This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 40 focal events and 8 molecular subtypes across 172 patients, 99 significant findings detected with P value < 0.05 and Q value < 0.25.
-
1p cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
1q cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
2p cnv correlated to 'CN_CNMF'.
-
2q cnv correlated to 'CN_CNMF'.
-
3p cnv correlated to 'CN_CNMF'.
-
3q cnv correlated to 'CN_CNMF'.
-
4p cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
4q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CNMF'.
-
5p cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
5q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
6p cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CNMF'.
-
6q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CNMF'.
-
7p cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
7q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
8q cnv correlated to 'METHLYATION_CNMF'.
-
9p cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
9q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
10p cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
10q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
11p cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
11q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
12p cnv correlated to 'CN_CNMF'.
-
12q cnv correlated to 'CN_CNMF'.
-
13q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
14q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
15q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
16p cnv correlated to 'CN_CNMF'.
-
16q cnv correlated to 'CN_CNMF'.
-
17p cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
17q cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
18p cnv correlated to 'METHLYATION_CNMF' and 'MRNASEQ_CNMF'.
-
18q cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
19p cnv correlated to 'CN_CNMF'.
-
19q cnv correlated to 'CN_CNMF'.
-
22q cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
xq cnv correlated to 'CN_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 40 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 99 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| 9q | 26 (15%) | 146 |
8.81e-07 (0.000249) |
1.32e-08 (4.02e-06) |
1.17e-08 (3.56e-06) |
1.09e-07 (3.24e-05) |
0.00475 (0.945) |
1.3e-05 (0.00346) |
0.0336 (1.00) |
1.83e-05 (0.00476) |
| 10q | 17 (10%) | 155 |
6.87e-07 (0.000196) |
2.4e-07 (6.96e-05) |
0.000451 (0.106) |
0.0932 (1.00) |
0.605 (1.00) |
0.000372 (0.0889) |
0.507 (1.00) |
0.000635 (0.146) |
| 14q | 37 (22%) | 135 |
2.82e-13 (8.87e-11) |
0.000432 (0.102) |
5.95e-05 (0.0151) |
4.08e-08 (1.22e-05) |
0.0941 (1.00) |
0.000843 (0.189) |
0.311 (1.00) |
0.0351 (1.00) |
| 22q | 42 (24%) | 130 |
7.5e-07 (0.000214) |
1.18e-07 (3.48e-05) |
1.29e-07 (3.8e-05) |
0.000958 (0.213) |
0.0988 (1.00) |
1.61e-05 (0.00424) |
0.0294 (1.00) |
0.0182 (1.00) |
| 1p | 26 (15%) | 146 |
1.74e-07 (5.11e-05) |
0.000735 (0.168) |
2.69e-05 (0.00697) |
0.000124 (0.0308) |
0.783 (1.00) |
0.677 (1.00) |
0.521 (1.00) |
0.87 (1.00) |
| 4p | 24 (14%) | 148 |
2.9e-06 (8e-04) |
9.7e-09 (2.97e-06) |
3.28e-06 (0.000898) |
0.000532 (0.124) |
0.562 (1.00) |
0.0494 (1.00) |
0.456 (1.00) |
0.00325 (0.666) |
| 7p | 101 (59%) | 71 |
1.14e-18 (3.66e-16) |
2.12e-07 (6.16e-05) |
1.43e-12 (4.48e-10) |
2.01e-07 (5.88e-05) |
0.0969 (1.00) |
0.00126 (0.277) |
0.17 (1.00) |
0.0408 (1.00) |
| 7q | 102 (59%) | 70 |
4e-18 (1.28e-15) |
5.75e-07 (0.000166) |
2.02e-12 (6.29e-10) |
3.02e-07 (8.73e-05) |
0.149 (1.00) |
0.0054 (1.00) |
0.234 (1.00) |
0.126 (1.00) |
| 9p | 25 (15%) | 147 |
7.59e-07 (0.000216) |
3.38e-08 (1.02e-05) |
3.66e-08 (1.1e-05) |
1.79e-05 (0.00469) |
0.00986 (1.00) |
0.00162 (0.35) |
0.045 (1.00) |
0.00197 (0.419) |
| 10p | 17 (10%) | 155 |
1.51e-08 (4.56e-06) |
1.14e-06 (0.000318) |
0.000451 (0.106) |
0.26 (1.00) |
0.876 (1.00) |
0.00078 (0.176) |
0.842 (1.00) |
0.00123 (0.27) |
| 17p | 110 (64%) | 62 |
1.05e-14 (3.32e-12) |
1.76e-05 (0.00462) |
9.26e-07 (0.000261) |
5.28e-06 (0.00142) |
0.0489 (1.00) |
0.00373 (0.754) |
0.101 (1.00) |
0.0854 (1.00) |
| 1q | 27 (16%) | 145 |
1.38e-06 (0.000384) |
0.00394 (0.792) |
0.000341 (0.0822) |
0.000526 (0.123) |
0.603 (1.00) |
0.177 (1.00) |
0.523 (1.00) |
0.602 (1.00) |
| 4q | 24 (14%) | 148 |
2.9e-06 (8e-04) |
2.21e-09 (6.79e-07) |
3.28e-06 (0.000898) |
0.00156 (0.338) |
0.843 (1.00) |
0.128 (1.00) |
0.355 (1.00) |
0.0125 (1.00) |
| 5p | 33 (19%) | 139 |
7.13e-05 (0.018) |
3.5e-06 (0.000952) |
0.00594 (1.00) |
0.000122 (0.0306) |
0.0217 (1.00) |
0.122 (1.00) |
0.192 (1.00) |
0.163 (1.00) |
| 5q | 33 (19%) | 139 |
0.000191 (0.0472) |
4.46e-05 (0.0114) |
0.00996 (1.00) |
0.000122 (0.0306) |
0.0217 (1.00) |
0.353 (1.00) |
0.192 (1.00) |
0.287 (1.00) |
| 6p | 20 (12%) | 152 |
0.000544 (0.126) |
0.000899 (0.201) |
0.000357 (0.0858) |
0.0275 (1.00) |
0.0407 (1.00) |
0.123 (1.00) |
0.0911 (1.00) |
0.106 (1.00) |
| 6q | 21 (12%) | 151 |
4.01e-06 (0.00109) |
3.3e-05 (0.00848) |
0.000423 (0.101) |
0.0121 (1.00) |
0.283 (1.00) |
0.173 (1.00) |
0.159 (1.00) |
0.152 (1.00) |
| 11p | 18 (10%) | 154 |
2.72e-06 (0.000752) |
8.58e-06 (0.0023) |
0.0305 (1.00) |
0.000324 (0.0783) |
0.0229 (1.00) |
0.165 (1.00) |
0.518 (1.00) |
0.00927 (1.00) |
| 11q | 21 (12%) | 151 |
3.45e-08 (1.04e-05) |
