This pipeline computes the correlation between cancer subtypes identified by different molecular patterns and selected clinical features.
Testing the association between subtypes identified by 8 different clustering approaches and 7 clinical features across 80 patients, 13 significant findings detected with P value < 0.05 and Q value < 0.25.
-
3 subtypes identified in current cancer cohort by 'Copy Number Ratio CNMF subtypes'. These subtypes correlate to 'Time to Death', 'YEARS_TO_BIRTH', and 'PATHOLOGY_M_STAGE'.
-
3 subtypes identified in current cancer cohort by 'METHLYATION CNMF'. These subtypes correlate to 'Time to Death'.
-
CNMF clustering analysis on sequencing-based mRNA expression data identified 3 subtypes that correlate to 'Time to Death' and 'PATHOLOGY_M_STAGE'.
-
Consensus hierarchical clustering analysis on sequencing-based mRNA expression data identified 3 subtypes that correlate to 'Time to Death' and 'PATHOLOGIC_STAGE'.
-
3 subtypes identified in current cancer cohort by 'MIRSEQ CNMF'. These subtypes correlate to 'Time to Death'.
-
4 subtypes identified in current cancer cohort by 'MIRSEQ CHIERARCHICAL'. These subtypes correlate to 'Time to Death'.
-
3 subtypes identified in current cancer cohort by 'MIRseq Mature CNMF subtypes'. These subtypes correlate to 'Time to Death' and 'PATHOLOGY_M_STAGE'.
-
6 subtypes identified in current cancer cohort by 'MIRseq Mature cHierClus subtypes'. These subtypes correlate to 'Time to Death'.
Table 1. Get Full Table Overview of the association between subtypes identified by 8 different clustering approaches and 7 clinical features. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 13 significant findings detected.
|
Clinical Features |
Time to Death |
YEARS TO BIRTH |
PATHOLOGIC STAGE |
PATHOLOGY T STAGE |
PATHOLOGY M STAGE |
GENDER |
RADIATION THERAPY |
| Statistical Tests | logrank test | Kruskal-Wallis (anova) | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test |
| Copy Number Ratio CNMF subtypes |
0.00172 (0.0161) |
0.0498 (0.212) |
0.0735 (0.248) |
0.468 (0.582) |
0.0482 (0.212) |
0.919 (0.953) |
0.342 (0.479) |
| METHLYATION CNMF |
0.000778 (0.00872) |
0.166 (0.32) |
0.165 (0.32) |
0.265 (0.391) |
0.11 (0.274) |
0.503 (0.612) |
0.789 (0.857) |
| RNAseq CNMF subtypes |
1.66e-05 (0.000466) |
0.198 (0.324) |
0.105 (0.274) |
0.131 (0.282) |
0.0376 (0.211) |
0.699 (0.799) |
0.781 (0.857) |
| RNAseq cHierClus subtypes |
1.04e-05 (0.000466) |
0.0852 (0.259) |
0.0414 (0.211) |
0.114 (0.274) |
0.0645 (0.241) |
0.878 (0.927) |
1 (1.00) |
| MIRSEQ CNMF |
0.0271 (0.168) |
0.559 (0.666) |
0.0878 (0.259) |
0.203 (0.324) |
0.181 (0.324) |
0.189 (0.324) |
0.796 (0.857) |
| MIRSEQ CHIERARCHICAL |
4.13e-05 (0.000771) |
0.125 (0.28) |
0.192 (0.324) |
0.0964 (0.27) |
0.117 (0.274) |
0.2 (0.324) |
1 (1.00) |
| MIRseq Mature CNMF subtypes |
0.00411 (0.0328) |
0.053 (0.212) |
0.138 (0.287) |
0.318 (0.457) |
0.0231 (0.162) |
0.0754 (0.248) |
0.406 (0.528) |
| MIRseq Mature cHierClus subtypes |
6.34e-05 (0.000888) |
0.42 (0.535) |
0.208 (0.324) |
0.394 (0.525) |
0.235 (0.356) |
0.381 (0.52) |
0.677 (0.79) |
Table S1. Description of clustering approach #1: 'Copy Number Ratio CNMF subtypes'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 22 | 36 | 22 |
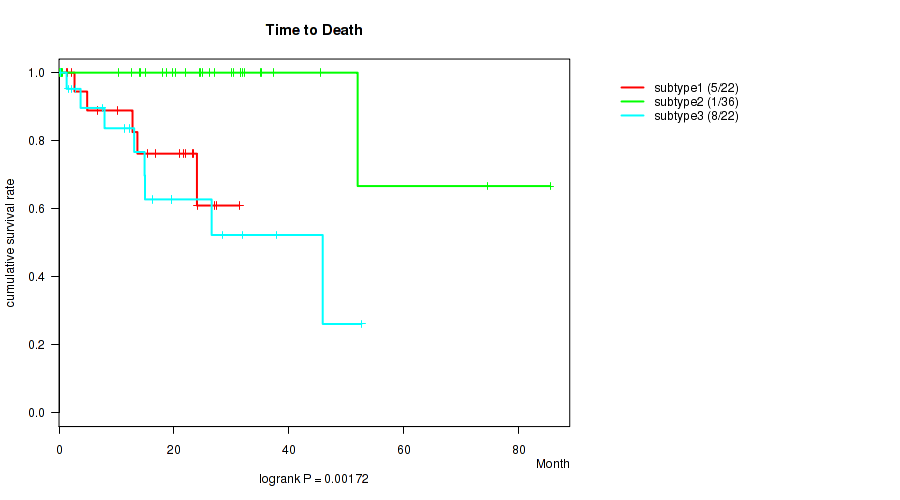
P value = 0.00172 (logrank test), Q value = 0.016
