This pipeline computes the correlation between significant arm-level copy number variations (cnvs) and subtypes.
Testing the association between copy number variation 45 arm-level results and 8 molecular subtypes across 497 patients, 102 significant findings detected with Q value < 0.25.
-
1q gain cnv correlated to 'CN_CNMF'.
-
4p gain cnv correlated to 'CN_CNMF'.
-
4q gain cnv correlated to 'CN_CNMF'.
-
5p gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
5q gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
7p gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
7q gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
11q gain cnv correlated to 'CN_CNMF'.
-
12p gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
12q gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
13q gain cnv correlated to 'CN_CNMF'.
-
14q gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
16p gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
16q gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
17p gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
17q gain cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
19p gain cnv correlated to 'CN_CNMF'.
-
19q gain cnv correlated to 'CN_CNMF'.
-
20p gain cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
20q gain cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
1p loss cnv correlated to 'CN_CNMF'.
-
2p loss cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
2q loss cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
3q loss cnv correlated to 'CN_CNMF'.
-
8p loss cnv correlated to 'CN_CNMF'.
-
8q loss cnv correlated to 'CN_CNMF'.
-
9p loss cnv correlated to 'CN_CNMF'.
-
9q loss cnv correlated to 'CN_CNMF'.
-
10p loss cnv correlated to 'CN_CNMF'.
-
10q loss cnv correlated to 'CN_CNMF'.
-
11p loss cnv correlated to 'CN_CNMF'.
-
11q loss cnv correlated to 'CN_CNMF'.
-
15q loss cnv correlated to 'CN_CNMF', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
18p loss cnv correlated to 'CN_CNMF'.
-
18q loss cnv correlated to 'CN_CNMF'.
-
21q loss cnv correlated to 'CN_CNMF'.
-
22q loss cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 45 arm-level results and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 102 significant findings detected.
|
Molecular subtypes |
CN CNMF |
METHLYATION CNMF |
RPPA CNMF |
RPPA CHIERARCHICAL |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| 7q gain | 0 (0%) | 478 |
1.76e-23 (5.65e-21) |
3.64e-05 (0.00969) |
0.689 (1.00) |
0.00298 (0.597) |
4.87e-05 (0.0128) |
6.26e-06 (0.00177) |
0.000289 (0.0691) |
2.87e-05 (0.00773) |
| 12p gain | 0 (0%) | 486 |
1.03e-12 (3.22e-10) |
3.14e-06 (0.000912) |
0.901 (1.00) |
0.0299 (1.00) |
1.59e-05 (0.00446) |
3.43e-06 (0.000987) |
5.19e-05 (0.0136) |
1.7e-05 (0.00472) |
| 12q gain | 0 (0%) | 485 |
1.27e-12 (3.98e-10) |
6.85e-07 (0.000204) |
0.901 (1.00) |
0.0299 (1.00) |
3.88e-06 (0.00111) |
7.94e-07 (0.000235) |
1.64e-05 (0.00459) |
4.85e-06 (0.00138) |
| 16p gain | 0 (0%) | 485 |
5.51e-14 (1.75e-11) |
2.82e-05 (0.0076) |
1 (1.00) |
0.0445 (1.00) |
0.000128 (0.0323) |
3.37e-05 (0.00899) |
0.000214 (0.052) |
5.88e-05 (0.0152) |
| 17p gain | 0 (0%) | 485 |
2e-09 (6.13e-07) |
6.85e-07 (0.000204) |
0.772 (1.00) |
0.0179 (1.00) |
3.88e-06 (0.00111) |
7.94e-07 (0.000235) |
5.19e-05 (0.0136) |
1.7e-05 (0.00472) |
| 17q gain | 0 (0%) | 484 |
3.83e-09 (1.17e-06) |
1.58e-07 (4.73e-05) |
0.795 (1.00) |
0.0206 (1.00) |
1.36e-06 (0.000398) |
1.92e-07 (5.74e-05) |
1.64e-05 (0.00459) |
4.85e-06 (0.00138) |
| 2p loss | 0 (0%) | 489 |
5.02e-09 (1.53e-06) |
8.94e-05 (0.0229) |
1 (1.00) |
0.0817 (1.00) |
0.000496 (0.114) |
0.000164 (0.041) |
5.53e-05 (0.0144) |
2.27e-05 (0.00615) |
| 22q loss | 0 (0%) | 431 |
6.79e-58 (2.19e-55) |
0.00013 (0.0328) |
0.114 (1.00) |
0.0835 (1.00) |
8.4e-11 (2.6e-08) |
1.98e-10 (6.11e-08) |
4.33e-05 (0.0114) |
9.51e-06 (0.00268) |
| 5q gain | 0 (0%) | 483 |
2.6e-18 (8.27e-16) |
3.16e-05 (0.00847) |
0.901 (1.00) |
0.0299 (1.00) |
0.000163 (0.0408) |
4.26e-05 (0.0113) |
0.00146 (0.309) |
0.000106 (0.0269) |
| 7p gain | 0 (0%) | 480 |
1.59e-22 (5.09e-20) |
0.000218 (0.0529) |
1 (1.00) |
0.0129 (1.00) |
0.000389 (0.0906) |
6.9e-05 (0.0178) |
0.00232 (0.478) |
0.000272 (0.0652) |
| 2q loss | 0 (0%) | 490 |
3.78e-09 (1.16e-06) |
0.000378 (0.0884) |
0.837 (1.00) |
0.0549 (1.00) |
0.00144 (0.307) |
0.00065 (0.146) |
0.000192 (0.0475) |
8.85e-05 (0.0227) |
| 5p gain | 0 (0%) | 480 |
6.97e-20 (2.22e-17) |
0.000747 (0.166) |
1 (1.00) |
0.125 (1.00) |
0.00247 (0.499) |
0.000522 (0.119) |
0.00691 (1.00) |
0.000541 (0.123) |
| 14q gain | 0 (0%) | 489 |
5.02e-09 (1.53e-06) |
8.94e-05 (0.0229) |
1 (1.00) |
0.256 (1.00) |
0.000496 (0.114) |
0.000164 (0.041) |
0.00467 (0.906) |
0.00229 (0.475) |
| 16q gain | 0 (0%) | 487 |
5.67e-13 (1.79e-10) |
0.000247 (0.0595) |
0.856 (1.00) |
0.129 (1.00) |
0.00144 (0.308) |
0.000429 (0.0995) |
0.00169 (0.355) |
0.000547 (0.124) |
| 15q loss | 0 (0%) | 490 |
0.000464 (0.107) |
0.00241 (0.494) |
0.458 (1.00) |
0.256 (1.00) |
0.00491 (0.948) |
0.00175 (0.367) |
0.000664 (0.149) |
