This report serves to describe the mutational landscape and properties of a given individual set, as well as rank genes and genesets according to mutational significance. MutSig v1.5 was used to generate the results found in this report.
Working with individual set: UCEC.
Number of patients in set: 239
The input for this pipeline is a set of individuals with the following files associated for each:
1. An annotated .maf file describing the mutations called for the respective individual, and their properties.
2. A .wig file that contains information about the coverage of the sample.
Significantly mutated genes (q ≤ 0.1): 240
Mutations seen in COSMIC: 1
Significantly mutated genes in COSMIC territory: 0
Genes with clustered mutations (&le 3 aa apart): 21
Significantly mutated genesets: 217
Significantly mutated genesets: (excluding sig. mutated genes): 50
Table 1. Get Full Table Table representing breakdown of mutations by type.
| type | count |
|---|---|
| Frame_Shift_Del | 2848 |
| Frame_Shift_Ins | 1040 |
| In_Frame_Del | 204 |
| In_Frame_Ins | 28 |
| Indel | 2 |
| Missense_Mutation | 112137 |
| Nonsense_Mutation | 11196 |
| Nonstop_Mutation | 160 |
| Silent | 36723 |
| Splice_Site | 2821 |
| Total | 167159 |
Table 2. Get Full Table A breakdown of mutation rates per category discovered for this individual set.
| category | n | N | rate | rate_per_mb | relative_rate |
|---|---|---|---|---|---|
| A->T | 34661 | 3330542914 | 1e-05 | 10 | 0.53 |
| C->(A/T) | 52348 | 3324649891 | 0.000016 | 16 | 0.8 |
| A->(C/G) | 18461 | 3330542914 | 5.5e-06 | 5.5 | 0.28 |
| C->G | 6658 | 3324649891 | 2e-06 | 2 | 0.1 |
| indel+null | 16724 | 6655193044 | 2.5e-06 | 2.5 | 0.13 |
| double_null | 1584 | 6655193044 | 2.4e-07 | 0.24 | 0.012 |
| Total | 130436 | 6655193044 | 2e-05 | 20 | 1 |
The x axis represents the samples. The y axis represents the exons, one row per exon, and they are sorted by average coverage across samples. For exons with exactly the same average coverage, they are sorted next by the %GC of the exon. (The secondary sort is especially useful for the zero-coverage exons at the bottom).
Figure 1.
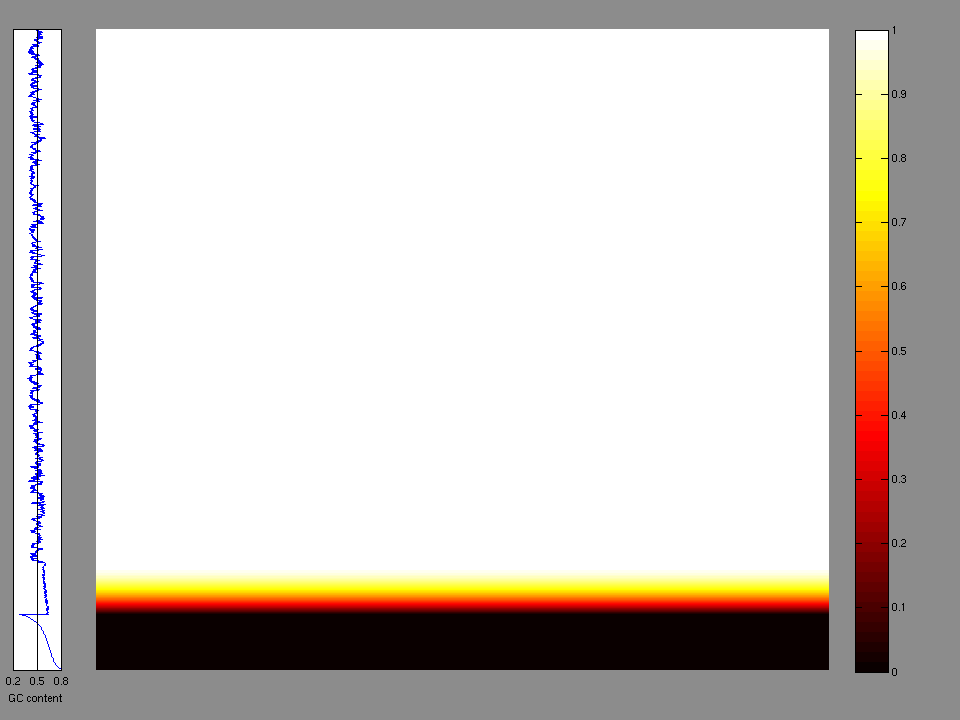

Figure 2.
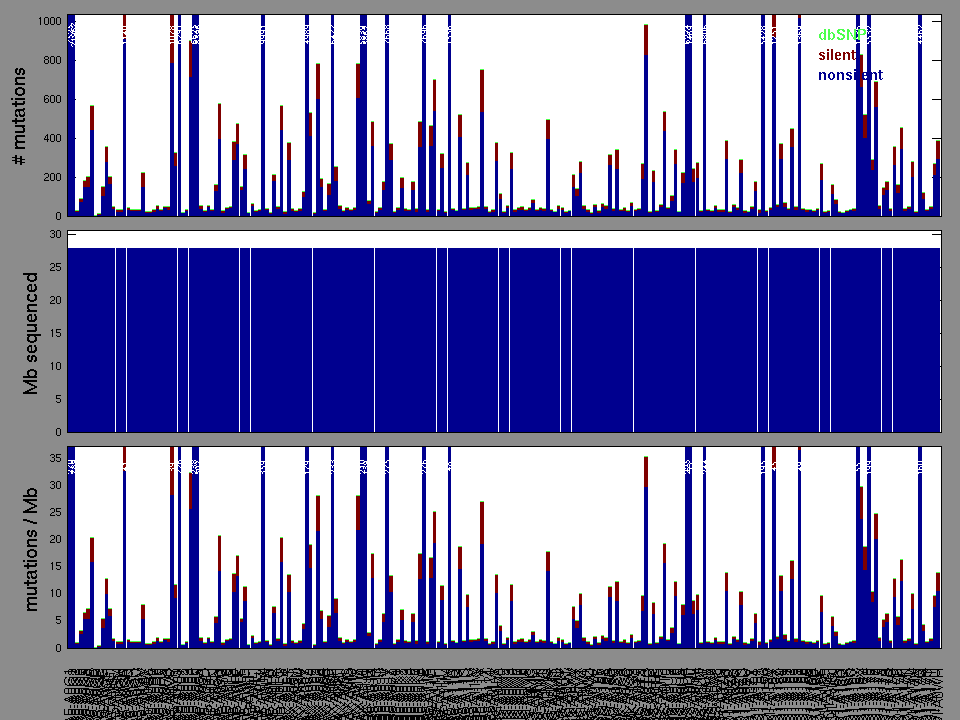
Table 3. Get Full Table A Ranked List of Significantly Mutated Genes. Number of significant genes found: 240. Number of genes displayed: 35
| rank | gene | description | N | n | npat | nsite | nsil | n1 | n2 | n3 | n4 | n5 | n6 | p | q |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | PTEN | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 298272 | 198 | 150 | 121 | 5 | 17 | 12 | 54 | 5 | 90 | 20 | <1.00e-15 | <1.73e-11 |
| 2 | ARID1A | AT rich interactive domain 1A (SWI-like) | 1385483 | 109 | 86 | 85 | 5 | 5 | 7 | 3 | 2 | 68 | 24 | 8.66e-15 | 7.48e-11 |
| 3 | KRAS | v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog | 168973 | 29 | 29 | 5 | 0 | 19 | 0 | 10 | 0 | 0 | 0 | 4.03e-14 | 9.78e-11 |
| 4 | RPL22 | ribosomal protein L22 | 92493 | 15 | 14 | 5 | 0 | 0 | 1 | 0 | 2 | 10 | 2 | 4.46e-14 | 9.78e-11 |
| 5 | TP53 | tumor protein p53 | 313329 | 40 | 36 | 30 | 2 | 12 | 9 | 4 | 3 | 12 | 0 | 4.50e-14 | 9.78e-11 |
| 6 | ARID5B | AT rich interactive domain 5B (MRF1-like) | 862073 | 36 | 29 | 32 | 8 | 7 | 8 | 4 | 3 | 12 | 2 | 4.64e-14 | 9.78e-11 |
| 7 | PIK3CA | phosphoinositide-3-kinase, catalytic, alpha polypeptide | 785593 | 138 | 110 | 61 | 2 | 49 | 31 | 44 | 8 | 6 | 0 | 4.93e-14 | 9.78e-11 |
| 8 | PIK3R1 | phosphoinositide-3-kinase, regulatory subunit 1 (alpha) | 565235 | 67 | 52 | 55 | 2 | 3 | 6 | 13 | 3 | 31 | 11 | 5.05e-14 | 9.78e-11 |
| 9 | CTNNB1 | catenin (cadherin-associated protein), beta 1, 88kDa | 574078 | 66 | 63 | 21 | 5 | 11 | 35 | 8 | 12 | 0 | 0 | 5.30e-14 | 9.78e-11 |
| 10 | CTCF | CCCTC-binding factor (zinc finger protein) | 531536 | 55 | 48 | 40 | 1 | 3 | 9 | 5 | 1 | 33 | 4 | 5.66e-14 | 9.78e-11 |
| 11 | PPP2R1A | protein phosphatase 2 (formerly 2A), regulatory subunit A , alpha isoform | 417772 | 27 | 24 | 20 | 3 | 6 | 15 | 1 | 3 | 2 | 0 | 3.19e-13 | 5.00e-10 |
| 12 | FGFR2 | fibroblast growth factor receptor 2 (bacteria-expressed kinase, keratinocyte growth factor receptor, craniofacial dysostosis 1, Crouzon syndrome, Pfeiffer syndrome, Jackson-Weiss syndrome) | 644344 | 27 | 25 | 15 | 2 | 2 | 9 | 5 | 10 | 1 | 0 | 1.65e-12 | 2.38e-09 |
| 13 | MLL4 | 1092469 | 53 | 35 | 50 | 8 | 11 | 18 | 8 | 2 | 14 | 0 | 2.61e-12 | 3.47e-09 | |
| 14 | MUC5B | mucin 5B, oligomeric mucus/gel-forming | 1730599 | 78 | 46 | 77 | 35 | 13 | 45 | 9 | 3 | 8 | 0 | 1.03e-11 | 1.27e-08 |
| 15 | FBXW7 | F-box and WD repeat domain containing 7 | 617576 | 34 | 24 | 28 | 2 | 5 | 15 | 1 | 2 | 9 | 2 | 2.16e-11 | 2.49e-08 |
| 16 | BCOR | BCL6 co-repressor | 1215076 | 52 | 38 | 41 | 10 | 6 | 27 | 14 | 1 | 4 | 0 | 2.61e-11 | 2.82e-08 |