Table S2. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 80 | 14 | 0.1 - 85.5 (19.1) |
| subtype1 | 22 | 5 | 0.1 - 31.4 (16.2) |
| subtype2 | 36 | 1 | 0.1 - 85.5 (23.2) |
| subtype3 | 22 | 8 | 0.3 - 52.6 (14.0) |
Figure S1. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
P value = 0.0498 (Kruskal-Wallis (anova)), Q value = 0.21
Table S3. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 80 | 61.6 (13.9) |
| subtype1 | 22 | 65.0 (12.5) |
| subtype2 | 36 | 57.1 (14.3) |
| subtype3 | 22 | 65.7 (13.1) |
Figure S2. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #2: 'YEARS_TO_BIRTH'

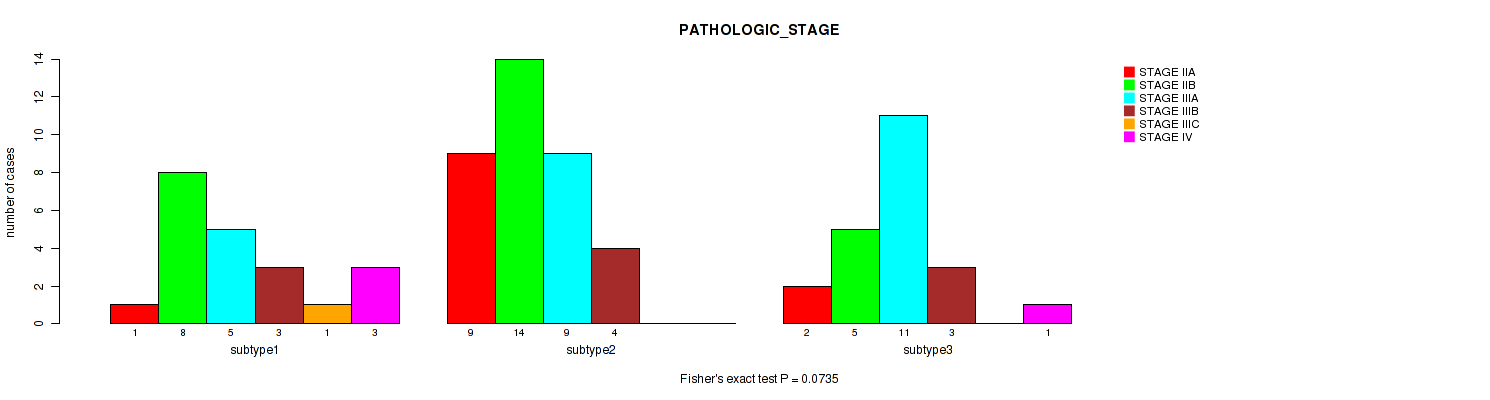
P value = 0.0735 (Fisher's exact test), Q value = 0.25
Table S4. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
| nPatients | STAGE IIA | STAGE IIB | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|
| ALL | 12 | 27 | 25 | 10 | 1 | 4 |
| subtype1 | 1 | 8 | 5 | 3 | 1 | 3 |
| subtype2 | 9 | 14 | 9 | 4 | 0 | 0 |
| subtype3 | 2 | 5 | 11 | 3 | 0 | 1 |
Figure S3. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
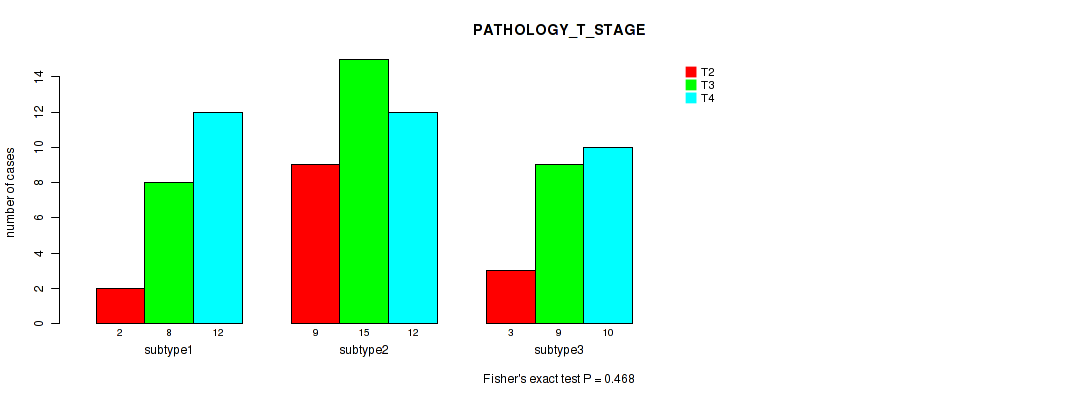
P value = 0.468 (Fisher's exact test), Q value = 0.58
Table S5. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
| nPatients | T2 | T3 | T4 |
|---|---|---|---|
| ALL | 14 | 32 | 34 |
| subtype1 | 2 | 8 | 12 |
| subtype2 | 9 | 15 | 12 |
| subtype3 | 3 | 9 | 10 |
Figure S4. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
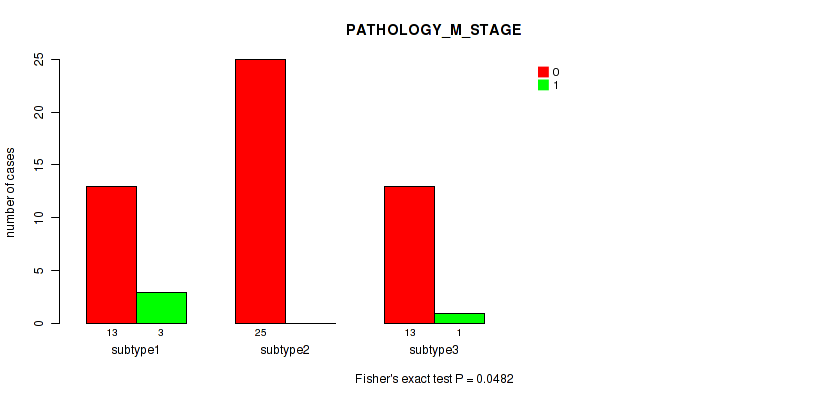
P value = 0.0482 (Fisher's exact test), Q value = 0.21
Table S6. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 51 | 4 |
| subtype1 | 13 | 3 |
| subtype2 | 25 | 0 |
| subtype3 | 13 | 1 |
Figure S5. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
P value = 0.919 (Fisher's exact test), Q value = 0.95
Table S7. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #6: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 35 | 45 |
| subtype1 | 10 | 12 |
| subtype2 | 15 | 21 |
| subtype3 | 10 | 12 |
Figure S6. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #6: 'GENDER'
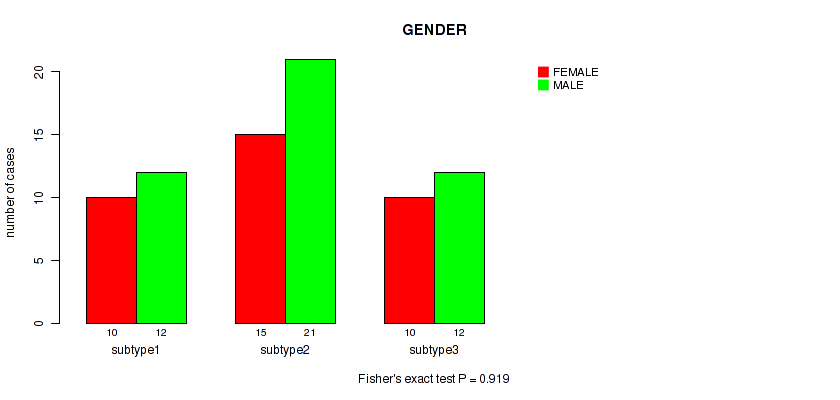
P value = 0.342 (Fisher's exact test), Q value = 0.48
Table S8. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #7: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 76 | 3 |
| subtype1 | 19 | 2 |
| subtype2 | 35 | 1 |
| subtype3 | 22 | 0 |
Figure S7. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #7: 'RADIATION_THERAPY'
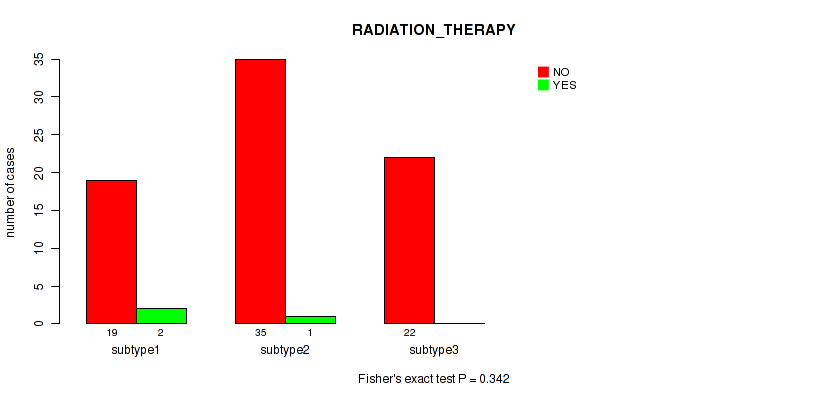
Table S9. Description of clustering approach #2: 'METHLYATION CNMF'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 33 | 18 | 29 |
P value = 0.000778 (logrank test), Q value = 0.0087
Table S10. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 80 | 14 | 0.1 - 85.5 (19.1) |
| subtype1 | 33 | 11 | 0.1 - 45.9 (15.0) |
| subtype2 | 18 | 2 | 0.2 - 74.5 (19.6) |
| subtype3 | 29 | 1 | 0.1 - 85.5 (24.4) |
Figure S8. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #1: 'Time to Death'
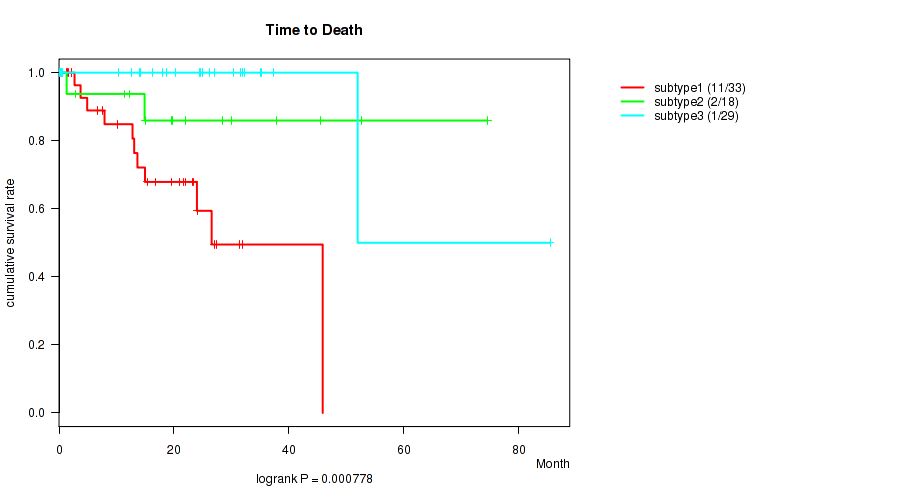
P value = 0.166 (Kruskal-Wallis (anova)), Q value = 0.32
Table S11. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 80 | 61.6 (13.9) |
| subtype1 | 33 | 65.2 (13.2) |
| subtype2 | 18 | 61.9 (11.9) |
| subtype3 | 29 | 57.4 (15.2) |
Figure S9. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