0.000342 (0.0805) |
| 20p gain | 0 (0%) | 488 |
1.11e-11 (3.44e-09) |
0.000767 (0.17) |
1 (1.00) |
0.0817 (1.00) |
0.00397 (0.789) |
0.00123 (0.27) |
0.0124 (1.00) |
0.00619 (1.00) |
| 20q gain | 0 (0%) | 488 |
1.11e-11 (3.44e-09) |
0.000767 (0.17) |
1 (1.00) |
0.0817 (1.00) |
0.00397 (0.789) |
0.00123 (0.27) |
0.0124 (1.00) |
0.00619 (1.00) |
| 1q gain | 0 (0%) | 476 |
3.29e-06 (0.000951) |
0.0812 (1.00) |
0.0153 (1.00) |
0.0826 (1.00) |
0.0148 (1.00) |
0.00133 (0.288) |
0.224 (1.00) |
0.204 (1.00) |
| 4p gain | 0 (0%) | 493 |
1.85e-05 (0.0051) |
0.0192 (1.00) |
0.0366 (1.00) |
0.0186 (1.00) |
0.112 (1.00) |
0.0975 (1.00) |
||
| 4q gain | 0 (0%) | 493 |
1.85e-05 (0.0051) |
0.0192 (1.00) |
0.0366 (1.00) |
0.0186 (1.00) |
0.112 (1.00) |
0.0975 (1.00) |
||
| 11q gain | 0 (0%) | 493 |
0.000204 (0.0503) |
0.0192 (1.00) |
0.0366 (1.00) |
0.0186 (1.00) |
0.00778 (1.00) |
0.00503 (0.965) |
||
| 13q gain | 0 (0%) | 494 |
0.000294 (0.07) |
0.0495 (1.00) |
0.101 (1.00) |
0.0769 (1.00) |
||||
| 19p gain | 0 (0%) | 493 |
1.85e-05 (0.0051) |
0.0192 (1.00) |
0.0366 (1.00) |
0.0186 (1.00) |
0.0264 (1.00) |
0.0191 (1.00) |
||
| 19q gain | 0 (0%) | 492 |
1.12e-06 (0.00033) |
0.00434 (0.855) |
0.636 (1.00) |
0.0551 (1.00) |
0.0147 (1.00) |
0.00587 (1.00) |
0.00778 (1.00) |
0.00503 (0.965) |
| 1p loss | 0 (0%) | 494 |
0.000294 (0.07) |
0.559 (1.00) |
0.495 (1.00) |
0.214 (1.00) |
0.3 (1.00) |
0.276 (1.00) |
||
| 3q loss | 0 (0%) | 494 |
0.000294 (0.07) |
0.0495 (1.00) |
0.101 (1.00) |
0.0769 (1.00) |
0.0264 (1.00) |
0.0191 (1.00) |
||
| 8p loss | 0 (0%) | 493 |
1.85e-05 (0.0051) |
0.047 (1.00) |
0.0366 (1.00) |
0.0186 (1.00) |
0.00778 (1.00) |
0.00503 (0.965) |
||
| 8q loss | 0 (0%) | 493 |
1.85e-05 (0.0051) |
0.047 (1.00) |
0.0366 (1.00) |
0.0186 (1.00) |
0.00778 (1.00) |
0.00503 (0.965) |
||
| 9p loss | 0 (0%) | 485 |
1.08e-11 (3.36e-09) |
0.272 (1.00) |
0.559 (1.00) |
0.206 (1.00) |
0.157 (1.00) |
0.206 (1.00) |
0.185 (1.00) |
0.159 (1.00) |
| 9q loss | 0 (0%) | 480 |
1.58e-13 (4.99e-11) |
0.204 (1.00) |
0.21 (1.00) |
0.00441 (0.86) |
0.268 (1.00) |
0.295 (1.00) |
0.235 (1.00) |
0.228 (1.00) |
| 10p loss | 0 (0%) | 491 |
6.62e-08 (2e-05) |
0.0979 (1.00) |
0.047 (1.00) |
0.0135 (1.00) |
0.0399 (1.00) |
0.0333 (1.00) |
||
| 10q loss | 0 (0%) | 492 |
1.12e-06 (0.00033) |
0.175 (1.00) |
0.102 (1.00) |
0.0405 (1.00) |
0.112 (1.00) |
0.0975 (1.00) |
||
| 11p loss | 0 (0%) | 490 |
7.74e-08 (2.33e-05) |
0.00947 (1.00) |
0.499 (1.00) |
1 (1.00) |
0.0148 (1.00) |
0.0419 (1.00) |
0.0399 (1.00) |
0.0333 (1.00) |
| 11q loss | 0 (0%) | 488 |
3.24e-06 (0.00094) |
0.0149 (1.00) |
0.388 (1.00) |
0.798 (1.00) |
0.044 (1.00) |
0.039 (1.00) |
0.0138 (1.00) |
0.0075 (1.00) |
| 18p loss | 0 (0%) | 493 |
0.000204 (0.0503) |
0.19 (1.00) |
1 (1.00) |
0.369 (1.00) |
0.22 (1.00) |
0.12 (1.00) |
0.0743 (1.00) |
0.0671 (1.00) |
| 18q loss | 0 (0%) | 493 |
0.000204 (0.0503) |
0.19 (1.00) |
1 (1.00) |
0.369 (1.00) |
0.22 (1.00) |
0.12 (1.00) |
0.0743 (1.00) |
0.0671 (1.00) |
| 21q loss | 0 (0%) | 491 |
0.000192 (0.0475) |
0.0979 (1.00) |
0.384 (1.00) |
0.0551 (1.00) |
0.0439 (1.00) |
0.112 (1.00) |
0.0399 (1.00) |
0.0333 (1.00) |
| 11p gain | 0 (0%) | 492 |
0.00134 (0.288) |
0.00434 (0.855) |
0.0147 (1.00) |
0.00587 (1.00) |
0.00228 (0.474) |
0.00132 (0.286) |
||
| 18p gain | 0 (0%) | 494 |
0.00245 (0.499) |
0.0495 (1.00) |
0.101 (1.00) |
0.0769 (1.00) |
||||
| 18q gain | 0 (0%) | 494 |
0.00245 (0.499) |
0.0495 (1.00) |
0.101 (1.00) |
0.0769 (1.00) |
||||
| 21q gain | 0 (0%) | 494 |
0.00245 (0.499) |
0.0495 (1.00) |
0.101 (1.00) |
0.0769 (1.00) |
0.0264 (1.00) |
0.0191 (1.00) |
||
| 6q loss | 0 (0%) | 494 |
0.0218 (1.00) |
0.0495 (1.00) |
0.101 (1.00) |
0.0769 (1.00) |
0.0264 (1.00) |
0.0191 (1.00) |
||
| 13q loss | 0 (0%) | 486 |
0.0123 (1.00) |
0.0182 (1.00) |
0.0779 (1.00) |
1 (1.00) |
0.0182 (1.00) |
0.00154 (0.325) |
0.00774 (1.00) |
0.0164 (1.00) |
| 17p loss | 0 (0%) | 490 |
0.00118 (0.26) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.79 (1.00) |
0.819 (1.00) |
0.518 (1.00) |
0.499 (1.00) |
| 19p loss | 0 (0%) | 493 |
0.132 (1.00) |
0.0192 (1.00) |
0.281 (1.00) |
0.369 (1.00) |
0.0366 (1.00) |
0.0186 (1.00) |
0.00778 (1.00) |
0.00503 (0.965) |
P value = 3.29e-06 (Fisher's exact test), Q value = 0.00095
Table S1. Gene #1: '1q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 1Q GAIN CNV | 9 | 11 | 1 |
| 1Q GAIN WILD-TYPE | 25 | 374 | 77 |
Figure S1. Get High-res Image Gene #1: '1q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.85e-05 (Fisher's exact test), Q value = 0.0051
Table S2. Gene #2: '4p gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 4P GAIN CNV | 4 | 0 | 0 |
| 4P GAIN WILD-TYPE | 30 | 385 | 78 |
Figure S2. Get High-res Image Gene #2: '4p gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.85e-05 (Fisher's exact test), Q value = 0.0051
Table S3. Gene #3: '4q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 4Q GAIN CNV | 4 | 0 | 0 |
| 4Q GAIN WILD-TYPE | 30 | 385 | 78 |
