This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 61 focal events and 8 molecular subtypes across 370 patients, 289 significant findings detected with P value < 0.05 and Q value < 0.25.
-
amp_1q21.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
amp_1q42.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
amp_2p24.1 cnv correlated to 'CN_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
amp_2q33.1 cnv correlated to 'CN_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_MATURE_CNMF'.
-
amp_3q26.31 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_4p14 cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_5p15.33 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
amp_5q35.3 cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
amp_6p25.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CNMF'.
-
amp_6p21.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CNMF'.
-
amp_6q12 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MIRSEQ_CNMF'.
-
amp_6q12 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
amp_7q31.2 cnv correlated to 'METHLYATION_CNMF'.
-
amp_8q24.21 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_9q34.2 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_10p15.1 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_11q13.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_13q32.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MIRSEQ_CNMF', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_13q33.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MIRSEQ_CNMF', and 'MIRSEQ_MATURE_CNMF'.
-
amp_15q26.3 cnv correlated to 'MRNASEQ_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
amp_17p11.2 cnv correlated to 'METHLYATION_CNMF'.
-
amp_17q25.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_19p13.12 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_19q13.11 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_20q13.13 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_20q13.33 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CNMF'.
-
amp_xq28 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_1p36.31 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_1p36.11 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_2q22.1 cnv correlated to 'CN_CNMF' and 'MRNASEQ_CNMF'.
-
del_2q37.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_3p13 cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_4p16.3 cnv correlated to 'CN_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_4q21.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_4q24 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_4q35.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_6q27 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_7q36.3 cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_8p23.2 cnv correlated to 'CN_CNMF'.
-
del_8p12 cnv correlated to 'CN_CNMF'.
-
del_9p21.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_9q31.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_10q23.31 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_10q26.13 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_11q14.1 cnv correlated to 'CN_CNMF', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_11q23.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_12p12.1 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_12q24.33 cnv correlated to 'CN_CNMF'.
-
del_13q14.2 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_14q23.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_MATURE_CNMF'.
-
del_14q32.33 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_15q26.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CNMF'.
-
del_16q23.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_17p13.1 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_17p11.2 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_18q21.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_19p13.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_22q12.1 cnv correlated to 'CN_CNMF' and 'MRNASEQ_CNMF'.
-
del_22q13.32 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_xq26.2 cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 61 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 289 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| amp 19p13 12 | 69 (19%) | 301 |
0.013 (0.0279) |
0.00118 (0.0042) |
0.00065 (0.00254) |
0.00055 (0.0022) |
0.00321 (0.00932) |
0.00223 (0.00676) |
0.00247 (0.00739) |
0.00129 (0.00453) |
| del 1p36 31 | 162 (44%) | 208 |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
0.00149 (0.00505) |
0.00113 (0.00408) |
0.00398 (0.0111) |
0.00469 (0.0127) |
| del 1p36 11 | 142 (38%) | 228 |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
2e-05 (0.000137) |
7e-05 (0.00038) |
0.00054 (0.00218) |
0.00026 (0.00117) |
| del 4q21 3 | 171 (46%) | 199 |
1e-05 (7.39e-05) |
0.00095 (0.00351) |
1e-05 (7.39e-05) |
0.00021 (0.000976) |
0.00133 (0.0046) |
0.00154 (0.00515) |
6e-05 (0.000337) |
6e-05 (0.000337) |
| del 4q24 | 173 (47%) | 197 |
1e-05 (7.39e-05) |
0.00081 (0.00309) |
1e-05 (7.39e-05) |
5e-05 (0.000298) |
0.00183 (0.0058) |
0.00021 (0.000976) |
2e-05 (0.000137) |
4e-05 (0.000257) |
| del 4q35 1 | 172 (46%) | 198 |
1e-05 (7.39e-05) |
0.00036 (0.00153) |
1e-05 (7.39e-05) |
0.00072 (0.00277) |
0.00188 (0.00592) |
0.00597 (0.0153) |
9e-05 (0.000453) |
0.00148 (0.00505) |
| del 9p21 3 | 150 (41%) | 220 |
1e-05 (7.39e-05) |
0.0242 (0.0469) |
1e-05 (7.39e-05) |
0.00175 (0.00566) |
4e-05 (0.000257) |
0.00215 (0.0066) |
0.0208 (0.0409) |
0.00478 (0.0127) |
| del 10q23 31 | 97 (26%) | 273 |
1e-05 (7.39e-05) |
0.015 (0.0314) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
0.0156 (0.0324) |
1e-05 (7.39e-05) |
| del 16q23 1 | 162 (44%) | 208 |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
5e-05 (0.000298) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
5e-05 (0.000298) |
| amp 17q25 3 | 145 (39%) | 225 |
1e-05 (7.39e-05) |
0.0478 (0.0809) |
0.0113 (0.0251) |
0.0106 (0.0239) |
0.00748 (0.0183) |
0.00131 (0.00457) |
0.0548 (0.0906) |
0.011 (0.0246) |
| amp 19q13 11 | 91 (25%) | 279 |
0.0164 (0.0336) |
0.0926 (0.137) |
0.00041 (0.00171) |
0.00066 (0.00256) |
0.0325 (0.0598) |
0.0018 (0.00574) |
1e-05 (7.39e-05) |
0.00027 (0.00121) |
| amp xq28 | 87 (24%) | 283 |
0.00178 (0.00571) |
0.0503 (0.0846) |
0.0275 (0.0521) |
0.00997 (0.0225) |
0.00011 (0.000537) |
0.00335 (0.00962) |
0.0158 (0.0326) |
0.00034 (0.00147) |
| del 3p13 | 73 (20%) | 297 |
0.0751 (0.115) |
0.00882 (0.0208) |
0.00011 (0.000537) |
1e-05 (7.39e-05) |
5e-05 (0.000298) |
0.0335 (0.0608) |
0.00993 (0.0225) |
0.00163 (0.00534) |
| del 9q31 3 | 119 (32%) | 251 |
1e-05 (7.39e-05) |
0.00897 (0.021) |
0.00115 (0.00413) |
0.00032 (0.00139) |
0.027 (0.0513) |
0.00696 (0.0173) |
0.251 (0.307) |
0.00022 (0.00101) |
| del 10q26 13 | 105 (28%) | 265 |
1e-05 (7.39e-05) |
0.0242 (0.0469) |
1e-05 (7.39e-05) |
1e-05 (7.39e-05) |
3e-05 (0.000201) |
1e-05 (7.39e-05) |
0.0728 (0.112) |
1e-05 (7.39e-05) |
| del 13q14 2 | 179 (48%) | 191 |
1e-05 (7.39e-05) |
0.134 (0.184) |
0.00028 (0.00124) |
1e-05 (7.39e-05) |
0.00175 (0.00566) |
0.00064 (0.00252) |
0.0179 (0.0359) |
8e-05 (0.00042) |
| amp 4p14 | 54 (15%) | 316 |
0.0636 (0.102) |
0.0188 (0.0376) |
4e-05 (0.000257) |
0.00122 (0.00431) |
0.0331 (0.0603) |
0.0555 (0.0912) |
0.00764 (0.0186) |
0.0123 (0.0268) |
| amp 8q24 21 | 231 (62%) | 139 |
1e-05 (7.39e-05) |
0.0474 (0.0808) |
0.0005 (0.00207) |
1e-05 (7.39e-05) |
0.0804 (0.122) |
0.00625 (0.0157) |
0.376 (0.44) |
7e-05 (0.00038) |
| amp 11q13 3 | 57 (15%) | 313 |
0.04 (0.0709) |
0.494 (0.546) |
0.00459 (0.0125) |
0.0177 (0.0358) |
0.0128 (0.0276) |
0.00331 (0.00956) |
0.181 (0.234) |
0.0001 (0.000498) |
| amp 20q13 13 | 124 (34%) | 246 |
1e-05 (7.39e-05) |
0.579 (0.62) |
0.0168 (0.0343) |
0.00035 (0.0015) |
0.377 (0.44) |
0.0108 (0.0241) |
5e-05 (0.000298) |
0.0477 (0.0809) |
| del 17p13 1 | 226 (61%) | 144 |
1e-05 (7.39e-05) |
0.0682 (0.108) |
2e-05 (0.000137) |
6e-05 (0.000337) |
1e-05 (7.39e-05) |
8e-05 (0.00042) |
0.187 (0.241) |
1e-05 (7.39e-05) |
| del 17p11 2 | 195 (53%) | 175 |
1e-05 (7.39e-05) |
0.326 (0.384) |
0.00016 (0.000765) |
0.00839 (0.02) |
0.0009 (0.00338) |
0.00024 (0.00109) |
0.231 (0.289) |
0.00039 (0.00164) |
| amp 1q21 3 | 271 (73%) | 99 |
1e-05 (7.39e-05) |
9e-05 (0.000453) |
3e-05 (0.000201) |
0.00398 (0.0111) |
0.136 (0.186) |
0.0299 (0.0557) |
0.106 (0.152) |
0.385 (0.448) |
| amp 1q42 2 | 268 (72%) | 102 |
1e-05 (7.39e-05) |
0.0021 (0.00653) |
1e-05 (7.39e-05) |
7e-05 (0.00038) |
0.249 (0.306) |
0.00852 (0.0202) |
0.0575 (0.0942) |
0.406 (0.466) |
| amp 3q26 31 | 77 (21%) | 293 |
1e-05 (7.39e-05) |
0.0995 (0.145) |
0.0206 (0.0409) |
0.0203 (0.0405) |
0.0703 (0.11) |
0.00491 (0.013) |
0.255 (0.31) |
0.0482 (0.0814) |
| amp 6p25 2 | 159 (43%) | 211 |
1e-05 (7.39e-05) |
0.00017 (0.000805) |
0.0434 (0.0745) |
0.0245 (0.0471) |
0.015 (0.0314) |
0.0634 (0.102) |
0.709 (0.741) |
0.426 (0.482) |
| amp 10p15 1 | 72 (19%) | 298 |
0.00415 (0.0115) |
0.902 (0.914) |
0.0359 (0.0644) |
0.00356 (0.0101) |
0.408 (0.468) |
0.243 (0.3) |
0.008 (0.0193) |
0.0236 (0.0463) |
| amp 13q32 3 | 67 (18%) | 303 |
1e-05 (7.39e-05) |
0.101 (0.146) |
0.00938 (0.0219) |
0.08 (0.121) |
0.0126 (0.0272) |
0.198 (0.253) |
0.0062 (0.0157) |
0.0351 (0.0634) |
| amp 13q33 3 | 67 (18%) | 303 |
1e-05 (7.39e-05) |
0.0464 (0.0795) |
0.00162 (0.00534) |
0.25 (0.306) |
0.00949 (0.0221) |
0.18 (0.234) |
0.00617 (0.0157) |
0.084 (0.125) |
| del 2q37 3 | 62 (17%) | 308 |
5e-05 (0.000298) |
0.535 (0.585) |
1e-05 (7.39e-05) |
0.00091 (0.00339) |
0.0507 (0.0847) |
0.00994 (0.0225) |
0.307 (0.365) |
0.00148 (0.00505) |
| del 14q23 3 | 131 (35%) | 239 |
1e-05 (7.39e-05) |
0.499 (0.55) |
0.00214 (0.0066) |
6e-05 (0.000337) |
0.012 (0.0263) |
0.0976 (0.143) |
0.00416 (0.0115) |
0.0504 (0.0846) |
| del 14q32 33 | 133 (36%) | 237 |
1e-05 (7.39e-05) |
0.143 (0.193) |
0.00474 (0.0127) |
0.00012 (0.00058) |
0.083 (0.125) |
0.175 (0.228) |
0.0125 (0.0271) |
0.015 (0.0314) |
| del 19p13 3 | 87 (24%) | 283 |
1e-05 (7.39e-05) |
0.424 (0.482) |
0.00498 (0.0131) |
0.0026 (0.00769) |
0.111 (0.159) |
0.132 (0.181) |
0.00217 (0.00662) |
0.00099 (0.00363) |
| amp 2p24 1 | 65 (18%) | 305 |
0.0354 (0.0638) |
1 (1.00) |
0.112 (0.159) |
0.00399 (0.0111) |
0.0173 (0.0351) |
0.0423 (0.073) |
0.0687 (0.108) |
0.173 (0.227) |
| amp 2q33 1 | 64 (17%) | 306 |
0.0277 (0.0522) |
0.938 (0.946) |
0.0515 (0.0858) |
0.0114 (0.0253) |
0.00157 (0.00521) |
0.121 (0.171) |
0.00596 (0.0153) |
0.192 (0.246) |
| amp 6q12 | 109 (29%) | 261 |
1e-05 (7.39e-05) |
0.0417 (0.0725) |
0.0318 (0.0588) |
0.108 (0.154) |
0.041 (0.0717) |
0.618 (0.659) |
0.235 (0.292) |
0.159 (0.211) |
| amp 6q12 | 83 (22%) | 287 |
2e-05 (0.000137) |
0.00725 (0.018) |
0.00813 (0.0195) |
0.00967 (0.0224) |
0.319 (0.378) |
0.549 (0.595) |
0.0836 (0.125) |
0.0872 (0.13) |
| amp 9q34 2 | 47 (13%) | 323 |
0.029 (0.0542) |
0.404 (0.464) |
0.0258 (0.0494) |
0.0927 (0.137) |
0.242 (0.3) |
0.0122 (0.0268) |
0.4 (0.461) |
0.0208 (0.0409) |
| amp 20q13 33 | 128 (35%) | 242 |
1e-05 (7.39e-05) |
0.149 (0.198) |
0.00482 (0.0128) |
6e-05 (0.000337) |
0.231 (0.289) |
0.0622 (0.101) |
1e-05 (7.39e-05) |
0.13 (0.18) |
| del 6q27 | 136 (37%) | 234 |
0.00152 (0.00512) |
0.0054 (0.0141) |
0.471 (0.526) |
0.48 (0.534) |
0.0289 (0.0542) |
0.0391 (0.07) |
0.559 (0.603) |
0.309 (0.367) |
| del 7q36 3 | 34 (9%) | 336 |
0.231 (0.289) |
0.0408 (0.0716) |
0.0414 (0.0722) |
0.131 (0.181) |
0.901 (0.914) |
0.0706 (0.11) |
0.00032 (0.00139) |
0.00772 (0.0187) |
| del 11q23 3 | 93 (25%) | 277 |
0.00054 (0.00218) |
0.488 (0.541) |
0.0587 (0.0955) |
0.0406 (0.0716) |
0.0761 (0.116) |
0.136 (0.186) |
0.0138 (0.0295) |
0.0021 (0.00653) |
| del 12p12 1 | 78 (21%) | 292 |
0.00084 (0.00318) |
0.453 (0.507) |
0.00254 (0.00756) |
0.229 (0.289) |
0.0641 (0.103) |
0.031 (0.0575) |
0.181 (0.234) |
0.0326 (0.0598) |
| del 15q26 3 | 79 (21%) | 291 |
0.00052 (0.00213) |
0.436 (0.491) |
0.00834 (0.02) |
0.00971 (0.0224) |
0.0793 (0.12) |
0.121 (0.171) |
0.00225 (0.00678) |
0.0649 (0.103) |
| amp 5p15 33 | 163 (44%) | 207 |
1e-05 (7.39e-05) |
0.0331 (0.0603) |
0.199 (0.253) |
0.607 (0.648) |
0.141 (0.191) |
0.0174 (0.0353) |
0.143 (0.193) |
0.127 (0.177) |
| amp 6p21 1 | 158 (43%) | 212 |
1e-05 (7.39e-05) |
0.00109 (0.00397) |
0.0141 (0.03) |
0.0655 (0.104) |
0.054 (0.0896) |
0.0623 (0.101) |
0.174 (0.228) |
0.529 (0.581) |
| del 4p16 3 | 85 (23%) | 285 |
1e-05 (7.39e-05) |
0.148 (0.198) |
0.105 (0.15) |
0.101 (0.146) |
0.0982 (0.144) |
0.027 (0.0513) |
0.0578 (0.0943) |
0.00557 (0.0145) |
| del 11q14 1 | 77 (21%) | 293 |
0.0161 (0.0331) |
0.357 (0.418) |
0.122 (0.172) |
0.144 (0.194) |
0.13 (0.18) |
0.263 (0.319) |
0.00743 (0.0183) |
0.0153 (0.0319) |
| del 18q21 2 | 94 (25%) | 276 |
0.0241 (0.0469) |
9e-05 (0.000453) |
0.0709 (0.11) |
2e-05 (0.000137) |
0.821 (0.837) |
0.538 (0.587) |
0.417 (0.474) |
0.299 (0.356) |
| del 22q13 32 | 83 (22%) | 287 |
0.00056 (0.00222) |
0.0819 (0.123) |
9e-05 (0.000453) |
0.00992 (0.0225) |
0.0552 (0.0911) |
0.264 (0.32) |
0.718 (0.746) |
0.285 (0.34) |
| amp 5q35 3 | 135 (36%) | 235 |
1e-05 (7.39e-05) |
0.0403 (0.0713) |
0.73 (0.754) |
0.957 (0.963) |
0.436 (0.491) |
0.999 (1.00) |
0.688 (0.725) |
0.62 (0.659) |
| amp 15q26 3 | 54 (15%) | 316 |
0.0741 (0.114) |
0.445 (0.5) |
0.0243 (0.0469) |
0.0395 (0.0703) |
0.415 (0.473) |
0.181 (0.234) |
0.497 (0.548) |
0.576 (0.619) |
| del 2q22 1 | 52 (14%) | 318 |
0.00308 (0.009) |
0.394 (0.455) |
0.00471 (0.0127) |
0.0659 (0.104) |
0.275 (0.332) |
0.167 (0.221) |
0.777 (0.793) |
0.0778 (0.119) |
| del 22q12 1 | 79 (21%) | 291 |
0.00622 (0.0157) |
0.323 (0.382) |
0.00288 (0.00847) |
0.106 (0.152) |
0.635 (0.673) |
0.194 (0.248) |
0.714 (0.744) |
0.771 (0.791) |
| del xq26 2 | 67 (18%) | 303 |
8e-05 (0.00042) |
0.00695 (0.0173) |
