This pipeline computes the correlation between significantly recurrent gene mutations and selected clinical features.
Testing the association between mutation status of 197 genes and 7 clinical features across 248 patients, 12 significant findings detected with Q value < 0.25.
-
PTEN mutation correlated to 'HISTOLOGICAL_TYPE' and 'RACE'.
-
PIK3R1 mutation correlated to 'HISTOLOGICAL_TYPE'.
-
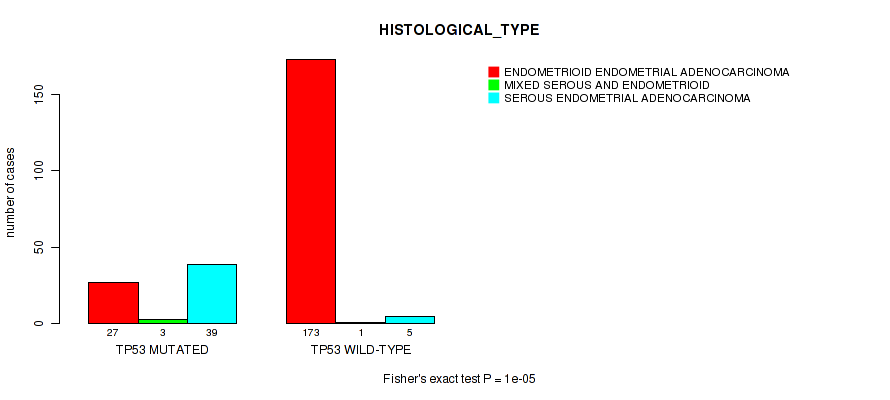
TP53 mutation correlated to 'YEARS_TO_BIRTH' and 'HISTOLOGICAL_TYPE'.
-
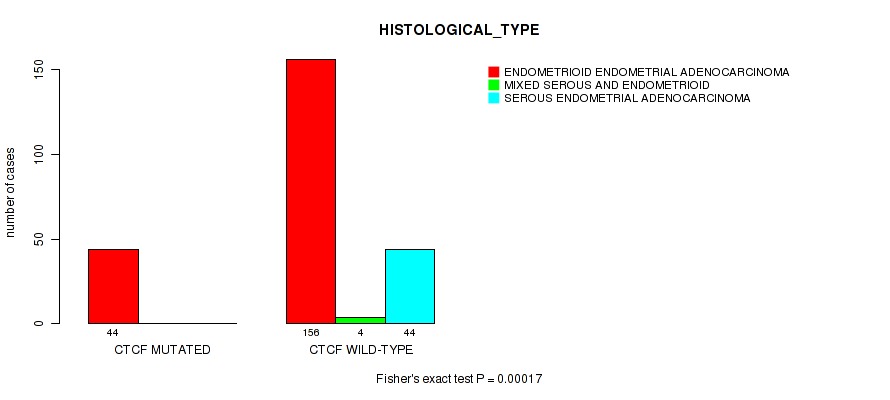
CTCF mutation correlated to 'HISTOLOGICAL_TYPE'.
-
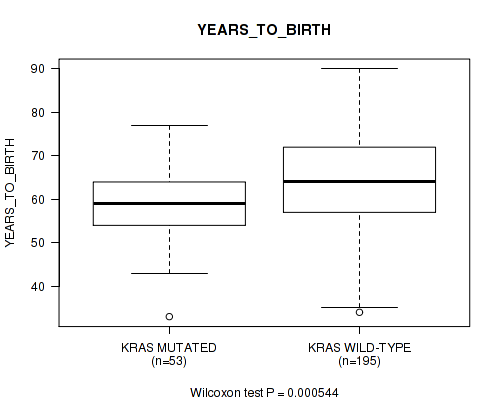
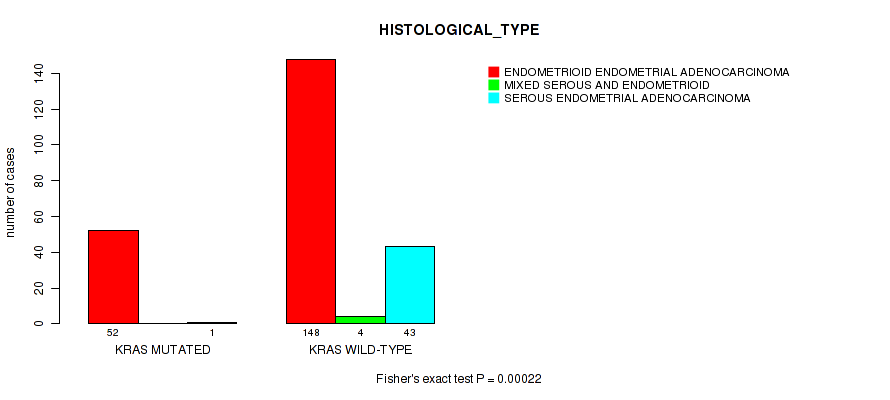
KRAS mutation correlated to 'YEARS_TO_BIRTH' and 'HISTOLOGICAL_TYPE'.
-
ARID1A mutation correlated to 'HISTOLOGICAL_TYPE'.
-
CTNNB1 mutation correlated to 'YEARS_TO_BIRTH' and 'HISTOLOGICAL_TYPE'.
-
PPP2R1A mutation correlated to 'HISTOLOGICAL_TYPE'.
Table 1. Get Full Table Overview of the association between mutation status of 197 genes and 7 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 12 significant findings detected.
|
Clinical Features |
Time to Death |
YEARS TO BIRTH |
HISTOLOGICAL TYPE |
RADIATIONS RADIATION REGIMENINDICATION |
COMPLETENESS OF RESECTION |
RACE | ETHNICITY | ||
| nMutated (%) | nWild-Type | logrank test | Wilcoxon-test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| PTEN | 161 (65%) | 87 |
0.0155 (0.485) |
0.00753 (0.447) |
1e-05 (0.00276) |
0.0359 (0.653) |
0.0465 (0.713) |
0.00063 (0.0724) |
1 (1.00) |
| TP53 | 69 (28%) | 179 |
0.0187 (0.528) |
5.85e-06 (0.00276) |
1e-05 (0.00276) |
0.237 (0.768) |
0.0851 (0.756) |
0.00753 (0.447) |
1 (1.00) |
| KRAS | 53 (21%) | 195 |
0.0846 (0.756) |
0.000544 (0.0682) |
0.00022 (0.0379) |
0.257 (0.772) |
0.458 (0.834) |
0.498 (0.854) |
0.583 (0.897) |
| CTNNB1 | 74 (30%) | 174 |
0.757 (0.993) |
0.000515 (0.0682) |
1e-05 (0.00276) |
0.31 (0.787) |
0.493 (0.854) |
0.728 (0.988) |
1 (1.00) |
| PIK3R1 | 83 (33%) | 165 |
0.903 (1.00) |
0.398 (0.826) |
1e-05 (0.00276) |
0.324 (0.802) |
0.954 (1.00) |
0.108 (0.756) |
0.136 (0.756) |
| CTCF | 44 (18%) | 204 |
0.206 (0.756) |
0.0368 (0.659) |
0.00017 (0.0335) |
1 (1.00) |
0.735 (0.993) |
0.777 (1.00) |
1 (1.00) |
| ARID1A | 83 (33%) | 165 |
0.00846 (0.447) |
0.00633 (0.447) |
9e-05 (0.0207) |
0.573 (0.896) |
0.265 (0.775) |
0.247 (0.772) |
1 (1.00) |
| PPP2R1A | 27 (11%) | 221 |
0.92 (1.00) |
0.007 (0.447) |
0.00042 (0.0644) |
0.523 (0.874) |
1 (1.00) |
0.402 (0.826) |
0.31 (0.787) |
| FBXW7 | 39 (16%) | 209 |
0.67 (0.956) |
0.888 (1.00) |
0.00343 (0.278) |
0.856 (1.00) |
0.528 (0.878) |
0.786 (1.00) |
0.528 (0.878) |
| ARHGAP35 | 36 (15%) | 212 |
0.671 (0.956) |
0.914 (1.00) |
0.726 (0.988) |
1 (1.00) |
0.928 (1.00) |
0.195 (0.756) |
0.457 (0.834) |
| PIK3CA | 132 (53%) | 116 |
0.0226 (0.566) |
0.185 (0.756) |
0.252 (0.772) |
0.689 (0.966) |
0.187 (0.756) |
0.72 (0.985) |
1 (1.00) |
| FGFR2 | 31 (12%) | 217 |
0.569 (0.895) |
0.983 (1.00) |
0.774 (1.00) |
1 (1.00) |
0.38 (0.824) |
0.631 (0.929) |
1 (1.00) |
| ZFHX3 | 44 (18%) | 204 |
0.52 (0.871) |
0.744 (0.993) |
0.0666 (0.756) |
0.862 (1.00) |
0.495 (0.854) |
0.1 (0.756) |
0.126 (0.756) |
| TCP11L2 | 14 (6%) | 234 |
0.164 (0.756) |
0.988 (1.00) |
0.171 (0.756) |
1 (1.00) |
0.247 (0.772) |
0.0902 (0.756) |
0.235 (0.766) |
| SPOP | 21 (8%) | 227 |
0.433 (0.826) |
0.451 (0.834) |
0.842 (1.00) |
0.812 (1.00) |
0.554 (0.882) |
0.728 (0.988) |
0.31 (0.787) |
| RBMX | 13 (5%) | 235 |
0.594 (0.904) |
0.233 (0.766) |
0.3 (0.779) |
0.146 (0.756) |
0.134 (0.756) |
0.213 (0.756) |
1 (1.00) |
| SOX17 | 7 (3%) | 241 |
0.453 (0.834) |
0.452 (0.834) |
0.428 (0.826) |
0.427 (0.826) |
0.686 (0.963) |
0.0642 (0.756) |
1 (1.00) |
| NFE2L2 | 15 (6%) | 233 |
0.687 (0.964) |
0.154 (0.756) |
0.14 (0.756) |
0.587 (0.897) |
0.779 (1.00) |
0.514 (0.869) |
1 (1.00) |
| CCND1 | 14 (6%) | 234 |
0.625 (0.926) |
0.0652 (0.756) |
0.171 (0.756) |
0.567 (0.895) |
0.304 (0.781) |
0.757 (0.993) |
0.235 (0.766) |
| GNPTAB | 20 (8%) | 228 |
0.149 (0.756) |
0.324 (0.802) |
0.268 (0.775) |
0.464 (0.839) |
0.648 (0.936) |
