This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 21 focal events and 8 molecular subtypes across 87 patients, 52 significant findings detected with P value < 0.05 and Q value < 0.25.
-
del_1p36.31 cnv correlated to 'CN_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
del_1p21.3 cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_2q35 cnv correlated to 'MRNASEQ_CHIERARCHICAL'.
-
del_3p21.1 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_4q26 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_4q34.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_5q23.2 cnv correlated to 'CN_CNMF'.
-
del_6q22.1 cnv correlated to 'CN_CNMF' and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_6q26 cnv correlated to 'CN_CNMF' and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_9p21.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_10p15.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_10q25.2 cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
del_11q23.2 cnv correlated to 'CN_CNMF'.
-
del_12p13.31 cnv correlated to 'METHLYATION_CNMF' and 'MRNASEQ_CNMF'.
-
del_13q14.11 cnv correlated to 'CN_CNMF'.
-
del_14q32.31 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CNMF'.
-
del_15q15.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_16q24.1 cnv correlated to 'CN_CNMF'.
-
del_22q12.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 21 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 52 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| del 14q32 31 | 39 (45%) | 48 |
0.00012 (0.00168) |
0.311 (0.496) |
0.00216 (0.0158) |
1e-05 (0.00042) |
0.0262 (0.0956) |
0.0089 (0.0468) |
0.0426 (0.14) |
0.0795 (0.224) |
| del 3p21 1 | 48 (55%) | 39 |
0.0004 (0.00448) |
0.123 (0.292) |
0.00014 (0.00181) |
2e-05 (0.00048) |
0.202 (0.381) |
0.0153 (0.0694) |
0.275 (0.446) |
0.00787 (0.0468) |
| del 9p21 3 | 51 (59%) | 36 |
1e-05 (0.00042) |
0.00933 (0.0475) |
0.0191 (0.0781) |
3e-05 (0.00063) |
0.96 (0.986) |
0.13 (0.299) |
0.477 (0.661) |
0.0172 (0.0722) |
| del 10p15 3 | 28 (32%) | 59 |
0.00176 (0.0134) |
0.00665 (0.043) |
0.0329 (0.111) |
0.00892 (0.0468) |
1 (1.00) |
0.445 (0.634) |
0.355 (0.54) |
0.444 (0.634) |
| del 15q15 1 | 28 (32%) | 59 |
2e-05 (0.00048) |
0.00118 (0.0104) |
0.0275 (0.0961) |
0.0851 (0.23) |
0.36 (0.54) |
0.525 (0.695) |
0.725 (0.823) |
0.0209 (0.0835) |
| del 1p36 31 | 33 (38%) | 54 |
1e-05 (0.00042) |
0.184 (0.359) |
0.329 (0.516) |
0.0216 (0.0845) |
0.0817 (0.225) |
0.00872 (0.0468) |
0.433 (0.632) |
0.0713 (0.206) |
| del 1p21 3 | 37 (43%) | 50 |
0.108 (0.267) |
0.0269 (0.0961) |
0.00062 (0.00613) |
0.0157 (0.0694) |
0.926 (0.972) |
0.313 (0.496) |
0.593 (0.755) |
0.554 (0.721) |
| del 4q26 | 38 (44%) | 49 |
2e-05 (0.00048) |
0.0561 (0.171) |
0.00236 (0.0165) |
0.00082 (0.00765) |
0.276 (0.446) |
0.0516 (0.164) |
0.591 (0.755) |
0.178 (0.353) |
| del 4q34 3 | 41 (47%) | 46 |
1e-05 (0.00042) |
0.0169 (0.0722) |
0.0612 (0.184) |
0.00012 (0.00168) |
0.177 (0.353) |
0.242 (0.415) |
0.961 (0.986) |
0.132 (0.299) |
| del 22q12 2 | 68 (78%) | 19 |
0.00129 (0.0108) |
5e-05 (0.00084) |
0.106 (0.265) |
0.014 (0.0672) |
0.896 (0.953) |
0.627 (0.776) |
0.0799 (0.224) |
0.679 (0.804) |
| del 6q22 1 | 41 (47%) | 46 |
0.00147 (0.0118) |
0.241 (0.415) |
0.157 (0.33) |
0.492 (0.663) |