0.145 (0.195) |
0.392 (0.454) |
0.281 (0.339) |
0.695 (0.731) |
0.285 (0.34) |
0.701 (0.734) |
| amp 7q31 2 | 120 (32%) | 250 |
0.225 (0.285) |
0.00454 (0.0124) |
0.637 (0.674) |
0.897 (0.911) |
0.171 (0.225) |
0.432 (0.489) |
0.722 (0.748) |
0.604 (0.647) |
| amp 17p11 2 | 46 (12%) | 324 |
0.188 (0.242) |
0.042 (0.0727) |
0.472 (0.526) |
0.231 (0.289) |
0.144 (0.194) |
0.252 (0.307) |
0.387 (0.45) |
0.244 (0.3) |
| del 8p23 2 | 247 (67%) | 123 |
1e-05 (7.39e-05) |
0.127 (0.178) |
0.0974 (0.143) |
0.652 (0.688) |
0.696 (0.731) |
0.17 (0.224) |
0.17 (0.225) |
0.0691 (0.108) |
| del 8p12 | 242 (65%) | 128 |
1e-05 (7.39e-05) |
0.128 (0.178) |
0.0725 (0.112) |
0.562 (0.606) |
0.34 (0.4) |
0.1 (0.145) |
0.129 (0.18) |
0.0884 (0.131) |
| del 12q24 33 | 59 (16%) | 311 |
0.00345 (0.00985) |
1 (1.00) |
0.544 (0.593) |
0.916 (0.925) |
0.502 (0.552) |
0.717 (0.746) |
0.776 (0.793) |
0.283 (0.34) |
| del 20p12 1 | 29 (8%) | 341 |
0.234 (0.292) |
0.548 (0.595) |
0.734 (0.758) |
0.549 (0.595) |
0.206 (0.261) |
0.737 (0.759) |
0.76 (0.781) |
0.409 (0.468) |
P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S1. Gene #1: 'amp_1q21.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 1(1Q21.3) MUTATED | 42 | 116 | 113 |
| AMP PEAK 1(1Q21.3) WILD-TYPE | 51 | 35 | 13 |
Figure S1. Get High-res Image Gene #1: 'amp_1q21.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 9e-05 (Fisher's exact test), Q value = 0.00045
Table S2. Gene #1: 'amp_1q21.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 1(1Q21.3) MUTATED | 54 | 119 | 98 |
| AMP PEAK 1(1Q21.3) WILD-TYPE | 39 | 42 | 18 |
Figure S2. Get High-res Image Gene #1: 'amp_1q21.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 3e-05 (Fisher's exact test), Q value = 2e-04
Table S3. Gene #1: 'amp_1q21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 1(1Q21.3) MUTATED | 49 | 51 | 66 | 46 | 54 |
| AMP PEAK 1(1Q21.3) WILD-TYPE | 18 | 35 | 5 | 25 | 15 |
Figure S3. Get High-res Image Gene #1: 'amp_1q21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00398 (Fisher's exact test), Q value = 0.011
Table S4. Gene #1: 'amp_1q21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 1(1Q21.3) MUTATED | 36 | 38 | 74 | 72 | 46 |
| AMP PEAK 1(1Q21.3) WILD-TYPE | 18 | 18 | 40 | 12 | 10 |
Figure S4. Get High-res Image Gene #1: 'amp_1q21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.0299 (Fisher's exact test), Q value = 0.056
Table S5. Gene #1: 'amp_1q21.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 1(1Q21.3) MUTATED | 37 | 60 | 45 | 86 | 39 |
| AMP PEAK 1(1Q21.3) WILD-TYPE | 7 | 35 | 11 | 36 | 9 |
Figure S5. Get High-res Image Gene #1: 'amp_1q21.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S6. Gene #2: 'amp_1q42.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 2(1Q42.2) MUTATED | 50 | 105 | 113 |
| AMP PEAK 2(1Q42.2) WILD-TYPE | 43 | 46 | 13 |
Figure S6. Get High-res Image Gene #2: 'amp_1q42.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0021 (Fisher's exact test), Q value = 0.0065
Table S7. Gene #2: 'amp_1q42.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 2(1Q42.2) MUTATED | 55 | 119 | 94 |
| AMP PEAK 2(1Q42.2) WILD-TYPE | 38 | 42 | 22 |
Figure S7. Get High-res Image Gene #2: 'amp_1q42.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S8. Gene #2: 'amp_1q42.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 2(1Q42.2) MUTATED | 46 | 46 | 67 | 49 | 55 |
| AMP PEAK 2(1Q42.2) WILD-TYPE | 21 | 40 | 4 | 22 | 14 |
Figure S8. Get High-res Image Gene #2: 'amp_1q42.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 7e-05 (Fisher's exact test), Q value = 0.00038
Table S9. Gene #2: 'amp_1q42.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 2(1Q42.2) MUTATED | 33 | 39 | 71 | 73 | 47 |
| AMP PEAK 2(1Q42.2) WILD-TYPE | 21 | 17 | 43 | 11 | 9 |
Figure S9. Get High-res Image Gene #2: 'amp_1q42.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00852 (Fisher's exact test), Q value = 0.02
Table S10. Gene #2: 'amp_1q42.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 2(1Q42.2) MUTATED | 32 | 58 | 45 | 87 | 42 |
| AMP PEAK 2(1Q42.2) WILD-TYPE | 12 | 37 | 11 | 35 | 6 |
Figure S10. Get High-res Image Gene #2: 'amp_1q42.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.0354 (Fisher's exact test), Q value = 0.064
Table S11. Gene #3: 'amp_2p24.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 3(2P24.1) MUTATED | 9 | 34 | 22 |
| AMP PEAK 3(2P24.1) WILD-TYPE | 84 | 117 | 104 |
Figure S11. Get High-res Image Gene #3: 'amp_2p24.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00399 (Fisher's exact test), Q value = 0.011
Table S12. Gene #3: 'amp_2p24.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 3(2P24.1) MUTATED | 17 | 3 | 21 | 16 | 6 |
| AMP PEAK 3(2P24.1) WILD-TYPE | 37 | 53 | 93 | 68 | 50 |
Figure S12. Get High-res Image Gene #3: 'amp_2p24.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.0173 (Fisher's exact test), Q value = 0.035
Table S13. Gene #3: 'amp_2p24.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 3(2P24.1) MUTATED | 12 | 15 | 8 | 13 | 6 | 10 |
| AMP PEAK 3(2P24.1) WILD-TYPE | 28 | 48 | 84 | 71 | 40 | 30 |
Figure S13. Get High-res Image Gene #3: 'amp_2p24.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.0423 (Fisher's exact test), Q value = 0.073
Table S14. Gene #3: 'amp_2p24.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 3(2P24.1) MUTATED | 14 | 20 | 9 | 16 | 5 |
| AMP PEAK 3(2P24.1) WILD-TYPE | 30 | 75 | 47 | 106 | 43 |
Figure S14. Get High-res Image Gene #3: 'amp_2p24.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.0277 (Fisher's exact test), Q value = 0.052
Table S15. Gene #4: 'amp_2q33.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 4(2Q33.1) MUTATED | 8 | 30 | 26 |
| AMP PEAK 4(2Q33.1) WILD-TYPE | 85 | 121 | 100 |
Figure S15. Get High-res Image Gene #4: 'amp_2q33.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0114 (Fisher's exact test), Q value = 0.025
Table S16. Gene #4: 'amp_2q33.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 4(2Q33.1) MUTATED | 15 | 4 | 14 | 20 | 9 |
| AMP PEAK 4(2Q33.1) WILD-TYPE | 39 | 52 | 100 | 64 | 47 |
Figure S16. Get High-res Image Gene #4: 'amp_2q33.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00157 (Fisher's exact test), Q value = 0.0052
Table S17. Gene #4: 'amp_2q33.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 4(2Q33.1) MUTATED | 9 | 17 | 6 | 10 | 10 | 11 |
| AMP PEAK 4(2Q33.1) WILD-TYPE | 31 | 46 | 86 | 74 | 36 | 29 |
Figure S17. Get High-res Image Gene #4: 'amp_2q33.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00596 (Fisher's exact test), Q value = 0.015
Table S18. Gene #4: 'amp_2q33.1' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| AMP PEAK 4(2Q33.1) MUTATED | 30 | 10 | 11 | 10 |
| AMP PEAK 4(2Q33.1) WILD-TYPE | 77 | 71 | 91 | 40 |
Figure S18. Get High-res Image Gene #4: 'amp_2q33.1' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S19. Gene #5: 'amp_3q26.31' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 5(3Q26.31) MUTATED | 7 | 51 | 19 |
| AMP PEAK 5(3Q26.31) WILD-TYPE | 86 | 100 | 107 |
Figure S19. Get High-res Image Gene #5: 'amp_3q26.31' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0206 (Fisher's exact test), Q value = 0.041
Table S20. Gene #5: 'amp_3q26.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 5(3Q26.31) MUTATED | 22 | 23 | 12 | 11 | 9 |
| AMP PEAK 5(3Q26.31) WILD-TYPE | 45 | 63 | 59 | 60 | 60 |
Figure S20. Get High-res Image Gene #5: 'amp_3q26.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.0203 (Fisher's exact test), Q value = 0.041
Table S21. Gene #5: 'amp_3q26.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 5(3Q26.31) MUTATED | 17 | 5 | 30 | 16 | 9 |
| AMP PEAK 5(3Q26.31) WILD-TYPE | 37 | 51 | 84 | 68 | 47 |
Figure S21. Get High-res Image Gene #5: 'amp_3q26.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00491 (Fisher's exact test), Q value = 0.013
Table S22. Gene #5: 'amp_3q26.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 5(3Q26.31) MUTATED | 17 | 24 | 9 | 22 | 4 |
| AMP PEAK 5(3Q26.31) WILD-TYPE | 27 | 71 | 47 | 100 | 44 |
Figure S22. Get High-res Image Gene #5: 'amp_3q26.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.0482 (Fisher's exact test), Q value = 0.081
Table S23. Gene #5: 'amp_3q26.31' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 5(3Q26.31) MUTATED | 14 | 20 | 20 | 12 | 5 |
| AMP PEAK 5(3Q26.31) WILD-TYPE | 56 | 62 | 80 | 22 | 49 |
Figure S23. Get High-res Image Gene #5: 'amp_3q26.31' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.0188 (Fisher's exact test), Q value = 0.038
Table S24. Gene #6: 'amp_4p14' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 6(4P14) MUTATED | 11 | 17 | 26 |
| AMP PEAK 6(4P14) WILD-TYPE | 82 | 144 | 90 |
Figure S24. Get High-res Image Gene #6: 'amp_4p14' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 4e-05 (Fisher's exact test), Q value = 0.00026
Table S25. Gene #6: 'amp_4p14' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 6(4P14) MUTATED | 12 | 11 | 21 | 8 | 1 |
| AMP PEAK 6(4P14) WILD-TYPE | 55 | 75 | 50 | 63 | 68 |
Figure S25. Get High-res Image Gene #6: 'amp_4p14' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00122 (Fisher's exact test), Q value = 0.0043
Table S26. Gene #6: 'amp_4p14' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 6(4P14) MUTATED | 10 | 8 | 13 | 21 | 1 |
| AMP PEAK 6(4P14) WILD-TYPE | 44 | 48 | 101 | 63 | 55 |
Figure S26. Get High-res Image Gene #6: 'amp_4p14' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.0331 (Fisher's exact test), Q value = 0.06
Table S27. Gene #6: 'amp_4p14' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 6(4P14) MUTATED | 7 | 14 | 9 | 6 | 10 | 8 |
| AMP PEAK 6(4P14) WILD-TYPE | 33 | 49 | 83 | 78 | 36 | 32 |
Figure S27. Get High-res Image Gene #6: 'amp_4p14' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00764 (Fisher's exact test), Q value = 0.019
Table S28. Gene #6: 'amp_4p14' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| AMP PEAK 6(4P14) MUTATED | 18 | 16 | 7 | 13 |
| AMP PEAK 6(4P14) WILD-TYPE | 89 | 65 | 95 | 37 |
Figure S28. Get High-res Image Gene #6: 'amp_4p14' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.0123 (Fisher's exact test), Q value = 0.027
Table S29. Gene #6: 'amp_4p14' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 6(4P14) MUTATED | 14 | 16 | 18 | 5 | 1 |
| AMP PEAK 6(4P14) WILD-TYPE | 56 | 66 | 82 | 29 | 53 |
Figure S29. Get High-res Image Gene #6: 'amp_4p14' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S30. Gene #7: 'amp_5p15.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 7(5P15.33) MUTATED | 16 | 80 | 67 |
| AMP PEAK 7(5P15.33) WILD-TYPE | 77 | 71 | 59 |
Figure S30. Get High-res Image Gene #7: 'amp_5p15.33' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0331 (Fisher's exact test), Q value = 0.06
Table S31. Gene #7: 'amp_5p15.33' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 7(5P15.33) MUTATED | 32 | 82 | 49 |
| AMP PEAK 7(5P15.33) WILD-TYPE | 61 | 79 | 67 |
Figure S31. Get High-res Image Gene #7: 'amp_5p15.33' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.0174 (Fisher's exact test), Q value = 0.035
Table S32. Gene #7: 'amp_5p15.33' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 7(5P15.33) MUTATED | 26 | 47 | 28 | 43 | 16 |
| AMP PEAK 7(5P15.33) WILD-TYPE | 18 | 48 | 28 | 79 | 32 |
Figure S32. Get High-res Image Gene #7: 'amp_5p15.33' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S33. Gene #8: 'amp_5q35.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 8(5Q35.3) MUTATED | 20 | 49 | 66 |
| AMP PEAK 8(5Q35.3) WILD-TYPE | 73 | 102 | 60 |
Figure S33. Get High-res Image Gene #8: 'amp_5q35.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0403 (Fisher's exact test), Q value = 0.071
Table S34. Gene #8: 'amp_5q35.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 8(5Q35.3) MUTATED | 24 | 66 | 45 |
| AMP PEAK 8(5Q35.3) WILD-TYPE | 69 | 95 | 71 |
Figure S34. Get High-res Image Gene #8: 'amp_5q35.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S35. Gene #9: 'amp_6p25.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 9(6P25.2) MUTATED | 15 | 79 | 65 |
| AMP PEAK 9(6P25.2) WILD-TYPE | 78 | 72 | 61 |
Figure S35. Get High-res Image Gene #9: 'amp_6p25.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00017 (Fisher's exact test), Q value = 0.00081
Table S36. Gene #9: 'amp_6p25.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 9(6P25.2) MUTATED | 23 | 79 | 57 |
| AMP PEAK 9(6P25.2) WILD-TYPE | 70 | 82 | 59 |
Figure S36. Get High-res Image Gene #9: 'amp_6p25.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.0434 (Fisher's exact test), Q value = 0.075
Table S37. Gene #9: 'amp_6p25.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 9(6P25.2) MUTATED | 32 | 33 | 38 | 22 | 34 |
| AMP PEAK 9(6P25.2) WILD-TYPE | 35 | 53 | 33 | 49 | 35 |
Figure S37. Get High-res Image Gene #9: 'amp_6p25.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.0245 (Fisher's exact test), Q value = 0.047
Table S38. Gene #9: 'amp_6p25.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 9(6P25.2) MUTATED | 24 | 16 | 45 | 46 | 28 |
| AMP PEAK 9(6P25.2) WILD-TYPE | 30 | 40 | 69 | 38 | 28 |
Figure S38. Get High-res Image Gene #9: 'amp_6p25.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.015 (Fisher's exact test), Q value = 0.031
Table S39. Gene #9: 'amp_6p25.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 9(6P25.2) MUTATED | 24 | 27 | 26 | 37 | 21 | 20 |
| AMP PEAK 9(6P25.2) WILD-TYPE | 16 | 36 | 66 | 47 | 25 | 20 |
Figure S39. Get High-res Image Gene #9: 'amp_6p25.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S40. Gene #10: 'amp_6p21.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 10(6P21.1) MUTATED | 17 | 79 | 62 |
| AMP PEAK 10(6P21.1) WILD-TYPE | 76 | 72 | 64 |