| 17 | SPOP | speckle-type POZ protein | 277479 | 19 | 18 | 15 | 0 | 11 | 1 | 2 | 2 | 3 | 0 | 5.06e-11 | 5.14e-08 |
| 18 | RNF43 | ring finger protein 43 | 567386 | 22 | 22 | 15 | 2 | 1 | 7 | 0 | 0 | 14 | 0 | 1.00e-10 | 9.60e-08 |
| 19 | ZFHX3 | zinc finger homeobox 3 | 2606295 | 80 | 45 | 75 | 11 | 16 | 33 | 5 | 2 | 18 | 6 | 1.30e-09 | 1.18e-06 |
| 20 | RB1 | retinoblastoma 1 (including osteosarcoma) | 658206 | 31 | 25 | 30 | 4 | 7 | 7 | 6 | 1 | 6 | 4 | 3.39e-09 | 2.93e-06 |
| 21 | RASA1 | RAS p21 protein activator (GTPase activating protein) 1 | 689993 | 29 | 22 | 27 | 2 | 6 | 6 | 3 | 1 | 11 | 2 | 1.00e-07 | 0.000082 |
| 22 | CCND1 | cyclin D1 | 118783 | 10 | 10 | 9 | 1 | 3 | 3 | 1 | 1 | 2 | 0 | 1.66e-07 | 0.00013 |
| 23 | PCGF3 | polycomb group ring finger 3 | 92971 | 10 | 10 | 8 | 1 | 1 | 6 | 0 | 0 | 3 | 0 | 2.87e-07 | 0.00022 |
| 24 | PXDN | peroxidasin homolog (Drosophila) | 790612 | 43 | 27 | 41 | 8 | 9 | 19 | 3 | 3 | 7 | 2 | 3.38e-07 | 0.00024 |
| 25 | SMTNL2 | smoothelin-like 2 | 152004 | 9 | 9 | 3 | 2 | 1 | 1 | 0 | 0 | 7 | 0 | 3.44e-07 | 0.00024 |
| 26 | ETV3 | ets variant gene 3 | 106116 | 12 | 11 | 12 | 0 | 3 | 2 | 5 | 0 | 2 | 0 | 5.05e-07 | 0.00034 |
| 27 | NFASC | neurofascin homolog (chicken) | 983246 | 33 | 27 | 33 | 3 | 10 | 13 | 4 | 1 | 3 | 2 | 6.96e-07 | 0.00044 |
| 28 | CSMD3 | CUB and Sushi multiple domains 3 | 2741330 | 106 | 40 | 101 | 20 | 33 | 37 | 15 | 5 | 7 | 9 | 7.14e-07 | 0.00044 |
| 29 | DCC | deleted in colorectal carcinoma | 1065940 | 35 | 27 | 34 | 3 | 11 | 14 | 3 | 2 | 5 | 0 | 1.86e-06 | 0.0011 |
| 30 | INPPL1 | inositol polyphosphate phosphatase-like 1 | 794914 | 23 | 19 | 20 | 7 | 4 | 6 | 2 | 0 | 9 | 2 | 2.15e-06 | 0.0012 |
| 31 | CHD3 | chromodomain helicase DNA binding protein 3 | 1425157 | 39 | 30 | 35 | 5 | 10 | 12 | 7 | 1 | 9 | 0 | 2.17e-06 | 0.0012 |
| 32 | MUC4 | mucin 4, cell surface associated | 1204082 | 43 | 29 | 42 | 17 | 9 | 20 | 5 | 5 | 4 | 0 | 3.14e-06 | 0.0017 |
| 33 | CNTN6 | contactin 6 | 758825 | 36 | 24 | 34 | 7 | 8 | 14 | 4 | 2 | 8 | 0 | 3.78e-06 | 0.0019 |
| 34 | LRCH2 | leucine-rich repeats and calponin homology (CH) domain containing 2 | 355871 | 23 | 19 | 22 | 2 | 10 | 6 | 3 | 0 | 4 | 0 | 3.83e-06 | 0.0019 |
| 35 | BRD3 | bromodomain containing 3 | 239000 | 15 | 14 | 13 | 7 | 6 | 3 | 1 | 0 | 5 | 0 | 3.94e-06 | 0.0019 |
Note:
N - number of sequenced bases in this gene across the individual set.
n - number of (nonsilent) mutations in this gene across the individual set.
npat - number of patients (individuals) with at least one nonsilent mutation.
nsite - number of unique sites having a non-silent mutation.
nsil - number of silent mutations in this gene across the individual set.
n1 - number of nonsilent mutations of type: A->T .
n2 - number of nonsilent mutations of type: C->(A/T) .
n3 - number of nonsilent mutations of type: A->(C/G) .
n4 - number of nonsilent mutations of type: C->G .
n5 - number of nonsilent mutations of type: indel+null .
null - mutation category that includes nonsense, frameshift, splice-site mutations
p_classic = p-value for the observed amount of nonsilent mutations being elevated in this gene
p_ns_s = p-value for the observed nonsilent/silent ratio being elevated in this gene
p = p-value (overall)
q = q-value, False Discovery Rate (Benjamini-Hochberg procedure)
In this analysis, COSMIC is used as a filter to increase power by restricting the territory of each gene. Cosmic version: v48.
Table 4. Get Full Table Significantly mutated genes (COSMIC territory only). To access the database please go to: COSMIC. Number of significant genes found: 0. Number of genes displayed: 10
| rank | gene | description | n | cos | n_cos | N_cos | cos_ev | p | q |
|---|---|---|---|---|---|---|---|---|---|
| 1 | SYNE1 | spectrin repeat containing, nuclear envelope 1 | 116 | 22 | 1 | 5258 | 1 | 0.098 | 1 |
| 2 | A4GNT | alpha-1,4-N-acetylglucosaminyltransferase | 6 | 0 | 0 | 0 | 0 | 1 | 1 |
| 3 | AACS | acetoacetyl-CoA synthetase | 9 | 0 | 0 | 0 | 0 | 1 | 1 |
| 4 | ABCA9 | ATP-binding cassette, sub-family A (ABC1), member 9 | 29 | 0 | 0 | 0 | 0 | 1 | 1 |
| 5 | ABCC10 | ATP-binding cassette, sub-family C (CFTR/MRP), member 10 | 14 | 0 | 0 | 0 | 0 | 1 | 1 |
| 6 | ABCF2 | ATP-binding cassette, sub-family F (GCN20), member 2 | 5 | 0 | 0 | 0 | 0 | 1 | 1 |
| 7 | ABHD2 | abhydrolase domain containing 2 | 4 | 0 | 0 | 0 | 0 | 1 | 1 |
| 8 | ABHD4 | abhydrolase domain containing 4 | 11 | 0 | 0 | 0 | 0 | 1 | 1 |
| 9 | ACADS | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 5 | 0 | 0 | 0 | 0 | 1 | 1 |
| 10 | ACOT11 | acyl-CoA thioesterase 11 | 6 | 0 | 0 | 0 | 0 | 1 | 1 |
Note:
n - number of (nonsilent) mutations in this gene across the individual set.
cos = number of unique mutated sites in this gene in COSMIC
n_cos = overlap between n and cos.
