This report serves to describe the mutational landscape and properties of a given individual set, as well as rank genes and genesets according to mutational significance. MutSig v2.0 was used to generate the results found in this report.
Working with individual set: OV.
Number of patients in set: 316
The input for this pipeline is a set of individuals with the following files associated for each:
1. An annotated .maf file describing the mutations called for the respective individual, and their properties.
2. A .wig file that contains information about the coverage of the sample.
MAF used for this analysis: OV.final_analysis_set.maf
Significantly mutated genes (q ≤ 0.1): 12
Mutations seen in COSMIC: 394
Significantly mutated genes in COSMIC territory: 39
Genes with clustered mutations (≤ 3 aa apart): 0
Significantly mutated genesets: 61
Significantly mutated genesets: (excluding sig. mutated genes): 13
Read 148 MAFs of type "Broad"
Read 88 MAFs of type "WashU"
Read 80 MAFs of type "Baylor"
Total number of mutations in input MAFs: 20219
Number of mutations before filtering: 20219
After removing 1 non-mutations: 20218
After removing 778 mutations outside gene set: 19440
After removing 9 mutations outside category set: 19431
After removing 4 "impossible" mutations in
gene-patient-category bins of zero coverage: 19427
Table 1. Get Full Table Table representing breakdown of mutations by type.
| type | count |
|---|---|
| Frame_Shift_Del | 441 |
| Frame_Shift_Ins | 155 |
| In_Frame_Del | 148 |
| In_Frame_Ins | 37 |
| Indel | 13 |
| Missense_Mutation | 13164 |
| Nonsense_Mutation | 800 |
| Nonstop_Mutation | 16 |
| Silent | 4231 |
| Splice_Site_Del | 33 |
| Splice_Site_Ins | 5 |
| Splice_Site_SNP | 388 |
| Total | 19431 |
Table 2. Get Full Table A breakdown of mutation rates per category discovered for this individual set.
| category | n | N | rate | rate_per_mb | relative_rate |
|---|---|---|---|---|---|
| *CpG->T | 1860 | 396729715 | 4.7e-06 | 4.7 | 2.5 |
| *Cp(A/C/T)->T | 2187 | 3661505903 | 6e-07 | 0.6 | 0.32 |
| C->(G/A) | 5212 | 4058235618 | 1.3e-06 | 1.3 | 0.69 |
| A->mut | 3901 | 4115893953 | 9.5e-07 | 0.95 | 0.51 |
| indel+null | 2030 | 8174129610 | 2.5e-07 | 0.25 | 0.13 |
| double_null | 7 | 8174129610 | 8.6e-10 | 0.00086 | 0.00046 |
| Total | 15197 | 8174129610 | 1.9e-06 | 1.9 | 1 |
The x axis represents the samples. The y axis represents the exons, one row per exon, and they are sorted by average coverage across samples. For exons with exactly the same average coverage, they are sorted next by the %GC of the exon. (The secondary sort is especially useful for the zero-coverage exons at the bottom).
Figure 1.