0.501 (0.854) |
0.273 (0.775) |
| ARID5B | 29 (12%) | 219 |
0.801 (1.00) |
0.884 (1.00) |
0.011 (0.46) |
1 (1.00) |
0.111 (0.756) |
0.00268 (0.264) |
0.411 (0.826) |
| DNER | 18 (7%) | 230 |
0.539 (0.882) |
0.0325 (0.633) |
0.416 (0.826) |
0.442 (0.829) |
0.645 (0.936) |
0.102 (0.756) |
0.235 (0.766) |
| EP300 | 21 (8%) | 227 |
0.133 (0.756) |
0.0984 (0.756) |
0.0663 (0.756) |
0.637 (0.929) |
1 (1.00) |
0.551 (0.882) |
0.0442 (0.687) |
| MAX | 11 (4%) | 237 |
0.904 (1.00) |
0.296 (0.775) |
0.349 (0.802) |
0.752 (0.993) |
0.405 (0.826) |
0.109 (0.756) |
1 (1.00) |
| SGK1 | 15 (6%) | 233 |
0.916 (1.00) |
0.529 (0.879) |
0.47 (0.842) |
0.16 (0.756) |
0.1 (0.756) |
0.364 (0.806) |
1 (1.00) |
| NRAS | 9 (4%) | 239 |
0.209 (0.756) |
0.785 (1.00) |
1 (1.00) |
0.723 (0.986) |
1 (1.00) |
0.0423 (0.687) |
0.155 (0.756) |
| KLHL8 | 12 (5%) | 236 |
0.269 (0.775) |
0.289 (0.775) |
0.29 (0.775) |
1 (1.00) |
0.227 (0.766) |
0.466 (0.84) |
0.176 (0.756) |
| MORC4 | 20 (8%) | 228 |
0.103 (0.756) |
0.0146 (0.481) |
0.268 (0.775) |
1 (1.00) |
1 (1.00) |
0.736 (0.993) |
0.273 (0.775) |
| ZNF781 | 10 (4%) | 238 |
0.344 (0.802) |
0.112 (0.756) |
0.337 (0.802) |
0.501 (0.854) |
0.491 (0.854) |
0.708 (0.98) |
0.134 (0.756) |
| MKI67 | 29 (12%) | 219 |
0.198 (0.756) |
0.499 (0.854) |
0.0109 (0.46) |
0.836 (1.00) |
0.294 (0.775) |
0.188 (0.756) |
0.395 (0.826) |
| ING1 | 13 (5%) | 235 |
0.308 (0.787) |
0.9 (1.00) |
0.3 (0.779) |
0.146 (0.756) |
0.843 (1.00) |
0.908 (1.00) |
1 (1.00) |
| INTS7 | 8 (3%) | 240 |
0.272 (0.775) |
0.536 (0.882) |
0.436 (0.826) |
1 (1.00) |
0.548 (0.882) |
0.178 (0.756) |
0.113 (0.756) |
| CCDC6 | 6 (2%) | 242 |
0.559 (0.889) |
0.749 (0.993) |
0.363 (0.806) |
1 (1.00) |
1 (1.00) |
0.534 (0.882) |
1 (1.00) |
| EIF2S2 | 9 (4%) | 239 |
0.334 (0.802) |
0.384 (0.826) |
0.458 (0.834) |
0.169 (0.756) |
0.427 (0.826) |
0.226 (0.766) |
0.155 (0.756) |
| RBBP6 | 22 (9%) | 226 |
0.418 (0.826) |
0.988 (1.00) |
0.0427 (0.687) |
1 (1.00) |
0.514 (0.869) |
0.282 (0.775) |
1 (1.00) |
| SOS1 | 12 (5%) | 236 |
0.252 (0.772) |
0.108 (0.756) |
0.291 (0.775) |
0.757 (0.993) |
0.843 (1.00) |
0.894 (1.00) |
1 (1.00) |
| NAT1 | 7 (3%) | 241 |
0.686 (0.963) |
0.983 (1.00) |
0.426 (0.826) |
1 (1.00) |
0.486 (0.854) |
0.199 (0.756) |
1 (1.00) |
| ADNP | 14 (6%) | 234 |
0.837 (1.00) |
0.794 (1.00) |
0.172 (0.756) |
1 (1.00) |
0.648 (0.936) |
0.908 (1.00) |
0.155 (0.756) |
| VPS11 | 12 (5%) | 236 |
0.843 (1.00) |
0.915 (1.00) |
1 (1.00) |
0.551 (0.882) |
0.845 (1.00) |
0.362 (0.805) |
1 (1.00) |
| L1TD1 | 16 (6%) | 232 |
0.201 (0.756) |
0.348 (0.802) |
0.119 (0.756) |
0.588 (0.897) |
0.886 (1.00) |
0.336 (0.802) |
0.235 (0.766) |
| MARK3 | 11 (4%) | 237 |
0.85 (1.00) |
0.0516 (0.736) |
0.353 (0.802) |
1 (1.00) |
0.214 (0.756) |
0.325 (0.802) |
0.134 (0.756) |
| CTNND1 | 19 (8%) | 229 |
0.797 (1.00) |
0.411 (0.826) |
0.427 (0.826) |
0.465 (0.839) |
0.426 (0.826) |
0.497 (0.854) |
1 (1.00) |
| GFAP | 9 (4%) | 239 |
0.426 (0.826) |
0.0727 (0.756) |
0.456 (0.834) |
1 (1.00) |
0.28 (0.775) |
0.351 (0.802) |
0.113 (0.756) |
| C9ORF102 | 16 (6%) | 232 |
0.167 (0.756) |
0.318 (0.799) |
0.12 (0.756) |
0.588 (0.897) |
0.68 (0.963) |
0.334 (0.802) |
0.235 (0.766) |
| EIF4A2 | 7 (3%) | 241 |
0.378 (0.824) |
0.205 (0.756) |
0.426 (0.826) |
0.427 (0.826) |
1 (1.00) |
0.177 (0.756) |
1 (1.00) |
| ZNF471 | 15 (6%) | 233 |
0.746 (0.993) |
0.0355 (0.653) |
0.141 (0.756) |
0.587 (0.897) |
0.38 (0.824) |
0.0615 (0.756) |
0.235 (0.766) |
| CDK17 | 14 (6%) | 234 |
0.58 (0.897) |
0.0852 (0.756) |
0.128 (0.756) |
0.776 (1.00) |
0.287 (0.775) |
0.359 (0.805) |
0.216 (0.756) |
| SIN3A | 21 (8%) | 227 |
0.366 (0.807) |
0.159 (0.756) |
0.278 (0.775) |
0.812 (1.00) |
0.9 (1.00) |
0.0975 (0.756) |
1 (1.00) |
| ZNF485 | 9 (4%) | 239 |
0.785 (1.00) |
0.361 (0.805) |
0.457 (0.834) |
0.169 (0.756) |
0.0903 (0.756) |
0.227 (0.766) |
0.176 (0.756) |
| RSBN1L | 12 (5%) | 236 |
0.181 (0.756) |
0.291 (0.775) |
0.752 (0.993) |
1 (1.00) |
0.195 (0.756) |
0.00331 (0.278) |
0.196 (0.756) |
| CUX1 | 23 (9%) | 225 |
0.406 (0.826) |
0.11 (0.756) |
0.172 (0.756) |
0.365 (0.806) |
0.829 (1.00) |
0.609 (0.911) |