Figure S3. Get High-res Image Gene #3: '4q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 6.97e-20 (Fisher's exact test), Q value = 2.2e-17
Table S4. Gene #4: '5p gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 5P GAIN CNV | 16 | 1 | 0 |
| 5P GAIN WILD-TYPE | 18 | 384 | 78 |
Figure S4. Get High-res Image Gene #4: '5p gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000747 (Fisher's exact test), Q value = 0.17
Table S5. Gene #4: '5p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 5P GAIN CNV | 12 | 2 | 3 |
| 5P GAIN WILD-TYPE | 137 | 72 | 271 |
Figure S5. Get High-res Image Gene #4: '5p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.000522 (Fisher's exact test), Q value = 0.12
Table S6. Gene #4: '5p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 5P GAIN CNV | 12 | 5 | 0 | 0 |
| 5P GAIN WILD-TYPE | 111 | 186 | 53 | 91 |
Figure S6. Get High-res Image Gene #4: '5p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 0.000541 (Fisher's exact test), Q value = 0.12
Table S7. Gene #4: '5p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 5P GAIN CNV | 10 | 6 | 0 |
| 5P GAIN WILD-TYPE | 104 | 162 | 142 |
Figure S7. Get High-res Image Gene #4: '5p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 2.6e-18 (Fisher's exact test), Q value = 8.3e-16
Table S8. Gene #5: '5q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 5Q GAIN CNV | 14 | 0 | 0 |
| 5Q GAIN WILD-TYPE | 20 | 385 | 78 |
Figure S8. Get High-res Image Gene #5: '5q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 3.16e-05 (Fisher's exact test), Q value = 0.0085
Table S9. Gene #5: '5q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 5Q GAIN CNV | 12 | 1 | 1 |
| 5Q GAIN WILD-TYPE | 137 | 73 | 273 |
Figure S9. Get High-res Image Gene #5: '5q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.000163 (Fisher's exact test), Q value = 0.041
Table S10. Gene #5: '5q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 5Q GAIN CNV | 12 | 0 | 0 | 2 |
| 5Q GAIN WILD-TYPE | 128 | 56 | 106 | 154 |
Figure S10. Get High-res Image Gene #5: '5q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 4.26e-05 (Fisher's exact test), Q value = 0.011
Table S11. Gene #5: '5q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 5Q GAIN CNV | 12 | 2 | 0 | 0 |
| 5Q GAIN WILD-TYPE | 111 | 189 | 53 | 91 |
Figure S11. Get High-res Image Gene #5: '5q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 0.000106 (Fisher's exact test), Q value = 0.027
Table S12. Gene #5: '5q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 5Q GAIN CNV | 10 | 3 | 0 |
| 5Q GAIN WILD-TYPE | 104 | 165 | 142 |
Figure S12. Get High-res Image Gene #5: '5q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 1.59e-22 (Fisher's exact test), Q value = 5.1e-20
Table S13. Gene #6: '7p gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 7P GAIN CNV | 17 | 0 | 0 |
| 7P GAIN WILD-TYPE | 17 | 385 | 78 |
Figure S13. Get High-res Image Gene #6: '7p gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000218 (Fisher's exact test), Q value = 0.053
Table S14. Gene #6: '7p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 7P GAIN CNV | 12 | 3 | 2 |
| 7P GAIN WILD-TYPE | 137 | 71 | 272 |
Figure S14. Get High-res Image Gene #6: '7p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.000389 (Fisher's exact test), Q value = 0.091
Table S15. Gene #6: '7p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 7P GAIN CNV | 13 | 0 | 1 | 2 |
| 7P GAIN WILD-TYPE | 127 | 56 | 105 | 154 |
Figure S15. Get High-res Image Gene #6: '7p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 6.9e-05 (Fisher's exact test), Q value = 0.018
Table S16. Gene #6: '7p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 7P GAIN CNV | 13 | 2 | 0 | 1 |
| 7P GAIN WILD-TYPE | 110 | 189 | 53 | 90 |
Figure S16. Get High-res Image Gene #6: '7p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 0.000272 (Fisher's exact test), Q value = 0.065
Table S17. Gene #6: '7p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 7P GAIN CNV | 12 | 4 | 1 |
| 7P GAIN WILD-TYPE | 102 | 164 | 141 |
Figure S17. Get High-res Image Gene #6: '7p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 1.76e-23 (Fisher's exact test), Q value = 5.7e-21
Table S18. Gene #7: '7q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 7Q GAIN CNV | 18 | 0 | 1 |
| 7Q GAIN WILD-TYPE | 16 | 385 | 77 |
Figure S18. Get High-res Image Gene #7: '7q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 3.64e-05 (Fisher's exact test), Q value = 0.0097
Table S19. Gene #7: '7q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 7Q GAIN CNV | 14 | 3 | 2 |
| 7Q GAIN WILD-TYPE | 135 | 71 | 272 |
Figure S19. Get High-res Image Gene #7: '7q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 4.87e-05 (Fisher's exact test), Q value = 0.013
Table S20. Gene #7: '7q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 7Q GAIN CNV | 15 | 0 | 1 | 2 |
| 7Q GAIN WILD-TYPE | 125 | 56 | 105 | 154 |
Figure S20. Get High-res Image Gene #7: '7q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 6.26e-06 (Fisher's exact test), Q value = 0.0018