N_cos = number of individuals times cos.
cos_ev = total evidence: number of reports in COSMIC for mutations seen in this gene.
p = p-value for seeing the observed amount of overlap in this gene)
q = q-value, False Discovery Rate (Benjamini-Hochberg procedure)
Table 5. Get Full Table Genes with Clustered Mutations
| num | gene | desc | n | mindist | npairs3 | npairs12 |
|---|---|---|---|---|---|---|
| 1878 | C3orf63 | chromosome 3 open reading frame 63 | 25 | 0 | 3 | 3 |
| 3911 | DMBT1 | deleted in malignant brain tumors 1 | 34 | 0 | 1 | 3 |
| 2601 | CDCP1 | CUB domain containing protein 1 | 13 | 0 | 1 | 1 |
| 3932 | DNAH11 | dynein, axonemal, heavy chain 11 | 84 | 0 | 1 | 1 |
| 7129 | KIAA1432 | KIAA1432 | 16 | 0 | 1 | 1 |
| 7178 | KIF20B | 47 | 0 | 1 | 1 | |
| 7472 | LARP1 | La ribonucleoprotein domain family, member 1 | 17 | 0 | 1 | 1 |
| 8159 | MDN1 | MDN1, midasin homolog (yeast) | 58 | 0 | 1 | 1 |
| 9070 | NLGN4X | neuroligin 4, X-linked | 11 | 0 | 1 | 1 |
| 10055 | PCM1 | pericentriolar material 1 | 25 | 0 | 1 | 1 |
Note:
n - number of mutations in this gene in the individual set.
mindist - distance (in aa) between closest pair of mutations in this gene
npairs3 - how many pairs of mutations are within 3 aa of each other.
npairs12 - how many pairs of mutations are within 12 aa of each other.
Table 6. Get Full Table A Ranked List of Significantly Mutated Genesets. (Source: MSigDB GSEA Cannonical Pathway Set).Number of significant genesets found: 217. Number of genesets displayed: 10
| rank | geneset | description | genes | N_genes | mut_tally | N | n | npat | nsite | nsil | n1 | n2 | n3 | n4 | n5 | n6 | p | q |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | SIG_CHEMOTAXIS | Genes related to chemotaxis | ACTR2, ACTR3, AKT1, AKT2, AKT3, ANGPTL2, ARHGAP1, ARHGAP4, ARHGEF11, BTK, CDC42, CFL1, CFL2, GDI1, GDI2, INPPL1, ITPR1, ITPR2, ITPR3, LIMK1, MYLK, MYLK2, P101-PI3K, PAK1, PAK2, PAK3, PAK4, PAK6, PAK7, PDK1, PIK3CA, PIK3CD, PIK3CG, PIK3R1, PITX2, PPP1R13B, PTEN, RACGAP1, RHO, ROCK1, ROCK2, RPS4X, SAG, WASF1, WASL | 44 | ACTR2(7), ACTR3(3), AKT1(8), AKT2(4), AKT3(8), ANGPTL2(9), ARHGAP1(2), ARHGAP4(15), ARHGEF11(22), BTK(11), CDC42(5), CFL1(1), CFL2(8), GDI1(4), GDI2(3), INPPL1(23), ITPR1(31), ITPR2(38), ITPR3(23), LIMK1(7), MYLK(17), MYLK2(7), PAK1(3), PAK2(6), PAK3(14), PAK4(2), PAK6(3), PAK7(12), PDK1(7), PIK3CA(138), PIK3CD(10), PIK3CG(25), PIK3R1(67), PITX2(10), PPP1R13B(7), PTEN(198), RACGAP1(8), RHO(8), ROCK1(27), ROCK2(15), RPS4X(4), SAG(5), WASF1(3), WASL(8) | 24593100 | 836 | 210 | 656 | 135 | 193 | 221 | 160 | 34 | 189 | 39 | <1.00e-15 | <2.46e-14 |
| 2 | EIF4PATHWAY | The eIF-4F complex recognizes 5' mRNA caps, recruits RNA helicases, and maintains mRNA-ribosome bridging. | AKT1, EIF4A1, EIF4A2, EIF4E, EIF4EBP1, EIF4G1, EIF4G2, EIF4G3, FRAP1, GHR, IRS1, MAPK1, MAPK14, MAPK3, MKNK1, PABPC1, PDK2, PDPK1, PIK3CA, PIK3R1, PRKCA, PRKCB1, PTEN, RPS6KB1 | 22 | AKT1(8), EIF4A1(4), EIF4A2(6), EIF4E(4), EIF4G1(15), EIF4G2(8), EIF4G3(26), GHR(10), IRS1(17), MAPK1(2), MAPK14(6), MAPK3(4), MKNK1(6), PABPC1(11), PDK2(5), PDPK1(3), PIK3CA(138), PIK3R1(67), PRKCA(10), PTEN(198), RPS6KB1(3) | 9912525 | 551 | 208 | 382 | 44 | 101 | 111 | 133 | 26 | 149 | 31 | <1.00e-15 | <2.46e-14 |
| 3 | IGF1MTORPATHWAY | Growth factor IGF-1 activates AKT, Gsk3-beta, and mTOR to promote muscle hypertrophy. | AKT1, EIF2B5, EIF2S1, EIF2S2, EIF2S3, EIF4E, EIF4EBP1, FRAP1, GSK3B, IGF1, IGF1R, INPPL1, PDK2, PDPK1, PIK3CA, PIK3R1, PPP2CA, PTEN, RPS6, RPS6KB1 | 19 | AKT1(8), EIF2B5(6), EIF2S1(1), EIF2S2(5), EIF2S3(6), EIF4E(4), GSK3B(11), IGF1(3), IGF1R(13), INPPL1(23), PDK2(5), PDPK1(3), PIK3CA(138), PIK3R1(67), PPP2CA(6), PTEN(198), RPS6(6), RPS6KB1(3) | 6967089 | 506 | 208 | 333 | 43 | 91 | 92 | 121 | 20 | 149 | 33 | <1.00e-15 | <2.46e-14 |
| 4 | MTORPATHWAY | Mammalian target of rapamycin (mTOR) senses mitogenic factors and nutrients, including ATP, and induces cell proliferation. | AKT1, EIF3S10, EIF4A1, EIF4A2, EIF4B, EIF4E, EIF4EBP1, EIF4G1, EIF4G2, EIF4G3, FKBP1A, FRAP1, MKNK1, PDK2, PDPK1, PIK3CA, PIK3R1, PPP2CA, PTEN, RPS6, RPS6KB1, TSC1, TSC2 | 21 | AKT1(8), EIF4A1(4), EIF4A2(6), EIF4B(6), EIF4E(4), EIF4G1(15), EIF4G2(8), EIF4G3(26), MKNK1(6), PDK2(5), PDPK1(3), PIK3CA(138), PIK3R1(67), PPP2CA(6), PTEN(198), RPS6(6), RPS6KB1(3), TSC1(13), TSC2(14) | 9012929 | 536 | 208 | 366 | 42 | 96 | 113 | 124 | 25 | 147 | 31 | <1.00e-15 | <2.46e-14 |
| 5 | SIG_INSULIN_RECEPTOR_PATHWAY_IN_CARDIAC_MYOCYTES | Genes related to the insulin receptor pathway | AKT1, AKT2, AKT3, BRD4, CAP1, CBL, CDC42, CDKN2A, F2RL2, FLOT1, FLOT2, FOXO1A, GRB2, GSK3A, GSK3B, IGFBP1, INPPL1, IRS1, IRS2, IRS4, LNPEP, MAPK1, MAPK3, PARD3, PARD6A, PDK1, PIK3CA, PIK3CD, PIK3R1, PPYR1, PSCD3, PTEN, PTPN1, RAF1, RPS6KA1, RPS6KA2, RPS6KA3, RPS6KB1, SERPINB6, SFN, SHC1, SLC2A4, SORBS1, SOS1, SOS2, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ | 48 | AKT1(8), AKT2(4), AKT3(8), BRD4(9), CAP1(7), CBL(10), CDC42(5), CDKN2A(1), F2RL2(3), FLOT1(1), FLOT2(3), GRB2(3), GSK3A(9), GSK3B(11), IGFBP1(5), INPPL1(23), IRS1(17), IRS4(20), LNPEP(9), MAPK1(2), MAPK3(4), PARD3(27), PARD6A(1), PDK1(7), PIK3CA(138), PIK3CD(10), PIK3R1(67), PPYR1(7), PTEN(198), PTPN1(4), RAF1(7), RPS6KA1(5), RPS6KA2(16), RPS6KA3(17), RPS6KB1(3), SERPINB6(4), SHC1(11), SLC2A4(4), SORBS1(22), SOS1(21), SOS2(22), YWHAB(1), YWHAE(1), YWHAG(3), YWHAH(4), YWHAQ(3), YWHAZ(1) | 20132165 | 766 | 208 | 583 | 117 | 159 | 183 | 167 | 32 | 188 | 37 | <1.00e-15 | <2.46e-14 |
