This pipeline computes the correlation between significant arm-level copy number variations (cnvs) and molecular subtypes.
Testing the association between copy number variation 52 arm-level events and 10 molecular subtypes across 48 patients, 2 significant findings detected with P value < 0.05 and Q value < 0.25.
-
7p gain cnv correlated to 'CN_CNMF'.
-
7q gain cnv correlated to 'CN_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 52 arm-level events and 10 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 2 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
RPPA CNMF |
RPPA CHIERARCHICAL |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| 7p gain | 15 (31%) | 33 |
1.04e-05 (0.0027) |
0.534 (1.00) |
0.699 (1.00) |
0.763 (1.00) |
1 (1.00) |
0.456 (1.00) |
0.924 (1.00) |
0.887 (1.00) |
1 (1.00) |
0.125 (1.00) |
| 7q gain | 13 (27%) | 35 |
7.71e-06 (0.0027) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.67 (1.00) |
0.122 (1.00) |
0.661 (1.00) |
0.988 (1.00) |
1 (1.00) |
0.382 (1.00) |
| 1q gain | 6 (12%) | 42 |
0.197 (1.00) |
0.666 (1.00) |
0.233 (1.00) |
0.276 (1.00) |
1 (1.00) |
0.638 (1.00) |
0.413 (1.00) |
0.313 (1.00) |
1 (1.00) |
0.842 (1.00) |
| 2p gain | 6 (12%) | 42 |
0.669 (1.00) |
0.666 (1.00) |
1 (1.00) |
0.826 (1.00) |
1 (1.00) |
1 (1.00) |
0.605 (1.00) |
1 (1.00) |
0.22 (1.00) |
|
| 2q gain | 6 (12%) | 42 |
0.669 (1.00) |
0.666 (1.00) |
1 (1.00) |
0.825 (1.00) |
1 (1.00) |
1 (1.00) |
0.607 (1.00) |
1 (1.00) |
0.22 (1.00) |
|
| 3p gain | 10 (21%) | 38 |
0.0363 (0.768) |
0.286 (1.00) |
1 (1.00) |
0.249 (1.00) |
1 (1.00) |
0.677 (1.00) |
0.633 (1.00) |
0.404 (1.00) |
0.707 (1.00) |
0.348 (1.00) |
| 3q gain | 13 (27%) | 35 |
0.0188 (0.691) |
0.193 (1.00) |
0.448 (1.00) |
0.0369 (0.768) |
1 (1.00) |
0.516 (1.00) |
0.765 (1.00) |
0.565 (1.00) |
1 (1.00) |
0.248 (1.00) |
| 5p gain | 7 (15%) | 41 |
0.687 (1.00) |
1 (1.00) |
0.639 (1.00) |
0.684 (1.00) |
1 (1.00) |
0.341 (1.00) |
0.641 (1.00) |
0.356 (1.00) |
0.119 (1.00) |
|
| 5q gain | 6 (12%) | 42 |
1 (1.00) |
1 (1.00) |
0.639 (1.00) |
0.686 (1.00) |
0.464 (1.00) |
0.66 (1.00) |
0.797 (1.00) |
0.613 (1.00) |
0.534 (1.00) |
|
| 6p gain | 6 (12%) | 42 |
0.197 (1.00) |
1 (1.00) |
0.346 (1.00) |
0.826 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.862 (1.00) |
0.644 (1.00) |
0.883 (1.00) |
| 6q gain | 4 (8%) | 44 |
0.0199 (0.691) |
0.609 (1.00) |
1 (1.00) |
0.769 (1.00) |
1 (1.00) |
0.71 (1.00) |
1 (1.00) |
0.0844 (1.00) |
0.874 (1.00) |
|
| 8p gain | 7 (15%) | 41 |
0.687 (1.00) |
0.416 (1.00) |
