This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 61 focal events and 8 molecular subtypes across 56 patients, 2 significant findings detected with P value < 0.05 and Q value < 0.25.
-
amp_19q12 cnv correlated to 'METHLYATION_CNMF'.
-
del_13q14.2 cnv correlated to 'METHLYATION_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 61 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 2 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| amp 19q12 | 45 (80%) | 11 |
0.00139 (0.673) |
0.00013 (0.0634) |
0.191 (1.00) |
0.0199 (1.00) |
0.642 (1.00) |
0.501 (1.00) |
0.232 (1.00) |
0.437 (1.00) |
| del 13q14 2 | 32 (57%) | 24 |
0.436 (1.00) |
0.00041 (0.2) |
0.0129 (1.00) |
0.0201 (1.00) |
0.48 (1.00) |
0.158 (1.00) |
0.0464 (1.00) |
0.193 (1.00) |
| amp 1q22 | 42 (75%) | 14 |
0.505 (1.00) |
0.224 (1.00) |
0.477 (1.00) |
0.294 (1.00) |
0.185 (1.00) |
0.187 (1.00) |
0.333 (1.00) |
0.267 (1.00) |
| amp 2p14 | 28 (50%) | 28 |
0.56 (1.00) |
0.0251 (1.00) |
0.00736 (1.00) |
0.0814 (1.00) |
0.489 (1.00) |
0.248 (1.00) |
0.223 (1.00) |
0.36 (1.00) |
| amp 2q13 | 29 (52%) | 27 |
0.532 (1.00) |
0.0156 (1.00) |
0.0465 (1.00) |
0.318 (1.00) |
1 (1.00) |
0.664 (1.00) |
0.738 (1.00) |
0.729 (1.00) |
| amp 3p25 1 | 20 (36%) | 36 |
0.0123 (1.00) |
0.515 (1.00) |
0.3 (1.00) |
0.112 (1.00) |
0.228 (1.00) |
0.0652 (1.00) |
0.122 (1.00) |
0.0663 (1.00) |
| amp 3q26 2 | 37 (66%) | 19 |
0.00515 (1.00) |
0.00499 (1.00) |
0.0908 (1.00) |
0.158 (1.00) |
0.349 (1.00) |
0.241 (1.00) |
0.462 (1.00) |
0.183 (1.00) |
| amp 4p16 3 | 25 (45%) | 31 |
0.0274 (1.00) |
0.13 (1.00) |
0.39 (1.00) |
0.187 (1.00) |
0.975 (1.00) |
0.288 (1.00) |
0.505 (1.00) |
0.531 (1.00) |
| amp 5p13 2 | 26 (46%) | 30 |
0.00172 (0.831) |
0.568 (1.00) |
0.245 (1.00) |
0.0569 (1.00) |
0.928 (1.00) |
0.0458 (1.00) |
0.478 (1.00) |
0.173 (1.00) |
| amp 6p24 2 | 37 (66%) | 19 |
0.109 (1.00) |
0.972 (1.00) |
0.688 (1.00) |
0.357 (1.00) |
0.877 (1.00) |
0.498 (1.00) |
0.875 (1.00) |
0.77 (1.00) |
| amp 8p11 21 | 33 (59%) | 23 |
0.551 (1.00) |
0.625 (1.00) |
0.679 (1.00) |
0.557 (1.00) |
0.624 (1.00) |
1 (1.00) |
0.779 (1.00) |
0.598 (1.00) |
| amp 8q11 23 | 38 (68%) | 18 |