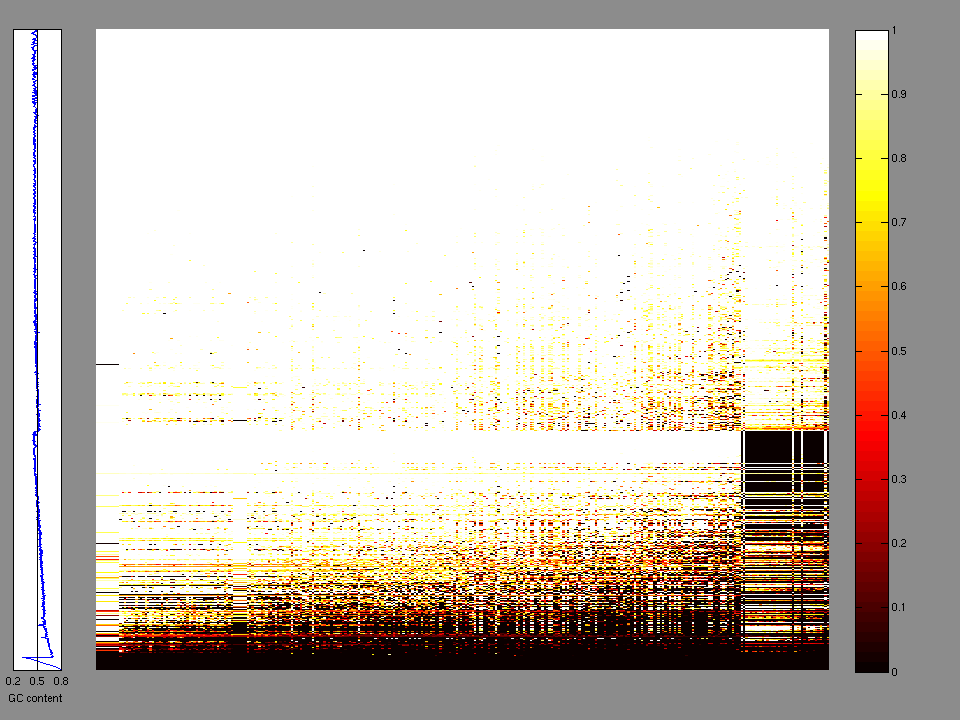

Figure 2. Patients counts and rates file used to generate this plot: OV.patients.counts_and_rates.txt
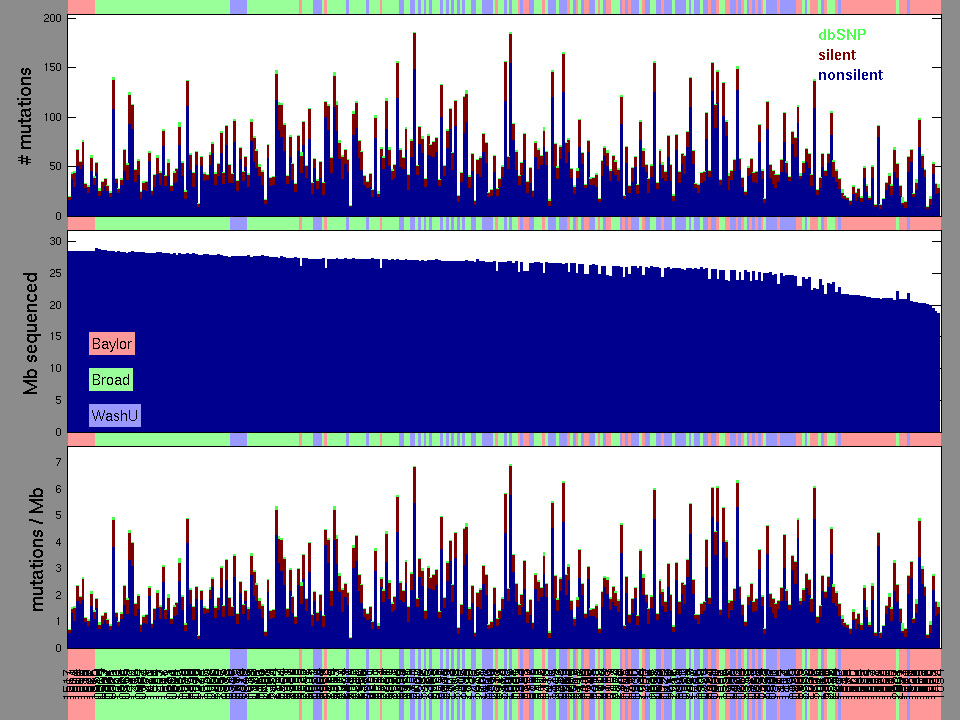
Figure 3. Get High-res Image The matrix in the center of the figure represents individual mutations in patient samples, color-coded by type of mutation, for the significantly mutated genes. The rate of synonymous and non-synonymous mutations is displayed at the top of the matrix. The barplot on the left of the matrix shows the number of mutations in each gene. The percentages represent the fraction of tumors with at least one mutation in the specified gene. The barplot to the right of the matrix displays the q-values for the most significantly mutated genes. The purple boxplots below the matrix (only displayed if required columns are present in the provided MAF) represent the distributions of allelic fractions observed in each sample. The plot at the bottom represents the base substitution distribution of individual samples, using the same categories that were used to calculate significance.