Figure S40. Get High-res Image Gene #10: 'amp_6p21.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00109 (Fisher's exact test), Q value = 0.004
Table S41. Gene #10: 'amp_6p21.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 10(6P21.1) MUTATED | 25 | 80 | 53 |
| AMP PEAK 10(6P21.1) WILD-TYPE | 68 | 81 | 63 |
Figure S41. Get High-res Image Gene #10: 'amp_6p21.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.0141 (Fisher's exact test), Q value = 0.03
Table S42. Gene #10: 'amp_6p21.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 10(6P21.1) MUTATED | 27 | 40 | 37 | 19 | 35 |
| AMP PEAK 10(6P21.1) WILD-TYPE | 40 | 46 | 34 | 52 | 34 |
Figure S42. Get High-res Image Gene #10: 'amp_6p21.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S43. Gene #11: 'amp_6q12' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 11(6Q12) MUTATED | 9 | 62 | 38 |
| AMP PEAK 11(6Q12) WILD-TYPE | 84 | 89 | 88 |
Figure S43. Get High-res Image Gene #11: 'amp_6q12' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0417 (Fisher's exact test), Q value = 0.073
Table S44. Gene #11: 'amp_6q12' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 11(6Q12) MUTATED | 18 | 52 | 39 |
| AMP PEAK 11(6Q12) WILD-TYPE | 75 | 109 | 77 |
Figure S44. Get High-res Image Gene #11: 'amp_6q12' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.0318 (Fisher's exact test), Q value = 0.059
Table S45. Gene #11: 'amp_6q12' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 11(6Q12) MUTATED | 24 | 22 | 30 | 14 | 19 |
| AMP PEAK 11(6Q12) WILD-TYPE | 43 | 64 | 41 | 57 | 50 |
Figure S45. Get High-res Image Gene #11: 'amp_6q12' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.041 (Fisher's exact test), Q value = 0.072
Table S46. Gene #11: 'amp_6q12' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 11(6Q12) MUTATED | 17 | 18 | 18 | 22 | 18 | 15 |
| AMP PEAK 11(6Q12) WILD-TYPE | 23 | 45 | 74 | 62 | 28 | 25 |
Figure S46. Get High-res Image Gene #11: 'amp_6q12' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 2e-05 (Fisher's exact test), Q value = 0.00014
Table S47. Gene #12: 'amp_6q12' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 12(6Q12) MUTATED | 5 | 47 | 31 |
| AMP PEAK 12(6Q12) WILD-TYPE | 88 | 104 | 95 |
Figure S47. Get High-res Image Gene #12: 'amp_6q12' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00725 (Fisher's exact test), Q value = 0.018
Table S48. Gene #12: 'amp_6q12' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 12(6Q12) MUTATED | 11 | 38 | 34 |
| AMP PEAK 12(6Q12) WILD-TYPE | 82 | 123 | 82 |
Figure S48. Get High-res Image Gene #12: 'amp_6q12' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.00813 (Fisher's exact test), Q value = 0.02
Table S49. Gene #12: 'amp_6q12' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 12(6Q12) MUTATED | 17 | 12 | 27 | 13 | 14 |
| AMP PEAK 12(6Q12) WILD-TYPE | 50 | 74 | 44 | 58 | 55 |
Figure S49. Get High-res Image Gene #12: 'amp_6q12' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00967 (Fisher's exact test), Q value = 0.022
Table S50. Gene #12: 'amp_6q12' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 12(6Q12) MUTATED | 15 | 10 | 17 | 30 | 11 |
| AMP PEAK 12(6Q12) WILD-TYPE | 39 | 46 | 97 | 54 | 45 |
Figure S50. Get High-res Image Gene #12: 'amp_6q12' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00454 (Fisher's exact test), Q value = 0.012
Table S51. Gene #13: 'amp_7q31.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 13(7Q31.2) MUTATED | 24 | 67 | 29 |
| AMP PEAK 13(7Q31.2) WILD-TYPE | 69 | 94 | 87 |
Figure S51. Get High-res Image Gene #13: 'amp_7q31.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S52. Gene #14: 'amp_8q24.21' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 14(8Q24.21) MUTATED | 45 | 79 | 107 |
| AMP PEAK 14(8Q24.21) WILD-TYPE | 48 | 72 | 19 |
Figure S52. Get High-res Image Gene #14: 'amp_8q24.21' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0474 (Fisher's exact test), Q value = 0.081
Table S53. Gene #14: 'amp_8q24.21' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 14(8Q24.21) MUTATED | 55 | 93 | 83 |
| AMP PEAK 14(8Q24.21) WILD-TYPE | 38 | 68 | 33 |
Figure S53. Get High-res Image Gene #14: 'amp_8q24.21' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 5e-04 (Fisher's exact test), Q value = 0.0021
Table S54. Gene #14: 'amp_8q24.21' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 14(8Q24.21) MUTATED | 44 | 36 | 50 | 47 | 49 |
| AMP PEAK 14(8Q24.21) WILD-TYPE | 23 | 50 | 21 | 24 | 20 |
Figure S54. Get High-res Image Gene #14: 'amp_8q24.21' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S55. Gene #14: 'amp_8q24.21' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 14(8Q24.21) MUTATED | 39 | 40 | 45 | 61 | 41 |
| AMP PEAK 14(8Q24.21) WILD-TYPE | 15 | 16 | 69 | 23 | 15 |
Figure S55. Get High-res Image Gene #14: 'amp_8q24.21' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00625 (Fisher's exact test), Q value = 0.016
Table S56. Gene #14: 'amp_8q24.21' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 14(8Q24.21) MUTATED | 24 | 46 | 37 | 85 | 35 |
| AMP PEAK 14(8Q24.21) WILD-TYPE | 20 | 49 | 19 | 37 | 13 |
Figure S56. Get High-res Image Gene #14: 'amp_8q24.21' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 7e-05 (Fisher's exact test), Q value = 0.00038
Table S57. Gene #14: 'amp_8q24.21' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 14(8Q24.21) MUTATED | 52 | 36 | 70 | 18 | 41 |
| AMP PEAK 14(8Q24.21) WILD-TYPE | 18 | 46 | 30 | 16 | 13 |
Figure S57. Get High-res Image Gene #14: 'amp_8q24.21' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.029 (Fisher's exact test), Q value = 0.054
Table S58. Gene #15: 'amp_9q34.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 15(9Q34.2) MUTATED | 5 | 25 | 17 |
| AMP PEAK 15(9Q34.2) WILD-TYPE | 88 | 126 | 109 |
Figure S58. Get High-res Image Gene #15: 'amp_9q34.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0258 (Fisher's exact test), Q value = 0.049
Table S59. Gene #15: 'amp_9q34.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 15(9Q34.2) MUTATED | 11 | 15 | 12 | 5 | 3 |
| AMP PEAK 15(9Q34.2) WILD-TYPE | 56 | 71 | 59 | 66 | 66 |
Figure S59. Get High-res Image Gene #15: 'amp_9q34.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.0122 (Fisher's exact test), Q value = 0.027
Table S60. Gene #15: 'amp_9q34.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 15(9Q34.2) MUTATED | 11 | 15 | 9 | 10 | 2 |
| AMP PEAK 15(9Q34.2) WILD-TYPE | 33 | 80 | 47 | 112 | 46 |
Figure S60. Get High-res Image Gene #15: 'amp_9q34.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.0208 (Fisher's exact test), Q value = 0.041
Table S61. Gene #15: 'amp_9q34.2' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 15(9Q34.2) MUTATED | 12 | 17 | 9 | 5 | 2 |
| AMP PEAK 15(9Q34.2) WILD-TYPE | 58 | 65 | 91 | 29 | 52 |
Figure S61. Get High-res Image Gene #15: 'amp_9q34.2' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.00415 (Fisher's exact test), Q value = 0.011
Table S62. Gene #16: 'amp_10p15.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 16(10P15.1) MUTATED | 8 | 38 | 26 |
| AMP PEAK 16(10P15.1) WILD-TYPE | 85 | 113 | 100 |
Figure S62. Get High-res Image Gene #16: 'amp_10p15.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0359 (Fisher's exact test), Q value = 0.064
Table S63. Gene #16: 'amp_10p15.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 16(10P15.1) MUTATED | 20 | 10 | 18 | 11 | 12 |
| AMP PEAK 16(10P15.1) WILD-TYPE | 47 | 76 | 53 | 60 | 57 |
Figure S63. Get High-res Image Gene #16: 'amp_10p15.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00356 (Fisher's exact test), Q value = 0.01
Table S64. Gene #16: 'amp_10p15.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 16(10P15.1) MUTATED | 19 | 8 | 14 | 22 | 8 |
| AMP PEAK 16(10P15.1) WILD-TYPE | 35 | 48 | 100 | 62 | 48 |
Figure S64. Get High-res Image Gene #16: 'amp_10p15.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.008 (Fisher's exact test), Q value = 0.019
Table S65. Gene #16: 'amp_10p15.1' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| AMP PEAK 16(10P15.1) MUTATED | 30 | 18 | 10 | 10 |
| AMP PEAK 16(10P15.1) WILD-TYPE | 77 | 63 | 92 | 40 |
Figure S65. Get High-res Image Gene #16: 'amp_10p15.1' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.0236 (Fisher's exact test), Q value = 0.046
Table S66. Gene #16: 'amp_10p15.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 16(10P15.1) MUTATED | 22 | 11 | 20 | 9 | 6 |
| AMP PEAK 16(10P15.1) WILD-TYPE | 48 | 71 | 80 | 25 | 48 |
Figure S66. Get High-res Image Gene #16: 'amp_10p15.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.04 (Fisher's exact test), Q value = 0.071
Table S67. Gene #17: 'amp_11q13.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 17(11Q13.3) MUTATED | 14 | 31 | 12 |
| AMP PEAK 17(11Q13.3) WILD-TYPE | 79 | 120 | 114 |
Figure S67. Get High-res Image Gene #17: 'amp_11q13.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00459 (Fisher's exact test), Q value = 0.013
Table S68. Gene #17: 'amp_11q13.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 17(11Q13.3) MUTATED | 17 | 18 | 10 | 6 | 4 |
| AMP PEAK 17(11Q13.3) WILD-TYPE | 50 | 68 | 61 | 65 | 65 |
Figure S68. Get High-res Image Gene #17: 'amp_11q13.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.0177 (Fisher's exact test), Q value = 0.036
Table S69. Gene #17: 'amp_11q13.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 17(11Q13.3) MUTATED | 10 | 7 | 25 | 11 | 2 |
| AMP PEAK 17(11Q13.3) WILD-TYPE | 44 | 49 | 89 | 73 | 54 |
Figure S69. Get High-res Image Gene #17: 'amp_11q13.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.0128 (Fisher's exact test), Q value = 0.028
Table S70. Gene #17: 'amp_11q13.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 17(11Q13.3) MUTATED | 10 | 16 | 13 | 5 | 6 | 7 |
| AMP PEAK 17(11Q13.3) WILD-TYPE | 30 | 47 | 79 | 79 | 40 | 33 |
Figure S70. Get High-res Image Gene #17: 'amp_11q13.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
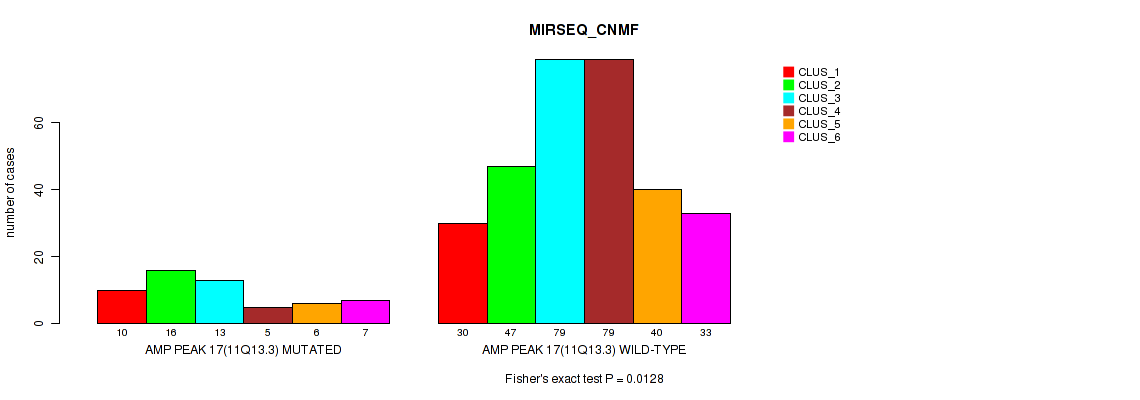
P value = 0.00331 (Fisher's exact test), Q value = 0.0096
Table S71. Gene #17: 'amp_11q13.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 17(11Q13.3) MUTATED | 11 | 23 | 5 | 16 | 2 |
| AMP PEAK 17(11Q13.3) WILD-TYPE | 33 | 72 | 51 | 106 | 46 |
Figure S71. Get High-res Image Gene #17: 'amp_11q13.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 1e-04 (Fisher's exact test), Q value = 5e-04
Table S72. Gene #17: 'amp_11q13.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 17(11Q13.3) MUTATED | 13 | 18 | 13 | 10 | 0 |
| AMP PEAK 17(11Q13.3) WILD-TYPE | 57 | 64 | 87 | 24 | 54 |
Figure S72. Get High-res Image Gene #17: 'amp_11q13.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S73. Gene #18: 'amp_13q32.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 18(13Q32.3) MUTATED | 1 | 42 | 24 |
| AMP PEAK 18(13Q32.3) WILD-TYPE | 92 | 109 | 102 |
Figure S73. Get High-res Image Gene #18: 'amp_13q32.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00938 (Fisher's exact test), Q value = 0.022
Table S74. Gene #18: 'amp_13q32.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 18(13Q32.3) MUTATED | 14 | 12 | 23 | 9 | 8 |
| AMP PEAK 18(13Q32.3) WILD-TYPE | 53 | 74 | 48 | 62 | 61 |
Figure S74. Get High-res Image Gene #18: 'amp_13q32.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.0126 (Fisher's exact test), Q value = 0.027
Table S75. Gene #18: 'amp_13q32.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 18(13Q32.3) MUTATED | 13 | 11 | 13 | 8 | 13 | 9 |
| AMP PEAK 18(13Q32.3) WILD-TYPE | 27 | 52 | 79 | 76 | 33 | 31 |
Figure S75. Get High-res Image Gene #18: 'amp_13q32.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.0062 (Fisher's exact test), Q value = 0.016
Table S76. Gene #18: 'amp_13q32.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| AMP PEAK 18(13Q32.3) MUTATED | 29 | 12 | 10 | 12 |
| AMP PEAK 18(13Q32.3) WILD-TYPE | 78 | 69 | 92 | 38 |
Figure S76. Get High-res Image Gene #18: 'amp_13q32.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.0351 (Fisher's exact test), Q value = 0.063
Table S77. Gene #18: 'amp_13q32.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 18(13Q32.3) MUTATED | 18 | 13 | 18 | 10 | 4 |
| AMP PEAK 18(13Q32.3) WILD-TYPE | 52 | 69 | 82 | 24 | 50 |
Figure S77. Get High-res Image Gene #18: 'amp_13q32.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S78. Gene #19: 'amp_13q33.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 19(13Q33.3) MUTATED | 3 | 39 | 25 |
| AMP PEAK 19(13Q33.3) WILD-TYPE | 90 | 112 | 101 |
Figure S78. Get High-res Image Gene #19: 'amp_13q33.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0464 (Fisher's exact test), Q value = 0.08
Table S79. Gene #19: 'amp_13q33.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 19(13Q33.3) MUTATED | 11 | 27 | 29 |
| AMP PEAK 19(13Q33.3) WILD-TYPE | 82 | 134 | 87 |
Figure S79. Get High-res Image Gene #19: 'amp_13q33.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.00162 (Fisher's exact test), Q value = 0.0053
Table S80. Gene #19: 'amp_13q33.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 19(13Q33.3) MUTATED | 13 | 10 | 25 | 8 | 10 |
| AMP PEAK 19(13Q33.3) WILD-TYPE | 54 | 76 | 46 | 63 | 59 |
Figure S80. Get High-res Image Gene #19: 'amp_13q33.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00949 (Fisher's exact test), Q value = 0.022
Table S81. Gene #19: 'amp_13q33.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 19(13Q33.3) MUTATED | 12 | 8 | 14 | 9 | 15 | 9 |
| AMP PEAK 19(13Q33.3) WILD-TYPE | 28 | 55 | 78 | 75 | 31 | 31 |
Figure S81. Get High-res Image Gene #19: 'amp_13q33.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00617 (Fisher's exact test), Q value = 0.016
Table S82. Gene #19: 'amp_13q33.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| AMP PEAK 19(13Q33.3) MUTATED | 28 | 12 | 10 | 13 |