| 6 | METPATHWAY | The hepatocyte growth factor receptor c-Met stimulates proliferation and alters cell motility and adhesion on binding the ligand HGF. | ACTA1, CRK, CRKL, DOCK1, ELK1, FOS, GAB1, GRB2, GRF2, HGF, HRAS, ITGA1, ITGB1, JUN, MAP2K1, MAP2K2, MAP4K1, MAPK1, MAPK3, MAPK8, MET, PAK1, PIK3CA, PIK3R1, PTEN, PTK2, PTK2B, PTPN11, PXN, RAF1, RAP1A, RAP1B, RASA1, SOS1, SRC, STAT3 | 34 | ACTA1(4), CRK(1), CRKL(5), DOCK1(32), ELK1(2), FOS(6), GAB1(10), GRB2(3), HGF(15), ITGA1(20), ITGB1(12), JUN(1), MAP2K1(2), MAP4K1(13), MAPK1(2), MAPK3(4), MAPK8(7), MET(10), PAK1(3), PIK3CA(138), PIK3R1(67), PTEN(198), PTK2(11), PTK2B(16), PTPN11(8), PXN(6), RAF1(7), RAP1A(3), RAP1B(2), RASA1(29), SOS1(21), SRC(2), STAT3(12) | 15458520 | 672 | 205 | 498 | 85 | 136 | 158 | 146 | 27 | 170 | 35 | <1.00e-15 | <2.46e-14 |
| 7 | PTENPATHWAY | PTEN suppresses AKT-induced cell proliferation and antagonizes the action of PI3K. | AKT1, BCAR1, CDKN1B, FOXO3A, GRB2, ILK, ITGB1, MAPK1, MAPK3, PDK2, PDPK1, PIK3CA, PIK3R1, PTEN, PTK2, SHC1, SOS1, TNFSF6 | 15 | AKT1(8), CDKN1B(5), GRB2(3), ILK(8), ITGB1(12), MAPK1(2), MAPK3(4), PDK2(5), PDPK1(3), PIK3CA(138), PIK3R1(67), PTEN(198), PTK2(11), SHC1(11), SOS1(21) | 6356683 | 496 | 203 | 321 | 30 | 86 | 92 | 124 | 20 | 141 | 33 | <1.00e-15 | <2.46e-14 |
| 8 | ST_PHOSPHOINOSITIDE_3_KINASE_PATHWAY | The phosphoinositide-3 kinase pathway produces the lipid second messenger PIP3 and regulates cell growth, survival, and movement. | A1BG, AKT1, AKT2, AKT3, BAD, BTK, CDKN2A, CSL4, DAF, DAPP1, FOXO1A, GRB2, GSK3A, GSK3B, IARS, IGFBP1, INPP5D, P14, PDK1, PIK3CA, PPP1R13B, PSCD3, PTEN, RPS6KA1, RPS6KA2, RPS6KA3, RPS6KB1, SFN, SHC1, SOS1, SOS2, TEC, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ | 33 | A1BG(3), AKT1(8), AKT2(4), AKT3(8), BAD(2), BTK(11), CDKN2A(1), DAPP1(7), GRB2(3), GSK3A(9), GSK3B(11), IARS(13), IGFBP1(5), INPP5D(14), PDK1(7), PIK3CA(138), PPP1R13B(7), PTEN(198), RPS6KA1(5), RPS6KA2(16), RPS6KA3(17), RPS6KB1(3), SHC1(11), SOS1(21), SOS2(22), TEC(5), YWHAB(1), YWHAE(1), YWHAG(3), YWHAH(4), YWHAQ(3), YWHAZ(1) | 12586696 | 562 | 201 | 396 | 60 | 115 | 137 | 132 | 22 | 132 | 24 | <1.00e-15 | <2.46e-14 |
| 9 | SA_PTEN_PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. | AKT1, AKT2, AKT3, BPNT1, GRB2, ILK, MAPK1, MAPK3, PDK1, PIK3CA, PIK3CD, PIP3-E, PTEN, PTK2B, RBL2, SHC1, SOS1 | 16 | AKT1(8), AKT2(4), AKT3(8), BPNT1(1), GRB2(3), ILK(8), MAPK1(2), MAPK3(4), PDK1(7), PIK3CA(138), PIK3CD(10), PTEN(198), PTK2B(16), RBL2(21), SHC1(11), SOS1(21) | 6977366 | 460 | 198 | 293 | 36 | 98 | 93 | 112 | 20 | 112 | 25 | <1.00e-15 | <2.46e-14 |
| 10 | HSA04210_APOPTOSIS | Genes involved in apoptosis | AIFM1, AKT1, AKT2, AKT3, APAF1, ATM, BAD, BAX, BCL2, BCL2L1, BID, BIRC2, BIRC3, BIRC4, CAPN1, CAPN2, CASP10, CASP3, CASP6, CASP7, CASP8, CASP9, CFLAR, CHP, CHUK, CSF2RB, CYCS, DFFA, DFFB, ENDOG, FADD, FAS, FASLG, IKBKB, IKBKG, IL1A, IL1B, IL1R1, IL1RAP, IL3, IL3RA, IRAK1, IRAK2, IRAK3, IRAK4, MAP3K14, MYD88, NFKB1, NFKB2, NFKBIA, NGFB, NTRK1, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PRKACA, PRKACB, PRKACG, PRKAR1A, PRKAR1B, PRKAR2A, PRKAR2B, RELA, RIPK1, TNF, TNFRSF10A, TNFRSF10B, TNFRSF10C, TNFRSF10D, TNFRSF1A, TNFSF10, TP53, TRADD, TRAF2 | 79 | AIFM1(12), AKT1(8), AKT2(4), AKT3(8), APAF1(19), ATM(54), BAD(2), BAX(4), BCL2L1(3), BID(3), BIRC2(13), BIRC3(3), CAPN1(4), CAPN2(6), CASP10(10), CASP3(4), CASP6(4), CASP7(4), CASP8(21), CASP9(3), CFLAR(7), CHUK(11), CSF2RB(11), CYCS(3), DFFA(1), DFFB(3), FADD(1), FAS(7), FASLG(6), IKBKB(15), IL1A(1), IL1B(2), IL1R1(5), IL1RAP(2), IL3RA(11), IRAK1(5), IRAK2(7), IRAK3(13), IRAK4(6), MAP3K14(12), MYD88(1), NFKB1(12), NFKB2(10), NFKBIA(3), NTRK1(11), PIK3CA(138), PIK3CB(19), PIK3CD(10), PIK3CG(25), PIK3R1(67), PIK3R2(4), PIK3R3(8), PIK3R5(8), PPP3CA(4), PPP3CB(7), PPP3CC(6), PPP3R1(2), PPP3R2(1), PRKACA(4), PRKACB(2), PRKACG(6), PRKAR1A(4), PRKAR1B(8), PRKAR2A(4), PRKAR2B(5), RELA(6), RIPK1(8), TNF(1), TNFRSF10A(3), TNFRSF10B(5), TNFRSF10C(3), TNFRSF10D(10), TNFRSF1A(6), TNFSF10(1), TP53(40), TRAF2(5) | 28018448 | 775 | 186 | 659 | 164 | 177 | 260 | 136 | 44 | 141 | 17 | <1.00e-15 | <2.46e-14 |
Table 7. Get Full Table A Ranked List of Significantly Mutated Genesets (Excluding Significantly Mutated Genes). Number of significant genesets found: 50. Number of genesets displayed: 10