4.06e-08 (1.21e-05) |
0.011 (1.00) |
0.000532 (0.124) |
0.00966 (1.00) |
0.128 (1.00) |
0.417 (1.00) |
0.0334 (1.00) |
| 13q | 36 (21%) | 136 |
5.57e-05 (0.0142) |
0.000199 (0.049) |
0.044 (1.00) |
0.000755 (0.171) |
0.0677 (1.00) |
0.0557 (1.00) |
0.123 (1.00) |
0.0675 (1.00) |
| 15q | 24 (14%) | 148 |
5.82e-07 (0.000167) |
0.000618 (0.142) |
0.00291 (0.602) |
1.31e-06 (0.000366) |
0.302 (1.00) |
0.533 (1.00) |
0.354 (1.00) |
0.308 (1.00) |
| 17q | 117 (68%) | 55 |
5.2e-12 (1.62e-09) |
0.00117 (0.259) |
9.37e-07 (0.000263) |
1.09e-05 (0.00291) |
0.293 (1.00) |
0.0743 (1.00) |
0.49 (1.00) |
0.122 (1.00) |
| 18q | 35 (20%) | 137 |
0.00181 (0.386) |
7.14e-06 (0.00192) |
1.97e-05 (0.00513) |
0.000811 (0.182) |
0.0308 (1.00) |
0.0329 (1.00) |
0.186 (1.00) |
0.342 (1.00) |
| 18p | 37 (22%) | 135 |
0.00142 (0.31) |
1.54e-05 (0.00409) |
3.04e-05 (0.00785) |
0.00212 (0.45) |
0.00974 (1.00) |
0.0299 (1.00) |
0.171 (1.00) |
0.189 (1.00) |
| 2p | 34 (20%) | 138 |
1.27e-07 (3.76e-05) |
0.00446 (0.893) |
0.195 (1.00) |
0.665 (1.00) |
0.0771 (1.00) |
0.782 (1.00) |
0.0338 (1.00) |
0.857 (1.00) |
| 2q | 36 (21%) | 136 |
7.07e-10 (2.18e-07) |
0.0187 (1.00) |
0.597 (1.00) |
0.455 (1.00) |
0.0859 (1.00) |
0.951 (1.00) |
0.0211 (1.00) |
0.863 (1.00) |
| 3p | 60 (35%) | 112 |
7.81e-16 (2.48e-13) |
0.518 (1.00) |
0.352 (1.00) |
0.118 (1.00) |
0.839 (1.00) |
0.729 (1.00) |
0.809 (1.00) |
1 (1.00) |
| 3q | 59 (34%) | 113 |
3.25e-17 (1.03e-14) |
0.223 (1.00) |
0.00179 (0.384) |
0.524 (1.00) |
0.348 (1.00) |
0.0266 (1.00) |
0.748 (1.00) |
0.124 (1.00) |
| 8q | 19 (11%) | 153 |
0.00254 (0.533) |
0.000104 (0.026) |
0.0033 (0.672) |
0.0656 (1.00) |
0.00783 (1.00) |
0.0388 (1.00) |
0.0582 (1.00) |
0.0122 (1.00) |
| 12p | 69 (40%) | 103 |
0.000301 (0.0731) |
0.605 (1.00) |
0.0287 (1.00) |
0.151 (1.00) |
0.389 (1.00) |
0.0801 (1.00) |
0.0806 (1.00) |
0.21 (1.00) |
| 12q | 68 (40%) | 104 |
0.000236 (0.0575) |
0.461 (1.00) |
0.0156 (1.00) |
0.131 (1.00) |
0.288 (1.00) |
0.069 (1.00) |
0.0597 (1.00) |
0.209 (1.00) |
| 16p | 93 (54%) | 79 |
1.68e-09 (5.17e-07) |
0.37 (1.00) |
0.00724 (1.00) |
0.00995 (1.00) |
0.0574 (1.00) |
0.0519 (1.00) |
0.0048 (0.951) |
0.0654 (1.00) |
| 16q | 90 (52%) | 82 |
4.87e-10 (1.51e-07) |
0.349 (1.00) |
0.0024 (0.506) |
0.0227 (1.00) |
0.0274 (1.00) |
0.00617 (1.00) |
0.0058 (1.00) |
0.0346 (1.00) |
| 19p | 15 (9%) | 157 |
0.000225 (0.0552) |
0.00306 (0.63) |
0.0297 (1.00) |
0.0114 (1.00) |
0.00737 (1.00) |
0.00343 (0.696) |
0.21 (1.00) |
0.00596 (1.00) |
| 19q | 14 (8%) | 158 |
4.27e-05 (0.0109) |
0.00821 (1.00) |
0.029 (1.00) |
0.0491 (1.00) |
0.0127 (1.00) |
0.00811 (1.00) |
0.0737 (1.00) |
0.0178 (1.00) |
| xq | 74 (43%) | 98 |
1.18e-13 (3.72e-11) |
0.0521 (1.00) |
0.0611 (1.00) |
0.0732 (1.00) |
0.159 (1.00) |
0.222 (1.00) |
0.204 (1.00) |
0.668 (1.00) |
| 8p | 18 (10%) | 154 |
0.138 (1.00) |
0.0188 (1.00) |
0.0903 (1.00) |
0.441 (1.00) |
0.0426 (1.00) |
0.0661 (1.00) |
0.107 (1.00) |
0.0215 (1.00) |
| 20p | 59 (34%) | 113 |
0.00274 (0.572) |
0.109 (1.00) |
0.107 (1.00) |
0.42 (1.00) |
0.301 (1.00) |
0.69 (1.00) |
0.862 (1.00) |
0.778 (1.00) |
| 20q | 60 (35%) | 112 |
0.00551 (1.00) |
0.109 (1.00) |
0.142 (1.00) |
0.452 (1.00) |
0.276 (1.00) |
0.729 (1.00) |
0.8 (1.00) |
0.747 (1.00) |
| 21q | 42 (24%) | 130 |
0.00284 (0.591) |
0.0771 (1.00) |
0.0995 (1.00) |
0.0596 (1.00) |
0.312 (1.00) |
0.216 (1.00) |
0.0575 (1.00) |
0.823 (1.00) |
P value = 1.74e-07 (Fisher's exact test), Q value = 5.1e-05
Table S1. Gene #1: '1p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 1P MUTATED | 1 | 9 | 0 | 16 |
| 1P WILD-TYPE | 32 | 59 | 37 | 18 |
Figure S1. Get High-res Image Gene #1: '1p' versus Molecular Subtype #1: 'CN_CNMF'
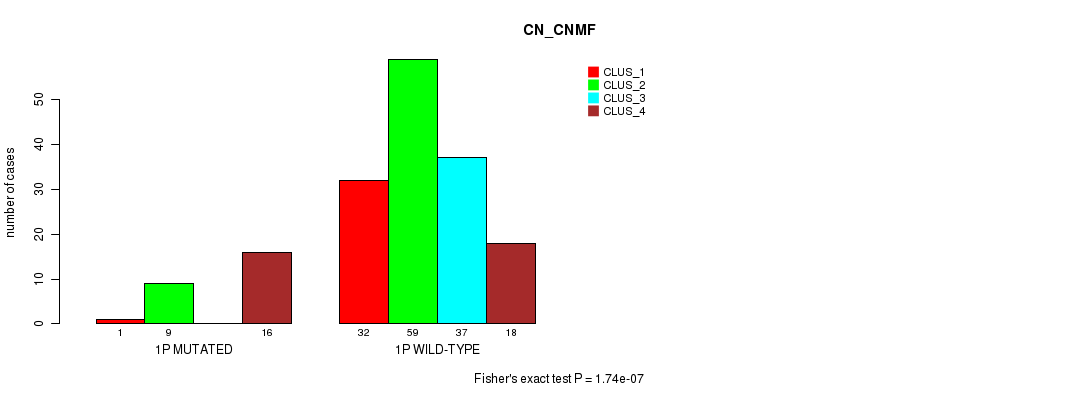
P value = 0.000735 (Fisher's exact test), Q value = 0.17
Table S2. Gene #1: '1p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 1P MUTATED | 5 | 15 | 5 |
| 1P WILD-TYPE | 38 | 28 | 65 |
Figure S2. Get High-res Image Gene #1: '1p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
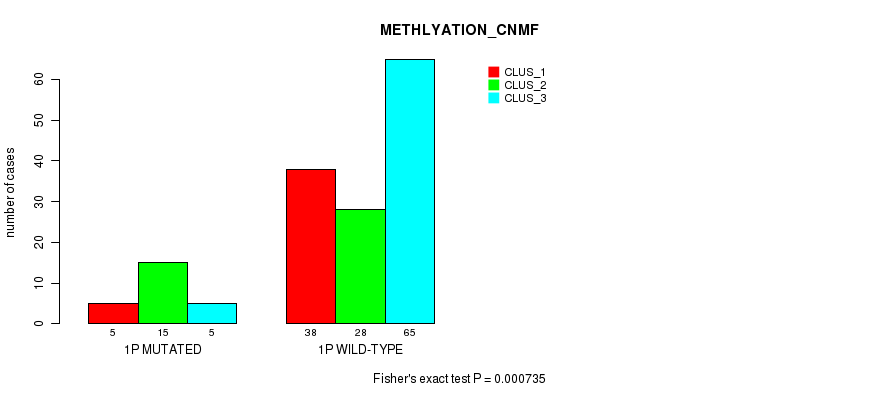
P value = 2.69e-05 (Fisher's exact test), Q value = 0.007
Table S3. Gene #1: '1p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 1P MUTATED | 19 | 3 | 1 | 0 |
| 1P WILD-TYPE | 39 | 46 | 30 | 23 |
Figure S3. Get High-res Image Gene #1: '1p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
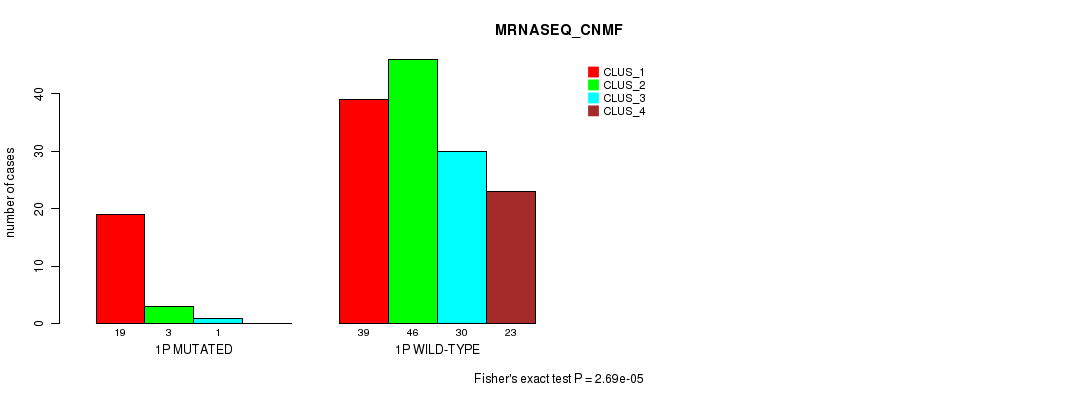
P value = 0.000124 (Fisher's exact test), Q value = 0.031
Table S4. Gene #1: '1p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 1P MUTATED | 12 | 8 | 3 |
| 1P WILD-TYPE | 19 | 58 | 61 |
Figure S4. Get High-res Image Gene #1: '1p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
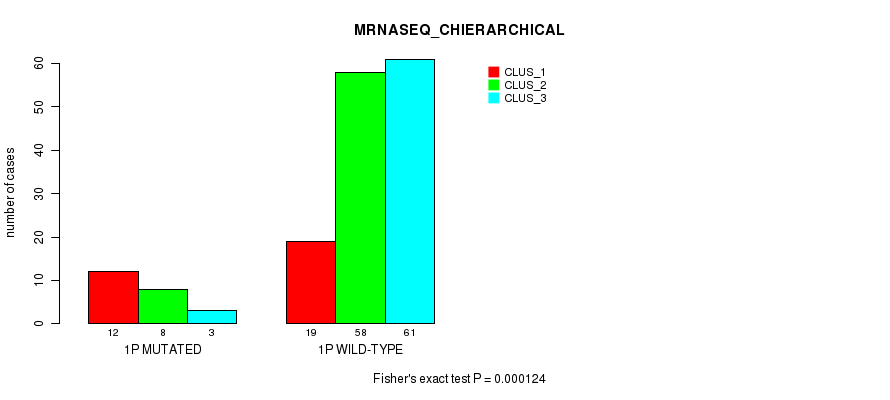
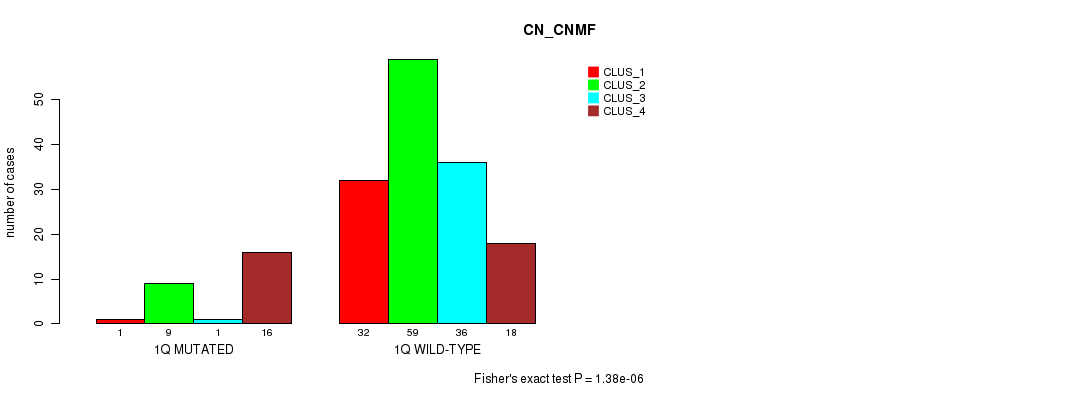
P value = 1.38e-06 (Fisher's exact test), Q value = 0.00038
Table S5. Gene #2: '1q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 1Q MUTATED | 1 | 9 | 1 | 16 |
| 1Q WILD-TYPE | 32 | 59 | 36 | 18 |
Figure S5. Get High-res Image Gene #2: '1q' versus Molecular Subtype #1: 'CN_CNMF'
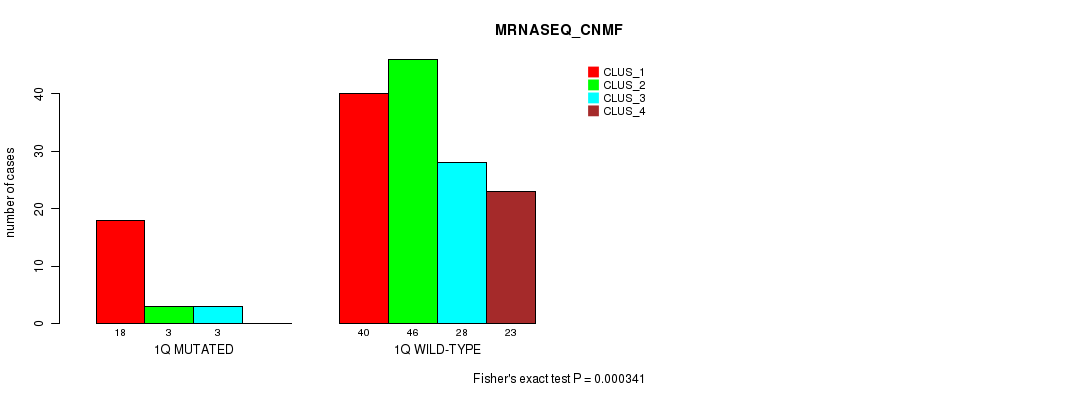
P value = 0.000341 (Fisher's exact test), Q value = 0.082
Table S6. Gene #2: '1q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 1Q MUTATED | 18 | 3 | 3 | 0 |
| 1Q WILD-TYPE | 40 | 46 | 28 | 23 |