0.0126 (1.00) |
0.227 (1.00) |
0.199 (1.00) |
0.168 (1.00) |
0.771 (1.00) |
0.922 (1.00) |
0.621 (1.00) |
0.857 (1.00) |
| amp 8q24 21 | 42 (75%) | 14 |
0.00206 (0.991) |
0.757 (1.00) |
0.19 (1.00) |
0.339 (1.00) |
0.0389 (1.00) |
0.0308 (1.00) |
0.00594 (1.00) |
0.015 (1.00) |
| amp 8q24 21 | 40 (71%) | 16 |
0.00053 (0.258) |
0.357 (1.00) |
0.0767 (1.00) |
0.118 (1.00) |
0.0474 (1.00) |
0.0634 (1.00) |
0.0077 (1.00) |
0.022 (1.00) |
| amp 10q22 2 | 26 (46%) | 30 |
0.312 (1.00) |
0.486 (1.00) |
0.963 (1.00) |
0.921 (1.00) |
0.725 (1.00) |
0.74 (1.00) |
1 (1.00) |
0.426 (1.00) |
| amp 11q13 1 | 19 (34%) | 37 |
0.777 (1.00) |
0.648 (1.00) |
0.327 (1.00) |
0.306 (1.00) |
0.762 (1.00) |
1 (1.00) |
0.823 (1.00) |
0.975 (1.00) |
| amp 12q12 | 22 (39%) | 34 |
0.031 (1.00) |
0.105 (1.00) |
0.258 (1.00) |
0.404 (1.00) |
0.926 (1.00) |
1 (1.00) |
0.883 (1.00) |
0.953 (1.00) |
| amp 12q15 | 17 (30%) | 39 |
0.586 (1.00) |
0.381 (1.00) |
0.254 (1.00) |
0.702 (1.00) |
0.8 (1.00) |
0.167 (1.00) |
1 (1.00) |
0.229 (1.00) |
| amp 13q31 3 | 25 (45%) | 31 |
0.525 (1.00) |
0.442 (1.00) |
0.135 (1.00) |
0.116 (1.00) |
0.0844 (1.00) |
0.0371 (1.00) |
0.00244 (1.00) |
0.0539 (1.00) |
| amp 16p11 2 | 20 (36%) | 36 |
0.0278 (1.00) |
0.283 (1.00) |
0.652 (1.00) |
0.0745 (1.00) |
0.759 (1.00) |
0.409 (1.00) |
0.515 (1.00) |
0.209 (1.00) |
| amp 17q12 | 27 (48%) | 29 |
0.445 (1.00) |
0.19 (1.00) |
0.333 (1.00) |
0.46 (1.00) |
0.343 (1.00) |
0.783 (1.00) |
0.608 (1.00) |
1 (1.00) |
| amp 17q25 1 | 32 (57%) | 24 |
0.496 (1.00) |
0.0123 (1.00) |
0.63 (1.00) |
0.307 (1.00) |
0.688 (1.00) |
0.717 (1.00) |
0.323 (1.00) |
0.658 (1.00) |
| amp 19p13 2 | 19 (34%) | 37 |
0.332 (1.00) |
0.273 (1.00) |
0.796 (1.00) |
0.323 (1.00) |
0.344 (1.00) |
0.277 (1.00) |
0.146 (1.00) |
0.267 (1.00) |
| amp 20q11 21 | 48 (86%) | 8 |
0.00432 (1.00) |
0.167 (1.00) |
0.546 (1.00) |
0.422 (1.00) |
0.162 (1.00) |
0.14 (1.00) |
0.0566 (1.00) |
0.274 (1.00) |
| amp 20q11 21 | 48 (86%) | 8 |
0.102 (1.00) |
0.255 (1.00) |
0.194 (1.00) |
0.314 (1.00) |
0.178 (1.00) |
0.35 (1.00) |
0.374 (1.00) |
0.559 (1.00) |
| amp xp11 21 | 28 (50%) | 28 |