0.559 (0.723) |
0.2 (0.381) |
0.258 (0.425) |
0.0144 (0.0672) |
| del 6q26 | 36 (41%) | 51 |
5e-05 (0.00084) |
0.254 (0.423) |
0.839 (0.909) |
0.799 (0.888) |
0.345 (0.534) |
0.493 (0.663) |
0.347 (0.534) |
0.0435 (0.141) |
| del 10q25 2 | 31 (36%) | 56 |
0.00272 (0.0183) |
0.00806 (0.0468) |
0.483 (0.661) |
0.455 (0.642) |
0.717 (0.823) |
0.194 (0.374) |
0.232 (0.415) |
0.148 (0.318) |
| del 12p13 31 | 11 (13%) | 76 |
0.117 (0.28) |
0.0125 (0.0617) |
0.0244 (0.093) |
0.15 (0.319) |
0.25 (0.42) |
0.104 (0.265) |
1 (1.00) |
0.23 (0.415) |
| del 2q35 | 17 (20%) | 70 |
0.0889 (0.233) |
0.0533 (0.166) |
0.553 (0.721) |
0.0082 (0.0468) |
0.484 (0.661) |
0.0861 (0.23) |
0.68 (0.804) |
0.36 (0.54) |
| del 5q23 2 | 17 (20%) | 70 |
0.00024 (0.00288) |
0.378 (0.557) |
0.719 (0.823) |
0.138 (0.308) |
0.742 (0.831) |
0.436 (0.632) |
1 (1.00) |
1 (1.00) |
| del 11q23 2 | 18 (21%) | 69 |
0.00056 (0.00588) |
0.715 (0.823) |
0.139 (0.308) |
0.215 (0.394) |
0.469 (0.657) |
0.131 (0.299) |
0.739 (0.831) |
0.111 (0.27) |
| del 13q14 11 | 43 (49%) | 44 |
0.0297 (0.102) |
0.179 (0.353) |
0.216 (0.394) |
0.247 (0.419) |
0.963 (0.986) |
0.525 (0.695) |
0.172 (0.353) |
0.91 (0.961) |
| del 16q24 1 | 22 (25%) | 65 |
0.026 (0.0956) |
0.675 (0.804) |
0.242 (0.415) |
0.365 (0.543) |
0.605 (0.764) |
0.165 (0.343) |
0.812 (0.898) |
0.212 (0.394) |
| del 8p23 2 | 18 (21%) | 69 |
0.865 (0.93) |
0.725 (0.823) |
0.702 (0.823) |
0.937 (0.977) |
0.63 (0.776) |
0.834 (0.909) |
0.839 (0.909) |
0.633 (0.776) |
| del 16p13 3 | 6 (7%) | 81 |
0.07 (0.206) |
0.238 (0.415) |
0.143 (0.313) |
0.0988 (0.255) |
0.656 (0.796) |
0.658 (0.796) |
0.869 (0.93) |
0.621 (0.776) |
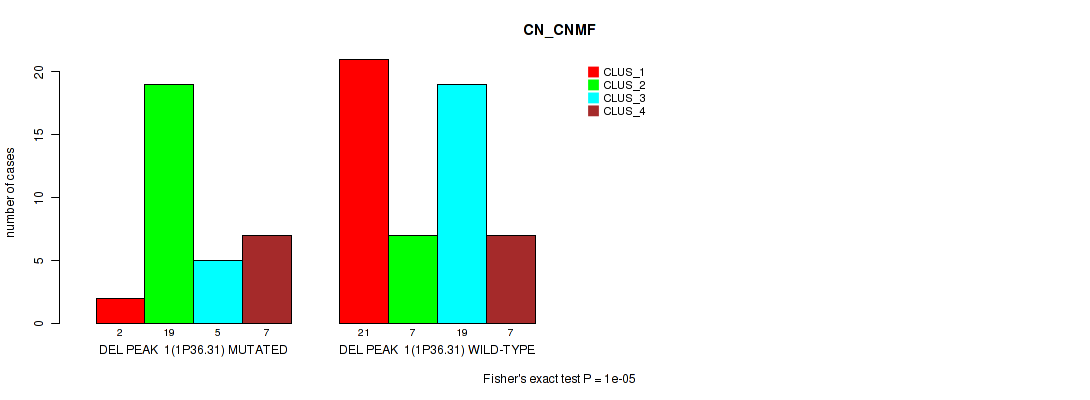
P value = 1e-05 (Fisher's exact test), Q value = 0.00042
Table S1. Gene #1: 'del_1p36.31' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 1(1P36.31) MUTATED | 2 | 19 | 5 | 7 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 21 | 7 | 19 | 7 |
Figure S1. Get High-res Image Gene #1: 'del_1p36.31' versus Molecular Subtype #1: 'CN_CNMF'
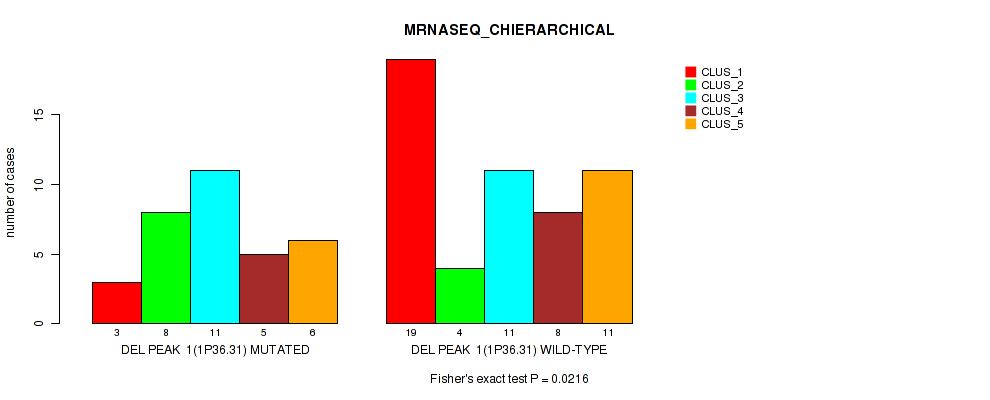
P value = 0.0216 (Fisher's exact test), Q value = 0.084
Table S2. Gene #1: 'del_1p36.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 22 | 12 | 22 | 13 | 17 |
| DEL PEAK 1(1P36.31) MUTATED | 3 | 8 | 11 | 5 | 6 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 19 | 4 | 11 | 8 | 11 |
Figure S2. Get High-res Image Gene #1: 'del_1p36.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
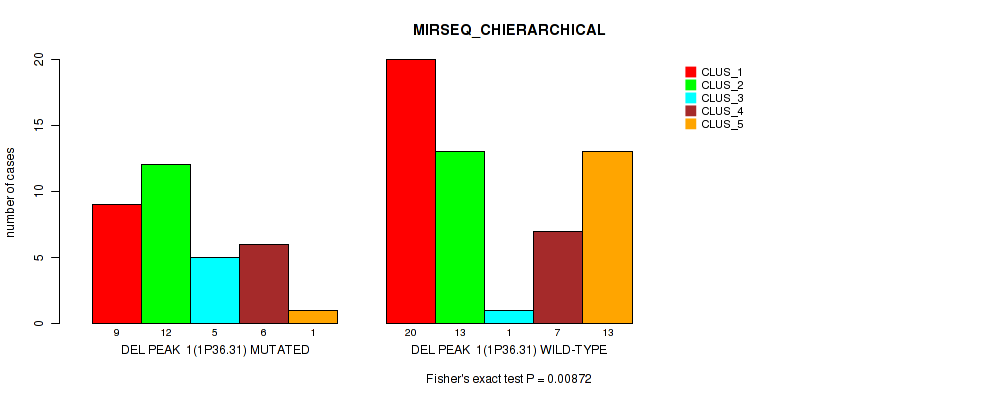
P value = 0.00872 (Fisher's exact test), Q value = 0.047
Table S3. Gene #1: 'del_1p36.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 29 | 25 | 6 | 13 | 14 |
| DEL PEAK 1(1P36.31) MUTATED | 9 | 12 | 5 | 6 | 1 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 20 | 13 | 1 | 7 | 13 |
Figure S3. Get High-res Image Gene #1: 'del_1p36.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
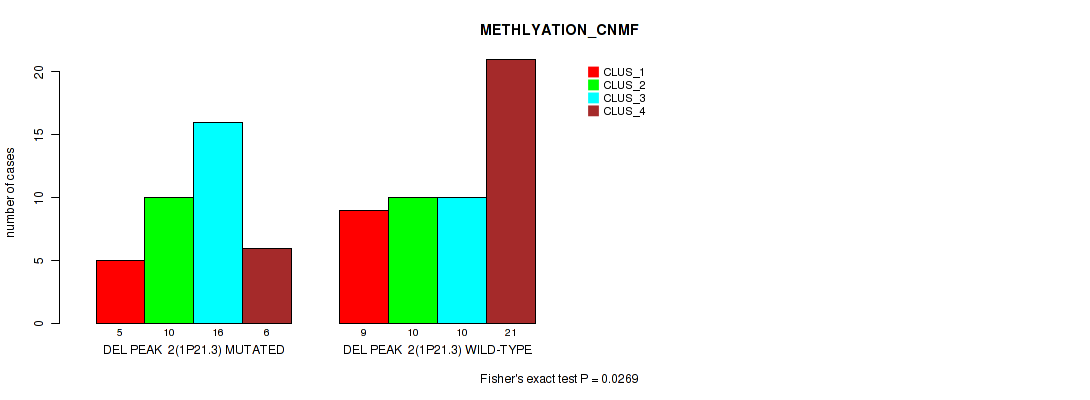
P value = 0.0269 (Fisher's exact test), Q value = 0.096
Table S4. Gene #2: 'del_1p21.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 14 | 20 | 26 | 27 |
| DEL PEAK 2(1P21.3) MUTATED | 5 | 10 | 16 | 6 |
| DEL PEAK 2(1P21.3) WILD-TYPE | 9 | 10 | 10 | 21 |
Figure S4. Get High-res Image Gene #2: 'del_1p21.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