Table S21. Gene #7: '7q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 7Q GAIN CNV | 15 | 2 | 0 | 1 |
| 7Q GAIN WILD-TYPE | 108 | 189 | 53 | 90 |
Figure S21. Get High-res Image Gene #7: '7q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 0.000289 (Fisher's exact test), Q value = 0.069
Table S22. Gene #7: '7q gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 148 | 149 |
| 7Q GAIN CNV | 14 | 2 | 3 |
| 7Q GAIN WILD-TYPE | 113 | 146 | 146 |
Figure S22. Get High-res Image Gene #7: '7q gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'

P value = 2.87e-05 (Fisher's exact test), Q value = 0.0077
Table S23. Gene #7: '7q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 7Q GAIN CNV | 14 | 4 | 1 |
| 7Q GAIN WILD-TYPE | 100 | 164 | 141 |
Figure S23. Get High-res Image Gene #7: '7q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 0.000204 (Fisher's exact test), Q value = 0.05
Table S24. Gene #9: '11q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 11Q GAIN CNV | 3 | 0 | 1 |
| 11Q GAIN WILD-TYPE | 31 | 385 | 77 |
Figure S24. Get High-res Image Gene #9: '11q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.03e-12 (Fisher's exact test), Q value = 3.2e-10
Table S25. Gene #10: '12p gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 12P GAIN CNV | 10 | 0 | 1 |
| 12P GAIN WILD-TYPE | 24 | 385 | 77 |
Figure S25. Get High-res Image Gene #10: '12p gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 3.14e-06 (Fisher's exact test), Q value = 0.00091
Table S26. Gene #10: '12p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 12P GAIN CNV | 11 | 0 | 0 |
| 12P GAIN WILD-TYPE | 138 | 74 | 274 |
Figure S26. Get High-res Image Gene #10: '12p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1.59e-05 (Fisher's exact test), Q value = 0.0045
Table S27. Gene #10: '12p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 12P GAIN CNV | 11 | 0 | 0 | 0 |
| 12P GAIN WILD-TYPE | 129 | 56 | 106 | 156 |
Figure S27. Get High-res Image Gene #10: '12p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 3.43e-06 (Fisher's exact test), Q value = 0.00099
Table S28. Gene #10: '12p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 12P GAIN CNV | 11 | 0 | 0 | 0 |
| 12P GAIN WILD-TYPE | 112 | 191 | 53 | 91 |
Figure S28. Get High-res Image Gene #10: '12p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 5.19e-05 (Fisher's exact test), Q value = 0.014
Table S29. Gene #10: '12p gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 148 | 149 |
| 12P GAIN CNV | 10 | 0 | 1 |
| 12P GAIN WILD-TYPE | 117 | 148 | 148 |
Figure S29. Get High-res Image Gene #10: '12p gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'

P value = 1.7e-05 (Fisher's exact test), Q value = 0.0047
Table S30. Gene #10: '12p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 12P GAIN CNV | 10 | 1 | 0 |
| 12P GAIN WILD-TYPE | 104 | 167 | 142 |
Figure S30. Get High-res Image Gene #10: '12p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 1.27e-12 (Fisher's exact test), Q value = 4e-10
Table S31. Gene #11: '12q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 12Q GAIN CNV | 10 | 0 | 2 |
| 12Q GAIN WILD-TYPE | 24 | 385 | 76 |
Figure S31. Get High-res Image Gene #11: '12q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 6.85e-07 (Fisher's exact test), Q value = 2e-04
Table S32. Gene #11: '12q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 12Q GAIN CNV | 12 | 0 | 0 |
| 12Q GAIN WILD-TYPE | 137 | 74 | 274 |
Figure S32. Get High-res Image Gene #11: '12q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 3.88e-06 (Fisher's exact test), Q value = 0.0011
Table S33. Gene #11: '12q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 12Q GAIN CNV | 12 | 0 | 0 | 0 |
| 12Q GAIN WILD-TYPE | 128 | 56 | 106 | 156 |
Figure S33. Get High-res Image Gene #11: '12q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 7.94e-07 (Fisher's exact test), Q value = 0.00023
Table S34. Gene #11: '12q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 12Q GAIN CNV | 12 | 0 | 0 | 0 |
| 12Q GAIN WILD-TYPE | 111 | 191 | 53 | 91 |
Figure S34. Get High-res Image Gene #11: '12q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 1.64e-05 (Fisher's exact test), Q value = 0.0046
Table S35. Gene #11: '12q gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 148 | 149 |
| 12Q GAIN CNV | 11 | 0 | 1 |
| 12Q GAIN WILD-TYPE | 116 | 148 | 148 |
Figure S35. Get High-res Image Gene #11: '12q gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'

P value = 4.85e-06 (Fisher's exact test), Q value = 0.0014
Table S36. Gene #11: '12q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 12Q GAIN CNV | 11 | 1 | 0 |
| 12Q GAIN WILD-TYPE | 103 | 167 | 142 |
Figure S36. Get High-res Image Gene #11: '12q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
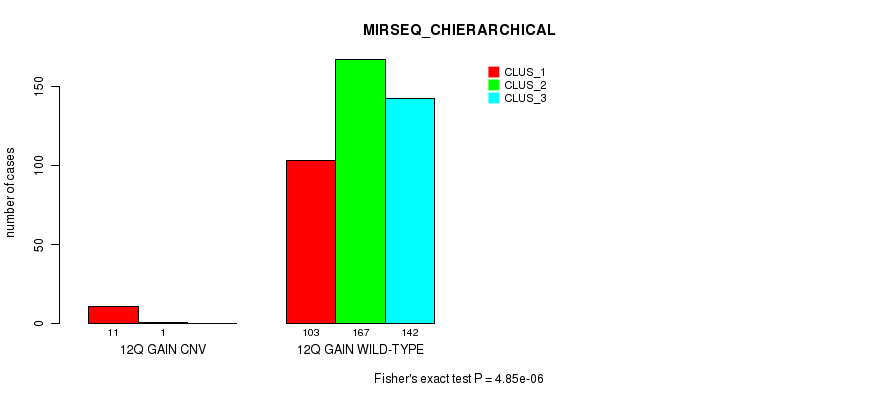
P value = 0.000294 (Fisher's exact test), Q value = 0.07
Table S37. Gene #12: '13q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 13Q GAIN CNV | 3 | 0 | 0 |
| 13Q GAIN WILD-TYPE | 31 | 385 | 78 |