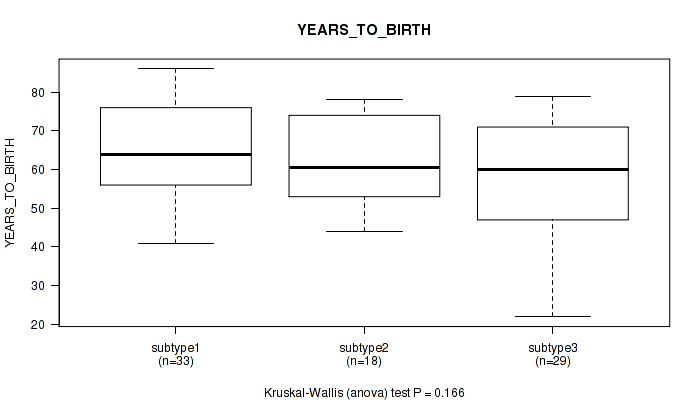
P value = 0.165 (Fisher's exact test), Q value = 0.32
Table S12. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
| nPatients | STAGE IIA | STAGE IIB | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|
| ALL | 12 | 27 | 25 | 10 | 1 | 4 |
| subtype1 | 2 | 11 | 11 | 3 | 1 | 4 |
| subtype2 | 2 | 5 | 7 | 4 | 0 | 0 |
| subtype3 | 8 | 11 | 7 | 3 | 0 | 0 |
Figure S10. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
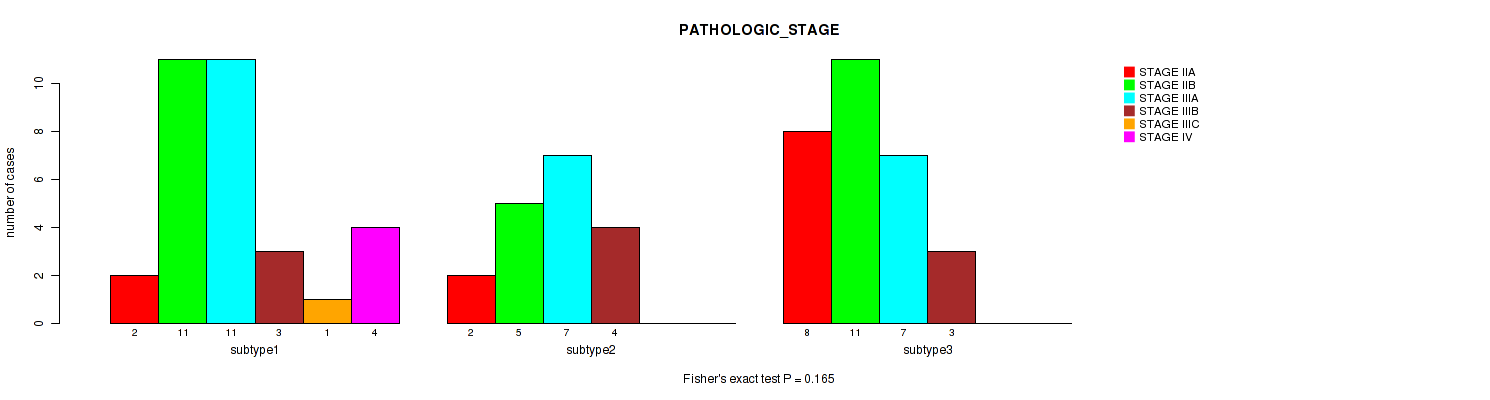
P value = 0.265 (Fisher's exact test), Q value = 0.39
Table S13. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
| nPatients | T2 | T3 | T4 |
|---|---|---|---|
| ALL | 14 | 32 | 34 |
| subtype1 | 3 | 12 | 18 |
| subtype2 | 3 | 8 | 7 |
| subtype3 | 8 | 12 | 9 |
Figure S11. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'

P value = 0.11 (Fisher's exact test), Q value = 0.27
Table S14. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 51 | 4 |
| subtype1 | 20 | 4 |
| subtype2 | 13 | 0 |
| subtype3 | 18 | 0 |
Figure S12. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
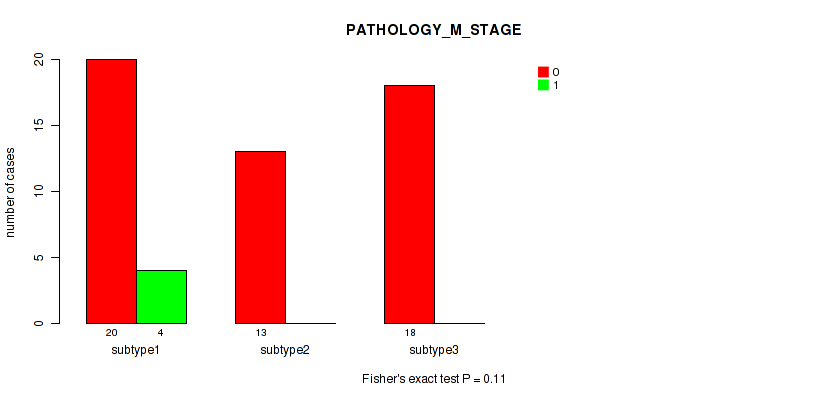
P value = 0.503 (Fisher's exact test), Q value = 0.61
Table S15. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #6: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 35 | 45 |
| subtype1 | 14 | 19 |
| subtype2 | 10 | 8 |
| subtype3 | 11 | 18 |
Figure S13. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #6: 'GENDER'
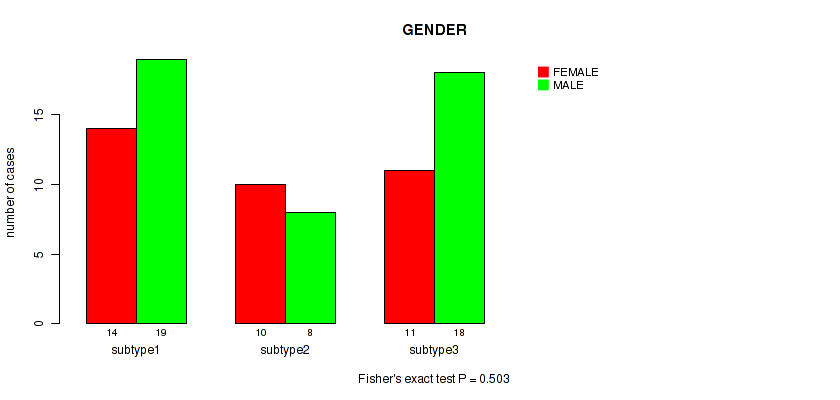
P value = 0.789 (Fisher's exact test), Q value = 0.86
Table S16. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #7: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 76 | 3 |
| subtype1 | 30 | 2 |
| subtype2 | 18 | 0 |
| subtype3 | 28 | 1 |
Figure S14. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #7: 'RADIATION_THERAPY'
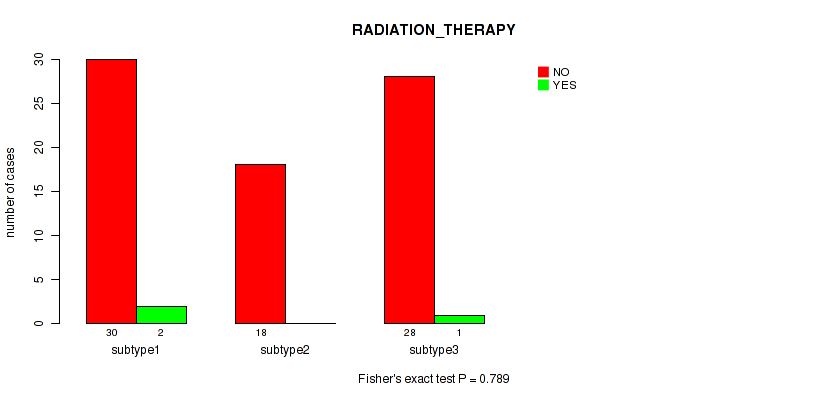
Table S17. Description of clustering approach #3: 'RNAseq CNMF subtypes'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 30 | 15 | 35 |
P value = 1.66e-05 (logrank test), Q value = 0.00047
Table S18. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 80 | 14 | 0.1 - 85.5 (19.1) |
| subtype1 | 30 | 11 | 0.1 - 37.8 (13.4) |
| subtype2 | 15 | 2 | 0.2 - 52.6 (21.0) |
| subtype3 | 35 | 1 | 0.1 - 85.5 (22.0) |
Figure S15. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
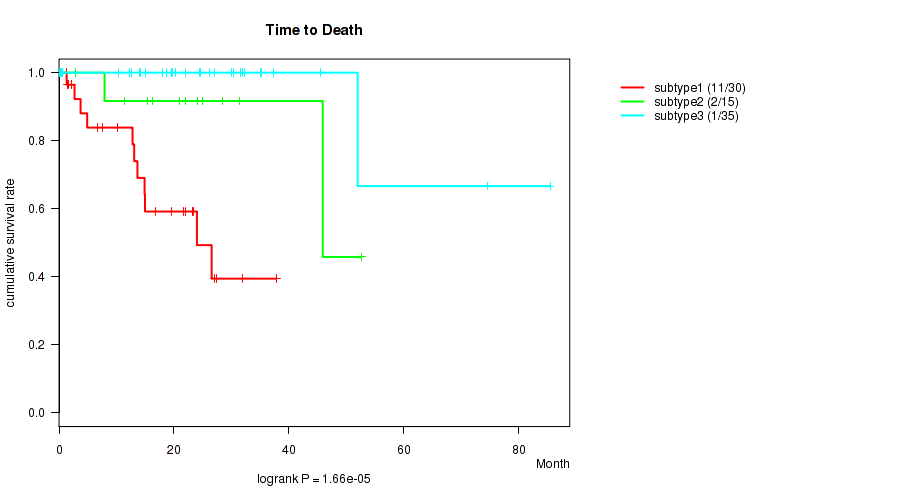
P value = 0.198 (Kruskal-Wallis (anova)), Q value = 0.32
Table S19. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 80 | 61.6 (13.9) |
| subtype1 | 30 | 65.5 (13.2) |
| subtype2 | 15 | 61.0 (13.1) |
| subtype3 | 35 | 58.6 (14.5) |
Figure S16. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
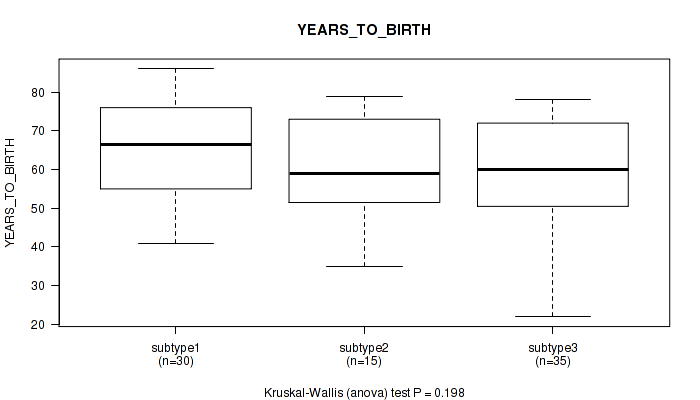
P value = 0.105 (Fisher's exact test), Q value = 0.27
Table S20. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
| nPatients | STAGE IIA | STAGE IIB | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|
| ALL | 12 | 27 | 25 | 10 | 1 | 4 |
| subtype1 | 3 | 8 | 11 | 3 | 1 | 4 |
| subtype2 | 0 | 6 | 5 | 3 | 0 | 0 |
| subtype3 | 9 | 13 | 9 | 4 | 0 | 0 |
Figure S17. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
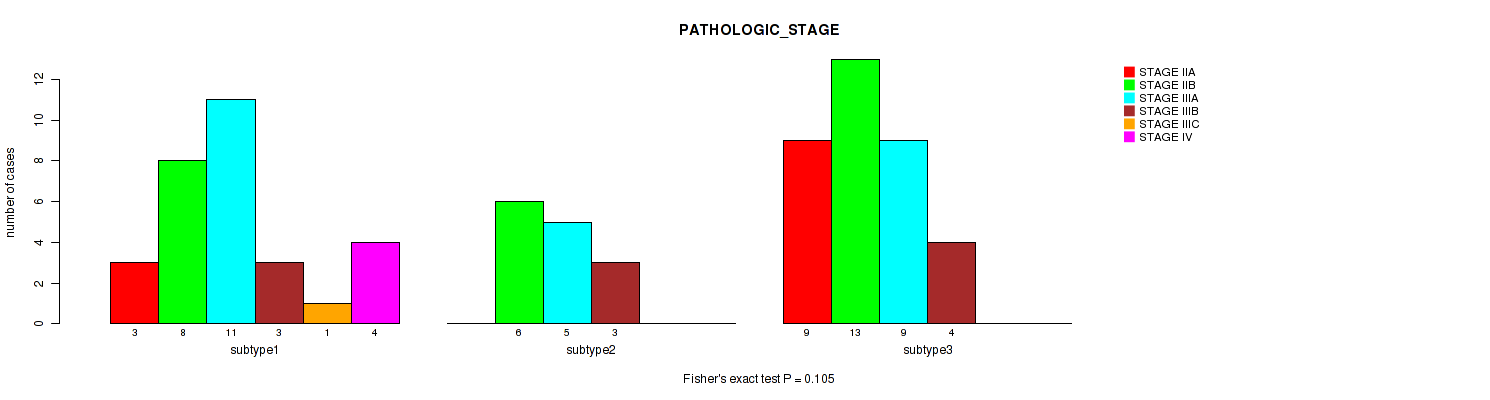
P value = 0.131 (Fisher's exact test), Q value = 0.28
Table S21. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