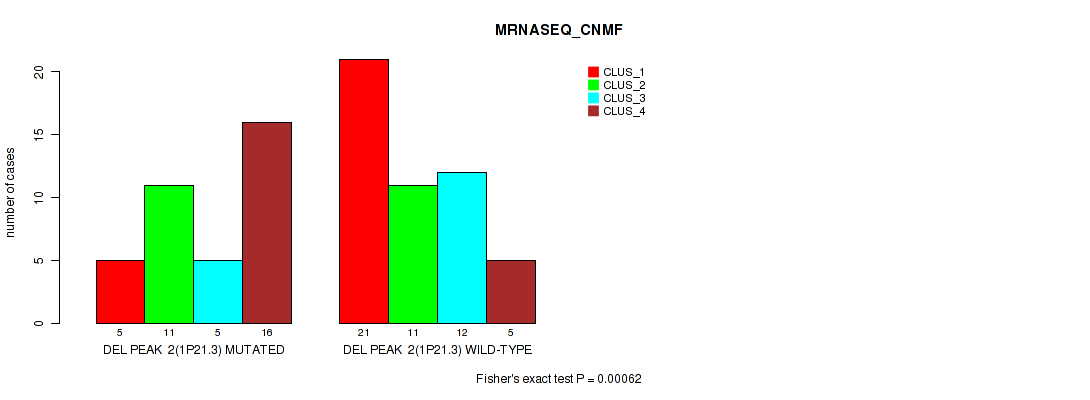
P value = 0.00062 (Fisher's exact test), Q value = 0.0061
Table S5. Gene #2: 'del_1p21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 26 | 22 | 17 | 21 |
| DEL PEAK 2(1P21.3) MUTATED | 5 | 11 | 5 | 16 |
| DEL PEAK 2(1P21.3) WILD-TYPE | 21 | 11 | 12 | 5 |
Figure S5. Get High-res Image Gene #2: 'del_1p21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
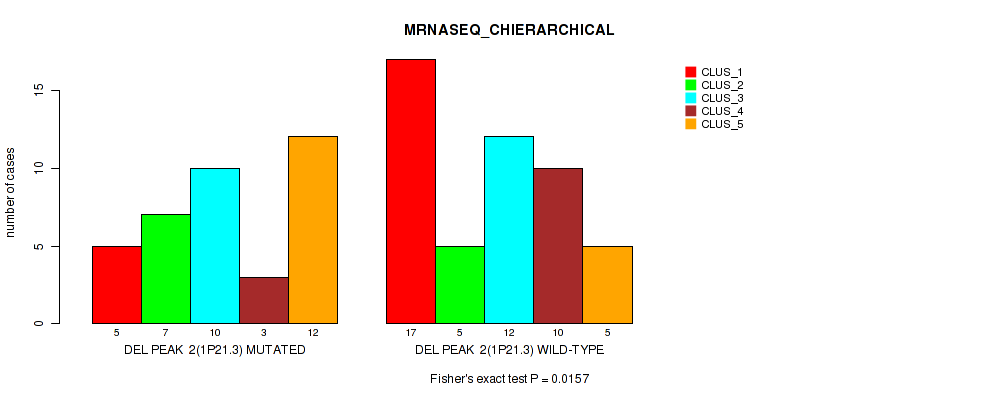
P value = 0.0157 (Fisher's exact test), Q value = 0.069
Table S6. Gene #2: 'del_1p21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 22 | 12 | 22 | 13 | 17 |
| DEL PEAK 2(1P21.3) MUTATED | 5 | 7 | 10 | 3 | 12 |
| DEL PEAK 2(1P21.3) WILD-TYPE | 17 | 5 | 12 | 10 | 5 |
Figure S6. Get High-res Image Gene #2: 'del_1p21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
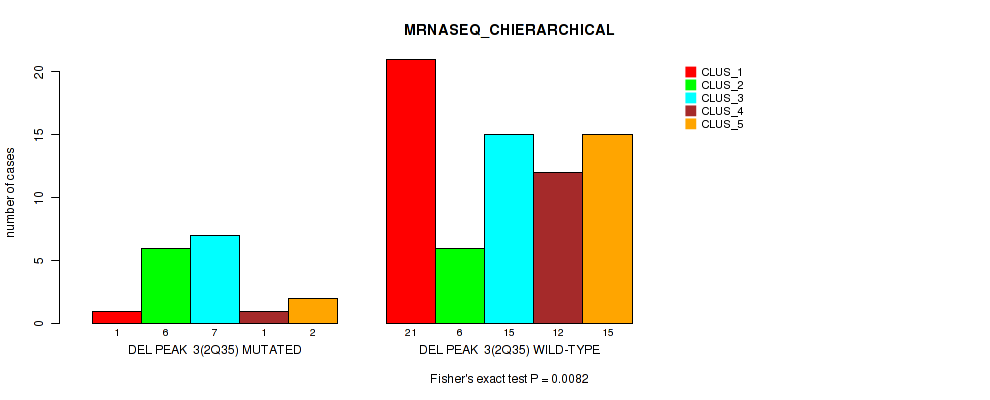
P value = 0.0082 (Fisher's exact test), Q value = 0.047
Table S7. Gene #3: 'del_2q35' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 22 | 12 | 22 | 13 | 17 |
| DEL PEAK 3(2Q35) MUTATED | 1 | 6 | 7 | 1 | 2 |
| DEL PEAK 3(2Q35) WILD-TYPE | 21 | 6 | 15 | 12 | 15 |
Figure S7. Get High-res Image Gene #3: 'del_2q35' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
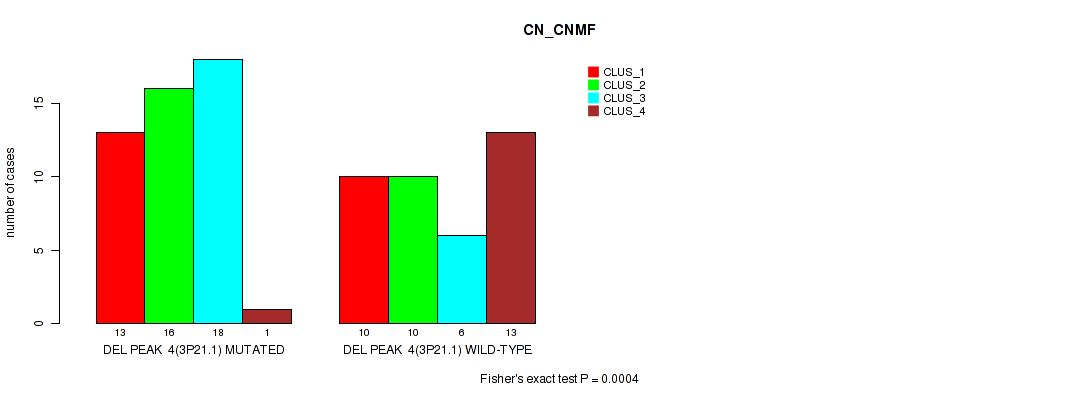
P value = 4e-04 (Fisher's exact test), Q value = 0.0045
Table S8. Gene #4: 'del_3p21.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 4(3P21.1) MUTATED | 13 | 16 | 18 | 1 |
| DEL PEAK 4(3P21.1) WILD-TYPE | 10 | 10 | 6 | 13 |
Figure S8. Get High-res Image Gene #4: 'del_3p21.1' versus Molecular Subtype #1: 'CN_CNMF'
P value = 0.00014 (Fisher's exact test), Q value = 0.0018
Table S9. Gene #4: 'del_3p21.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 26 | 22 | 17 | 21 |
| DEL PEAK 4(3P21.1) MUTATED | 21 | 4 | 10 | 13 |
| DEL PEAK 4(3P21.1) WILD-TYPE | 5 | 18 | 7 | 8 |
Figure S9. Get High-res Image Gene #4: 'del_3p21.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
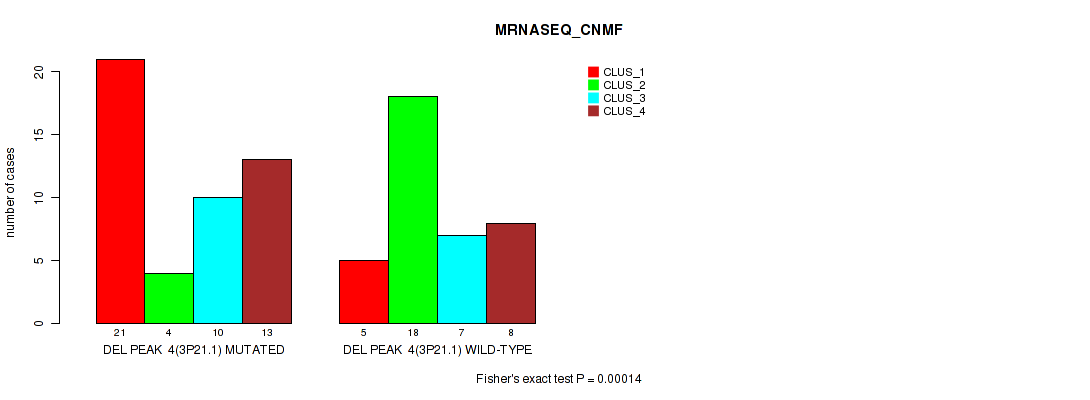
P value = 2e-05 (Fisher's exact test), Q value = 0.00048
Table S10. Gene #4: 'del_3p21.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 22 | 12 | 22 | 13 | 17 |
| DEL PEAK 4(3P21.1) MUTATED | 16 | 12 | 4 | 7 | 9 |
| DEL PEAK 4(3P21.1) WILD-TYPE | 6 | 0 | 18 | 6 | 8 |
Figure S10. Get High-res Image Gene #4: 'del_3p21.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
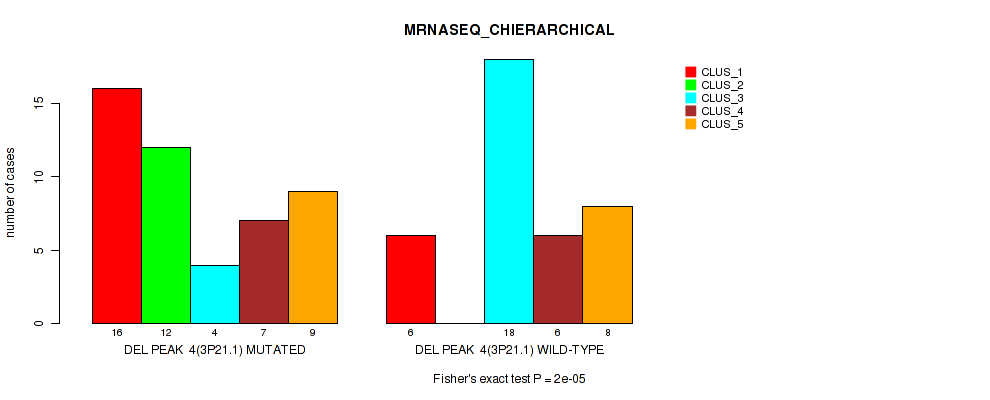
P value = 0.0153 (Fisher's exact test), Q value = 0.069
Table S11. Gene #4: 'del_3p21.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 29 | 25 | 6 | 13 | 14 |
| DEL PEAK 4(3P21.1) MUTATED | 16 | 8 | 3 | 9 | 12 |
| DEL PEAK 4(3P21.1) WILD-TYPE | 13 | 17 | 3 | 4 | 2 |
Figure S11. Get High-res Image Gene #4: 'del_3p21.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
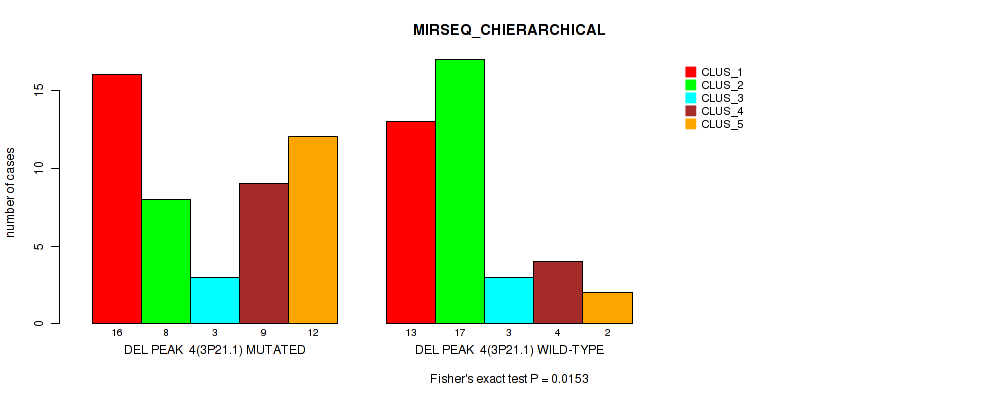
P value = 0.00787 (Fisher's exact test), Q value = 0.047
Table S12. Gene #4: 'del_3p21.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 16 | 17 | 21 | 31 |
| DEL PEAK 4(3P21.1) MUTATED | 13 | 4 | 11 | 19 |
| DEL PEAK 4(3P21.1) WILD-TYPE | 3 | 13 | 10 | 12 |
Figure S12. Get High-res Image Gene #4: 'del_3p21.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
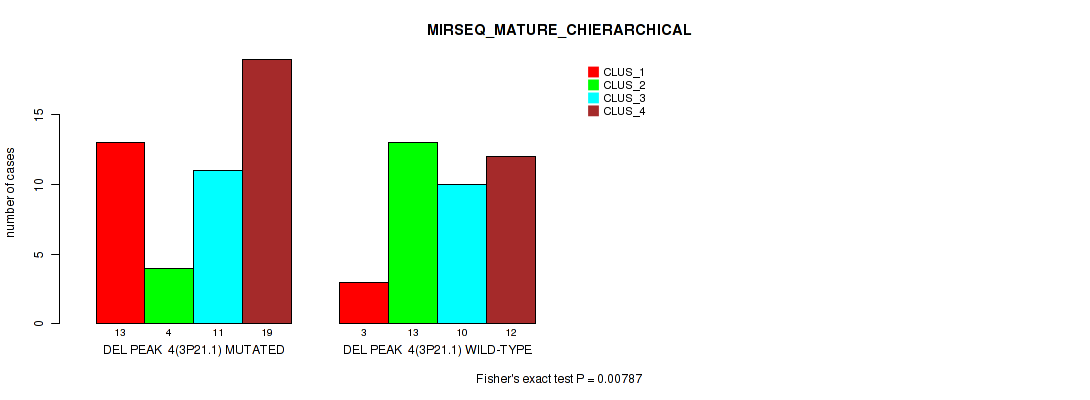