| AMP PEAK 19(13Q33.3) WILD-TYPE | 79 | 69 | 92 | 37 |
Figure S82. Get High-res Image Gene #19: 'amp_13q33.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.0243 (Fisher's exact test), Q value = 0.047
Table S83. Gene #20: 'amp_15q26.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 20(15Q26.3) MUTATED | 8 | 15 | 18 | 7 | 5 |
| AMP PEAK 20(15Q26.3) WILD-TYPE | 59 | 71 | 53 | 64 | 64 |
Figure S83. Get High-res Image Gene #20: 'amp_15q26.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.0395 (Fisher's exact test), Q value = 0.07
Table S84. Gene #20: 'amp_15q26.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 20(15Q26.3) MUTATED | 9 | 7 | 17 | 18 | 2 |
| AMP PEAK 20(15Q26.3) WILD-TYPE | 45 | 49 | 97 | 66 | 54 |
Figure S84. Get High-res Image Gene #20: 'amp_15q26.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.042 (Fisher's exact test), Q value = 0.073
Table S85. Gene #21: 'amp_17p11.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 21(17P11.2) MUTATED | 5 | 23 | 18 |
| AMP PEAK 21(17P11.2) WILD-TYPE | 88 | 138 | 98 |
Figure S85. Get High-res Image Gene #21: 'amp_17p11.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S86. Gene #22: 'amp_17q25.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 22(17Q25.3) MUTATED | 18 | 82 | 45 |
| AMP PEAK 22(17Q25.3) WILD-TYPE | 75 | 69 | 81 |
Figure S86. Get High-res Image Gene #22: 'amp_17q25.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0478 (Fisher's exact test), Q value = 0.081
Table S87. Gene #22: 'amp_17q25.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 22(17Q25.3) MUTATED | 29 | 74 | 42 |
| AMP PEAK 22(17Q25.3) WILD-TYPE | 64 | 87 | 74 |
Figure S87. Get High-res Image Gene #22: 'amp_17q25.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.0113 (Fisher's exact test), Q value = 0.025
Table S88. Gene #22: 'amp_17q25.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 22(17Q25.3) MUTATED | 34 | 36 | 24 | 17 | 31 |
| AMP PEAK 22(17Q25.3) WILD-TYPE | 33 | 50 | 47 | 54 | 38 |
Figure S88. Get High-res Image Gene #22: 'amp_17q25.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.0106 (Fisher's exact test), Q value = 0.024
Table S89. Gene #22: 'amp_17q25.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 22(17Q25.3) MUTATED | 28 | 12 | 49 | 29 | 24 |
| AMP PEAK 22(17Q25.3) WILD-TYPE | 26 | 44 | 65 | 55 | 32 |
Figure S89. Get High-res Image Gene #22: 'amp_17q25.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00748 (Fisher's exact test), Q value = 0.018
Table S90. Gene #22: 'amp_17q25.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 22(17Q25.3) MUTATED | 22 | 30 | 29 | 38 | 10 | 15 |
| AMP PEAK 22(17Q25.3) WILD-TYPE | 18 | 33 | 63 | 46 | 36 | 25 |
Figure S90. Get High-res Image Gene #22: 'amp_17q25.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00131 (Fisher's exact test), Q value = 0.0046
Table S91. Gene #22: 'amp_17q25.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 22(17Q25.3) MUTATED | 26 | 40 | 24 | 32 | 22 |
| AMP PEAK 22(17Q25.3) WILD-TYPE | 18 | 55 | 32 | 90 | 26 |
Figure S91. Get High-res Image Gene #22: 'amp_17q25.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.011 (Fisher's exact test), Q value = 0.025
Table S92. Gene #22: 'amp_17q25.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 22(17Q25.3) MUTATED | 33 | 35 | 26 | 18 | 24 |
| AMP PEAK 22(17Q25.3) WILD-TYPE | 37 | 47 | 74 | 16 | 30 |
Figure S92. Get High-res Image Gene #22: 'amp_17q25.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.013 (Fisher's exact test), Q value = 0.028
Table S93. Gene #23: 'amp_19p13.12' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 23(19P13.12) MUTATED | 10 | 26 | 33 |
| AMP PEAK 23(19P13.12) WILD-TYPE | 83 | 125 | 93 |
Figure S93. Get High-res Image Gene #23: 'amp_19p13.12' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00118 (Fisher's exact test), Q value = 0.0042
Table S94. Gene #23: 'amp_19p13.12' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| AMP PEAK 23(19P13.12) MUTATED | 16 | 19 | 34 |
| AMP PEAK 23(19P13.12) WILD-TYPE | 77 | 142 | 82 |
Figure S94. Get High-res Image Gene #23: 'amp_19p13.12' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.00065 (Fisher's exact test), Q value = 0.0025
Table S95. Gene #23: 'amp_19p13.12' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 23(19P13.12) MUTATED | 15 | 10 | 20 | 19 | 4 |
| AMP PEAK 23(19P13.12) WILD-TYPE | 52 | 76 | 51 | 52 | 65 |
Figure S95. Get High-res Image Gene #23: 'amp_19p13.12' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00055 (Fisher's exact test), Q value = 0.0022
Table S96. Gene #23: 'amp_19p13.12' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 23(19P13.12) MUTATED | 16 | 13 | 12 | 23 | 4 |
| AMP PEAK 23(19P13.12) WILD-TYPE | 38 | 43 | 102 | 61 | 52 |
Figure S96. Get High-res Image Gene #23: 'amp_19p13.12' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00321 (Fisher's exact test), Q value = 0.0093
Table S97. Gene #23: 'amp_19p13.12' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 23(19P13.12) MUTATED | 4 | 8 | 23 | 9 | 9 | 15 |
| AMP PEAK 23(19P13.12) WILD-TYPE | 36 | 55 | 69 | 75 | 37 | 25 |
Figure S97. Get High-res Image Gene #23: 'amp_19p13.12' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00223 (Fisher's exact test), Q value = 0.0068
Table S98. Gene #23: 'amp_19p13.12' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 23(19P13.12) MUTATED | 3 | 20 | 14 | 29 | 2 |
| AMP PEAK 23(19P13.12) WILD-TYPE | 41 | 75 | 42 | 93 | 46 |
Figure S98. Get High-res Image Gene #23: 'amp_19p13.12' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.00247 (Fisher's exact test), Q value = 0.0074
Table S99. Gene #23: 'amp_19p13.12' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| AMP PEAK 23(19P13.12) MUTATED | 24 | 22 | 8 | 10 |
| AMP PEAK 23(19P13.12) WILD-TYPE | 83 | 59 | 94 | 40 |
Figure S99. Get High-res Image Gene #23: 'amp_19p13.12' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00129 (Fisher's exact test), Q value = 0.0045
Table S100. Gene #23: 'amp_19p13.12' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 23(19P13.12) MUTATED | 18 | 14 | 27 | 2 | 3 |
| AMP PEAK 23(19P13.12) WILD-TYPE | 52 | 68 | 73 | 32 | 51 |
Figure S100. Get High-res Image Gene #23: 'amp_19p13.12' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.0164 (Fisher's exact test), Q value = 0.034
Table S101. Gene #24: 'amp_19q13.11' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 24(19Q13.11) MUTATED | 14 | 47 | 30 |
| AMP PEAK 24(19Q13.11) WILD-TYPE | 79 | 104 | 96 |
Figure S101. Get High-res Image Gene #24: 'amp_19q13.11' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00041 (Fisher's exact test), Q value = 0.0017
Table S102. Gene #24: 'amp_19q13.11' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 24(19Q13.11) MUTATED | 22 | 16 | 25 | 21 | 6 |
| AMP PEAK 24(19Q13.11) WILD-TYPE | 45 | 70 | 46 | 50 | 63 |
Figure S102. Get High-res Image Gene #24: 'amp_19q13.11' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00066 (Fisher's exact test), Q value = 0.0026
Table S103. Gene #24: 'amp_19q13.11' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 24(19Q13.11) MUTATED | 16 | 17 | 21 | 31 | 5 |
| AMP PEAK 24(19Q13.11) WILD-TYPE | 38 | 39 | 93 | 53 | 51 |
Figure S103. Get High-res Image Gene #24: 'amp_19q13.11' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.0325 (Fisher's exact test), Q value = 0.06
Table S104. Gene #24: 'amp_19q13.11' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 24(19Q13.11) MUTATED | 12 | 15 | 28 | 10 | 12 | 13 |
| AMP PEAK 24(19Q13.11) WILD-TYPE | 28 | 48 | 64 | 74 | 34 | 27 |
Figure S104. Get High-res Image Gene #24: 'amp_19q13.11' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.0018 (Fisher's exact test), Q value = 0.0057
Table S105. Gene #24: 'amp_19q13.11' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 24(19Q13.11) MUTATED | 14 | 23 | 18 | 33 | 2 |
| AMP PEAK 24(19Q13.11) WILD-TYPE | 30 | 72 | 38 | 89 | 46 |
Figure S105. Get High-res Image Gene #24: 'amp_19q13.11' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S106. Gene #24: 'amp_19q13.11' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| AMP PEAK 24(19Q13.11) MUTATED | 36 | 23 | 7 | 19 |
| AMP PEAK 24(19Q13.11) WILD-TYPE | 71 | 58 | 95 | 31 |
Figure S106. Get High-res Image Gene #24: 'amp_19q13.11' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00027 (Fisher's exact test), Q value = 0.0012
Table S107. Gene #24: 'amp_19q13.11' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 24(19Q13.11) MUTATED | 24 | 20 | 29 | 10 | 2 |
| AMP PEAK 24(19Q13.11) WILD-TYPE | 46 | 62 | 71 | 24 | 52 |
Figure S107. Get High-res Image Gene #24: 'amp_19q13.11' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S108. Gene #25: 'amp_20q13.13' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 25(20Q13.13) MUTATED | 11 | 69 | 44 |
| AMP PEAK 25(20Q13.13) WILD-TYPE | 82 | 82 | 82 |
Figure S108. Get High-res Image Gene #25: 'amp_20q13.13' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0168 (Fisher's exact test), Q value = 0.034
Table S109. Gene #25: 'amp_20q13.13' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 25(20Q13.13) MUTATED | 26 | 29 | 32 | 23 | 13 |
| AMP PEAK 25(20Q13.13) WILD-TYPE | 41 | 57 | 39 | 48 | 56 |
Figure S109. Get High-res Image Gene #25: 'amp_20q13.13' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00035 (Fisher's exact test), Q value = 0.0015
Table S110. Gene #25: 'amp_20q13.13' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 25(20Q13.13) MUTATED | 19 | 19 | 37 | 41 | 7 |
| AMP PEAK 25(20Q13.13) WILD-TYPE | 35 | 37 | 77 | 43 | 49 |
Figure S110. Get High-res Image Gene #25: 'amp_20q13.13' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.0108 (Fisher's exact test), Q value = 0.024
Table S111. Gene #25: 'amp_20q13.13' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 25(20Q13.13) MUTATED | 21 | 29 | 24 | 41 | 8 |
| AMP PEAK 25(20Q13.13) WILD-TYPE | 23 | 66 | 32 | 81 | 40 |
Figure S111. Get High-res Image Gene #25: 'amp_20q13.13' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 5e-05 (Fisher's exact test), Q value = 3e-04
Table S112. Gene #25: 'amp_20q13.13' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| AMP PEAK 25(20Q13.13) MUTATED | 45 | 25 | 19 | 27 |
| AMP PEAK 25(20Q13.13) WILD-TYPE | 62 | 56 | 83 | 23 |
Figure S112. Get High-res Image Gene #25: 'amp_20q13.13' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.0477 (Fisher's exact test), Q value = 0.081
Table S113. Gene #25: 'amp_20q13.13' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 25(20Q13.13) MUTATED | 28 | 24 | 37 | 16 | 11 |
| AMP PEAK 25(20Q13.13) WILD-TYPE | 42 | 58 | 63 | 18 | 43 |
Figure S113. Get High-res Image Gene #25: 'amp_20q13.13' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S114. Gene #26: 'amp_20q13.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 26(20Q13.33) MUTATED | 10 | 69 | 49 |
| AMP PEAK 26(20Q13.33) WILD-TYPE | 83 | 82 | 77 |
Figure S114. Get High-res Image Gene #26: 'amp_20q13.33' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00482 (Fisher's exact test), Q value = 0.013
Table S115. Gene #26: 'amp_20q13.33' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 26(20Q13.33) MUTATED | 25 | 30 | 36 | 22 | 14 |
| AMP PEAK 26(20Q13.33) WILD-TYPE | 42 | 56 | 35 | 49 | 55 |
Figure S115. Get High-res Image Gene #26: 'amp_20q13.33' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 6e-05 (Fisher's exact test), Q value = 0.00034
Table S116. Gene #26: 'amp_20q13.33' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 26(20Q13.33) MUTATED | 21 | 17 | 39 | 43 | 7 |
| AMP PEAK 26(20Q13.33) WILD-TYPE | 33 | 39 | 75 | 41 | 49 |
Figure S116. Get High-res Image Gene #26: 'amp_20q13.33' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S117. Gene #26: 'amp_20q13.33' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| AMP PEAK 26(20Q13.33) MUTATED | 48 | 25 | 20 | 27 |
| AMP PEAK 26(20Q13.33) WILD-TYPE | 59 | 56 | 82 | 23 |
Figure S117. Get High-res Image Gene #26: 'amp_20q13.33' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00178 (Fisher's exact test), Q value = 0.0057
Table S118. Gene #27: 'amp_xq28' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| AMP PEAK 27(XQ28) MUTATED | 22 | 23 | 42 |
| AMP PEAK 27(XQ28) WILD-TYPE | 71 | 128 | 84 |
Figure S118. Get High-res Image Gene #27: 'amp_xq28' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0275 (Fisher's exact test), Q value = 0.052
Table S119. Gene #27: 'amp_xq28' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| AMP PEAK 27(XQ28) MUTATED | 14 | 22 | 12 | 12 | 26 |
| AMP PEAK 27(XQ28) WILD-TYPE | 53 | 64 | 59 | 59 | 43 |
Figure S119. Get High-res Image Gene #27: 'amp_xq28' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00997 (Fisher's exact test), Q value = 0.023
Table S120. Gene #27: 'amp_xq28' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| AMP PEAK 27(XQ28) MUTATED | 13 | 11 | 29 | 11 | 22 |
| AMP PEAK 27(XQ28) WILD-TYPE | 41 | 45 | 85 | 73 | 34 |
Figure S120. Get High-res Image Gene #27: 'amp_xq28' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00011 (Fisher's exact test), Q value = 0.00054
Table S121. Gene #27: 'amp_xq28' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| AMP PEAK 27(XQ28) MUTATED | 4 | 17 | 15 | 35 | 5 | 9 |
| AMP PEAK 27(XQ28) WILD-TYPE | 36 | 46 | 77 | 49 | 41 | 31 |
Figure S121. Get High-res Image Gene #27: 'amp_xq28' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00335 (Fisher's exact test), Q value = 0.0096
Table S122. Gene #27: 'amp_xq28' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| AMP PEAK 27(XQ28) MUTATED | 5 | 23 | 14 | 22 | 21 |
| AMP PEAK 27(XQ28) WILD-TYPE | 39 | 72 | 42 | 100 | 27 |