| rank | geneset | description | genes | N_genes | mut_tally | N | n | npat | nsite | nsil | n1 | n2 | n3 | n4 | n5 | n6 | p | q |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HSA04020_CALCIUM_SIGNALING_PATHWAY | Genes involved in calcium signaling pathway | ADCY1, ADCY2, ADCY3, ADCY4, ADCY7, ADCY8, ADCY9, ADORA2A, ADORA2B, ADRA1A, ADRA1B, ADRA1D, ADRB1, ADRB2, ADRB3, AGTR1, ATP2A1, ATP2A2, ATP2A3, ATP2B1, ATP2B2, ATP2B3, ATP2B4, AVPR1A, AVPR1B, BDKRB1, BDKRB2, BST1, CACNA1A, CACNA1B, CACNA1C, CACNA1D, CACNA1E, CACNA1F, CACNA1G, CACNA1H, CACNA1I, CACNA1S, CALM1, CALM2, CALM3, CALML3, CALML6, CAMK2A, CAMK2B, CAMK2D, CAMK2G, CAMK4, CCKAR, CCKBR, CD38, CHP, CHRM1, CHRM2, CHRM3, CHRM5, CHRNA7, CYSLTR1, CYSLTR2, DRD1, EDNRA, EDNRB, EGFR, ERBB2, ERBB3, ERBB4, F2R, GNA11, GNA14, GNA15, GNAL, GNAQ, GNAS, GRIN1, GRIN2A, GRIN2C, GRIN2D, GRM1, GRM5, GRPR, HRH1, HRH2, HTR2A, HTR2B, HTR2C, HTR4, HTR5A, HTR6, HTR7, ITPKA, ITPKB, ITPR1, ITPR2, ITPR3, LHCGR, LTB4R2, MLCK, MYLK, MYLK2, NOS1, NOS2A, NOS3, NTSR1, OXTR, P2RX1, P2RX2, P2RX3, P2RX4, P2RX5, P2RX7, P2RXL1, PDE1A, PDE1B, PDE1C, PDGFRA, PDGFRB, PHKA1, PHKA2, PHKB, PHKG1, PHKG2, PLCB1, PLCB2, PLCB3, PLCB4, PLCD1, PLCD3, PLCD4, PLCE1, PLCG1, PLCG2, PLCZ1, PLN, PPID, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PRKACA, PRKACB, PRKACG, PRKCA, PRKCB1, PRKCG, PRKX, PRKY, PTAFR, PTGER1, PTGER3, PTGFR, PTK2B, RYR1, RYR2, RYR3, SLC25A4, SLC25A5, SLC25A6, SLC8A1, SLC8A2, SLC8A3, SPHK1, SPHK2, TACR1, TACR2, TACR3, TBXA2R, TNNC1, TNNC2, TRHR, TRPC1, VDAC1, VDAC2, VDAC3 | 162 | ADCY1(17), ADCY2(17), ADCY3(8), ADCY4(8), ADCY7(6), ADCY8(16), ADCY9(25), ADORA2A(5), ADORA2B(2), ADRA1A(11), ADRA1B(7), ADRB1(2), ADRB2(8), ATP2A1(16), ATP2A2(9), ATP2A3(12), ATP2B1(17), ATP2B2(12), ATP2B3(21), ATP2B4(18), AVPR1A(8), AVPR1B(3), BDKRB1(5), BDKRB2(9), BST1(5), CACNA1A(19), CACNA1B(1), CACNA1C(30), CACNA1D(34), CACNA1E(47), CACNA1F(36), CACNA1G(20), CACNA1I(13), CACNA1S(33), CALM2(3), CALM3(1), CALML6(4), CAMK2A(8), CAMK2B(2), CAMK2D(7), CAMK2G(8), CAMK4(5), CCKAR(12), CCKBR(7), CD38(7), CHRM1(6), CHRM2(9), CHRM3(9), CHRM5(13), CHRNA7(6), CYSLTR1(5), CYSLTR2(6), DRD1(6), EDNRA(6), EDNRB(2), EGFR(20), ERBB2(6), ERBB3(16), ERBB4(29), F2R(8), GNA11(3), GNA14(5), GNA15(1), GNAL(6), GNAS(25), GRIN1(5), GRIN2A(25), GRIN2C(6), GRIN2D(8), GRM1(22), GRM5(22), GRPR(4), HRH1(9), HRH2(3), HTR2A(15), HTR2B(2), HTR4(7), HTR5A(9), HTR6(4), HTR7(4), ITPKB(6), ITPR1(31), ITPR2(38), ITPR3(23), LHCGR(16), LTB4R2(3), MYLK(17), MYLK2(7), NOS1(21), NOS3(11), NTSR1(4), OXTR(4), P2RX1(9), P2RX3(9), P2RX4(2), P2RX5(8), P2RX7(3), PDE1A(12), PDE1B(9), PDE1C(14), PDGFRA(29), PDGFRB(12), PHKA1(17), PHKA2(15), PHKB(14), PHKG1(6), PHKG2(3), PLCB1(6), PLCB2(9), PLCB3(9), PLCB4(20), PLCD1(5), PLCD3(7), PLCD4(13), PLCE1(36), PLCG1(12), PLCG2(27), PLCZ1(16), PLN(1), PPID(6), PPP3CA(4), PPP3CB(7), PPP3CC(6), PPP3R1(2), PPP3R2(1), PRKACA(4), PRKACB(2), PRKACG(6), PRKCA(10), PRKCG(10), PRKX(4), PTAFR(6), PTGER3(9), PTGFR(10), PTK2B(16), RYR1(80), RYR2(111), RYR3(87), SLC25A4(3), SLC25A5(4), SLC25A6(6), SLC8A1(20), SLC8A2(6), SLC8A3(14), SPHK1(2), SPHK2(1), TACR1(6), TACR2(4), TACR3(9), TNNC1(2), TNNC2(4), TRHR(5), TRPC1(15), VDAC1(2), VDAC3(4) | 91603920 | 1888 | 150 | 1836 | 659 | 546 | 777 | 260 | 92 | 190 | 23 | 3.2e-15 | 2e-12 |
| 2 | CALCIUM_REGULATION_IN_CARDIAC_CELLS | ADCY1, ADCY2, ADCY3, ADCY4, ADCY5, ADCY6, ADCY7, ADCY8, ADCY9, ADRA1A, ADRA1B, ADRA1D, ADRB1, ADRB2, ADRB3, ANXA6, ARRB1, ARRB2, ATP1A4, ATP1B1, ATP1B2, ATP1B3, ATP2A2, ATP2A3, ATP2B1, ATP2B2, ATP2B3, CACNA1A, CACNA1B, CACNA1C, CACNA1D, CACNA1E, CACNA1S, CACNB1, CACNB3, CALM1, CALM2, CALM3, CALR, CAMK1, CAMK2A, CAMK2B, CAMK2D, CAMK2G, CAMK4, CASQ1, CASQ2, CHRM1, CHRM2, CHRM3, CHRM4, CHRM5, FXYD2, GJA1, GJA12, GJA4, GJA5, GJB1, GJB2, GJB3, GJB4, GJB5, GJB6, GNA11, GNAI2, GNAI3, GNAO1, GNAQ, GNAZ, GNB1, GNB2, GNB3, GNB4, GNB5, GNG12, GNG13, GNG2, GNG3, GNG4, GNG5, GNG7, GNGT1, GRK4, GRK5, GRK6, ITPR1, ITPR2, ITPR3, KCNB1, KCNJ3, KCNJ5, MGC11266, MYCBP, NME7, PEA15, PKIA, PKIB, PKIG, PLCB3, PLN, PRKACA, PRKACB, PRKAR1A, PRKAR1B, PRKAR2A, PRKAR2B, PRKCA, PRKCB1, PRKCD, PRKCE, PRKCG, PRKCH, PRKCQ, PRKCZ, PRKD1, RGS1, RGS10, RGS11, RGS14, RGS16, RGS17, RGS18, RGS19, RGS2, RGS20, RGS3, RGS4, RGS5, RGS6, RGS7, RGS9, RYR1, RYR2, RYR3, SARA1, SFN, SLC8A1, SLC8A3, USP5, YWHAB, YWHAH, YWHAQ, YWHAQ, MIB1 | 133 | ADCY1(17), ADCY2(17), ADCY3(8), ADCY4(8), ADCY5(11), ADCY6(16), ADCY7(6), ADCY8(16), ADCY9(25), ADRA1A(11), ADRA1B(7), ADRB1(2), ADRB2(8), ANXA6(10), ARRB1(7), ARRB2(3), ATP1A4(24), ATP1B1(3), ATP1B2(7), ATP1B3(4), ATP2A2(9), ATP2A3(12), ATP2B1(17), ATP2B2(12), ATP2B3(21), CACNA1A(19), CACNA1B(1), CACNA1C(30), CACNA1D(34), CACNA1E(47), CACNA1S(33), CACNB1(9), CACNB3(6), CALM2(3), CALM3(1), CALR(4), CAMK1(9), CAMK2A(8), CAMK2B(2), CAMK2D(7), CAMK2G(8), CAMK4(5), CASQ1(7), CASQ2(8), CHRM1(6), CHRM2(9), CHRM3(9), CHRM4(9), CHRM5(13), GJA1(6), GJA4(2), GJB1(1), GJB2(3), GJB3(2), GJB4(5), GJB5(3), GJB6(4), GNA11(3), GNAI2(7), GNAI3(3), GNAO1(5), GNAZ(7), GNB1(3), GNB2(5), GNB3(4), GNB4(3), GNB5(3), GNG12(2), GNG13(2), GNG2(2), GNG3(3), GRK4(14), GRK5(10), GRK6(3), ITPR1(31), ITPR2(38), ITPR3(23), KCNB1(17), KCNJ3(15), KCNJ5(6), MIB1(8), MYCBP(1), NME7(4), PEA15(2), PKIG(2), PLCB3(9), PLN(1), PRKACA(4), PRKACB(2), PRKAR1A(4), PRKAR2A(4), PRKAR2B(5), PRKCA(10), PRKCD(7), PRKCE(11), PRKCG(10), PRKCH(9), PRKCQ(13), PRKCZ(4), PRKD1(8), RGS1(6), RGS10(3), RGS14(5), RGS16(3), RGS17(5), RGS18(2), RGS2(3), RGS20(5), RGS3(15), RGS4(6), RGS5(4), RGS6(7), RGS7(9), RGS9(19), RYR1(80), RYR2(111), RYR3(87), SLC8A1(20), SLC8A3(14), USP5(13), YWHAB(1), YWHAH(4), YWHAQ(3) | 65550530 | 1346 | 138 | 1309 | 463 | 395 | 564 | 151 | 72 | 145 | 19 | 3.4e-11 | 1e-08 | |
