 (1.00) |
0.488 (1.00) |
0.866 (1.00) |
0.239 (1.00) |
0.169 (1.00) |
0.961 (1.00) |
0.374 (1.00) |
0.765 (1.00) |
|
| PIK3CA | 8 (3%) | 239 |
0.266 (1.00) |
0.402 (1.00) |
0.498 (1.00) |
0.153 (1.00) |
0.0267 (1.00) |
0.685 (1.00) |
0.0142 (1.00) |
0.138 (1.00) |
0.068 (1.00) |
0.025 (1.00) |
0.687 (1.00) |
0.441 (1.00) |
1 (1.00) |
| IDH1 | 4 (2%) | 243 |
0.0173 (1.00) |
0.656 (1.00) |
0.367 (1.00) |
1 (1.00) |
0.0811 (1.00) |
0.398 (1.00) |
0.00795 (1.00) |
0.0508 (1.00) |
0.0407 (1.00) |
0.0114 (1.00) |
0.0713 (1.00) |
0.339 (1.00) |
|
| FAM47C | 8 (3%) | 239 |
0.547 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.761 (1.00) |
0.392 (1.00) |
0.117 (1.00) |
0.902 (1.00) |
0.0386 (1.00) |
0.267 (1.00) |
0.711 (1.00) |
0.864 (1.00) |
1 (1.00) |
| TP53BP1 | 4 (2%) | 243 |
0.28 (1.00) |
0.654 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.551 (1.00) |
0.236 (1.00) |
0.135 (1.00) |
0.735 (1.00) |
0.298 (1.00) |
0.929 (1.00) |
0.8 (1.00) |
|
| CNTNAP1 | 5 (2%) | 242 |
0.656 (1.00) |
1 (1.00) |
1 (1.00) |
0.098 (1.00) |
0.32 (1.00) |
0.551 (1.00) |
0.101 (1.00) |
0.746 (1.00) |
0.0468 (1.00) |
0.431 (1.00) |
0.762 (1.00) |
0.815 (1.00) |
|
| MLLT10 | 4 (2%) | 243 |
0.164 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.634 (1.00) |
0.488 (1.00) |
0.716 (1.00) |
0.688 (1.00) |
0.497 (1.00) |
0.799 (1.00) |
0.677 (1.00) |
0.124 (1.00) |
1 (1.00) |
| ZMYM3 | 4 (2%) | 243 |
0.816 (1.00) |
0.195 (1.00) |
0.367 (1.00) |
1 (1.00) |
0.082 (1.00) |
0.398 (1.00) |
0.0124 (1.00) |
0.0467 (1.00) |
0.0606 (1.00) |
0.0178 (1.00) |
0.106 (1.00) |
0.122 (1.00) |
1 (1.00) |
| SMG7 | 4 (2%) | 243 |
0.665 (1.00) |
0.655 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.488 (1.00) |
0.716 (1.00) |
0.933 (1.00) |
0.611 (1.00) |
0.799 (1.00) |
0.251 (1.00) |
0.22 (1.00) |
1 (1.00) |
| EDC4 | 3 (1%) | 244 |
0.329 (1.00) |
1 (1.00) |
0.29 (1.00) |
1 (1.00) |
1 (1.00) |
0.24 (1.00) |
0.548 (1.00) |
0.64 (1.00) |
0.697 (1.00) |
0.493 (1.00) |
0.174 (1.00) |
0.459 (1.00) |
0.163 (1.00) |
| ZNF709 | 4 (2%) | 243 |
0.486 (1.00) |
0.197 (1.00) |
1 (1.00) |
1 (1.00) |
0.118 (1.00) |
0.488 (1.00) |
0.236 (1.00) |
0.0467 (1.00) |
0.735 (1.00) |
0.19 (1.00) |
0.481 (1.00) |
0.637 (1.00) |
|
| CDK12 | 7 (3%) | 240 |