Figure S6. Get High-res Image Gene #2: '1q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
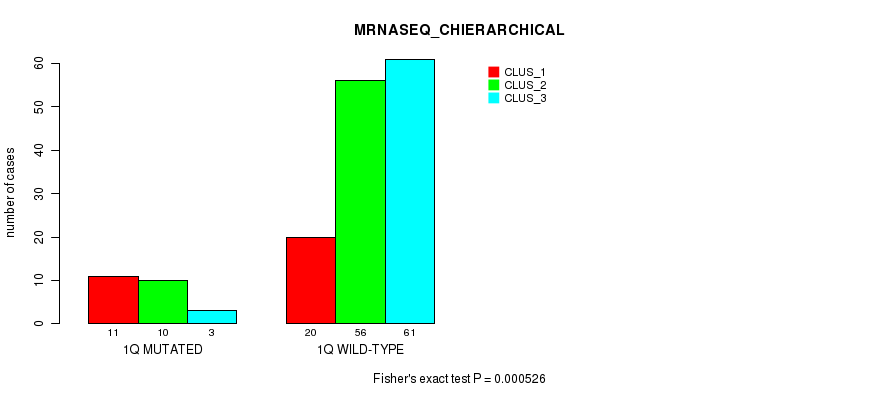
P value = 0.000526 (Fisher's exact test), Q value = 0.12
Table S7. Gene #2: '1q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 1Q MUTATED | 11 | 10 | 3 |
| 1Q WILD-TYPE | 20 | 56 | 61 |
Figure S7. Get High-res Image Gene #2: '1q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
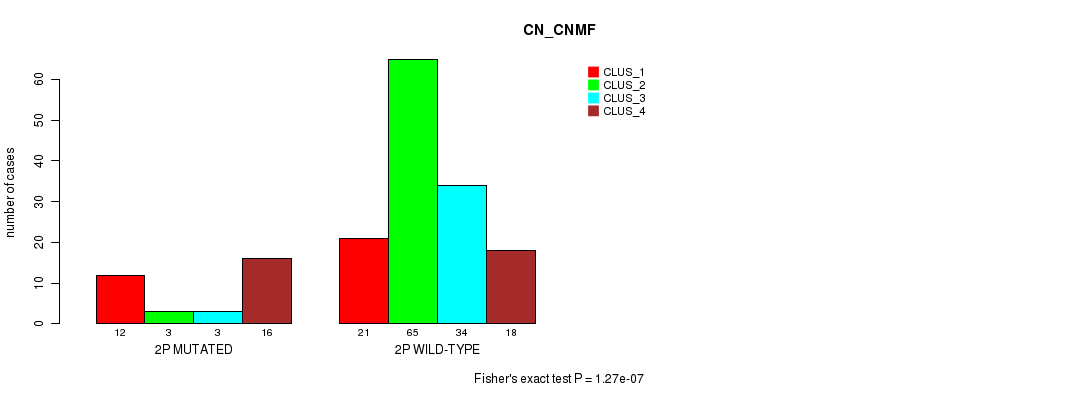
P value = 1.27e-07 (Fisher's exact test), Q value = 3.8e-05
Table S8. Gene #3: '2p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 2P MUTATED | 12 | 3 | 3 | 16 |
| 2P WILD-TYPE | 21 | 65 | 34 | 18 |
Figure S8. Get High-res Image Gene #3: '2p' versus Molecular Subtype #1: 'CN_CNMF'
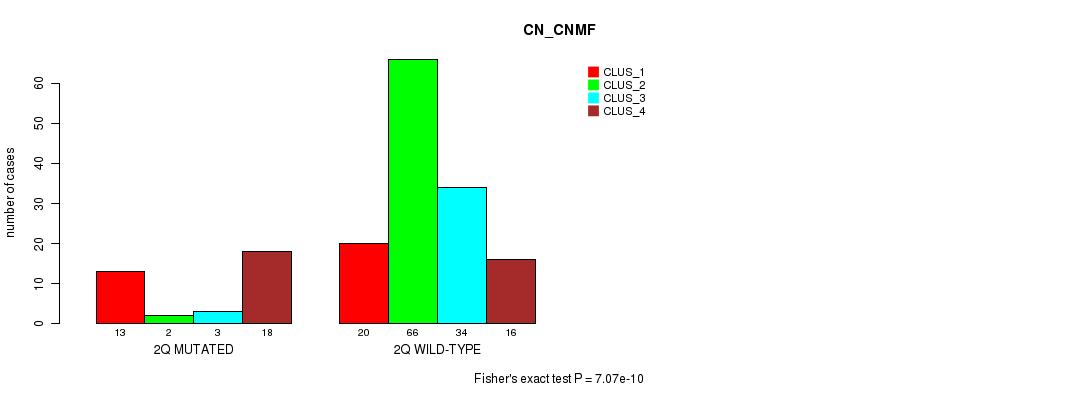
P value = 7.07e-10 (Fisher's exact test), Q value = 2.2e-07
Table S9. Gene #4: '2q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 2Q MUTATED | 13 | 2 | 3 | 18 |
| 2Q WILD-TYPE | 20 | 66 | 34 | 16 |
Figure S9. Get High-res Image Gene #4: '2q' versus Molecular Subtype #1: 'CN_CNMF'
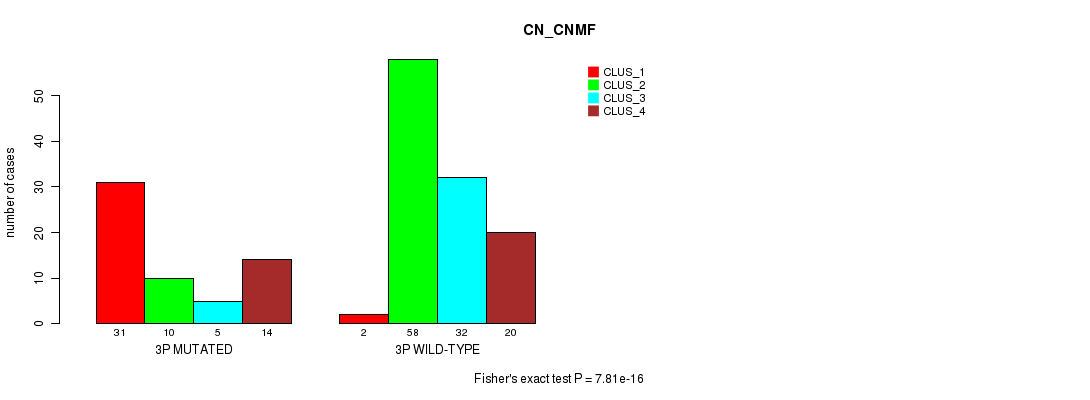
P value = 7.81e-16 (Fisher's exact test), Q value = 2.5e-13
Table S10. Gene #5: '3p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 3P MUTATED | 31 | 10 | 5 | 14 |
| 3P WILD-TYPE | 2 | 58 | 32 | 20 |
Figure S10. Get High-res Image Gene #5: '3p' versus Molecular Subtype #1: 'CN_CNMF'
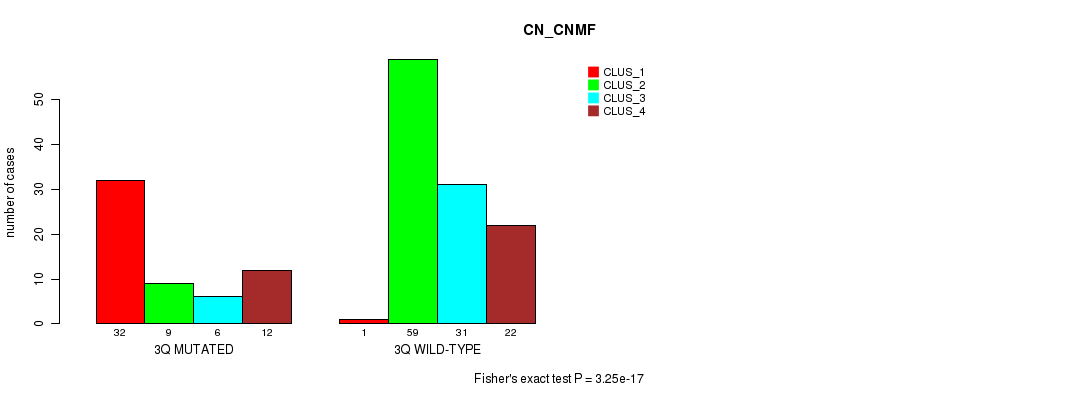
P value = 3.25e-17 (Fisher's exact test), Q value = 1e-14
Table S11. Gene #6: '3q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 3Q MUTATED | 32 | 9 | 6 | 12 |
| 3Q WILD-TYPE | 1 | 59 | 31 | 22 |
Figure S11. Get High-res Image Gene #6: '3q' versus Molecular Subtype #1: 'CN_CNMF'
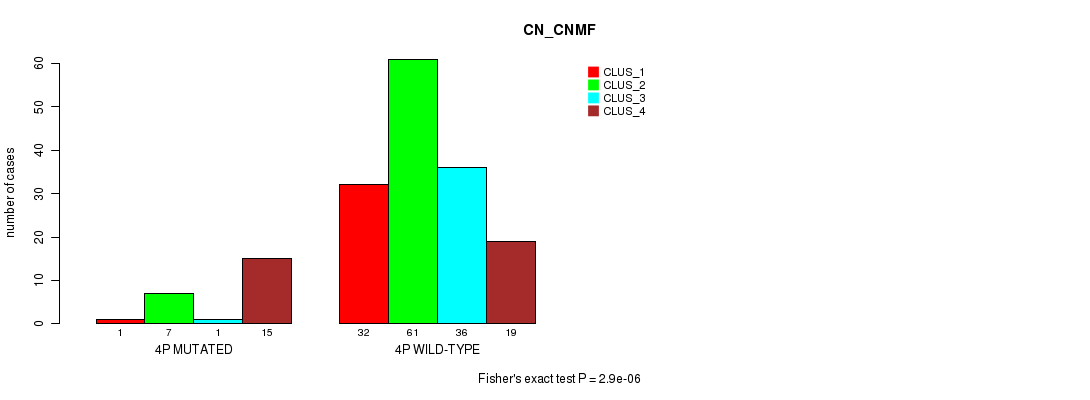
P value = 2.9e-06 (Fisher's exact test), Q value = 8e-04
Table S12. Gene #7: '4p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 4P MUTATED | 1 | 7 | 1 | 15 |
| 4P WILD-TYPE | 32 | 61 | 36 | 19 |
Figure S12. Get High-res Image Gene #7: '4p' versus Molecular Subtype #1: 'CN_CNMF'
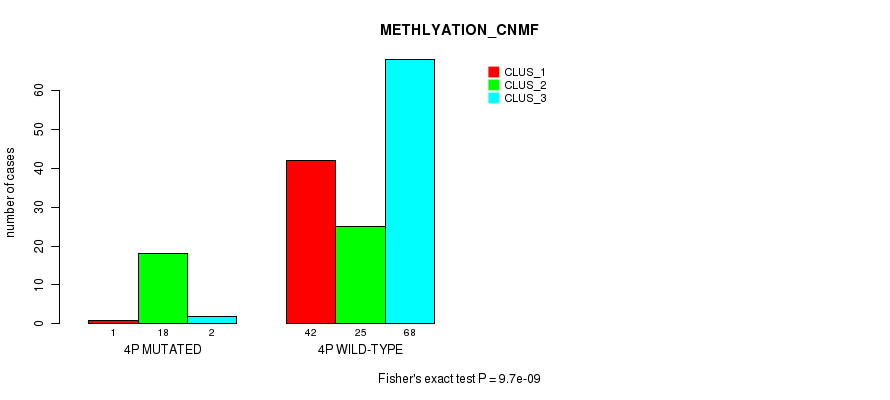
P value = 9.7e-09 (Fisher's exact test), Q value = 3e-06
Table S13. Gene #7: '4p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 4P MUTATED | 1 | 18 | 2 |
| 4P WILD-TYPE | 42 | 25 | 68 |
Figure S13. Get High-res Image Gene #7: '4p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
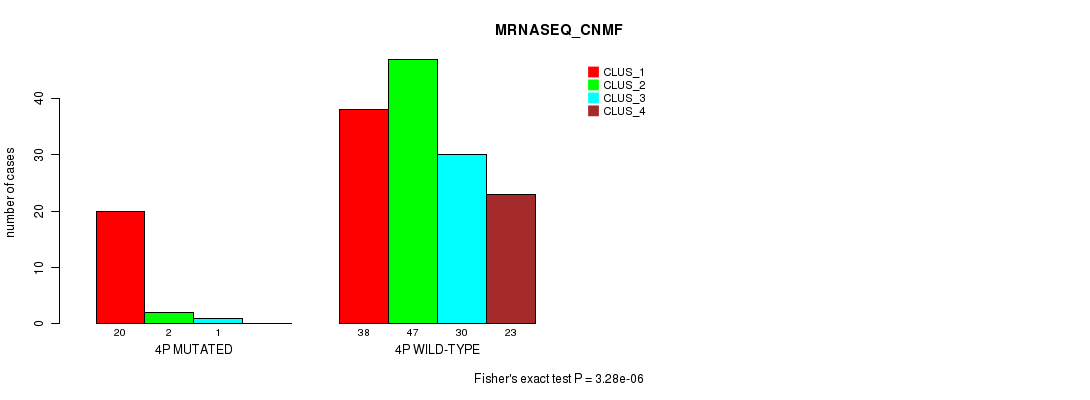
P value = 3.28e-06 (Fisher's exact test), Q value = 9e-04
Table S14. Gene #7: '4p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 4P MUTATED | 20 | 2 | 1 | 0 |
| 4P WILD-TYPE | 38 | 47 | 30 | 23 |
Figure S14. Get High-res Image Gene #7: '4p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
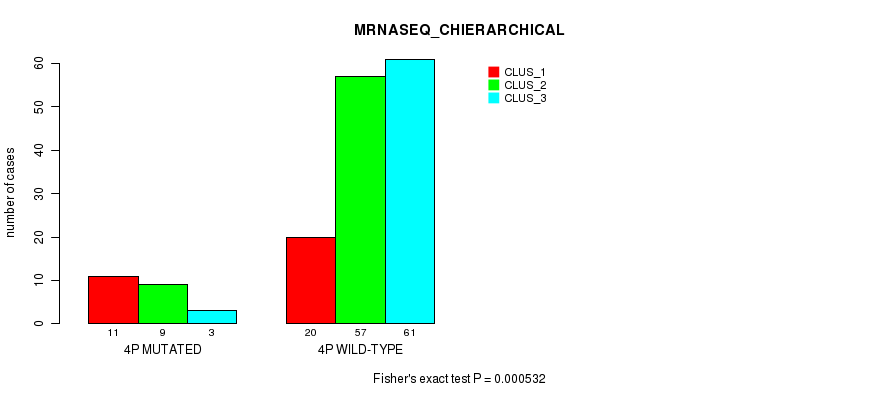
P value = 0.000532 (Fisher's exact test), Q value = 0.12
Table S15. Gene #7: '4p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 4P MUTATED | 11 | 9 | 3 |
| 4P WILD-TYPE | 20 | 57 | 61 |
Figure S15. Get High-res Image Gene #7: '4p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
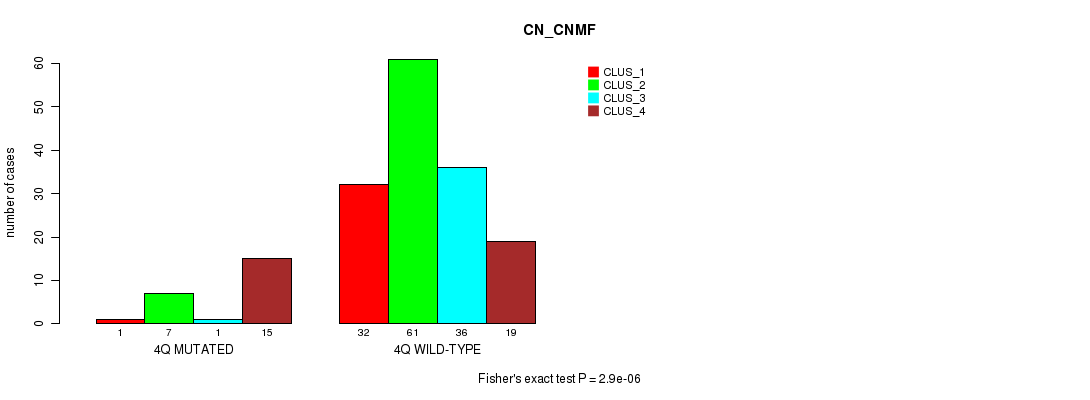
P value = 2.9e-06 (Fisher's exact test), Q value = 8e-04
Table S16. Gene #8: '4q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 4Q MUTATED | 1 | 7 | 1 | 15 |
| 4Q WILD-TYPE | 32 | 61 | 36 | 19 |
Figure S16. Get High-res Image Gene #8: '4q' versus Molecular Subtype #1: 'CN_CNMF'
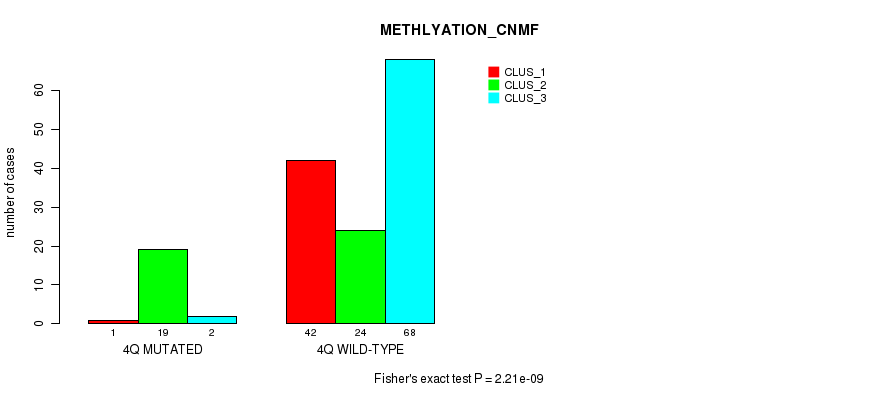
P value = 2.21e-09 (Fisher's exact test), Q value = 6.8e-07
Table S17. Gene #8: '4q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 4Q MUTATED | 1 | 19 | 2 |
| 4Q WILD-TYPE | 42 | 24 | 68 |
Figure S17. Get High-res Image Gene #8: '4q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
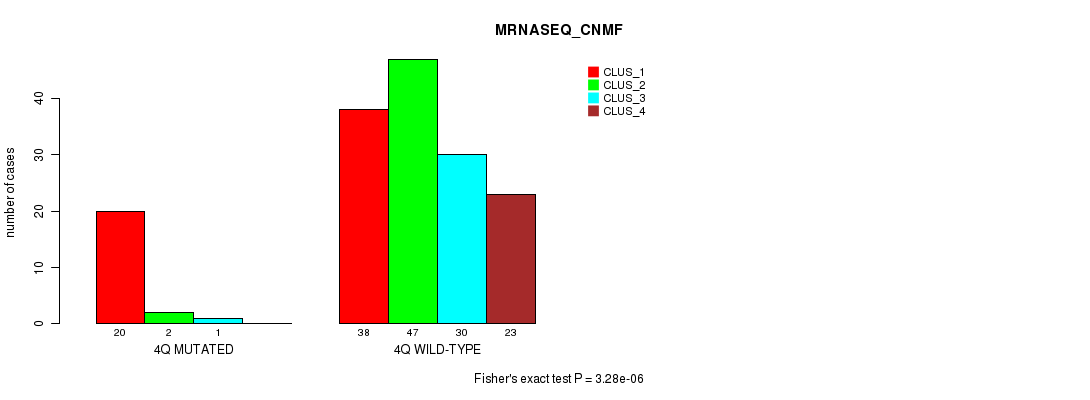
P value = 3.28e-06 (Fisher's exact test), Q value = 9e-04
Table S18. Gene #8: '4q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 4Q MUTATED | 20 | 2 | 1 | 0 |
| 4Q WILD-TYPE | 38 | 47 | 30 | 23 |
Figure S18. Get High-res Image Gene #8: '4q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
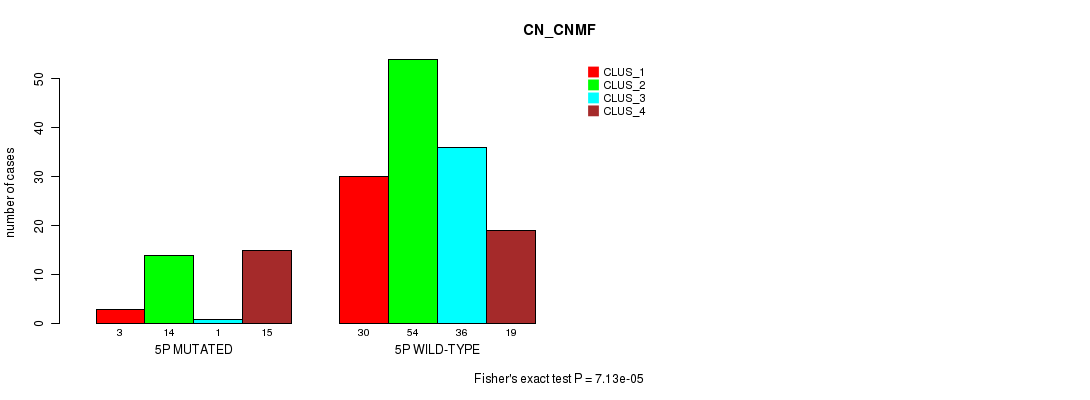
P value = 7.13e-05 (Fisher's exact test), Q value = 0.018
Table S19. Gene #9: '5p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 5P MUTATED | 3 | 14 | 1 | 15 |
| 5P WILD-TYPE | 30 | 54 | 36 | 19 |
Figure S19. Get High-res Image Gene #9: '5p' versus Molecular Subtype #1: 'CN_CNMF'
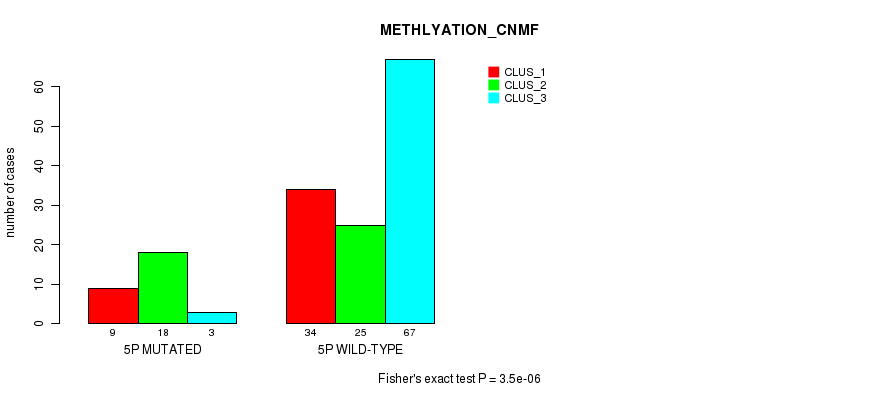
P value = 3.5e-06 (Fisher's exact test), Q value = 0.00095
Table S20. Gene #9: '5p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 5P MUTATED | 9 | 18 | 3 |
| 5P WILD-TYPE | 34 | 25 | 67 |
Figure S20. Get High-res Image Gene #9: '5p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
P value = 0.000122 (Fisher's exact test), Q value = 0.031
Table S21. Gene #9: '5p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 5P MUTATED | 10 | 19 | 3 |
| 5P WILD-TYPE | 21 | 47 | 61 |
Figure S21. Get High-res Image Gene #9: '5p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
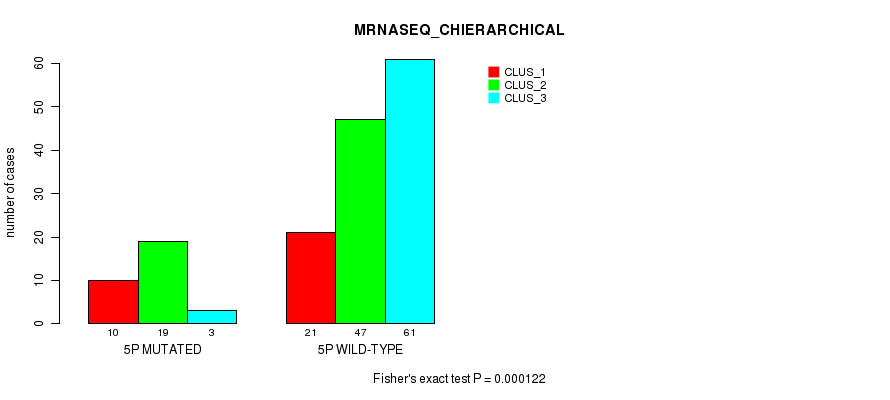
P value = 0.000191 (Fisher's exact test), Q value = 0.047
Table S22. Gene #10: '5q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 5Q MUTATED | 3 | 15 | 1 | 14 |
| 5Q WILD-TYPE | 30 | 53 | 36 | 20 |