| nPatients | T2 | T3 | T4 |
|---|---|---|---|
| ALL | 14 | 32 | 34 |
| subtype1 | 3 | 9 | 18 |
| subtype2 | 2 | 8 | 5 |
| subtype3 | 9 | 15 | 11 |
Figure S18. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
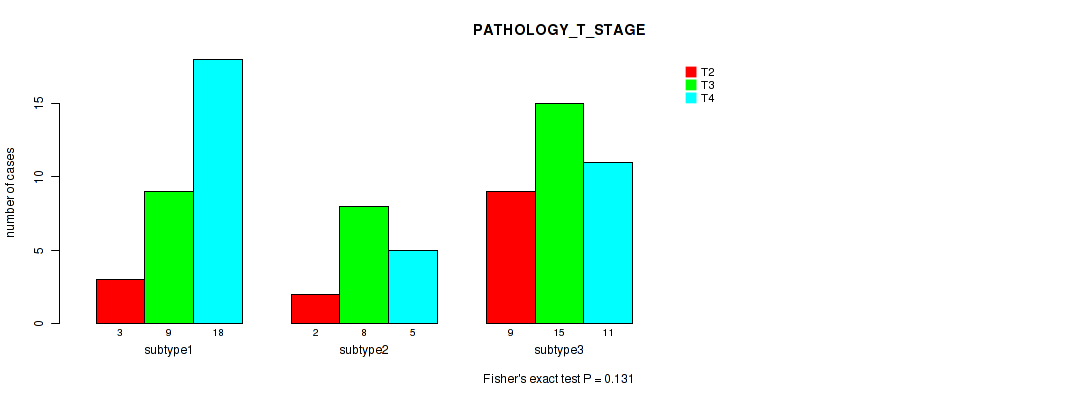
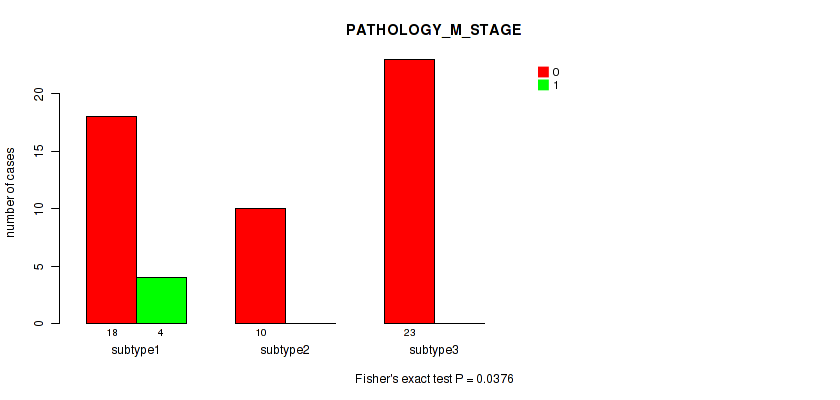
P value = 0.0376 (Fisher's exact test), Q value = 0.21
Table S22. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 51 | 4 |
| subtype1 | 18 | 4 |
| subtype2 | 10 | 0 |
| subtype3 | 23 | 0 |
Figure S19. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
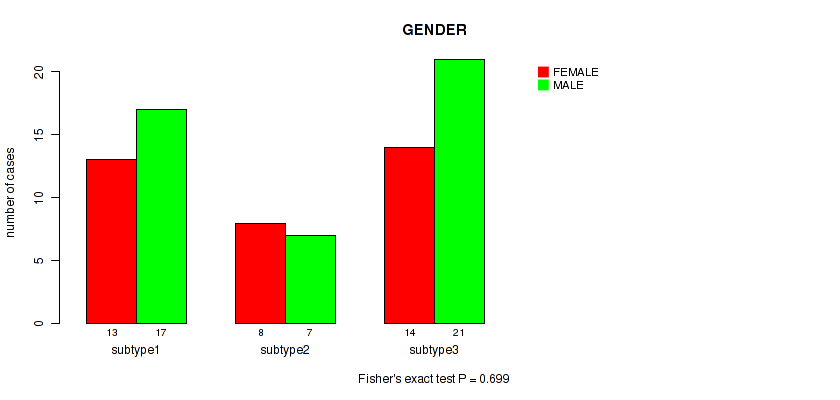
P value = 0.699 (Fisher's exact test), Q value = 0.8
Table S23. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #6: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 35 | 45 |
| subtype1 | 13 | 17 |
| subtype2 | 8 | 7 |
| subtype3 | 14 | 21 |
Figure S20. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #6: 'GENDER'
P value = 0.781 (Fisher's exact test), Q value = 0.86
Table S24. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #7: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 76 | 3 |
| subtype1 | 28 | 1 |
| subtype2 | 14 | 1 |
| subtype3 | 34 | 1 |
Figure S21. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #7: 'RADIATION_THERAPY'
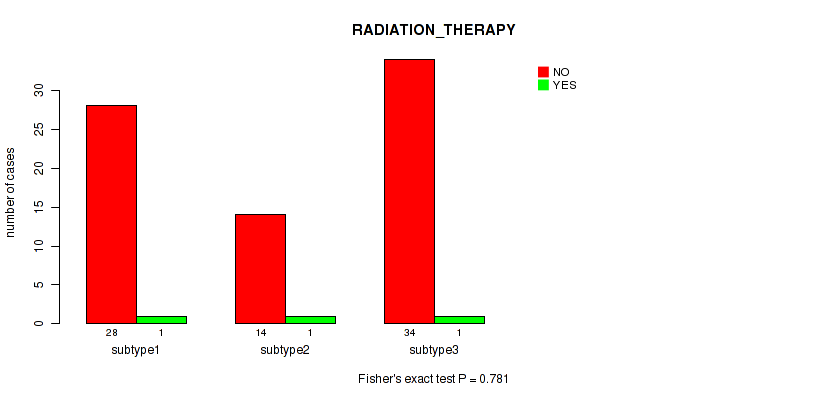
Table S25. Description of clustering approach #4: 'RNAseq cHierClus subtypes'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 29 | 33 | 18 |
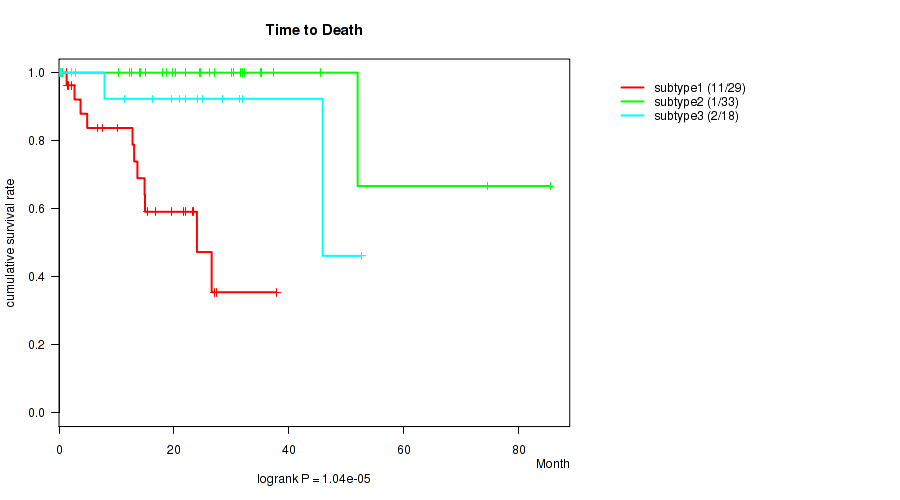
P value = 1.04e-05 (logrank test), Q value = 0.00047
Table S26. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 80 | 14 | 0.1 - 85.5 (19.1) |
| subtype1 | 29 | 11 | 0.1 - 37.8 (13.6) |
| subtype2 | 33 | 1 | 0.1 - 85.5 (24.4) |
| subtype3 | 18 | 2 | 0.2 - 52.6 (20.3) |
Figure S22. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
P value = 0.0852 (Kruskal-Wallis (anova)), Q value = 0.26
Table S27. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 80 | 61.6 (13.9) |
| subtype1 | 29 | 66.4 (12.5) |
| subtype2 | 33 | 57.8 (14.5) |
| subtype3 | 18 | 60.9 (13.4) |
Figure S23. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
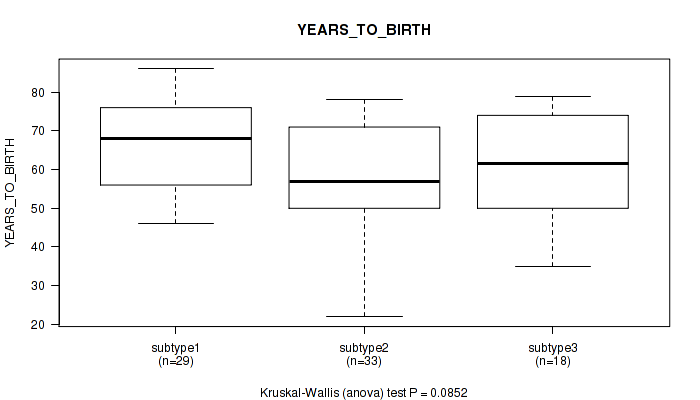
P value = 0.0414 (Fisher's exact test), Q value = 0.21
Table S28. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
| nPatients | STAGE IIA | STAGE IIB | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|
| ALL | 12 | 27 | 25 | 10 | 1 | 4 |
| subtype1 | 3 | 9 | 9 | 3 | 1 | 4 |
| subtype2 | 9 | 13 | 8 | 3 | 0 | 0 |
| subtype3 | 0 | 5 | 8 | 4 | 0 | 0 |
Figure S24. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
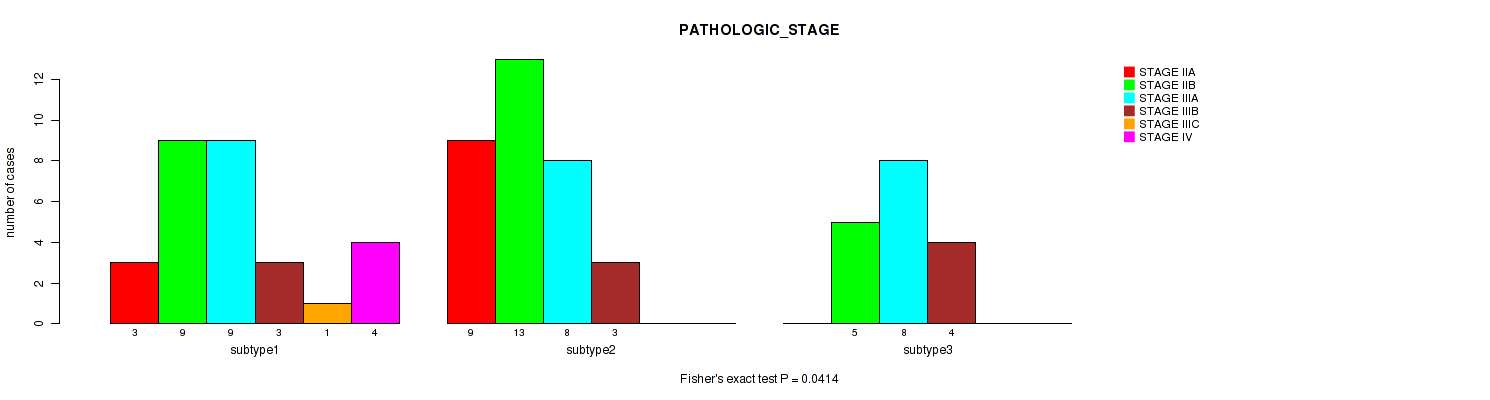
P value = 0.114 (Fisher's exact test), Q value = 0.27
Table S29. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
| nPatients | T2 | T3 | T4 |
|---|---|---|---|
| ALL | 14 | 32 | 34 |
| subtype1 | 4 | 9 | 16 |
| subtype2 | 9 | 15 | 9 |
| subtype3 | 1 | 8 | 9 |
Figure S25. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
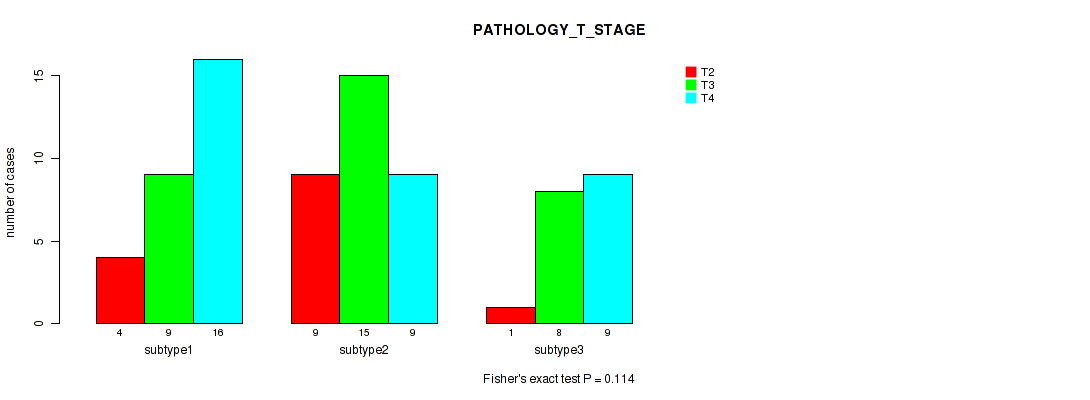
P value = 0.0645 (Fisher's exact test), Q value = 0.24
Table S30. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 51 | 4 |
| subtype1 | 18 | 4 |
| subtype2 | 22 | 0 |
| subtype3 | 11 | 0 |
Figure S26. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
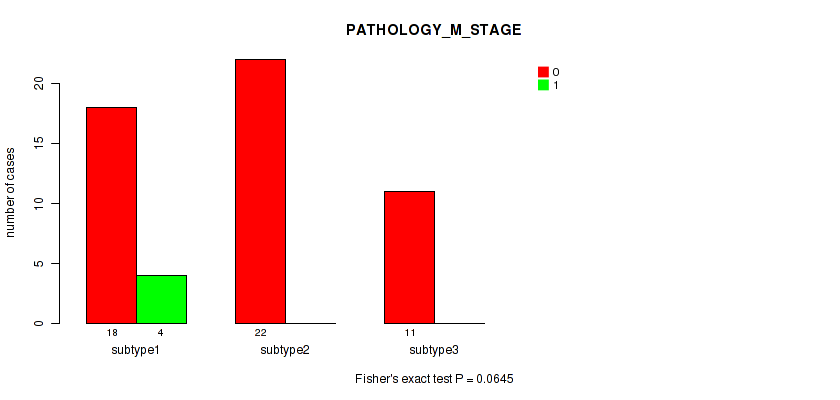