| 3 | HSA04080_NEUROACTIVE_LIGAND_RECEPTOR_INTERACTION | Genes involved in neuroactive ligand-receptor interaction | ADCYAP1R1, ADORA1, ADORA2A, ADORA2B, ADORA3, ADRA1A, ADRA1B, ADRA2A, ADRA2B, ADRA2C, ADRB1, ADRB2, ADRB3, AGTR1, AGTR2, AGTRL1, AVPR1A, AVPR1B, AVPR2, BDKRB1, BDKRB2, BRS3, C3AR1, C5AR1, CALCR, CALCRL, CCKAR, CCKBR, CGA, CHRM1, CHRM2, CHRM3, CHRM4, CHRM5, CNR1, CNR2, CRHR1, CRHR2, CTSG, CYSLTR1, CYSLTR2, DRD1, DRD2, DRD3, DRD4, DRD5, EDG1, EDG2, EDG3, EDG4, EDG5, EDG6, EDG7, EDG8, EDNRA, EDNRB, F2, F2R, F2RL1, F2RL2, F2RL3, FPR1, FPRL1, FPRL2, FSHB, FSHR, GABBR1, GABBR2, GABRA1, GABRA2, GABRA3, GABRA4, GABRA5, GABRA6, GABRB1, GABRB2, GABRB3, GABRD, GABRE, GABRG1, GABRG2, GABRG3, GABRP, GABRQ, GABRR1, GABRR2, GALR1, GALR2, GALR3, GCGR, GH1, GH2, GHR, GHRHR, GHSR, GIPR, GLP1R, GLP2R, GLRA1, GLRA2, GLRA3, GLRB, GNRHR, GPR156, GPR23, GPR35, GPR50, GPR63, GPR83, GRIA1, GRIA2, GRIA3, GRIA4, GRID1, GRID2, GRIK1, GRIK2, GRIK3, GRIK4, GRIK5, GRIN1, GRIN2A, GRIN2B, GRIN2C, GRIN2D, GRIN3A, GRIN3B, GRM1, GRM2, GRM3, GRM4, GRM5, GRM6, GRM7, GRM8, GRPR, GZMA, HCRTR1, HCRTR2, HRH1, HRH2, HRH3, HRH4, HTR1A, HTR1B, HTR1D, HTR1E, HTR1F, HTR2A, HTR2B, HTR2C, HTR4, HTR5A, HTR6, HTR7, KISS1R, LEP, LEPR, LHB, LHCGR, LTB4R, LTB4R2, MAS1, MC1R, MC2R, MC3R, MC4R, MC5R, MCHR1, MCHR2, MLNR, MTNR1A, MTNR1B, NMBR, NMUR1, NMUR2, NPBWR1, NPBWR2, NPFFR1, NPFFR2, NPY1R, NPY2R, NPY5R, NR3C1, NTSR1, NTSR2, OPRD1, OPRK1, OPRL1, OPRM1, OXTR, P2RX1, P2RX2, P2RX3, P2RX4, P2RX5, P2RX7, P2RXL1, P2RY1, P2RY10, P2RY11, P2RY13, P2RY14, P2RY2, P2RY4, P2RY5, P2RY6, P2RY8, PARD3, PPYR1, PRL, PRLHR, PRLR, PRSS1, PRSS2, PRSS3, PTAFR, PTGDR, PTGER1, PTGER2, PTGER3, PTGER4, PTGFR, PTGIR, PTH2R, PTHR1, RXFP1, RXFP2, SCTR, SSTR1, SSTR2, SSTR3, SSTR4, SSTR5, TAAR1, TAAR2, TAAR5, TAAR6, TAAR8, TAAR9, TACR1, TACR2, TACR3, TBXA2R, THRA, THRB, TRHR, TRPV1, TSHB, TSHR, TSPO, UTS2R, VIPR1, VIPR2 | 214 | ADCYAP1R1(7), ADORA1(6), ADORA2A(5), ADORA2B(2), ADORA3(5), ADRA1A(11), ADRA1B(7), ADRA2B(5), ADRB1(2), ADRB2(8), AGTR2(8), AVPR1A(8), AVPR1B(3), AVPR2(5), BDKRB1(5), BDKRB2(9), BRS3(13), C3AR1(7), C5AR1(5), CALCR(5), CALCRL(9), CCKAR(12), CCKBR(7), CGA(1), CHRM1(6), CHRM2(9), CHRM3(9), CHRM4(9), CHRM5(13), CNR1(8), CNR2(3), CRHR1(2), CRHR2(2), CTSG(3), CYSLTR1(5), CYSLTR2(6), DRD1(6), DRD2(10), DRD3(5), DRD5(11), EDNRA(6), EDNRB(2), F2(9), F2R(8), F2RL1(3), F2RL2(3), FPR1(4), FSHR(17), GABBR1(17), GABBR2(14), GABRA1(10), GABRA2(12), GABRA3(15), GABRA4(17), GABRA5(8), GABRA6(8), GABRB1(8), GABRB2(16), GABRB3(15), GABRE(7), GABRG1(8), GABRG2(19), GABRG3(14), GABRP(5), GABRQ(7), GABRR1(5), GABRR2(2), GALR1(2), GALR3(3), GH1(6), GH2(5), GHR(10), GHRHR(2), GHSR(7), GIPR(4), GLP1R(8), GLP2R(17), GLRA1(9), GLRA2(6), GLRA3(11), GLRB(8), GNRHR(4), GPR156(6), GPR50(8), GPR63(5), GPR83(5), GRIA1(12), GRIA2(25), GRIA4(23), GRID1(16), GRID2(13), GRIK1(16), GRIK2(15), GRIK3(17), GRIK4(8), GRIK5(19), GRIN1(5), GRIN2A(25), GRIN2B(23), GRIN2C(6), GRIN2D(8), GRIN3A(15), GRM1(22), GRM2(7), GRM3(20), GRM4(16), GRM5(22), GRM7(15), GRM8(24), GRPR(4), GZMA(6), HCRTR1(1), HCRTR2(8), HRH1(9), HRH2(3), HRH4(4), HTR1A(13), HTR1B(6), HTR1D(6), HTR1E(6), HTR2A(15), HTR2B(2), HTR4(7), HTR5A(9), HTR6(4), HTR7(4), LEP(2), LEPR(15), LHB(3), LHCGR(16), LTB4R(3), LTB4R2(3), MAS1(2), MC2R(2), MC3R(8), MC4R(7), MC5R(12), MCHR1(7), MCHR2(1), MLNR(3), MTNR1A(6), MTNR1B(4), NMBR(4), NMUR1(8), NMUR2(7), NPBWR1(2), NPFFR1(2), NPFFR2(12), NPY1R(6), NPY2R(4), NPY5R(10), NR3C1(13), NTSR1(4), NTSR2(2), OPRD1(2), OPRK1(9), OPRM1(11), OXTR(4), P2RX1(9), P2RX3(9), P2RX4(2), P2RX5(8), P2RX7(3), P2RY1(2), P2RY10(6), P2RY11(7), P2RY13(3), P2RY14(5), P2RY2(6), P2RY4(7), P2RY6(3), PARD3(27), PPYR1(7), PRL(6), PRLHR(2), PRLR(6), PRSS1(7), PRSS3(2), PTAFR(6), PTGDR(6), PTGER2(5), PTGER3(9), PTGER4(8), PTGFR(10), PTGIR(3), PTH2R(7), RXFP1(16), RXFP2(17), SCTR(4), SSTR2(3), SSTR3(6), TAAR1(6), TAAR2(5), TAAR5(8), TAAR6(5), TAAR8(3), TAAR9(8), TACR1(6), TACR2(4), TACR3(9), THRA(5), THRB(4), TRHR(5), TRPV1(6), TSHB(3), TSHR(12), VIPR1(3) | 71058285 | 1639 | 145 | 1582 | 539 | 480 | 673 | 258 | 73 | 153 | 2 | 4.3e-10 | 8.8e-08 |
| 4 | HSA04010_MAPK_SIGNALING_PATHWAY | Genes involved in MAPK signaling pathway | ACVR1B, ACVR1C, AKT1, AKT2, AKT3, ARRB1, ARRB2, ATF2, ATF4, BDNF, BRAF, CACNA1A, CACNA1B, CACNA1C, CACNA1D, CACNA1E, CACNA1F, CACNA1G, CACNA1H, CACNA1I, CACNA1S, CACNA2D1, CACNA2D2, CACNA2D3, CACNA2D4, CACNB1, CACNB2, CACNB3, CACNB4, CACNG1, CACNG2, CACNG3, CACNG4, CACNG5, CACNG6, CACNG7, CACNG8, CASP3, CD14, CDC25B, CDC42, CHP, CHUK, CRK, CRKL, DAXX, DDIT3, DUSP1, DUSP10, DUSP14, DUSP16, DUSP2, DUSP3, DUSP4, DUSP5, DUSP6, DUSP7, DUSP8, DUSP9, ECSIT, EGF, EGFR, ELK1, ELK4, EVI1, FAS, FASLG, FGF1, FGF10, FGF11, FGF12, FGF13, FGF14, FGF16, FGF17, FGF18, FGF19, FGF2, FGF20, FGF21, FGF22, FGF23, FGF3, FGF4, FGF5, FGF6, FGF7, FGF8, FGF9, FGFR1, FGFR2, FGFR3, FGFR4, FLNA, FLNB, FLNC, FOS, GADD45A, GADD45B, GADD45G, GNA12, GNG12, GRB2, HRAS, IKBKB, IKBKG, IL1A, IL1B, IL1R1, IL1R2, JUN, JUND, KRAS, LOC653852, MAP2K1, MAP2K1IP1, MAP2K2, MAP2K3, MAP2K4, MAP2K5, MAP2K6, MAP2K7, MAP3K1, MAP3K10, MAP3K12, MAP3K13, MAP3K14, MAP3K2, MAP3K3, MAP3K4, MAP3K5, MAP3K6, MAP3K7, MAP3K7IP1, MAP3K7IP2, MAP3K8, MAP4K1, MAP4K2, MAP4K3, MAP4K4, MAPK1, MAPK10, MAPK11, MAPK12, MAPK13, MAPK14, MAPK3, MAPK7, MAPK8, MAPK8IP1, MAPK8IP2, MAPK8IP3, MAPK9, MAPKAPK2, MAPKAPK3, MAPKAPK5, MAPT, MAX, MEF2C, MKNK1, MKNK2, MOS, MRAS, MYC, NF1, NFATC2, NFATC4, NFKB1, NFKB2, NGFB, NLK, NR4A1, NRAS, NTF3, NTF5, NTRK1, NTRK2, PAK1, PAK2, PDGFA, PDGFB, PDGFRA, PDGFRB, PLA2G10, PLA2G12A, PLA2G12B, PLA2G1B, PLA2G2A, PLA2G2D, PLA2G2E, PLA2G2F, PLA2G3, PLA2G4A, PLA2G5, PLA2G6, PPM1A, PPM1B, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PPP5C, PRKACA, PRKACB, PRKACG, PRKCA, PRKCB1, PRKCG, PRKX, PRKY, PTPN5, PTPN7, PTPRR, RAC1, RAC2, RAC3, RAF1, RAP1A, RAP1B, RAPGEF2, RASA1, RASA2, RASGRF1, RASGRF2, RASGRP1, RASGRP2, RASGRP3, RASGRP4, RPS6KA1, RPS6KA2, RPS6KA3, RPS6KA4, RPS6KA5, RPS6KA6, RRAS, RRAS2, SOS1, SOS2, SRF, STK3, STK4, STMN1, TAOK1, TAOK2, TAOK3, TGFB1, TGFB2, TGFB3, TGFBR1, TGFBR2, TNF, TNFRSF1A, TP53, TRAF2, TRAF6, ZAK | 225 | ACVR1B(6), ACVR1C(7), AKT1(8), AKT2(4), AKT3(8), ARRB1(7), ARRB2(3), ATF2(6), ATF4(4), BDNF(8), BRAF(12), CACNA1A(19), CACNA1B(1), CACNA1C(30), CACNA1D(34), CACNA1E(47), CACNA1F(36), CACNA1G(20), CACNA1I(13), CACNA1S(33), CACNA2D1(14), CACNA2D2(9), CACNA2D3(20), CACNA2D4(18), CACNB1(9), CACNB2(11), CACNB3(6), CACNB4(14), CACNG2(3), CACNG3(9), CACNG4(6), CACNG5(4), CACNG6(6), CACNG7(10), CASP3(4), CD14(2), CDC25B(3), CDC42(5), CHUK(11), CRK(1), CRKL(5), DAXX(11), DDIT3(1), DUSP1(2), DUSP10(7), DUSP14(3), DUSP16(13), DUSP2(4), DUSP3(2), DUSP4(1), DUSP5(5), DUSP6(1), DUSP7(4), DUSP9(8), ECSIT(5), EGF(24), EGFR(20), ELK1(2), ELK4(3), FAS(7), FASLG(6), FGF1(1), FGF10(1), FGF11(2), FGF12(4), FGF13(6), FGF14(3), FGF16(1), FGF17(1), FGF18(2), FGF21(3), FGF23(6), FGF5(8), FGF6(2), FGF7(6), FGF8(2), FGF9(2), FGFR1(10), FGFR4(11), FLNA(31), FLNB(38), FLNC(34), FOS(6), GNA12(10), GNG12(2), GRB2(3), IKBKB(15), IL1A(1), IL1B(2), IL1R1(5), IL1R2(7), JUN(1), MAP2K1(2), MAP2K3(8), MAP2K4(6), MAP2K5(2), MAP2K6(9), MAP3K10(7), MAP3K12(8), MAP3K13(13), MAP3K14(12), MAP3K2(8), MAP3K3(9), MAP3K4(23), MAP3K5(21), MAP3K6(7), MAP3K7(8), MAP3K8(12), MAP4K1(13), MAP4K2(6), MAP4K3(21), MAP4K4(14), MAPK1(2), MAPK10(9), MAPK13(2), MAPK14(6), MAPK3(4), MAPK7(17), MAPK8(7), MAPK8IP1(5), MAPK9(7), MAPKAPK2(6), MAPKAPK3(4), MAPKAPK5(6), MAPT(11), MAX(8), MEF2C(11), MKNK1(6), MKNK2(4), MOS(8), MRAS(3), MYC(7), NF1(40), NFATC2(15), NFATC4(8), NFKB1(12), NFKB2(10), NLK(5), NR4A1(4), NTF3(6), NTRK1(11), NTRK2(15), PAK1(3), PAK2(6), PDGFB(4), PDGFRA(29), PDGFRB(12), PLA2G10(1), PLA2G12A(1), PLA2G12B(3), PLA2G1B(1), PLA2G2D(2), PLA2G2E(3), PLA2G2F(3), PLA2G3(5), PLA2G4A(11), PLA2G5(2), PLA2G6(6), PPM1A(7), PPM1B(6), PPP3CA(4), PPP3CB(7), PPP3CC(6), PPP3R1(2), PPP3R2(1), PPP5C(7), PRKACA(4), PRKACB(2), PRKACG(6), PRKCA(10), PRKCG(10), PRKX(4), PTPN5(5), PTPN7(5), PTPRR(18), RAC2(2), RAF1(7), RAP1A(3), RAP1B(2), RAPGEF2(38), RASA2(8), RASGRF1(13), RASGRF2(17), RASGRP1(8), RASGRP2(5), RASGRP3(8), RASGRP4(7), RPS6KA1(5), RPS6KA2(16), RPS6KA3(17), RPS6KA4(5), RPS6KA5(7), RPS6KA6(15), RRAS(1), RRAS2(5), SOS1(21), SOS2(22), SRF(3), STK3(9), STK4(6), STMN1(6), TAOK1(10), TAOK2(10), TAOK3(4), TGFB1(4), TGFB2(6), TGFB3(9), TGFBR1(8), TGFBR2(6), TNF(1), TNFRSF1A(6), TRAF2(5), TRAF6(4), ZAK(12) | 92476270 | 1836 | 141 | 1785 | 566 | 501 | 733 | 230 | 111 | 239 | 22 | 1.1e-08 | 1.8e-06 |
| 5 | HSA04530_TIGHT_JUNCTION | Genes involved in tight junction | ACTB, ACTG1, ACTN1, ACTN2, ACTN3, ACTN4, AKT1, AKT2, AKT3, AMOTL1, ASH1L, CASK, CDC42, CDK4, CGN, CLDN1, CLDN10, CLDN11, CLDN14, CLDN15, CLDN16, CLDN17, CLDN18, CLDN19, CLDN2, CLDN20, CLDN22, CLDN23, CLDN3, CLDN4, CLDN5, CLDN6, CLDN7, CLDN8, CLDN9, CRB3, CSDA, CSNK2A1, CSNK2A2, CSNK2B, CTNNA1, CTNNA2, CTNNA3, CTNNB1, CTTN, EPB41, EPB41L1, EPB41L2, EPB41L3, EXOC3, EXOC4, F11R, GNAI1, GNAI2, GNAI3, HCLS1, HRAS, IGSF5, INADL, JAM2, JAM3, KRAS, LLGL1, LLGL2, MAGI1, MAGI2, MAGI3, MLLT4, MPDZ, MPP5, MRAS, MRCL3, MRLC2, MYH1, MYH10, MYH11, MYH13, MYH14, MYH15, MYH2, MYH3, MYH4, MYH6, MYH7, MYH7B, MYH8, MYH9, MYL2, MYL5, MYL7, MYL8P, MYL9, MYLC2PL, MYLPF, NRAS, OCLN, PARD3, PARD6A, PARD6B, PARD6G, PPM1J, PPP2CA, PPP2CB, PPP2R1A, PPP2R1B, PPP2R2A, PPP2R2B, PPP2R2C, PPP2R3A, PPP2R3B, PPP2R4, PRKCA, PRKCB1, PRKCD, PRKCE, PRKCG, PRKCH, PRKCI, PRKCQ, PRKCZ, PTEN, RAB13, RAB3B, RHOA, RRAS, RRAS2, SPTAN1, SRC, SYMPK, TJAP1, TJP1, TJP2, TJP3, VAPA, YES1, ZAK | 121 | ACTB(5), ACTN1(13), ACTN2(9), ACTN3(10), ACTN4(11), AKT1(8), AKT2(4), AKT3(8), AMOTL1(14), ASH1L(40), CASK(20), CDC42(5), CDK4(4), CGN(15), CLDN1(2), CLDN10(9), CLDN11(1), CLDN15(4), CLDN16(6), CLDN17(2), CLDN18(4), CLDN19(2), CLDN2(4), CLDN20(4), CLDN22(3), CLDN6(1), CLDN7(1), CLDN8(5), CLDN9(2), CRB3(1), CSDA(6), CSNK2A1(9), CSNK2A2(5), CTNNA1(18), CTNNA2(13), CTNNA3(13), CTTN(8), EPB41(11), EPB41L1(11), EPB41L2(13), EPB41L3(18), EXOC3(12), EXOC4(4), F11R(2), GNAI1(4), GNAI2(7), GNAI3(3), HCLS1(8), IGSF5(9), INADL(17), JAM2(9), JAM3(3), LLGL1(14), LLGL2(5), MAGI1(32), MAGI2(26), MAGI3(17), MLLT4(32), MPDZ(22), MPP5(5), MRAS(3), MYH1(44), MYH10(29), MYH11(39), MYH13(27), MYH14(22), MYH15(22), MYH2(37), MYH3(30), MYH4(36), MYH6(23), MYH7(28), MYH7B(15), MYH8(35), MYH9(26), MYL2(1), MYL5(3), MYL7(4), MYL9(3), MYLPF(2), OCLN(5), PARD3(27), PARD6A(1), PARD6B(3), PPM1J(3), PPP2CA(6), PPP2CB(2), PPP2R1B(9), PPP2R2A(15), PPP2R2B(12), PPP2R2C(3), PPP2R3A(13), PPP2R4(7), PRKCA(10), PRKCD(7), PRKCE(11), PRKCG(10), PRKCH(9), PRKCI(8), PRKCQ(13), PRKCZ(4), RAB13(3), RAB3B(1), RHOA(4), RRAS(1), RRAS2(5), SPTAN1(29), SRC(2), SYMPK(20), TJAP1(8), TJP1(33), TJP2(16), TJP3(6), VAPA(5), YES1(5), ZAK(12) | 64754182 | 1330 | 139 | 1292 | 409 | 366 | 531 | 194 | 61 | 161 | 17 | 3.6e-08 | 4.4e-06 |
