This pipeline computes the correlation between significantly recurrent gene mutations and molecular subtypes.
Testing the association between mutation status of 19 genes and 10 molecular subtypes across 161 patients, 2 significant findings detected with P value < 0.05 and Q value < 0.25.
-
MET mutation correlated to 'MRNASEQ_CHIERARCHICAL'.
-
SETD2 mutation correlated to 'METHLYATION_CNMF'.
Table 1. Get Full Table Overview of the association between mutation status of 19 genes and 10 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 2 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
RPPA CNMF |
RPPA CHIERARCHICAL |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nMutated (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| MET | 15 (9%) | 146 |
0.198 (1.00) |
0.0895 (1.00) |
0.297 (1.00) |
0.0374 (1.00) |
0.108 (1.00) |
0.00033 (0.0604) |
0.838 (1.00) |
0.815 (1.00) |
0.642 (1.00) |
0.141 (1.00) |
| SETD2 | 10 (6%) | 151 |
0.132 (1.00) |
6e-05 (0.011) |
0.193 (1.00) |
0.526 (1.00) |
0.00764 (1.00) |
0.0332 (1.00) |
0.0177 (1.00) |
0.0202 (1.00) |
0.0728 (1.00) |
0.343 (1.00) |
| HNRNPM | 10 (6%) | 151 |
0.0749 (1.00) |
0.176 (1.00) |
1 (1.00) |
0.263 (1.00) |
0.507 (1.00) |
0.573 (1.00) |
0.713 (1.00) |
0.317 (1.00) |
0.35 (1.00) |
0.513 (1.00) |
| NEFH | 10 (6%) | 151 |
0.432 (1.00) |
0.91 (1.00) |
0.427 (1.00) |
0.346 (1.00) |
0.222 (1.00) |
0.661 (1.00) |
0.0147 (1.00) |
0.03 (1.00) |
0.197 (1.00) |
0.328 (1.00) |
| ZNF598 | 10 (6%) | 151 |
0.513 (1.00) |
0.831 (1.00) |
0.512 (1.00) |
0.26 (1.00) |
0.92 (1.00) |
1 (1.00) |
0.394 (1.00) |
0.238 (1.00) |
0.102 (1.00) |
0.119 (1.00) |
| NF2 | 10 (6%) | 151 |
0.132 (1.00) |
0.0241 (1.00) |
0.828 (1.00) |
0.577 (1.00) |
0.0963 (1.00) |
0.345 (1.00) |
0.655 (1.00) |
0.55 (1.00) |
0.493 (1.00) |
0.0385 (1.00) |
| TDG | 5 (3%) | 156 |
0.101 (1.00) |
0.276 (1.00) |
0.634 (1.00) |
0.369 (1.00) |
0.74 (1.00) |
0.666 (1.00) |
0.439 (1.00) |
0.279 (1.00) |
0.444 (1.00) |
0.91 (1.00) |
| SKI | 6 (4%) | 155 |
0.108 (1.00) |
0.0607 (1.00) |
0.738 (1.00) |
0.707 (1.00) |
0.88 (1.00) |
1 (1.00) |
0.511 (1.00) |
0.458 (1.00) |
0.862 (1.00) |
0.373 (1.00) |
| MUC5B | 14 (9%) | 147 |
0.116 (1.00) |
0.537 (1.00) |
0.463 (1.00) |
0.413 (1.00) |
0.883 (1.00) |
0.317 (1.00) |
0.885 (1.00) |
0.288 (1.00) |
0.441 (1.00) |
0.965 (1.00) |
| ZNF814 | 8 (5%) | 153 |
0.685 (1.00) |
0.43 (1.00) |
1 (1.00) |
0.799 (1.00) |
0.899 (1.00) |
0.603 (1.00) |
0.436 (1.00) |
0.642 (1.00) |
0.193 (1.00) |
0.79 (1.00) |
| SMARCB1 | 4 (2%) | 157 |
0.825 (1.00) |
0.216 (1.00) |
1 (1.00) |
0.584 (1.00) |
0.453 (1.00) |
0.413 (1.00) |
0.0352 (1.00) |
0.463 (1.00) |
0.108 (1.00) |
0.33 (1.00) |
| KDM6A | 7 (4%) | 154 |
1 (1.00) |
0.555 (1.00) |
0.632 (1.00) |
0.613 (1.00) |
0.0302 (1.00) |
0.735 (1.00) |
0.387 (1.00) |
0.162 (1.00) |
0.396 (1.00) |
0.22 (1.00) |
| AHNAK2 | 7 (4%) | 154 |
0.951 (1.00) |
0.764 (1.00) |
0.657 (1.00) |
0.973 (1.00) |
0.794 (1.00) |
0.235 (1.00) |
0.295 (1.00) |
0.213 (1.00) |
0.257 (1.00) |
0.3 (1.00) |
| AHCY | 4 (2%) | 157 |
0.0589 (1.00) |
0.473 (1.00) |
0.559 (1.00) |
0.889 (1.00) |
0.261 (1.00) |
0.646 (1.00) |
0.196 (1.00) |
1 (1.00) |
||
| IDUA | 5 (3%) | 156 |
0.29 (1.00) |
0.731 (1.00) |
0.3 (1.00) |
0.459 (1.00) |
0.739 (1.00) |
0.267 (1.00) |
0.444 (1.00) |
0.0702 (1.00) |
0.83 (1.00) |
0.868 (1.00) |
| OR2L8 | 4 (2%) | 157 |
0.391 (1.00) |
0.294 (1.00) |
0.676 (1.00) |
0.196 (1.00) |
0.375 (1.00) |
0.0398 (1.00) |
0.39 (1.00) |
0.508 (1.00) |
0.549 (1.00) |
0.866 (1.00) |
| HOXD8 | 4 (2%) | 157 |
0.181 (1.00) |
0.785 (1.00) |
0.0183 (1.00) |
0.034 (1.00) |
0.375 (1.00) |
0.153 (1.00) |
0.321 (1.00) |
0.834 (1.00) |
0.0173 (1.00) |
0.0106 (1.00) |
| PAM | 3 (2%) | 158 |
0.585 (1.00) |
0.787 (1.00) |
0.115 (1.00) |
0.104 (1.00) |
0.189 (1.00) |
1 (1.00) |
0.392 (1.00) |
0.0291 (1.00) |
||
| CSGALNACT2 | 5 (3%) | 156 |
0.137 (1.00) |
0.327 (1.00) |
0.831 (1.00) |
0.774 (1.00) |
0.741 (1.00) |
0.618 (1.00) |
0.525 (1.00) |
0.728 (1.00) |
P value = 0.0749 (Fisher's exact test), Q value = 1
Table S1. Gene #1: 'HNRNPM MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| HNRNPM MUTATED | 7 | 0 | 2 | 1 |
| HNRNPM WILD-TYPE | 54 | 39 | 23 | 35 |
P value = 0.176 (Fisher's exact test), Q value = 1
Table S2. Gene #1: 'HNRNPM MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| HNRNPM MUTATED | 5 | 3 | 2 |
| HNRNPM WILD-TYPE | 35 | 42 | 61 |
P value = 1 (Fisher's exact test), Q value = 1
Table S3. Gene #1: 'HNRNPM MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| HNRNPM MUTATED | 4 | 4 | 2 |
| HNRNPM WILD-TYPE | 44 | 40 | 30 |
P value = 0.263 (Fisher's exact test), Q value = 1
Table S4. Gene #1: 'HNRNPM MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| HNRNPM MUTATED | 3 | 1 | 1 | 0 | 1 | 1 | 2 | 1 |
| HNRNPM WILD-TYPE | 8 | 11 | 15 | 16 | 16 | 24 | 10 | 14 |
P value = 0.507 (Fisher's exact test), Q value = 1
Table S5. Gene #1: 'HNRNPM MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| HNRNPM MUTATED | 1 | 4 | 5 |
| HNRNPM WILD-TYPE | 42 | 52 | 57 |
P value = 0.573 (Fisher's exact test), Q value = 1
Table S6. Gene #1: 'HNRNPM MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| HNRNPM MUTATED | 5 | 2 | 0 | 3 |
| HNRNPM WILD-TYPE | 68 | 37 | 20 | 26 |
P value = 0.713 (Fisher's exact test), Q value = 1
Table S7. Gene #1: 'HNRNPM MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| HNRNPM MUTATED | 2 | 3 | 5 |
| HNRNPM WILD-TYPE | 45 | 51 | 55 |