Figure S22. Get High-res Image Gene #10: '5q' versus Molecular Subtype #1: 'CN_CNMF'
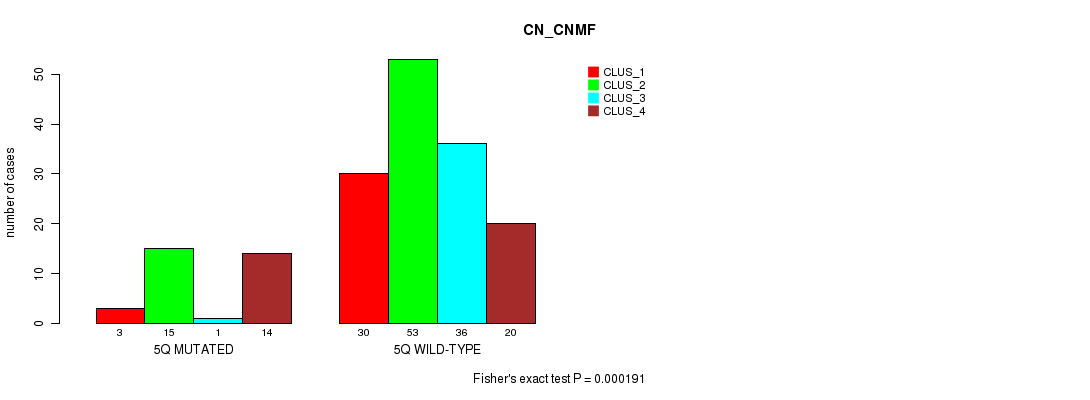
P value = 4.46e-05 (Fisher's exact test), Q value = 0.011
Table S23. Gene #10: '5q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 5Q MUTATED | 9 | 17 | 4 |
| 5Q WILD-TYPE | 34 | 26 | 66 |
Figure S23. Get High-res Image Gene #10: '5q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
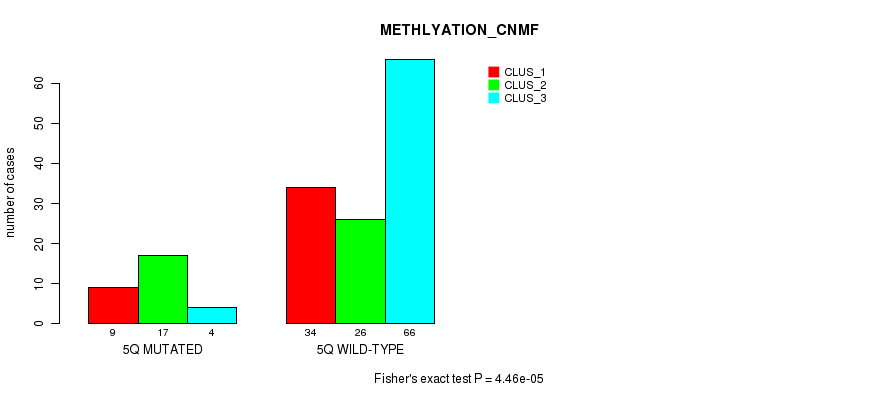
P value = 0.000122 (Fisher's exact test), Q value = 0.031
Table S24. Gene #10: '5q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 5Q MUTATED | 10 | 19 | 3 |
| 5Q WILD-TYPE | 21 | 47 | 61 |
Figure S24. Get High-res Image Gene #10: '5q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
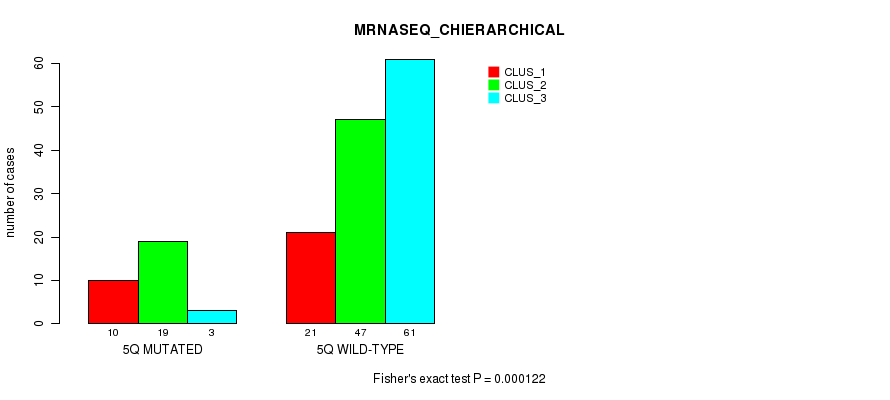
P value = 0.000544 (Fisher's exact test), Q value = 0.13
Table S25. Gene #11: '6p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 6P MUTATED | 1 | 7 | 1 | 11 |
| 6P WILD-TYPE | 32 | 61 | 36 | 23 |
Figure S25. Get High-res Image Gene #11: '6p' versus Molecular Subtype #1: 'CN_CNMF'
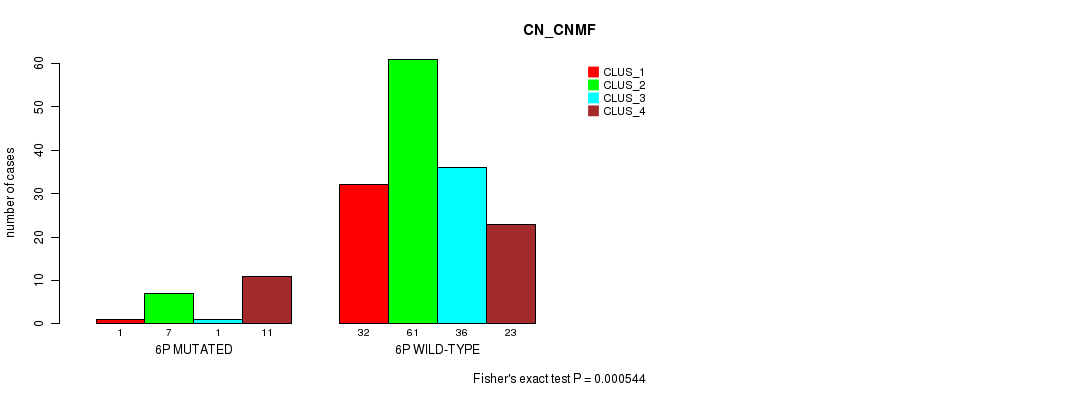
P value = 0.000899 (Fisher's exact test), Q value = 0.2
Table S26. Gene #11: '6p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 6P MUTATED | 2 | 12 | 4 |
| 6P WILD-TYPE | 41 | 31 | 66 |
Figure S26. Get High-res Image Gene #11: '6p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
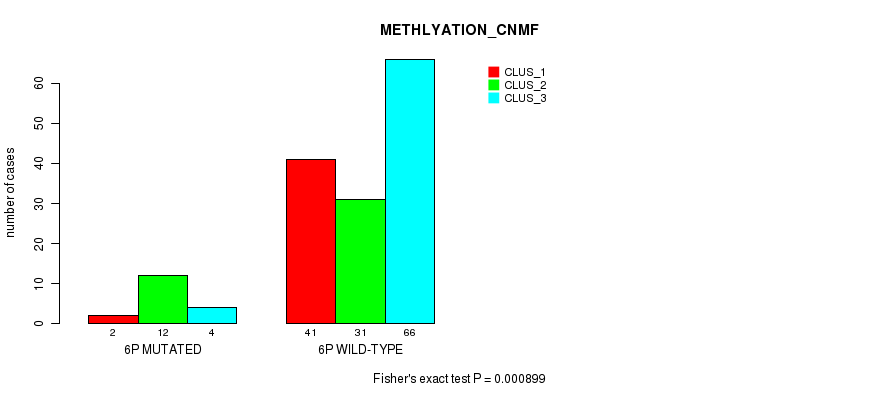
P value = 0.000357 (Fisher's exact test), Q value = 0.086
Table S27. Gene #11: '6p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 6P MUTATED | 15 | 4 | 0 | 0 |
| 6P WILD-TYPE | 43 | 45 | 31 | 23 |
Figure S27. Get High-res Image Gene #11: '6p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 4.01e-06 (Fisher's exact test), Q value = 0.0011
Table S28. Gene #12: '6q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 6Q MUTATED | 1 | 5 | 1 | 14 |
| 6Q WILD-TYPE | 32 | 63 | 36 | 20 |
Figure S28. Get High-res Image Gene #12: '6q' versus Molecular Subtype #1: 'CN_CNMF'
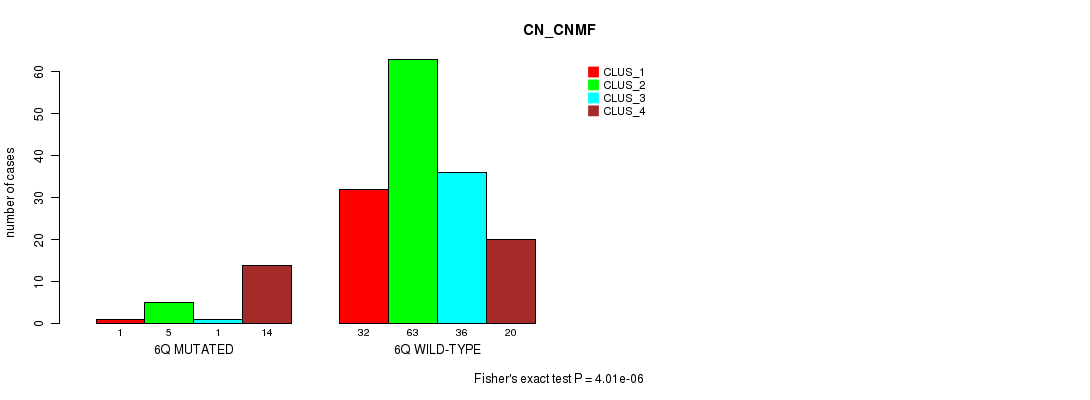
P value = 3.3e-05 (Fisher's exact test), Q value = 0.0085
Table S29. Gene #12: '6q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 6Q MUTATED | 2 | 14 | 3 |
| 6Q WILD-TYPE | 41 | 29 | 67 |
Figure S29. Get High-res Image Gene #12: '6q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
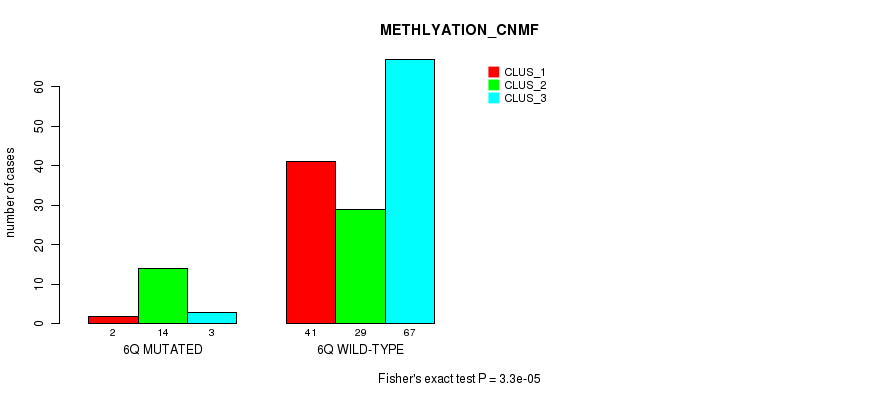
P value = 0.000423 (Fisher's exact test), Q value = 0.1
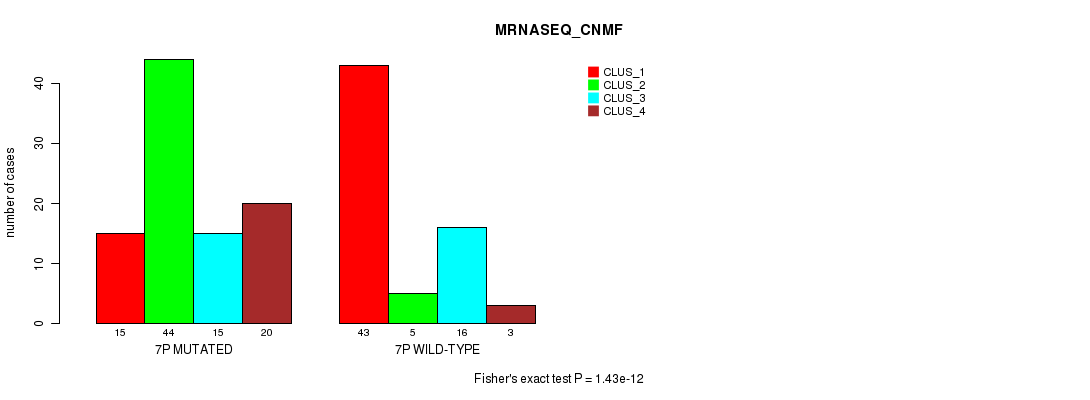
Table S30. Gene #12: '6q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 6Q MUTATED | 16 | 3 | 1 | 0 |
| 6Q WILD-TYPE | 42 | 46 | 30 | 23 |
Figure S30. Get High-res Image Gene #12: '6q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
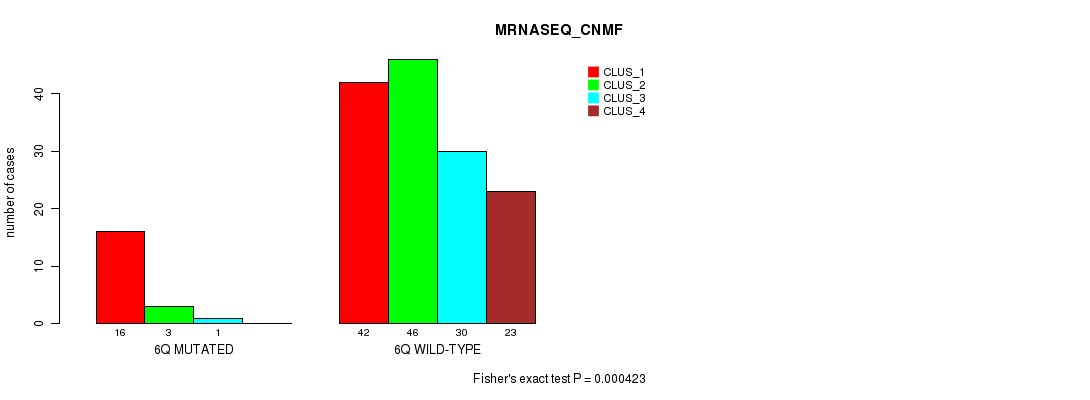
P value = 1.14e-18 (Fisher's exact test), Q value = 3.7e-16
Table S31. Gene #13: '7p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 7P MUTATED | 31 | 19 | 37 | 14 |
| 7P WILD-TYPE | 2 | 49 | 0 | 20 |
Figure S31. Get High-res Image Gene #13: '7p' versus Molecular Subtype #1: 'CN_CNMF'
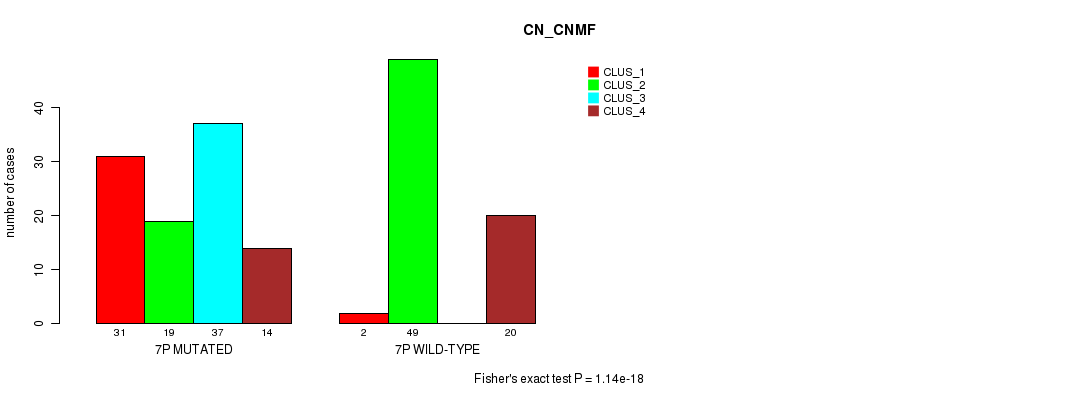
P value = 2.12e-07 (Fisher's exact test), Q value = 6.2e-05
Table S32. Gene #13: '7p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 7P MUTATED | 20 | 14 | 57 |
| 7P WILD-TYPE | 23 | 29 | 13 |
Figure S32. Get High-res Image Gene #13: '7p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
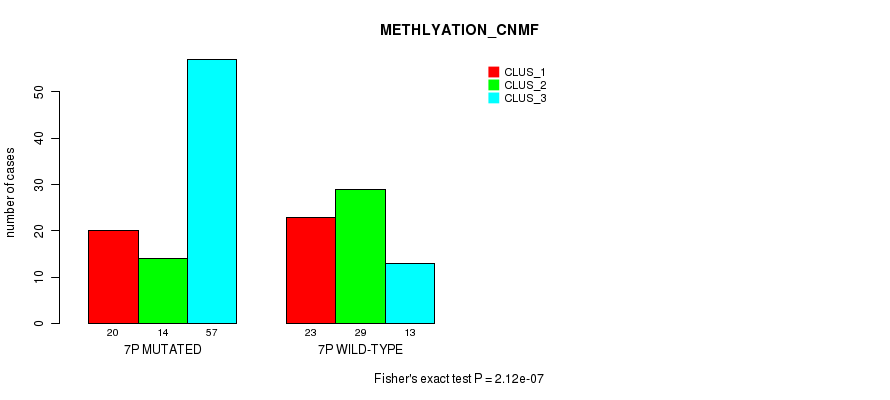
P value = 1.43e-12 (Fisher's exact test), Q value = 4.5e-10
Table S33. Gene #13: '7p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 7P MUTATED | 15 | 44 | 15 | 20 |
| 7P WILD-TYPE | 43 | 5 | 16 | 3 |
Figure S33. Get High-res Image Gene #13: '7p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
P value = 2.01e-07 (Fisher's exact test), Q value = 5.9e-05
Table S34. Gene #13: '7p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 7P MUTATED | 9 | 32 | 53 |
| 7P WILD-TYPE | 22 | 34 | 11 |
Figure S34. Get High-res Image Gene #13: '7p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
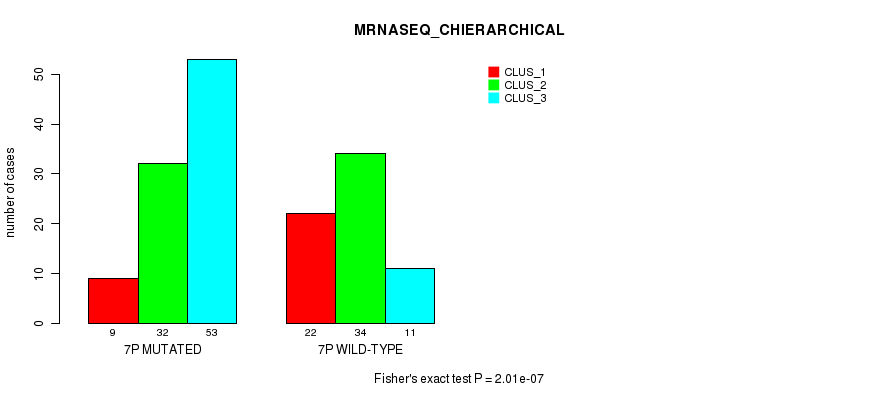
P value = 4e-18 (Fisher's exact test), Q value = 1.3e-15
Table S35. Gene #14: '7q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 7Q MUTATED | 31 | 20 | 37 | 14 |
| 7Q WILD-TYPE | 2 | 48 | 0 | 20 |
Figure S35. Get High-res Image Gene #14: '7q' versus Molecular Subtype #1: 'CN_CNMF'
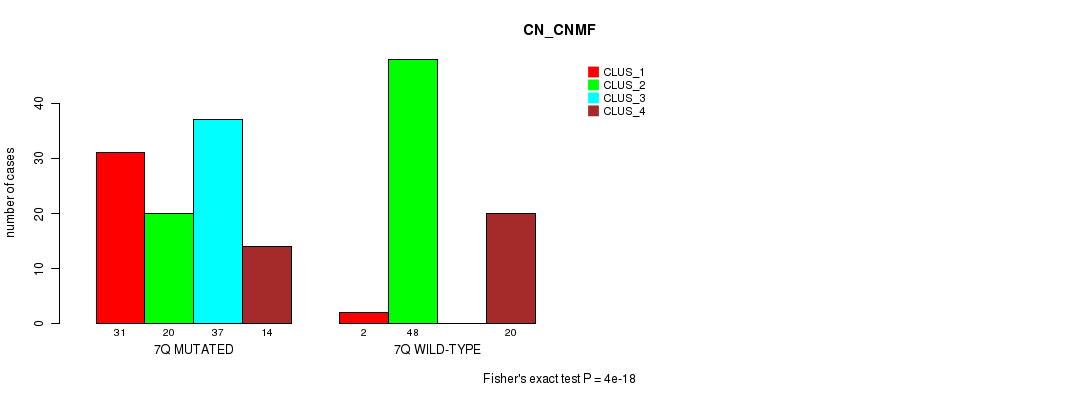
P value = 5.75e-07 (Fisher's exact test), Q value = 0.00017
Table S36. Gene #14: '7q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 7Q MUTATED | 20 | 15 | 57 |
| 7Q WILD-TYPE | 23 | 28 | 13 |
Figure S36. Get High-res Image Gene #14: '7q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
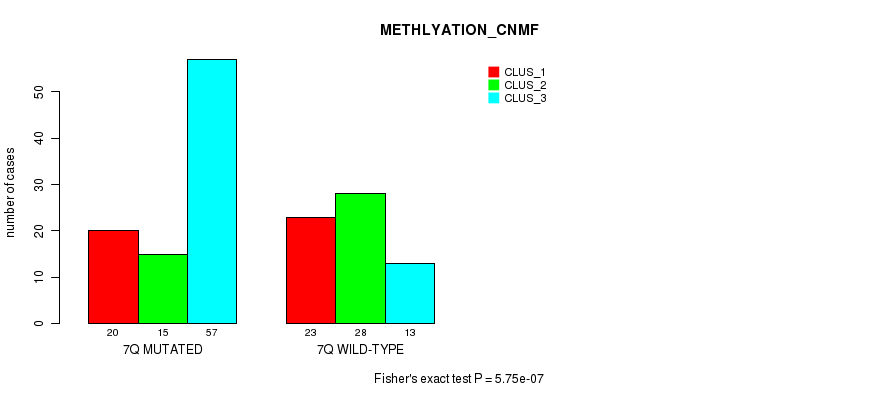
P value = 2.02e-12 (Fisher's exact test), Q value = 6.3e-10
Table S37. Gene #14: '7q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 7Q MUTATED | 15 | 44 | 16 | 20 |
| 7Q WILD-TYPE | 43 | 5 | 15 | 3 |
Figure S37. Get High-res Image Gene #14: '7q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
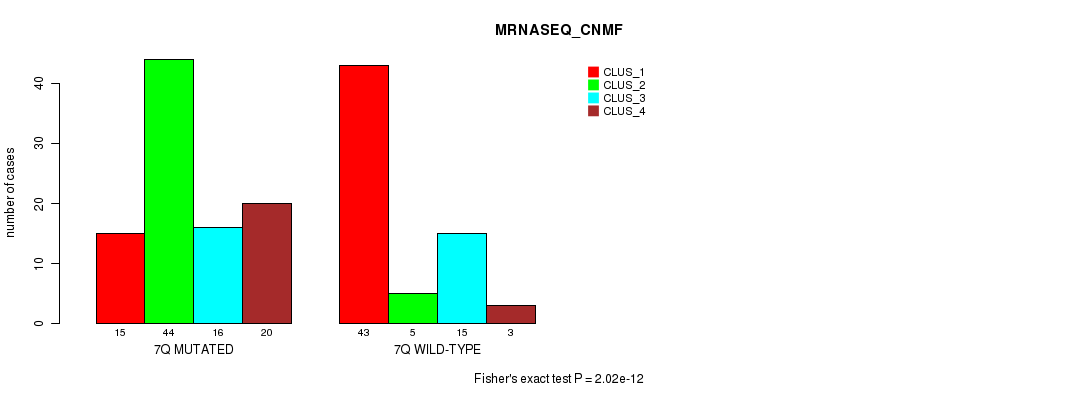
P value = 3.02e-07 (Fisher's exact test), Q value = 8.7e-05
Table S38. Gene #14: '7q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 7Q MUTATED | 9 | 33 | 53 |
| 7Q WILD-TYPE | 22 | 33 | 11 |