1 (1.00) |
| BCOR | 30 (12%) | 218 |
0.0583 (0.756) |
0.0735 (0.756) |
0.00843 (0.447) |
1 (1.00) |
0.379 (0.824) |
0.0295 (0.623) |
1 (1.00) |
| WDR45 | 11 (4%) | 237 |
0.937 (1.00) |
0.551 (0.882) |
0.351 (0.802) |
0.521 (0.871) |
0.546 (0.882) |
0.0421 (0.687) |
1 (1.00) |
| CAB39L | 8 (3%) | 240 |
0.294 (0.775) |
0.789 (1.00) |
0.437 (0.826) |
0.268 (0.775) |
0.55 (0.882) |
0.00918 (0.452) |
1 (1.00) |
| TAB3 | 18 (7%) | 230 |
0.128 (0.756) |
0.0402 (0.687) |
0.0862 (0.756) |
0.614 (0.914) |
0.296 (0.775) |
0.0309 (0.623) |
0.31 (0.787) |
| OAZ3 | 8 (3%) | 240 |
0.319 (0.799) |
0.165 (0.756) |
1 (1.00) |
0.454 (0.834) |
0.547 (0.882) |
0.109 (0.756) |
0.113 (0.756) |
| AHCYL1 | 6 (2%) | 242 |
0.426 (0.826) |
0.0311 (0.623) |
0.632 (0.929) |
1 (1.00) |
0.416 (0.826) |
0.171 (0.756) |
0.0689 (0.756) |
| ATM | 29 (12%) | 219 |
0.0425 (0.687) |
0.265 (0.775) |
0.0753 (0.756) |
0.534 (0.882) |
0.904 (1.00) |
0.222 (0.761) |
0.378 (0.824) |
| MSH4 | 15 (6%) | 233 |
0.186 (0.756) |
0.155 (0.756) |
0.141 (0.756) |
1 (1.00) |
0.649 (0.936) |
0.0705 (0.756) |
0.216 (0.756) |
| FAM65B | 16 (6%) | 232 |
0.175 (0.756) |
0.049 (0.725) |
0.479 (0.853) |
0.792 (1.00) |
0.778 (1.00) |
0.251 (0.772) |
0.216 (0.756) |
| FN1 | 24 (10%) | 224 |
0.383 (0.826) |
0.714 (0.983) |
0.0216 (0.551) |
1 (1.00) |
0.568 (0.895) |
0.777 (1.00) |
0.345 (0.802) |
| JAKMIP2 | 12 (5%) | 236 |
0.724 (0.986) |
0.0301 (0.623) |
0.289 (0.775) |
1 (1.00) |
0.275 (0.775) |
0.0294 (0.623) |
0.155 (0.756) |
| WBP4 | 8 (3%) | 240 |
0.462 (0.839) |
0.133 (0.756) |
0.436 (0.826) |
0.718 (0.983) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
| RASA1 | 22 (9%) | 226 |
0.0855 (0.756) |
0.745 (0.993) |
0.575 (0.896) |
0.493 (0.854) |
1 (1.00) |
0.0929 (0.756) |
0.0442 (0.687) |
| ALPK2 | 19 (8%) | 229 |
0.0894 (0.756) |
0.233 (0.766) |
0.43 (0.826) |
0.315 (0.796) |
0.255 (0.772) |
0.127 (0.756) |
0.255 (0.772) |
| POLE | 27 (11%) | 221 |
0.611 (0.911) |
0.552 (0.882) |
0.127 (0.756) |
0.199 (0.756) |
0.457 (0.834) |
0.37 (0.812) |
0.362 (0.805) |
| KIF20B | 21 (8%) | 227 |
0.143 (0.756) |
0.142 (0.756) |
0.0645 (0.756) |
0.637 (0.929) |
0.913 (1.00) |
0.552 (0.882) |
0.255 (0.772) |
| C14ORF166B | 10 (4%) | 238 |
0.285 (0.775) |
0.903 (1.00) |
0.335 (0.802) |
0.172 (0.756) |
1 (1.00) |
0.12 (0.756) |
1 (1.00) |
| SLC26A8 | 12 (5%) | 236 |
0.298 (0.778) |
0.0329 (0.633) |
0.287 (0.775) |
0.551 (0.882) |
0.7 (0.975) |
0.406 (0.826) |
0.196 (0.756) |
| ZNF334 | 17 (7%) | 231 |
0.819 (1.00) |
0.00468 (0.358) |
0.809 (1.00) |
1 (1.00) |
0.68 (0.963) |
0.734 (0.992) |
0.255 (0.772) |
| RRAS2 | 4 (2%) | 244 |
0.637 (0.929) |
0.0213 (0.551) |
1 (1.00) |
0.611 (0.911) |
0.437 (0.826) |
1 (1.00) |
1 (1.00) |
| PPM1N | 7 (3%) | 241 |
0.346 (0.802) |
0.981 (1.00) |
0.426 (0.826) |
1 (1.00) |
0.426 (0.826) |
0.428 (0.826) |
0.113 (0.756) |
| FCN1 | 8 (3%) | 240 |
0.824 (1.00) |
0.627 (0.928) |
0.438 (0.826) |
0.718 (0.983) |
1 (1.00) |
0.168 (0.756) |
1 (1.00) |
| TIAL1 | 10 (4%) | 238 |
0.322 (0.802) |
0.272 (0.775) |
0.337 (0.802) |
1 (1.00) |
0.604 (0.908) |
0.156 (0.756) |
1 (1.00) |
| PSMC4 | 11 (4%) | 237 |
0.252 (0.772) |
0.831 (1.00) |
0.352 (0.802) |
0.521 (0.871) |
0.857 (1.00) |
0.15 (0.756) |
1 (1.00) |
| MFAP5 | 9 (4%) | 239 |
0.358 (0.805) |
0.0104 (0.46) |
1 (1.00) |
1 (1.00) |
0.547 (0.882) |
0.569 (0.895) |
0.134 (0.756) |
| RAB3GAP1 | 16 (6%) | 232 |
0.771 (1.00) |
0.466 (0.84) |
0.12 (0.756) |
1 (1.00) |
0.193 (0.756) |
0.186 (0.756) |
0.011 (0.46) |
| MSH6 | 17 (7%) | 231 |
0.533 (0.882) |
0.124 (0.756) |
0.492 (0.854) |
0.296 (0.775) |
0.679 (0.963) |
0.576 (0.896) |
0.216 (0.756) |
| BMP2K | 13 (5%) | 235 |
0.318 (0.799) |
0.904 (1.00) |
0.302 (0.779) |
0.551 (0.882) |
0.703 (0.976) |
0.732 (0.991) |
0.196 (0.756) |
| ZNF606 | 16 (6%) | 232 |
0.196 (0.756) |
0.446 (0.834) |
0.12 (0.756) |
0.588 (0.897) |
0.515 (0.869) |
0.13 (0.756) |
0.0211 (0.551) |
| ZNF263 | 7 (3%) | 241 |
0.364 (0.806) |
0.0566 (0.756) |
0.426 (0.826) |
1 (1.00) |
0.209 (0.756) |
0.355 (0.802) |
0.0689 (0.756) |
| CHEK2 | 13 (5%) | 235 |
0.709 (0.98) |
0.0191 (0.528) |
0.575 (0.896) |