P value = 0.317 (Fisher's exact test), Q value = 1
Table S8. Gene #1: 'HNRNPM MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| HNRNPM MUTATED | 0 | 4 | 2 | 4 |
| HNRNPM WILD-TYPE | 30 | 31 | 37 | 53 |
P value = 0.35 (Fisher's exact test), Q value = 1
Table S9. Gene #1: 'HNRNPM MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| HNRNPM MUTATED | 3 | 1 | 5 |
| HNRNPM WILD-TYPE | 34 | 38 | 41 |
P value = 0.513 (Fisher's exact test), Q value = 1
Table S10. Gene #1: 'HNRNPM MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| HNRNPM MUTATED | 4 | 5 | 0 | 0 |
| HNRNPM WILD-TYPE | 43 | 42 | 12 | 16 |
P value = 0.432 (Fisher's exact test), Q value = 1
Table S11. Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| NEFH MUTATED | 3 | 1 | 2 | 4 |
| NEFH WILD-TYPE | 58 | 38 | 23 | 32 |
P value = 0.91 (Fisher's exact test), Q value = 1
Table S12. Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| NEFH MUTATED | 3 | 2 | 4 |
| NEFH WILD-TYPE | 37 | 43 | 59 |
P value = 0.427 (Fisher's exact test), Q value = 1
Table S13. Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| NEFH MUTATED | 3 | 1 | 3 |
| NEFH WILD-TYPE | 45 | 43 | 29 |
P value = 0.346 (Fisher's exact test), Q value = 1
Table S14. Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| NEFH MUTATED | 0 | 1 | 0 | 1 | 0 | 1 | 2 | 2 |
| NEFH WILD-TYPE | 11 | 11 | 16 | 15 | 17 | 24 | 10 | 13 |
P value = 0.222 (Fisher's exact test), Q value = 1
Table S15. Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| NEFH MUTATED | 5 | 3 | 2 |
| NEFH WILD-TYPE | 38 | 53 | 60 |
P value = 0.661 (Fisher's exact test), Q value = 1
Table S16. Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| NEFH MUTATED | 6 | 3 | 0 | 1 |
| NEFH WILD-TYPE | 67 | 36 | 20 | 28 |
P value = 0.0147 (Fisher's exact test), Q value = 1
Table S17. Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| NEFH MUTATED | 5 | 5 | 0 |
| NEFH WILD-TYPE | 42 | 49 | 60 |
Figure S1. Get High-res Image Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
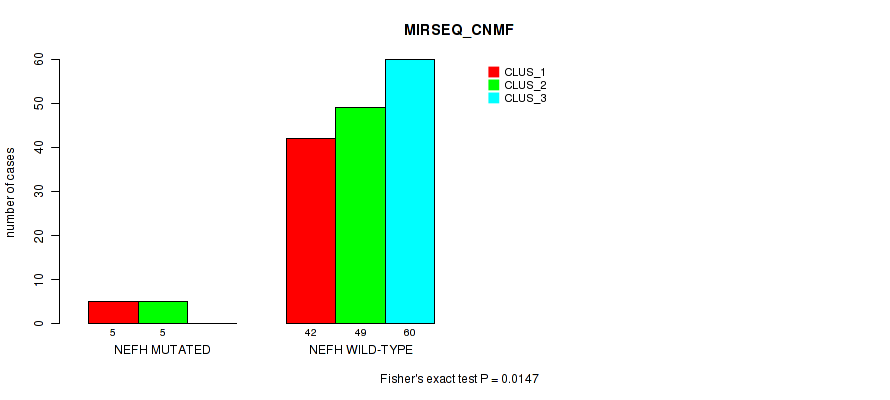
P value = 0.03 (Fisher's exact test), Q value = 1
Table S18. Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| NEFH MUTATED | 3 | 4 | 3 | 0 |
| NEFH WILD-TYPE | 27 | 31 | 36 | 57 |
Figure S2. Get High-res Image Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
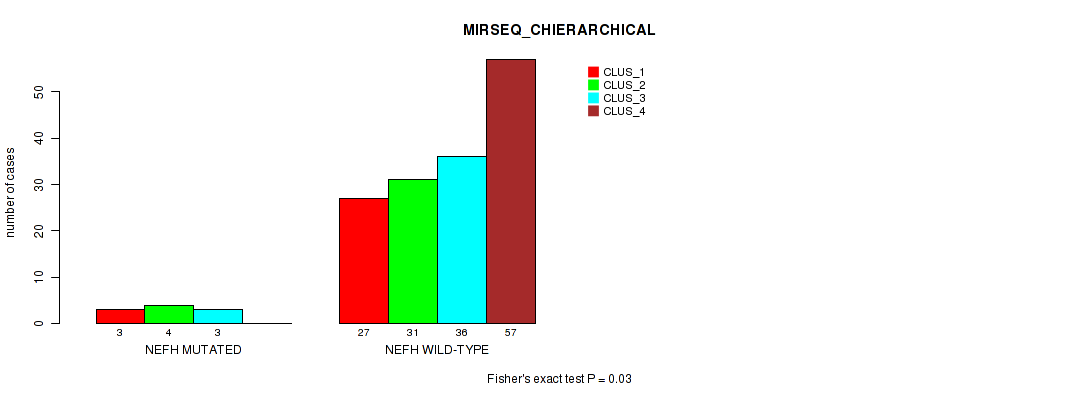
P value = 0.197 (Fisher's exact test), Q value = 1
Table S19. Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| NEFH MUTATED | 2 | 1 | 0 |
| NEFH WILD-TYPE | 35 | 38 | 46 |
P value = 0.328 (Fisher's exact test), Q value = 1
Table S20. Gene #2: 'NEFH MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| NEFH MUTATED | 3 | 0 | 0 | 0 |
| NEFH WILD-TYPE | 44 | 47 | 12 | 16 |
P value = 0.513 (Fisher's exact test), Q value = 1
Table S21. Gene #3: 'ZNF598 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| ZNF598 MUTATED | 4 | 1 | 1 | 4 |
| ZNF598 WILD-TYPE | 57 | 38 | 24 | 32 |
P value = 0.831 (Fisher's exact test), Q value = 1
Table S22. Gene #3: 'ZNF598 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| ZNF598 MUTATED | 3 | 3 | 3 |
| ZNF598 WILD-TYPE | 37 | 42 | 60 |
P value = 0.512 (Fisher's exact test), Q value = 1
Table S23. Gene #3: 'ZNF598 MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| ZNF598 MUTATED | 5 | 3 | 1 |
| ZNF598 WILD-TYPE | 43 | 41 | 31 |
P value = 0.26 (Fisher's exact test), Q value = 1
Table S24. Gene #3: 'ZNF598 MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| ZNF598 MUTATED | 2 | 1 | 1 | 1 | 3 | 0 | 0 | 1 |
| ZNF598 WILD-TYPE | 9 | 11 | 15 | 15 | 14 | 25 | 12 | 14 |
P value = 0.92 (Fisher's exact test), Q value = 1
Table S25. Gene #3: 'ZNF598 MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| ZNF598 MUTATED | 2 | 4 | 4 |
| ZNF598 WILD-TYPE | 41 | 52 | 58 |
P value = 1 (Fisher's exact test), Q value = 1
Table S26. Gene #3: 'ZNF598 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| ZNF598 MUTATED | 5 | 2 | 1 | 2 |
| ZNF598 WILD-TYPE | 68 | 37 | 19 | 27 |
P value = 0.394 (Fisher's exact test), Q value = 1
Table S27. Gene #3: 'ZNF598 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| ZNF598 MUTATED | 2 | 2 | 6 |
| ZNF598 WILD-TYPE | 45 | 52 | 54 |
P value = 0.238 (Fisher's exact test), Q value = 1
Table S28. Gene #3: 'ZNF598 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| ZNF598 MUTATED | 2 | 0 | 2 | 6 |
| ZNF598 WILD-TYPE | 28 | 35 | 37 | 51 |
P value = 0.102 (Fisher's exact test), Q value = 1
Table S29. Gene #3: 'ZNF598 MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| ZNF598 MUTATED | 2 | 1 | 7 |
| ZNF598 WILD-TYPE | 35 | 38 | 39 |
P value = 0.119 (Fisher's exact test), Q value = 1
Table S30. Gene #3: 'ZNF598 MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| ZNF598 MUTATED | 1 | 7 | 1 | 1 |
| ZNF598 WILD-TYPE | 46 | 40 | 11 | 15 |
P value = 0.132 (Fisher's exact test), Q value = 1
Table S31. Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| NF2 MUTATED | 4 | 1 | 0 | 5 |
| NF2 WILD-TYPE | 57 | 38 | 25 | 31 |
P value = 0.0241 (Fisher's exact test), Q value = 1
Table S32. Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| NF2 MUTATED | 0 | 6 | 3 |
| NF2 WILD-TYPE | 40 | 39 | 60 |
Figure S3. Get High-res Image Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
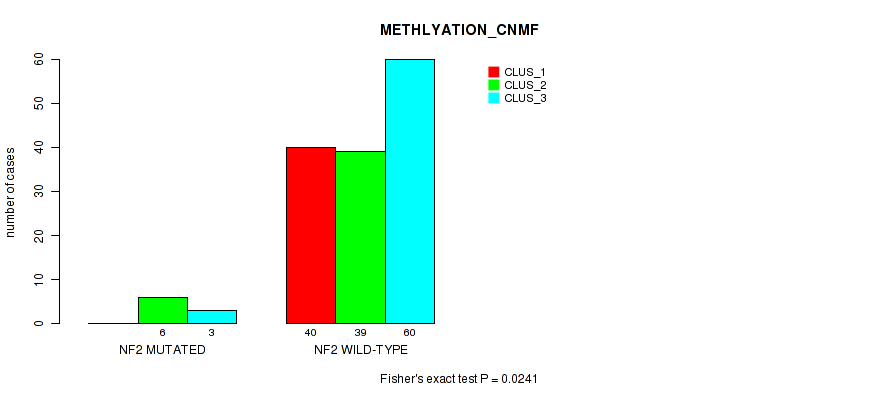
P value = 0.828 (Fisher's exact test), Q value = 1
Table S33. Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| NF2 MUTATED | 3 | 3 | 3 |
| NF2 WILD-TYPE | 45 | 41 | 29 |