1 (1.00) |
1 (1.00) |
0.206 (1.00) |
0.102 (1.00) |
0.00548 (0.57) |
0.428 (1.00) |
0.258 (1.00) |
|
| 8q gain | 8 (17%) | 40 |
0.451 (1.00) |
0.245 (1.00) |
0.639 (1.00) |
0.827 (1.00) |
0.206 (1.00) |
0.147 (1.00) |
0.0277 (0.768) |
0.428 (1.00) |
0.258 (1.00) |
|
| 9p gain | 7 (15%) | 41 |
0.687 (1.00) |
1 (1.00) |
1 (1.00) |
0.438 (1.00) |
1 (1.00) |
0.775 (1.00) |
0.64 (1.00) |
1 (1.00) |
0.465 (1.00) |
|
| 9q gain | 7 (15%) | 41 |
0.687 (1.00) |
1 (1.00) |
1 (1.00) |
0.438 (1.00) |
1 (1.00) |
0.485 (1.00) |
0.293 (1.00) |
1 (1.00) |
0.267 (1.00) |
|
| 10p gain | 4 (8%) | 44 |
0.0199 (0.691) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.581 (1.00) |
0.64 (1.00) |
||
| 10q gain | 4 (8%) | 44 |
0.286 (1.00) |
0.609 (1.00) |
1 (1.00) |
0.768 (1.00) |
0.464 (1.00) |
0.425 (1.00) |
0.777 (1.00) |
0.0844 (1.00) |
0.26 (1.00) |
|
| 11p gain | 9 (19%) | 39 |
0.451 (1.00) |
0.461 (1.00) |
1 (1.00) |
0.379 (1.00) |
0.6 (1.00) |
0.145 (1.00) |
0.72 (1.00) |
0.163 (1.00) |
0.433 (1.00) |
0.228 (1.00) |
| 11q gain | 13 (27%) | 35 |
0.0959 (1.00) |
0.193 (1.00) |
0.665 (1.00) |
0.357 (1.00) |
0.686 (1.00) |
0.0861 (1.00) |
0.448 (1.00) |
0.0729 (1.00) |
0.468 (1.00) |
0.401 (1.00) |
| 12p gain | 7 (15%) | 41 |
1 (1.00) |
0.416 (1.00) |
1 (1.00) |
0.3 (1.00) |
0.583 (1.00) |
0.11 (1.00) |
0.815 (1.00) |
0.273 (1.00) |
0.384 (1.00) |
0.863 (1.00) |
| 12q gain | 9 (19%) | 39 |
1 (1.00) |
0.137 (1.00) |
0.674 (1.00) |
0.475 (1.00) |
1 (1.00) |
0.148 (1.00) |
0.646 (1.00) |
0.136 (1.00) |
0.433 (1.00) |
0.491 (1.00) |
| 13q gain | 5 (10%) | 43 |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.206 (1.00) |
0.135 (1.00) |
0.66 (1.00) |
0.0346 (0.768) |
0.838 (1.00) |
|
| 16p gain | 7 (15%) | 41 |
0.0114 (0.691) |
0.416 (1.00) |
1 (1.00) |
1 (1.00) |
0.6 (1.00) |
0.21 (1.00) |
0.326 (1.00) |
0.0282 (0.768) |
0.384 (1.00) |
0.512 (1.00) |
| 16q gain | 7 (15%) | 41 |
0.0967 (1.00) |
0.0971 (1.00) |
1 (1.00) |
1 (1.00) |
0.6 (1.00) |
1 (1.00) |
0.327 (1.00) |
0.0138 (0.691) |
0.384 (1.00) |
0.706 (1.00) |
| 17q gain | 3 (6%) | 45 |
0.554 (1.00) |
0.234 (1.00) |
1 (1.00) |
0.77 (1.00) |
0.464 (1.00) |
0.608 (1.00) |
0.0358 (0.768) |
0.581 (1.00) |
0.496 (1.00) |
|
| 18p gain | 13 (27%) | 35 |
0.0959 (1.00) |
1 (1.00) |
0.134 (1.00) |
0.779 (1.00) |
1 (1.00) |
0.677 (1.00) |
0.364 (1.00) |
0.122 (1.00) |
0.504 (1.00) |
0.524 (1.00) |
| 18q gain | 14 (29%) | 34 |
0.0491 (0.913) |
1 (1.00) |
0.0696 (1.00) |
0.687 (1.00) |
0.655 (1.00) |
1 (1.00) |
0.452 (1.00) |
0.116 (1.00) |
0.323 (1.00) |