0.057 (1.00) |
0.565 (1.00) |
0.0471 (1.00) |
0.302 (1.00) |
0.0274 (1.00) |
0.101 (1.00) |
0.113 (1.00) |
0.0647 (1.00) |
| del 1p36 21 | 19 (34%) | 37 |
0.728 (1.00) |
0.377 (1.00) |
0.346 (1.00) |
0.0817 (1.00) |
0.134 (1.00) |
0.0233 (1.00) |
0.0719 (1.00) |
0.0598 (1.00) |
| del 2q22 1 | 11 (20%) | 45 |
0.917 (1.00) |
0.542 (1.00) |
0.527 (1.00) |
0.171 (1.00) |
0.552 (1.00) |
0.557 (1.00) |
0.257 (1.00) |
0.921 (1.00) |
| del 3p14 2 | 37 (66%) | 19 |
0.73 (1.00) |
0.7 (1.00) |
0.811 (1.00) |
0.473 (1.00) |
0.626 (1.00) |
0.766 (1.00) |
0.87 (1.00) |
0.902 (1.00) |
| del 3q13 31 | 22 (39%) | 34 |
0.206 (1.00) |
0.0455 (1.00) |
0.88 (1.00) |
0.0683 (1.00) |
1 (1.00) |
0.461 (1.00) |
0.354 (1.00) |
0.821 (1.00) |
| del 4q22 1 | 37 (66%) | 19 |
0.164 (1.00) |
0.0561 (1.00) |
0.0244 (1.00) |
0.154 (1.00) |
0.0126 (1.00) |
0.127 (1.00) |
0.0319 (1.00) |
0.0573 (1.00) |
| del 4q34 3 | 37 (66%) | 19 |
0.0176 (1.00) |
0.0198 (1.00) |
0.0121 (1.00) |
0.0409 (1.00) |
0.00311 (1.00) |
0.199 (1.00) |
0.0048 (1.00) |
0.124 (1.00) |
| del 5q12 1 | 26 (46%) | 30 |
0.629 (1.00) |
0.0924 (1.00) |
0.587 (1.00) |
0.453 (1.00) |
0.548 (1.00) |
0.296 (1.00) |
0.255 (1.00) |
0.444 (1.00) |
| del 6q26 | 10 (18%) | 46 |
0.417 (1.00) |
0.544 (1.00) |
0.123 (1.00) |
0.7 (1.00) |
0.331 (1.00) |
0.301 (1.00) |
0.817 (1.00) |
0.316 (1.00) |
| del 7q11 22 | 12 (21%) | 44 |
0.714 (1.00) |
0.506 (1.00) |
0.76 (1.00) |
0.835 (1.00) |
0.861 (1.00) |
0.533 (1.00) |
0.841 (1.00) |
0.751 (1.00) |
| del 7q36 2 | 20 (36%) | 36 |
0.326 (1.00) |
0.169 (1.00) |
0.497 (1.00) |
0.276 (1.00) |
0.402 (1.00) |
0.0438 (1.00) |
0.324 (1.00) |
0.0822 (1.00) |
| del 8p23 2 | 33 (59%) | 23 |
0.0018 (0.868) |
0.429 (1.00) |
0.352 (1.00) |
0.359 (1.00) |
0.739 (1.00) |
0.329 (1.00) |
0.598 (1.00) |
0.561 (1.00) |
| del 9p23 | 34 (61%) | 22 |
0.296 (1.00) |
0.122 (1.00) |
0.636 (1.00) |
0.32 (1.00) |
0.501 (1.00) |
0.425 (1.00) |
0.489 (1.00) |
0.121 (1.00) |
| del 9q21 13 | 39 (70%) | 17 |
0.34 (1.00) |
0.325 (1.00) |
0.209 (1.00) |
0.258 (1.00) |
0.479 (1.00) |
0.319 (1.00) |
0.744 (1.00) |
0.399 (1.00) |
| del 9q33 3 | 39 (70%) | 17 |
0.668 (1.00) |