P value = 2e-05 (Fisher's exact test), Q value = 0.00048
Table S13. Gene #5: 'del_4q26' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 5(4Q26) MUTATED | 3 | 21 | 6 | 8 |
| DEL PEAK 5(4Q26) WILD-TYPE | 20 | 5 | 18 | 6 |
Figure S13. Get High-res Image Gene #5: 'del_4q26' versus Molecular Subtype #1: 'CN_CNMF'
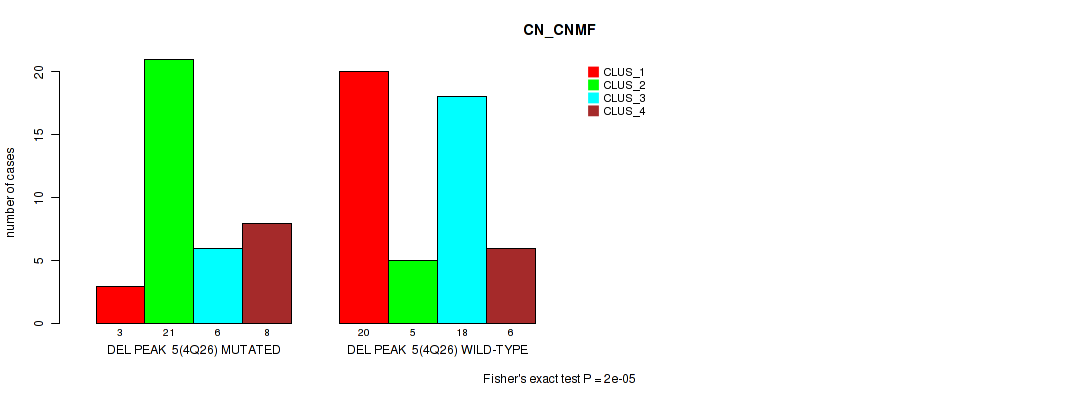
P value = 0.00236 (Fisher's exact test), Q value = 0.017
Table S14. Gene #5: 'del_4q26' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 26 | 22 | 17 | 21 |
| DEL PEAK 5(4Q26) MUTATED | 4 | 11 | 9 | 14 |
| DEL PEAK 5(4Q26) WILD-TYPE | 22 | 11 | 8 | 7 |
Figure S14. Get High-res Image Gene #5: 'del_4q26' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
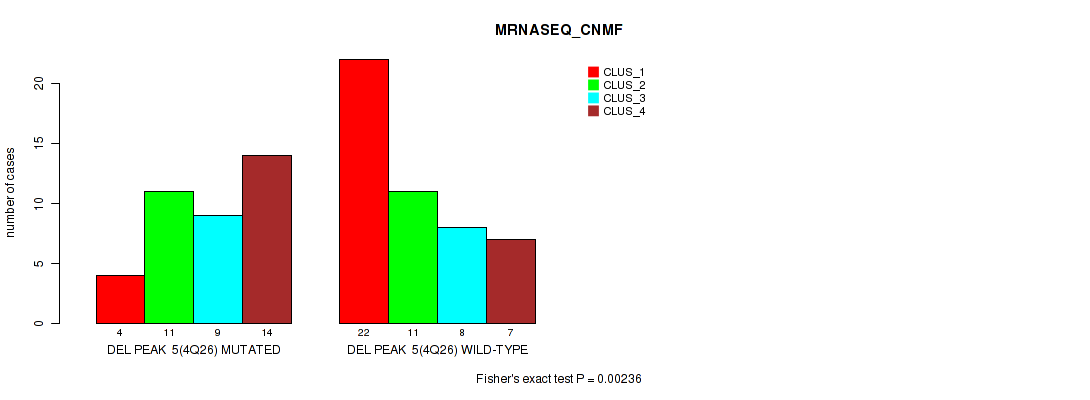
P value = 0.00082 (Fisher's exact test), Q value = 0.0077
Table S15. Gene #5: 'del_4q26' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 22 | 12 | 22 | 13 | 17 |
| DEL PEAK 5(4Q26) MUTATED | 2 | 8 | 12 | 5 | 11 |
| DEL PEAK 5(4Q26) WILD-TYPE | 20 | 4 | 10 | 8 | 6 |
Figure S15. Get High-res Image Gene #5: 'del_4q26' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
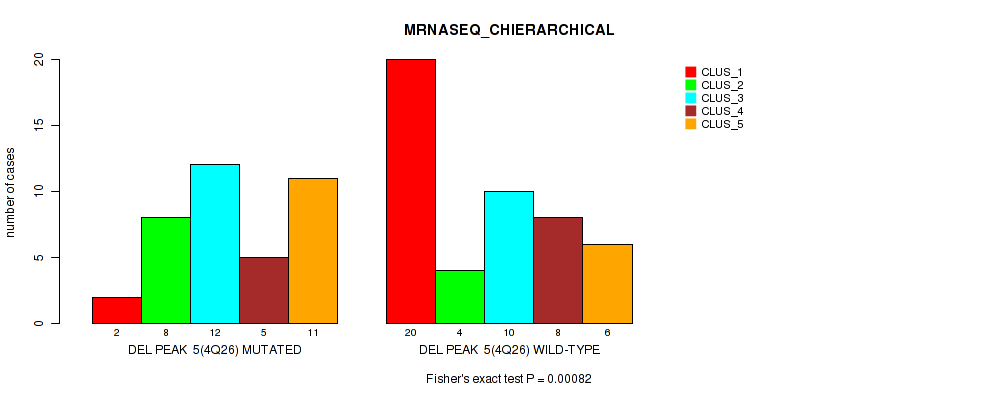
P value = 1e-05 (Fisher's exact test), Q value = 0.00042
Table S16. Gene #6: 'del_4q34.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 6(4Q34.3) MUTATED | 3 | 22 | 7 | 9 |
| DEL PEAK 6(4Q34.3) WILD-TYPE | 20 | 4 | 17 | 5 |
Figure S16. Get High-res Image Gene #6: 'del_4q34.3' versus Molecular Subtype #1: 'CN_CNMF'
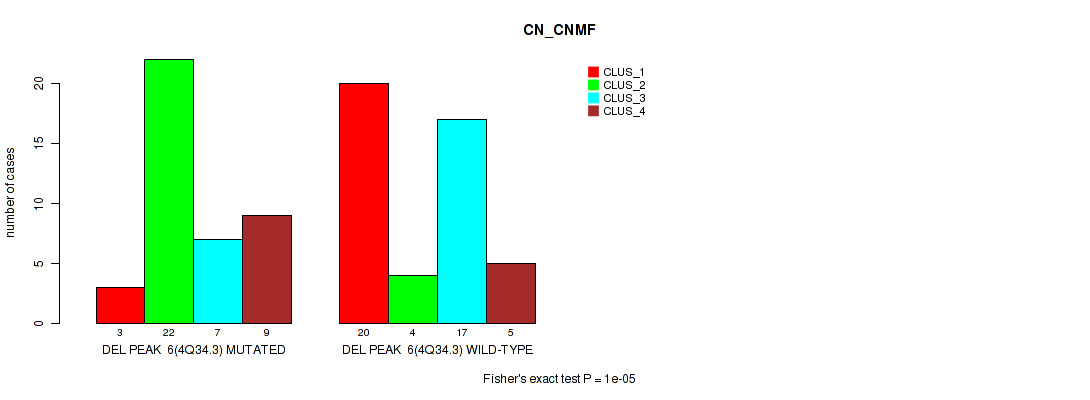
P value = 0.0169 (Fisher's exact test), Q value = 0.072
Table S17. Gene #6: 'del_4q34.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 14 | 20 | 26 | 27 |
| DEL PEAK 6(4Q34.3) MUTATED | 4 | 15 | 13 | 9 |
| DEL PEAK 6(4Q34.3) WILD-TYPE | 10 | 5 | 13 | 18 |
Figure S17. Get High-res Image Gene #6: 'del_4q34.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
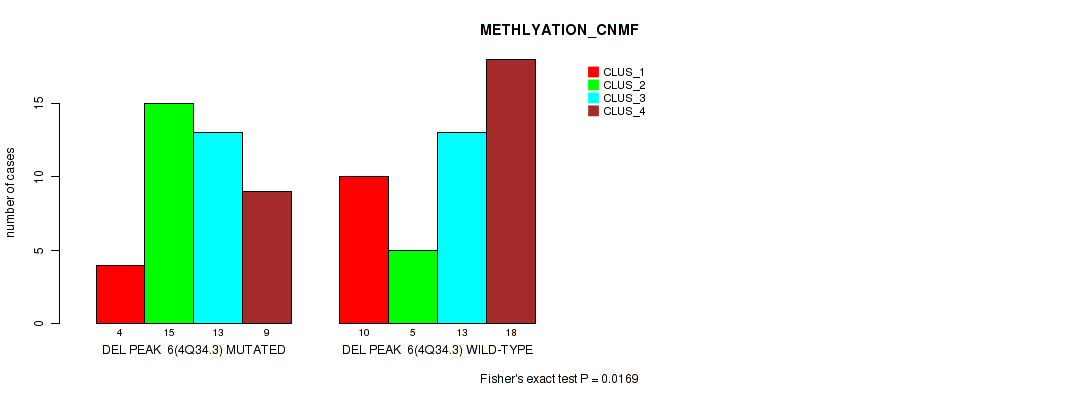
P value = 0.00012 (Fisher's exact test), Q value = 0.0017
Table S18. Gene #6: 'del_4q34.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 22 | 12 | 22 | 13 | 17 |
| DEL PEAK 6(4Q34.3) MUTATED | 2 | 9 | 14 | 5 | 11 |
| DEL PEAK 6(4Q34.3) WILD-TYPE | 20 | 3 | 8 | 8 | 6 |
Figure S18. Get High-res Image Gene #6: 'del_4q34.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
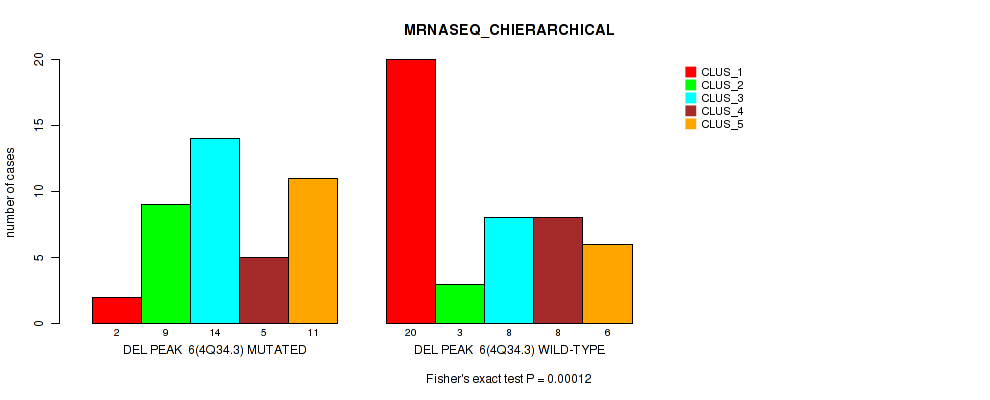
P value = 0.00024 (Fisher's exact test), Q value = 0.0029
Table S19. Gene #7: 'del_5q23.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 7(5Q23.2) MUTATED | 0 | 10 | 7 | 0 |
| DEL PEAK 7(5Q23.2) WILD-TYPE | 23 | 16 | 17 | 14 |
Figure S19. Get High-res Image Gene #7: 'del_5q23.2' versus Molecular Subtype #1: 'CN_CNMF'
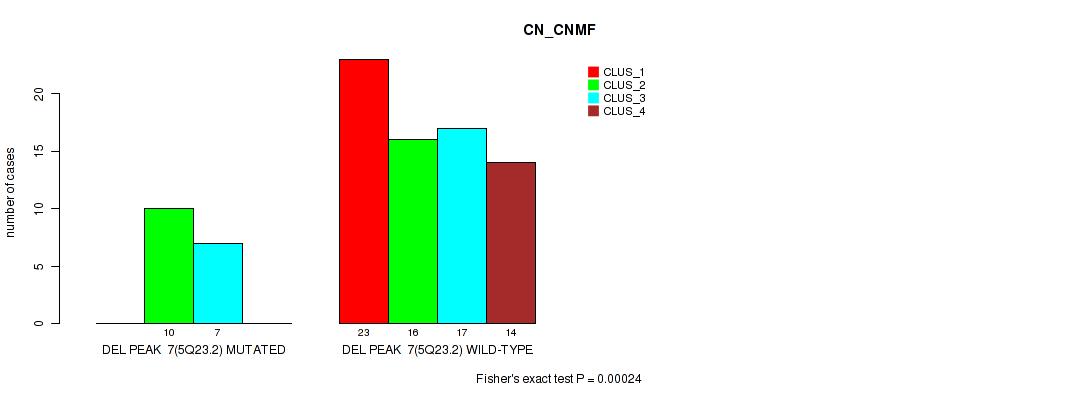
P value = 0.00147 (Fisher's exact test), Q value = 0.012
Table S20. Gene #8: 'del_6q22.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 8(6Q22.1) MUTATED | 12 | 19 | 8 | 2 |
| DEL PEAK 8(6Q22.1) WILD-TYPE | 11 | 7 | 16 | 12 |
Figure S20. Get High-res Image Gene #8: 'del_6q22.1' versus Molecular Subtype #1: 'CN_CNMF'
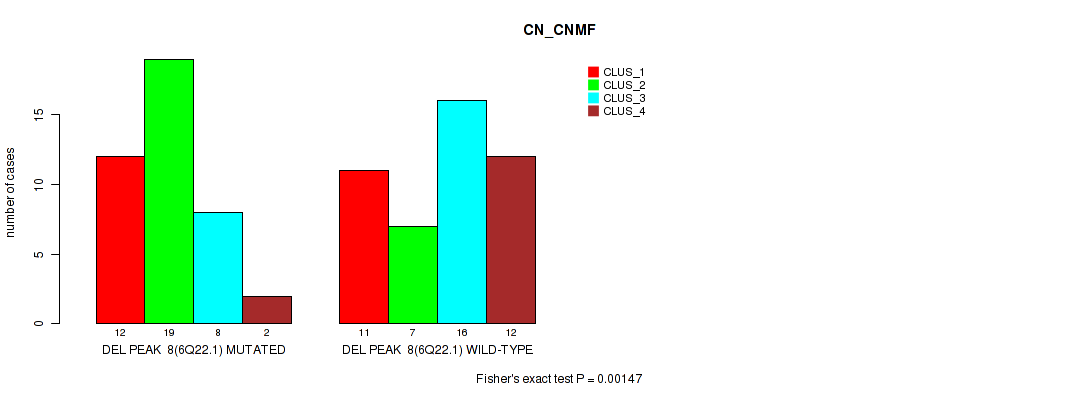
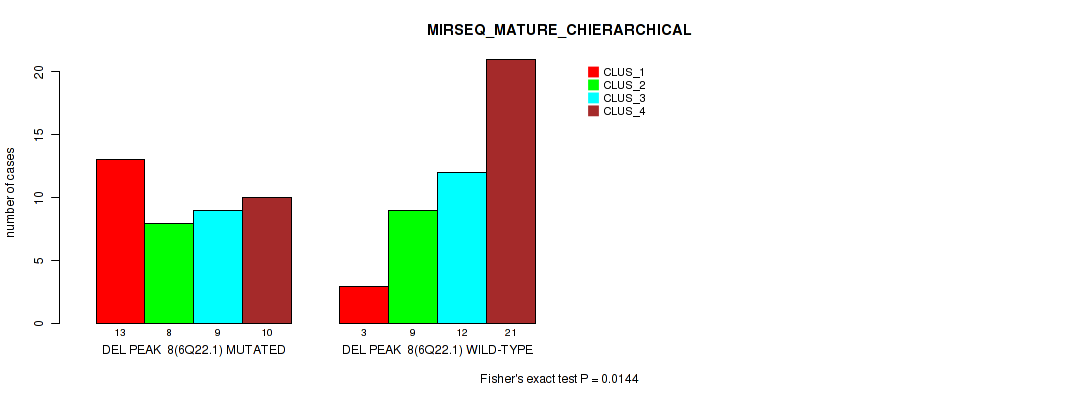
P value = 0.0144 (Fisher's exact test), Q value = 0.067
Table S21. Gene #8: 'del_6q22.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 16 | 17 | 21 | 31 |