P value = 0.878 (Fisher's exact test), Q value = 0.93
Table S31. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #6: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 35 | 45 |
| subtype1 | 12 | 17 |
| subtype2 | 14 | 19 |
| subtype3 | 9 | 9 |
Figure S27. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #6: 'GENDER'
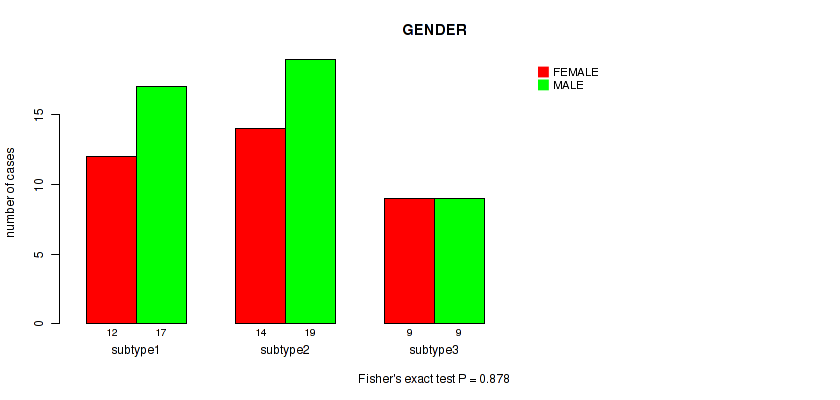
P value = 1 (Fisher's exact test), Q value = 1
Table S32. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #7: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 76 | 3 |
| subtype1 | 27 | 1 |
| subtype2 | 32 | 1 |
| subtype3 | 17 | 1 |
Figure S28. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #7: 'RADIATION_THERAPY'
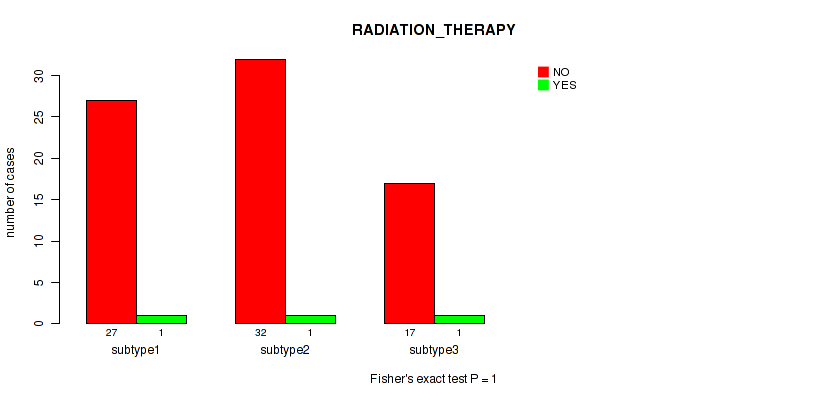
Table S33. Description of clustering approach #5: 'MIRSEQ CNMF'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 34 | 17 | 29 |
P value = 0.0271 (logrank test), Q value = 0.17
Table S34. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 80 | 14 | 0.1 - 85.5 (19.1) |
| subtype1 | 34 | 8 | 0.1 - 52.6 (17.5) |
| subtype2 | 17 | 5 | 0.1 - 85.5 (15.0) |
| subtype3 | 29 | 1 | 0.2 - 74.5 (20.2) |
Figure S29. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #1: 'Time to Death'
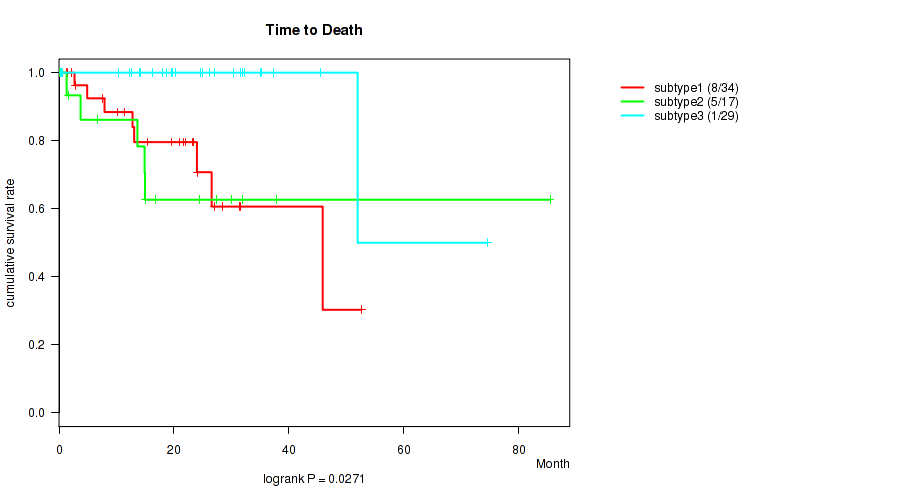
P value = 0.559 (Kruskal-Wallis (anova)), Q value = 0.67
Table S35. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 80 | 61.6 (13.9) |
| subtype1 | 34 | 63.2 (12.4) |
| subtype2 | 17 | 63.6 (14.2) |
| subtype3 | 29 | 58.7 (15.5) |
Figure S30. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
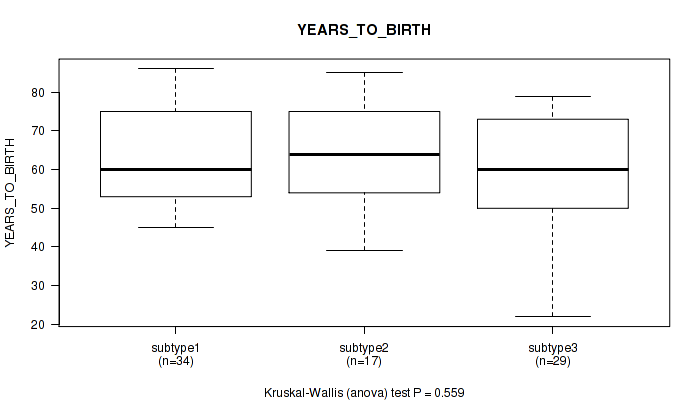
P value = 0.0878 (Fisher's exact test), Q value = 0.26
Table S36. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
| nPatients | STAGE IIA | STAGE IIB | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|
| ALL | 12 | 27 | 25 | 10 | 1 | 4 |
| subtype1 | 1 | 14 | 10 | 5 | 1 | 2 |
| subtype2 | 2 | 5 | 7 | 1 | 0 | 2 |
| subtype3 | 9 | 8 | 8 | 4 | 0 | 0 |
Figure S31. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
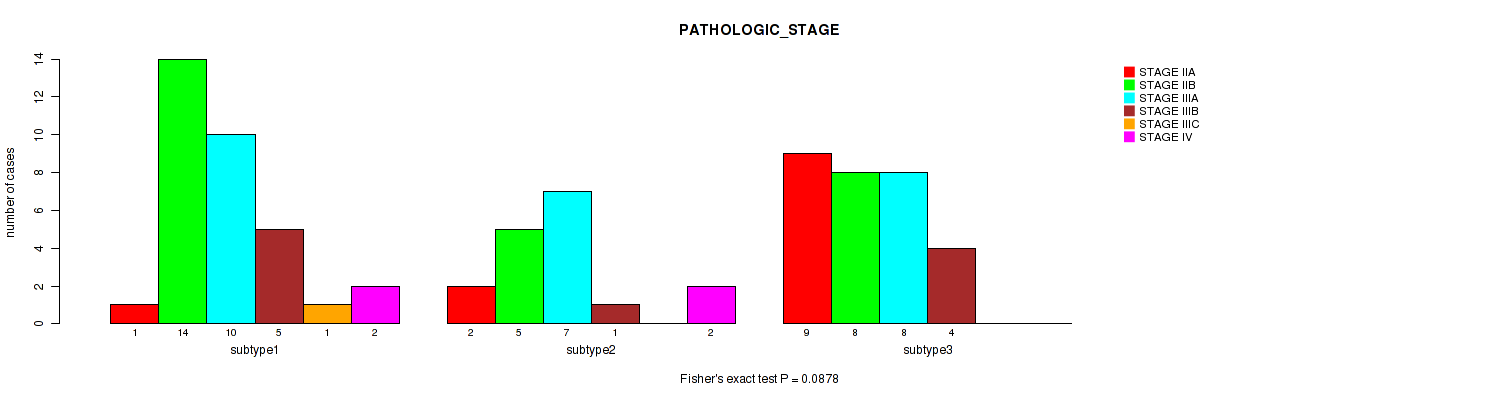
P value = 0.203 (Fisher's exact test), Q value = 0.32
Table S37. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
| nPatients | T2 | T3 | T4 |
|---|---|---|---|
| ALL | 14 | 32 | 34 |
| subtype1 | 3 | 16 | 15 |
| subtype2 | 2 | 6 | 9 |
| subtype3 | 9 | 10 | 10 |
Figure S32. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
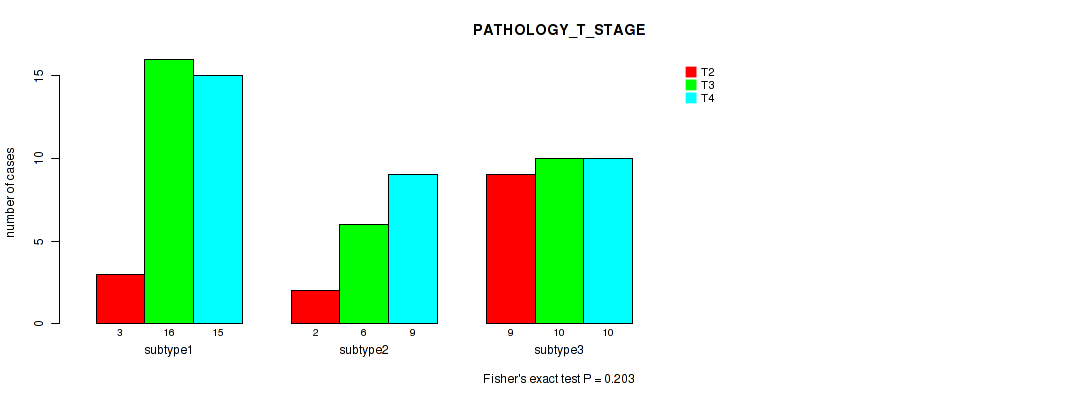
P value = 0.181 (Fisher's exact test), Q value = 0.32
Table S38. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 51 | 4 |
| subtype1 | 22 | 2 |
| subtype2 | 9 | 2 |
| subtype3 | 20 | 0 |
Figure S33. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'

P value = 0.189 (Fisher's exact test), Q value = 0.32
Table S39. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #6: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 35 | 45 |
| subtype1 | 11 | 23 |
| subtype2 | 10 | 7 |
| subtype3 | 14 | 15 |
Figure S34. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #6: 'GENDER'
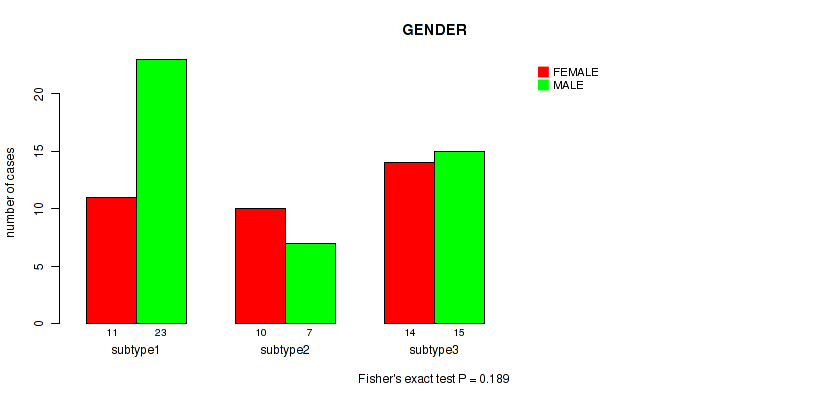
P value = 0.796 (Fisher's exact test), Q value = 0.86
Table S40. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #7: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 76 | 3 |
| subtype1 | 31 | 2 |
| subtype2 | 17 | 0 |
| subtype3 | 28 | 1 |