0.0741 (1.00) |
0.113 (1.00) |
0.14 (1.00) |
0.967 (1.00) |
0.113 (1.00) |
1 (1.00) |
0.26 (1.00) |
| del 10q23 31 | 20 (36%) | 36 |
0.151 (1.00) |
0.106 (1.00) |
0.123 (1.00) |
0.241 (1.00) |
0.763 (1.00) |
0.853 (1.00) |
0.769 (1.00) |
0.826 (1.00) |
| del 11p15 5 | 25 (45%) | 31 |
0.59 (1.00) |
0.769 (1.00) |
0.899 (1.00) |
0.896 (1.00) |
0.724 (1.00) |
0.896 (1.00) |
0.828 (1.00) |
0.917 (1.00) |
| del 11q14 1 | 26 (46%) | 30 |
0.444 (1.00) |
0.17 (1.00) |
1 (1.00) |
0.923 (1.00) |
0.27 (1.00) |
0.264 (1.00) |
0.227 (1.00) |
0.074 (1.00) |
| del 11q23 2 | 26 (46%) | 30 |
1 (1.00) |
0.114 (1.00) |
0.343 (1.00) |
0.87 (1.00) |
0.358 (1.00) |
0.86 (1.00) |
0.397 (1.00) |
0.359 (1.00) |
| del 12q23 1 | 20 (36%) | 36 |
0.882 (1.00) |
0.9 (1.00) |
0.655 (1.00) |
0.753 (1.00) |
0.95 (1.00) |
0.342 (1.00) |
1 (1.00) |
0.544 (1.00) |
| del 12q24 31 | 20 (36%) | 36 |
0.443 (1.00) |
0.824 (1.00) |
0.154 (1.00) |
0.259 (1.00) |
0.561 (1.00) |
0.889 (1.00) |
0.368 (1.00) |
0.931 (1.00) |
| del 13q12 11 | 30 (54%) | 26 |
0.294 (1.00) |
0.0197 (1.00) |
0.375 (1.00) |
0.208 (1.00) |
0.727 (1.00) |
0.0254 (1.00) |
0.346 (1.00) |
0.0629 (1.00) |
| del 14q21 1 | 28 (50%) | 28 |
0.891 (1.00) |
0.0508 (1.00) |
0.0635 (1.00) |
0.0554 (1.00) |
0.378 (1.00) |
0.384 (1.00) |
0.835 (1.00) |
0.365 (1.00) |
| del 15q11 2 | 41 (73%) | 15 |
0.748 (1.00) |
0.0969 (1.00) |
0.552 (1.00) |
0.622 (1.00) |
0.143 (1.00) |
0.908 (1.00) |
0.325 (1.00) |
0.705 (1.00) |
| del 15q15 1 | 43 (77%) | 13 |
0.165 (1.00) |
0.0308 (1.00) |
0.348 (1.00) |
0.768 (1.00) |
0.12 (1.00) |
0.349 (1.00) |
0.0668 (1.00) |
0.208 (1.00) |
| del 16p13 3 | 33 (59%) | 23 |
0.663 (1.00) |
0.55 (1.00) |
0.572 (1.00) |
0.228 (1.00) |
0.35 (1.00) |
0.391 (1.00) |
0.383 (1.00) |
0.716 (1.00) |
| del 16q23 1 | 39 (70%) | 17 |
0.626 (1.00) |
0.424 (1.00) |
0.396 (1.00) |
0.394 (1.00) |
0.864 (1.00) |
0.812 (1.00) |
0.171 (1.00) |
0.622 (1.00) |
| del 17p13 1 | 42 (75%) | 14 |
0.139 (1.00) |
0.0553 (1.00) |
0.0568 (1.00) |
0.42 (1.00) |
0.133 (1.00) |
0.183 (1.00) |
0.0198 (1.00) |
0.0854 (1.00) |
| del 17q21 32 | 19 (34%) | 37 |
0.148 (1.00) |
0.764 (1.00) |
0.179 (1.00) |