Figure S38. Get High-res Image Gene #14: '7q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
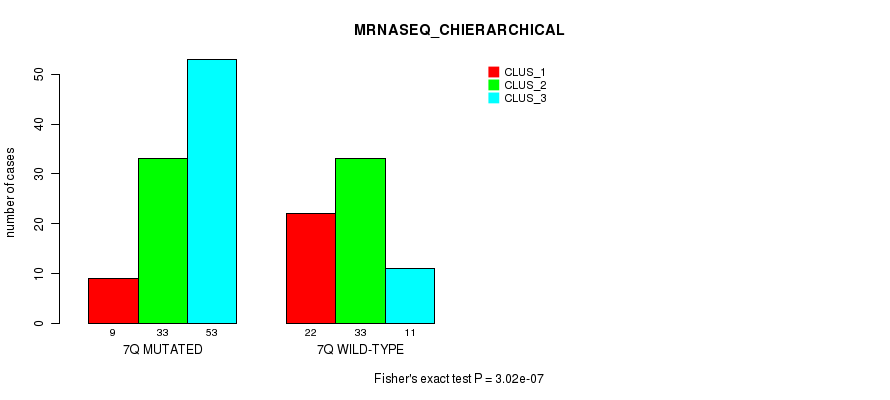
P value = 0.000104 (Fisher's exact test), Q value = 0.026
Table S39. Gene #16: '8q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 8Q MUTATED | 2 | 12 | 2 |
| 8Q WILD-TYPE | 41 | 31 | 68 |
Figure S39. Get High-res Image Gene #16: '8q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
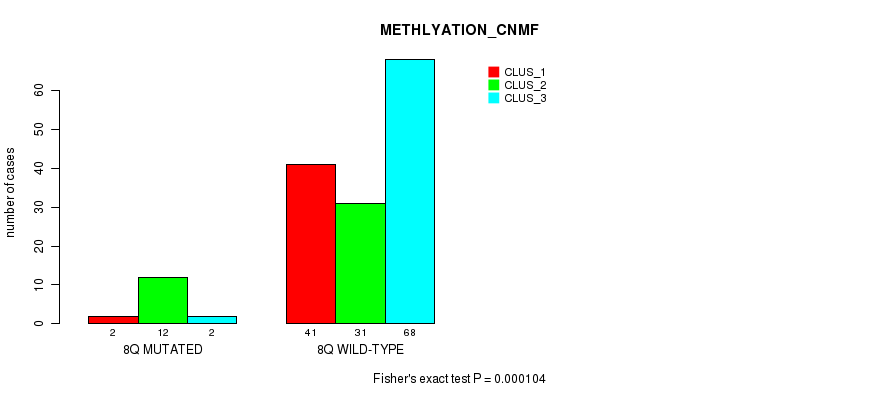
P value = 7.59e-07 (Fisher's exact test), Q value = 0.00022
Table S40. Gene #17: '9p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 9P MUTATED | 1 | 9 | 0 | 15 |
| 9P WILD-TYPE | 32 | 59 | 37 | 19 |
Figure S40. Get High-res Image Gene #17: '9p' versus Molecular Subtype #1: 'CN_CNMF'
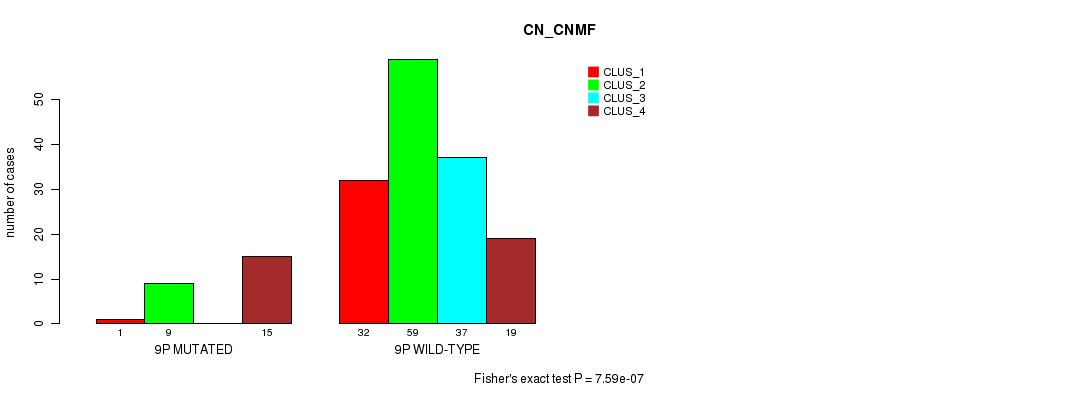
P value = 3.38e-08 (Fisher's exact test), Q value = 1e-05
Table S41. Gene #17: '9p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 9P MUTATED | 5 | 16 | 0 |
| 9P WILD-TYPE | 38 | 27 | 70 |
Figure S41. Get High-res Image Gene #17: '9p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
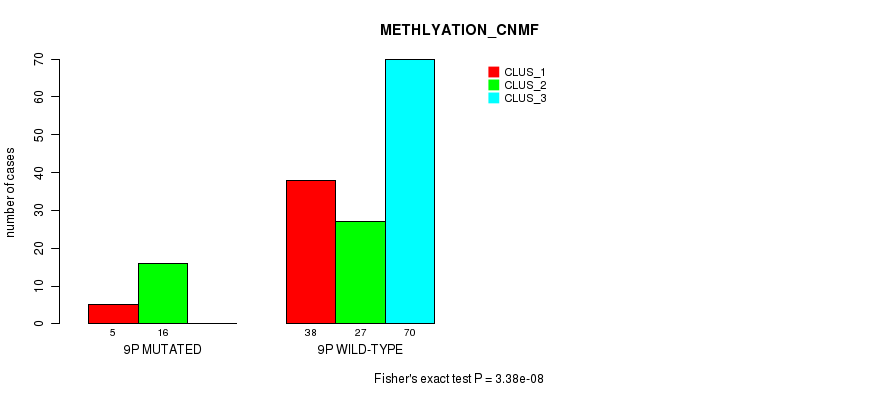
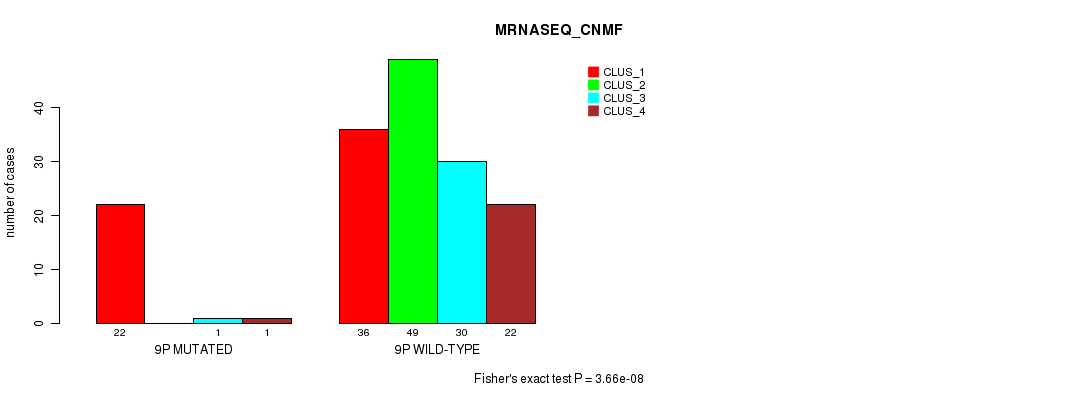
P value = 3.66e-08 (Fisher's exact test), Q value = 1.1e-05
Table S42. Gene #17: '9p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 9P MUTATED | 22 | 0 | 1 | 1 |
| 9P WILD-TYPE | 36 | 49 | 30 | 22 |
Figure S42. Get High-res Image Gene #17: '9p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
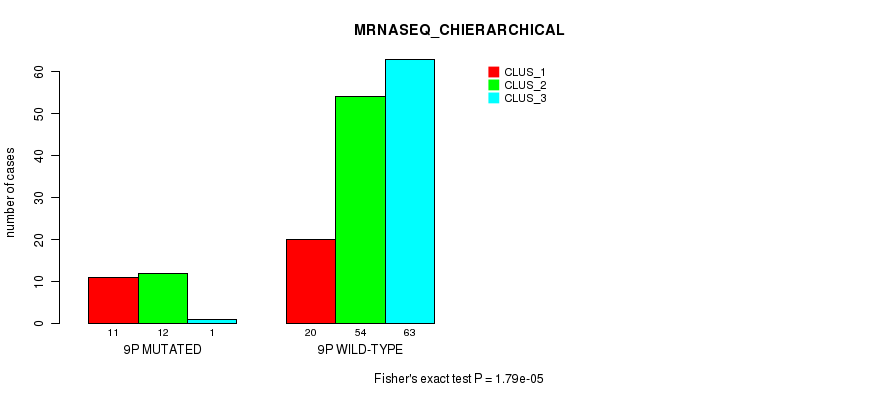
P value = 1.79e-05 (Fisher's exact test), Q value = 0.0047
Table S43. Gene #17: '9p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 9P MUTATED | 11 | 12 | 1 |
| 9P WILD-TYPE | 20 | 54 | 63 |
Figure S43. Get High-res Image Gene #17: '9p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
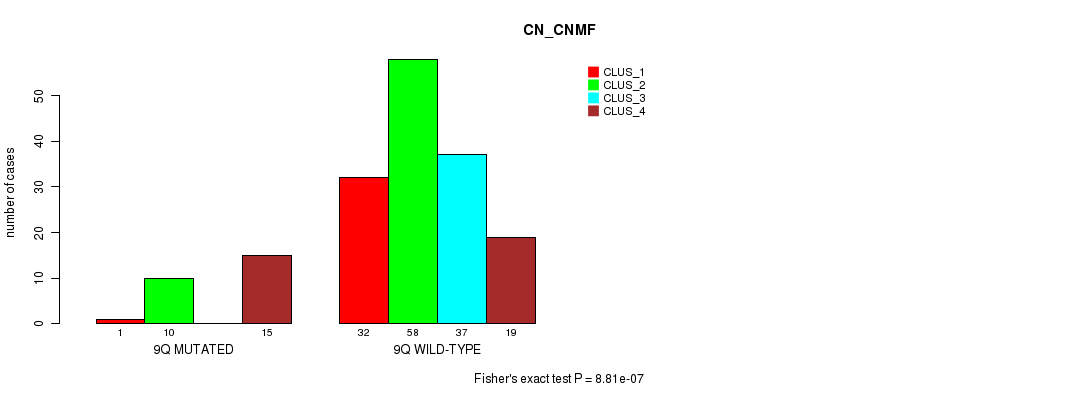
P value = 8.81e-07 (Fisher's exact test), Q value = 0.00025
Table S44. Gene #18: '9q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 9Q MUTATED | 1 | 10 | 0 | 15 |
| 9Q WILD-TYPE | 32 | 58 | 37 | 19 |
Figure S44. Get High-res Image Gene #18: '9q' versus Molecular Subtype #1: 'CN_CNMF'
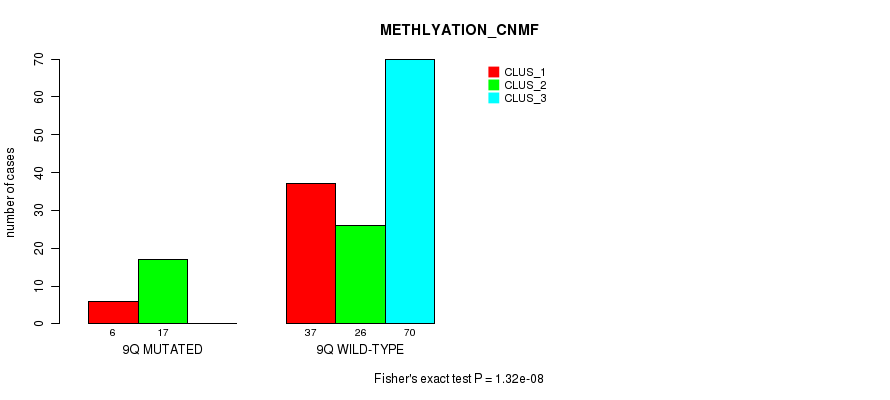
P value = 1.32e-08 (Fisher's exact test), Q value = 4e-06
Table S45. Gene #18: '9q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 9Q MUTATED | 6 | 17 | 0 |
| 9Q WILD-TYPE | 37 | 26 | 70 |
Figure S45. Get High-res Image Gene #18: '9q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
P value = 1.17e-08 (Fisher's exact test), Q value = 3.6e-06
Table S46. Gene #18: '9q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 9Q MUTATED | 23 | 0 | 1 | 1 |
| 9Q WILD-TYPE | 35 | 49 | 30 | 22 |
Figure S46. Get High-res Image Gene #18: '9q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
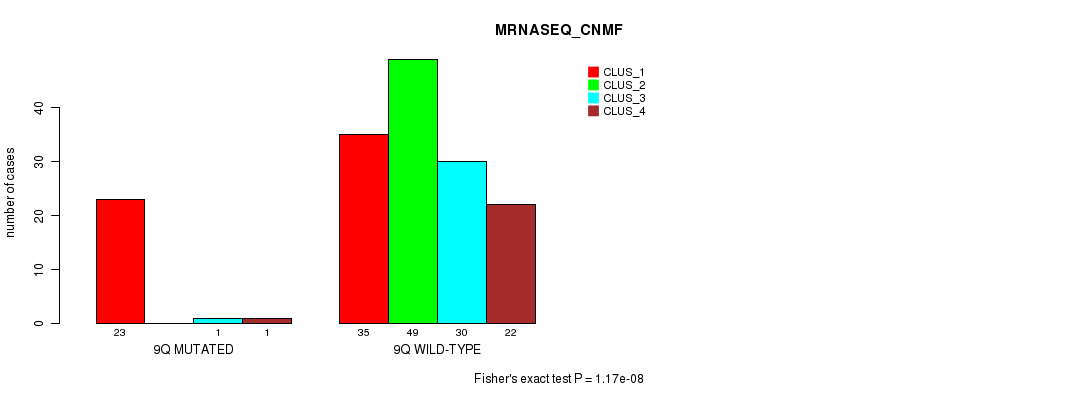
P value = 1.09e-07 (Fisher's exact test), Q value = 3.2e-05
Table S47. Gene #18: '9q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 9Q MUTATED | 13 | 12 | 0 |
| 9Q WILD-TYPE | 18 | 54 | 64 |
Figure S47. Get High-res Image Gene #18: '9q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
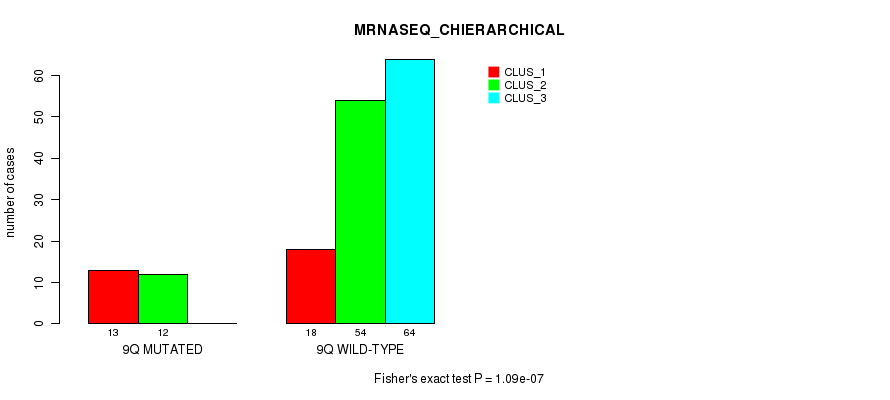
P value = 1.3e-05 (Fisher's exact test), Q value = 0.0035
Table S48. Gene #18: '9q' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 80 | 74 |
| 9Q MUTATED | 6 | 2 | 18 |
| 9Q WILD-TYPE | 12 | 78 | 56 |
Figure S48. Get High-res Image Gene #18: '9q' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
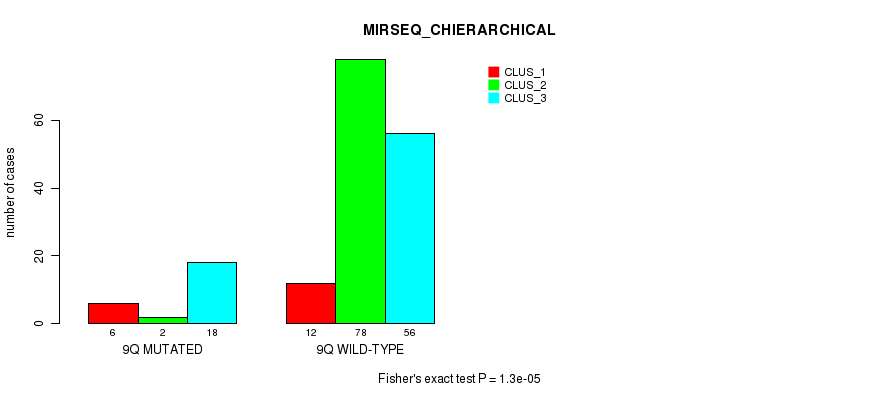
P value = 1.83e-05 (Fisher's exact test), Q value = 0.0048
Table S49. Gene #18: '9q' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 14 | 87 | 71 |
| 9Q MUTATED | 5 | 3 | 18 |
| 9Q WILD-TYPE | 9 | 84 | 53 |
Figure S49. Get High-res Image Gene #18: '9q' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
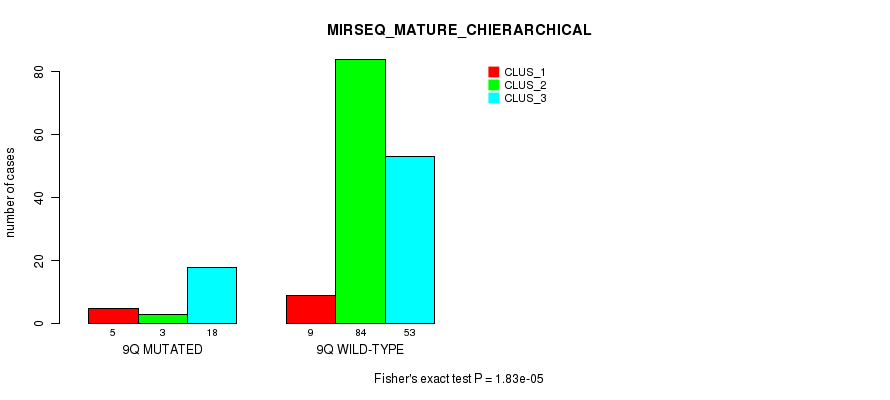
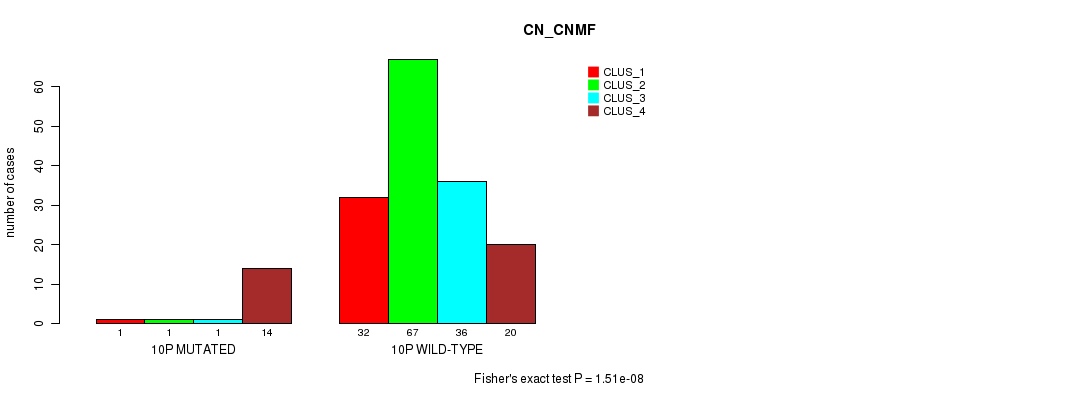
P value = 1.51e-08 (Fisher's exact test), Q value = 4.6e-06
Table S50. Gene #19: '10p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 10P MUTATED | 1 | 1 | 1 | 14 |
| 10P WILD-TYPE | 32 | 67 | 36 | 20 |
Figure S50. Get High-res Image Gene #19: '10p' versus Molecular Subtype #1: 'CN_CNMF'
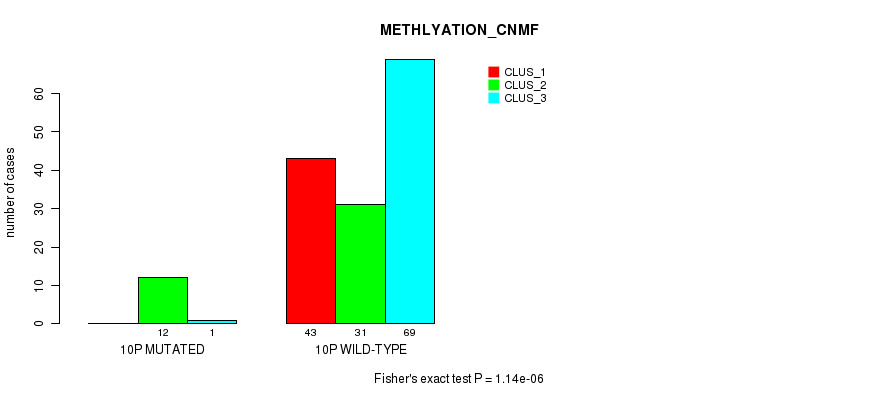
P value = 1.14e-06 (Fisher's exact test), Q value = 0.00032
Table S51. Gene #19: '10p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 10P MUTATED | 0 | 12 | 1 |
| 10P WILD-TYPE | 43 | 31 | 69 |
Figure S51. Get High-res Image Gene #19: '10p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
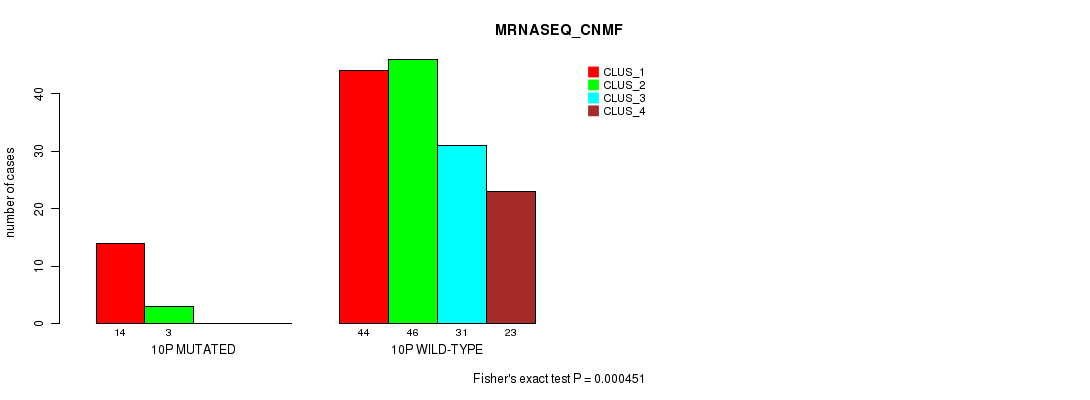
P value = 0.000451 (Fisher's exact test), Q value = 0.11
Table S52. Gene #19: '10p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 10P MUTATED | 14 | 3 | 0 | 0 |
| 10P WILD-TYPE | 44 | 46 | 31 | 23 |
Figure S52. Get High-res Image Gene #19: '10p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
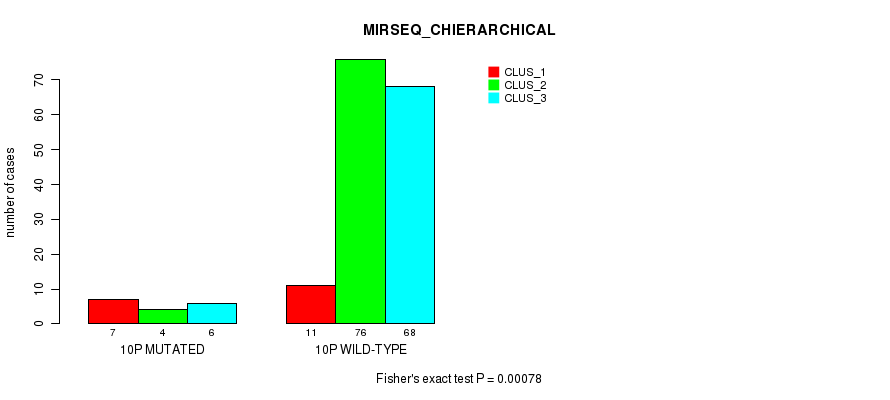
P value = 0.00078 (Fisher's exact test), Q value = 0.18
Table S53. Gene #19: '10p' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 80 | 74 |
| 10P MUTATED | 7 | 4 | 6 |
| 10P WILD-TYPE | 11 | 76 | 68 |
Figure S53. Get High-res Image Gene #19: '10p' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
P value = 6.87e-07 (Fisher's exact test), Q value = 2e-04