Figure S35. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #7: 'RADIATION_THERAPY'
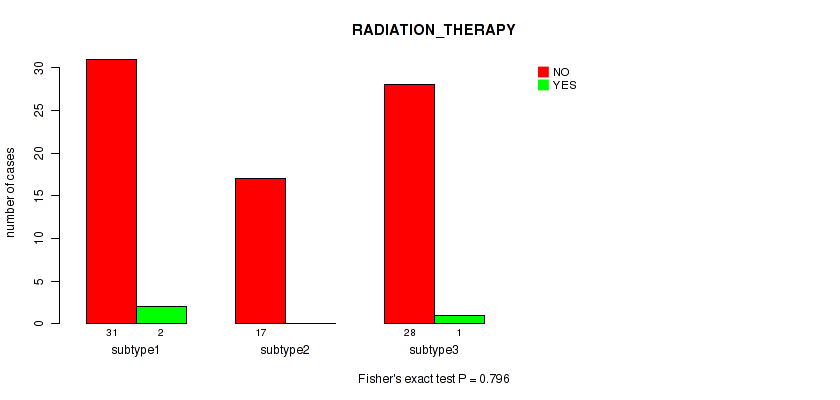
Table S41. Description of clustering approach #6: 'MIRSEQ CHIERARCHICAL'
| Cluster Labels | 1 | 2 | 3 | 4 |
|---|---|---|---|---|
| Number of samples | 17 | 13 | 28 | 22 |
P value = 4.13e-05 (logrank test), Q value = 0.00077
Table S42. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 80 | 14 | 0.1 - 85.5 (19.1) |
| subtype1 | 17 | 3 | 1.4 - 52.6 (21.7) |
| subtype2 | 13 | 0 | 0.1 - 85.5 (22.0) |
| subtype3 | 28 | 2 | 0.2 - 74.5 (20.0) |
| subtype4 | 22 | 9 | 0.1 - 37.8 (12.9) |
Figure S36. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #1: 'Time to Death'

P value = 0.125 (Kruskal-Wallis (anova)), Q value = 0.28
Table S43. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 80 | 61.6 (13.9) |
| subtype1 | 17 | 60.9 (14.9) |
| subtype2 | 13 | 59.2 (12.2) |
| subtype3 | 28 | 58.4 (15.7) |
| subtype4 | 22 | 67.8 (10.2) |
Figure S37. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
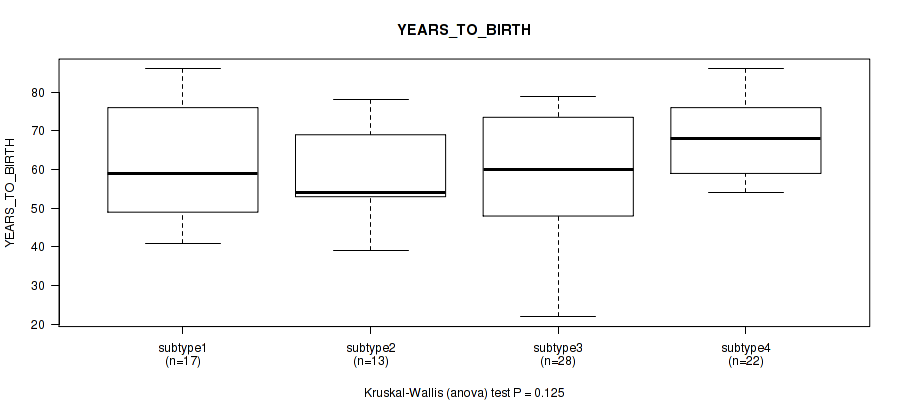
P value = 0.192 (Fisher's exact test), Q value = 0.32
Table S44. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
| nPatients | STAGE IIA | STAGE IIB | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|
| ALL | 12 | 27 | 25 | 10 | 1 | 4 |
| subtype1 | 1 | 7 | 5 | 1 | 1 | 1 |
| subtype2 | 0 | 6 | 5 | 2 | 0 | 0 |
| subtype3 | 9 | 8 | 7 | 4 | 0 | 0 |
| subtype4 | 2 | 6 | 8 | 3 | 0 | 3 |
Figure S38. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
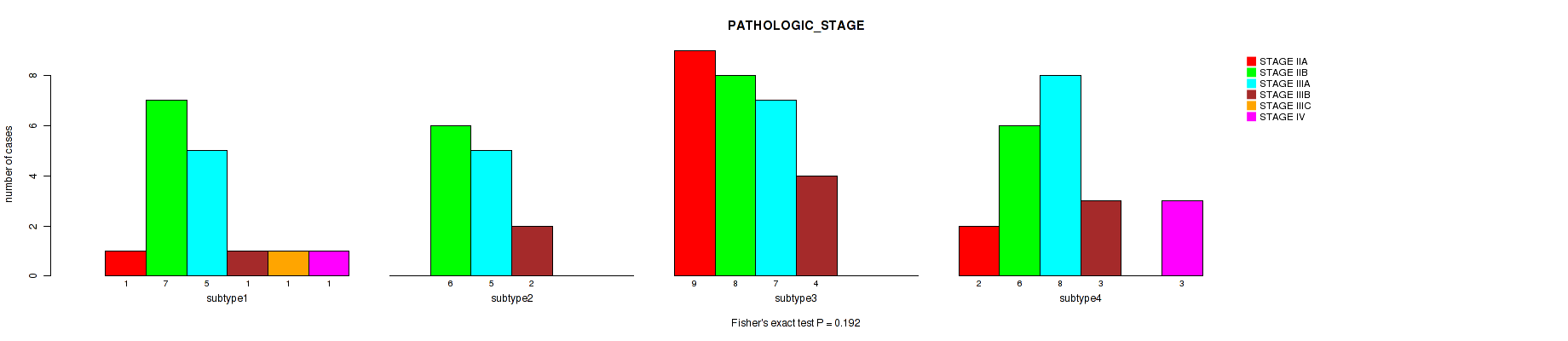
P value = 0.0964 (Fisher's exact test), Q value = 0.27
Table S45. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
| nPatients | T2 | T3 | T4 |
|---|---|---|---|
| ALL | 14 | 32 | 34 |
| subtype1 | 3 | 9 | 5 |
| subtype2 | 0 | 6 | 7 |
| subtype3 | 9 | 10 | 9 |
| subtype4 | 2 | 7 | 13 |
Figure S39. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
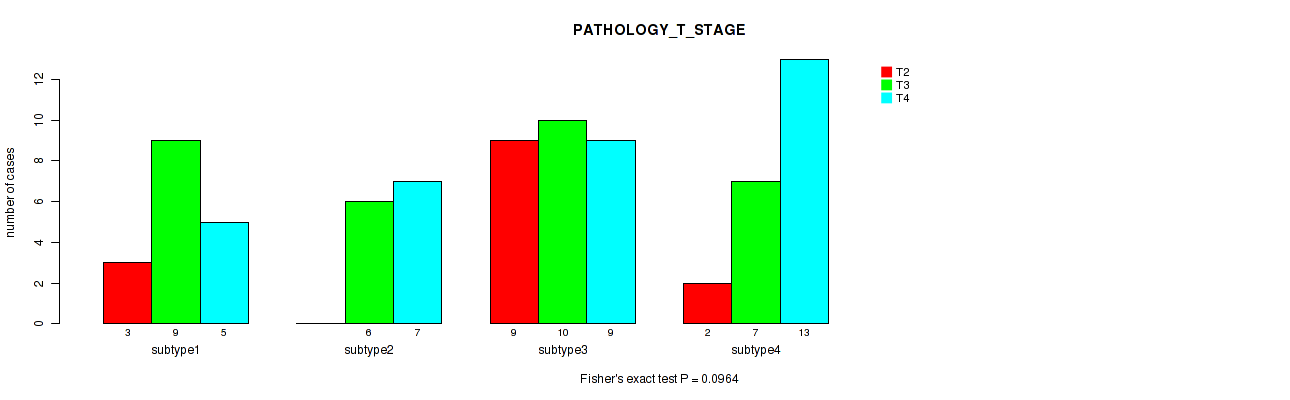
P value = 0.117 (Fisher's exact test), Q value = 0.27
Table S46. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 51 | 4 |
| subtype1 | 10 | 1 |
| subtype2 | 9 | 0 |
| subtype3 | 19 | 0 |
| subtype4 | 13 | 3 |
Figure S40. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
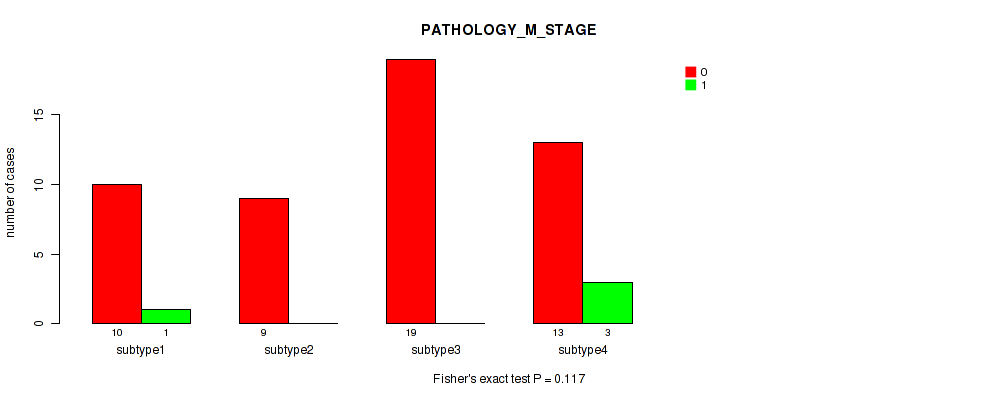
P value = 0.2 (Fisher's exact test), Q value = 0.32
Table S47. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #6: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 35 | 45 |
| subtype1 | 10 | 7 |
| subtype2 | 3 | 10 |
| subtype3 | 14 | 14 |
| subtype4 | 8 | 14 |
Figure S41. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #6: 'GENDER'
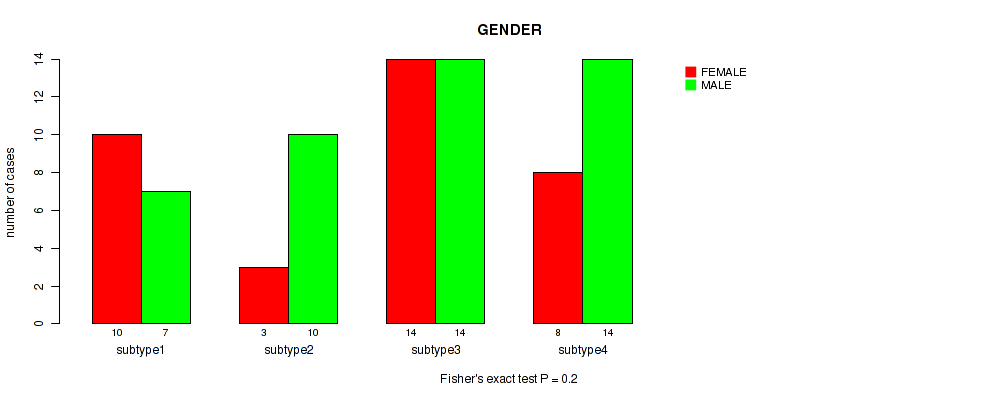
P value = 1 (Fisher's exact test), Q value = 1
Table S48. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #7: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 76 | 3 |
| subtype1 | 15 | 1 |
| subtype2 | 13 | 0 |
| subtype3 | 27 | 1 |
| subtype4 | 21 | 1 |
Figure S42. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #7: 'RADIATION_THERAPY'
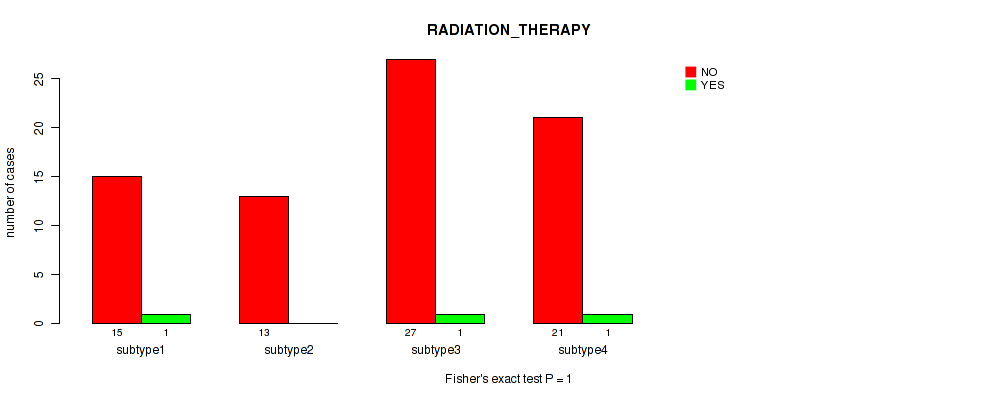
Table S49. Description of clustering approach #7: 'MIRseq Mature CNMF subtypes'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 32 | 28 | 14 |
P value = 0.00411 (logrank test), Q value = 0.033
Table S50. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 74 | 14 | 0.1 - 85.5 (19.1) |