1 (1.00) |
0.207 (0.756) |
0.479 (0.853) |
0.176 (0.756) |
| MGA | 26 (10%) | 222 |
0.708 (0.98) |
0.67 (0.956) |
0.35 (0.802) |
0.392 (0.826) |
0.154 (0.756) |
0.355 (0.802) |
0.292 (0.775) |
| RHBDD3 | 4 (2%) | 244 |
0.546 (0.882) |
0.128 (0.756) |
1 (1.00) |
0.611 (0.911) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
| TAP1 | 8 (3%) | 240 |
0.86 (1.00) |
0.942 (1.00) |
0.438 (0.826) |
1 (1.00) |
1 (1.00) |
0.404 (0.826) |
1 (1.00) |
| RB1 | 20 (8%) | 228 |
0.333 (0.802) |
0.124 (0.756) |
0.0685 (0.756) |
0.464 (0.839) |
0.93 (1.00) |
0.0784 (0.756) |
1 (1.00) |
| SLC1A3 | 12 (5%) | 236 |
0.768 (1.00) |
0.755 (0.993) |
1 (1.00) |
1 (1.00) |
0.656 (0.943) |
0.00814 (0.447) |
1 (1.00) |
| ATAD5 | 17 (7%) | 231 |
0.154 (0.756) |
0.252 (0.772) |
0.49 (0.854) |
1 (1.00) |
0.793 (1.00) |
0.101 (0.756) |
1 (1.00) |
| ALG8 | 10 (4%) | 238 |
0.292 (0.775) |
0.854 (1.00) |
0.336 (0.802) |
0.501 (0.854) |
0.549 (0.882) |
0.327 (0.802) |
0.0141 (0.481) |
| TIGD4 | 13 (5%) | 235 |
0.218 (0.756) |
0.0517 (0.736) |
0.305 (0.781) |
0.771 (1.00) |
0.682 (0.963) |
0.48 (0.853) |
0.216 (0.756) |
| PARG | 9 (4%) | 239 |
0.312 (0.789) |
0.554 (0.882) |
0.459 (0.834) |
1 (1.00) |
1 (1.00) |
0.712 (0.981) |
1 (1.00) |
| FAT1 | 40 (16%) | 208 |
0.0305 (0.623) |
0.875 (1.00) |
0.0443 (0.687) |
0.279 (0.775) |
0.802 (1.00) |
0.408 (0.826) |
0.118 (0.756) |
| DYM | 10 (4%) | 238 |
0.37 (0.812) |
0.732 (0.991) |
0.74 (0.993) |
0.742 (0.993) |
1 (1.00) |
0.586 (0.897) |
1 (1.00) |
| PSMD3 | 11 (4%) | 237 |
0.253 (0.772) |
0.373 (0.816) |
0.746 (0.993) |
1 (1.00) |
1 (1.00) |
0.402 (0.826) |
1 (1.00) |
| PPM1D | 11 (4%) | 237 |
0.234 (0.766) |
0.351 (0.802) |
0.351 (0.802) |
0.339 (0.802) |
0.605 (0.908) |
0.254 (0.772) |
0.155 (0.756) |
| ZMYM2 | 17 (7%) | 231 |
0.171 (0.756) |
0.0287 (0.623) |
0.102 (0.756) |
1 (1.00) |
0.56 (0.889) |
0.278 (0.775) |
1 (1.00) |
| INPP4B | 12 (5%) | 236 |
0.586 (0.897) |
0.178 (0.756) |
0.29 (0.775) |
0.0621 (0.756) |
0.207 (0.756) |
0.145 (0.756) |
0.196 (0.756) |
| USP28 | 10 (4%) | 238 |
0.228 (0.766) |
0.502 (0.854) |
0.334 (0.802) |
0.501 (0.854) |
0.79 (1.00) |
0.352 (0.802) |
1 (1.00) |
| EMR1 | 13 (5%) | 235 |
0.235 (0.766) |
0.127 (0.756) |
0.301 (0.779) |
0.551 (0.882) |
0.701 (0.975) |
0.389 (0.826) |
0.176 (0.756) |
| ZNF385B | 7 (3%) | 241 |
0.419 (0.826) |
0.204 (0.756) |
0.428 (0.826) |
1 (1.00) |
1 (1.00) |
0.501 (0.854) |
1 (1.00) |
| RAE1 | 11 (4%) | 237 |
0.221 (0.761) |
0.729 (0.989) |
0.745 (0.993) |
0.339 (0.802) |
1 (1.00) |
0.321 (0.802) |
1 (1.00) |
| TRIM59 | 9 (4%) | 239 |
0.283 (0.775) |
0.876 (1.00) |
0.454 (0.834) |
1 (1.00) |
1 (1.00) |
0.0396 (0.687) |
1 (1.00) |
| ZNF721 | 13 (5%) | 235 |
0.263 (0.775) |
0.0494 (0.725) |
0.302 (0.779) |
0.551 (0.882) |
0.646 (0.936) |
0.187 (0.756) |
0.196 (0.756) |
| MCTP1 | 13 (5%) | 235 |
0.238 (0.769) |
0.0153 (0.485) |
0.576 (0.896) |
0.382 (0.826) |
0.647 (0.936) |
0.482 (0.854) |
0.196 (0.756) |
| ZNF774 | 10 (4%) | 238 |
0.36 (0.805) |
0.581 (0.897) |
0.741 (0.993) |
1 (1.00) |
0.606 (0.908) |
0.711 (0.981) |
0.134 (0.756) |
| FAM9A | 14 (6%) | 234 |
0.337 (0.802) |
0.836 (1.00) |
0.433 (0.826) |
0.567 (0.895) |
0.842 (1.00) |
0.683 (0.963) |
1 (1.00) |
| CFP | 8 (3%) | 240 |
0.0127 (0.472) |
0.932 (1.00) |
0.435 (0.826) |
1 (1.00) |
0.211 (0.756) |
0.353 (0.802) |
1 (1.00) |
| PER3 | 12 (5%) | 236 |
0.273 (0.775) |
0.521 (0.871) |
0.287 (0.775) |
1 (1.00) |
1 (1.00) |
0.56 (0.889) |
1 (1.00) |
| ZRANB3 | 8 (3%) | 240 |
0.257 (0.772) |
0.918 (1.00) |
1 (1.00) |
0.454 (0.834) |
1 (1.00) |
0.273 (0.775) |
1 (1.00) |
| OMA1 | 10 (4%) | 238 |
0.35 (0.802) |
0.09 (0.756) |
0.334 (0.802) |
0.172 (0.756) |
0.547 (0.882) |
0.274 (0.775) |
0.134 (0.756) |
| CLDN15 | 5 (2%) | 243 |
0.55 (0.882) |
0.327 (0.802) |
0.619 (0.92) |
0.661 (0.949) |
0.416 (0.826) |
0.265 (0.775) |
0.113 (0.756) |
| TTC39C | 7 (3%) | 241 |
0.343 (0.802) |
0.344 (0.802) |
0.428 (0.826) |
1 (1.00) |
1 (1.00) |
0.597 (0.907) |
1 (1.00) |
| TXNRD1 | 8 (3%) | 240 |
0.296 (0.775) |
0.632 (0.929) |
0.439 (0.827) |
0.718 (0.983) |
0.361 (0.805) |
0.825 (1.00) |
1 (1.00) |