Table S54. Gene #20: '10q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 10Q MUTATED | 1 | 2 | 1 | 13 |
| 10Q WILD-TYPE | 32 | 66 | 36 | 21 |
Figure S54. Get High-res Image Gene #20: '10q' versus Molecular Subtype #1: 'CN_CNMF'
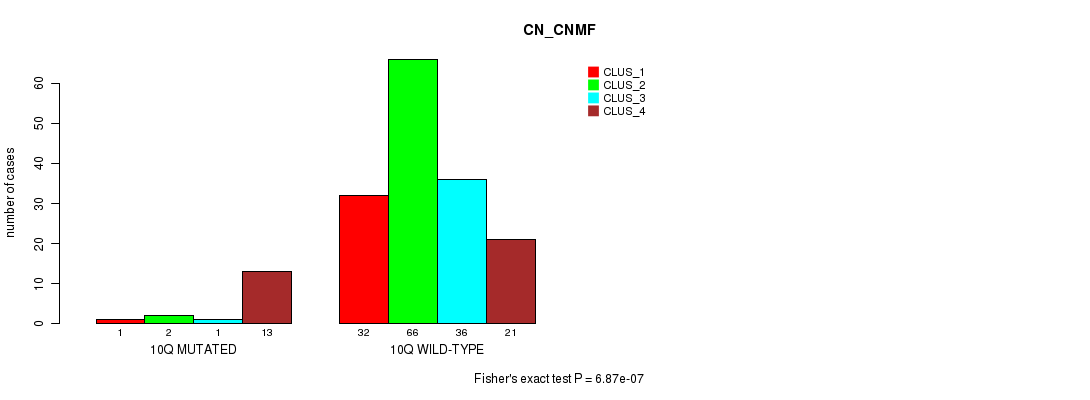
P value = 2.4e-07 (Fisher's exact test), Q value = 7e-05
Table S55. Gene #20: '10q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 10Q MUTATED | 0 | 13 | 1 |
| 10Q WILD-TYPE | 43 | 30 | 69 |
Figure S55. Get High-res Image Gene #20: '10q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
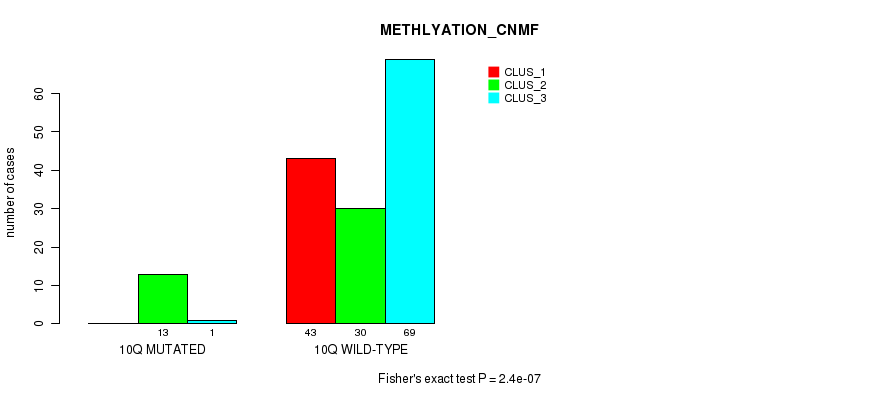
P value = 0.000451 (Fisher's exact test), Q value = 0.11
Table S56. Gene #20: '10q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 10Q MUTATED | 14 | 3 | 0 | 0 |
| 10Q WILD-TYPE | 44 | 46 | 31 | 23 |
Figure S56. Get High-res Image Gene #20: '10q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
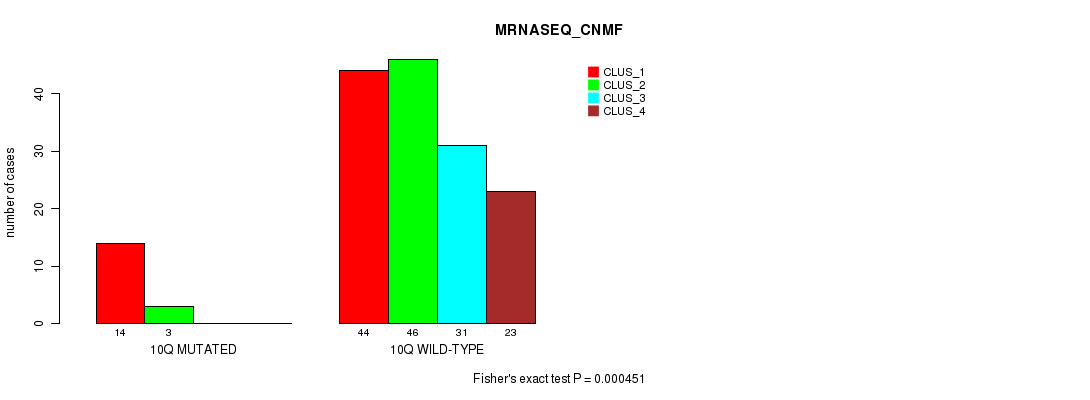
P value = 0.000372 (Fisher's exact test), Q value = 0.089
Table S57. Gene #20: '10q' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 80 | 74 |
| 10Q MUTATED | 7 | 3 | 7 |
| 10Q WILD-TYPE | 11 | 77 | 67 |
Figure S57. Get High-res Image Gene #20: '10q' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
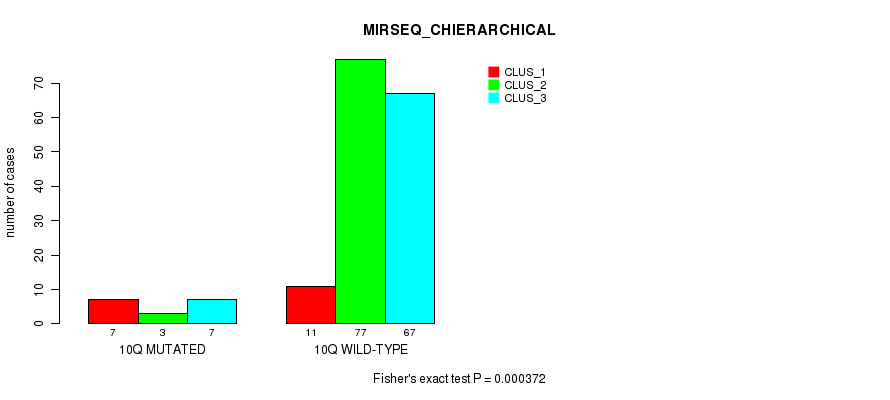
P value = 0.000635 (Fisher's exact test), Q value = 0.15
Table S58. Gene #20: '10q' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 14 | 87 | 71 |
| 10Q MUTATED | 6 | 4 | 7 |
| 10Q WILD-TYPE | 8 | 83 | 64 |
Figure S58. Get High-res Image Gene #20: '10q' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
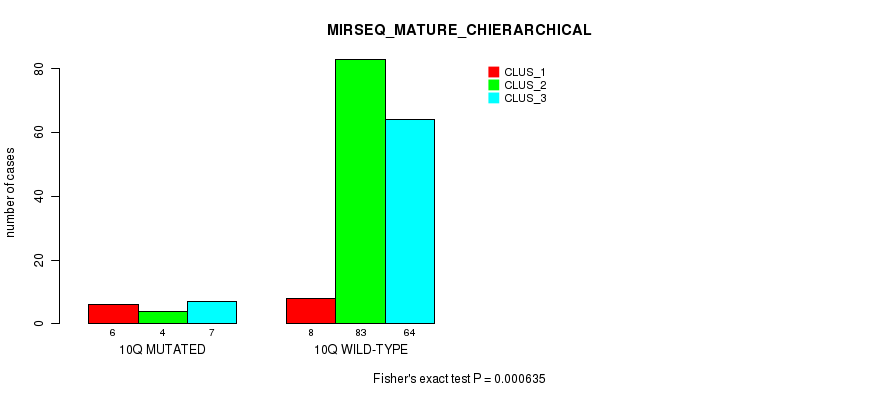
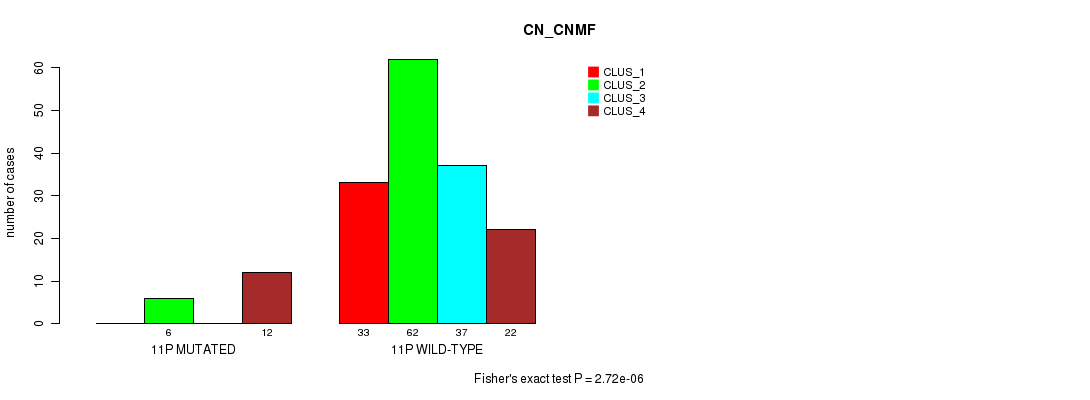
P value = 2.72e-06 (Fisher's exact test), Q value = 0.00075
Table S59. Gene #21: '11p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 11P MUTATED | 0 | 6 | 0 | 12 |
| 11P WILD-TYPE | 33 | 62 | 37 | 22 |
Figure S59. Get High-res Image Gene #21: '11p' versus Molecular Subtype #1: 'CN_CNMF'
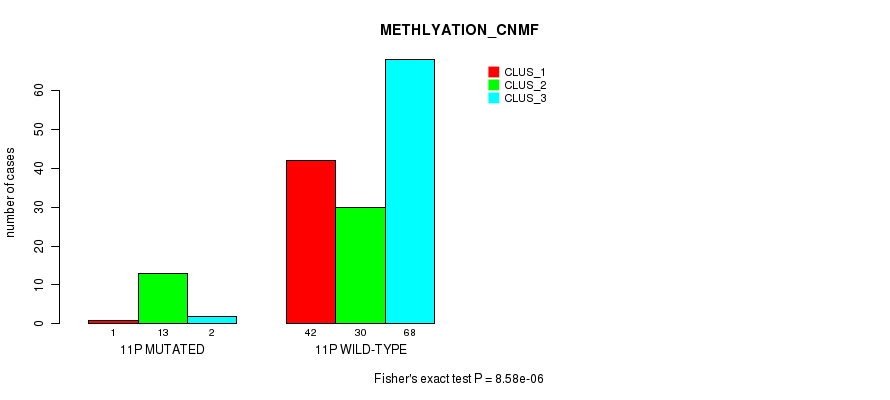
P value = 8.58e-06 (Fisher's exact test), Q value = 0.0023
Table S60. Gene #21: '11p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 11P MUTATED | 1 | 13 | 2 |
| 11P WILD-TYPE | 42 | 30 | 68 |
Figure S60. Get High-res Image Gene #21: '11p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
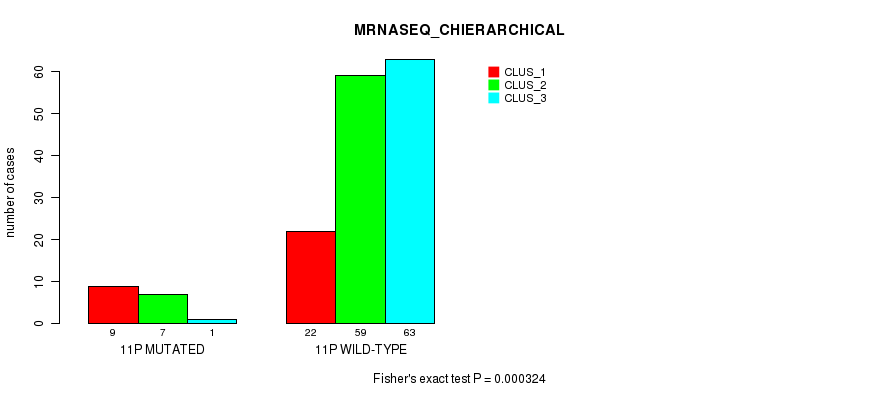
P value = 0.000324 (Fisher's exact test), Q value = 0.078
Table S61. Gene #21: '11p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 11P MUTATED | 9 | 7 | 1 |
| 11P WILD-TYPE | 22 | 59 | 63 |
Figure S61. Get High-res Image Gene #21: '11p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
P value = 3.45e-08 (Fisher's exact test), Q value = 1e-05
Table S62. Gene #22: '11q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 11Q MUTATED | 0 | 6 | 0 | 15 |
| 11Q WILD-TYPE | 33 | 62 | 37 | 19 |
Figure S62. Get High-res Image Gene #22: '11q' versus Molecular Subtype #1: 'CN_CNMF'
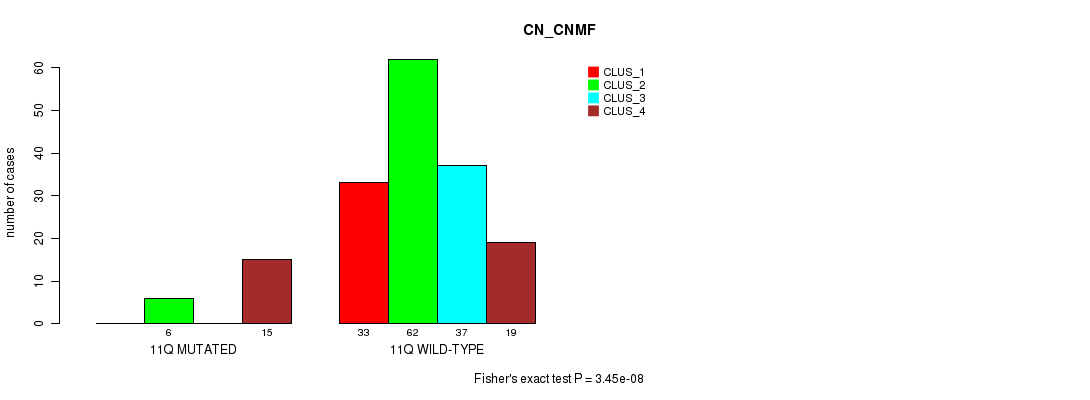
P value = 4.06e-08 (Fisher's exact test), Q value = 1.2e-05
Table S63. Gene #22: '11q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 11Q MUTATED | 1 | 17 | 2 |
| 11Q WILD-TYPE | 42 | 26 | 68 |
Figure S63. Get High-res Image Gene #22: '11q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
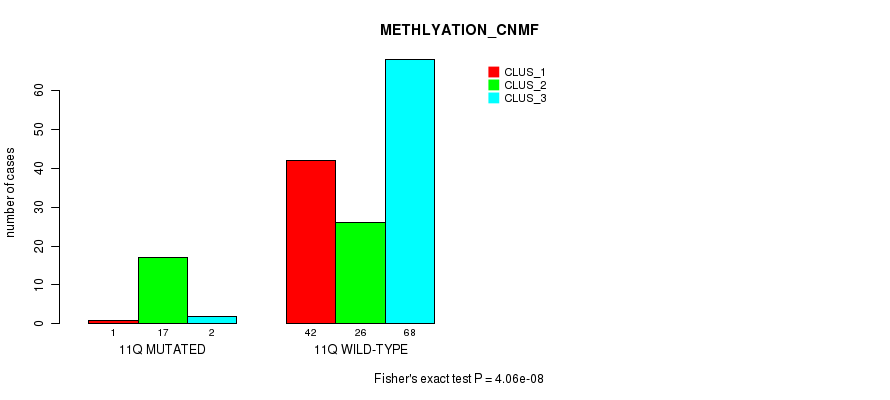
P value = 0.000532 (Fisher's exact test), Q value = 0.12
Table S64. Gene #22: '11q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 11Q MUTATED | 8 | 11 | 1 |
| 11Q WILD-TYPE | 23 | 55 | 63 |
Figure S64. Get High-res Image Gene #22: '11q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
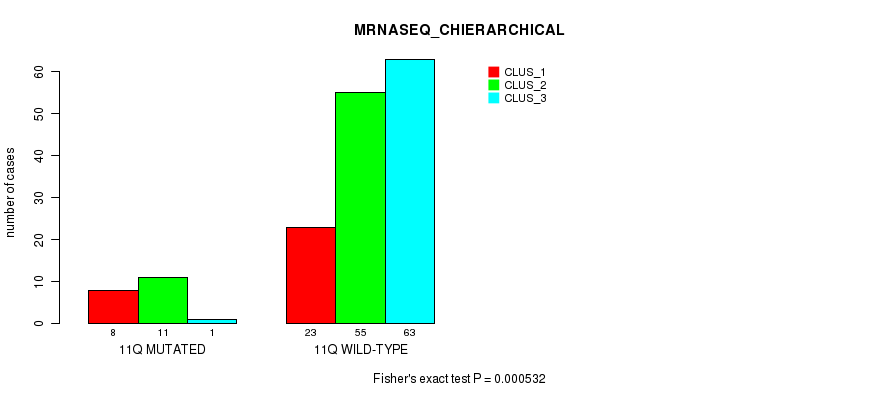
P value = 0.000301 (Fisher's exact test), Q value = 0.073
Table S65. Gene #23: '12p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 12P MUTATED | 21 | 16 | 20 | 12 |
| 12P WILD-TYPE | 12 | 52 | 17 | 22 |
Figure S65. Get High-res Image Gene #23: '12p' versus Molecular Subtype #1: 'CN_CNMF'
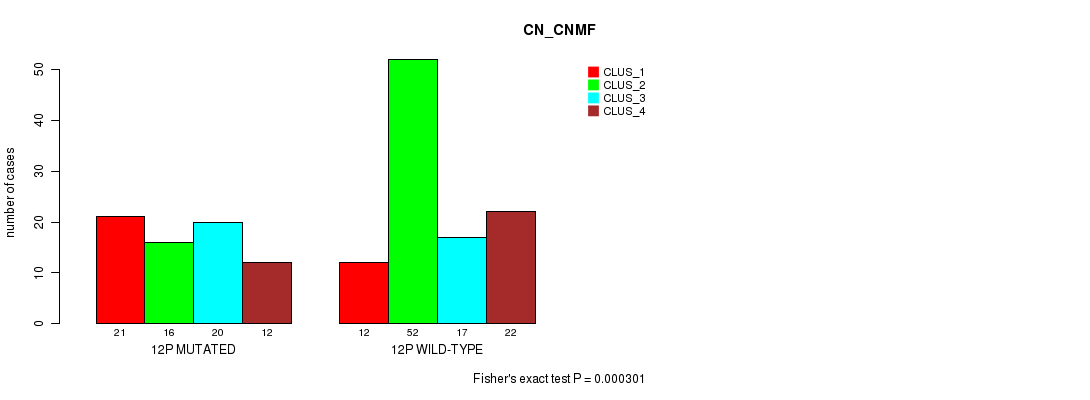
P value = 0.000236 (Fisher's exact test), Q value = 0.058
Table S66. Gene #24: '12q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 12Q MUTATED | 21 | 16 | 20 | 11 |
| 12Q WILD-TYPE | 12 | 52 | 17 | 23 |
Figure S66. Get High-res Image Gene #24: '12q' versus Molecular Subtype #1: 'CN_CNMF'
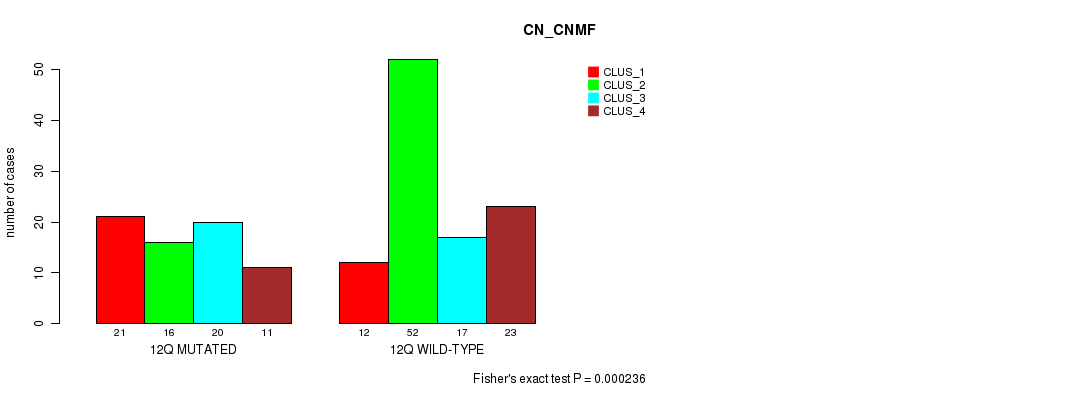
P value = 5.57e-05 (Fisher's exact test), Q value = 0.014
Table S67. Gene #25: '13q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 13Q MUTATED | 7 | 10 | 2 | 17 |
| 13Q WILD-TYPE | 26 | 58 | 35 | 17 |
Figure S67. Get High-res Image Gene #25: '13q' versus Molecular Subtype #1: 'CN_CNMF'
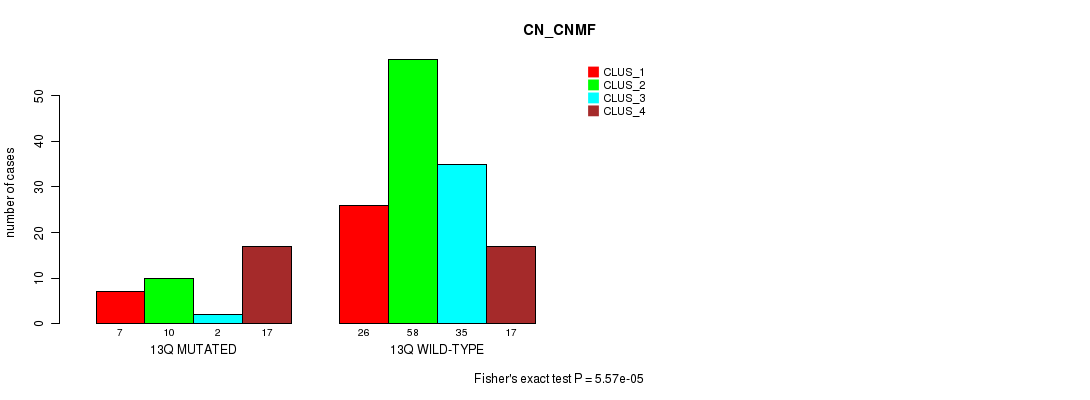
P value = 0.000199 (Fisher's exact test), Q value = 0.049
Table S68. Gene #25: '13q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 13Q MUTATED | 5 | 19 | 9 |
| 13Q WILD-TYPE | 38 | 24 | 61 |
Figure S68. Get High-res Image Gene #25: '13q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
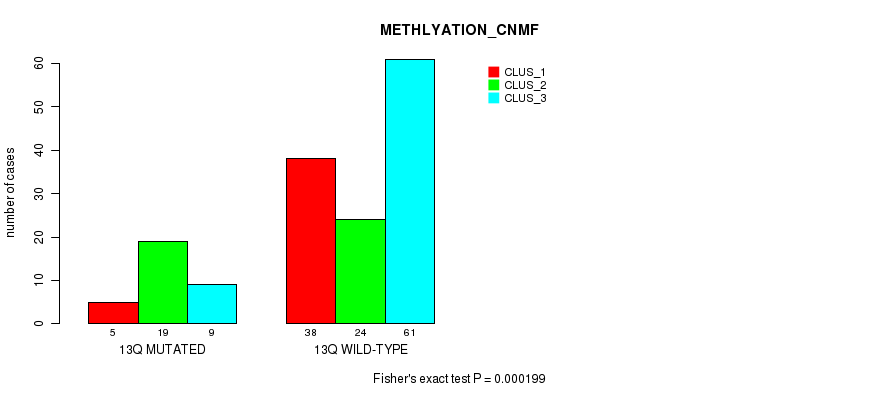
P value = 0.000755 (Fisher's exact test), Q value = 0.17
Table S69. Gene #25: '13q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 13Q MUTATED | 15 | 13 | 8 |
| 13Q WILD-TYPE | 16 | 53 | 56 |
Figure S69. Get High-res Image Gene #25: '13q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