P value = 0.577 (Fisher's exact test), Q value = 1
Table S34. Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| NF2 MUTATED | 0 | 1 | 2 | 0 | 3 | 2 | 0 | 1 |
| NF2 WILD-TYPE | 11 | 11 | 14 | 16 | 14 | 23 | 12 | 14 |
P value = 0.0963 (Fisher's exact test), Q value = 1
Table S35. Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| NF2 MUTATED | 0 | 4 | 6 |
| NF2 WILD-TYPE | 43 | 52 | 56 |
P value = 0.345 (Fisher's exact test), Q value = 1
Table S36. Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| NF2 MUTATED | 3 | 2 | 1 | 4 |
| NF2 WILD-TYPE | 70 | 37 | 19 | 25 |
P value = 0.655 (Fisher's exact test), Q value = 1
Table S37. Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| NF2 MUTATED | 3 | 2 | 5 |
| NF2 WILD-TYPE | 44 | 52 | 55 |
P value = 0.55 (Fisher's exact test), Q value = 1
Table S38. Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| NF2 MUTATED | 1 | 3 | 1 | 5 |
| NF2 WILD-TYPE | 29 | 32 | 38 | 52 |
P value = 0.493 (Fisher's exact test), Q value = 1
Table S39. Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| NF2 MUTATED | 2 | 1 | 4 |
| NF2 WILD-TYPE | 35 | 38 | 42 |
P value = 0.0385 (Fisher's exact test), Q value = 1
Table S40. Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| NF2 MUTATED | 3 | 1 | 3 | 0 |
| NF2 WILD-TYPE | 44 | 46 | 9 | 16 |
Figure S4. Get High-res Image Gene #4: 'NF2 MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
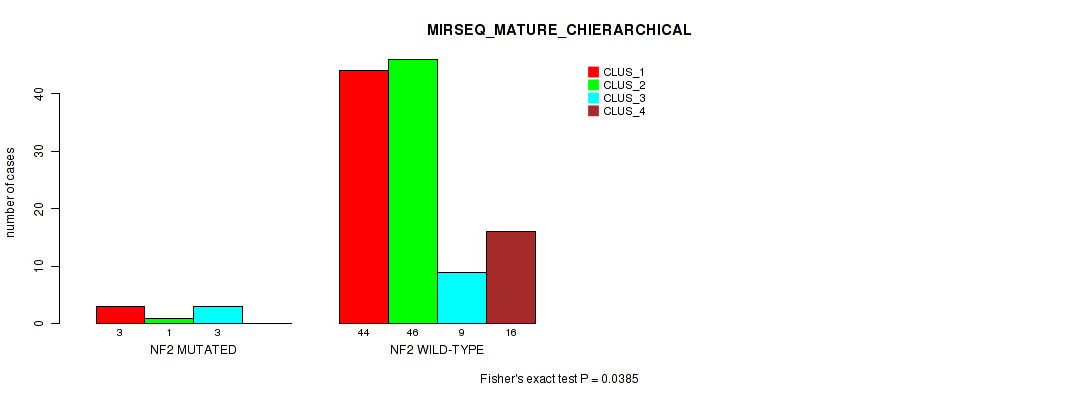
P value = 0.101 (Fisher's exact test), Q value = 1
Table S41. Gene #5: 'TDG MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| TDG MUTATED | 0 | 2 | 2 | 1 |
| TDG WILD-TYPE | 61 | 37 | 23 | 35 |
P value = 0.276 (Fisher's exact test), Q value = 1
Table S42. Gene #5: 'TDG MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| TDG MUTATED | 0 | 3 | 2 |
| TDG WILD-TYPE | 40 | 42 | 61 |
P value = 0.634 (Fisher's exact test), Q value = 1
Table S43. Gene #5: 'TDG MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| TDG MUTATED | 1 | 2 | 2 |
| TDG WILD-TYPE | 47 | 42 | 30 |
P value = 0.369 (Fisher's exact test), Q value = 1
Table S44. Gene #5: 'TDG MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| TDG MUTATED | 0 | 0 | 0 | 2 | 1 | 0 | 1 | 1 |
| TDG WILD-TYPE | 11 | 12 | 16 | 14 | 16 | 25 | 11 | 14 |
P value = 0.74 (Fisher's exact test), Q value = 1
Table S45. Gene #5: 'TDG MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| TDG MUTATED | 1 | 1 | 3 |
| TDG WILD-TYPE | 42 | 55 | 59 |
P value = 0.666 (Fisher's exact test), Q value = 1
Table S46. Gene #5: 'TDG MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| TDG MUTATED | 4 | 1 | 0 | 0 |
| TDG WILD-TYPE | 69 | 38 | 20 | 29 |
P value = 0.439 (Fisher's exact test), Q value = 1
Table S47. Gene #5: 'TDG MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| TDG MUTATED | 3 | 1 | 1 |
| TDG WILD-TYPE | 44 | 53 | 59 |
P value = 0.279 (Fisher's exact test), Q value = 1
Table S48. Gene #5: 'TDG MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| TDG MUTATED | 2 | 2 | 0 | 1 |
| TDG WILD-TYPE | 28 | 33 | 39 | 56 |
P value = 0.444 (Fisher's exact test), Q value = 1
Table S49. Gene #5: 'TDG MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| TDG MUTATED | 3 | 1 | 1 |
| TDG WILD-TYPE | 34 | 38 | 45 |
P value = 0.91 (Fisher's exact test), Q value = 1
Table S50. Gene #5: 'TDG MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| TDG MUTATED | 3 | 2 | 0 | 0 |
| TDG WILD-TYPE | 44 | 45 | 12 | 16 |
P value = 0.108 (Fisher's exact test), Q value = 1
Table S51. Gene #6: 'SKI MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| SKI MUTATED | 5 | 0 | 1 | 0 |
| SKI WILD-TYPE | 56 | 39 | 24 | 36 |
P value = 0.0607 (Fisher's exact test), Q value = 1
Table S52. Gene #6: 'SKI MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| SKI MUTATED | 4 | 0 | 2 |
| SKI WILD-TYPE | 36 | 45 | 61 |
P value = 0.738 (Fisher's exact test), Q value = 1
Table S53. Gene #6: 'SKI MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| SKI MUTATED | 3 | 1 | 1 |
| SKI WILD-TYPE | 45 | 43 | 31 |
P value = 0.707 (Fisher's exact test), Q value = 1
Table S54. Gene #6: 'SKI MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| SKI MUTATED | 1 | 1 | 0 | 0 | 1 | 1 | 1 | 0 |
| SKI WILD-TYPE | 10 | 11 | 16 | 16 | 16 | 24 | 11 | 15 |
P value = 0.88 (Fisher's exact test), Q value = 1
Table S55. Gene #6: 'SKI MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| SKI MUTATED | 1 | 2 | 3 |
| SKI WILD-TYPE | 42 | 54 | 59 |
P value = 1 (Fisher's exact test), Q value = 1
Table S56. Gene #6: 'SKI MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| SKI MUTATED | 3 | 1 | 1 | 1 |
| SKI WILD-TYPE | 70 | 38 | 19 | 28 |
P value = 0.511 (Fisher's exact test), Q value = 1
Table S57. Gene #6: 'SKI MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| SKI MUTATED | 3 | 1 | 2 |
| SKI WILD-TYPE | 44 | 53 | 58 |
P value = 0.458 (Fisher's exact test), Q value = 1
Table S58. Gene #6: 'SKI MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| SKI MUTATED | 2 | 0 | 1 | 3 |
| SKI WILD-TYPE | 28 | 35 | 38 | 54 |
P value = 0.862 (Fisher's exact test), Q value = 1
Table S59. Gene #6: 'SKI MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| SKI MUTATED | 2 | 1 | 2 |
| SKI WILD-TYPE | 35 | 38 | 44 |
P value = 0.373 (Fisher's exact test), Q value = 1
Table S60. Gene #6: 'SKI MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| SKI MUTATED | 1 | 4 | 0 | 0 |
| SKI WILD-TYPE | 46 | 43 | 12 | 16 |
P value = 0.116 (Fisher's exact test), Q value = 1