0.404 (1.00) |
| 19p gain | 3 (6%) | 45 |
0.554 (1.00) |
1 (1.00) |
1 (1.00) |
0.464 (1.00) |
0.687 (1.00) |
0.689 (1.00) |
0.452 (1.00) |
|||
| 19q gain | 3 (6%) | 45 |
0.554 (1.00) |
1 (1.00) |
1 (1.00) |
0.464 (1.00) |
0.688 (1.00) |
0.691 (1.00) |
0.452 (1.00) |
|||
| 20p gain | 5 (10%) | 43 |
0.372 (1.00) |
0.348 (1.00) |
0.607 (1.00) |
0.154 (1.00) |
1 (1.00) |
0.289 (1.00) |
0.377 (1.00) |
1 (1.00) |
0.7 (1.00) |
|
| 20q gain | 4 (8%) | 44 |
1 (1.00) |
0.609 (1.00) |
1 (1.00) |
0.117 (1.00) |
0.464 (1.00) |
0.656 (1.00) |
0.818 (1.00) |
0.581 (1.00) |
0.305 (1.00) |
|
| 21q gain | 10 (21%) | 38 |
1 (1.00) |
1 (1.00) |
0.665 (1.00) |
0.0346 (0.768) |
1 (1.00) |
0.422 (1.00) |
0.0286 (0.768) |
0.185 (1.00) |
0.258 (1.00) |
0.374 (1.00) |
| xp gain | 6 (12%) | 42 |
0.0297 (0.768) |
0.666 (1.00) |
1 (1.00) |
0.816 (1.00) |
0.484 (1.00) |
0.521 (1.00) |
0.18 (1.00) |
0.158 (1.00) |
0.0134 (0.691) |
|
| xq gain | 6 (12%) | 42 |
0.197 (1.00) |
0.666 (1.00) |
1 (1.00) |
0.815 (1.00) |
1 (1.00) |
0.385 (1.00) |
0.784 (1.00) |
0.0644 (1.00) |
0.384 (1.00) |
0.112 (1.00) |
| 1p loss | 3 (6%) | 45 |
0.554 (1.00) |
1 (1.00) |
0.579 (1.00) |
0.573 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.849 (1.00) |
1 (1.00) |
|
| 3p loss | 5 (10%) | 43 |
1 (1.00) |
1 (1.00) |
0.579 (1.00) |
1 (1.00) |
1 (1.00) |
0.855 (1.00) |
0.959 (1.00) |
1 (1.00) |
0.665 (1.00) |
|
| 3q loss | 4 (8%) | 44 |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.581 (1.00) |
0.527 (1.00) |
||
| 4p loss | 3 (6%) | 45 |
0.554 (1.00) |
1 (1.00) |
0.199 (1.00) |
0.464 (1.00) |
0.151 (1.00) |
0.689 (1.00) |
1 (1.00) |
|||
| 4q loss | 4 (8%) | 44 |
0.286 (1.00) |
0.609 (1.00) |
0.579 (1.00) |
0.571 (1.00) |
1 (1.00) |
0.466 (1.00) |
0.235 (1.00) |
0.613 (1.00) |
0.876 (1.00) |
|
| 6q loss | 7 (15%) | 41 |
1 (1.00) |
0.416 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.387 (1.00) |
0.706 (1.00) |
0.726 (1.00) |
0.0754 (1.00) |
0.516 (1.00) |
| 8p loss | 8 (17%) | 40 |
0.0446 (0.86) |
1 (1.00) |
0.00452 (0.57) |
0.0445 (0.86) |
0.311 (1.00) |
0.357 (1.00) |
0.2 (1.00) |
0.0254 (0.768) |
1 (1.00) |
0.934 (1.00) |
| 8q loss | 4 (8%) | 44 |
0.286 (1.00) |
1 (1.00) |
0.0834 (1.00) |
0.0767 (1.00) |
1 (1.00) |
0.654 (1.00) |
0.158 (1.00) |
1 (1.00) |
||
| 13q loss | 3 (6%) | 45 |
0.554 (1.00) |
1 (1.00) |
0.455 (1.00) |
1 (1.00) |
1 (1.00) |
0.851 (1.00) |
1 (1.00) |
|||
| 15q loss | 7 (15%) | 41 |
0.0114 (0.691) |
0.416 (1.00) |
0.579 (1.00) |
0.57 (1.00) |
0.639 (1.00) |
0.452 (1.00) |
0.127 (1.00) |