| DEL PEAK 8(6Q22.1) MUTATED | 13 | 8 | 9 | 10 |
| DEL PEAK 8(6Q22.1) WILD-TYPE | 3 | 9 | 12 | 21 |
Figure S21. Get High-res Image Gene #8: 'del_6q22.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
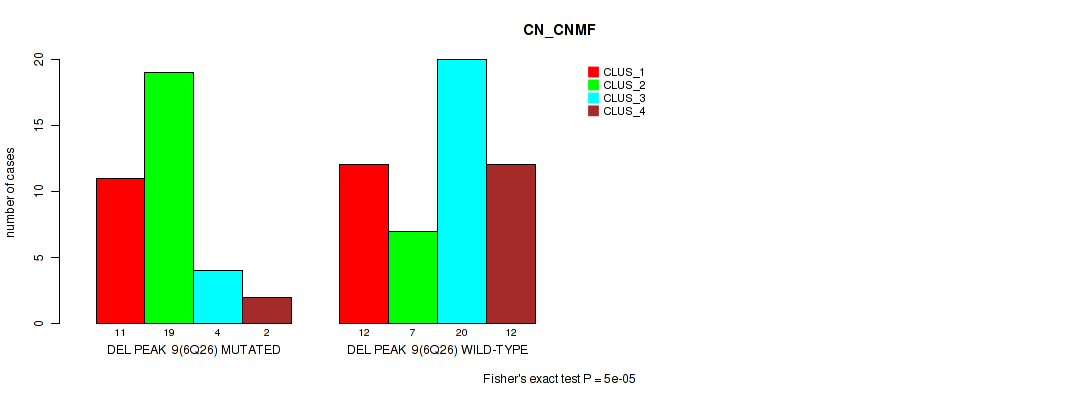
P value = 5e-05 (Fisher's exact test), Q value = 0.00084
Table S22. Gene #9: 'del_6q26' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 9(6Q26) MUTATED | 11 | 19 | 4 | 2 |
| DEL PEAK 9(6Q26) WILD-TYPE | 12 | 7 | 20 | 12 |
Figure S22. Get High-res Image Gene #9: 'del_6q26' versus Molecular Subtype #1: 'CN_CNMF'
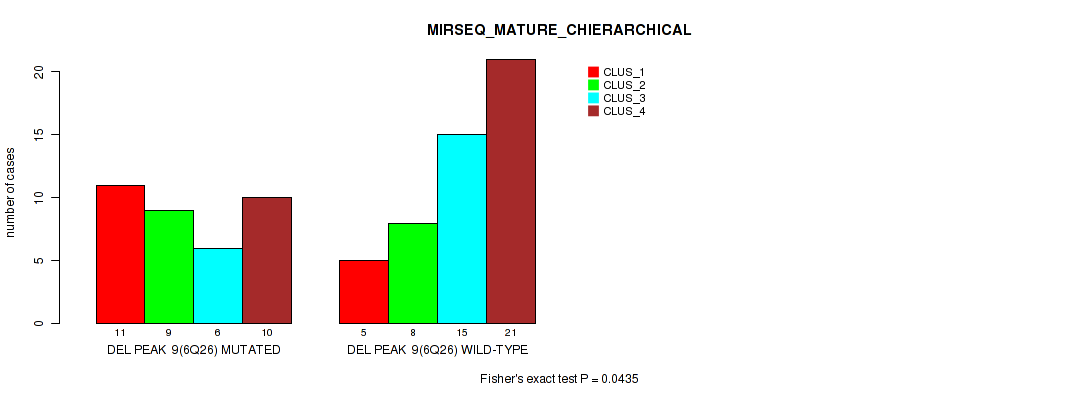
P value = 0.0435 (Fisher's exact test), Q value = 0.14
Table S23. Gene #9: 'del_6q26' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 16 | 17 | 21 | 31 |
| DEL PEAK 9(6Q26) MUTATED | 11 | 9 | 6 | 10 |
| DEL PEAK 9(6Q26) WILD-TYPE | 5 | 8 | 15 | 21 |
Figure S23. Get High-res Image Gene #9: 'del_6q26' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
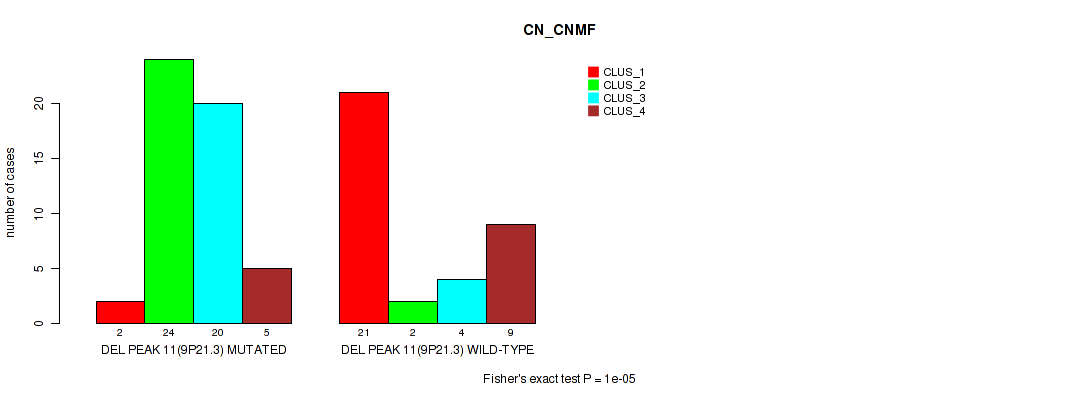
P value = 1e-05 (Fisher's exact test), Q value = 0.00042
Table S24. Gene #11: 'del_9p21.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 11(9P21.3) MUTATED | 2 | 24 | 20 | 5 |
| DEL PEAK 11(9P21.3) WILD-TYPE | 21 | 2 | 4 | 9 |
Figure S24. Get High-res Image Gene #11: 'del_9p21.3' versus Molecular Subtype #1: 'CN_CNMF'
P value = 0.00933 (Fisher's exact test), Q value = 0.047
Table S25. Gene #11: 'del_9p21.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 14 | 20 | 26 | 27 |
| DEL PEAK 11(9P21.3) MUTATED | 6 | 17 | 17 | 11 |
| DEL PEAK 11(9P21.3) WILD-TYPE | 8 | 3 | 9 | 16 |
Figure S25. Get High-res Image Gene #11: 'del_9p21.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
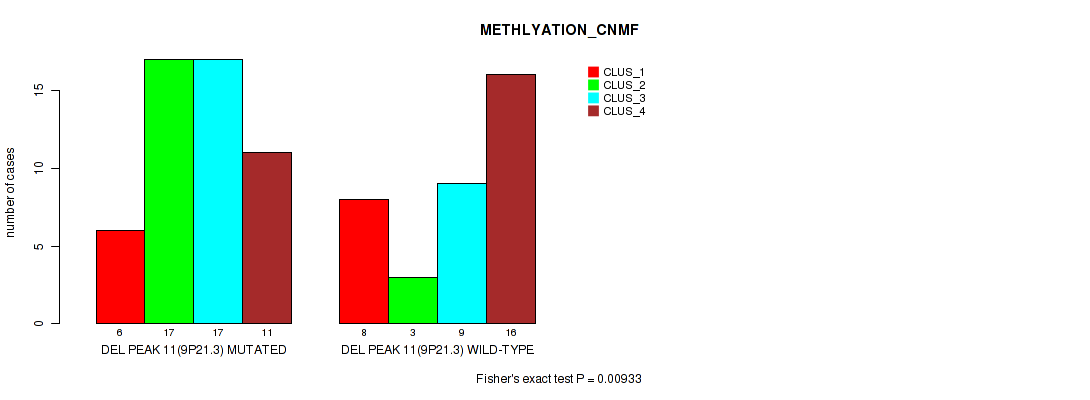
P value = 0.0191 (Fisher's exact test), Q value = 0.078
Table S26. Gene #11: 'del_9p21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 26 | 22 | 17 | 21 |
| DEL PEAK 11(9P21.3) MUTATED | 9 | 14 | 12 | 16 |
| DEL PEAK 11(9P21.3) WILD-TYPE | 17 | 8 | 5 | 5 |
Figure S26. Get High-res Image Gene #11: 'del_9p21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
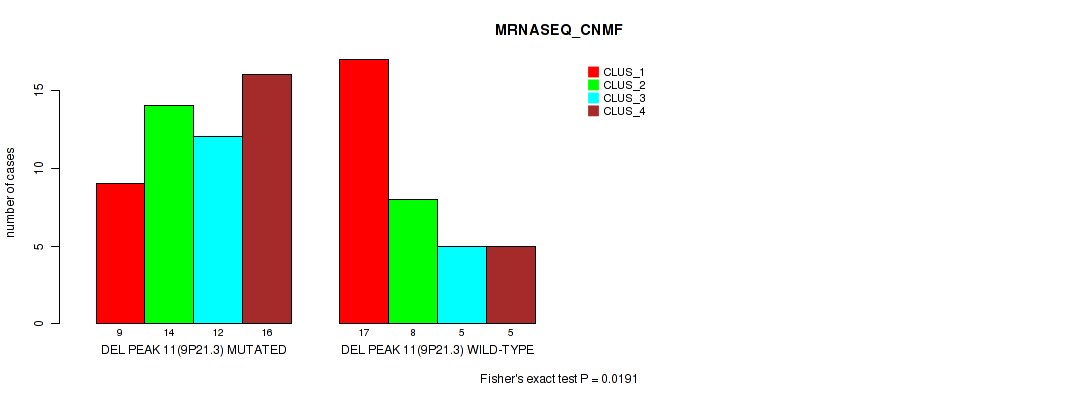
P value = 3e-05 (Fisher's exact test), Q value = 0.00063
Table S27. Gene #11: 'del_9p21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 22 | 12 | 22 | 13 | 17 |
| DEL PEAK 11(9P21.3) MUTATED | 4 | 12 | 14 | 9 | 12 |
| DEL PEAK 11(9P21.3) WILD-TYPE | 18 | 0 | 8 | 4 | 5 |
Figure S27. Get High-res Image Gene #11: 'del_9p21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
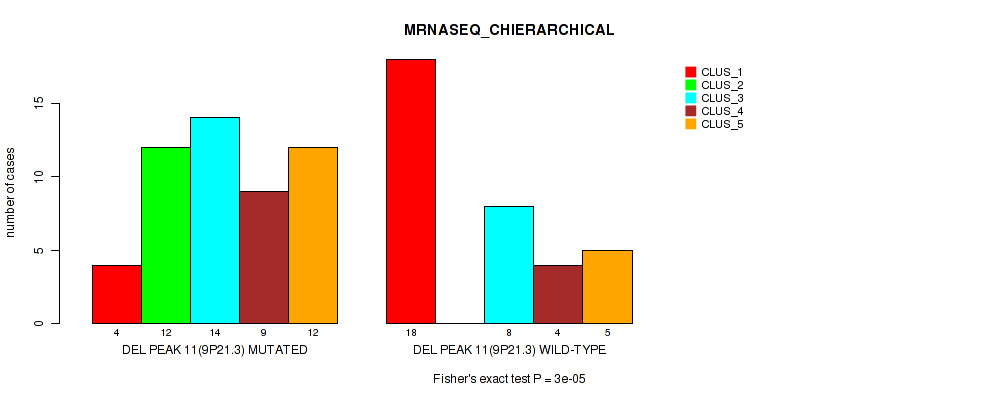
P value = 0.0172 (Fisher's exact test), Q value = 0.072
Table S28. Gene #11: 'del_9p21.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 16 | 17 | 21 | 31 |
| DEL PEAK 11(9P21.3) MUTATED | 4 | 12 | 12 | 22 |
| DEL PEAK 11(9P21.3) WILD-TYPE | 12 | 5 | 9 | 9 |
Figure S28. Get High-res Image Gene #11: 'del_9p21.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
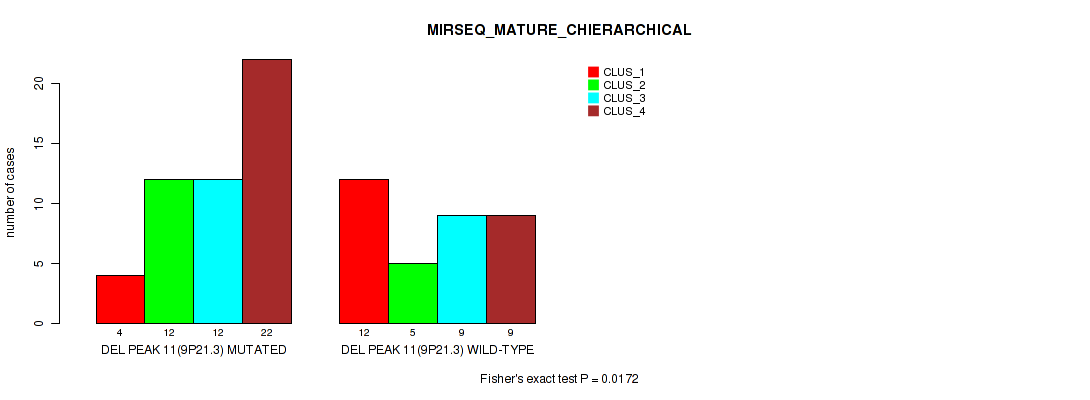
P value = 0.00176 (Fisher's exact test), Q value = 0.013
Table S29. Gene #12: 'del_10p15.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 12(10P15.3) MUTATED | 2 | 14 | 10 | 2 |
| DEL PEAK 12(10P15.3) WILD-TYPE | 21 | 12 | 14 | 12 |
Figure S29. Get High-res Image Gene #12: 'del_10p15.3' versus Molecular Subtype #1: 'CN_CNMF'