| MECOM | 12 (5%) | 236 |
0.198 (0.756) |
0.0117 (0.46) |
0.286 (0.775) |
1 (1.00) |
1 (1.00) |
0.811 (1.00) |
1 (1.00) |
| CCDC147 | 15 (6%) | 233 |
0.2 (0.756) |
0.0312 (0.623) |
0.47 (0.842) |
1 (1.00) |
0.684 (0.963) |
0.28 (0.775) |
0.196 (0.756) |
| SACS | 26 (10%) | 222 |
0.0694 (0.756) |
0.855 (1.00) |
0.12 (0.756) |
1 (1.00) |
0.709 (0.98) |
0.0494 (0.725) |
0.00311 (0.278) |
| MUTED | 7 (3%) | 241 |
0.352 (0.802) |
0.748 (0.993) |
0.427 (0.826) |
1 (1.00) |
1 (1.00) |
0.0962 (0.756) |
1 (1.00) |
| SLC44A3 | 6 (2%) | 242 |
0.469 (0.842) |
0.0571 (0.756) |
0.634 (0.929) |
1 (1.00) |
0.62 (0.92) |
0.424 (0.826) |
1 (1.00) |
| ZNF674 | 14 (6%) | 234 |
0.162 (0.756) |
0.101 (0.756) |
0.588 (0.897) |
0.567 (0.895) |
0.532 (0.882) |
0.196 (0.756) |
1 (1.00) |
| CCDC144A | 18 (7%) | 230 |
0.516 (0.869) |
0.128 (0.756) |
0.661 (0.949) |
0.614 (0.914) |
0.346 (0.802) |
0.036 (0.653) |
0.255 (0.772) |
| OR8B8 | 7 (3%) | 241 |
0.425 (0.826) |
0.0145 (0.481) |
0.429 (0.826) |
0.427 (0.826) |
0.414 (0.826) |
0.288 (0.775) |
0.113 (0.756) |
| LNX2 | 14 (6%) | 234 |
0.215 (0.756) |
0.843 (1.00) |
0.171 (0.756) |
0.147 (0.756) |
0.875 (1.00) |
0.184 (0.756) |
0.235 (0.766) |
| TMEM62 | 9 (4%) | 239 |
0.343 (0.802) |
0.506 (0.857) |
0.457 (0.834) |
0.723 (0.986) |
0.602 (0.908) |
0.355 (0.802) |
0.155 (0.756) |
| ATF7IP | 17 (7%) | 231 |
0.135 (0.756) |
0.195 (0.756) |
0.0991 (0.756) |
0.431 (0.826) |
0.887 (1.00) |
0.699 (0.975) |
0.255 (0.772) |
| IL20 | 7 (3%) | 241 |
0.338 (0.802) |
0.169 (0.756) |
0.426 (0.826) |
0.24 (0.77) |
0.132 (0.756) |
0.0136 (0.481) |
0.0911 (0.756) |
| C3AR1 | 8 (3%) | 240 |
0.254 (0.772) |
0.878 (1.00) |
0.438 (0.826) |
1 (1.00) |
0.605 (0.908) |
0.179 (0.756) |
0.134 (0.756) |
| PKD2 | 8 (3%) | 240 |
0.37 (0.812) |
0.154 (0.756) |
0.436 (0.826) |
0.268 (0.775) |
0.548 (0.882) |
0.347 (0.802) |
0.134 (0.756) |
| BMP5 | 13 (5%) | 235 |
0.174 (0.756) |
0.146 (0.756) |
0.573 (0.896) |
0.551 (0.882) |
0.649 (0.936) |
0.0436 (0.687) |
0.216 (0.756) |
| OR52I2 | 8 (3%) | 240 |
0.999 (1.00) |
0.679 (0.963) |
0.157 (0.756) |
0.454 (0.834) |
0.788 (1.00) |
0.35 (0.802) |
1 (1.00) |
| C1ORF100 | 9 (4%) | 239 |
0.277 (0.775) |
0.139 (0.756) |
0.454 (0.834) |
0.502 (0.854) |
0.482 (0.854) |
0.352 (0.802) |
0.0911 (0.756) |
| CCDC160 | 11 (4%) | 237 |
0.224 (0.766) |
0.204 (0.756) |
0.354 (0.802) |
1 (1.00) |
0.843 (1.00) |
0.401 (0.826) |
1 (1.00) |
| NAA15 | 14 (6%) | 234 |
0.587 (0.897) |
0.991 (1.00) |
0.586 (0.897) |
0.252 (0.772) |
0.874 (1.00) |
0.232 (0.766) |
0.196 (0.756) |
| RNF31 | 17 (7%) | 231 |
0.218 (0.756) |
0.692 (0.969) |
0.807 (1.00) |
0.602 (0.908) |
0.258 (0.772) |
0.138 (0.756) |
0.216 (0.756) |
| NFE2L3 | 12 (5%) | 236 |
0.254 (0.772) |
0.781 (1.00) |
0.216 (0.756) |
0.117 (0.756) |
0.424 (0.826) |
0.206 (0.756) |
1 (1.00) |
| ZKSCAN1 | 7 (3%) | 241 |
0.41 (0.826) |
0.968 (1.00) |
1 (1.00) |
0.696 (0.973) |
0.742 (0.993) |
0.599 (0.907) |
1 (1.00) |
| CDKL1 | 7 (3%) | 241 |
0.427 (0.826) |
0.0112 (0.46) |
0.427 (0.826) |
1 (1.00) |
0.427 (0.826) |
0.0747 (0.756) |
0.113 (0.756) |
| ERBB3 | 17 (7%) | 231 |
0.185 (0.756) |
0.598 (0.907) |
0.491 (0.854) |
0.602 (0.908) |
0.292 (0.775) |
0.0904 (0.756) |
1 (1.00) |
| SELP | 9 (4%) | 239 |
0.983 (1.00) |
0.105 (0.756) |
0.456 (0.834) |
1 (1.00) |
0.212 (0.756) |
0.06 (0.756) |
0.134 (0.756) |
| PPIG | 16 (6%) | 232 |
0.256 (0.772) |
0.0726 (0.756) |
0.119 (0.756) |
0.187 (0.756) |
0.913 (1.00) |
0.931 (1.00) |
0.235 (0.766) |
| GPRASP1 | 21 (8%) | 227 |
0.144 (0.756) |
0.17 (0.756) |
0.275 (0.775) |
0.812 (1.00) |
0.643 (0.936) |
0.675 (0.96) |
0.273 (0.775) |
| REV3L | 20 (8%) | 228 |
0.121 (0.756) |
0.503 (0.854) |
0.0683 (0.756) |
0.464 (0.839) |
1 (1.00) |
0.152 (0.756) |
0.273 (0.775) |
| CTNNA1 | 13 (5%) | 235 |
0.203 (0.756) |
0.428 (0.826) |
0.573 (0.896) |
0.771 (1.00) |
0.416 (0.826) |
0.0306 (0.623) |
1 (1.00) |
| MRPL47 | 6 (2%) | 242 |
0.487 (0.854) |
0.92 (1.00) |
1 (1.00) |
0.421 (0.826) |
0.481 (0.854) |
1 (1.00) |
0.0911 (0.756) |
| DEPDC1B | 11 (4%) | 237 |
0.992 (1.00) |
0.592 (0.902) |
0.745 (0.993) |
0.521 (0.871) |
0.193 (0.756) |