Table S61. Gene #7: 'MUC5B MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| MUC5B MUTATED | 2 | 3 | 4 | 5 |
| MUC5B WILD-TYPE | 59 | 36 | 21 | 31 |
P value = 0.537 (Fisher's exact test), Q value = 1
Table S62. Gene #7: 'MUC5B MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| MUC5B MUTATED | 5 | 4 | 4 |
| MUC5B WILD-TYPE | 35 | 41 | 59 |
P value = 0.463 (Fisher's exact test), Q value = 1
Table S63. Gene #7: 'MUC5B MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| MUC5B MUTATED | 2 | 5 | 2 |
| MUC5B WILD-TYPE | 46 | 39 | 30 |
P value = 0.413 (Fisher's exact test), Q value = 1
Table S64. Gene #7: 'MUC5B MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| MUC5B MUTATED | 2 | 1 | 2 | 1 | 0 | 3 | 0 | 0 |
| MUC5B WILD-TYPE | 9 | 11 | 14 | 15 | 17 | 22 | 12 | 15 |
P value = 0.883 (Fisher's exact test), Q value = 1
Table S65. Gene #7: 'MUC5B MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| MUC5B MUTATED | 4 | 4 | 6 |
| MUC5B WILD-TYPE | 39 | 52 | 56 |
P value = 0.317 (Fisher's exact test), Q value = 1
Table S66. Gene #7: 'MUC5B MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| MUC5B MUTATED | 9 | 2 | 0 | 3 |
| MUC5B WILD-TYPE | 64 | 37 | 20 | 26 |
P value = 0.885 (Fisher's exact test), Q value = 1
Table S67. Gene #7: 'MUC5B MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| MUC5B MUTATED | 5 | 4 | 5 |
| MUC5B WILD-TYPE | 42 | 50 | 55 |
P value = 0.288 (Fisher's exact test), Q value = 1
Table S68. Gene #7: 'MUC5B MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| MUC5B MUTATED | 4 | 4 | 4 | 2 |
| MUC5B WILD-TYPE | 26 | 31 | 35 | 55 |
P value = 0.441 (Fisher's exact test), Q value = 1
Table S69. Gene #7: 'MUC5B MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| MUC5B MUTATED | 5 | 2 | 5 |
| MUC5B WILD-TYPE | 32 | 37 | 41 |
P value = 0.965 (Fisher's exact test), Q value = 1
Table S70. Gene #7: 'MUC5B MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| MUC5B MUTATED | 4 | 5 | 1 | 2 |
| MUC5B WILD-TYPE | 43 | 42 | 11 | 14 |
P value = 0.685 (Fisher's exact test), Q value = 1
Table S71. Gene #8: 'ZNF814 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| ZNF814 MUTATED | 2 | 2 | 1 | 3 |
| ZNF814 WILD-TYPE | 59 | 37 | 24 | 33 |
P value = 0.43 (Fisher's exact test), Q value = 1
Table S72. Gene #8: 'ZNF814 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| ZNF814 MUTATED | 1 | 4 | 3 |
| ZNF814 WILD-TYPE | 39 | 41 | 60 |
P value = 1 (Fisher's exact test), Q value = 1
Table S73. Gene #8: 'ZNF814 MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| ZNF814 MUTATED | 3 | 2 | 2 |
| ZNF814 WILD-TYPE | 45 | 42 | 30 |
P value = 0.799 (Fisher's exact test), Q value = 1
Table S74. Gene #8: 'ZNF814 MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| ZNF814 MUTATED | 0 | 0 | 1 | 1 | 0 | 2 | 1 | 2 |
| ZNF814 WILD-TYPE | 11 | 12 | 15 | 15 | 17 | 23 | 11 | 13 |
P value = 0.899 (Fisher's exact test), Q value = 1
Table S75. Gene #8: 'ZNF814 MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| ZNF814 MUTATED | 2 | 2 | 4 |
| ZNF814 WILD-TYPE | 41 | 54 | 58 |
P value = 0.603 (Fisher's exact test), Q value = 1
Table S76. Gene #8: 'ZNF814 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| ZNF814 MUTATED | 5 | 2 | 1 | 0 |
| ZNF814 WILD-TYPE | 68 | 37 | 19 | 29 |
P value = 0.436 (Fisher's exact test), Q value = 1
Table S77. Gene #8: 'ZNF814 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| ZNF814 MUTATED | 3 | 1 | 4 |
| ZNF814 WILD-TYPE | 44 | 53 | 56 |
P value = 0.642 (Fisher's exact test), Q value = 1
Table S78. Gene #8: 'ZNF814 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| ZNF814 MUTATED | 2 | 3 | 1 | 2 |
| ZNF814 WILD-TYPE | 28 | 32 | 38 | 55 |
P value = 0.193 (Fisher's exact test), Q value = 1
Table S79. Gene #8: 'ZNF814 MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| ZNF814 MUTATED | 0 | 3 | 4 |
| ZNF814 WILD-TYPE | 37 | 36 | 42 |
P value = 0.79 (Fisher's exact test), Q value = 1
Table S80. Gene #8: 'ZNF814 MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| ZNF814 MUTATED | 2 | 4 | 0 | 1 |
| ZNF814 WILD-TYPE | 45 | 43 | 12 | 15 |
P value = 0.198 (Fisher's exact test), Q value = 1
Table S81. Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| MET MUTATED | 5 | 6 | 0 | 4 |
| MET WILD-TYPE | 56 | 33 | 25 | 32 |
P value = 0.0895 (Fisher's exact test), Q value = 1
Table S82. Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| MET MUTATED | 3 | 1 | 9 |
| MET WILD-TYPE | 37 | 44 | 54 |
P value = 0.297 (Fisher's exact test), Q value = 1
Table S83. Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| MET MUTATED | 7 | 5 | 1 |
| MET WILD-TYPE | 41 | 39 | 31 |
P value = 0.0374 (Fisher's exact test), Q value = 1
Table S84. Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| MET MUTATED | 3 | 0 | 4 | 2 | 2 | 0 | 0 | 2 |
| MET WILD-TYPE | 8 | 12 | 12 | 14 | 15 | 25 | 12 | 13 |
Figure S5. Get High-res Image Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
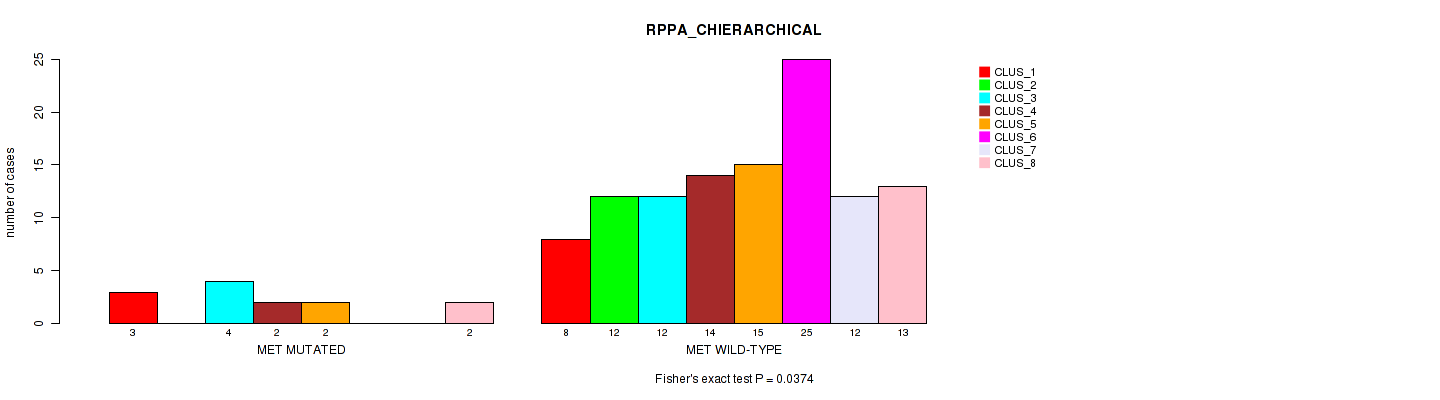
P value = 0.108 (Fisher's exact test), Q value = 1
Table S85. Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| MET MUTATED | 3 | 9 | 3 |
| MET WILD-TYPE | 40 | 47 | 59 |
P value = 0.00033 (Fisher's exact test), Q value = 0.06
Table S86. Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| MET MUTATED | 4 | 1 | 8 | 2 |
| MET WILD-TYPE | 69 | 38 | 12 | 27 |
Figure S6. Get High-res Image Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
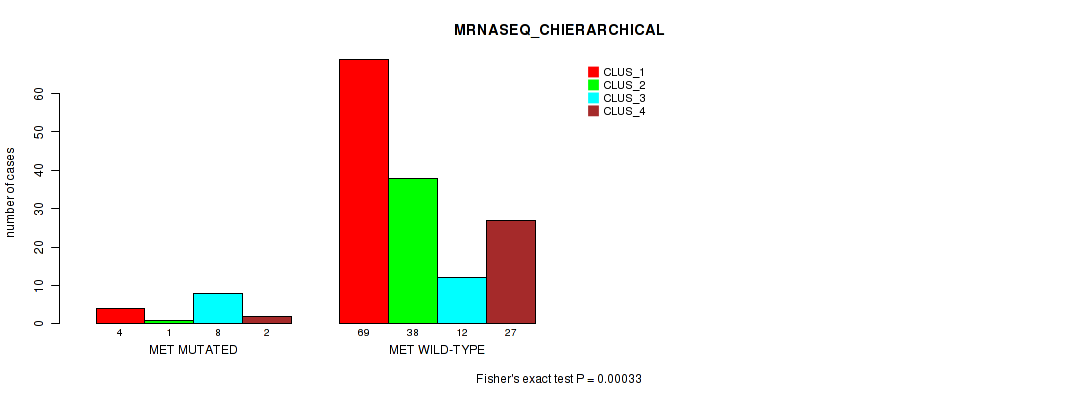