Figure S37. Get High-res Image Gene #12: '13q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 5.02e-09 (Fisher's exact test), Q value = 1.5e-06
Table S38. Gene #13: '14q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 14Q GAIN CNV | 7 | 0 | 1 |
| 14Q GAIN WILD-TYPE | 27 | 385 | 77 |
Figure S38. Get High-res Image Gene #13: '14q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 8.94e-05 (Fisher's exact test), Q value = 0.023
Table S39. Gene #13: '14q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 14Q GAIN CNV | 8 | 0 | 0 |
| 14Q GAIN WILD-TYPE | 141 | 74 | 274 |
Figure S39. Get High-res Image Gene #13: '14q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.000496 (Fisher's exact test), Q value = 0.11
Table S40. Gene #13: '14q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 14Q GAIN CNV | 8 | 0 | 0 | 0 |
| 14Q GAIN WILD-TYPE | 132 | 56 | 106 | 156 |
Figure S40. Get High-res Image Gene #13: '14q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 0.000164 (Fisher's exact test), Q value = 0.041
Table S41. Gene #13: '14q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 14Q GAIN CNV | 8 | 0 | 0 | 0 |
| 14Q GAIN WILD-TYPE | 115 | 191 | 53 | 91 |
Figure S41. Get High-res Image Gene #13: '14q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 5.51e-14 (Fisher's exact test), Q value = 1.7e-11
Table S42. Gene #14: '16p gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 16P GAIN CNV | 11 | 0 | 1 |
| 16P GAIN WILD-TYPE | 23 | 385 | 77 |
Figure S42. Get High-res Image Gene #14: '16p gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 2.82e-05 (Fisher's exact test), Q value = 0.0076
Table S43. Gene #14: '16p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 16P GAIN CNV | 10 | 2 | 0 |
| 16P GAIN WILD-TYPE | 139 | 72 | 274 |
Figure S43. Get High-res Image Gene #14: '16p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.000128 (Fisher's exact test), Q value = 0.032
Table S44. Gene #14: '16p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 16P GAIN CNV | 11 | 0 | 0 | 1 |
| 16P GAIN WILD-TYPE | 129 | 56 | 106 | 155 |
Figure S44. Get High-res Image Gene #14: '16p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 3.37e-05 (Fisher's exact test), Q value = 0.009
Table S45. Gene #14: '16p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 16P GAIN CNV | 11 | 1 | 0 | 0 |
| 16P GAIN WILD-TYPE | 112 | 190 | 53 | 91 |
Figure S45. Get High-res Image Gene #14: '16p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 0.000214 (Fisher's exact test), Q value = 0.052
Table S46. Gene #14: '16p gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 148 | 149 |
| 16P GAIN CNV | 10 | 0 | 2 |
| 16P GAIN WILD-TYPE | 117 | 148 | 147 |
Figure S46. Get High-res Image Gene #14: '16p gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'

P value = 5.88e-05 (Fisher's exact test), Q value = 0.015
Table S47. Gene #14: '16p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 16P GAIN CNV | 10 | 2 | 0 |
| 16P GAIN WILD-TYPE | 104 | 166 | 142 |
Figure S47. Get High-res Image Gene #14: '16p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 5.67e-13 (Fisher's exact test), Q value = 1.8e-10
Table S48. Gene #15: '16q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 16Q GAIN CNV | 10 | 0 | 0 |
| 16Q GAIN WILD-TYPE | 24 | 385 | 78 |
Figure S48. Get High-res Image Gene #15: '16q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000247 (Fisher's exact test), Q value = 0.059
Table S49. Gene #15: '16q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 16Q GAIN CNV | 8 | 2 | 0 |
| 16Q GAIN WILD-TYPE | 141 | 72 | 274 |
Figure S49. Get High-res Image Gene #15: '16q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.000429 (Fisher's exact test), Q value = 0.099
Table S50. Gene #15: '16q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 16Q GAIN CNV | 9 | 1 | 0 | 0 |
| 16Q GAIN WILD-TYPE | 114 | 190 | 53 | 91 |
Figure S50. Get High-res Image Gene #15: '16q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 0.000547 (Fisher's exact test), Q value = 0.12
Table S51. Gene #15: '16q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 16Q GAIN CNV | 8 | 2 | 0 |
| 16Q GAIN WILD-TYPE | 106 | 166 | 142 |
Figure S51. Get High-res Image Gene #15: '16q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 2e-09 (Fisher's exact test), Q value = 6.1e-07
Table S52. Gene #16: '17p gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 17P GAIN CNV | 9 | 2 | 1 |
| 17P GAIN WILD-TYPE | 25 | 383 | 77 |
Figure S52. Get High-res Image Gene #16: '17p gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 6.85e-07 (Fisher's exact test), Q value = 2e-04
Table S53. Gene #16: '17p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 17P GAIN CNV | 12 | 0 | 0 |
| 17P GAIN WILD-TYPE | 137 | 74 | 274 |
Figure S53. Get High-res Image Gene #16: '17p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 3.88e-06 (Fisher's exact test), Q value = 0.0011
Table S54. Gene #16: '17p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 17P GAIN CNV | 12 | 0 | 0 | 0 |
| 17P GAIN WILD-TYPE | 128 | 56 | 106 | 156 |