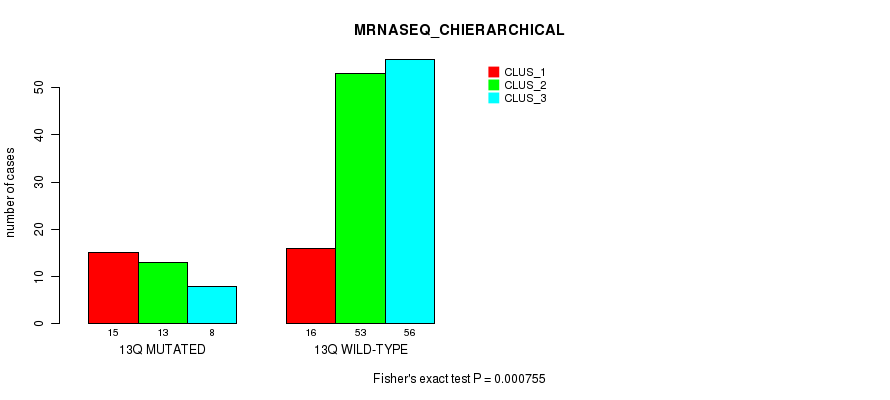
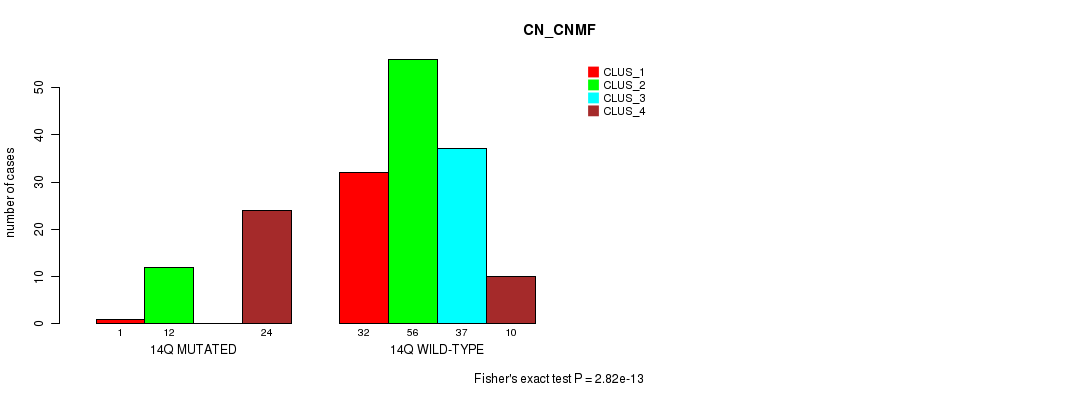
P value = 2.82e-13 (Fisher's exact test), Q value = 8.9e-11
Table S70. Gene #26: '14q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 14Q MUTATED | 1 | 12 | 0 | 24 |
| 14Q WILD-TYPE | 32 | 56 | 37 | 10 |
Figure S70. Get High-res Image Gene #26: '14q' versus Molecular Subtype #1: 'CN_CNMF'
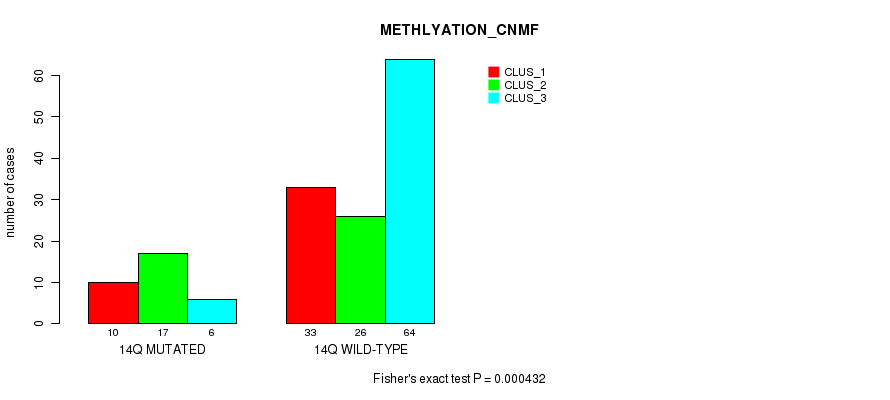
P value = 0.000432 (Fisher's exact test), Q value = 0.1
Table S71. Gene #26: '14q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 14Q MUTATED | 10 | 17 | 6 |
| 14Q WILD-TYPE | 33 | 26 | 64 |
Figure S71. Get High-res Image Gene #26: '14q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
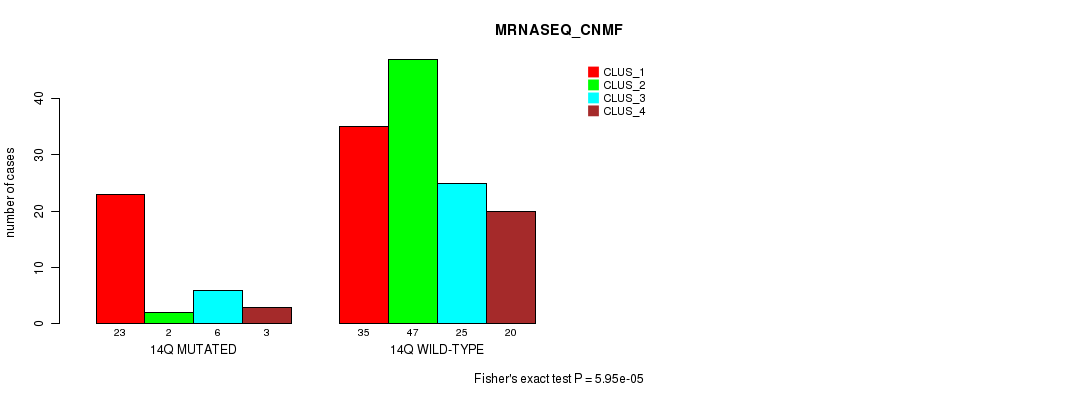
P value = 5.95e-05 (Fisher's exact test), Q value = 0.015
Table S72. Gene #26: '14q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 14Q MUTATED | 23 | 2 | 6 | 3 |
| 14Q WILD-TYPE | 35 | 47 | 25 | 20 |
Figure S72. Get High-res Image Gene #26: '14q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
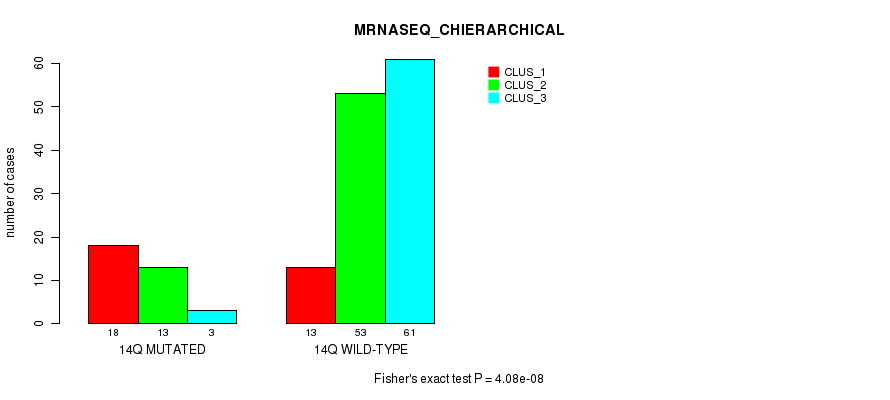
P value = 4.08e-08 (Fisher's exact test), Q value = 1.2e-05
Table S73. Gene #26: '14q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 14Q MUTATED | 18 | 13 | 3 |
| 14Q WILD-TYPE | 13 | 53 | 61 |
Figure S73. Get High-res Image Gene #26: '14q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
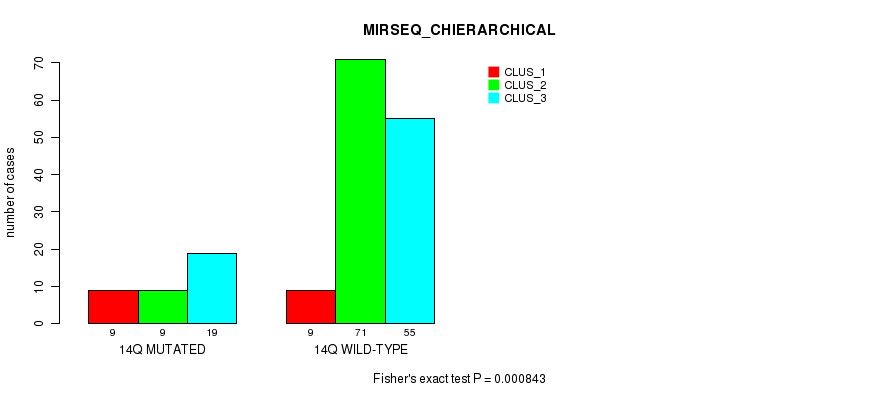
P value = 0.000843 (Fisher's exact test), Q value = 0.19
Table S74. Gene #26: '14q' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 80 | 74 |
| 14Q MUTATED | 9 | 9 | 19 |
| 14Q WILD-TYPE | 9 | 71 | 55 |
Figure S74. Get High-res Image Gene #26: '14q' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
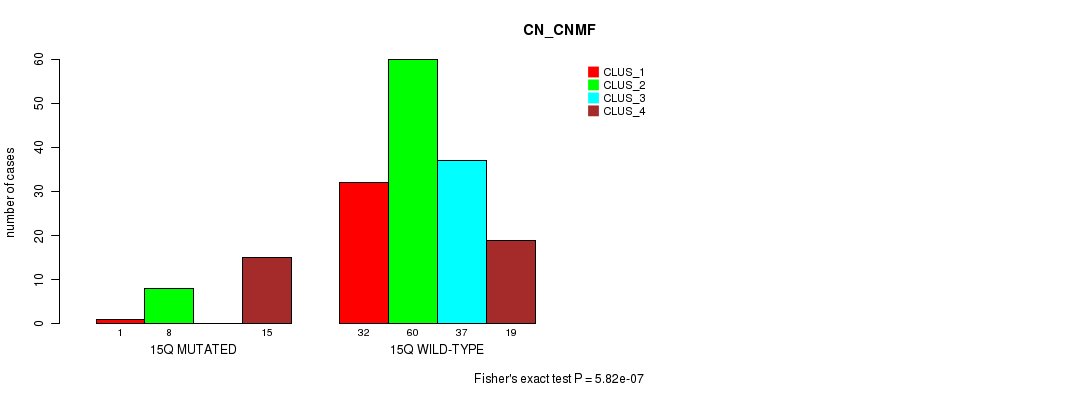
P value = 5.82e-07 (Fisher's exact test), Q value = 0.00017
Table S75. Gene #27: '15q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 15Q MUTATED | 1 | 8 | 0 | 15 |
| 15Q WILD-TYPE | 32 | 60 | 37 | 19 |
Figure S75. Get High-res Image Gene #27: '15q' versus Molecular Subtype #1: 'CN_CNMF'
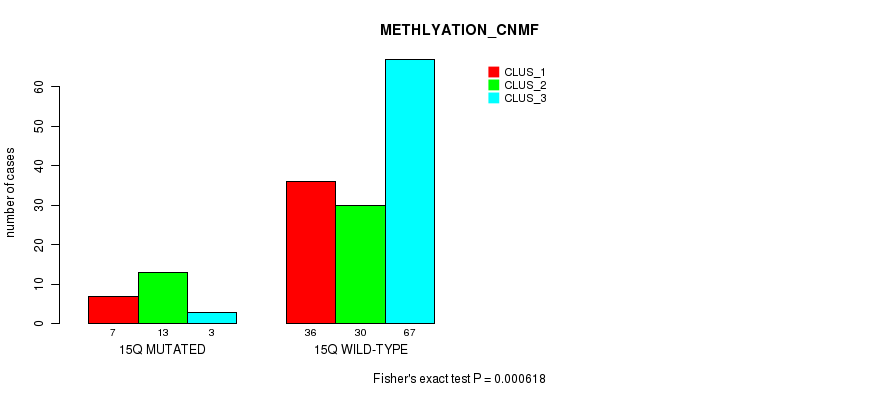
P value = 0.000618 (Fisher's exact test), Q value = 0.14
Table S76. Gene #27: '15q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 15Q MUTATED | 7 | 13 | 3 |
| 15Q WILD-TYPE | 36 | 30 | 67 |
Figure S76. Get High-res Image Gene #27: '15q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
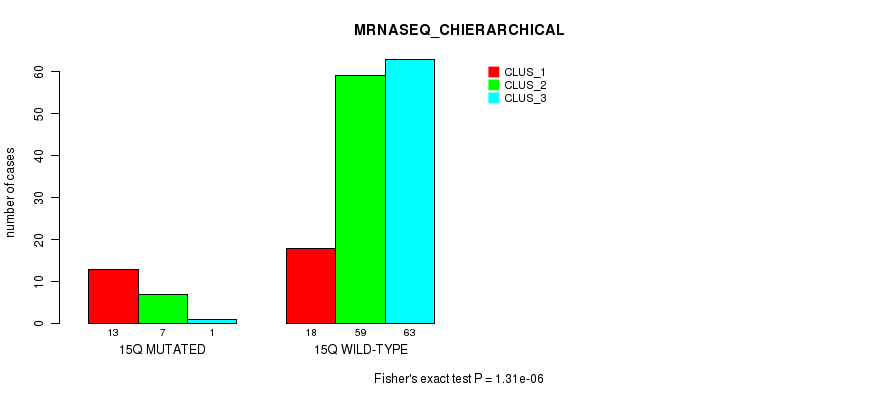
P value = 1.31e-06 (Fisher's exact test), Q value = 0.00037
Table S77. Gene #27: '15q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 15Q MUTATED | 13 | 7 | 1 |
| 15Q WILD-TYPE | 18 | 59 | 63 |
Figure S77. Get High-res Image Gene #27: '15q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
P value = 1.68e-09 (Fisher's exact test), Q value = 5.2e-07
Table S78. Gene #28: '16p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 16P MUTATED | 33 | 29 | 15 | 16 |
| 16P WILD-TYPE | 0 | 39 | 22 | 18 |
Figure S78. Get High-res Image Gene #28: '16p' versus Molecular Subtype #1: 'CN_CNMF'
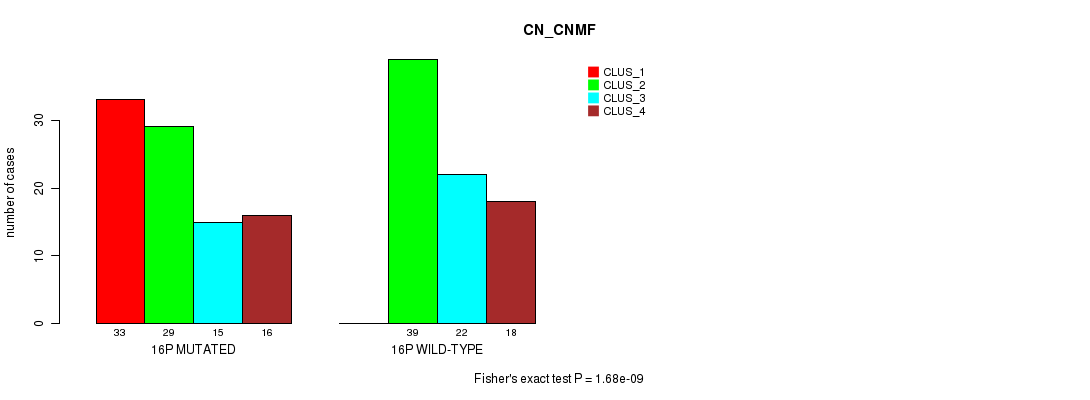
P value = 4.87e-10 (Fisher's exact test), Q value = 1.5e-07
Table S79. Gene #29: '16q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 16Q MUTATED | 33 | 29 | 14 | 14 |
| 16Q WILD-TYPE | 0 | 39 | 23 | 20 |
Figure S79. Get High-res Image Gene #29: '16q' versus Molecular Subtype #1: 'CN_CNMF'
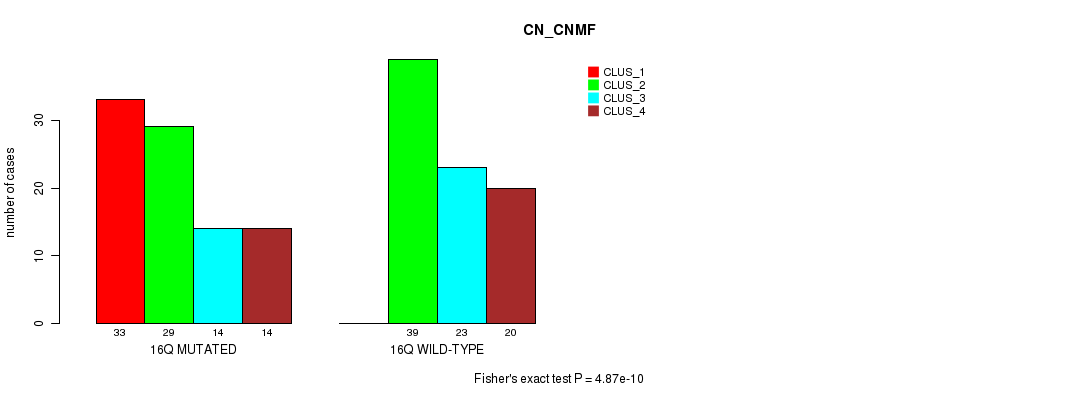
P value = 1.05e-14 (Fisher's exact test), Q value = 3.3e-12
Table S80. Gene #30: '17p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 17P MUTATED | 32 | 24 | 36 | 18 |
| 17P WILD-TYPE | 1 | 44 | 1 | 16 |
Figure S80. Get High-res Image Gene #30: '17p' versus Molecular Subtype #1: 'CN_CNMF'
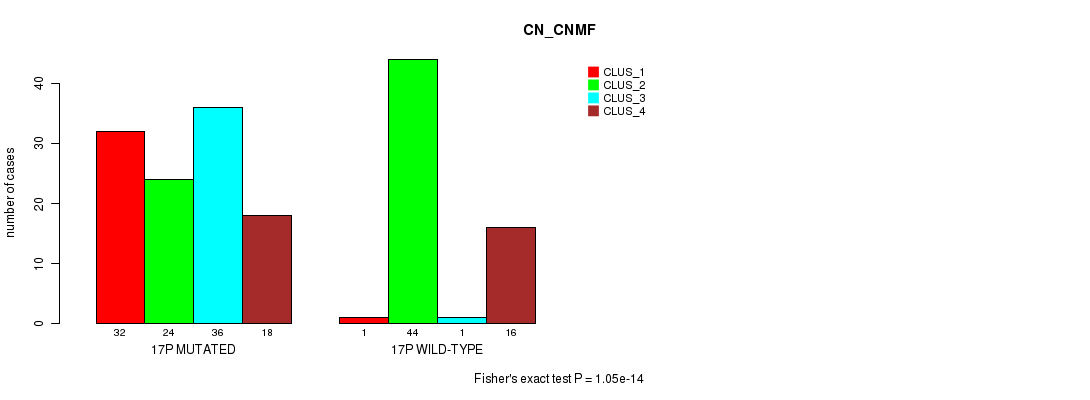
P value = 1.76e-05 (Fisher's exact test), Q value = 0.0046
Table S81. Gene #30: '17p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 17P MUTATED | 19 | 22 | 58 |
| 17P WILD-TYPE | 24 | 21 | 12 |
Figure S81. Get High-res Image Gene #30: '17p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
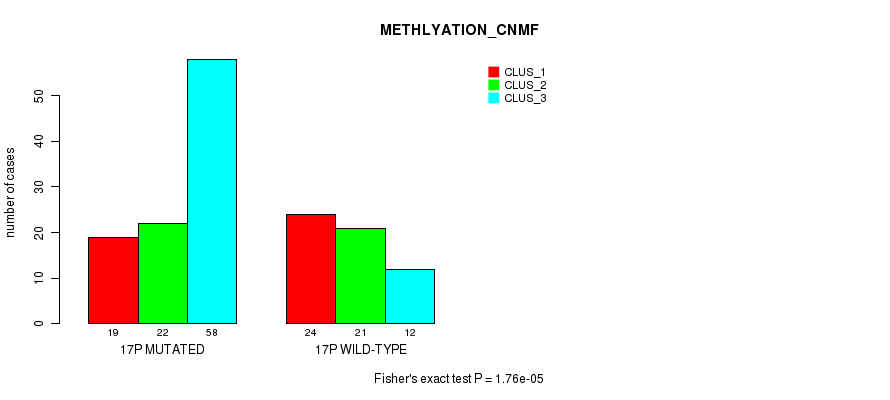
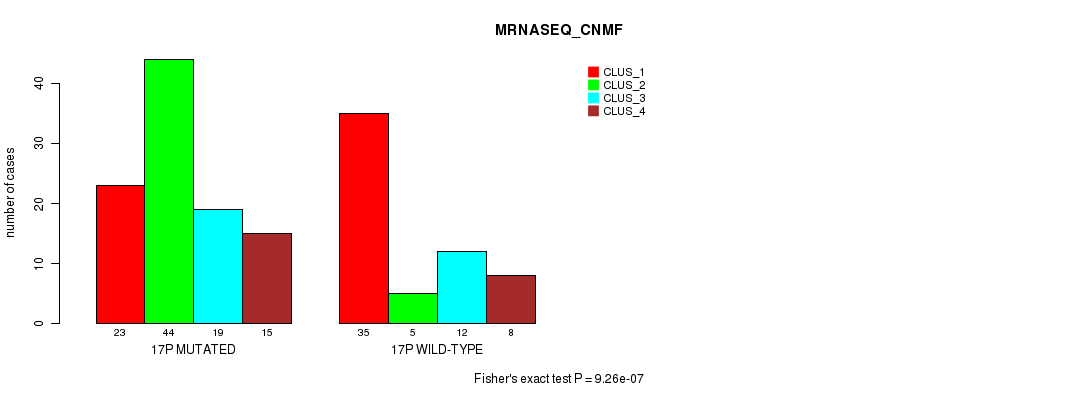
P value = 9.26e-07 (Fisher's exact test), Q value = 0.00026
Table S82. Gene #30: '17p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 17P MUTATED | 23 | 44 | 19 | 15 |
| 17P WILD-TYPE | 35 | 5 | 12 | 8 |
Figure S82. Get High-res Image Gene #30: '17p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
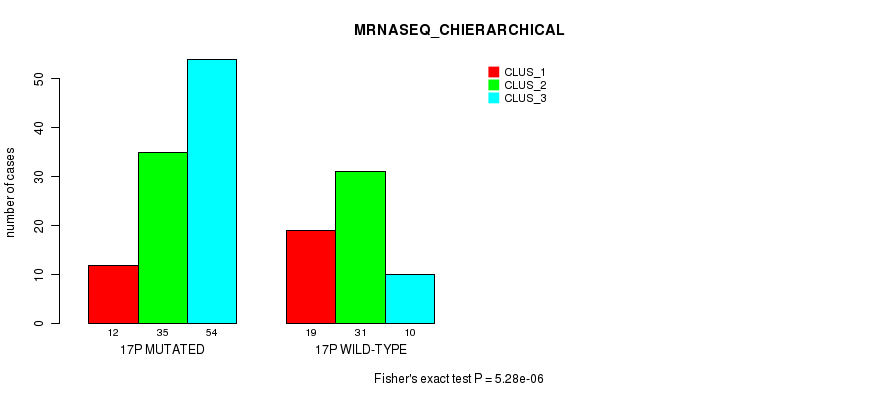
P value = 5.28e-06 (Fisher's exact test), Q value = 0.0014
Table S83. Gene #30: '17p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 17P MUTATED | 12 | 35 | 54 |
| 17P WILD-TYPE | 19 | 31 | 10 |
Figure S83. Get High-res Image Gene #30: '17p' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
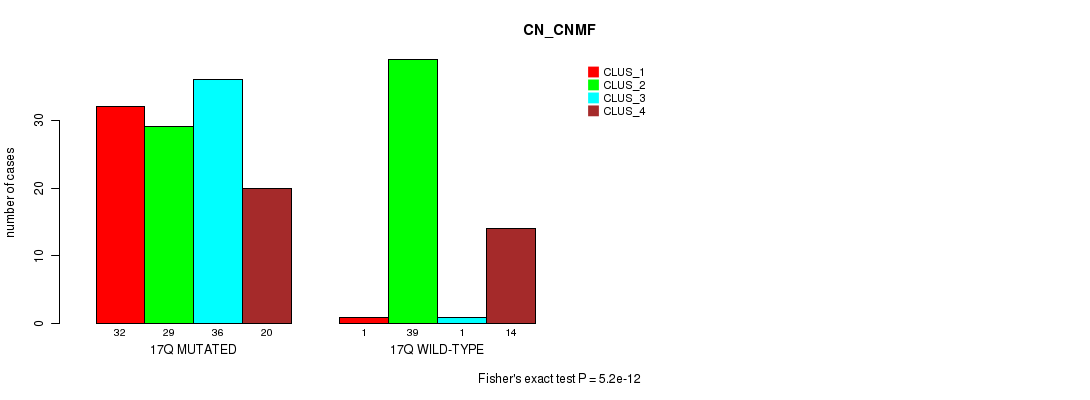
P value = 5.2e-12 (Fisher's exact test), Q value = 1.6e-09
Table S84. Gene #31: '17q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 17Q MUTATED | 32 | 29 | 36 | 20 |
| 17Q WILD-TYPE | 1 | 39 | 1 | 14 |
Figure S84. Get High-res Image Gene #31: '17q' versus Molecular Subtype #1: 'CN_CNMF'
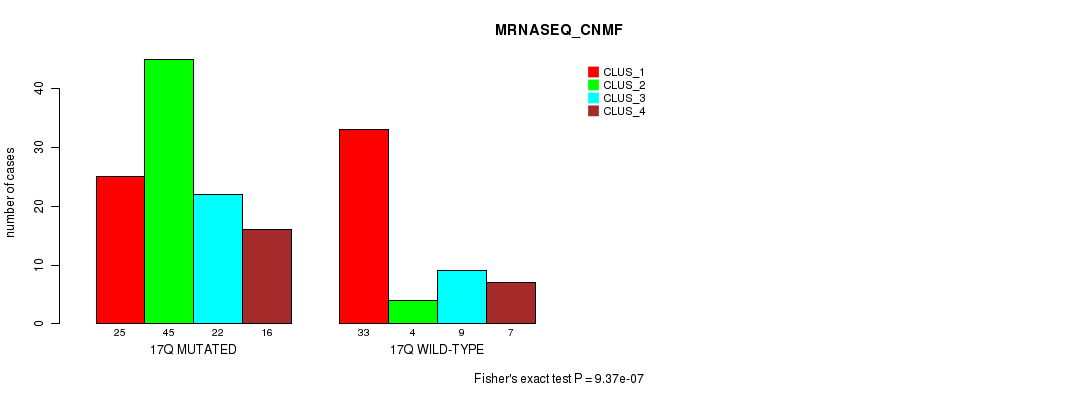
P value = 9.37e-07 (Fisher's exact test), Q value = 0.00026