P value = 0.838 (Fisher's exact test), Q value = 1
Table S87. Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| MET MUTATED | 5 | 4 | 6 |
| MET WILD-TYPE | 42 | 50 | 54 |
P value = 0.815 (Fisher's exact test), Q value = 1
Table S88. Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| MET MUTATED | 3 | 2 | 3 | 7 |
| MET WILD-TYPE | 27 | 33 | 36 | 50 |
P value = 0.642 (Fisher's exact test), Q value = 1
Table S89. Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| MET MUTATED | 4 | 5 | 3 |
| MET WILD-TYPE | 33 | 34 | 43 |
P value = 0.141 (Fisher's exact test), Q value = 1
Table S90. Gene #9: 'MET MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| MET MUTATED | 5 | 3 | 0 | 4 |
| MET WILD-TYPE | 42 | 44 | 12 | 12 |
P value = 0.825 (Fisher's exact test), Q value = 1
Table S91. Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| SMARCB1 MUTATED | 2 | 1 | 1 | 0 |
| SMARCB1 WILD-TYPE | 59 | 38 | 24 | 36 |
P value = 0.216 (Fisher's exact test), Q value = 1
Table S92. Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| SMARCB1 MUTATED | 0 | 3 | 1 |
| SMARCB1 WILD-TYPE | 40 | 42 | 62 |
P value = 1 (Fisher's exact test), Q value = 1
Table S93. Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| SMARCB1 MUTATED | 1 | 1 | 1 |
| SMARCB1 WILD-TYPE | 47 | 43 | 31 |
P value = 0.584 (Fisher's exact test), Q value = 1
Table S94. Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| SMARCB1 MUTATED | 0 | 1 | 0 | 0 | 0 | 1 | 1 | 0 |
| SMARCB1 WILD-TYPE | 11 | 11 | 16 | 16 | 17 | 24 | 11 | 15 |
P value = 0.453 (Fisher's exact test), Q value = 1
Table S95. Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| SMARCB1 MUTATED | 0 | 1 | 3 |
| SMARCB1 WILD-TYPE | 43 | 55 | 59 |
P value = 0.413 (Fisher's exact test), Q value = 1
Table S96. Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| SMARCB1 MUTATED | 3 | 0 | 1 | 0 |
| SMARCB1 WILD-TYPE | 70 | 39 | 19 | 29 |
P value = 0.0352 (Fisher's exact test), Q value = 1
Table S97. Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| SMARCB1 MUTATED | 0 | 0 | 4 |
| SMARCB1 WILD-TYPE | 47 | 54 | 56 |
Figure S7. Get High-res Image Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
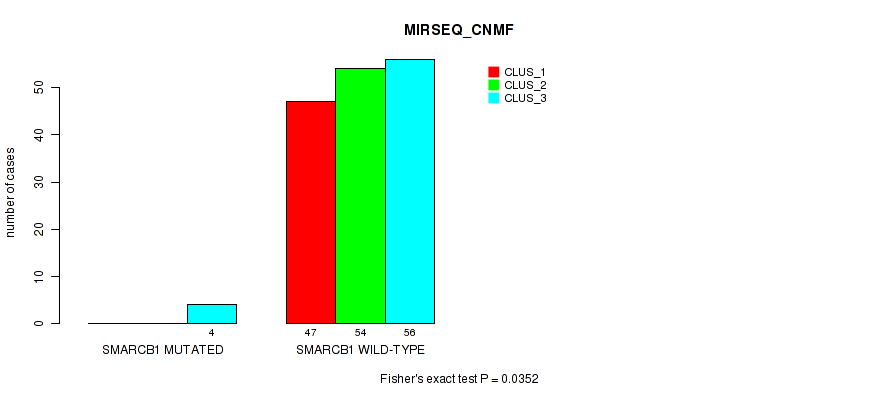
P value = 0.463 (Fisher's exact test), Q value = 1
Table S98. Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| SMARCB1 MUTATED | 0 | 1 | 0 | 3 |
| SMARCB1 WILD-TYPE | 30 | 34 | 39 | 54 |
P value = 0.108 (Fisher's exact test), Q value = 1
Table S99. Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| SMARCB1 MUTATED | 0 | 0 | 3 |
| SMARCB1 WILD-TYPE | 37 | 39 | 43 |
P value = 0.33 (Fisher's exact test), Q value = 1
Table S100. Gene #10: 'SMARCB1 MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| SMARCB1 MUTATED | 0 | 3 | 0 | 0 |
| SMARCB1 WILD-TYPE | 47 | 44 | 12 | 16 |
P value = 1 (Fisher's exact test), Q value = 1
Table S101. Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| KDM6A MUTATED | 3 | 2 | 1 | 1 |
| KDM6A WILD-TYPE | 58 | 37 | 24 | 35 |
P value = 0.555 (Fisher's exact test), Q value = 1
Table S102. Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| KDM6A MUTATED | 3 | 2 | 2 |
| KDM6A WILD-TYPE | 37 | 43 | 61 |
P value = 0.632 (Fisher's exact test), Q value = 1
Table S103. Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| KDM6A MUTATED | 1 | 2 | 2 |
| KDM6A WILD-TYPE | 47 | 42 | 30 |
P value = 0.613 (Fisher's exact test), Q value = 1
Table S104. Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| KDM6A MUTATED | 0 | 0 | 1 | 1 | 0 | 3 | 0 | 0 |
| KDM6A WILD-TYPE | 11 | 12 | 15 | 15 | 17 | 22 | 12 | 15 |
P value = 0.0302 (Fisher's exact test), Q value = 1
Table S105. Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| KDM6A MUTATED | 5 | 1 | 1 |
| KDM6A WILD-TYPE | 38 | 55 | 61 |
Figure S8. Get High-res Image Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
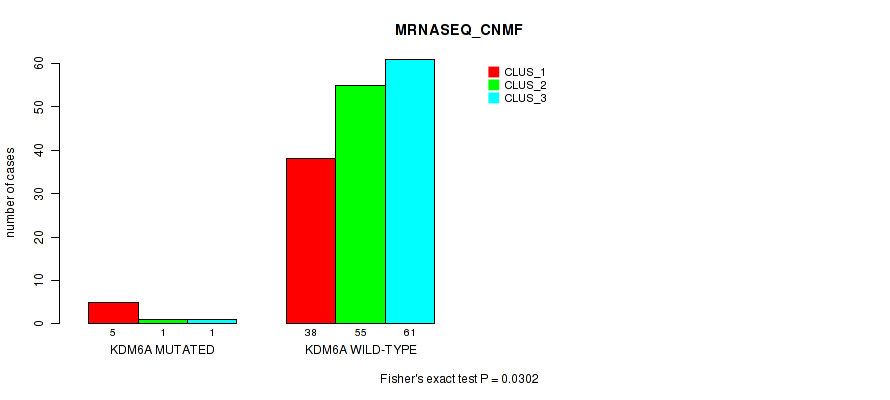
P value = 0.735 (Fisher's exact test), Q value = 1
Table S106. Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| KDM6A MUTATED | 5 | 1 | 0 | 1 |
| KDM6A WILD-TYPE | 68 | 38 | 20 | 28 |
P value = 0.387 (Fisher's exact test), Q value = 1
Table S107. Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| KDM6A MUTATED | 3 | 3 | 1 |
| KDM6A WILD-TYPE | 44 | 51 | 59 |
P value = 0.162 (Fisher's exact test), Q value = 1
Table S108. Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| KDM6A MUTATED | 1 | 4 | 1 | 1 |
| KDM6A WILD-TYPE | 29 | 31 | 38 | 56 |
P value = 0.396 (Fisher's exact test), Q value = 1
Table S109. Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| KDM6A MUTATED | 2 | 0 | 1 |
| KDM6A WILD-TYPE | 35 | 39 | 45 |
P value = 0.22 (Fisher's exact test), Q value = 1
Table S110. Gene #11: 'KDM6A MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| KDM6A MUTATED | 2 | 0 | 1 | 0 |
| KDM6A WILD-TYPE | 45 | 47 | 11 | 16 |
P value = 0.132 (Fisher's exact test), Q value = 1
Table S111. Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| SETD2 MUTATED | 4 | 1 | 0 | 5 |
| SETD2 WILD-TYPE | 57 | 38 | 25 | 31 |
P value = 6e-05 (Fisher's exact test), Q value = 0.011
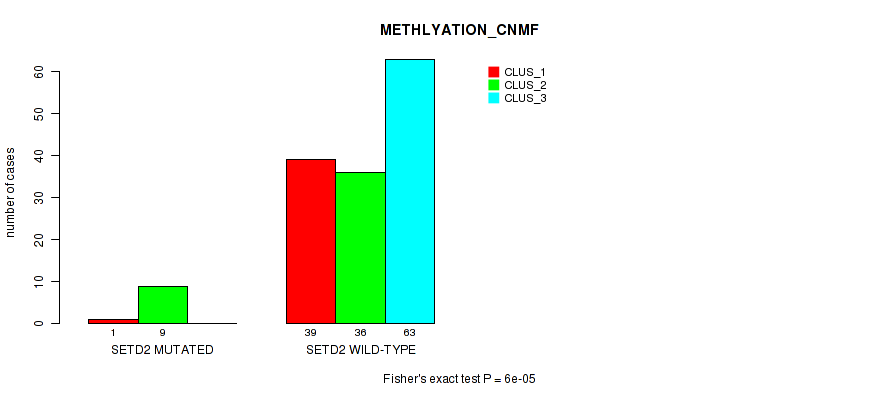
Table S112. Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| SETD2 MUTATED | 1 | 9 | 0 |
| SETD2 WILD-TYPE | 39 | 36 | 63 |
Figure S9. Get High-res Image Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
P value = 0.193 (Fisher's exact test), Q value = 1
Table S113. Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| SETD2 MUTATED | 2 | 1 | 4 |