0.707 (1.00) |
1 (1.00) |
0.498 (1.00) |
0.135 (1.00) |
0.37 (1.00) |
0.601 (1.00) |
0.0527 (1.00) |
0.23 (1.00) |
0.243 (1.00) |
0.0821 (1.00) |
0.36 (1.00) |
0.728 (1.00) |
|
| RPTN | 6 (2%) | 241 |
0.362 (1.00) |
0.49 (1.00) |
1 (1.00) |
1 (1.00) |
0.122 (1.00) |
0.436 (1.00) |
0.175 (1.00) |
0.375 (1.00) |
0.486 (1.00) |
0.164 (1.00) |
0.558 (1.00) |
0.128 (1.00) |
0.163 (1.00) |
| MED15 | 5 (2%) | 242 |
0.534 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.436 (1.00) |
0.0456 (1.00) |
0.074 (1.00) |
0.222 (1.00) |
0.0647 (1.00) |
0.409 (1.00) |
0.23 (1.00) |
0.16 (1.00) |
| NKX3-1 | 5 (2%) | 242 |
0.396 (1.00) |
0.266 (1.00) |
1 (1.00) |
1 (1.00) |
0.655 (1.00) |
0.436 (1.00) |
0.435 (1.00) |
0.601 (1.00) |
0.126 (1.00) |
0.118 (1.00) |
0.738 (1.00) |
0.794 (1.00) |
1 (1.00) |
| KDM6A | 5 (2%) | 242 |
0.635 (1.00) |
0.0338 (1.00) |
1 (1.00) |
1 (1.00) |
0.63 (1.00) |
0.436 (1.00) |
0.435 (1.00) |
0.0422 (1.00) |
0.446 (1.00) |
0.852 (1.00) |
1 (1.00) |
0.374 (1.00) |
1 (1.00) |
| RNF31 | 4 (2%) | 243 |
0.461 (1.00) |
0.657 (1.00) |
1 (1.00) |
1 (1.00) |
0.12 (1.00) |
0.488 (1.00) |
0.152 (1.00) |
0.3 (1.00) |
0.233 (1.00) |
0.19 (1.00) |
0.664 (1.00) |
0.765 (1.00) |
|
| ESCO1 | 4 (2%) | 243 |
0.293 (1.00) |
0.655 (1.00) |
1 (1.00) |
1 (1.00) |
0.583 (1.00) |
0.488 (1.00) |
0.716 (1.00) |
0.39 (1.00) |
0.173 (1.00) |
0.799 (1.00) |
0.545 (1.00) |
0.408 (1.00) |
1 (1.00) |
| LMOD2 | 3 (1%) | 244 |
0.28 (1.00) |
0.621 (1.00) |
1 (1.00) |
1 (1.00) |
0.564 (1.00) |
0.64 (1.00) |
0.697 (1.00) |
0.52 (1.00) |
0.0897 (1.00) |
0.356 (1.00) |
|||
| FAM120B | 3 (1%) | 244 |
0.977 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.548 (1.00) |
0.535 (1.00) |
0.169 (1.00) |
0.493 (1.00) |
0.703 (1.00) |
0.765 (1.00) |
1 (1.00) |
|
| CLEC1A | 4 (2%) | 243 |
0.699 (1.00) |
0.0796 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.551 (1.00) |
0.135 (1.00) |
0.39 (1.00) |
0.307 (1.00) |
0.122 (1.00) |
0.969 (1.00) |
0.143 (1.00) |
|
| RNF17 | 3 (1%) | 244 |
0.412 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.551 (1.00) |
0.548 (1.00) |
0.118 (1.00) |
0.558 (1.00) |
0.493 (1.00) |
0.519 (1.00) |
1 (1.00) |
|
| RP1L1 | 6 (2%) | 241 |
0.345 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.168 (1.00) |