| subtype1 | 32 | 9 | 0.1 - 52.6 (14.5) |
| subtype2 | 28 | 1 | 0.1 - 85.5 (23.2) |
| subtype3 | 14 | 4 | 0.5 - 37.8 (13.2) |
Figure S43. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #1: 'Time to Death'

P value = 0.053 (Kruskal-Wallis (anova)), Q value = 0.21
Table S51. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 74 | 61.9 (14.1) |
| subtype1 | 32 | 63.2 (13.6) |
| subtype2 | 28 | 57.0 (15.4) |
| subtype3 | 14 | 68.9 (9.2) |
Figure S44. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
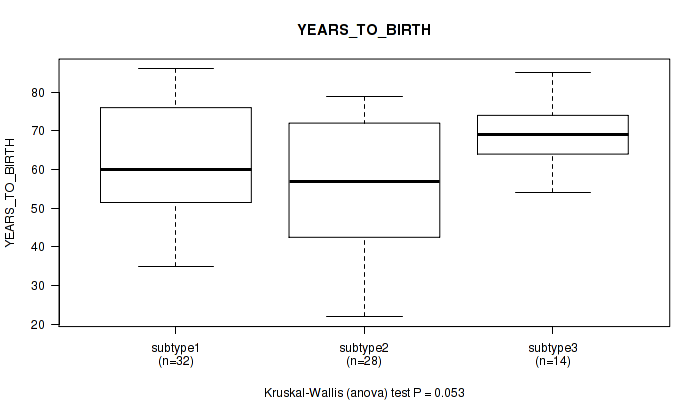
P value = 0.138 (Fisher's exact test), Q value = 0.29
Table S52. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
| nPatients | STAGE IIA | STAGE IIB | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|
| ALL | 10 | 25 | 24 | 9 | 1 | 4 |
| subtype1 | 1 | 12 | 11 | 4 | 1 | 2 |
| subtype2 | 4 | 10 | 10 | 4 | 0 | 0 |
| subtype3 | 5 | 3 | 3 | 1 | 0 | 2 |
Figure S45. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
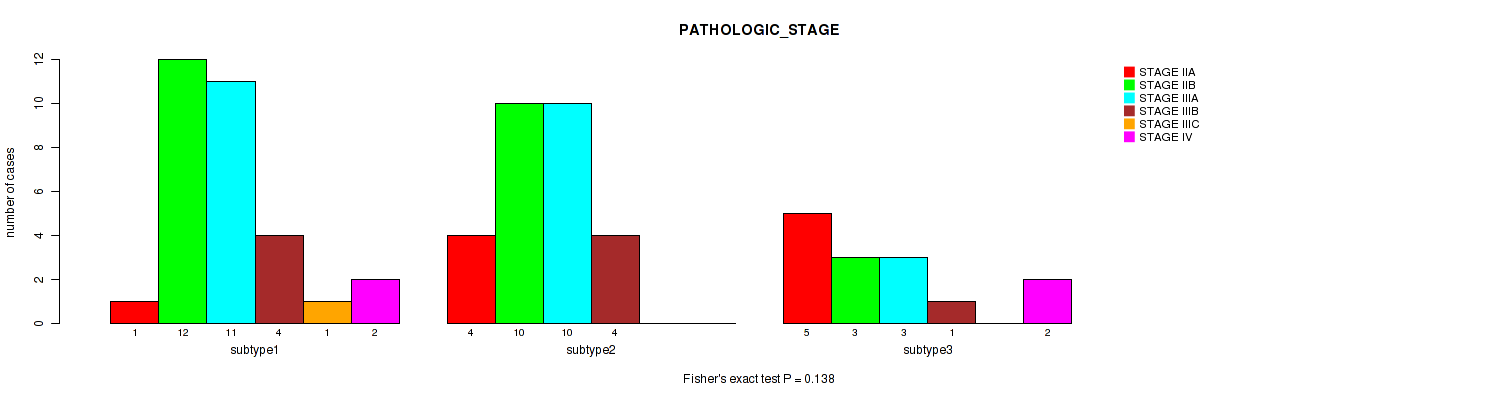
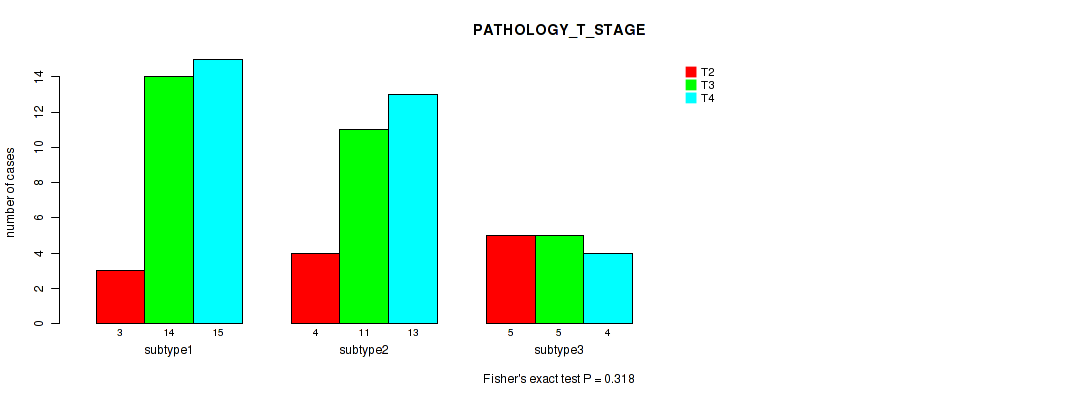
P value = 0.318 (Fisher's exact test), Q value = 0.46
Table S53. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
| nPatients | T2 | T3 | T4 |
|---|---|---|---|
| ALL | 12 | 30 | 32 |
| subtype1 | 3 | 14 | 15 |
| subtype2 | 4 | 11 | 13 |
| subtype3 | 5 | 5 | 4 |
Figure S46. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
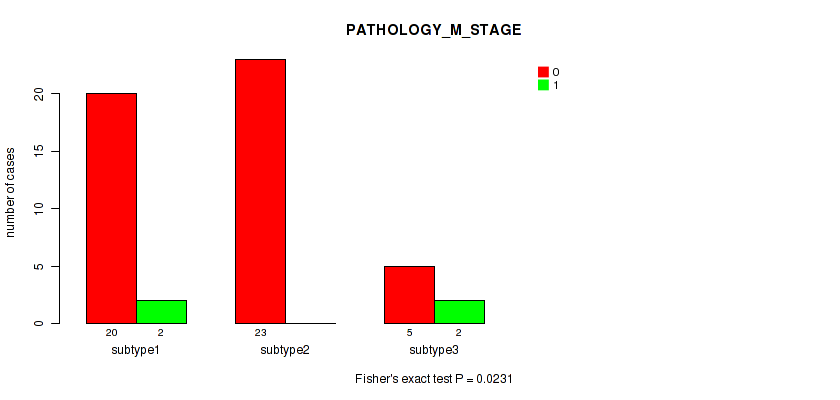
P value = 0.0231 (Fisher's exact test), Q value = 0.16
Table S54. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 48 | 4 |
| subtype1 | 20 | 2 |
| subtype2 | 23 | 0 |
| subtype3 | 5 | 2 |
Figure S47. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
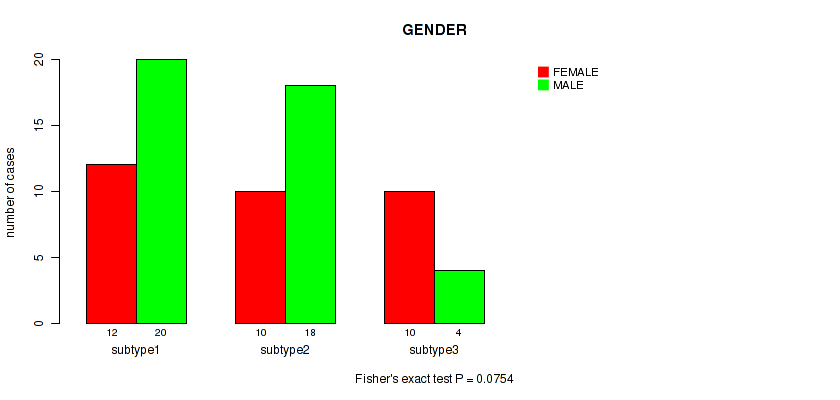
P value = 0.0754 (Fisher's exact test), Q value = 0.25
Table S55. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #6: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 32 | 42 |
| subtype1 | 12 | 20 |
| subtype2 | 10 | 18 |
| subtype3 | 10 | 4 |
Figure S48. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #6: 'GENDER'
P value = 0.406 (Fisher's exact test), Q value = 0.53
Table S56. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #7: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 70 | 3 |
| subtype1 | 29 | 2 |
| subtype2 | 28 | 0 |
| subtype3 | 13 | 1 |
Figure S49. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #7: 'RADIATION_THERAPY'
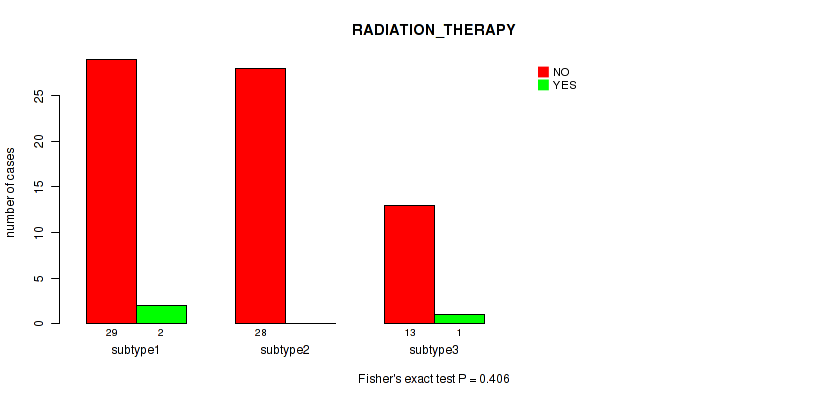
Table S57. Description of clustering approach #8: 'MIRseq Mature cHierClus subtypes'
| Cluster Labels | 1 | 2 | 3 | 4 | 5 | 6 |
|---|---|---|---|---|---|---|
| Number of samples | 5 | 12 | 16 | 16 | 15 | 10 |
P value = 6.34e-05 (logrank test), Q value = 0.00089
Table S58. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 74 | 14 | 0.1 - 85.5 (19.1) |
| subtype1 | 5 | 2 | 1.4 - 23.3 (7.9) |
| subtype2 | 12 | 1 | 0.3 - 52.6 (20.3) |
| subtype3 | 16 | 0 | 0.1 - 85.5 (23.2) |
| subtype4 | 16 | 1 | 0.2 - 74.5 (23.6) |
| subtype5 | 15 | 6 | 0.1 - 37.8 (13.1) |
| subtype6 | 10 | 4 | 1.2 - 28.4 (18.1) |
Figure S50. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
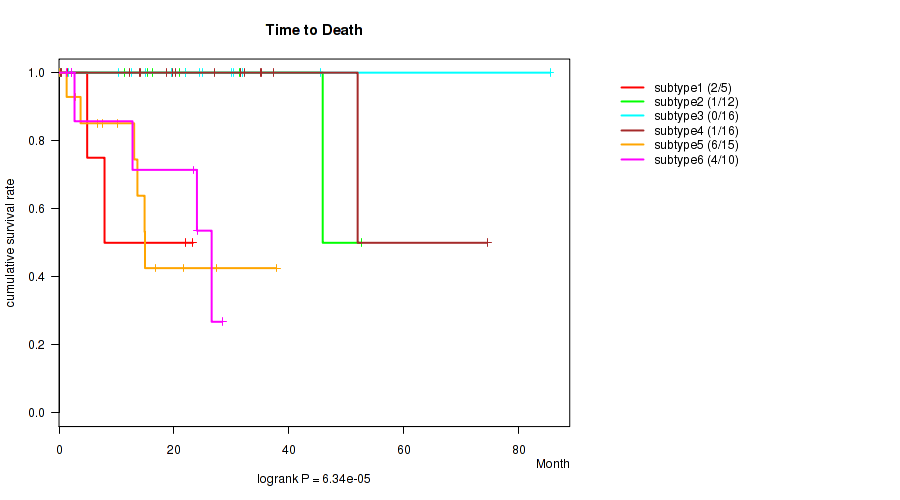
P value = 0.42 (Kruskal-Wallis (anova)), Q value = 0.54
Table S59. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 74 | 61.9 (14.1) |
| subtype1 | 5 | 68.2 (18.8) |