0.369 (1.00) |
0.197 (1.00) |
0.746 (1.00) |
| 16q loss | 4 (8%) | 44 |
0.0199 (0.691) |
1 (1.00) |
0.579 (1.00) |
0.572 (1.00) |
1 (1.00) |
0.639 (1.00) |
0.0962 (1.00) |
0.126 (1.00) |
0.239 (1.00) |
0.662 (1.00) |
| 17p loss | 9 (19%) | 39 |
0.0195 (0.691) |
1 (1.00) |
0.639 (1.00) |
1 (1.00) |
1 (1.00) |
0.76 (1.00) |
0.887 (1.00) |
0.503 (1.00) |
0.384 (1.00) |
0.268 (1.00) |
| 17q loss | 4 (8%) | 44 |
0.286 (1.00) |
0.609 (1.00) |
0.199 (1.00) |
1 (1.00) |
0.71 (1.00) |
0.492 (1.00) |
0.0844 (1.00) |
0.464 (1.00) |
||
| 18p loss | 5 (10%) | 43 |
0.372 (1.00) |
0.348 (1.00) |
0.233 (1.00) |
0.275 (1.00) |
0.484 (1.00) |
0.555 (1.00) |
0.252 (1.00) |
0.158 (1.00) |
0.219 (1.00) |
|
| 18q loss | 4 (8%) | 44 |
0.286 (1.00) |
0.609 (1.00) |
1 (1.00) |
1 (1.00) |
0.656 (1.00) |
1 (1.00) |
0.581 (1.00) |
0.524 (1.00) |
||
| 22q loss | 3 (6%) | 45 |
0.056 (1.00) |
1 (1.00) |
1 (1.00) |
0.769 (1.00) |
1 (1.00) |
1 (1.00) |
0.436 (1.00) |
1 (1.00) |
||
| xp loss | 5 (10%) | 43 |
0.372 (1.00) |
0.348 (1.00) |
0.579 (1.00) |
0.569 (1.00) |
0.583 (1.00) |
0.561 (1.00) |
0.334 (1.00) |
0.0135 (0.691) |
0.356 (1.00) |
0.281 (1.00) |
| xq loss | 4 (8%) | 44 |
0.286 (1.00) |
0.109 (1.00) |
0.579 (1.00) |
0.57 (1.00) |
0.583 (1.00) |
0.561 (1.00) |
0.143 (1.00) |
0.00215 (0.373) |
0.613 (1.00) |
0.323 (1.00) |
P value = 1.04e-05 (Fisher's exact test), Q value = 0.0027
Table S1. Gene #10: '7p gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 29 | 19 |
| 7P GAIN MUTATED | 2 | 13 |
| 7P GAIN WILD-TYPE | 27 | 6 |
Figure S1. Get High-res Image Gene #10: '7p gain' versus Molecular Subtype #1: 'CN_CNMF'

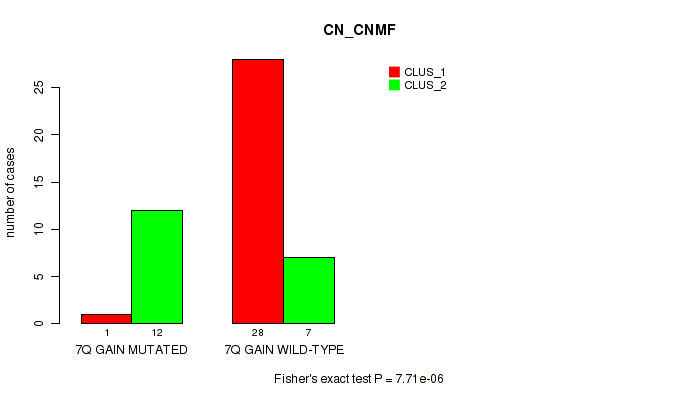
P value = 7.71e-06 (Fisher's exact test), Q value = 0.0027
Table S2. Gene #11: '7q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 |
|---|---|---|
| ALL | 29 | 19 |
| 7Q GAIN MUTATED | 1 | 12 |
| 7Q GAIN WILD-TYPE | 28 | 7 |
Figure S2. Get High-res Image Gene #11: '7q gain' versus Molecular Subtype #1: 'CN_CNMF'
-
Copy number data file = broad_values_by_arm.txt from GISTIC pipeline
-
Processed Copy number data file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_Correlate_Genomic_Events_Preprocess/DLBC-TP/19780780/transformed.cor.cli.txt
-
Molecular subtypes file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_mergedClustering/DLBC-TP/20125263/DLBC-TP.transferedmergedcluster.txt
-
Number of patients = 48
-
Number of significantly arm-level cnvs = 52
-
Number of molecular subtypes = 10
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.