Figure S54. Get High-res Image Gene #16: '17p gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 7.94e-07 (Fisher's exact test), Q value = 0.00023
Table S55. Gene #16: '17p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 17P GAIN CNV | 12 | 0 | 0 | 0 |
| 17P GAIN WILD-TYPE | 111 | 191 | 53 | 91 |
Figure S55. Get High-res Image Gene #16: '17p gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 5.19e-05 (Fisher's exact test), Q value = 0.014
Table S56. Gene #16: '17p gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 148 | 149 |
| 17P GAIN CNV | 10 | 0 | 1 |
| 17P GAIN WILD-TYPE | 117 | 148 | 148 |
Figure S56. Get High-res Image Gene #16: '17p gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'

P value = 1.7e-05 (Fisher's exact test), Q value = 0.0047
Table S57. Gene #16: '17p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 17P GAIN CNV | 10 | 1 | 0 |
| 17P GAIN WILD-TYPE | 104 | 167 | 142 |
Figure S57. Get High-res Image Gene #16: '17p gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 3.83e-09 (Fisher's exact test), Q value = 1.2e-06
Table S58. Gene #17: '17q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 17Q GAIN CNV | 9 | 2 | 2 |
| 17Q GAIN WILD-TYPE | 25 | 383 | 76 |
Figure S58. Get High-res Image Gene #17: '17q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.58e-07 (Fisher's exact test), Q value = 4.7e-05
Table S59. Gene #17: '17q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 17Q GAIN CNV | 13 | 0 | 0 |
| 17Q GAIN WILD-TYPE | 136 | 74 | 274 |
Figure S59. Get High-res Image Gene #17: '17q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1.36e-06 (Fisher's exact test), Q value = 4e-04
Table S60. Gene #17: '17q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 17Q GAIN CNV | 13 | 0 | 0 | 0 |
| 17Q GAIN WILD-TYPE | 127 | 56 | 106 | 156 |
Figure S60. Get High-res Image Gene #17: '17q gain' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 1.92e-07 (Fisher's exact test), Q value = 5.7e-05
Table S61. Gene #17: '17q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 17Q GAIN CNV | 13 | 0 | 0 | 0 |
| 17Q GAIN WILD-TYPE | 110 | 191 | 53 | 91 |
Figure S61. Get High-res Image Gene #17: '17q gain' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 1.64e-05 (Fisher's exact test), Q value = 0.0046
Table S62. Gene #17: '17q gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 148 | 149 |
| 17Q GAIN CNV | 11 | 0 | 1 |
| 17Q GAIN WILD-TYPE | 116 | 148 | 148 |
Figure S62. Get High-res Image Gene #17: '17q gain' versus Molecular Subtype #7: 'MIRSEQ_CNMF'

P value = 4.85e-06 (Fisher's exact test), Q value = 0.0014
Table S63. Gene #17: '17q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 17Q GAIN CNV | 11 | 1 | 0 |
| 17Q GAIN WILD-TYPE | 103 | 167 | 142 |
Figure S63. Get High-res Image Gene #17: '17q gain' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 1.85e-05 (Fisher's exact test), Q value = 0.0051
Table S64. Gene #20: '19p gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 19P GAIN CNV | 4 | 0 | 0 |
| 19P GAIN WILD-TYPE | 30 | 385 | 78 |
Figure S64. Get High-res Image Gene #20: '19p gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.12e-06 (Fisher's exact test), Q value = 0.00033
Table S65. Gene #21: '19q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 19Q GAIN CNV | 5 | 0 | 0 |
| 19Q GAIN WILD-TYPE | 29 | 385 | 78 |
Figure S65. Get High-res Image Gene #21: '19q gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.11e-11 (Fisher's exact test), Q value = 3.4e-09
Table S66. Gene #22: '20p gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 20P GAIN CNV | 9 | 0 | 0 |
| 20P GAIN WILD-TYPE | 25 | 385 | 78 |
Figure S66. Get High-res Image Gene #22: '20p gain' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000767 (Fisher's exact test), Q value = 0.17
Table S67. Gene #22: '20p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 20P GAIN CNV | 7 | 2 | 0 |
| 20P GAIN WILD-TYPE | 142 | 72 | 274 |
Figure S67. Get High-res Image Gene #22: '20p gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1.11e-11 (Fisher's exact test), Q value = 3.4e-09
Table S68. Gene #23: '20q gain' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 20Q GAIN CNV | 9 | 0 | 0 |
| 20Q GAIN WILD-TYPE | 25 | 385 | 78 |
Figure S68. Get High-res Image Gene #23: '20q gain' versus Molecular Subtype #1: 'CN_CNMF'
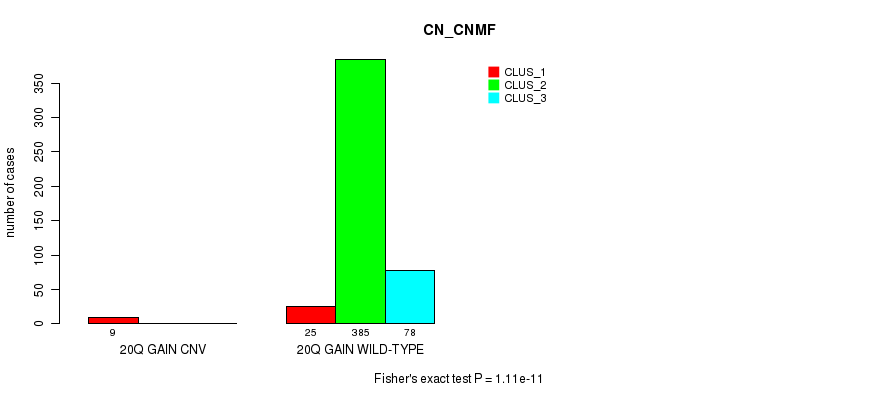
P value = 0.000767 (Fisher's exact test), Q value = 0.17
Table S69. Gene #23: '20q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 20Q GAIN CNV | 7 | 2 | 0 |
| 20Q GAIN WILD-TYPE | 142 | 72 | 274 |
Figure S69. Get High-res Image Gene #23: '20q gain' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.000294 (Fisher's exact test), Q value = 0.07
Table S70. Gene #25: '1p loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 1P LOSS CNV | 3 | 0 | 0 |
| 1P LOSS WILD-TYPE | 31 | 385 | 78 |
Figure S70. Get High-res Image Gene #25: '1p loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 5.02e-09 (Fisher's exact test), Q value = 1.5e-06