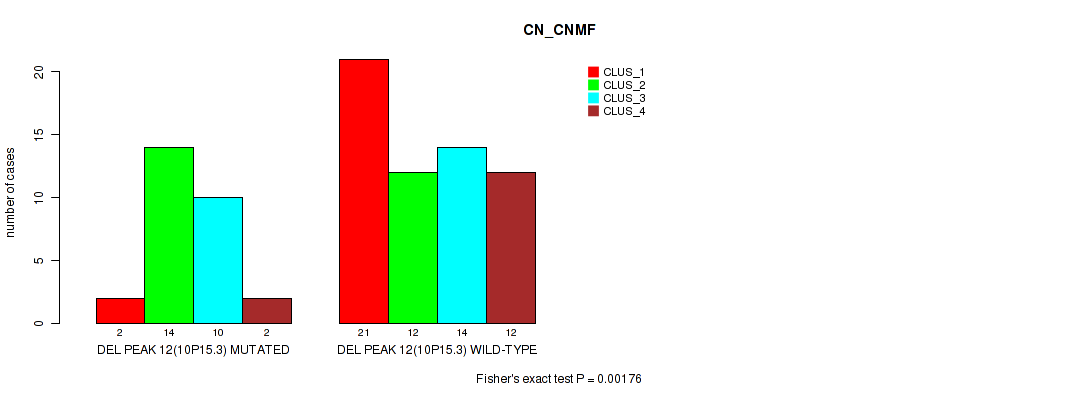
P value = 0.00665 (Fisher's exact test), Q value = 0.043
Table S30. Gene #12: 'del_10p15.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 14 | 20 | 26 | 27 |
| DEL PEAK 12(10P15.3) MUTATED | 1 | 12 | 9 | 6 |
| DEL PEAK 12(10P15.3) WILD-TYPE | 13 | 8 | 17 | 21 |
Figure S30. Get High-res Image Gene #12: 'del_10p15.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
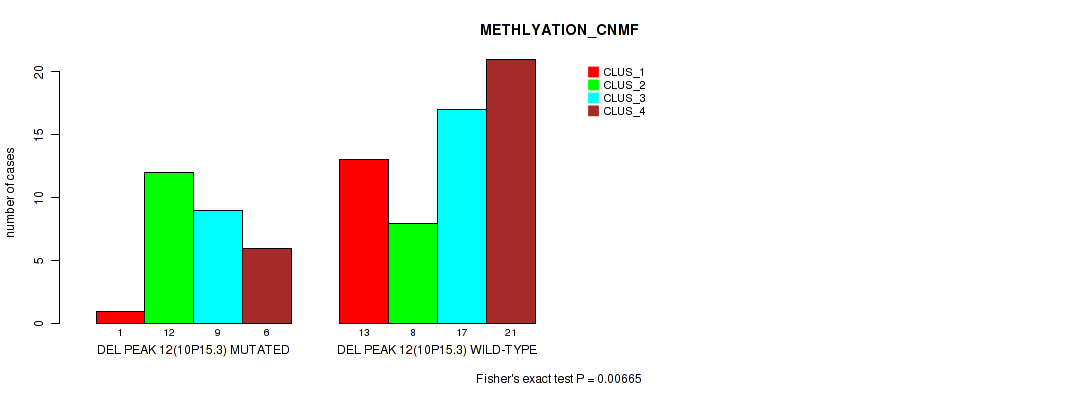
P value = 0.0329 (Fisher's exact test), Q value = 0.11
Table S31. Gene #12: 'del_10p15.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 26 | 22 | 17 | 21 |
| DEL PEAK 12(10P15.3) MUTATED | 3 | 8 | 7 | 10 |
| DEL PEAK 12(10P15.3) WILD-TYPE | 23 | 14 | 10 | 11 |
Figure S31. Get High-res Image Gene #12: 'del_10p15.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
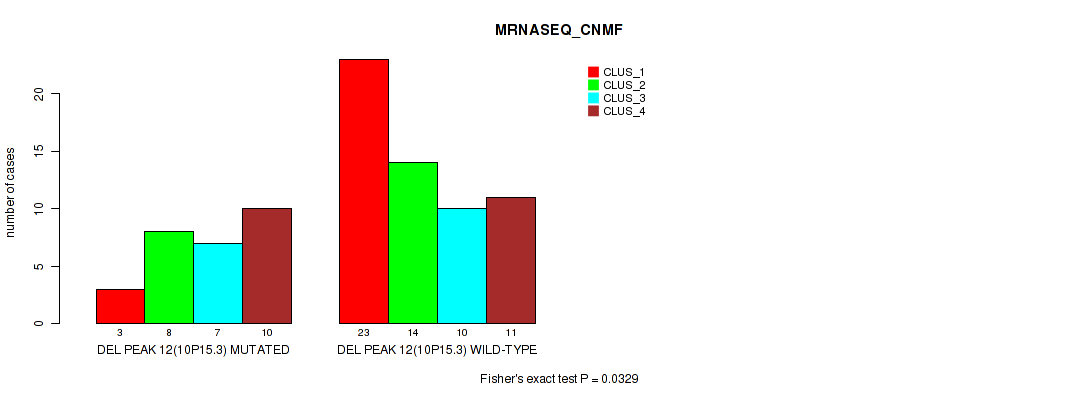
P value = 0.00892 (Fisher's exact test), Q value = 0.047
Table S32. Gene #12: 'del_10p15.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 22 | 12 | 22 | 13 | 17 |
| DEL PEAK 12(10P15.3) MUTATED | 1 | 6 | 8 | 5 | 8 |
| DEL PEAK 12(10P15.3) WILD-TYPE | 21 | 6 | 14 | 8 | 9 |
Figure S32. Get High-res Image Gene #12: 'del_10p15.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
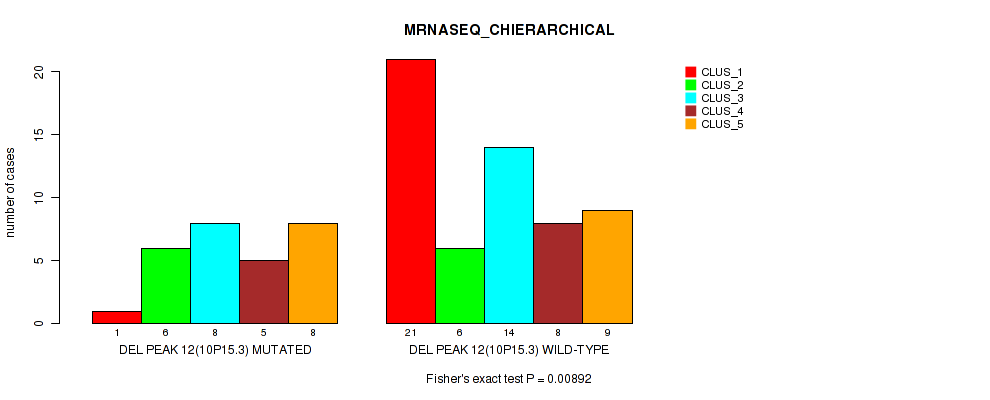
P value = 0.00272 (Fisher's exact test), Q value = 0.018
Table S33. Gene #13: 'del_10q25.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 13(10Q25.2) MUTATED | 3 | 10 | 15 | 3 |
| DEL PEAK 13(10Q25.2) WILD-TYPE | 20 | 16 | 9 | 11 |
Figure S33. Get High-res Image Gene #13: 'del_10q25.2' versus Molecular Subtype #1: 'CN_CNMF'
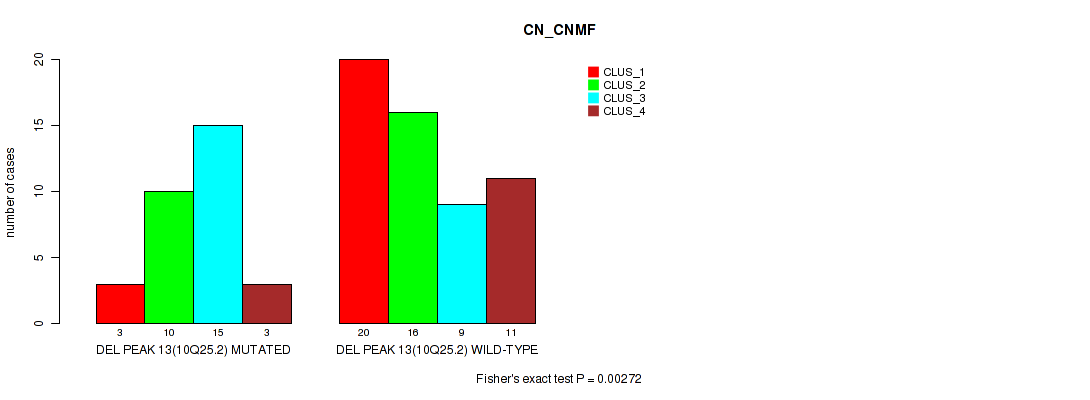
P value = 0.00806 (Fisher's exact test), Q value = 0.047
Table S34. Gene #13: 'del_10q25.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 14 | 20 | 26 | 27 |
| DEL PEAK 13(10Q25.2) MUTATED | 2 | 11 | 13 | 5 |
| DEL PEAK 13(10Q25.2) WILD-TYPE | 12 | 9 | 13 | 22 |
Figure S34. Get High-res Image Gene #13: 'del_10q25.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
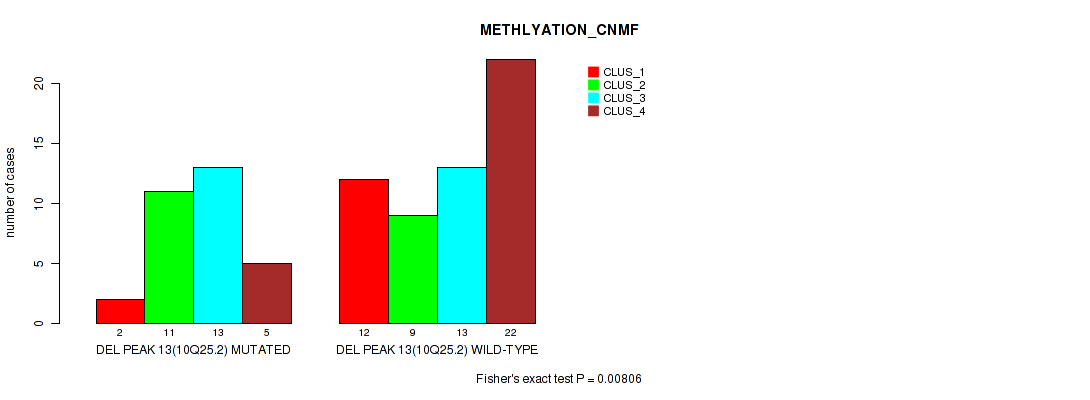
P value = 0.00056 (Fisher's exact test), Q value = 0.0059
Table S35. Gene #14: 'del_11q23.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 14(11Q23.2) MUTATED | 0 | 3 | 10 | 5 |
| DEL PEAK 14(11Q23.2) WILD-TYPE | 23 | 23 | 14 | 9 |
Figure S35. Get High-res Image Gene #14: 'del_11q23.2' versus Molecular Subtype #1: 'CN_CNMF'
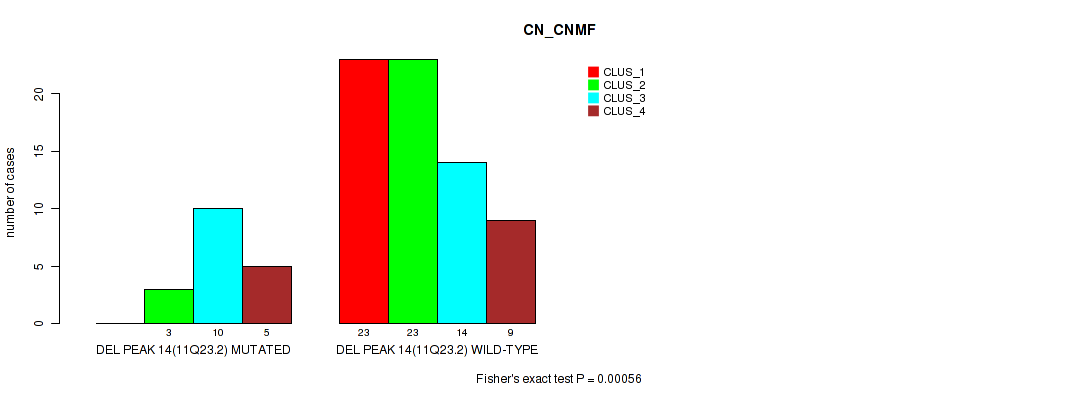
P value = 0.0125 (Fisher's exact test), Q value = 0.062
Table S36. Gene #15: 'del_12p13.31' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 14 | 20 | 26 | 27 |
| DEL PEAK 15(12P13.31) MUTATED | 1 | 7 | 2 | 1 |
| DEL PEAK 15(12P13.31) WILD-TYPE | 13 | 13 | 24 | 26 |
Figure S36. Get High-res Image Gene #15: 'del_12p13.31' versus Molecular Subtype #2: 'METHLYATION_CNMF'
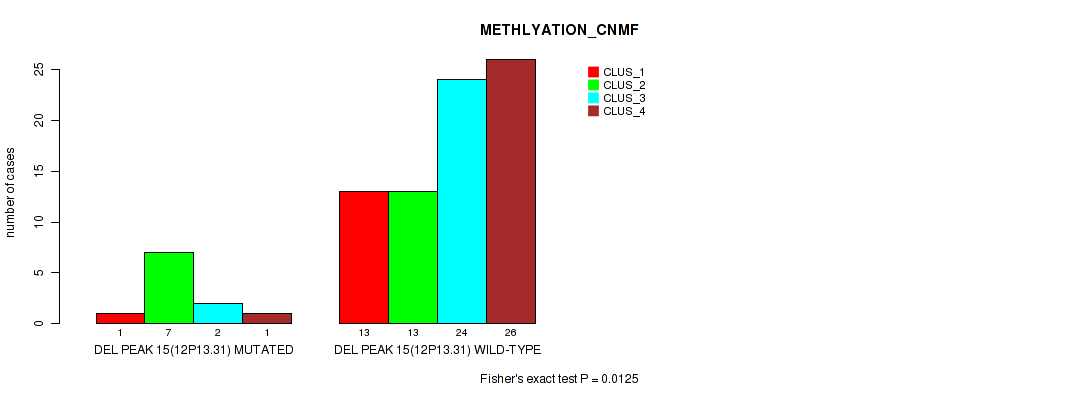
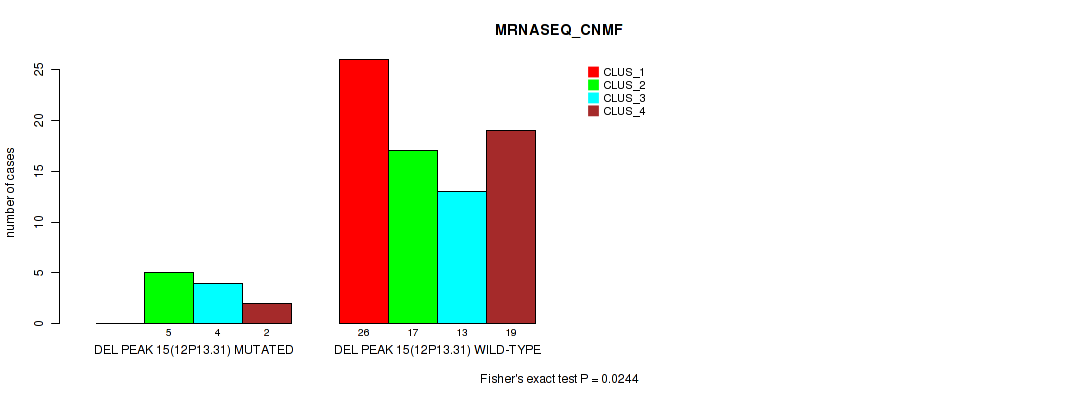
P value = 0.0244 (Fisher's exact test), Q value = 0.093
Table S37. Gene #15: 'del_12p13.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 26 | 22 | 17 | 21 |
| DEL PEAK 15(12P13.31) MUTATED | 0 | 5 | 4 | 2 |
| DEL PEAK 15(12P13.31) WILD-TYPE | 26 | 17 | 13 | 19 |
Figure S37. Get High-res Image Gene #15: 'del_12p13.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