| SETD2 WILD-TYPE | 46 | 43 | 28 |
P value = 0.526 (Fisher's exact test), Q value = 1
Table S114. Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| SETD2 MUTATED | 0 | 0 | 0 | 1 | 1 | 4 | 0 | 1 |
| SETD2 WILD-TYPE | 11 | 12 | 16 | 15 | 16 | 21 | 12 | 14 |
P value = 0.00764 (Fisher's exact test), Q value = 1
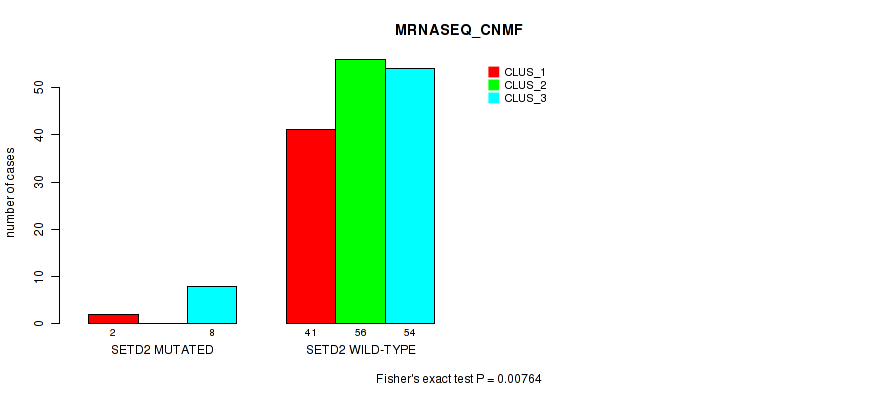
Table S115. Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| SETD2 MUTATED | 2 | 0 | 8 |
| SETD2 WILD-TYPE | 41 | 56 | 54 |
Figure S10. Get High-res Image Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
P value = 0.0332 (Fisher's exact test), Q value = 1
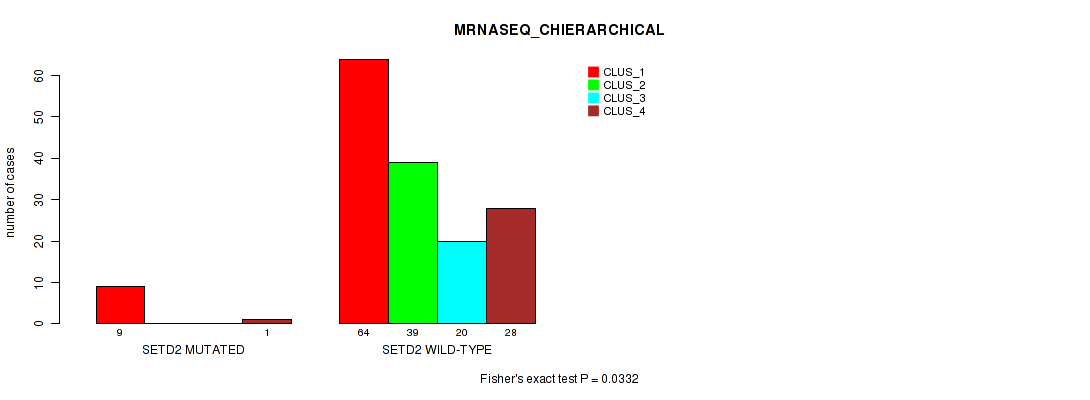
Table S116. Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| SETD2 MUTATED | 9 | 0 | 0 | 1 |
| SETD2 WILD-TYPE | 64 | 39 | 20 | 28 |
Figure S11. Get High-res Image Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
P value = 0.0177 (Fisher's exact test), Q value = 1
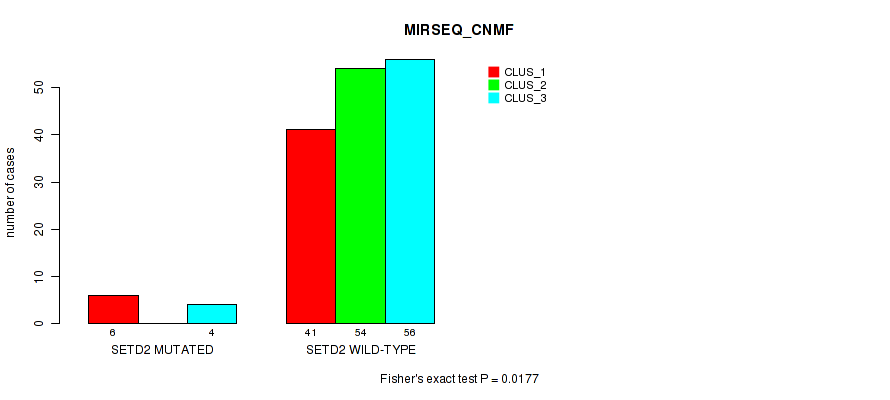
Table S117. Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| SETD2 MUTATED | 6 | 0 | 4 |
| SETD2 WILD-TYPE | 41 | 54 | 56 |
Figure S12. Get High-res Image Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
P value = 0.0202 (Fisher's exact test), Q value = 1
Table S118. Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| SETD2 MUTATED | 5 | 3 | 0 | 2 |
| SETD2 WILD-TYPE | 25 | 32 | 39 | 55 |
Figure S13. Get High-res Image Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
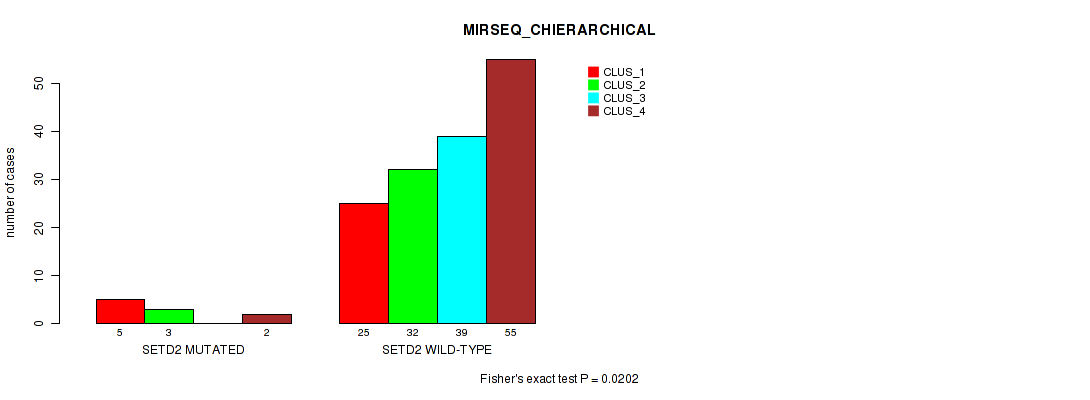
P value = 0.0728 (Fisher's exact test), Q value = 1
Table S119. Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| SETD2 MUTATED | 4 | 0 | 5 |
| SETD2 WILD-TYPE | 33 | 39 | 41 |
P value = 0.343 (Fisher's exact test), Q value = 1
Table S120. Gene #12: 'SETD2 MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| SETD2 MUTATED | 3 | 6 | 0 | 0 |
| SETD2 WILD-TYPE | 44 | 41 | 12 | 16 |
P value = 0.951 (Fisher's exact test), Q value = 1
Table S121. Gene #13: 'AHNAK2 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| AHNAK2 MUTATED | 3 | 1 | 1 | 2 |
| AHNAK2 WILD-TYPE | 58 | 38 | 24 | 34 |
P value = 0.764 (Fisher's exact test), Q value = 1
Table S122. Gene #13: 'AHNAK2 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| AHNAK2 MUTATED | 2 | 1 | 3 |
| AHNAK2 WILD-TYPE | 38 | 44 | 60 |
P value = 0.657 (Fisher's exact test), Q value = 1
Table S123. Gene #13: 'AHNAK2 MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| AHNAK2 MUTATED | 3 | 1 | 2 |
| AHNAK2 WILD-TYPE | 45 | 43 | 30 |
P value = 0.973 (Fisher's exact test), Q value = 1
Table S124. Gene #13: 'AHNAK2 MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| AHNAK2 MUTATED | 0 | 1 | 1 | 0 | 1 | 2 | 0 | 1 |
| AHNAK2 WILD-TYPE | 11 | 11 | 15 | 16 | 16 | 23 | 12 | 14 |
P value = 0.794 (Fisher's exact test), Q value = 1
Table S125. Gene #13: 'AHNAK2 MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| AHNAK2 MUTATED | 1 | 3 | 3 |
| AHNAK2 WILD-TYPE | 42 | 53 | 59 |
P value = 0.235 (Fisher's exact test), Q value = 1
Table S126. Gene #13: 'AHNAK2 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| AHNAK2 MUTATED | 4 | 0 | 2 | 1 |
| AHNAK2 WILD-TYPE | 69 | 39 | 18 | 28 |
P value = 0.295 (Fisher's exact test), Q value = 1
Table S127. Gene #13: 'AHNAK2 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| AHNAK2 MUTATED | 4 | 1 | 2 |
| AHNAK2 WILD-TYPE | 43 | 53 | 58 |
P value = 0.213 (Fisher's exact test), Q value = 1
Table S128. Gene #13: 'AHNAK2 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| AHNAK2 MUTATED | 2 | 3 | 0 | 2 |
| AHNAK2 WILD-TYPE | 28 | 32 | 39 | 55 |
P value = 0.257 (Fisher's exact test), Q value = 1
Table S129. Gene #13: 'AHNAK2 MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| AHNAK2 MUTATED | 4 | 2 | 1 |
| AHNAK2 WILD-TYPE | 33 | 37 | 45 |
P value = 0.3 (Fisher's exact test), Q value = 1
Table S130. Gene #13: 'AHNAK2 MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| AHNAK2 MUTATED | 5 | 1 | 0 | 1 |
| AHNAK2 WILD-TYPE | 42 | 46 | 12 | 15 |
P value = 0.0589 (Fisher's exact test), Q value = 1
Table S131. Gene #14: 'AHCY MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| AHCY MUTATED | 0 | 1 | 0 | 3 |
| AHCY WILD-TYPE | 61 | 38 | 25 | 33 |
P value = 0.473 (Fisher's exact test), Q value = 1
Table S132. Gene #14: 'AHCY MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| AHCY MUTATED | 0 | 2 | 1 |
| AHCY WILD-TYPE | 40 | 43 | 62 |
P value = 0.559 (Fisher's exact test), Q value = 1
Table S133. Gene #14: 'AHCY MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| AHCY MUTATED | 0 | 2 | 2 |
| AHCY WILD-TYPE | 43 | 54 | 60 |
P value = 0.889 (Fisher's exact test), Q value = 1
Table S134. Gene #14: 'AHCY MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| AHCY MUTATED | 3 | 1 | 0 | 0 |
| AHCY WILD-TYPE | 70 | 38 | 20 | 29 |
P value = 0.261 (Fisher's exact test), Q value = 1
Table S135. Gene #14: 'AHCY MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| AHCY MUTATED | 2 | 2 | 0 |
| AHCY WILD-TYPE | 45 | 52 | 60 |
P value = 0.646 (Fisher's exact test), Q value = 1
Table S136. Gene #14: 'AHCY MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| AHCY MUTATED | 0 | 2 | 1 | 1 |
| AHCY WILD-TYPE | 30 | 33 | 38 | 56 |
P value = 0.196 (Fisher's exact test), Q value = 1
Table S137. Gene #14: 'AHCY MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| AHCY MUTATED | 2 | 1 | 0 |
| AHCY WILD-TYPE | 35 | 38 | 46 |
P value = 1 (Fisher's exact test), Q value = 1