| 6 | HSA04910_INSULIN_SIGNALING_PATHWAY | Genes involved in insulin signaling pathway | ACACA, ACACB, AKT1, AKT2, AKT3, ARAF, BAD, BRAF, CALM1, CALM2, CALM3, CALML3, CALML6, CBL, CBLB, CBLC, CRK, CRKL, EIF4EBP1, ELK1, EXOC7, FASN, FBP1, FBP2, FLOT1, FLOT2, FOXO1, FRAP1, G6PC, G6PC2, GCK, GRB2, GSK3B, GYS1, GYS2, HRAS, IKBKB, INPP5D, INS, INSR, IRS1, IRS2, IRS4, KIAA1303, KRAS, LIPE, MAP2K1, MAP2K2, MAPK1, MAPK10, MAPK3, MAPK8, MAPK9, MKNK1, MKNK2, NRAS, PCK1, PCK2, PDE3A, PDE3B, PDPK1, PFKL, PFKM, PFKP, PHKA1, PHKA2, PHKB, PHKG1, PHKG2, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PKLR, PKM2, PPARGC1A, PPP1CA, PPP1CB, PPP1CC, PPP1R3A, PPP1R3B, PPP1R3C, PPP1R3D, PRKAA1, PRKAA2, PRKAB1, PRKAB2, PRKACA, PRKACB, PRKACG, PRKAG1, PRKAG2, PRKAG3, PRKAR1A, PRKAR1B, PRKAR2A, PRKAR2B, PRKCI, PRKCZ, PRKX, PRKY, PTPN1, PTPRF, PYGB, PYGL, PYGM, RAF1, RAPGEF1, RHEB, RHOQ, RPS6, RPS6KB1, RPS6KB2, SH2B2, SHC1, SHC2, SHC3, SHC4, SKIP, SLC2A4, SOCS1, SOCS2, SOCS3, SOCS4, SORBS1, SOS1, SOS2, SREBF1, TRIP10, TSC1, TSC2 | 118 | ACACA(27), ACACB(31), AKT1(8), AKT2(4), AKT3(8), ARAF(8), BAD(2), BRAF(12), CALM2(3), CALM3(1), CALML6(4), CBL(10), CBLB(15), CBLC(5), CRK(1), CRKL(5), ELK1(2), EXOC7(8), FBP1(4), FLOT1(1), FLOT2(3), FOXO1(11), G6PC(6), G6PC2(9), GCK(5), GRB2(3), GSK3B(11), GYS1(12), GYS2(19), IKBKB(15), INPP5D(14), INSR(18), IRS1(17), IRS4(20), LIPE(7), MAP2K1(2), MAPK1(2), MAPK10(9), MAPK3(4), MAPK8(7), MAPK9(7), MKNK1(6), MKNK2(4), PCK1(15), PCK2(7), PDE3A(17), PDE3B(15), PDPK1(3), PFKM(7), PFKP(11), PHKA1(17), PHKA2(15), PHKB(14), PHKG1(6), PHKG2(3), PIK3CB(19), PIK3CD(10), PIK3CG(25), PIK3R2(4), PIK3R3(8), PIK3R5(8), PKLR(10), PKM2(5), PPARGC1A(13), PPP1CA(6), PPP1CB(7), PPP1CC(2), PPP1R3A(23), PPP1R3B(3), PPP1R3C(3), PRKAA1(6), PRKAA2(9), PRKAB1(7), PRKAB2(2), PRKACA(4), PRKACB(2), PRKACG(6), PRKAG1(2), PRKAG2(4), PRKAG3(6), PRKAR1A(4), PRKAR2A(4), PRKAR2B(5), PRKCI(8), PRKCZ(4), PRKX(4), PTPN1(4), PTPRF(34), PYGB(10), PYGL(9), PYGM(10), RAF1(7), RAPGEF1(18), RHEB(2), RHOQ(1), RPS6(6), RPS6KB1(3), RPS6KB2(6), SH2B2(2), SHC1(11), SHC3(9), SHC4(7), SLC2A4(4), SOCS2(7), SOCS4(4), SORBS1(22), SOS1(21), SOS2(22), SREBF1(6), TRIP10(6), TSC1(13), TSC2(14) | 49120714 | 981 | 128 | 952 | 281 | 250 | 412 | 126 | 43 | 138 | 12 | 4.8e-06 | 0.0005 |
| 7 | ERK5PATHWAY | Signaling between a tissue and its innervating axon stimulates retrograde transport via Trk receptors, which activate Erk5, which induces transcription of anti-apoptotic factors. | AKT1, CREB1, GRB2, HRAS, MAPK1, MAPK3, MAPK7, MEF2A, MEF2B, MEF2C, MEF2D, NTRK1, PIK3CA, PIK3R1, PLCG1, RPS6KA1, SHC1 | 15 | AKT1(8), CREB1(5), GRB2(3), MAPK1(2), MAPK3(4), MAPK7(17), MEF2A(5), MEF2B(3), MEF2C(11), MEF2D(7), NTRK1(11), PLCG1(12), RPS6KA1(5), SHC1(11) | 5187495 | 104 | 52 | 96 | 30 | 23 | 51 | 9 | 5 | 16 | 0 | 6.6e-06 | 0.00058 |
| 8 | WNTPATHWAY | The Wnt glycoprotein binds to membrane-bound receptors such as Frizzled to activate a number of signaling pathways, including that of beta-catenin. | APC, AXIN1, BTRC, CCND1, CREBBP, CSNK1A1, CSNK1D, CSNK2A1, CTBP1, CTNNB1, DVL1, FRAT1, FZD1, GSK3B, HDAC1, MADH4, MAP3K7, MAP3K7IP1, MYC, NLK, PPARD, PPP2CA, TCF1, TLE1, WIF1, WNT1 | 17 | AXIN1(8), BTRC(13), CREBBP(37), CSNK1A1(5), CSNK2A1(9), FZD1(2), GSK3B(11), HDAC1(4), MAP3K7(8), MYC(7), NLK(5), PPARD(6), PPP2CA(6), TLE1(12), WIF1(5), WNT1(1) | 6567242 | 139 | 59 | 136 | 28 | 43 | 56 | 17 | 3 | 18 | 2 | 0.000018 | 0.0014 |
| 9 | PITX2PATHWAY | The bicoid-related transcription factor Pitx2 is activated by Wnt binding to the Frizzled receptor and induces tissue-specific cell proliferation. | APC, AXIN1, CREBBP, CTNNB1, DVL1, EP300, FZD1, GSK3B, HDAC1, HTATIP, LDB1, LEF1, PITX2, PPARBP, TRRAP, WNT1 | 10 | AXIN1(8), CREBBP(37), EP300(29), FZD1(2), GSK3B(11), HDAC1(4), LDB1(3), LEF1(9), TRRAP(72), WNT1(1) | 8072942 | 176 | 62 | 170 | 33 | 49 | 68 | 23 | 6 | 28 | 2 | 0.000028 | 0.0019 |
| 10 | TGFBPATHWAY | The TGF-beta receptor responds to ligand binding by activating the SMAD family of transcriptional regulations, commonly blocking cell growth. | APC, CDH1, CREBBP, EP300, MADH2, MADH3, MADH4, MADH7, MADHIP, MAP2K1, MAP3K7, MAP3K7IP1, MAPK3, SKIL, TGFB1, TGFB2, TGFB3, TGFBR1, TGFBR2 | 12 | CDH1(15), CREBBP(37), EP300(29), MAP2K1(2), MAP3K7(8), MAPK3(4), SKIL(13), TGFB1(4), TGFB2(6), TGFB3(9), TGFBR1(8), TGFBR2(6) | 6902081 | 141 | 57 | 136 | 37 | 35 | 58 | 20 | 6 | 22 | 0 | 0.000054 | 0.0032 |
In brief, we tabulate the number of mutations and the number of covered bases for each gene. The counts are broken down by mutation context category: four context categories that are discovered by MutSig, and one for indel and 'null' mutations, which include indels, nonsense mutations, splice-site mutations, and non-stop (read-through) mutations. For each gene, we calculate the probability of seeing the observed constellation of mutations, i.e. the product P1 x P2 x ... x Pm, or a more extreme one, given the background mutation rates calculated across the dataset.[1]
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.