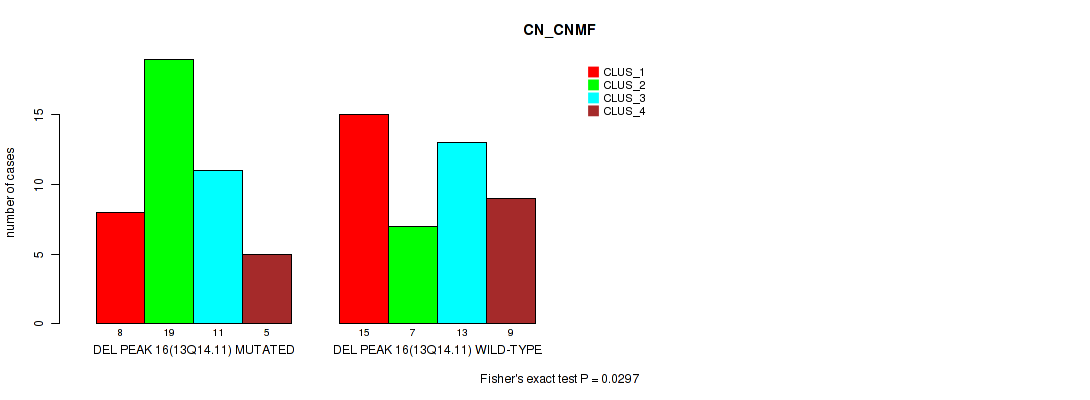
P value = 0.0297 (Fisher's exact test), Q value = 0.1
Table S38. Gene #16: 'del_13q14.11' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 16(13Q14.11) MUTATED | 8 | 19 | 11 | 5 |
| DEL PEAK 16(13Q14.11) WILD-TYPE | 15 | 7 | 13 | 9 |
Figure S38. Get High-res Image Gene #16: 'del_13q14.11' versus Molecular Subtype #1: 'CN_CNMF'
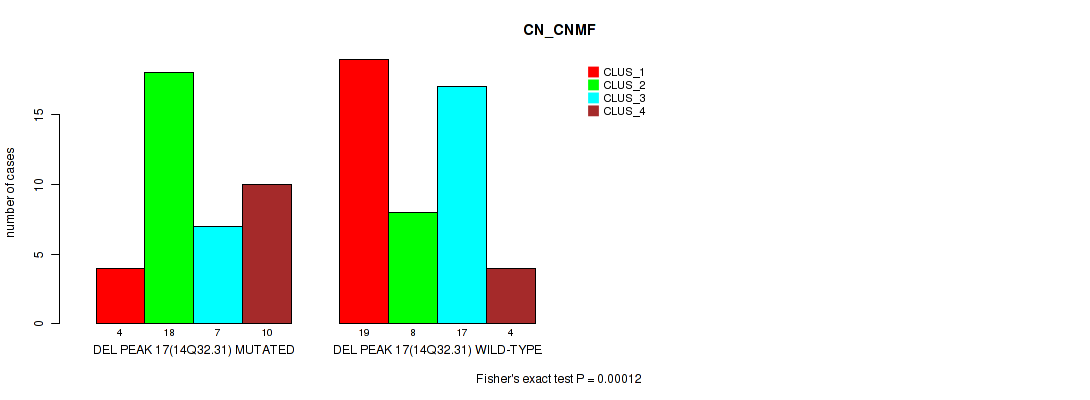
P value = 0.00012 (Fisher's exact test), Q value = 0.0017
Table S39. Gene #17: 'del_14q32.31' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 17(14Q32.31) MUTATED | 4 | 18 | 7 | 10 |
| DEL PEAK 17(14Q32.31) WILD-TYPE | 19 | 8 | 17 | 4 |
Figure S39. Get High-res Image Gene #17: 'del_14q32.31' versus Molecular Subtype #1: 'CN_CNMF'
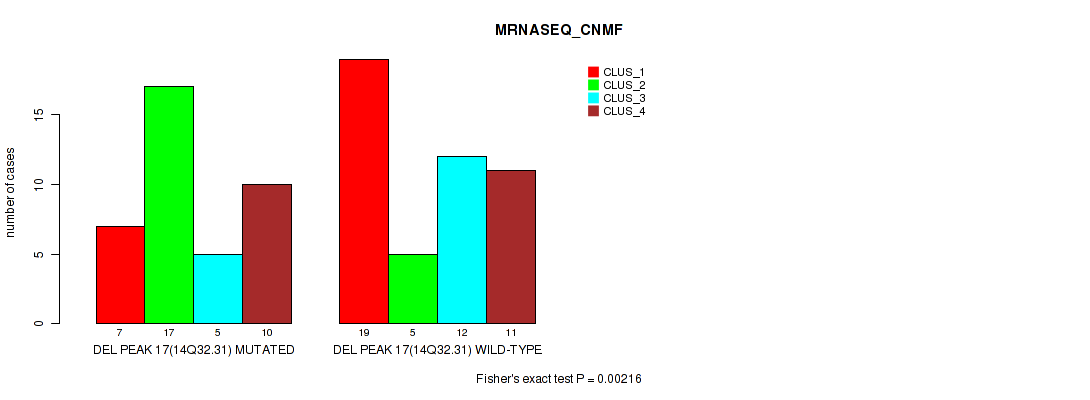
P value = 0.00216 (Fisher's exact test), Q value = 0.016
Table S40. Gene #17: 'del_14q32.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 26 | 22 | 17 | 21 |
| DEL PEAK 17(14Q32.31) MUTATED | 7 | 17 | 5 | 10 |
| DEL PEAK 17(14Q32.31) WILD-TYPE | 19 | 5 | 12 | 11 |
Figure S40. Get High-res Image Gene #17: 'del_14q32.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
P value = 1e-05 (Fisher's exact test), Q value = 0.00042
Table S41. Gene #17: 'del_14q32.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 22 | 12 | 22 | 13 | 17 |
| DEL PEAK 17(14Q32.31) MUTATED | 4 | 9 | 18 | 1 | 7 |
| DEL PEAK 17(14Q32.31) WILD-TYPE | 18 | 3 | 4 | 12 | 10 |
Figure S41. Get High-res Image Gene #17: 'del_14q32.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
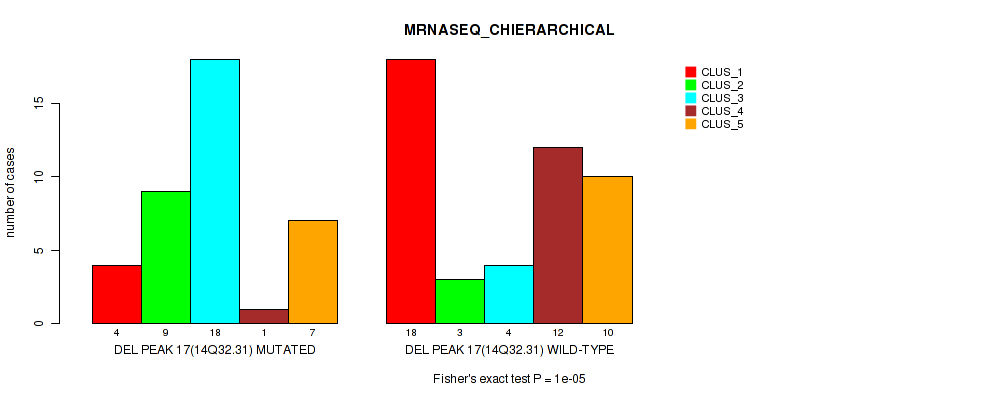
P value = 0.0262 (Fisher's exact test), Q value = 0.096
Table S42. Gene #17: 'del_14q32.31' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 36 | 29 | 22 |
| DEL PEAK 17(14Q32.31) MUTATED | 10 | 16 | 13 |
| DEL PEAK 17(14Q32.31) WILD-TYPE | 26 | 13 | 9 |
Figure S42. Get High-res Image Gene #17: 'del_14q32.31' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
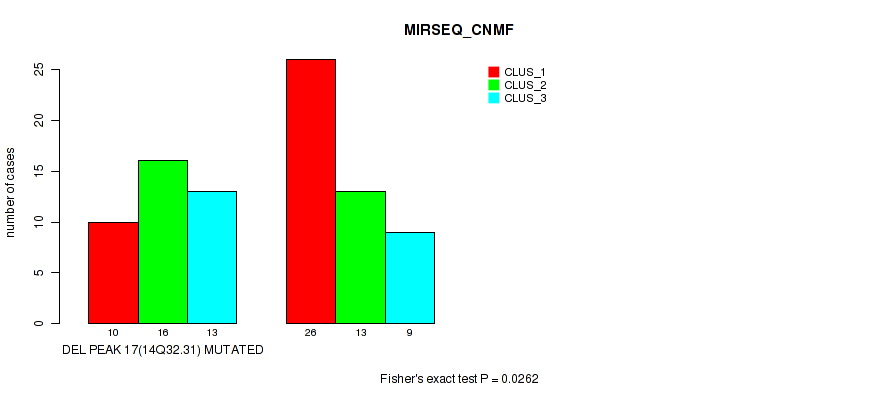
P value = 0.0089 (Fisher's exact test), Q value = 0.047
Table S43. Gene #17: 'del_14q32.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 29 | 25 | 6 | 13 | 14 |
| DEL PEAK 17(14Q32.31) MUTATED | 8 | 15 | 4 | 9 | 3 |
| DEL PEAK 17(14Q32.31) WILD-TYPE | 21 | 10 | 2 | 4 | 11 |
Figure S43. Get High-res Image Gene #17: 'del_14q32.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
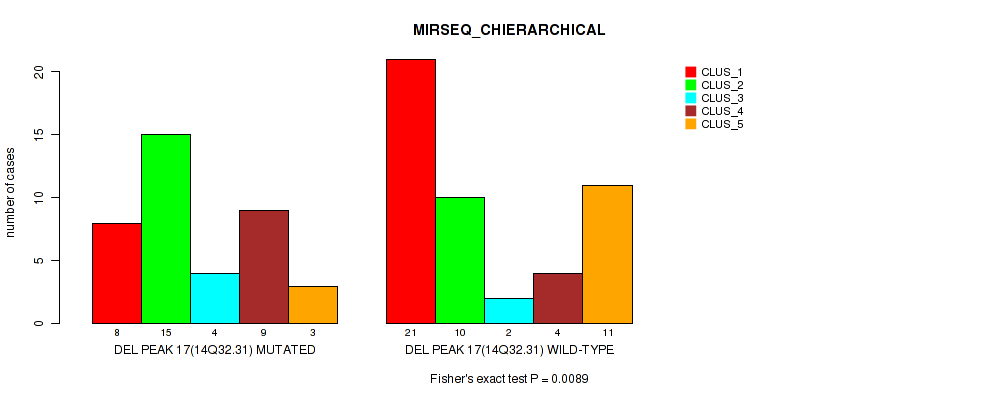
P value = 0.0426 (Fisher's exact test), Q value = 0.14
Table S44. Gene #17: 'del_14q32.31' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 27 | 21 |
| DEL PEAK 17(14Q32.31) MUTATED | 21 | 7 | 10 |
| DEL PEAK 17(14Q32.31) WILD-TYPE | 16 | 20 | 11 |
Figure S44. Get High-res Image Gene #17: 'del_14q32.31' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
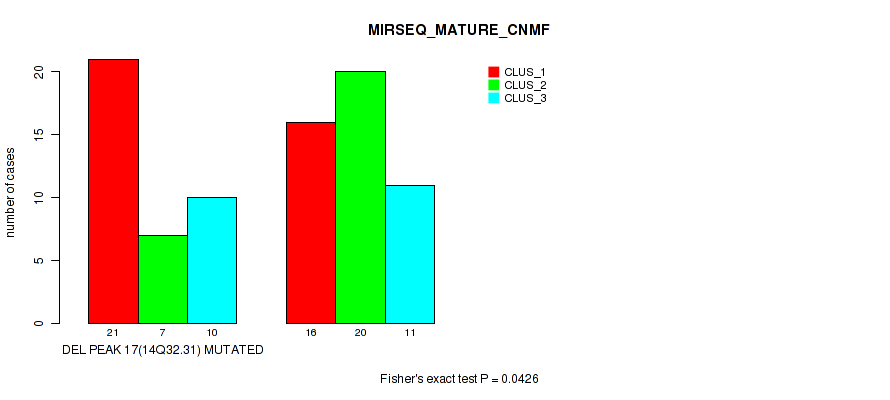
P value = 2e-05 (Fisher's exact test), Q value = 0.00048
Table S45. Gene #18: 'del_15q15.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 18(15Q15.1) MUTATED | 0 | 9 | 15 | 4 |
| DEL PEAK 18(15Q15.1) WILD-TYPE | 23 | 17 | 9 | 10 |
Figure S45. Get High-res Image Gene #18: 'del_15q15.1' versus Molecular Subtype #1: 'CN_CNMF'
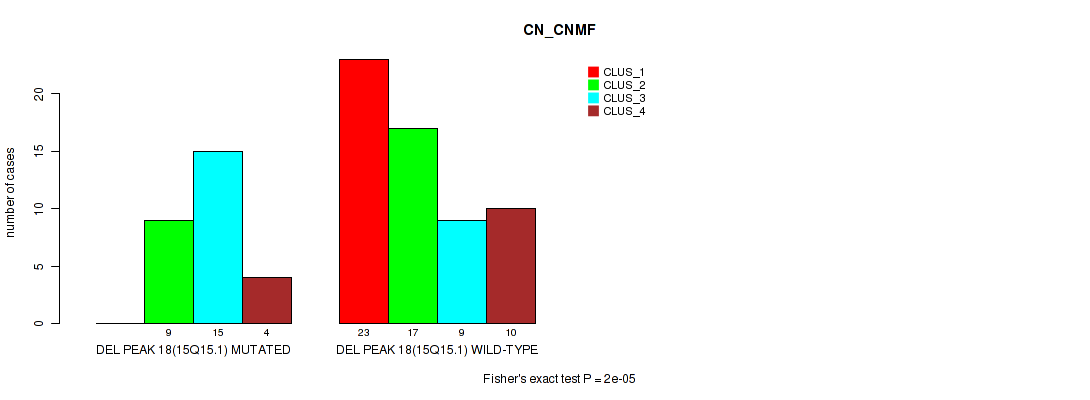
P value = 0.00118 (Fisher's exact test), Q value = 0.01