Table S138. Gene #14: 'AHCY MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| AHCY MUTATED | 2 | 1 | 0 | 0 |
| AHCY WILD-TYPE | 45 | 46 | 12 | 16 |
P value = 0.29 (Fisher's exact test), Q value = 1
Table S139. Gene #15: 'IDUA MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| IDUA MUTATED | 4 | 0 | 0 | 1 |
| IDUA WILD-TYPE | 57 | 39 | 25 | 35 |
P value = 0.731 (Fisher's exact test), Q value = 1
Table S140. Gene #15: 'IDUA MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| IDUA MUTATED | 2 | 1 | 2 |
| IDUA WILD-TYPE | 38 | 44 | 61 |
P value = 0.3 (Fisher's exact test), Q value = 1
Table S141. Gene #15: 'IDUA MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| IDUA MUTATED | 1 | 1 | 3 |
| IDUA WILD-TYPE | 47 | 43 | 29 |
P value = 0.459 (Fisher's exact test), Q value = 1
Table S142. Gene #15: 'IDUA MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| IDUA MUTATED | 0 | 1 | 0 | 2 | 0 | 2 | 0 | 0 |
| IDUA WILD-TYPE | 11 | 11 | 16 | 14 | 17 | 23 | 12 | 15 |
P value = 0.739 (Fisher's exact test), Q value = 1
Table S143. Gene #15: 'IDUA MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| IDUA MUTATED | 1 | 1 | 3 |
| IDUA WILD-TYPE | 42 | 55 | 59 |
P value = 0.267 (Fisher's exact test), Q value = 1
Table S144. Gene #15: 'IDUA MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| IDUA MUTATED | 2 | 0 | 1 | 2 |
| IDUA WILD-TYPE | 71 | 39 | 19 | 27 |
P value = 0.444 (Fisher's exact test), Q value = 1
Table S145. Gene #15: 'IDUA MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| IDUA MUTATED | 3 | 1 | 1 |
| IDUA WILD-TYPE | 44 | 53 | 59 |
P value = 0.0702 (Fisher's exact test), Q value = 1
Table S146. Gene #15: 'IDUA MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| IDUA MUTATED | 3 | 0 | 0 | 2 |
| IDUA WILD-TYPE | 27 | 35 | 39 | 55 |
P value = 0.83 (Fisher's exact test), Q value = 1
Table S147. Gene #15: 'IDUA MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| IDUA MUTATED | 1 | 2 | 1 |
| IDUA WILD-TYPE | 36 | 37 | 45 |
P value = 0.868 (Fisher's exact test), Q value = 1
Table S148. Gene #15: 'IDUA MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| IDUA MUTATED | 1 | 2 | 0 | 1 |
| IDUA WILD-TYPE | 46 | 45 | 12 | 15 |
P value = 0.391 (Fisher's exact test), Q value = 1
Table S149. Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| OR2L8 MUTATED | 1 | 2 | 1 | 0 |
| OR2L8 WILD-TYPE | 60 | 37 | 24 | 36 |
P value = 0.294 (Fisher's exact test), Q value = 1
Table S150. Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| OR2L8 MUTATED | 2 | 0 | 2 |
| OR2L8 WILD-TYPE | 38 | 45 | 61 |
P value = 0.676 (Fisher's exact test), Q value = 1
Table S151. Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| OR2L8 MUTATED | 2 | 2 | 0 |
| OR2L8 WILD-TYPE | 46 | 42 | 32 |
P value = 0.196 (Fisher's exact test), Q value = 1
Table S152. Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| OR2L8 MUTATED | 0 | 1 | 2 | 0 | 0 | 0 | 0 | 1 |
| OR2L8 WILD-TYPE | 11 | 11 | 14 | 16 | 17 | 25 | 12 | 14 |
P value = 0.375 (Fisher's exact test), Q value = 1
Table S153. Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| OR2L8 MUTATED | 0 | 3 | 1 |
| OR2L8 WILD-TYPE | 43 | 53 | 61 |
P value = 0.0398 (Fisher's exact test), Q value = 1
Table S154. Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| OR2L8 MUTATED | 0 | 1 | 2 | 1 |
| OR2L8 WILD-TYPE | 73 | 38 | 18 | 28 |
Figure S14. Get High-res Image Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
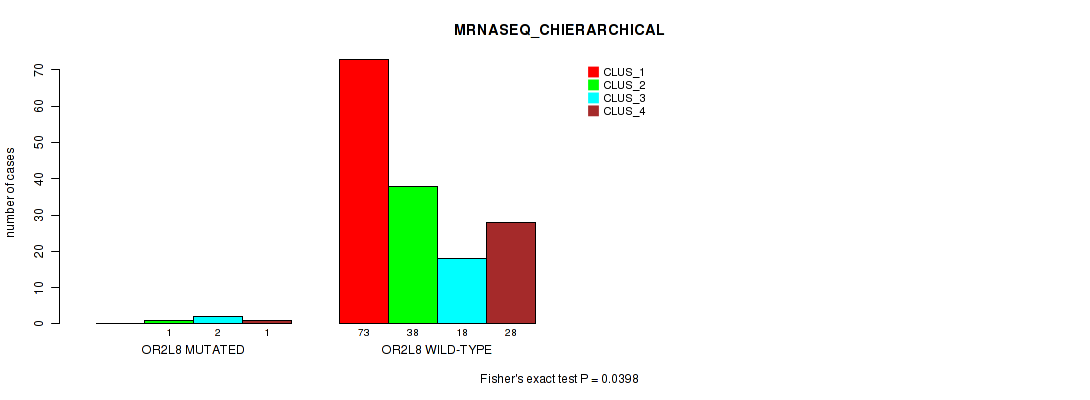
P value = 0.39 (Fisher's exact test), Q value = 1
Table S155. Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| OR2L8 MUTATED | 0 | 1 | 3 |
| OR2L8 WILD-TYPE | 47 | 53 | 57 |
P value = 0.508 (Fisher's exact test), Q value = 1
Table S156. Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| OR2L8 MUTATED | 0 | 0 | 1 | 3 |
| OR2L8 WILD-TYPE | 30 | 35 | 38 | 54 |
P value = 0.549 (Fisher's exact test), Q value = 1
Table S157. Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| OR2L8 MUTATED | 0 | 2 | 2 |
| OR2L8 WILD-TYPE | 37 | 37 | 44 |
P value = 0.866 (Fisher's exact test), Q value = 1
Table S158. Gene #16: 'OR2L8 MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| OR2L8 MUTATED | 1 | 2 | 0 | 1 |
| OR2L8 WILD-TYPE | 46 | 45 | 12 | 15 |
P value = 0.181 (Fisher's exact test), Q value = 1
Table S159. Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| HOXD8 MUTATED | 1 | 3 | 0 | 0 |
| HOXD8 WILD-TYPE | 60 | 36 | 25 | 36 |
P value = 0.785 (Fisher's exact test), Q value = 1
Table S160. Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| HOXD8 MUTATED | 0 | 1 | 2 |
| HOXD8 WILD-TYPE | 40 | 44 | 61 |
P value = 0.0183 (Fisher's exact test), Q value = 1
Table S161. Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| HOXD8 MUTATED | 0 | 4 | 0 |
| HOXD8 WILD-TYPE | 48 | 40 | 32 |
Figure S15. Get High-res Image Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
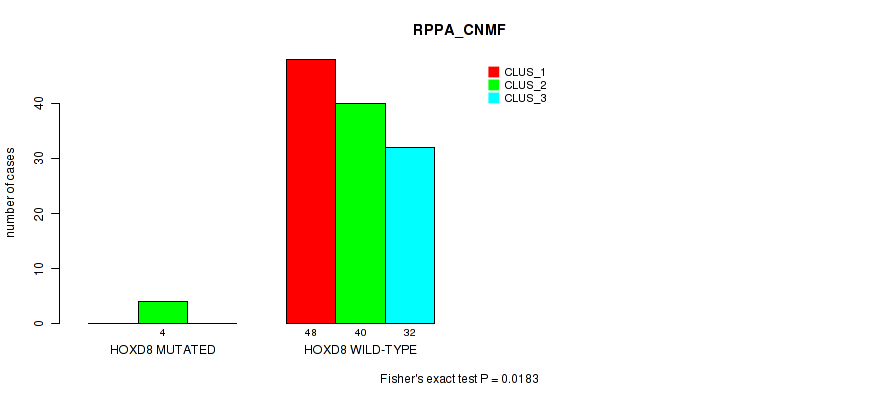
P value = 0.034 (Fisher's exact test), Q value = 1
Table S162. Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| HOXD8 MUTATED | 0 | 0 | 3 | 1 | 0 | 0 | 0 | 0 |
| HOXD8 WILD-TYPE | 11 | 12 | 13 | 15 | 17 | 25 | 12 | 15 |
Figure S16. Get High-res Image Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
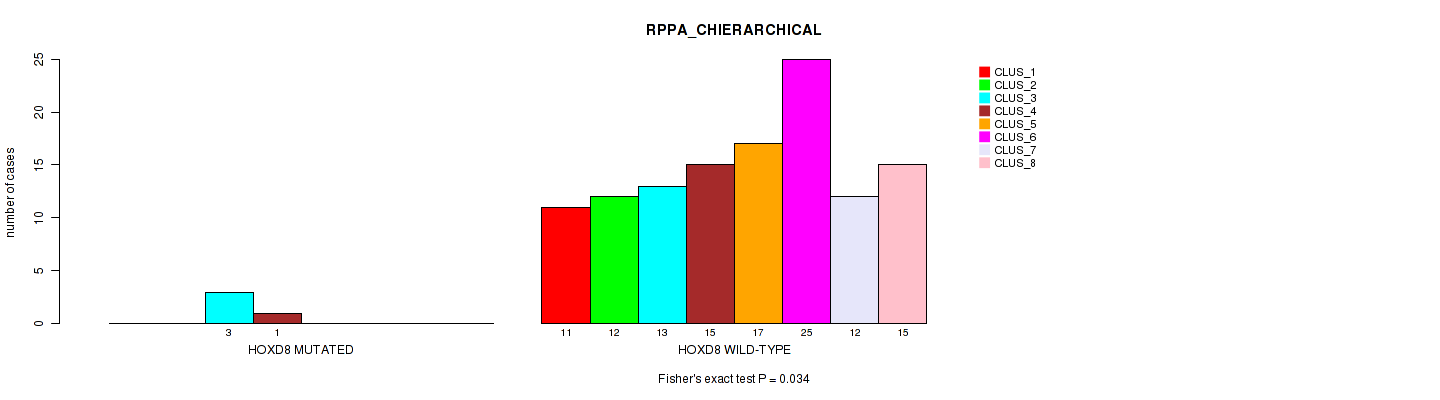
P value = 0.375 (Fisher's exact test), Q value = 1
Table S163. Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| HOXD8 MUTATED | 0 | 3 | 1 |
| HOXD8 WILD-TYPE | 43 | 53 | 61 |