0.049 (0.725) |
1 (1.00) |
| CASP8 | 17 (7%) | 231 |
0.561 (0.889) |
0.748 (0.993) |
0.489 (0.854) |
1 (1.00) |
0.545 (0.882) |
0.028 (0.623) |
0.0295 (0.623) |
| CACNB4 | 10 (4%) | 238 |
0.353 (0.802) |
0.19 (0.756) |
0.336 (0.802) |
0.501 (0.854) |
0.492 (0.854) |
0.112 (0.756) |
1 (1.00) |
| BHLHB9 | 8 (3%) | 240 |
0.388 (0.826) |
0.398 (0.826) |
0.436 (0.826) |
1 (1.00) |
0.743 (0.993) |
0.404 (0.826) |
1 (1.00) |
| C22ORF25 | 6 (2%) | 242 |
0.495 (0.854) |
0.984 (1.00) |
0.636 (0.929) |
0.0201 (0.544) |
0.675 (0.96) |
0.0689 (0.756) |
|
| GALNT7 | 11 (4%) | 237 |
0.684 (0.963) |
0.435 (0.826) |
0.744 (0.993) |
0.103 (0.756) |
0.0243 (0.588) |
0.0167 (0.5) |
0.176 (0.756) |
| LGMN | 7 (3%) | 241 |
0.0572 (0.756) |
0.987 (1.00) |
0.427 (0.826) |
0.24 (0.77) |
0.288 (0.775) |
1 (1.00) |
1 (1.00) |
| EPS8 | 12 (5%) | 236 |
0.927 (1.00) |
0.124 (0.756) |
0.751 (0.993) |
1 (1.00) |
0.702 (0.975) |
0.403 (0.826) |
0.196 (0.756) |
| TGOLN2 | 3 (1%) | 245 |
0.575 (0.896) |
0.0514 (0.736) |
1 (1.00) |
0.553 (0.882) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
| TPX2 | 6 (2%) | 242 |
0.567 (0.895) |
0.0114 (0.46) |
0.635 (0.929) |
1 (1.00) |
0.141 (0.756) |
0.672 (0.957) |
0.0689 (0.756) |
| C14ORF118 | 15 (6%) | 233 |
0.237 (0.768) |
0.0161 (0.492) |
0.141 (0.756) |
0.587 (0.897) |
0.872 (1.00) |
0.502 (0.854) |
0.196 (0.756) |
| SLC34A3 | 6 (2%) | 242 |
0.445 (0.834) |
0.108 (0.756) |
0.633 (0.929) |
0.421 (0.826) |
1 (1.00) |
0.233 (0.766) |
1 (1.00) |
| ZNF662 | 13 (5%) | 235 |
0.221 (0.761) |
0.493 (0.854) |
0.574 (0.896) |
0.771 (1.00) |
0.645 (0.936) |
0.278 (0.775) |
0.176 (0.756) |
| ZDBF2 | 18 (7%) | 230 |
0.159 (0.756) |
0.186 (0.756) |
0.0856 (0.756) |
0.311 (0.787) |
0.763 (0.999) |
0.725 (0.987) |
0.255 (0.772) |
| SFRP4 | 8 (3%) | 240 |
0.317 (0.799) |
0.124 (0.756) |
0.437 (0.826) |
1 (1.00) |
0.428 (0.826) |
0.346 (0.802) |
0.113 (0.756) |
| CCDC146 | 14 (6%) | 234 |
0.217 (0.756) |
0.033 (0.633) |
0.173 (0.756) |
0.391 (0.826) |
0.646 (0.936) |
0.229 (0.766) |
0.196 (0.756) |
| C7ORF60 | 10 (4%) | 238 |
0.302 (0.779) |
0.172 (0.756) |
0.337 (0.802) |
0.322 (0.802) |
0.605 (0.908) |
1 (1.00) |
0.155 (0.756) |
| ZNF649 | 14 (6%) | 234 |
0.233 (0.766) |
0.185 (0.756) |
0.173 (0.756) |
0.776 (1.00) |
0.875 (1.00) |
0.909 (1.00) |
0.196 (0.756) |
| RASSF9 | 9 (4%) | 239 |
0.264 (0.775) |
0.0134 (0.481) |
0.458 (0.834) |
0.723 (0.986) |
0.603 (0.908) |
0.167 (0.756) |
0.134 (0.756) |
| NIPA2 | 8 (3%) | 240 |
0.35 (0.802) |
0.986 (1.00) |
1 (1.00) |
0.718 (0.983) |
1 (1.00) |
0.0338 (0.638) |
1 (1.00) |
| FILIP1 | 16 (6%) | 232 |
0.149 (0.756) |
0.596 (0.906) |
0.48 (0.853) |
0.187 (0.756) |
0.682 (0.963) |
0.802 (1.00) |
0.176 (0.756) |
| PHKA2 | 16 (6%) | 232 |
0.152 (0.756) |
0.538 (0.882) |
0.479 (0.853) |
0.792 (1.00) |
0.795 (1.00) |
0.0806 (0.756) |
1 (1.00) |
| COBLL1 | 15 (6%) | 233 |
0.207 (0.756) |
0.979 (1.00) |
0.139 (0.756) |
0.587 (0.897) |
0.372 (0.814) |
0.503 (0.854) |
0.216 (0.756) |
| AGPAT9 | 8 (3%) | 240 |
0.295 (0.775) |
0.63 (0.929) |
0.435 (0.826) |
0.0535 (0.752) |
0.278 (0.775) |
0.176 (0.756) |
0.134 (0.756) |
| NOP56 | 11 (4%) | 237 |
0.248 (0.772) |
0.137 (0.756) |
0.352 (0.802) |
0.752 (0.993) |
0.829 (1.00) |
0.398 (0.826) |
1 (1.00) |
| RBL2 | 12 (5%) | 236 |
0.766 (1.00) |
0.211 (0.756) |
0.287 (0.775) |
1 (1.00) |
0.193 (0.756) |
0.152 (0.756) |
0.196 (0.756) |
| ABCC6 | 12 (5%) | 236 |
0.476 (0.852) |
0.545 (0.882) |
0.0583 (0.756) |
0.117 (0.756) |
0.287 (0.775) |
0.876 (1.00) |
0.176 (0.756) |
| CCDC104 | 10 (4%) | 238 |
0.284 (0.775) |
0.0971 (0.756) |
0.337 (0.802) |
0.501 (0.854) |
1 (1.00) |
0.585 (0.897) |
1 (1.00) |
| MLL2 | 33 (13%) | 215 |
0.0883 (0.756) |
0.55 (0.882) |
0.146 (0.756) |
1 (1.00) |
0.867 (1.00) |
0.092 (0.756) |
0.442 (0.829) |
| SSH2 | 12 (5%) | 236 |
0.239 (0.769) |
0.208 (0.756) |
0.753 (0.993) |
0.757 (0.993) |
0.686 (0.963) |
0.00772 (0.447) |
0.0174 (0.511) |
| ZNF709 | 12 (5%) | 236 |
0.206 (0.756) |
0.246 (0.772) |
0.287 (0.775) |
1 (1.00) |
0.705 (0.978) |
0.15 (0.756) |
0.155 (0.756) |
| BRDT | 14 (6%) | 234 |
0.213 (0.756) |
0.0842 (0.756) |