Figure S122. Get High-res Image Gene #27: 'amp_xq28' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.0158 (Fisher's exact test), Q value = 0.033
Table S123. Gene #27: 'amp_xq28' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| AMP PEAK 27(XQ28) MUTATED | 23 | 15 | 34 | 6 |
| AMP PEAK 27(XQ28) WILD-TYPE | 84 | 66 | 68 | 44 |
Figure S123. Get High-res Image Gene #27: 'amp_xq28' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00034 (Fisher's exact test), Q value = 0.0015
Table S124. Gene #27: 'amp_xq28' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| AMP PEAK 27(XQ28) MUTATED | 16 | 19 | 17 | 2 | 24 |
| AMP PEAK 27(XQ28) WILD-TYPE | 54 | 63 | 83 | 32 | 30 |
Figure S124. Get High-res Image Gene #27: 'amp_xq28' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S125. Gene #28: 'del_1p36.31' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 1(1P36.31) MUTATED | 18 | 62 | 82 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 75 | 89 | 44 |
Figure S125. Get High-res Image Gene #28: 'del_1p36.31' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S126. Gene #28: 'del_1p36.31' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 1(1P36.31) MUTATED | 28 | 61 | 73 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 65 | 100 | 43 |
Figure S126. Get High-res Image Gene #28: 'del_1p36.31' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S127. Gene #28: 'del_1p36.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 1(1P36.31) MUTATED | 17 | 33 | 59 | 32 | 18 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 50 | 53 | 12 | 39 | 51 |
Figure S127. Get High-res Image Gene #28: 'del_1p36.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S128. Gene #28: 'del_1p36.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 1(1P36.31) MUTATED | 24 | 24 | 40 | 57 | 14 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 30 | 32 | 74 | 27 | 42 |
Figure S128. Get High-res Image Gene #28: 'del_1p36.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00149 (Fisher's exact test), Q value = 0.005
Table S129. Gene #28: 'del_1p36.31' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 1(1P36.31) MUTATED | 12 | 26 | 39 | 30 | 23 | 29 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 28 | 37 | 53 | 54 | 23 | 11 |
Figure S129. Get High-res Image Gene #28: 'del_1p36.31' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00113 (Fisher's exact test), Q value = 0.0041
Table S130. Gene #28: 'del_1p36.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 1(1P36.31) MUTATED | 14 | 37 | 31 | 65 | 12 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 30 | 58 | 25 | 57 | 36 |
Figure S130. Get High-res Image Gene #28: 'del_1p36.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.00398 (Fisher's exact test), Q value = 0.011
Table S131. Gene #28: 'del_1p36.31' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 1(1P36.31) MUTATED | 56 | 39 | 30 | 25 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 51 | 42 | 72 | 25 |
Figure S131. Get High-res Image Gene #28: 'del_1p36.31' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00469 (Fisher's exact test), Q value = 0.013
Table S132. Gene #28: 'del_1p36.31' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 1(1P36.31) MUTATED | 33 | 34 | 57 | 11 | 15 |
| DEL PEAK 1(1P36.31) WILD-TYPE | 37 | 48 | 43 | 23 | 39 |
Figure S132. Get High-res Image Gene #28: 'del_1p36.31' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S133. Gene #29: 'del_1p36.11' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 2(1P36.11) MUTATED | 11 | 56 | 75 |
| DEL PEAK 2(1P36.11) WILD-TYPE | 82 | 95 | 51 |
Figure S133. Get High-res Image Gene #29: 'del_1p36.11' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S134. Gene #29: 'del_1p36.11' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 2(1P36.11) MUTATED | 24 | 48 | 70 |
| DEL PEAK 2(1P36.11) WILD-TYPE | 69 | 113 | 46 |
Figure S134. Get High-res Image Gene #29: 'del_1p36.11' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S135. Gene #29: 'del_1p36.11' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 2(1P36.11) MUTATED | 17 | 23 | 58 | 25 | 17 |
| DEL PEAK 2(1P36.11) WILD-TYPE | 50 | 63 | 13 | 46 | 52 |
Figure S135. Get High-res Image Gene #29: 'del_1p36.11' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S136. Gene #29: 'del_1p36.11' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 2(1P36.11) MUTATED | 22 | 19 | 31 | 54 | 14 |
| DEL PEAK 2(1P36.11) WILD-TYPE | 32 | 37 | 83 | 30 | 42 |
Figure S136. Get High-res Image Gene #29: 'del_1p36.11' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 2e-05 (Fisher's exact test), Q value = 0.00014
Table S137. Gene #29: 'del_1p36.11' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 2(1P36.11) MUTATED | 8 | 18 | 33 | 28 | 25 | 27 |
| DEL PEAK 2(1P36.11) WILD-TYPE | 32 | 45 | 59 | 56 | 21 | 13 |
Figure S137. Get High-res Image Gene #29: 'del_1p36.11' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 7e-05 (Fisher's exact test), Q value = 0.00038
Table S138. Gene #29: 'del_1p36.11' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 2(1P36.11) MUTATED | 9 | 30 | 25 | 64 | 11 |
| DEL PEAK 2(1P36.11) WILD-TYPE | 35 | 65 | 31 | 58 | 37 |
Figure S138. Get High-res Image Gene #29: 'del_1p36.11' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.00054 (Fisher's exact test), Q value = 0.0022
Table S139. Gene #29: 'del_1p36.11' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 2(1P36.11) MUTATED | 52 | 34 | 23 | 23 |
| DEL PEAK 2(1P36.11) WILD-TYPE | 55 | 47 | 79 | 27 |
Figure S139. Get High-res Image Gene #29: 'del_1p36.11' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00026 (Fisher's exact test), Q value = 0.0012
Table S140. Gene #29: 'del_1p36.11' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 2(1P36.11) MUTATED | 30 | 26 | 55 | 7 | 14 |
| DEL PEAK 2(1P36.11) WILD-TYPE | 40 | 56 | 45 | 27 | 40 |
Figure S140. Get High-res Image Gene #29: 'del_1p36.11' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.00308 (Fisher's exact test), Q value = 0.009
Table S141. Gene #30: 'del_2q22.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 3(2Q22.1) MUTATED | 6 | 32 | 14 |
| DEL PEAK 3(2Q22.1) WILD-TYPE | 87 | 119 | 112 |
Figure S141. Get High-res Image Gene #30: 'del_2q22.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00471 (Fisher's exact test), Q value = 0.013
Table S142. Gene #30: 'del_2q22.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 3(2Q22.1) MUTATED | 14 | 16 | 13 | 7 | 2 |
| DEL PEAK 3(2Q22.1) WILD-TYPE | 53 | 70 | 58 | 64 | 67 |
Figure S142. Get High-res Image Gene #30: 'del_2q22.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 5e-05 (Fisher's exact test), Q value = 3e-04
Table S143. Gene #31: 'del_2q37.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 4(2Q37.3) MUTATED | 5 | 39 | 18 |
| DEL PEAK 4(2Q37.3) WILD-TYPE | 88 | 112 | 108 |
Figure S143. Get High-res Image Gene #31: 'del_2q37.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S144. Gene #31: 'del_2q37.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 4(2Q37.3) MUTATED | 18 | 22 | 14 | 8 | 0 |
| DEL PEAK 4(2Q37.3) WILD-TYPE | 49 | 64 | 57 | 63 | 69 |
Figure S144. Get High-res Image Gene #31: 'del_2q37.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00091 (Fisher's exact test), Q value = 0.0034
Table S145. Gene #31: 'del_2q37.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 4(2Q37.3) MUTATED | 12 | 9 | 28 | 12 | 1 |
| DEL PEAK 4(2Q37.3) WILD-TYPE | 42 | 47 | 86 | 72 | 55 |
Figure S145. Get High-res Image Gene #31: 'del_2q37.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00994 (Fisher's exact test), Q value = 0.023
Table S146. Gene #31: 'del_2q37.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 4(2Q37.3) MUTATED | 11 | 16 | 16 | 15 | 3 |
| DEL PEAK 4(2Q37.3) WILD-TYPE | 33 | 79 | 40 | 107 | 45 |
Figure S146. Get High-res Image Gene #31: 'del_2q37.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.00148 (Fisher's exact test), Q value = 0.005
Table S147. Gene #31: 'del_2q37.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 4(2Q37.3) MUTATED | 21 | 14 | 10 | 9 | 4 |
| DEL PEAK 4(2Q37.3) WILD-TYPE | 49 | 68 | 90 | 25 | 50 |
Figure S147. Get High-res Image Gene #31: 'del_2q37.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.00882 (Fisher's exact test), Q value = 0.021
Table S148. Gene #32: 'del_3p13' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 5(3P13) MUTATED | 20 | 21 | 32 |
| DEL PEAK 5(3P13) WILD-TYPE | 73 | 140 | 84 |
Figure S148. Get High-res Image Gene #32: 'del_3p13' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.00011 (Fisher's exact test), Q value = 0.00054
Table S149. Gene #32: 'del_3p13' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 5(3P13) MUTATED | 13 | 10 | 26 | 18 | 5 |
| DEL PEAK 5(3P13) WILD-TYPE | 54 | 76 | 45 | 53 | 64 |
Figure S149. Get High-res Image Gene #32: 'del_3p13' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S150. Gene #32: 'del_3p13' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 5(3P13) MUTATED | 23 | 5 | 15 | 24 | 5 |
| DEL PEAK 5(3P13) WILD-TYPE | 31 | 51 | 99 | 60 | 51 |
Figure S150. Get High-res Image Gene #32: 'del_3p13' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 5e-05 (Fisher's exact test), Q value = 3e-04
Table S151. Gene #32: 'del_3p13' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 5(3P13) MUTATED | 5 | 8 | 20 | 9 | 9 | 20 |
| DEL PEAK 5(3P13) WILD-TYPE | 35 | 55 | 72 | 75 | 37 | 20 |
Figure S151. Get High-res Image Gene #32: 'del_3p13' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.0335 (Fisher's exact test), Q value = 0.061
Table S152. Gene #32: 'del_3p13' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 5(3P13) MUTATED | 5 | 17 | 12 | 33 | 4 |
| DEL PEAK 5(3P13) WILD-TYPE | 39 | 78 | 44 | 89 | 44 |
Figure S152. Get High-res Image Gene #32: 'del_3p13' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.00993 (Fisher's exact test), Q value = 0.023
Table S153. Gene #32: 'del_3p13' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 5(3P13) MUTATED | 29 | 19 | 10 | 9 |
| DEL PEAK 5(3P13) WILD-TYPE | 78 | 62 | 92 | 41 |
Figure S153. Get High-res Image Gene #32: 'del_3p13' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00163 (Fisher's exact test), Q value = 0.0053
Table S154. Gene #32: 'del_3p13' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 5(3P13) MUTATED | 20 | 12 | 28 | 3 | 4 |
| DEL PEAK 5(3P13) WILD-TYPE | 50 | 70 | 72 | 31 | 50 |
Figure S154. Get High-res Image Gene #32: 'del_3p13' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S155. Gene #33: 'del_4p16.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 6(4P16.3) MUTATED | 4 | 51 | 30 |
| DEL PEAK 6(4P16.3) WILD-TYPE | 89 | 100 | 96 |
Figure S155. Get High-res Image Gene #33: 'del_4p16.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.027 (Fisher's exact test), Q value = 0.051
Table S156. Gene #33: 'del_4p16.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 6(4P16.3) MUTATED | 6 | 20 | 20 | 32 | 6 |
| DEL PEAK 6(4P16.3) WILD-TYPE | 38 | 75 | 36 | 90 | 42 |
Figure S156. Get High-res Image Gene #33: 'del_4p16.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.00557 (Fisher's exact test), Q value = 0.014
Table S157. Gene #33: 'del_4p16.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 6(4P16.3) MUTATED | 26 | 20 | 22 | 4 | 6 |
| DEL PEAK 6(4P16.3) WILD-TYPE | 44 | 62 | 78 | 30 | 48 |
Figure S157. Get High-res Image Gene #33: 'del_4p16.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S158. Gene #34: 'del_4q21.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 7(4Q21.3) MUTATED | 7 | 108 | 56 |
| DEL PEAK 7(4Q21.3) WILD-TYPE | 86 | 43 | 70 |
Figure S158. Get High-res Image Gene #34: 'del_4q21.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00095 (Fisher's exact test), Q value = 0.0035
Table S159. Gene #34: 'del_4q21.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 7(4Q21.3) MUTATED | 35 | 66 | 70 |
| DEL PEAK 7(4Q21.3) WILD-TYPE | 58 | 95 | 46 |
Figure S159. Get High-res Image Gene #34: 'del_4q21.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S160. Gene #34: 'del_4q21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 7(4Q21.3) MUTATED | 39 | 43 | 52 | 20 | 15 |
| DEL PEAK 7(4Q21.3) WILD-TYPE | 28 | 43 | 19 | 51 | 54 |
Figure S160. Get High-res Image Gene #34: 'del_4q21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00021 (Fisher's exact test), Q value = 0.00098
Table S161. Gene #34: 'del_4q21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 7(4Q21.3) MUTATED | 28 | 22 | 55 | 51 | 13 |
| DEL PEAK 7(4Q21.3) WILD-TYPE | 26 | 34 | 59 | 33 | 43 |
Figure S161. Get High-res Image Gene #34: 'del_4q21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00133 (Fisher's exact test), Q value = 0.0046
Table S162. Gene #34: 'del_4q21.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 7(4Q21.3) MUTATED | 20 | 28 | 39 | 27 | 30 | 26 |
| DEL PEAK 7(4Q21.3) WILD-TYPE | 20 | 35 | 53 | 57 | 16 | 14 |
Figure S162. Get High-res Image Gene #34: 'del_4q21.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00154 (Fisher's exact test), Q value = 0.0051
Table S163. Gene #34: 'del_4q21.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 7(4Q21.3) MUTATED | 20 | 45 | 32 | 63 | 10 |
| DEL PEAK 7(4Q21.3) WILD-TYPE | 24 | 50 | 24 | 59 | 38 |
Figure S163. Get High-res Image Gene #34: 'del_4q21.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 6e-05 (Fisher's exact test), Q value = 0.00034
Table S164. Gene #34: 'del_4q21.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 7(4Q21.3) MUTATED | 62 | 33 | 34 | 34 |
| DEL PEAK 7(4Q21.3) WILD-TYPE | 45 | 48 | 68 | 16 |
Figure S164. Get High-res Image Gene #34: 'del_4q21.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 6e-05 (Fisher's exact test), Q value = 0.00034
Table S165. Gene #34: 'del_4q21.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 7(4Q21.3) MUTATED | 45 | 42 | 50 | 14 | 12 |