Table S85. Gene #31: '17q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 17Q MUTATED | 25 | 45 | 22 | 16 |
| 17Q WILD-TYPE | 33 | 4 | 9 | 7 |
Figure S85. Get High-res Image Gene #31: '17q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
P value = 1.09e-05 (Fisher's exact test), Q value = 0.0029
Table S86. Gene #31: '17q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 17Q MUTATED | 12 | 41 | 55 |
| 17Q WILD-TYPE | 19 | 25 | 9 |
Figure S86. Get High-res Image Gene #31: '17q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
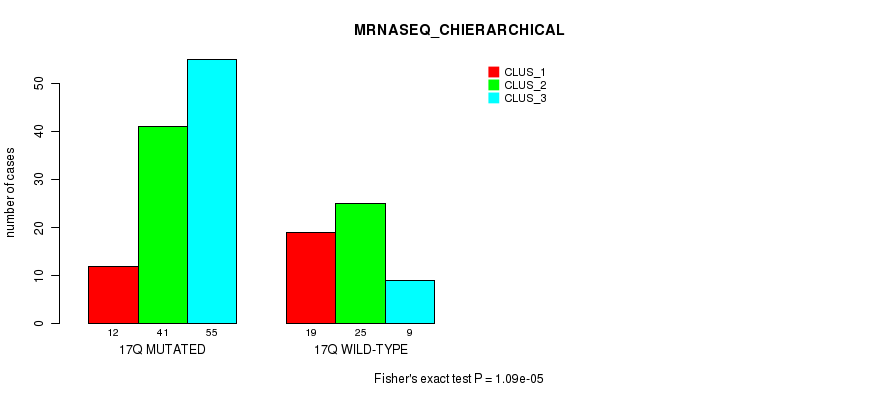
P value = 1.54e-05 (Fisher's exact test), Q value = 0.0041
Table S87. Gene #32: '18p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 18P MUTATED | 4 | 20 | 8 |
| 18P WILD-TYPE | 39 | 23 | 62 |
Figure S87. Get High-res Image Gene #32: '18p' versus Molecular Subtype #2: 'METHLYATION_CNMF'
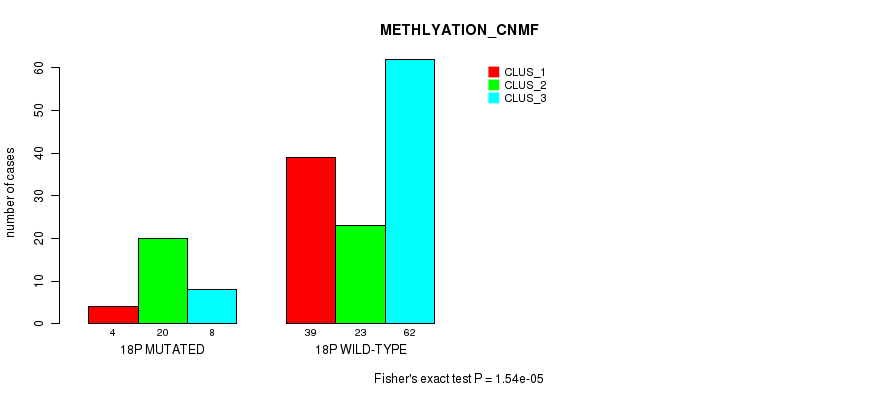
P value = 3.04e-05 (Fisher's exact test), Q value = 0.0078
Table S88. Gene #32: '18p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 18P MUTATED | 24 | 3 | 4 | 2 |
| 18P WILD-TYPE | 34 | 46 | 27 | 21 |
Figure S88. Get High-res Image Gene #32: '18p' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
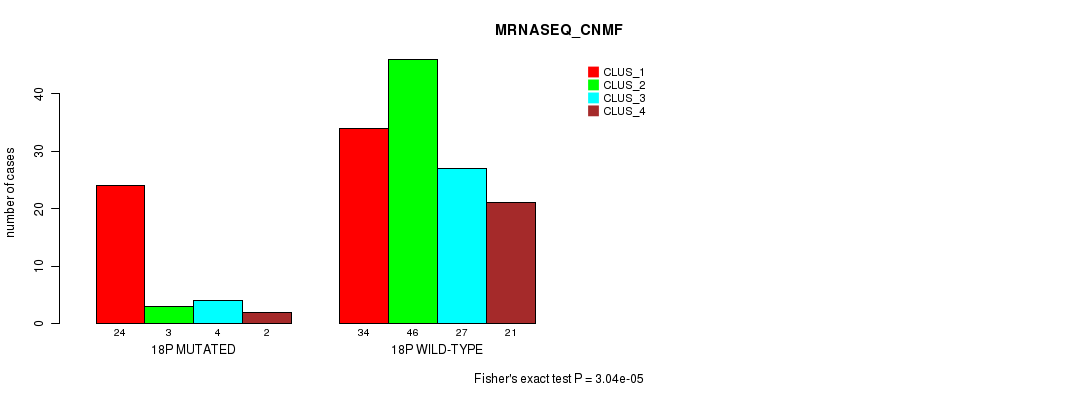
P value = 7.14e-06 (Fisher's exact test), Q value = 0.0019
Table S89. Gene #33: '18q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 18Q MUTATED | 4 | 20 | 7 |
| 18Q WILD-TYPE | 39 | 23 | 63 |
Figure S89. Get High-res Image Gene #33: '18q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
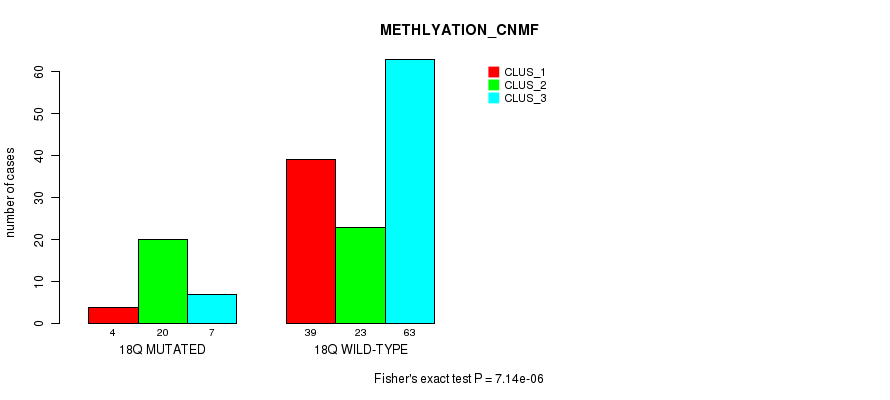
P value = 1.97e-05 (Fisher's exact test), Q value = 0.0051
Table S90. Gene #33: '18q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 18Q MUTATED | 23 | 2 | 4 | 2 |
| 18Q WILD-TYPE | 35 | 47 | 27 | 21 |
Figure S90. Get High-res Image Gene #33: '18q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
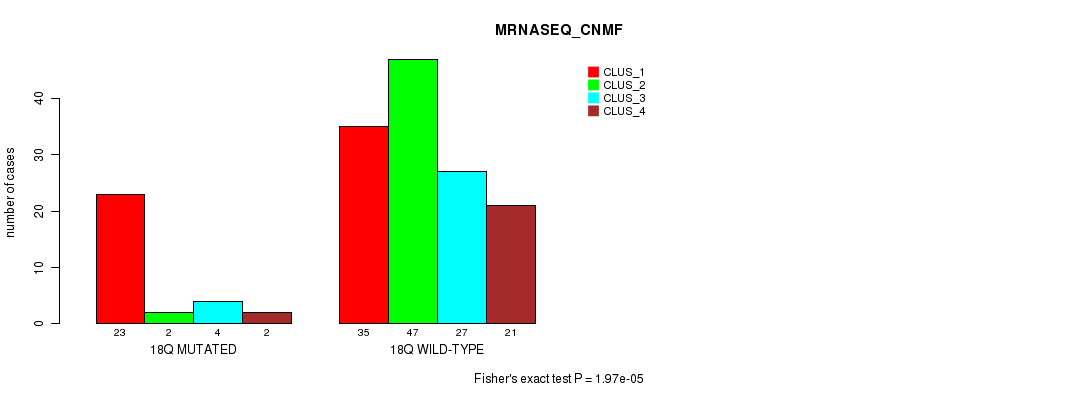
P value = 0.000811 (Fisher's exact test), Q value = 0.18
Table S91. Gene #33: '18q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 18Q MUTATED | 11 | 16 | 4 |
| 18Q WILD-TYPE | 20 | 50 | 60 |
Figure S91. Get High-res Image Gene #33: '18q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
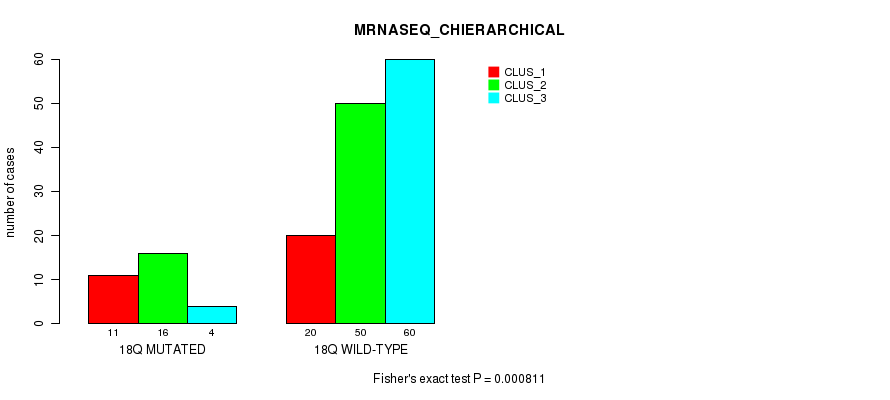
P value = 0.000225 (Fisher's exact test), Q value = 0.055
Table S92. Gene #34: '19p' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 19P MUTATED | 1 | 2 | 2 | 10 |
| 19P WILD-TYPE | 32 | 66 | 35 | 24 |
Figure S92. Get High-res Image Gene #34: '19p' versus Molecular Subtype #1: 'CN_CNMF'
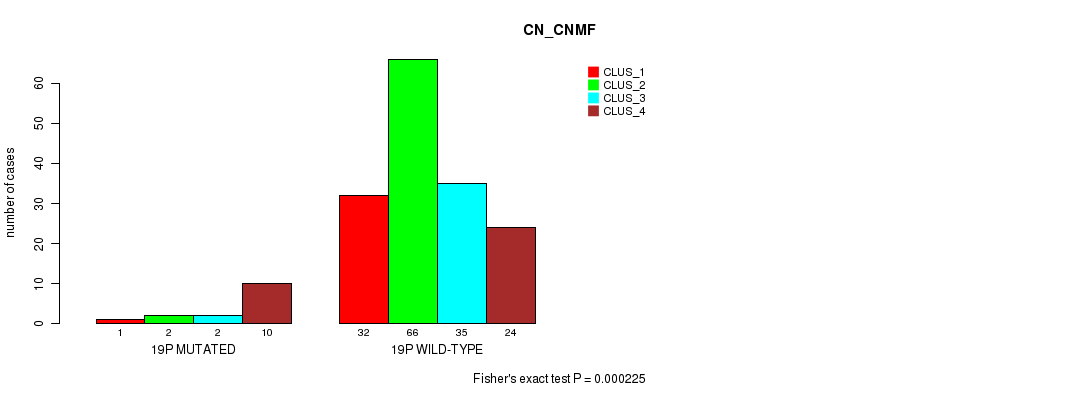
P value = 4.27e-05 (Fisher's exact test), Q value = 0.011
Table S93. Gene #35: '19q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 19Q MUTATED | 1 | 1 | 2 | 10 |
| 19Q WILD-TYPE | 32 | 67 | 35 | 24 |
Figure S93. Get High-res Image Gene #35: '19q' versus Molecular Subtype #1: 'CN_CNMF'
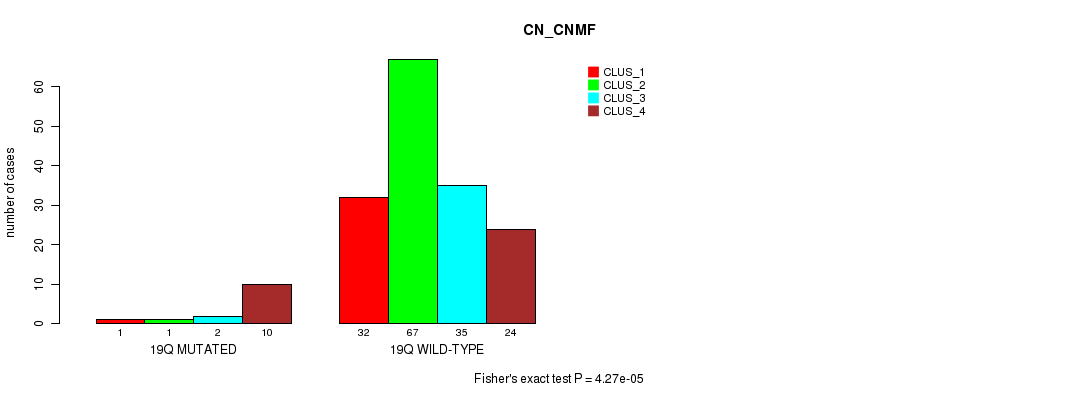
P value = 7.5e-07 (Fisher's exact test), Q value = 0.00021
Table S94. Gene #39: '22q' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| 22Q MUTATED | 1 | 16 | 5 | 20 |
| 22Q WILD-TYPE | 32 | 52 | 32 | 14 |
Figure S94. Get High-res Image Gene #39: '22q' versus Molecular Subtype #1: 'CN_CNMF'
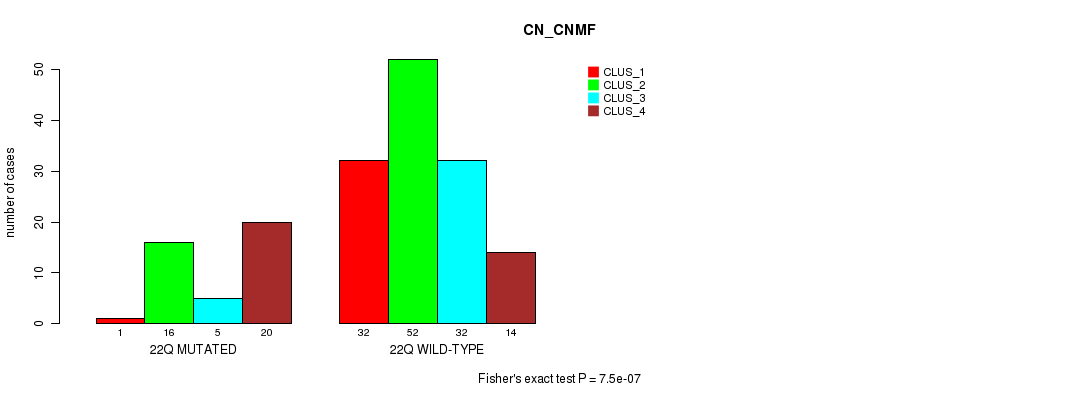
P value = 1.18e-07 (Fisher's exact test), Q value = 3.5e-05
Table S95. Gene #39: '22q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 43 | 43 | 70 |
| 22Q MUTATED | 5 | 25 | 9 |
| 22Q WILD-TYPE | 38 | 18 | 61 |
Figure S95. Get High-res Image Gene #39: '22q' versus Molecular Subtype #2: 'METHLYATION_CNMF'
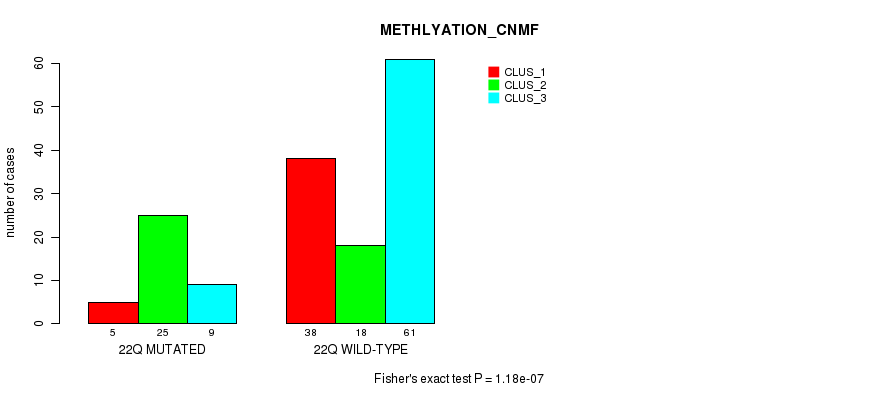
P value = 1.29e-07 (Fisher's exact test), Q value = 3.8e-05
Table S96. Gene #39: '22q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 49 | 31 | 23 |
| 22Q MUTATED | 29 | 5 | 5 | 0 |
| 22Q WILD-TYPE | 29 | 44 | 26 | 23 |
Figure S96. Get High-res Image Gene #39: '22q' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
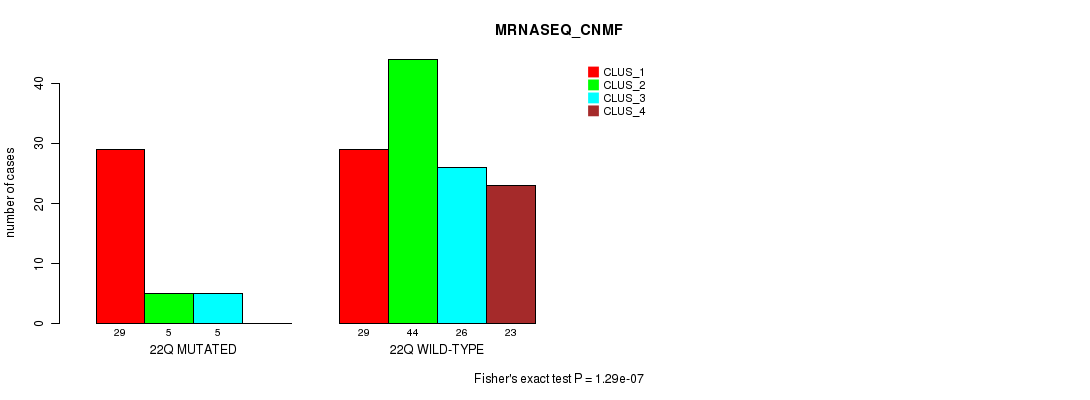
P value = 0.000958 (Fisher's exact test), Q value = 0.21
Table S97. Gene #39: '22q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 31 | 66 | 64 |
| 22Q MUTATED | 11 | 22 | 6 |
| 22Q WILD-TYPE | 20 | 44 | 58 |
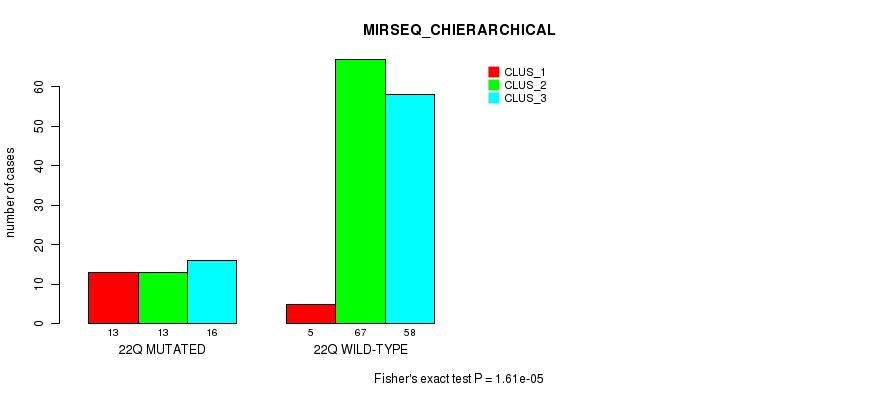
Figure S97. Get High-res Image Gene #39: '22q' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
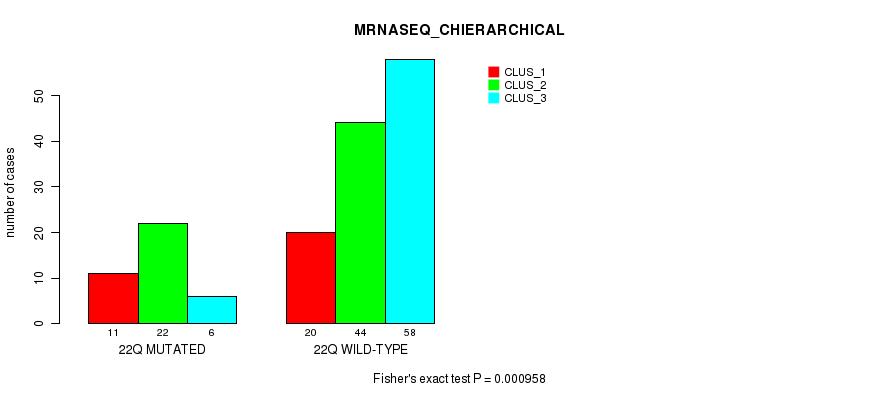
P value = 1.61e-05 (Fisher's exact test), Q value = 0.0042
Table S98. Gene #39: '22q' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 18 | 80 | 74 |
| 22Q MUTATED | 13 | 13 | 16 |
| 22Q WILD-TYPE | 5 | 67 | 58 |
Figure S98. Get High-res Image Gene #39: '22q' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
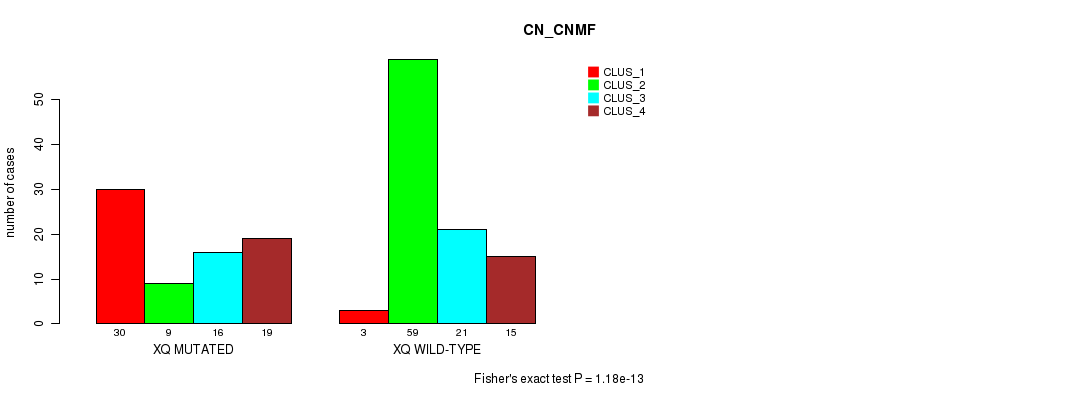
P value = 1.18e-13 (Fisher's exact test), Q value = 3.7e-11
Table S99. Gene #40: 'xq' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 68 | 37 | 34 |
| XQ MUTATED | 30 | 9 | 16 | 19 |
| XQ WILD-TYPE | 3 | 59 | 21 | 15 |
Figure S99. Get High-res Image Gene #40: 'xq' versus Molecular Subtype #1: 'CN_CNMF'
-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtype file = KIRP-TP.transferedmergedcluster.txt
-
Number of patients = 172
-
Number of significantly focal cnvs = 40
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.