0.171 (0.756) |
0.391 (0.826) |
0.648 (0.936) |
0.185 (0.756) |
0.196 (0.756) |
| SH3BGRL | 6 (2%) | 242 |
0.391 (0.826) |
0.26 (0.775) |
0.634 (0.929) |
0.668 (0.956) |
1 (1.00) |
0.029 (0.623) |
1 (1.00) |
| MLH3 | 17 (7%) | 231 |
0.668 (0.956) |
0.00875 (0.447) |
0.491 (0.854) |
1 (1.00) |
0.257 (0.772) |
0.174 (0.756) |
0.255 (0.772) |
| CCDC150 | 12 (5%) | 236 |
0.273 (0.775) |
0.931 (1.00) |
0.756 (0.993) |
1 (1.00) |
0.858 (1.00) |
0.894 (1.00) |
1 (1.00) |
| NT5C3 | 7 (3%) | 241 |
0.321 (0.802) |
0.506 (0.857) |
0.425 (0.826) |
0.696 (0.973) |
0.549 (0.882) |
0.128 (0.756) |
0.113 (0.756) |
| APAF1 | 13 (5%) | 235 |
0.635 (0.929) |
0.44 (0.827) |
0.301 (0.779) |
0.551 (0.882) |
0.206 (0.756) |
0.158 (0.756) |
0.176 (0.756) |
| KANK4 | 11 (4%) | 237 |
0.902 (1.00) |
0.043 (0.687) |
0.744 (0.993) |
0.521 (0.871) |
1 (1.00) |
0.516 (0.869) |
1 (1.00) |
| NRIP1 | 13 (5%) | 235 |
0.244 (0.772) |
0.117 (0.756) |
0.576 (0.896) |
1 (1.00) |
0.422 (0.826) |
0.36 (0.805) |
1 (1.00) |
| GTF2H1 | 8 (3%) | 240 |
0.351 (0.802) |
0.946 (1.00) |
0.435 (0.826) |
0.718 (0.983) |
0.548 (0.882) |
0.5 (0.854) |
0.134 (0.756) |
| ASXL2 | 14 (6%) | 234 |
0.208 (0.756) |
0.381 (0.824) |
0.584 (0.897) |
1 (1.00) |
0.221 (0.761) |
0.524 (0.874) |
0.176 (0.756) |
| ZNF611 | 12 (5%) | 236 |
0.913 (1.00) |
0.244 (0.772) |
0.291 (0.775) |
1 (1.00) |
0.205 (0.756) |
0.206 (0.756) |
0.196 (0.756) |
| SUN3 | 6 (2%) | 242 |
0.426 (0.826) |
0.0127 (0.472) |
0.633 (0.929) |
1 (1.00) |
0.416 (0.826) |
0.202 (0.756) |
0.00244 (0.259) |
| ZNF195 | 11 (4%) | 237 |
0.283 (0.775) |
0.023 (0.566) |
0.355 (0.802) |
0.752 (0.993) |
0.656 (0.943) |
0.119 (0.756) |
0.176 (0.756) |
| EXOSC9 | 10 (4%) | 238 |
0.288 (0.775) |
0.78 (1.00) |
0.334 (0.802) |
1 (1.00) |
0.603 (0.908) |
0.412 (0.826) |
0.155 (0.756) |
| MFN2 | 9 (4%) | 239 |
0.402 (0.826) |
0.357 (0.804) |
0.454 (0.834) |
0.502 (0.854) |
1 (1.00) |
0.483 (0.854) |
0.134 (0.756) |
| LIMA1 | 8 (3%) | 240 |
0.407 (0.826) |
0.888 (1.00) |
0.437 (0.826) |
0.718 (0.983) |
1 (1.00) |
0.662 (0.949) |
1 (1.00) |
| PTPN12 | 10 (4%) | 238 |
0.907 (1.00) |
0.829 (1.00) |
0.337 (0.802) |
0.501 (0.854) |
0.606 (0.908) |
0.232 (0.766) |
0.155 (0.756) |
| MTF2 | 9 (4%) | 239 |
0.369 (0.812) |
0.0734 (0.756) |
0.456 (0.834) |
1 (1.00) |
1 (1.00) |
0.191 (0.756) |
1 (1.00) |
| MLL4 | 30 (12%) | 218 |
0.0597 (0.756) |
0.245 (0.772) |
0.0425 (0.687) |
0.839 (1.00) |
0.651 (0.939) |
0.295 (0.775) |
0.427 (0.826) |
| C1ORF101 | 12 (5%) | 236 |
0.258 (0.772) |
0.0191 (0.528) |
0.287 (0.775) |
1 (1.00) |
0.742 (0.993) |
0.149 (0.756) |
0.196 (0.756) |
| MBOAT2 | 7 (3%) | 241 |
0.394 (0.826) |
0.0885 (0.756) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.291 (0.775) |
1 (1.00) |
| CHD4 | 35 (14%) | 213 |
0.428 (0.826) |
0.425 (0.826) |
0.453 (0.834) |
0.45 (0.834) |
1 (1.00) |
0.736 (0.993) |
1 (1.00) |
| SESN3 | 13 (5%) | 235 |
0.219 (0.758) |
0.468 (0.842) |
0.304 (0.781) |
0.771 (1.00) |
0.7 (0.975) |
0.896 (1.00) |
0.196 (0.756) |
P value = 1e-05 (Fisher's exact test), Q value = 0.0028
Table S1. Gene #1: 'PTEN MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|
| ALL | 200 | 4 | 44 |
| PTEN MUTATED | 159 | 1 | 1 |
| PTEN WILD-TYPE | 41 | 3 | 43 |
Figure S1. Get High-res Image Gene #1: 'PTEN MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
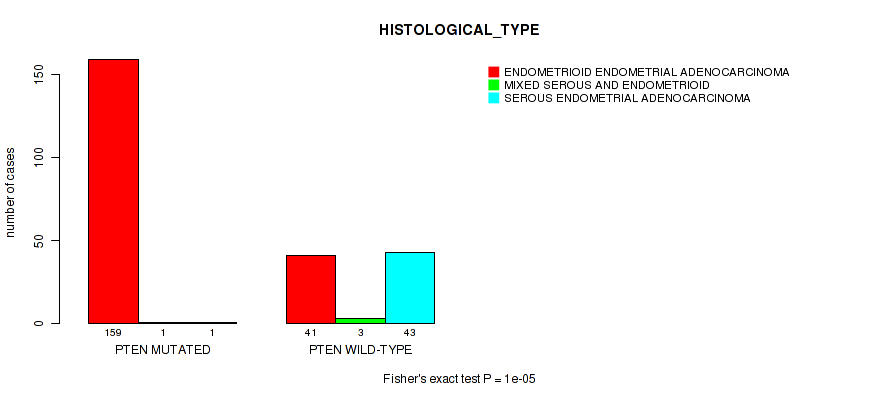
P value = 0.00063 (Fisher's exact test), Q value = 0.072
Table S2. Gene #1: 'PTEN MUTATION STATUS' versus Clinical Feature #6: 'RACE'
| nPatients | AMERICAN INDIAN OR ALASKA NATIVE | ASIAN | BLACK OR AFRICAN AMERICAN | NATIVE HAWAIIAN OR OTHER PACIFIC ISLANDER | WHITE |
|---|---|---|---|---|---|
| ALL | 3 | 13 | 25 | 7 | 193 |
| PTEN MUTATED | 3 | 10 | 7 | 4 | 133 |
| PTEN WILD-TYPE | 0 | 3 | 18 | 3 | 60 |
Figure S2. Get High-res Image Gene #1: 'PTEN MUTATION STATUS' versus Clinical Feature #6: 'RACE'
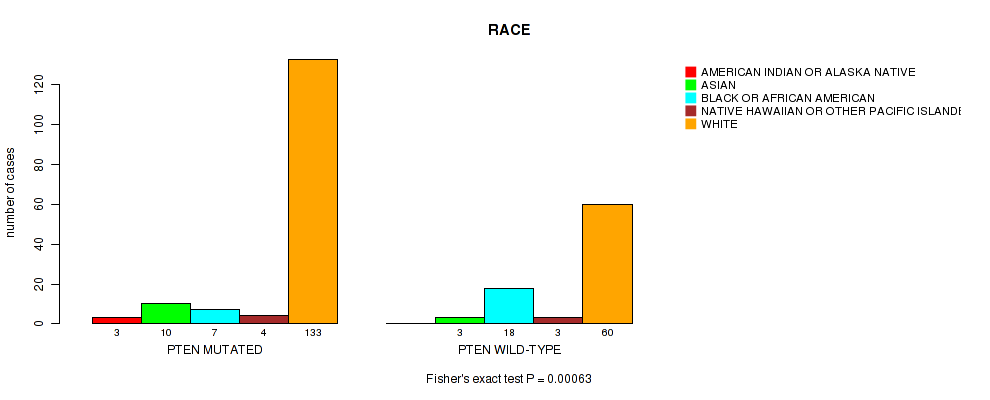
P value = 1e-05 (Fisher's exact test), Q value = 0.0028
Table S3. Gene #2: 'PIK3R1 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|
| ALL | 200 | 4 | 44 |
| PIK3R1 MUTATED | 80 | 1 | 2 |