| DEL PEAK 7(4Q21.3) WILD-TYPE | 25 | 40 | 50 | 20 | 42 |
Figure S165. Get High-res Image Gene #34: 'del_4q21.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S166. Gene #35: 'del_4q24' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 8(4Q24) MUTATED | 5 | 111 | 57 |
| DEL PEAK 8(4Q24) WILD-TYPE | 88 | 40 | 69 |
Figure S166. Get High-res Image Gene #35: 'del_4q24' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00081 (Fisher's exact test), Q value = 0.0031
Table S167. Gene #35: 'del_4q24' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 8(4Q24) MUTATED | 36 | 66 | 71 |
| DEL PEAK 8(4Q24) WILD-TYPE | 57 | 95 | 45 |
Figure S167. Get High-res Image Gene #35: 'del_4q24' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S168. Gene #35: 'del_4q24' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 8(4Q24) MUTATED | 38 | 43 | 52 | 22 | 15 |
| DEL PEAK 8(4Q24) WILD-TYPE | 29 | 43 | 19 | 49 | 54 |
Figure S168. Get High-res Image Gene #35: 'del_4q24' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 5e-05 (Fisher's exact test), Q value = 3e-04
Table S169. Gene #35: 'del_4q24' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 8(4Q24) MUTATED | 29 | 23 | 54 | 52 | 12 |
| DEL PEAK 8(4Q24) WILD-TYPE | 25 | 33 | 60 | 32 | 44 |
Figure S169. Get High-res Image Gene #35: 'del_4q24' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00183 (Fisher's exact test), Q value = 0.0058
Table S170. Gene #35: 'del_4q24' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 8(4Q24) MUTATED | 21 | 27 | 40 | 28 | 30 | 26 |
| DEL PEAK 8(4Q24) WILD-TYPE | 19 | 36 | 52 | 56 | 16 | 14 |
Figure S170. Get High-res Image Gene #35: 'del_4q24' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00021 (Fisher's exact test), Q value = 0.00098
Table S171. Gene #35: 'del_4q24' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 8(4Q24) MUTATED | 23 | 43 | 33 | 64 | 9 |
| DEL PEAK 8(4Q24) WILD-TYPE | 21 | 52 | 23 | 58 | 39 |
Figure S171. Get High-res Image Gene #35: 'del_4q24' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 2e-05 (Fisher's exact test), Q value = 0.00014
Table S172. Gene #35: 'del_4q24' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 8(4Q24) MUTATED | 61 | 34 | 34 | 36 |
| DEL PEAK 8(4Q24) WILD-TYPE | 46 | 47 | 68 | 14 |
Figure S172. Get High-res Image Gene #35: 'del_4q24' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 4e-05 (Fisher's exact test), Q value = 0.00026
Table S173. Gene #35: 'del_4q24' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 8(4Q24) MUTATED | 45 | 42 | 50 | 17 | 11 |
| DEL PEAK 8(4Q24) WILD-TYPE | 25 | 40 | 50 | 17 | 43 |
Figure S173. Get High-res Image Gene #35: 'del_4q24' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S174. Gene #36: 'del_4q35.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 9(4Q35.1) MUTATED | 9 | 102 | 61 |
| DEL PEAK 9(4Q35.1) WILD-TYPE | 84 | 49 | 65 |
Figure S174. Get High-res Image Gene #36: 'del_4q35.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00036 (Fisher's exact test), Q value = 0.0015
Table S175. Gene #36: 'del_4q35.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 9(4Q35.1) MUTATED | 33 | 68 | 71 |
| DEL PEAK 9(4Q35.1) WILD-TYPE | 60 | 93 | 45 |
Figure S175. Get High-res Image Gene #36: 'del_4q35.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S176. Gene #36: 'del_4q35.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 9(4Q35.1) MUTATED | 39 | 39 | 50 | 21 | 20 |
| DEL PEAK 9(4Q35.1) WILD-TYPE | 28 | 47 | 21 | 50 | 49 |
Figure S176. Get High-res Image Gene #36: 'del_4q35.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00072 (Fisher's exact test), Q value = 0.0028
Table S177. Gene #36: 'del_4q35.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 9(4Q35.1) MUTATED | 28 | 22 | 48 | 54 | 17 |
| DEL PEAK 9(4Q35.1) WILD-TYPE | 26 | 34 | 66 | 30 | 39 |
Figure S177. Get High-res Image Gene #36: 'del_4q35.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00188 (Fisher's exact test), Q value = 0.0059
Table S178. Gene #36: 'del_4q35.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 9(4Q35.1) MUTATED | 22 | 26 | 36 | 31 | 29 | 27 |
| DEL PEAK 9(4Q35.1) WILD-TYPE | 18 | 37 | 56 | 53 | 17 | 13 |
Figure S178. Get High-res Image Gene #36: 'del_4q35.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00597 (Fisher's exact test), Q value = 0.015
Table S179. Gene #36: 'del_4q35.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 9(4Q35.1) MUTATED | 21 | 41 | 32 | 65 | 12 |
| DEL PEAK 9(4Q35.1) WILD-TYPE | 23 | 54 | 24 | 57 | 36 |
Figure S179. Get High-res Image Gene #36: 'del_4q35.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 9e-05 (Fisher's exact test), Q value = 0.00045
Table S180. Gene #36: 'del_4q35.1' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 9(4Q35.1) MUTATED | 60 | 36 | 32 | 33 |
| DEL PEAK 9(4Q35.1) WILD-TYPE | 47 | 45 | 70 | 17 |
Figure S180. Get High-res Image Gene #36: 'del_4q35.1' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00148 (Fisher's exact test), Q value = 0.005
Table S181. Gene #36: 'del_4q35.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 9(4Q35.1) MUTATED | 42 | 39 | 51 | 16 | 13 |
| DEL PEAK 9(4Q35.1) WILD-TYPE | 28 | 43 | 49 | 18 | 41 |
Figure S181. Get High-res Image Gene #36: 'del_4q35.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.00152 (Fisher's exact test), Q value = 0.0051
Table S182. Gene #37: 'del_6q27' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 10(6Q27) MUTATED | 20 | 64 | 52 |
| DEL PEAK 10(6Q27) WILD-TYPE | 73 | 87 | 74 |
Figure S182. Get High-res Image Gene #37: 'del_6q27' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0054 (Fisher's exact test), Q value = 0.014
Table S183. Gene #37: 'del_6q27' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 10(6Q27) MUTATED | 23 | 72 | 41 |
| DEL PEAK 10(6Q27) WILD-TYPE | 70 | 89 | 75 |
Figure S183. Get High-res Image Gene #37: 'del_6q27' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.0289 (Fisher's exact test), Q value = 0.054
Table S184. Gene #37: 'del_6q27' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 10(6Q27) MUTATED | 19 | 20 | 23 | 35 | 15 | 20 |
| DEL PEAK 10(6Q27) WILD-TYPE | 21 | 43 | 69 | 49 | 31 | 20 |
Figure S184. Get High-res Image Gene #37: 'del_6q27' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.0391 (Fisher's exact test), Q value = 0.07
Table S185. Gene #37: 'del_6q27' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 10(6Q27) MUTATED | 20 | 25 | 18 | 45 | 24 |
| DEL PEAK 10(6Q27) WILD-TYPE | 24 | 70 | 38 | 77 | 24 |
Figure S185. Get High-res Image Gene #37: 'del_6q27' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.0408 (Fisher's exact test), Q value = 0.072
Table S186. Gene #38: 'del_7q36.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 11(7Q36.3) MUTATED | 8 | 9 | 17 |
| DEL PEAK 11(7Q36.3) WILD-TYPE | 85 | 152 | 99 |
Figure S186. Get High-res Image Gene #38: 'del_7q36.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.0414 (Fisher's exact test), Q value = 0.072
Table S187. Gene #38: 'del_7q36.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 11(7Q36.3) MUTATED | 10 | 4 | 11 | 6 | 3 |
| DEL PEAK 11(7Q36.3) WILD-TYPE | 57 | 82 | 60 | 65 | 66 |
Figure S187. Get High-res Image Gene #38: 'del_7q36.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00032 (Fisher's exact test), Q value = 0.0014
Table S188. Gene #38: 'del_7q36.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 11(7Q36.3) MUTATED | 19 | 5 | 2 | 7 |
| DEL PEAK 11(7Q36.3) WILD-TYPE | 88 | 76 | 100 | 43 |
Figure S188. Get High-res Image Gene #38: 'del_7q36.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00772 (Fisher's exact test), Q value = 0.019
Table S189. Gene #38: 'del_7q36.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 11(7Q36.3) MUTATED | 11 | 8 | 8 | 6 | 0 |
| DEL PEAK 11(7Q36.3) WILD-TYPE | 59 | 74 | 92 | 28 | 54 |
Figure S189. Get High-res Image Gene #38: 'del_7q36.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S190. Gene #39: 'del_8p23.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 12(8P23.2) MUTATED | 46 | 96 | 105 |
| DEL PEAK 12(8P23.2) WILD-TYPE | 47 | 55 | 21 |
Figure S190. Get High-res Image Gene #39: 'del_8p23.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S191. Gene #40: 'del_8p12' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 13(8P12) MUTATED | 44 | 91 | 107 |
| DEL PEAK 13(8P12) WILD-TYPE | 49 | 60 | 19 |
Figure S191. Get High-res Image Gene #40: 'del_8p12' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S192. Gene #41: 'del_9p21.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 14(9P21.3) MUTATED | 13 | 76 | 61 |
| DEL PEAK 14(9P21.3) WILD-TYPE | 80 | 75 | 65 |
Figure S192. Get High-res Image Gene #41: 'del_9p21.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0242 (Fisher's exact test), Q value = 0.047
Table S193. Gene #41: 'del_9p21.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 14(9P21.3) MUTATED | 34 | 57 | 59 |
| DEL PEAK 14(9P21.3) WILD-TYPE | 59 | 104 | 57 |
Figure S193. Get High-res Image Gene #41: 'del_9p21.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S194. Gene #41: 'del_9p21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 14(9P21.3) MUTATED | 31 | 29 | 45 | 27 | 15 |
| DEL PEAK 14(9P21.3) WILD-TYPE | 36 | 57 | 26 | 44 | 54 |
Figure S194. Get High-res Image Gene #41: 'del_9p21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00175 (Fisher's exact test), Q value = 0.0057
Table S195. Gene #41: 'del_9p21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 14(9P21.3) MUTATED | 28 | 20 | 38 | 46 | 15 |
| DEL PEAK 14(9P21.3) WILD-TYPE | 26 | 36 | 76 | 38 | 41 |
Figure S195. Get High-res Image Gene #41: 'del_9p21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 4e-05 (Fisher's exact test), Q value = 0.00026
Table S196. Gene #41: 'del_9p21.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 14(9P21.3) MUTATED | 20 | 16 | 35 | 26 | 31 | 21 |
| DEL PEAK 14(9P21.3) WILD-TYPE | 20 | 47 | 57 | 58 | 15 | 19 |
Figure S196. Get High-res Image Gene #41: 'del_9p21.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00215 (Fisher's exact test), Q value = 0.0066
Table S197. Gene #41: 'del_9p21.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 14(9P21.3) MUTATED | 20 | 31 | 29 | 59 | 10 |
| DEL PEAK 14(9P21.3) WILD-TYPE | 24 | 64 | 27 | 63 | 38 |
Figure S197. Get High-res Image Gene #41: 'del_9p21.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.0208 (Fisher's exact test), Q value = 0.041
Table S198. Gene #41: 'del_9p21.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 14(9P21.3) MUTATED | 49 | 30 | 34 | 29 |
| DEL PEAK 14(9P21.3) WILD-TYPE | 58 | 51 | 68 | 21 |
Figure S198. Get High-res Image Gene #41: 'del_9p21.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00478 (Fisher's exact test), Q value = 0.013
Table S199. Gene #41: 'del_9p21.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 14(9P21.3) MUTATED | 40 | 28 | 45 | 15 | 14 |
| DEL PEAK 14(9P21.3) WILD-TYPE | 30 | 54 | 55 | 19 | 40 |
Figure S199. Get High-res Image Gene #41: 'del_9p21.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S200. Gene #42: 'del_9q31.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 15(9Q31.3) MUTATED | 10 | 66 | 43 |
| DEL PEAK 15(9Q31.3) WILD-TYPE | 83 | 85 | 83 |
Figure S200. Get High-res Image Gene #42: 'del_9q31.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00897 (Fisher's exact test), Q value = 0.021
Table S201. Gene #42: 'del_9q31.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 15(9Q31.3) MUTATED | 23 | 46 | 50 |
| DEL PEAK 15(9Q31.3) WILD-TYPE | 70 | 115 | 66 |
Figure S201. Get High-res Image Gene #42: 'del_9q31.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.00115 (Fisher's exact test), Q value = 0.0041
Table S202. Gene #42: 'del_9q31.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 15(9Q31.3) MUTATED | 25 | 29 | 31 | 24 | 9 |
| DEL PEAK 15(9Q31.3) WILD-TYPE | 42 | 57 | 40 | 47 | 60 |
Figure S202. Get High-res Image Gene #42: 'del_9q31.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00032 (Fisher's exact test), Q value = 0.0014
Table S203. Gene #42: 'del_9q31.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 15(9Q31.3) MUTATED | 21 | 16 | 38 | 37 | 6 |
| DEL PEAK 15(9Q31.3) WILD-TYPE | 33 | 40 | 76 | 47 | 50 |
Figure S203. Get High-res Image Gene #42: 'del_9q31.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.027 (Fisher's exact test), Q value = 0.051
Table S204. Gene #42: 'del_9q31.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 15(9Q31.3) MUTATED | 17 | 19 | 29 | 16 | 17 | 18 |
| DEL PEAK 15(9Q31.3) WILD-TYPE | 23 | 44 | 63 | 68 | 29 | 22 |
Figure S204. Get High-res Image Gene #42: 'del_9q31.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00696 (Fisher's exact test), Q value = 0.017
Table S205. Gene #42: 'del_9q31.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 15(9Q31.3) MUTATED | 17 | 30 | 21 | 43 | 5 |
| DEL PEAK 15(9Q31.3) WILD-TYPE | 27 | 65 | 35 | 79 | 43 |
Figure S205. Get High-res Image Gene #42: 'del_9q31.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.00022 (Fisher's exact test), Q value = 0.001
Table S206. Gene #42: 'del_9q31.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 15(9Q31.3) MUTATED | 29 | 25 | 36 | 16 | 5 |
| DEL PEAK 15(9Q31.3) WILD-TYPE | 41 | 57 | 64 | 18 | 49 |
Figure S206. Get High-res Image Gene #42: 'del_9q31.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S207. Gene #43: 'del_10q23.31' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 16(10Q23.31) MUTATED | 11 | 71 | 15 |
| DEL PEAK 16(10Q23.31) WILD-TYPE | 82 | 80 | 111 |
Figure S207. Get High-res Image Gene #43: 'del_10q23.31' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.015 (Fisher's exact test), Q value = 0.031