0.0675 (1.00) |
0.223 (1.00) |
0.212 (1.00) |
0.387 (1.00) |
0.737 (1.00) |
| del 18q22 2 | 26 (46%) | 30 |
0.332 (1.00) |
0.341 (1.00) |
0.505 (1.00) |
0.361 (1.00) |
0.0215 (1.00) |
0.0777 (1.00) |
0.0134 (1.00) |
0.024 (1.00) |
| del 19p13 3 | 47 (84%) | 9 |
0.0613 (1.00) |
0.0203 (1.00) |
0.0973 (1.00) |
0.376 (1.00) |
0.0148 (1.00) |
0.0855 (1.00) |
0.0094 (1.00) |
0.112 (1.00) |
| del 19q13 33 | 17 (30%) | 39 |
0.316 (1.00) |
0.00906 (1.00) |
0.00066 (0.32) |
0.0346 (1.00) |
0.222 (1.00) |
0.0384 (1.00) |
0.368 (1.00) |
0.0128 (1.00) |
| del 20p12 1 | 9 (16%) | 47 |
0.538 (1.00) |
0.703 (1.00) |
0.391 (1.00) |
0.526 (1.00) |
0.219 (1.00) |
0.341 (1.00) |
0.366 (1.00) |
0.466 (1.00) |
| del 22q13 31 | 40 (71%) | 16 |
0.298 (1.00) |
0.41 (1.00) |
0.894 (1.00) |
0.461 (1.00) |
0.58 (1.00) |
0.351 (1.00) |
0.858 (1.00) |
0.589 (1.00) |
| del xp21 1 | 18 (32%) | 38 |
0.937 (1.00) |
0.421 (1.00) |
0.819 (1.00) |
0.868 (1.00) |
0.361 (1.00) |
1 (1.00) |
1 (1.00) |
0.9 (1.00) |
| del xq25 | 23 (41%) | 33 |
0.626 (1.00) |
0.31 (1.00) |
0.114 (1.00) |
0.898 (1.00) |
0.73 (1.00) |
0.568 (1.00) |
0.279 (1.00) |
0.384 (1.00) |
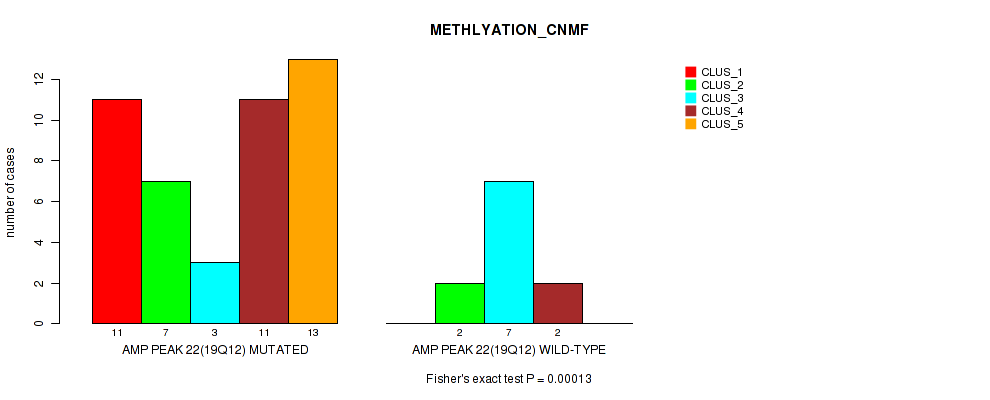
P value = 0.00013 (Fisher's exact test), Q value = 0.063
Table S1. Gene #22: 'amp_19q12' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 11 | 9 | 10 | 13 | 13 |
| AMP PEAK 22(19Q12) MUTATED | 11 | 7 | 3 | 11 | 13 |
| AMP PEAK 22(19Q12) WILD-TYPE | 0 | 2 | 7 | 2 | 0 |
Figure S1. Get High-res Image Gene #22: 'amp_19q12' versus Molecular Subtype #2: 'METHLYATION_CNMF'
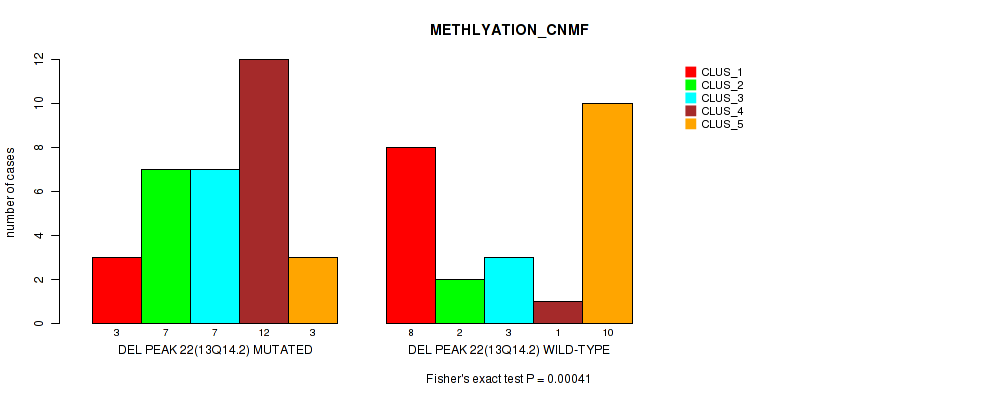
P value = 0.00041 (Fisher's exact test), Q value = 0.2
Table S2. Gene #47: 'del_13q14.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 11 | 9 | 10 | 13 | 13 |
| DEL PEAK 22(13Q14.2) MUTATED | 3 | 7 | 7 | 12 | 3 |
| DEL PEAK 22(13Q14.2) WILD-TYPE | 8 | 2 | 3 | 1 | 10 |
Figure S2. Get High-res Image Gene #47: 'del_13q14.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtype file = UCS-TP.transferedmergedcluster.txt
-
Number of patients = 56
-
Number of significantly focal cnvs = 61
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.