| subtype2 | 12 | 60.3 (13.3) |
| subtype3 | 16 | 58.2 (13.7) |
| subtype4 | 16 | 60.3 (17.4) |
| subtype5 | 15 | 67.9 (11.3) |
| subtype6 | 10 | 60.3 (10.6) |
Figure S51. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
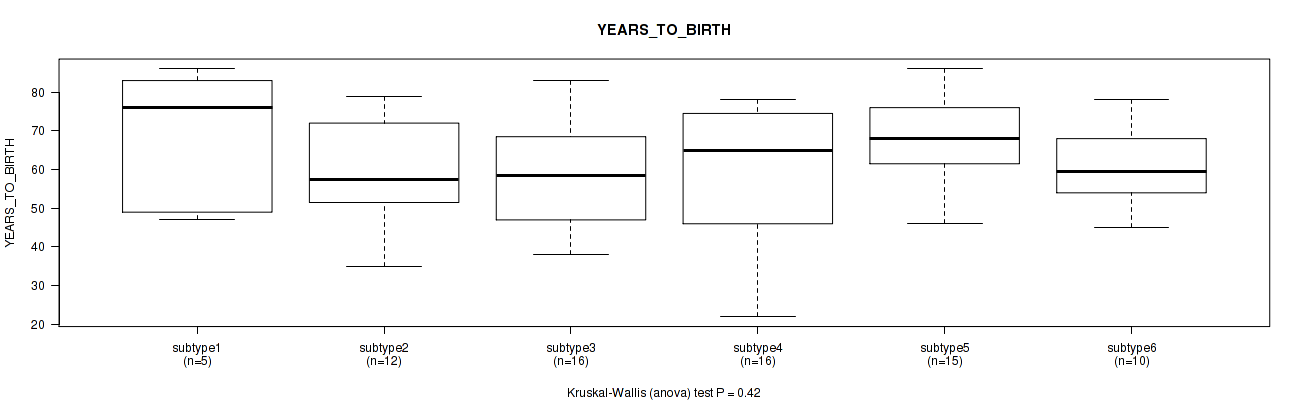
P value = 0.208 (Fisher's exact test), Q value = 0.32
Table S60. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
| nPatients | STAGE IIA | STAGE IIB | STAGE IIIA | STAGE IIIB | STAGE IIIC | STAGE IV |
|---|---|---|---|---|---|---|
| ALL | 10 | 25 | 24 | 9 | 1 | 4 |
| subtype1 | 0 | 1 | 2 | 0 | 1 | 1 |
| subtype2 | 0 | 5 | 3 | 3 | 0 | 0 |
| subtype3 | 3 | 5 | 7 | 1 | 0 | 0 |
| subtype4 | 5 | 6 | 3 | 2 | 0 | 0 |
| subtype5 | 1 | 7 | 4 | 1 | 0 | 2 |
| subtype6 | 1 | 1 | 5 | 2 | 0 | 1 |
Figure S52. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #3: 'PATHOLOGIC_STAGE'
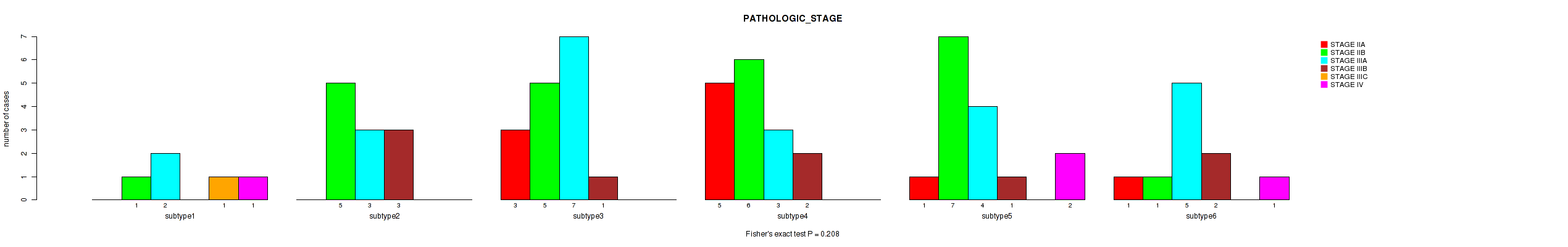
P value = 0.394 (Fisher's exact test), Q value = 0.53
Table S61. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
| nPatients | T2 | T3 | T4 |
|---|---|---|---|
| ALL | 12 | 30 | 32 |
| subtype1 | 0 | 2 | 3 |
| subtype2 | 2 | 5 | 5 |
| subtype3 | 3 | 5 | 8 |
| subtype4 | 5 | 8 | 3 |
| subtype5 | 1 | 8 | 6 |
| subtype6 | 1 | 2 | 7 |
Figure S53. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #4: 'PATHOLOGY_T_STAGE'
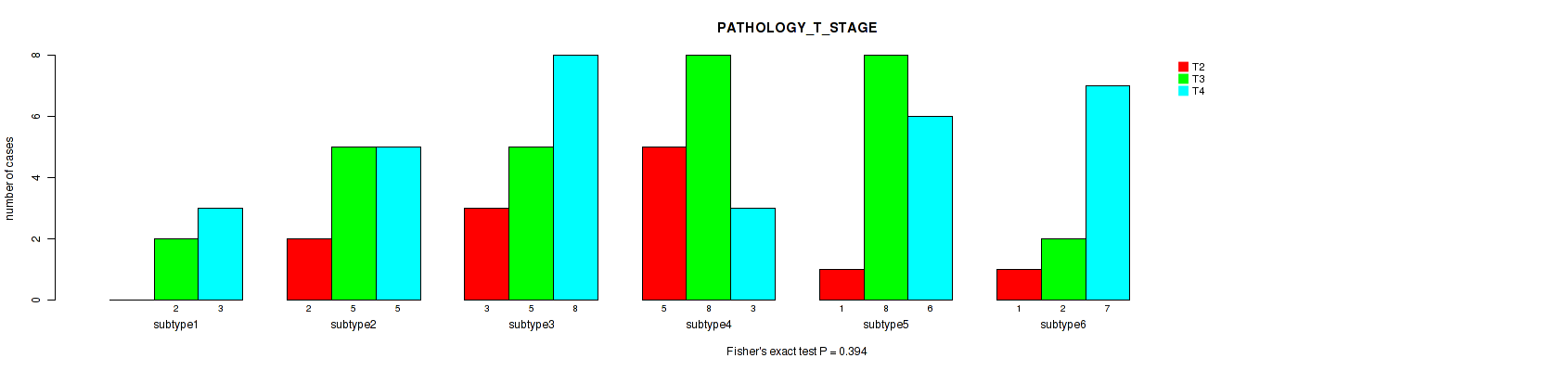
P value = 0.235 (Fisher's exact test), Q value = 0.36
Table S62. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 48 | 4 |
| subtype1 | 4 | 1 |
| subtype2 | 5 | 0 |
| subtype3 | 13 | 0 |
| subtype4 | 11 | 0 |
| subtype5 | 9 | 2 |
| subtype6 | 6 | 1 |
Figure S54. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #5: 'PATHOLOGY_M_STAGE'
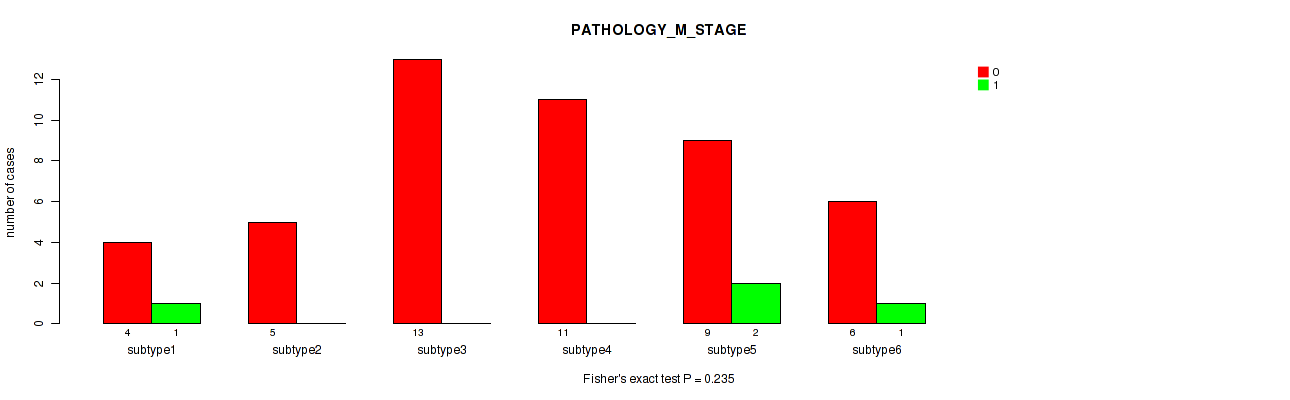
P value = 0.381 (Fisher's exact test), Q value = 0.52
Table S63. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #6: 'GENDER'
| nPatients | FEMALE | MALE |
|---|---|---|
| ALL | 32 | 42 |
| subtype1 | 4 | 1 |
| subtype2 | 5 | 7 |
| subtype3 | 6 | 10 |
| subtype4 | 8 | 8 |
| subtype5 | 7 | 8 |
| subtype6 | 2 | 8 |
Figure S55. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #6: 'GENDER'
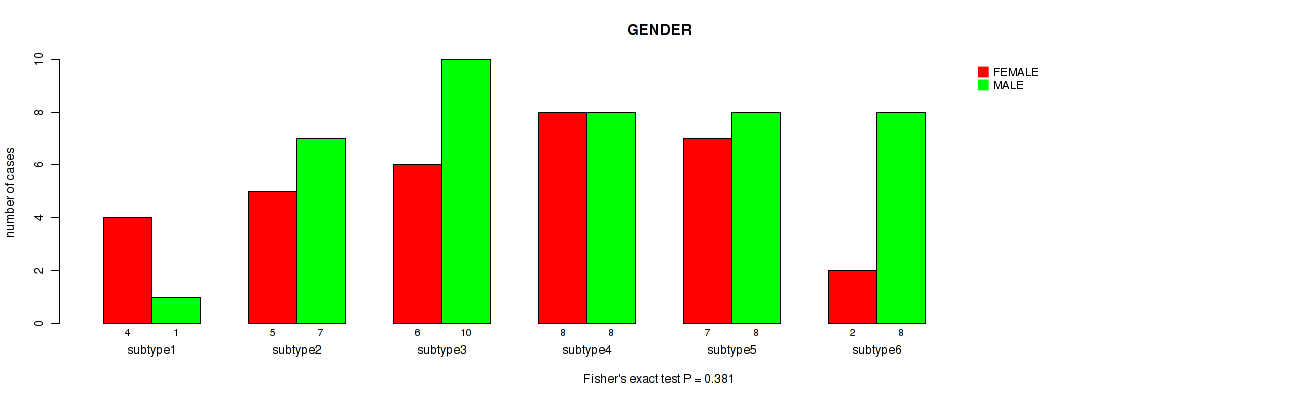
P value = 0.677 (Fisher's exact test), Q value = 0.79
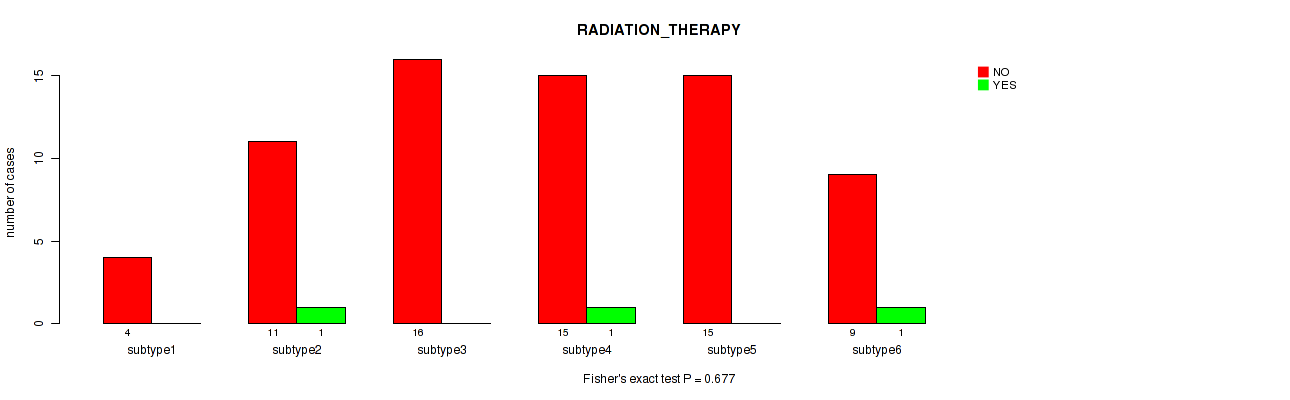
Table S64. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #7: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 70 | 3 |
| subtype1 | 4 | 0 |
| subtype2 | 11 | 1 |
| subtype3 | 16 | 0 |
| subtype4 | 15 | 1 |
| subtype5 | 15 | 0 |
| subtype6 | 9 | 1 |
Figure S56. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #7: 'RADIATION_THERAPY'
-
Cluster data file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_mergedClustering/UVM-TP/19785804/UVM-TP.mergedcluster.txt
-
Clinical data file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/Append_Data/UVM-TP/19775655/UVM-TP.merged_data.txt
-
Number of patients = 80
-
Number of clustering approaches = 8
-
Number of selected clinical features = 7
-
Exclude small clusters that include fewer than K patients, K = 3
consensus non-negative matrix factorization clustering approach (Brunet et al. 2004)
Resampling-based clustering method (Monti et al. 2003)
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For binary clinical features, two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.