Table S208. Gene #43: 'del_10q23.31' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 16(10Q23.31) MUTATED | 32 | 45 | 20 |
| DEL PEAK 16(10Q23.31) WILD-TYPE | 61 | 116 | 96 |
Figure S208. Get High-res Image Gene #43: 'del_10q23.31' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S209. Gene #43: 'del_10q23.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 16(10Q23.31) MUTATED | 38 | 29 | 6 | 16 | 5 |
| DEL PEAK 16(10Q23.31) WILD-TYPE | 29 | 57 | 65 | 55 | 64 |
Figure S209. Get High-res Image Gene #43: 'del_10q23.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S210. Gene #43: 'del_10q23.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 16(10Q23.31) MUTATED | 27 | 9 | 41 | 11 | 6 |
| DEL PEAK 16(10Q23.31) WILD-TYPE | 27 | 47 | 73 | 73 | 50 |
Figure S210. Get High-res Image Gene #43: 'del_10q23.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S211. Gene #43: 'del_10q23.31' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 16(10Q23.31) MUTATED | 15 | 33 | 15 | 11 | 9 | 14 |
| DEL PEAK 16(10Q23.31) WILD-TYPE | 25 | 30 | 77 | 73 | 37 | 26 |
Figure S211. Get High-res Image Gene #43: 'del_10q23.31' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S212. Gene #43: 'del_10q23.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 16(10Q23.31) MUTATED | 15 | 41 | 13 | 25 | 3 |
| DEL PEAK 16(10Q23.31) WILD-TYPE | 29 | 54 | 43 | 97 | 45 |
Figure S212. Get High-res Image Gene #43: 'del_10q23.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.0156 (Fisher's exact test), Q value = 0.032
Table S213. Gene #43: 'del_10q23.31' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 16(10Q23.31) MUTATED | 42 | 19 | 21 | 12 |
| DEL PEAK 16(10Q23.31) WILD-TYPE | 65 | 62 | 81 | 38 |
Figure S213. Get High-res Image Gene #43: 'del_10q23.31' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S214. Gene #43: 'del_10q23.31' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 16(10Q23.31) MUTATED | 21 | 38 | 20 | 13 | 2 |
| DEL PEAK 16(10Q23.31) WILD-TYPE | 49 | 44 | 80 | 21 | 52 |
Figure S214. Get High-res Image Gene #43: 'del_10q23.31' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S215. Gene #44: 'del_10q26.13' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 17(10Q26.13) MUTATED | 15 | 75 | 15 |
| DEL PEAK 17(10Q26.13) WILD-TYPE | 78 | 76 | 111 |
Figure S215. Get High-res Image Gene #44: 'del_10q26.13' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0242 (Fisher's exact test), Q value = 0.047
Table S216. Gene #44: 'del_10q26.13' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 17(10Q26.13) MUTATED | 36 | 44 | 25 |
| DEL PEAK 17(10Q26.13) WILD-TYPE | 57 | 117 | 91 |
Figure S216. Get High-res Image Gene #44: 'del_10q26.13' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S217. Gene #44: 'del_10q26.13' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 17(10Q26.13) MUTATED | 37 | 32 | 10 | 18 | 5 |
| DEL PEAK 17(10Q26.13) WILD-TYPE | 30 | 54 | 61 | 53 | 64 |
Figure S217. Get High-res Image Gene #44: 'del_10q26.13' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S218. Gene #44: 'del_10q26.13' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 17(10Q26.13) MUTATED | 30 | 11 | 41 | 14 | 6 |
| DEL PEAK 17(10Q26.13) WILD-TYPE | 24 | 45 | 73 | 70 | 50 |
Figure S218. Get High-res Image Gene #44: 'del_10q26.13' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 3e-05 (Fisher's exact test), Q value = 2e-04
Table S219. Gene #44: 'del_10q26.13' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 17(10Q26.13) MUTATED | 13 | 32 | 19 | 13 | 11 | 17 |
| DEL PEAK 17(10Q26.13) WILD-TYPE | 27 | 31 | 73 | 71 | 35 | 23 |
Figure S219. Get High-res Image Gene #44: 'del_10q26.13' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S220. Gene #44: 'del_10q26.13' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 17(10Q26.13) MUTATED | 14 | 45 | 12 | 31 | 3 |
| DEL PEAK 17(10Q26.13) WILD-TYPE | 30 | 50 | 44 | 91 | 45 |
Figure S220. Get High-res Image Gene #44: 'del_10q26.13' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S221. Gene #44: 'del_10q26.13' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 17(10Q26.13) MUTATED | 22 | 40 | 25 | 13 | 2 |
| DEL PEAK 17(10Q26.13) WILD-TYPE | 48 | 42 | 75 | 21 | 52 |
Figure S221. Get High-res Image Gene #44: 'del_10q26.13' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.0161 (Fisher's exact test), Q value = 0.033
Table S222. Gene #45: 'del_11q14.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 18(11Q14.1) MUTATED | 10 | 37 | 30 |
| DEL PEAK 18(11Q14.1) WILD-TYPE | 83 | 114 | 96 |
Figure S222. Get High-res Image Gene #45: 'del_11q14.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00743 (Fisher's exact test), Q value = 0.018
Table S223. Gene #45: 'del_11q14.1' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 18(11Q14.1) MUTATED | 33 | 13 | 13 | 12 |
| DEL PEAK 18(11Q14.1) WILD-TYPE | 74 | 68 | 89 | 38 |
Figure S223. Get High-res Image Gene #45: 'del_11q14.1' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.0153 (Fisher's exact test), Q value = 0.032
Table S224. Gene #45: 'del_11q14.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 18(11Q14.1) MUTATED | 23 | 21 | 15 | 6 | 6 |
| DEL PEAK 18(11Q14.1) WILD-TYPE | 47 | 61 | 85 | 28 | 48 |
Figure S224. Get High-res Image Gene #45: 'del_11q14.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.00054 (Fisher's exact test), Q value = 0.0022
Table S225. Gene #46: 'del_11q23.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 19(11Q23.3) MUTATED | 10 | 45 | 38 |
| DEL PEAK 19(11Q23.3) WILD-TYPE | 83 | 106 | 88 |
Figure S225. Get High-res Image Gene #46: 'del_11q23.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.0406 (Fisher's exact test), Q value = 0.072
Table S226. Gene #46: 'del_11q23.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 19(11Q23.3) MUTATED | 16 | 8 | 31 | 28 | 9 |
| DEL PEAK 19(11Q23.3) WILD-TYPE | 38 | 48 | 83 | 56 | 47 |
Figure S226. Get High-res Image Gene #46: 'del_11q23.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.0138 (Fisher's exact test), Q value = 0.03
Table S227. Gene #46: 'del_11q23.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 19(11Q23.3) MUTATED | 39 | 17 | 18 | 12 |
| DEL PEAK 19(11Q23.3) WILD-TYPE | 68 | 64 | 84 | 38 |
Figure S227. Get High-res Image Gene #46: 'del_11q23.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.0021 (Fisher's exact test), Q value = 0.0065
Table S228. Gene #46: 'del_11q23.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 19(11Q23.3) MUTATED | 27 | 27 | 20 | 5 | 7 |
| DEL PEAK 19(11Q23.3) WILD-TYPE | 43 | 55 | 80 | 29 | 47 |
Figure S228. Get High-res Image Gene #46: 'del_11q23.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.00084 (Fisher's exact test), Q value = 0.0032
Table S229. Gene #47: 'del_12p12.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 20(12P12.1) MUTATED | 9 | 44 | 25 |
| DEL PEAK 20(12P12.1) WILD-TYPE | 84 | 107 | 101 |
Figure S229. Get High-res Image Gene #47: 'del_12p12.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00254 (Fisher's exact test), Q value = 0.0076
Table S230. Gene #47: 'del_12p12.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 20(12P12.1) MUTATED | 24 | 24 | 10 | 11 | 9 |
| DEL PEAK 20(12P12.1) WILD-TYPE | 43 | 62 | 61 | 60 | 60 |
Figure S230. Get High-res Image Gene #47: 'del_12p12.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.031 (Fisher's exact test), Q value = 0.058
Table S231. Gene #47: 'del_12p12.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 20(12P12.1) MUTATED | 14 | 27 | 8 | 21 | 6 |
| DEL PEAK 20(12P12.1) WILD-TYPE | 30 | 68 | 48 | 101 | 42 |
Figure S231. Get High-res Image Gene #47: 'del_12p12.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.0326 (Fisher's exact test), Q value = 0.06
Table S232. Gene #47: 'del_12p12.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 20(12P12.1) MUTATED | 11 | 26 | 16 | 11 | 10 |
| DEL PEAK 20(12P12.1) WILD-TYPE | 59 | 56 | 84 | 23 | 44 |
Figure S232. Get High-res Image Gene #47: 'del_12p12.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.00345 (Fisher's exact test), Q value = 0.0098
Table S233. Gene #48: 'del_12q24.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 21(12Q24.33) MUTATED | 7 | 35 | 17 |
| DEL PEAK 21(12Q24.33) WILD-TYPE | 86 | 116 | 109 |
Figure S233. Get High-res Image Gene #48: 'del_12q24.33' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S234. Gene #49: 'del_13q14.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 22(13Q14.2) MUTATED | 21 | 109 | 49 |
| DEL PEAK 22(13Q14.2) WILD-TYPE | 72 | 42 | 77 |
Figure S234. Get High-res Image Gene #49: 'del_13q14.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00028 (Fisher's exact test), Q value = 0.0012
Table S235. Gene #49: 'del_13q14.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 22(13Q14.2) MUTATED | 41 | 45 | 41 | 30 | 19 |
| DEL PEAK 22(13Q14.2) WILD-TYPE | 26 | 41 | 30 | 41 | 50 |
Figure S235. Get High-res Image Gene #49: 'del_13q14.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S236. Gene #49: 'del_13q14.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 22(13Q14.2) MUTATED | 34 | 15 | 67 | 46 | 14 |
| DEL PEAK 22(13Q14.2) WILD-TYPE | 20 | 41 | 47 | 38 | 42 |
Figure S236. Get High-res Image Gene #49: 'del_13q14.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00175 (Fisher's exact test), Q value = 0.0057
Table S237. Gene #49: 'del_13q14.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 22(13Q14.2) MUTATED | 26 | 36 | 45 | 26 | 20 | 24 |
| DEL PEAK 22(13Q14.2) WILD-TYPE | 14 | 27 | 47 | 58 | 26 | 16 |
Figure S237. Get High-res Image Gene #49: 'del_13q14.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00064 (Fisher's exact test), Q value = 0.0025
Table S238. Gene #49: 'del_13q14.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 22(13Q14.2) MUTATED | 28 | 53 | 26 | 59 | 11 |
| DEL PEAK 22(13Q14.2) WILD-TYPE | 16 | 42 | 30 | 63 | 37 |
Figure S238. Get High-res Image Gene #49: 'del_13q14.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.0179 (Fisher's exact test), Q value = 0.036
Table S239. Gene #49: 'del_13q14.2' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 22(13Q14.2) MUTATED | 63 | 42 | 38 | 25 |
| DEL PEAK 22(13Q14.2) WILD-TYPE | 44 | 39 | 64 | 25 |
Figure S239. Get High-res Image Gene #49: 'del_13q14.2' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 8e-05 (Fisher's exact test), Q value = 0.00042
Table S240. Gene #49: 'del_13q14.2' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 22(13Q14.2) MUTATED | 37 | 48 | 49 | 22 | 12 |
| DEL PEAK 22(13Q14.2) WILD-TYPE | 33 | 34 | 51 | 12 | 42 |
Figure S240. Get High-res Image Gene #49: 'del_13q14.2' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S241. Gene #50: 'del_14q23.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 23(14Q23.3) MUTATED | 9 | 83 | 39 |
| DEL PEAK 23(14Q23.3) WILD-TYPE | 84 | 68 | 87 |
Figure S241. Get High-res Image Gene #50: 'del_14q23.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00214 (Fisher's exact test), Q value = 0.0066
Table S242. Gene #50: 'del_14q23.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 23(14Q23.3) MUTATED | 36 | 23 | 29 | 24 | 17 |
| DEL PEAK 23(14Q23.3) WILD-TYPE | 31 | 63 | 42 | 47 | 52 |
Figure S242. Get High-res Image Gene #50: 'del_14q23.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 6e-05 (Fisher's exact test), Q value = 0.00034
Table S243. Gene #50: 'del_14q23.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 23(14Q23.3) MUTATED | 34 | 12 | 38 | 32 | 13 |
| DEL PEAK 23(14Q23.3) WILD-TYPE | 20 | 44 | 76 | 52 | 43 |
Figure S243. Get High-res Image Gene #50: 'del_14q23.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.012 (Fisher's exact test), Q value = 0.026
Table S244. Gene #50: 'del_14q23.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 23(14Q23.3) MUTATED | 21 | 26 | 26 | 21 | 17 | 19 |
| DEL PEAK 23(14Q23.3) WILD-TYPE | 19 | 37 | 66 | 63 | 29 | 21 |
Figure S244. Get High-res Image Gene #50: 'del_14q23.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00416 (Fisher's exact test), Q value = 0.011
Table S245. Gene #50: 'del_14q23.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 23(14Q23.3) MUTATED | 52 | 29 | 25 | 19 |
| DEL PEAK 23(14Q23.3) WILD-TYPE | 55 | 52 | 77 | 31 |
Figure S245. Get High-res Image Gene #50: 'del_14q23.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S246. Gene #51: 'del_14q32.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 24(14Q32.33) MUTATED | 10 | 83 | 40 |
| DEL PEAK 24(14Q32.33) WILD-TYPE | 83 | 68 | 86 |
Figure S246. Get High-res Image Gene #51: 'del_14q32.33' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00474 (Fisher's exact test), Q value = 0.013
Table S247. Gene #51: 'del_14q32.33' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 24(14Q32.33) MUTATED | 35 | 24 | 30 | 25 | 17 |
| DEL PEAK 24(14Q32.33) WILD-TYPE | 32 | 62 | 41 | 46 | 52 |
Figure S247. Get High-res Image Gene #51: 'del_14q32.33' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00012 (Fisher's exact test), Q value = 0.00058
Table S248. Gene #51: 'del_14q32.33' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 24(14Q32.33) MUTATED | 33 | 12 | 37 | 35 | 14 |
| DEL PEAK 24(14Q32.33) WILD-TYPE | 21 | 44 | 77 | 49 | 42 |
Figure S248. Get High-res Image Gene #51: 'del_14q32.33' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.0125 (Fisher's exact test), Q value = 0.027
Table S249. Gene #51: 'del_14q32.33' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 24(14Q32.33) MUTATED | 48 | 32 | 25 | 21 |
| DEL PEAK 24(14Q32.33) WILD-TYPE | 59 | 49 | 77 | 29 |
Figure S249. Get High-res Image Gene #51: 'del_14q32.33' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.015 (Fisher's exact test), Q value = 0.031
Table S250. Gene #51: 'del_14q32.33' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 24(14Q32.33) MUTATED | 32 | 23 | 41 | 17 | 13 |