Table S71. Gene #26: '2p loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 2P LOSS CNV | 7 | 0 | 1 |
| 2P LOSS WILD-TYPE | 27 | 385 | 77 |
Figure S71. Get High-res Image Gene #26: '2p loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 8.94e-05 (Fisher's exact test), Q value = 0.023
Table S72. Gene #26: '2p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 2P LOSS CNV | 8 | 0 | 0 |
| 2P LOSS WILD-TYPE | 141 | 74 | 274 |
Figure S72. Get High-res Image Gene #26: '2p loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.000496 (Fisher's exact test), Q value = 0.11
Table S73. Gene #26: '2p loss' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 2P LOSS CNV | 8 | 0 | 0 | 0 |
| 2P LOSS WILD-TYPE | 132 | 56 | 106 | 156 |
Figure S73. Get High-res Image Gene #26: '2p loss' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 0.000164 (Fisher's exact test), Q value = 0.041
Table S74. Gene #26: '2p loss' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 2P LOSS CNV | 8 | 0 | 0 | 0 |
| 2P LOSS WILD-TYPE | 115 | 191 | 53 | 91 |
Figure S74. Get High-res Image Gene #26: '2p loss' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 5.53e-05 (Fisher's exact test), Q value = 0.014
Table S75. Gene #26: '2p loss' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 148 | 149 |
| 2P LOSS CNV | 8 | 0 | 0 |
| 2P LOSS WILD-TYPE | 119 | 148 | 149 |
Figure S75. Get High-res Image Gene #26: '2p loss' versus Molecular Subtype #7: 'MIRSEQ_CNMF'

P value = 2.27e-05 (Fisher's exact test), Q value = 0.0062
Table S76. Gene #26: '2p loss' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 2P LOSS CNV | 8 | 0 | 0 |
| 2P LOSS WILD-TYPE | 106 | 168 | 142 |
Figure S76. Get High-res Image Gene #26: '2p loss' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 3.78e-09 (Fisher's exact test), Q value = 1.2e-06
Table S77. Gene #27: '2q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 2Q LOSS CNV | 7 | 0 | 0 |
| 2Q LOSS WILD-TYPE | 27 | 385 | 78 |
Figure S77. Get High-res Image Gene #27: '2q loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000378 (Fisher's exact test), Q value = 0.088
Table S78. Gene #27: '2q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 2Q LOSS CNV | 7 | 0 | 0 |
| 2Q LOSS WILD-TYPE | 142 | 74 | 274 |
Figure S78. Get High-res Image Gene #27: '2q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.00065 (Fisher's exact test), Q value = 0.15
Table S79. Gene #27: '2q loss' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 2Q LOSS CNV | 7 | 0 | 0 | 0 |
| 2Q LOSS WILD-TYPE | 116 | 191 | 53 | 91 |
Figure S79. Get High-res Image Gene #27: '2q loss' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 0.000192 (Fisher's exact test), Q value = 0.048
Table S80. Gene #27: '2q loss' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 148 | 149 |
| 2Q LOSS CNV | 7 | 0 | 0 |
| 2Q LOSS WILD-TYPE | 120 | 148 | 149 |
Figure S80. Get High-res Image Gene #27: '2q loss' versus Molecular Subtype #7: 'MIRSEQ_CNMF'

P value = 8.85e-05 (Fisher's exact test), Q value = 0.023
Table S81. Gene #27: '2q loss' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 2Q LOSS CNV | 7 | 0 | 0 |
| 2Q LOSS WILD-TYPE | 107 | 168 | 142 |
Figure S81. Get High-res Image Gene #27: '2q loss' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 0.000294 (Fisher's exact test), Q value = 0.07
Table S82. Gene #28: '3q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 3Q LOSS CNV | 3 | 0 | 0 |
| 3Q LOSS WILD-TYPE | 31 | 385 | 78 |
Figure S82. Get High-res Image Gene #28: '3q loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.85e-05 (Fisher's exact test), Q value = 0.0051
Table S83. Gene #30: '8p loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 8P LOSS CNV | 4 | 0 | 0 |
| 8P LOSS WILD-TYPE | 30 | 385 | 78 |
Figure S83. Get High-res Image Gene #30: '8p loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.85e-05 (Fisher's exact test), Q value = 0.0051
Table S84. Gene #31: '8q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 8Q LOSS CNV | 4 | 0 | 0 |
| 8Q LOSS WILD-TYPE | 30 | 385 | 78 |
Figure S84. Get High-res Image Gene #31: '8q loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.08e-11 (Fisher's exact test), Q value = 3.4e-09
Table S85. Gene #32: '9p loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 9P LOSS CNV | 10 | 1 | 1 |
| 9P LOSS WILD-TYPE | 24 | 384 | 77 |
Figure S85. Get High-res Image Gene #32: '9p loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.58e-13 (Fisher's exact test), Q value = 5e-11
Table S86. Gene #33: '9q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 9Q LOSS CNV | 13 | 3 | 1 |
| 9Q LOSS WILD-TYPE | 21 | 382 | 77 |
Figure S86. Get High-res Image Gene #33: '9q loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 6.62e-08 (Fisher's exact test), Q value = 2e-05
Table S87. Gene #34: '10p loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 10P LOSS CNV | 6 | 0 | 0 |
| 10P LOSS WILD-TYPE | 28 | 385 | 78 |
Figure S87. Get High-res Image Gene #34: '10p loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1.12e-06 (Fisher's exact test), Q value = 0.00033