| PIK3R1 WILD-TYPE | 120 | 3 | 42 |
Figure S3. Get High-res Image Gene #2: 'PIK3R1 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
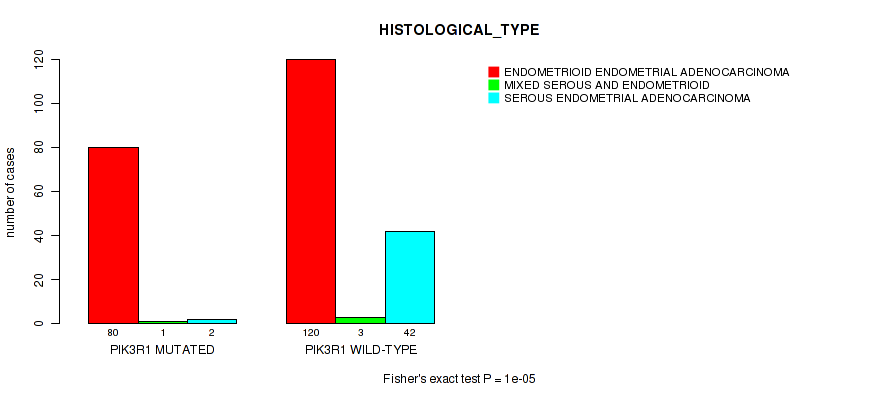
P value = 5.85e-06 (Wilcoxon-test), Q value = 0.0028
Table S4. Gene #3: 'TP53 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 248 | 63.1 (11.1) |
| TP53 MUTATED | 69 | 67.9 (9.4) |
| TP53 WILD-TYPE | 179 | 61.3 (11.2) |
Figure S4. Get High-res Image Gene #3: 'TP53 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
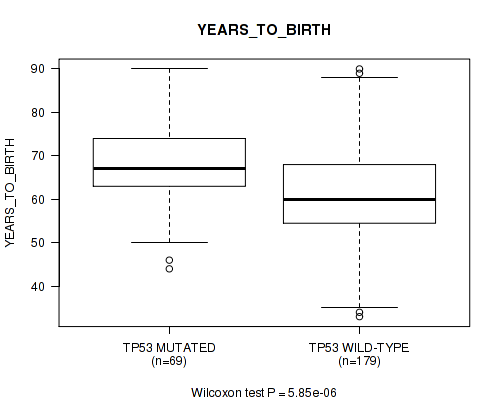
P value = 1e-05 (Fisher's exact test), Q value = 0.0028
Table S5. Gene #3: 'TP53 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|
| ALL | 200 | 4 | 44 |
| TP53 MUTATED | 27 | 3 | 39 |
| TP53 WILD-TYPE | 173 | 1 | 5 |
Figure S5. Get High-res Image Gene #3: 'TP53 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
P value = 0.00017 (Fisher's exact test), Q value = 0.033
Table S6. Gene #4: 'CTCF MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|
| ALL | 200 | 4 | 44 |
| CTCF MUTATED | 44 | 0 | 0 |
| CTCF WILD-TYPE | 156 | 4 | 44 |
Figure S6. Get High-res Image Gene #4: 'CTCF MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
P value = 0.000544 (Wilcoxon-test), Q value = 0.068
Table S7. Gene #6: 'KRAS MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 248 | 63.1 (11.1) |
| KRAS MUTATED | 53 | 58.8 (8.7) |
| KRAS WILD-TYPE | 195 | 64.3 (11.4) |
Figure S7. Get High-res Image Gene #6: 'KRAS MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
P value = 0.00022 (Fisher's exact test), Q value = 0.038
Table S8. Gene #6: 'KRAS MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|
| ALL | 200 | 4 | 44 |
| KRAS MUTATED | 52 | 0 | 1 |
| KRAS WILD-TYPE | 148 | 4 | 43 |
Figure S8. Get High-res Image Gene #6: 'KRAS MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
P value = 9e-05 (Fisher's exact test), Q value = 0.021
Table S9. Gene #9: 'ARID1A MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|
| ALL | 200 | 4 | 44 |
| ARID1A MUTATED | 78 | 1 | 4 |
| ARID1A WILD-TYPE | 122 | 3 | 40 |
Figure S9. Get High-res Image Gene #9: 'ARID1A MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
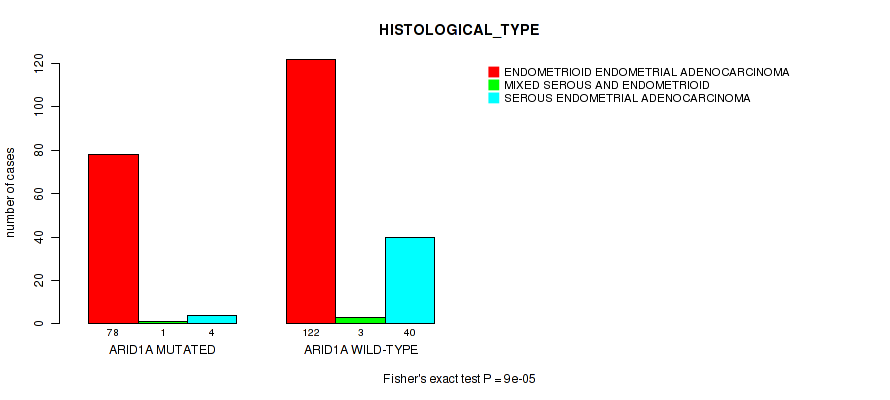
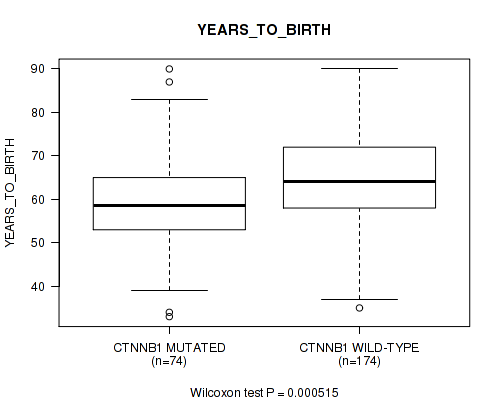
P value = 0.000515 (Wilcoxon-test), Q value = 0.068
Table S10. Gene #10: 'CTNNB1 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 248 | 63.1 (11.1) |
| CTNNB1 MUTATED | 74 | 59.6 (11.6) |
| CTNNB1 WILD-TYPE | 174 | 64.6 (10.6) |
Figure S10. Get High-res Image Gene #10: 'CTNNB1 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
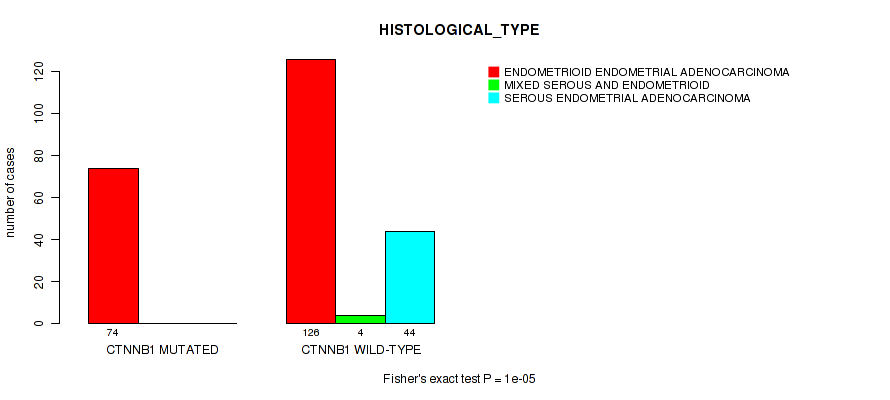
P value = 1e-05 (Fisher's exact test), Q value = 0.0028
Table S11. Gene #10: 'CTNNB1 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|
| ALL | 200 | 4 | 44 |
| CTNNB1 MUTATED | 74 | 0 | 0 |
| CTNNB1 WILD-TYPE | 126 | 4 | 44 |
Figure S11. Get High-res Image Gene #10: 'CTNNB1 MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
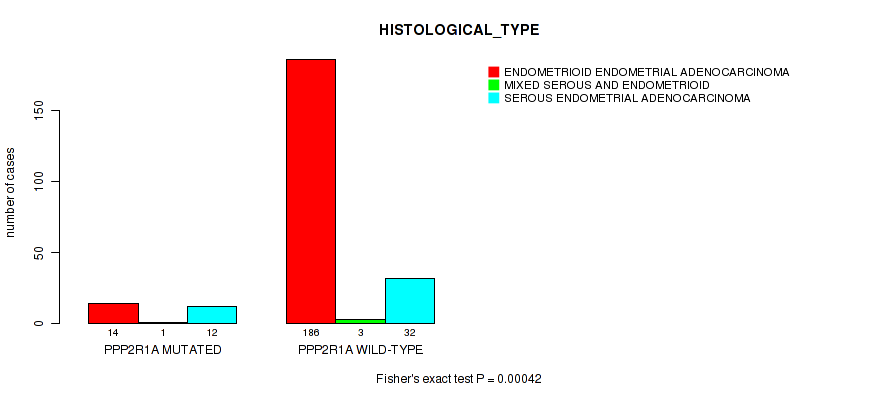
P value = 0.00042 (Fisher's exact test), Q value = 0.064
Table S12. Gene #43: 'PPP2R1A MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
| nPatients | ENDOMETRIOID ENDOMETRIAL ADENOCARCINOMA | MIXED SEROUS AND ENDOMETRIOID | SEROUS ENDOMETRIAL ADENOCARCINOMA |
|---|---|---|---|
| ALL | 200 | 4 | 44 |
| PPP2R1A MUTATED | 14 | 1 | 12 |
| PPP2R1A WILD-TYPE | 186 | 3 | 32 |
Figure S12. Get High-res Image Gene #43: 'PPP2R1A MUTATION STATUS' versus Clinical Feature #3: 'HISTOLOGICAL_TYPE'
-
Mutation data file = sample_sig_gene_table.txt from Mutsig_2CV pipeline
-
Processed Mutation data file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_Correlate_Genomic_Events_Preprocess/UCEC-TP/15165175/transformed.cor.cli.txt
-
Clinical data file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/Append_Data/UCEC-TP/15094079/UCEC-TP.merged_data.txt
-
Number of patients = 248
-
Number of significantly mutated genes = 197
-
Number of selected clinical features = 7
-
Exclude genes that fewer than K tumors have mutations, K = 3
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.