Table S46. Gene #18: 'del_15q15.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 14 | 20 | 26 | 27 |
| DEL PEAK 18(15Q15.1) MUTATED | 1 | 12 | 11 | 4 |
| DEL PEAK 18(15Q15.1) WILD-TYPE | 13 | 8 | 15 | 23 |
Figure S46. Get High-res Image Gene #18: 'del_15q15.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
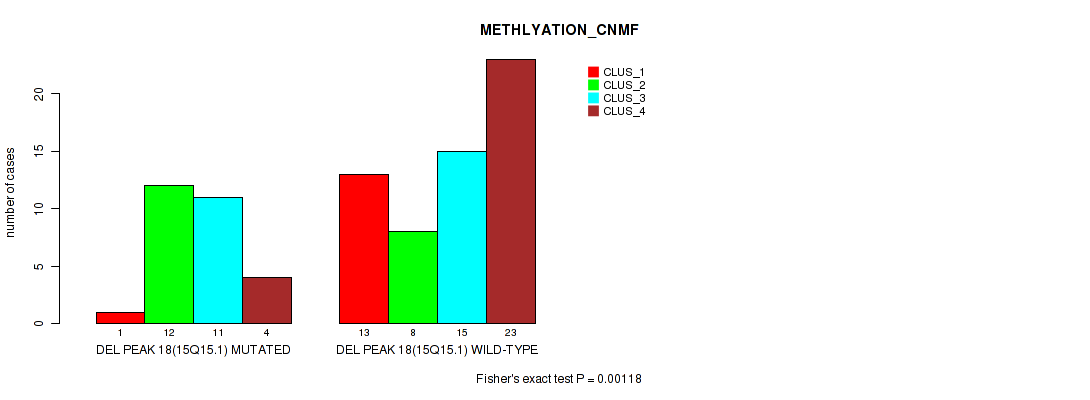
P value = 0.0275 (Fisher's exact test), Q value = 0.096
Table S47. Gene #18: 'del_15q15.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 26 | 22 | 17 | 21 |
| DEL PEAK 18(15Q15.1) MUTATED | 3 | 11 | 6 | 8 |
| DEL PEAK 18(15Q15.1) WILD-TYPE | 23 | 11 | 11 | 13 |
Figure S47. Get High-res Image Gene #18: 'del_15q15.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
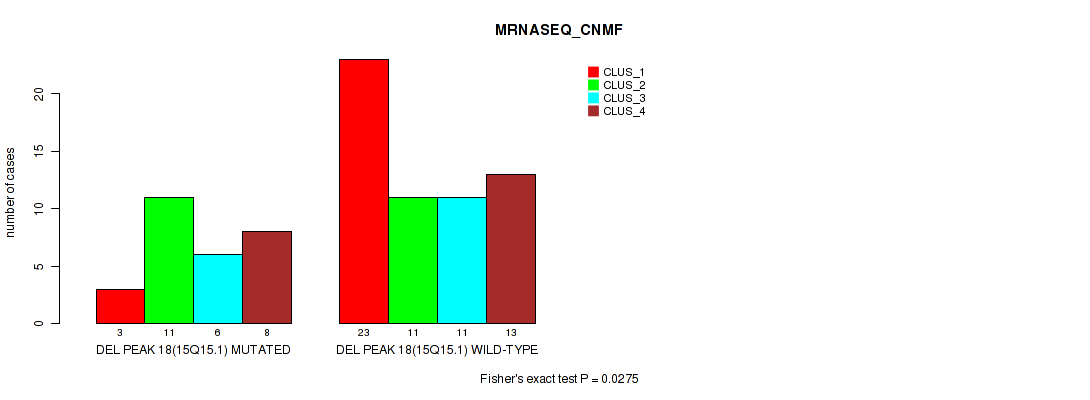
P value = 0.0209 (Fisher's exact test), Q value = 0.084
Table S48. Gene #18: 'del_15q15.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 16 | 17 | 21 | 31 |
| DEL PEAK 18(15Q15.1) MUTATED | 3 | 8 | 2 | 13 |
| DEL PEAK 18(15Q15.1) WILD-TYPE | 13 | 9 | 19 | 18 |
Figure S48. Get High-res Image Gene #18: 'del_15q15.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
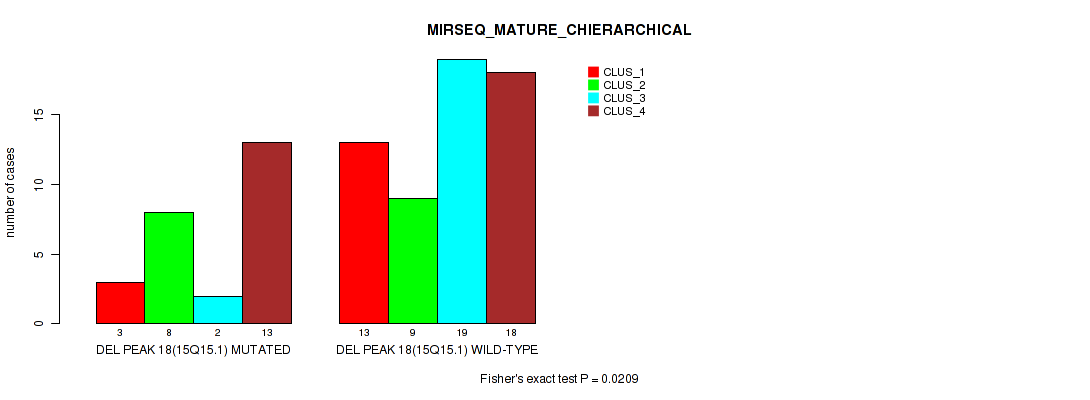
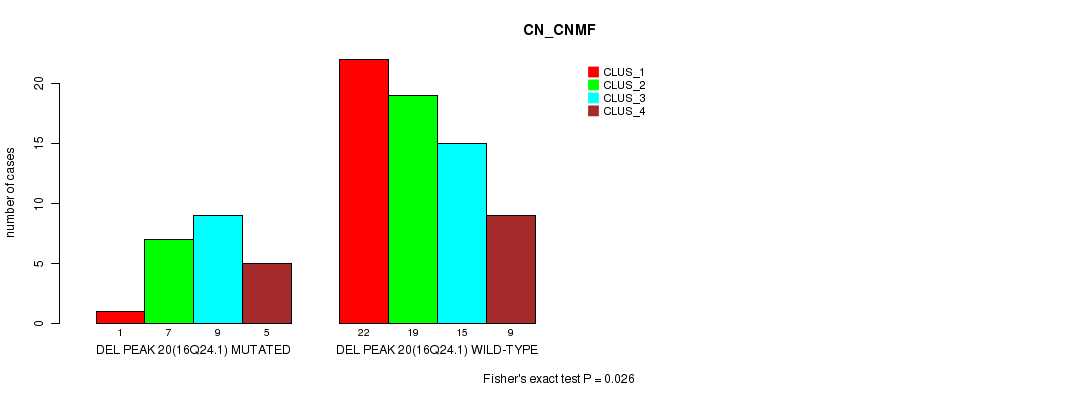
P value = 0.026 (Fisher's exact test), Q value = 0.096
Table S49. Gene #20: 'del_16q24.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 20(16Q24.1) MUTATED | 1 | 7 | 9 | 5 |
| DEL PEAK 20(16Q24.1) WILD-TYPE | 22 | 19 | 15 | 9 |
Figure S49. Get High-res Image Gene #20: 'del_16q24.1' versus Molecular Subtype #1: 'CN_CNMF'
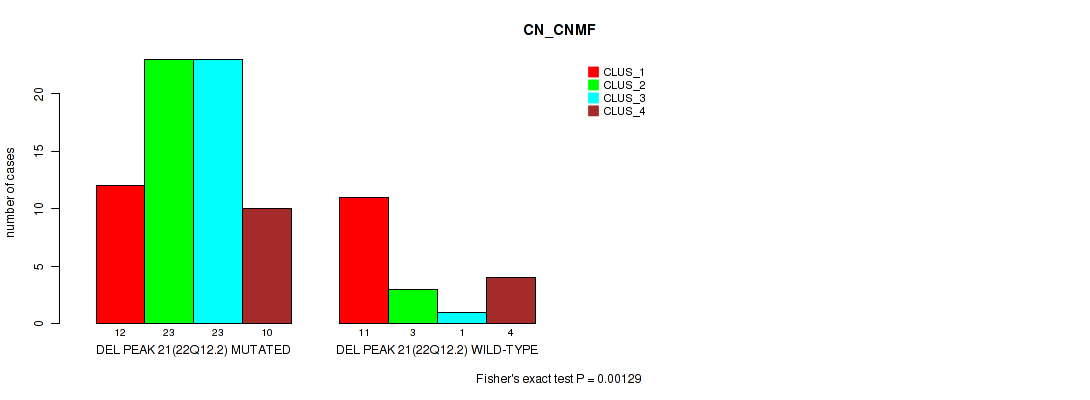
P value = 0.00129 (Fisher's exact test), Q value = 0.011
Table S50. Gene #21: 'del_22q12.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 26 | 24 | 14 |
| DEL PEAK 21(22Q12.2) MUTATED | 12 | 23 | 23 | 10 |
| DEL PEAK 21(22Q12.2) WILD-TYPE | 11 | 3 | 1 | 4 |
Figure S50. Get High-res Image Gene #21: 'del_22q12.2' versus Molecular Subtype #1: 'CN_CNMF'
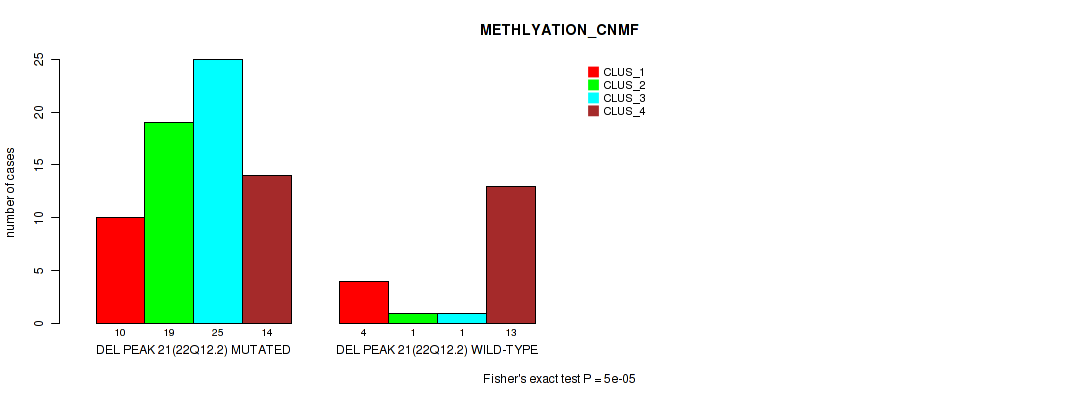
P value = 5e-05 (Fisher's exact test), Q value = 0.00084
Table S51. Gene #21: 'del_22q12.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 14 | 20 | 26 | 27 |
| DEL PEAK 21(22Q12.2) MUTATED | 10 | 19 | 25 | 14 |
| DEL PEAK 21(22Q12.2) WILD-TYPE | 4 | 1 | 1 | 13 |
Figure S51. Get High-res Image Gene #21: 'del_22q12.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
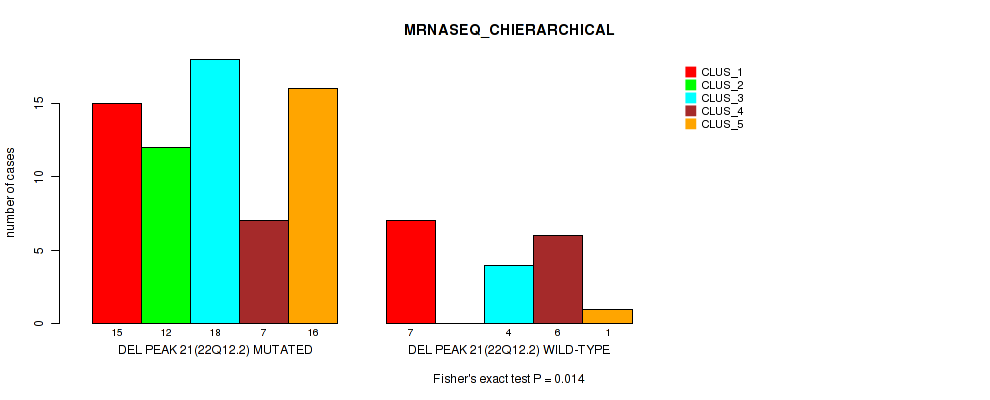
P value = 0.014 (Fisher's exact test), Q value = 0.067
Table S52. Gene #21: 'del_22q12.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 22 | 12 | 22 | 13 | 17 |
| DEL PEAK 21(22Q12.2) MUTATED | 15 | 12 | 18 | 7 | 16 |
| DEL PEAK 21(22Q12.2) WILD-TYPE | 7 | 0 | 4 | 6 | 1 |
Figure S52. Get High-res Image Gene #21: 'del_22q12.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
-
Copy number data file = all_lesions.txt from GISTIC pipeline
-
Processed Copy number data file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_Correlate_Genomic_Events_Preprocess/MESO-TP/15089884/transformed.cor.cli.txt
-
Molecular subtype file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_mergedClustering/MESO-TP/15104504/MESO-TP.transferedmergedcluster.txt
-
Number of patients = 87
-
Number of significantly focal cnvs = 21
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.