Table S88. Gene #35: '10q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 10Q LOSS CNV | 5 | 0 | 0 |
| 10Q LOSS WILD-TYPE | 29 | 385 | 78 |
Figure S88. Get High-res Image Gene #35: '10q loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 7.74e-08 (Fisher's exact test), Q value = 2.3e-05
Table S89. Gene #36: '11p loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 11P LOSS CNV | 6 | 0 | 1 |
| 11P LOSS WILD-TYPE | 28 | 385 | 77 |
Figure S89. Get High-res Image Gene #36: '11p loss' versus Molecular Subtype #1: 'CN_CNMF'
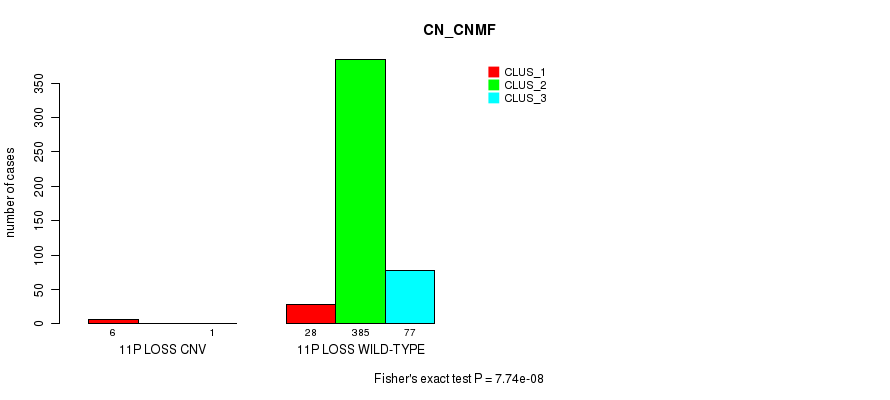
P value = 3.24e-06 (Fisher's exact test), Q value = 0.00094
Table S90. Gene #37: '11q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 11Q LOSS CNV | 6 | 2 | 1 |
| 11Q LOSS WILD-TYPE | 28 | 383 | 77 |
Figure S90. Get High-res Image Gene #37: '11q loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000464 (Fisher's exact test), Q value = 0.11
Table S91. Gene #39: '15q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 15Q LOSS CNV | 4 | 2 | 1 |
| 15Q LOSS WILD-TYPE | 30 | 383 | 77 |
Figure S91. Get High-res Image Gene #39: '15q loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000664 (Fisher's exact test), Q value = 0.15
Table S92. Gene #39: '15q loss' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 148 | 149 |
| 15Q LOSS CNV | 6 | 0 | 0 |
| 15Q LOSS WILD-TYPE | 121 | 148 | 149 |
Figure S92. Get High-res Image Gene #39: '15q loss' versus Molecular Subtype #7: 'MIRSEQ_CNMF'

P value = 0.000342 (Fisher's exact test), Q value = 0.08
Table S93. Gene #39: '15q loss' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 15Q LOSS CNV | 6 | 0 | 0 |
| 15Q LOSS WILD-TYPE | 108 | 168 | 142 |
Figure S93. Get High-res Image Gene #39: '15q loss' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

P value = 0.000204 (Fisher's exact test), Q value = 0.05
Table S94. Gene #41: '18p loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 18P LOSS CNV | 3 | 0 | 1 |
| 18P LOSS WILD-TYPE | 31 | 385 | 77 |
Figure S94. Get High-res Image Gene #41: '18p loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000204 (Fisher's exact test), Q value = 0.05
Table S95. Gene #42: '18q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 18Q LOSS CNV | 3 | 0 | 1 |
| 18Q LOSS WILD-TYPE | 31 | 385 | 77 |
Figure S95. Get High-res Image Gene #42: '18q loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.000192 (Fisher's exact test), Q value = 0.048
Table S96. Gene #44: '21q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 21Q LOSS CNV | 4 | 1 | 1 |
| 21Q LOSS WILD-TYPE | 30 | 384 | 77 |
Figure S96. Get High-res Image Gene #44: '21q loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 6.79e-58 (Fisher's exact test), Q value = 2.2e-55
Table S97. Gene #45: '22q loss' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 34 | 385 | 78 |
| 22Q LOSS CNV | 8 | 0 | 58 |
| 22Q LOSS WILD-TYPE | 26 | 385 | 20 |
Figure S97. Get High-res Image Gene #45: '22q loss' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00013 (Fisher's exact test), Q value = 0.033
Table S98. Gene #45: '22q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 149 | 74 | 274 |
| 22Q LOSS CNV | 35 | 7 | 24 |
| 22Q LOSS WILD-TYPE | 114 | 67 | 250 |
Figure S98. Get High-res Image Gene #45: '22q loss' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 8.4e-11 (Fisher's exact test), Q value = 2.6e-08
Table S99. Gene #45: '22q loss' versus Molecular Subtype #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 140 | 56 | 106 | 156 |
| 22Q LOSS CNV | 33 | 5 | 20 | 1 |
| 22Q LOSS WILD-TYPE | 107 | 51 | 86 | 155 |
Figure S99. Get High-res Image Gene #45: '22q loss' versus Molecular Subtype #5: 'MRNASEQ_CNMF'

P value = 1.98e-10 (Fisher's exact test), Q value = 6.1e-08
Table S100. Gene #45: '22q loss' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 123 | 191 | 53 | 91 |
| 22Q LOSS CNV | 33 | 6 | 2 | 18 |
| 22Q LOSS WILD-TYPE | 90 | 185 | 51 | 73 |
Figure S100. Get High-res Image Gene #45: '22q loss' versus Molecular Subtype #6: 'MRNASEQ_CHIERARCHICAL'

P value = 4.33e-05 (Fisher's exact test), Q value = 0.011
Table S101. Gene #45: '22q loss' versus Molecular Subtype #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 127 | 148 | 149 |
| 22Q LOSS CNV | 31 | 14 | 10 |
| 22Q LOSS WILD-TYPE | 96 | 134 | 139 |
Figure S101. Get High-res Image Gene #45: '22q loss' versus Molecular Subtype #7: 'MIRSEQ_CNMF'

P value = 9.51e-06 (Fisher's exact test), Q value = 0.0027
Table S102. Gene #45: '22q loss' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 114 | 168 | 142 |
| 22Q LOSS CNV | 30 | 11 | 14 |
| 22Q LOSS WILD-TYPE | 84 | 157 | 128 |
Figure S102. Get High-res Image Gene #45: '22q loss' versus Molecular Subtype #8: 'MIRSEQ_CHIERARCHICAL'

-
Mutation data file = broad_values_by_arm.mutsig.cluster.txt
-
Molecular subtypes file = THCA-TP.transferedmergedcluster.txt
-
Number of patients = 497
-
Number of significantly arm-level cnvs = 45
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.