0.436 (1.00) |
0.375 (1.00) |
0.916 (1.00) |
0.208 (1.00) |
0.894 (1.00) |
0.119 (1.00) |
0.693 (1.00) |
|
| NTM | 5 (2%) | 242 |
0.412 (1.00) |
0.439 (1.00) |
1 (1.00) |
1 (1.00) |
0.425 (1.00) |
0.436 (1.00) |
0.682 (1.00) |
0.601 (1.00) |
0.648 (1.00) |
0.745 (1.00) |
0.188 (1.00) |
0.301 (1.00) |
1 (1.00) |
| FNBP4 | 3 (1%) | 244 |
0.723 (1.00) |
0.297 (1.00) |
0.29 (1.00) |
1 (1.00) |
0.214 (1.00) |
0.24 (1.00) |
0.425 (1.00) |
0.251 (1.00) |
0.697 (1.00) |
0.478 (1.00) |
0.133 (1.00) |
||
| CDKN1B | 4 (2%) | 243 |
0.22 (1.00) |
0.0791 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.488 (1.00) |
0.236 (1.00) |
0.3 (1.00) |
0.611 (1.00) |
0.0359 (1.00) |
0.972 (1.00) |
0.656 (1.00) |
1 (1.00) |
| EIF2AK3 | 4 (2%) | 243 |
0.415 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.12 (1.00) |
0.908 (1.00) |
0.933 (1.00) |
0.867 (1.00) |
0.799 (1.00) |
0.803 (1.00) |
0.98 (1.00) |
1 (1.00) |
|
| LARP1 | 3 (1%) | 244 |
0.52 (1.00) |
1 (1.00) |
0.29 (1.00) |
1 (1.00) |
0.16 (1.00) |
0.564 (1.00) |
0.64 (1.00) |
0.697 (1.00) |
0.52 (1.00) |
0.87 (1.00) |
0.891 (1.00) |
P value = 9.8e-05 (Wilcoxon-test), Q value = 0.041
Table S1. Gene #2: 'TP53 MUTATION STATUS' versus Clinical Feature #7: 'GLEASON_SCORE_COMBINED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 247 | 7.3 (0.8) |
| TP53 MUTATED | 22 | 8.0 (0.9) |
| TP53 WILD-TYPE | 225 | 7.3 (0.8) |
Figure S1. Get High-res Image Gene #2: 'TP53 MUTATION STATUS' versus Clinical Feature #7: 'GLEASON_SCORE_COMBINED'
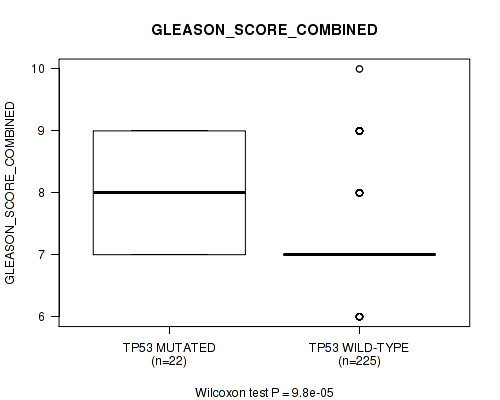
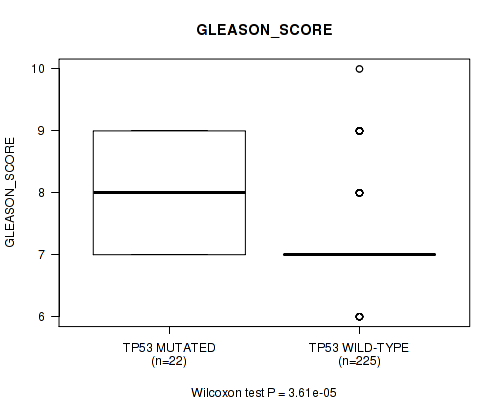
P value = 3.61e-05 (Wilcoxon-test), Q value = 0.015
Table S2. Gene #2: 'TP53 MUTATION STATUS' versus Clinical Feature #10: 'GLEASON_SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 247 | 7.4 (0.9) |
| TP53 MUTATED | 22 | 8.1 (0.9) |
| TP53 WILD-TYPE | 225 | 7.3 (0.8) |
Figure S2. Get High-res Image Gene #2: 'TP53 MUTATION STATUS' versus Clinical Feature #10: 'GLEASON_SCORE'
-
Mutation data file = transformed.cor.cli.txt
-
Clinical data file = PRAD-TP.merged_data.txt
-
Number of patients = 247
-
Number of significantly mutated genes = 34
-
Number of selected clinical features = 13
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.