P value = 0.153 (Fisher's exact test), Q value = 1
Table S164. Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| HOXD8 MUTATED | 1 | 1 | 2 | 0 |
| HOXD8 WILD-TYPE | 72 | 38 | 18 | 29 |
P value = 0.321 (Fisher's exact test), Q value = 1
Table S165. Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| HOXD8 MUTATED | 1 | 0 | 3 |
| HOXD8 WILD-TYPE | 46 | 54 | 57 |
P value = 0.834 (Fisher's exact test), Q value = 1
Table S166. Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| HOXD8 MUTATED | 1 | 0 | 1 | 2 |
| HOXD8 WILD-TYPE | 29 | 35 | 38 | 55 |
P value = 0.0173 (Fisher's exact test), Q value = 1
Table S167. Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 39 | 46 |
| HOXD8 MUTATED | 0 | 4 | 0 |
| HOXD8 WILD-TYPE | 37 | 35 | 46 |
Figure S17. Get High-res Image Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #9: 'MIRSEQ_MATURE_CNMF'
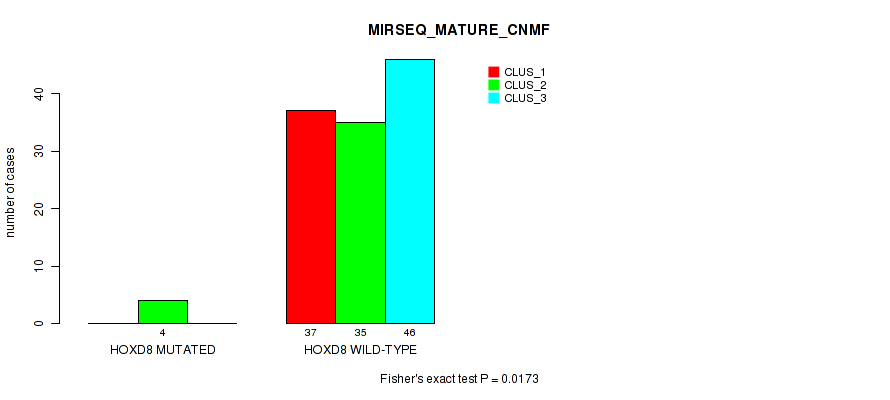
P value = 0.0106 (Fisher's exact test), Q value = 1
Table S168. Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 47 | 47 | 12 | 16 |
| HOXD8 MUTATED | 1 | 0 | 0 | 3 |
| HOXD8 WILD-TYPE | 46 | 47 | 12 | 13 |
Figure S18. Get High-res Image Gene #17: 'HOXD8 MUTATION STATUS' versus Molecular Subtype #10: 'MIRSEQ_MATURE_CHIERARCHICAL'
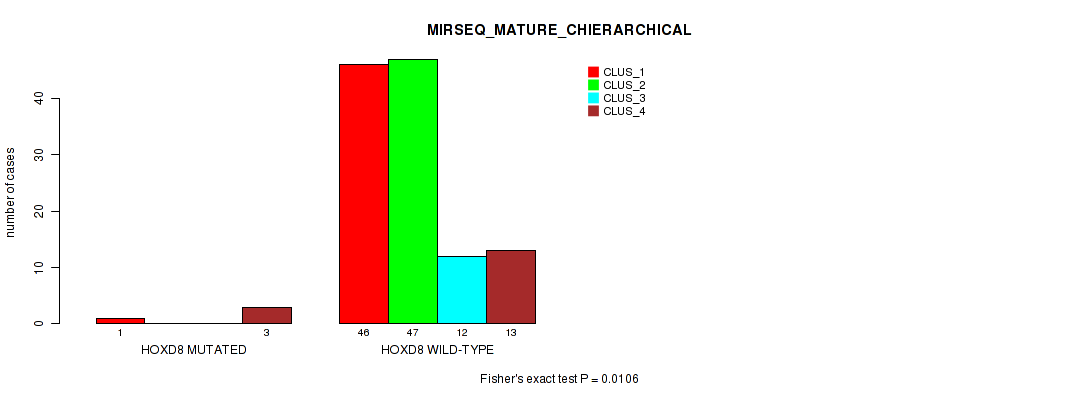
P value = 0.585 (Fisher's exact test), Q value = 1
Table S169. Gene #18: 'PAM MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| PAM MUTATED | 1 | 0 | 1 | 1 |
| PAM WILD-TYPE | 60 | 39 | 24 | 35 |
P value = 0.787 (Fisher's exact test), Q value = 1
Table S170. Gene #18: 'PAM MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| PAM MUTATED | 0 | 1 | 2 |
| PAM WILD-TYPE | 40 | 44 | 61 |
P value = 0.115 (Fisher's exact test), Q value = 1
Table S171. Gene #18: 'PAM MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| PAM MUTATED | 3 | 0 | 0 |
| PAM WILD-TYPE | 45 | 44 | 32 |
P value = 0.104 (Fisher's exact test), Q value = 1
Table S172. Gene #18: 'PAM MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| PAM MUTATED | 0 | 0 | 0 | 2 | 0 | 0 | 1 | 0 |
| PAM WILD-TYPE | 11 | 12 | 16 | 14 | 17 | 25 | 11 | 15 |
P value = 0.189 (Fisher's exact test), Q value = 1
Table S173. Gene #18: 'PAM MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| PAM MUTATED | 2 | 1 | 0 |
| PAM WILD-TYPE | 41 | 55 | 62 |
P value = 1 (Fisher's exact test), Q value = 1
Table S174. Gene #18: 'PAM MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| PAM MUTATED | 2 | 1 | 0 | 0 |
| PAM WILD-TYPE | 71 | 38 | 20 | 29 |
P value = 0.392 (Fisher's exact test), Q value = 1
Table S175. Gene #18: 'PAM MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| PAM MUTATED | 1 | 2 | 0 |
| PAM WILD-TYPE | 46 | 52 | 60 |
P value = 0.0291 (Fisher's exact test), Q value = 1
Table S176. Gene #18: 'PAM MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| PAM MUTATED | 0 | 0 | 3 | 0 |
| PAM WILD-TYPE | 30 | 35 | 36 | 57 |
Figure S19. Get High-res Image Gene #18: 'PAM MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
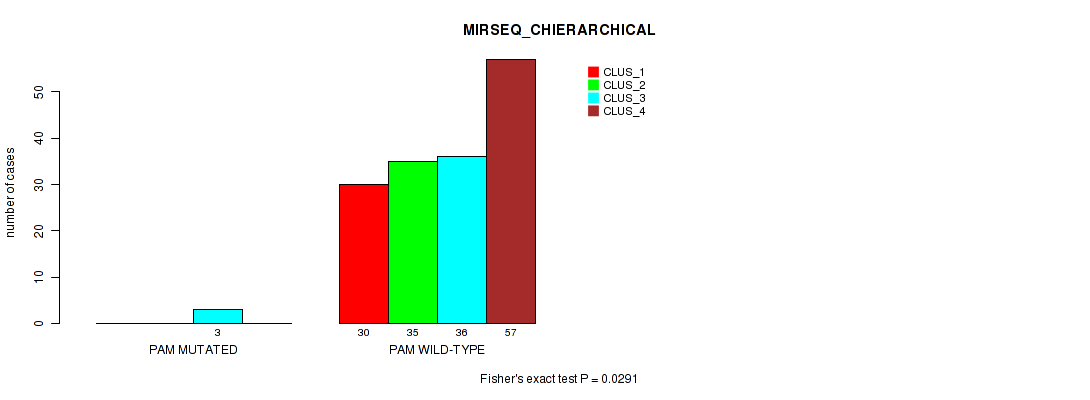
P value = 0.137 (Fisher's exact test), Q value = 1
Table S177. Gene #19: 'CSGALNACT2 MUTATION STATUS' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 61 | 39 | 25 | 36 |
| CSGALNACT2 MUTATED | 1 | 0 | 1 | 3 |
| CSGALNACT2 WILD-TYPE | 60 | 39 | 24 | 33 |
P value = 0.327 (Fisher's exact test), Q value = 1
Table S178. Gene #19: 'CSGALNACT2 MUTATION STATUS' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 45 | 63 |
| CSGALNACT2 MUTATED | 3 | 1 | 1 |
| CSGALNACT2 WILD-TYPE | 37 | 44 | 62 |
P value = 0.831 (Fisher's exact test), Q value = 1
Table S179. Gene #19: 'CSGALNACT2 MUTATION STATUS' versus Molecular Subtype #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 32 |
| CSGALNACT2 MUTATED | 1 | 2 | 1 |
| CSGALNACT2 WILD-TYPE | 47 | 42 | 31 |
P value = 0.774 (Fisher's exact test), Q value = 1
Table S180. Gene #19: 'CSGALNACT2 MUTATION STATUS' versus Molecular Subtype #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 | CLUS_8 |
|---|---|---|---|---|---|---|---|---|
| ALL | 11 | 12 | 16 | 16 | 17 | 25 | 12 | 15 |
| CSGALNACT2 MUTATED | 1 | 0 | 0 | 1 | 0 | 1 | 0 | 1 |
| CSGALNACT2 WILD-TYPE | 10 | 12 | 16 | 15 | 17 | 24 | 12 | 14 |
P value = 0.741 (Fisher's exact test), Q value = 1
Table S181. Gene #19: 'CSGALNACT2 MUTATION STATUS' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 56 | 62 |
| CSGALNACT2 MUTATED | 1 | 1 | 3 |
| CSGALNACT2 WILD-TYPE | 42 | 55 | 59 |
P value = 0.618 (Fisher's exact test), Q value = 1
Table S182. Gene #19: 'CSGALNACT2 MUTATION STATUS' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 73 | 39 | 20 | 29 |
| CSGALNACT2 MUTATED | 2 | 1 | 0 | 2 |
| CSGALNACT2 WILD-TYPE | 71 | 38 | 20 | 27 |
P value = 0.525 (Fisher's exact test), Q value = 1
Table S183. Gene #19: 'CSGALNACT2 MUTATION STATUS' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 47 | 54 | 60 |
| CSGALNACT2 MUTATED | 1 | 3 | 1 |
| CSGALNACT2 WILD-TYPE | 46 | 51 | 59 |
P value = 0.728 (Fisher's exact test), Q value = 1
Table S184. Gene #19: 'CSGALNACT2 MUTATION STATUS' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 30 | 35 | 39 | 57 |
| CSGALNACT2 MUTATED | 0 | 2 | 1 | 2 |
| CSGALNACT2 WILD-TYPE | 30 | 33 | 38 | 55 |
-
Mutation data file = transformed.cor.cli.txt
-
Molecular subtypes file = KIRP-TP.transferedmergedcluster.txt
-
Number of patients = 161
-
Number of significantly mutated genes = 19
-
Number of Molecular subtypes = 10
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.