| DEL PEAK 24(14Q32.33) WILD-TYPE | 38 | 59 | 59 | 17 | 41 |
Figure S250. Get High-res Image Gene #51: 'del_14q32.33' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.00052 (Fisher's exact test), Q value = 0.0021
Table S251. Gene #52: 'del_15q26.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 25(15Q26.3) MUTATED | 11 | 47 | 21 |
| DEL PEAK 25(15Q26.3) WILD-TYPE | 82 | 104 | 105 |
Figure S251. Get High-res Image Gene #52: 'del_15q26.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00834 (Fisher's exact test), Q value = 0.02
Table S252. Gene #52: 'del_15q26.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 25(15Q26.3) MUTATED | 25 | 17 | 16 | 11 | 9 |
| DEL PEAK 25(15Q26.3) WILD-TYPE | 42 | 69 | 55 | 60 | 60 |
Figure S252. Get High-res Image Gene #52: 'del_15q26.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00971 (Fisher's exact test), Q value = 0.022
Table S253. Gene #52: 'del_15q26.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 25(15Q26.3) MUTATED | 21 | 6 | 22 | 19 | 10 |
| DEL PEAK 25(15Q26.3) WILD-TYPE | 33 | 50 | 92 | 65 | 46 |
Figure S253. Get High-res Image Gene #52: 'del_15q26.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00225 (Fisher's exact test), Q value = 0.0068
Table S254. Gene #52: 'del_15q26.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 25(15Q26.3) MUTATED | 32 | 11 | 15 | 17 |
| DEL PEAK 25(15Q26.3) WILD-TYPE | 75 | 70 | 87 | 33 |
Figure S254. Get High-res Image Gene #52: 'del_15q26.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S255. Gene #53: 'del_16q23.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 26(16Q23.1) MUTATED | 8 | 86 | 68 |
| DEL PEAK 26(16Q23.1) WILD-TYPE | 85 | 65 | 58 |
Figure S255. Get High-res Image Gene #53: 'del_16q23.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S256. Gene #53: 'del_16q23.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 26(16Q23.1) MUTATED | 30 | 56 | 76 |
| DEL PEAK 26(16Q23.1) WILD-TYPE | 63 | 105 | 40 |
Figure S256. Get High-res Image Gene #53: 'del_16q23.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S257. Gene #53: 'del_16q23.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 26(16Q23.1) MUTATED | 36 | 34 | 60 | 20 | 10 |
| DEL PEAK 26(16Q23.1) WILD-TYPE | 31 | 52 | 11 | 51 | 59 |
Figure S257. Get High-res Image Gene #53: 'del_16q23.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S258. Gene #53: 'del_16q23.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 26(16Q23.1) MUTATED | 25 | 18 | 52 | 57 | 8 |
| DEL PEAK 26(16Q23.1) WILD-TYPE | 29 | 38 | 62 | 27 | 48 |
Figure S258. Get High-res Image Gene #53: 'del_16q23.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 5e-05 (Fisher's exact test), Q value = 3e-04
Table S259. Gene #53: 'del_16q23.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 26(16Q23.1) MUTATED | 19 | 28 | 32 | 24 | 32 | 25 |
| DEL PEAK 26(16Q23.1) WILD-TYPE | 21 | 35 | 60 | 60 | 14 | 15 |
Figure S259. Get High-res Image Gene #53: 'del_16q23.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S260. Gene #53: 'del_16q23.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 26(16Q23.1) MUTATED | 19 | 40 | 33 | 62 | 6 |
| DEL PEAK 26(16Q23.1) WILD-TYPE | 25 | 55 | 23 | 60 | 42 |
Figure S260. Get High-res Image Gene #53: 'del_16q23.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S261. Gene #53: 'del_16q23.1' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 26(16Q23.1) MUTATED | 61 | 33 | 28 | 32 |
| DEL PEAK 26(16Q23.1) WILD-TYPE | 46 | 48 | 74 | 18 |
Figure S261. Get High-res Image Gene #53: 'del_16q23.1' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 5e-05 (Fisher's exact test), Q value = 3e-04
Table S262. Gene #53: 'del_16q23.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 26(16Q23.1) MUTATED | 44 | 36 | 50 | 14 | 10 |
| DEL PEAK 26(16Q23.1) WILD-TYPE | 26 | 46 | 50 | 20 | 44 |
Figure S262. Get High-res Image Gene #53: 'del_16q23.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S263. Gene #54: 'del_17p13.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 27(17P13.1) MUTATED | 37 | 137 | 52 |
| DEL PEAK 27(17P13.1) WILD-TYPE | 56 | 14 | 74 |
Figure S263. Get High-res Image Gene #54: 'del_17p13.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 2e-05 (Fisher's exact test), Q value = 0.00014
Table S264. Gene #54: 'del_17p13.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 27(17P13.1) MUTATED | 53 | 63 | 37 | 35 | 33 |
| DEL PEAK 27(17P13.1) WILD-TYPE | 14 | 23 | 34 | 36 | 36 |
Figure S264. Get High-res Image Gene #54: 'del_17p13.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 6e-05 (Fisher's exact test), Q value = 0.00034
Table S265. Gene #54: 'del_17p13.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 27(17P13.1) MUTATED | 40 | 28 | 85 | 42 | 26 |
| DEL PEAK 27(17P13.1) WILD-TYPE | 14 | 28 | 29 | 42 | 30 |
Figure S265. Get High-res Image Gene #54: 'del_17p13.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S266. Gene #54: 'del_17p13.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 27(17P13.1) MUTATED | 35 | 49 | 48 | 40 | 27 | 25 |
| DEL PEAK 27(17P13.1) WILD-TYPE | 5 | 14 | 44 | 44 | 19 | 15 |
Figure S266. Get High-res Image Gene #54: 'del_17p13.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 8e-05 (Fisher's exact test), Q value = 0.00042
Table S267. Gene #54: 'del_17p13.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 27(17P13.1) MUTATED | 38 | 67 | 30 | 66 | 23 |
| DEL PEAK 27(17P13.1) WILD-TYPE | 6 | 28 | 26 | 56 | 25 |
Figure S267. Get High-res Image Gene #54: 'del_17p13.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S268. Gene #54: 'del_17p13.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 27(17P13.1) MUTATED | 39 | 65 | 55 | 30 | 25 |
| DEL PEAK 27(17P13.1) WILD-TYPE | 31 | 17 | 45 | 4 | 29 |
Figure S268. Get High-res Image Gene #54: 'del_17p13.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S269. Gene #55: 'del_17p11.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 28(17P11.2) MUTATED | 32 | 119 | 44 |
| DEL PEAK 28(17P11.2) WILD-TYPE | 61 | 32 | 82 |
Figure S269. Get High-res Image Gene #55: 'del_17p11.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00016 (Fisher's exact test), Q value = 0.00077
Table S270. Gene #55: 'del_17p11.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 28(17P11.2) MUTATED | 49 | 52 | 32 | 32 | 27 |
| DEL PEAK 28(17P11.2) WILD-TYPE | 18 | 34 | 39 | 39 | 42 |
Figure S270. Get High-res Image Gene #55: 'del_17p11.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00839 (Fisher's exact test), Q value = 0.02
Table S271. Gene #55: 'del_17p11.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 28(17P11.2) MUTATED | 37 | 27 | 68 | 37 | 23 |
| DEL PEAK 28(17P11.2) WILD-TYPE | 17 | 29 | 46 | 47 | 33 |
Figure S271. Get High-res Image Gene #55: 'del_17p11.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 9e-04 (Fisher's exact test), Q value = 0.0034
Table S272. Gene #55: 'del_17p11.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 40 | 63 | 92 | 84 | 46 | 40 |
| DEL PEAK 28(17P11.2) MUTATED | 32 | 39 | 41 | 36 | 23 | 22 |
| DEL PEAK 28(17P11.2) WILD-TYPE | 8 | 24 | 51 | 48 | 23 | 18 |
Figure S272. Get High-res Image Gene #55: 'del_17p11.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00024 (Fisher's exact test), Q value = 0.0011
Table S273. Gene #55: 'del_17p11.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 44 | 95 | 56 | 122 | 48 |
| DEL PEAK 28(17P11.2) MUTATED | 36 | 54 | 25 | 56 | 22 |
| DEL PEAK 28(17P11.2) WILD-TYPE | 8 | 41 | 31 | 66 | 26 |
Figure S273. Get High-res Image Gene #55: 'del_17p11.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.00039 (Fisher's exact test), Q value = 0.0016
Table S274. Gene #55: 'del_17p11.2' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 28(17P11.2) MUTATED | 34 | 52 | 48 | 28 | 22 |
| DEL PEAK 28(17P11.2) WILD-TYPE | 36 | 30 | 52 | 6 | 32 |
Figure S274. Get High-res Image Gene #55: 'del_17p11.2' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.0241 (Fisher's exact test), Q value = 0.047
Table S275. Gene #56: 'del_18q21.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 29(18Q21.2) MUTATED | 14 | 45 | 35 |
| DEL PEAK 29(18Q21.2) WILD-TYPE | 79 | 106 | 91 |
Figure S275. Get High-res Image Gene #56: 'del_18q21.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 9e-05 (Fisher's exact test), Q value = 0.00045
Table S276. Gene #56: 'del_18q21.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 29(18Q21.2) MUTATED | 14 | 34 | 46 |
| DEL PEAK 29(18Q21.2) WILD-TYPE | 79 | 127 | 70 |
Figure S276. Get High-res Image Gene #56: 'del_18q21.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 2e-05 (Fisher's exact test), Q value = 0.00014
Table S277. Gene #56: 'del_18q21.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 29(18Q21.2) MUTATED | 9 | 5 | 28 | 39 | 13 |
| DEL PEAK 29(18Q21.2) WILD-TYPE | 45 | 51 | 86 | 45 | 43 |
Figure S277. Get High-res Image Gene #56: 'del_18q21.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 7.4e-05
Table S278. Gene #57: 'del_19p13.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 30(19P13.3) MUTATED | 6 | 61 | 20 |
| DEL PEAK 30(19P13.3) WILD-TYPE | 87 | 90 | 106 |
Figure S278. Get High-res Image Gene #57: 'del_19p13.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00498 (Fisher's exact test), Q value = 0.013
Table S279. Gene #57: 'del_19p13.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 30(19P13.3) MUTATED | 28 | 17 | 17 | 12 | 12 |
| DEL PEAK 30(19P13.3) WILD-TYPE | 39 | 69 | 54 | 59 | 57 |
Figure S279. Get High-res Image Gene #57: 'del_19p13.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.0026 (Fisher's exact test), Q value = 0.0077
Table S280. Gene #57: 'del_19p13.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 30(19P13.3) MUTATED | 22 | 9 | 23 | 25 | 7 |
| DEL PEAK 30(19P13.3) WILD-TYPE | 32 | 47 | 91 | 59 | 49 |
Figure S280. Get High-res Image Gene #57: 'del_19p13.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00217 (Fisher's exact test), Q value = 0.0066
Table S281. Gene #57: 'del_19p13.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 107 | 81 | 102 | 50 |
| DEL PEAK 30(19P13.3) MUTATED | 38 | 15 | 15 | 15 |
| DEL PEAK 30(19P13.3) WILD-TYPE | 69 | 66 | 87 | 35 |
Figure S281. Get High-res Image Gene #57: 'del_19p13.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'

P value = 0.00099 (Fisher's exact test), Q value = 0.0036
Table S282. Gene #57: 'del_19p13.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 70 | 82 | 100 | 34 | 54 |
| DEL PEAK 30(19P13.3) MUTATED | 26 | 21 | 17 | 13 | 6 |
| DEL PEAK 30(19P13.3) WILD-TYPE | 44 | 61 | 83 | 21 | 48 |
Figure S282. Get High-res Image Gene #57: 'del_19p13.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'

P value = 0.00622 (Fisher's exact test), Q value = 0.016
Table S283. Gene #59: 'del_22q12.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 32(22Q12.1) MUTATED | 18 | 44 | 17 |
| DEL PEAK 32(22Q12.1) WILD-TYPE | 75 | 107 | 109 |
Figure S283. Get High-res Image Gene #59: 'del_22q12.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00288 (Fisher's exact test), Q value = 0.0085
Table S284. Gene #59: 'del_22q12.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 32(22Q12.1) MUTATED | 26 | 19 | 9 | 15 | 10 |
| DEL PEAK 32(22Q12.1) WILD-TYPE | 41 | 67 | 62 | 56 | 59 |
Figure S284. Get High-res Image Gene #59: 'del_22q12.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00056 (Fisher's exact test), Q value = 0.0022
Table S285. Gene #60: 'del_22q13.32' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 33(22Q13.32) MUTATED | 19 | 48 | 16 |
| DEL PEAK 33(22Q13.32) WILD-TYPE | 74 | 103 | 110 |
Figure S285. Get High-res Image Gene #60: 'del_22q13.32' versus Molecular Subtype #1: 'CN_CNMF'

P value = 9e-05 (Fisher's exact test), Q value = 0.00045
Table S286. Gene #60: 'del_22q13.32' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 67 | 86 | 71 | 71 | 69 |
| DEL PEAK 33(22Q13.32) MUTATED | 28 | 24 | 7 | 14 | 10 |
| DEL PEAK 33(22Q13.32) WILD-TYPE | 39 | 62 | 64 | 57 | 59 |
Figure S286. Get High-res Image Gene #60: 'del_22q13.32' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 0.00992 (Fisher's exact test), Q value = 0.023
Table S287. Gene #60: 'del_22q13.32' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 54 | 56 | 114 | 84 | 56 |
| DEL PEAK 33(22Q13.32) MUTATED | 16 | 6 | 36 | 16 | 9 |
| DEL PEAK 33(22Q13.32) WILD-TYPE | 38 | 50 | 78 | 68 | 47 |
Figure S287. Get High-res Image Gene #60: 'del_22q13.32' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 8e-05 (Fisher's exact test), Q value = 0.00042
Table S288. Gene #61: 'del_xq26.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 151 | 126 |
| DEL PEAK 34(XQ26.2) MUTATED | 6 | 42 | 19 |
| DEL PEAK 34(XQ26.2) WILD-TYPE | 87 | 109 | 107 |
Figure S288. Get High-res Image Gene #61: 'del_xq26.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00695 (Fisher's exact test), Q value = 0.017
Table S289. Gene #61: 'del_xq26.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 93 | 161 | 116 |
| DEL PEAK 34(XQ26.2) MUTATED | 8 | 30 | 29 |
| DEL PEAK 34(XQ26.2) WILD-TYPE | 85 | 131 | 87 |
Figure S289. Get High-res Image Gene #61: 'del_xq26.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'

-
Copy number data file = all_lesions.txt from GISTIC pipeline
-
Processed Copy number data file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_Correlate_Genomic_Events_Preprocess/LIHC-TP/22531241/transformed.cor.cli.txt
-
Molecular subtype file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_mergedClustering/LIHC-TP/22542932/LIHC-TP.transferedmergedcluster.txt
-
Number of patients = 370
-
Number of significantly focal cnvs = 61
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.