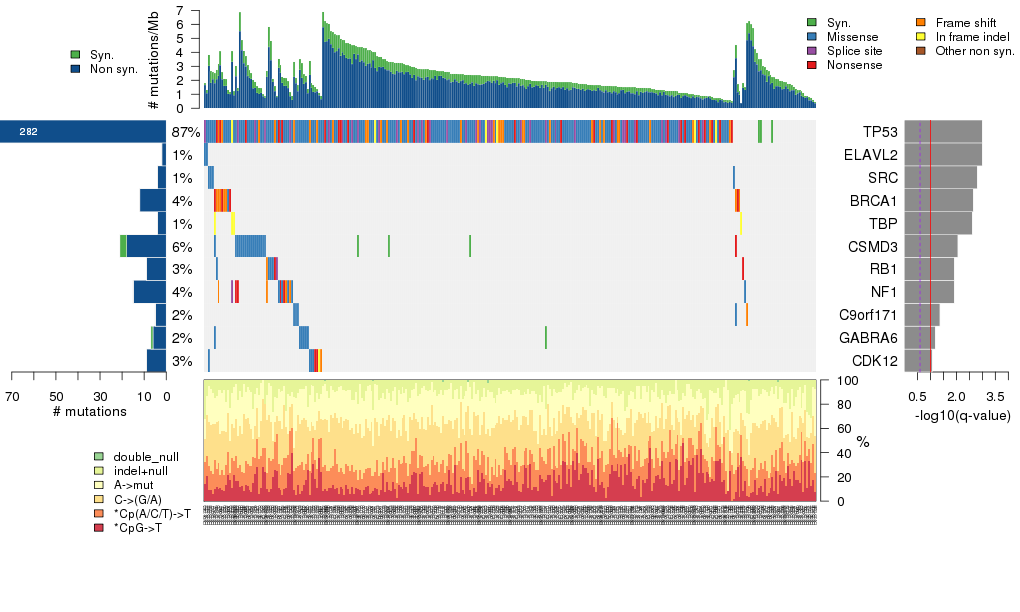
Table 3. Get Full Table A Ranked List of Significantly Mutated Genes. Number of significant genes found: 12. Number of genes displayed: 35. Click on a gene name to display its stick figure depicting the distribution of mutations and mutation types across the chosen gene (this feature may not be available for all significant genes).
| rank | gene | description | N | n | npat | nsite | nsil | n1 | n2 | n3 | n4 | n5 | n6 | p_classic | p_ks | p_cons | p_joint | p | q |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | TP53 | tumor protein p53 | 401524 | 279 | 276 | 143 | 3 | 47 | 31 | 39 | 61 | 101 | 0 | <1.00e-15 | 2e-07 | 2e-07 | 0 | <1.00e-15 | <8.54e-12 |
| 2 | ELAVL2 | ELAV (embryonic lethal, abnormal vision, Drosophila)-like 2 (Hu antigen B) | 347710 | 2 | 2 | 2 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0.041 | 0.048 | 0.0044 | 0 | <1.00e-15 | <8.54e-12 |
| 3 | SRC | v-src sarcoma (Schmidt-Ruppin A-2) viral oncogene homolog (avian) | 372331 | 4 | 4 | 2 | 0 | 0 | 0 | 4 | 0 | 0 | 0 | 0.0026 | 1.6e-06 | 0.017 | 1.8e-06 | 9.54e-08 | 0.00054 |
| 4 | BRCA1 | breast cancer 1, early onset | 1798045 | 12 | 12 | 12 | 0 | 0 | 0 | 1 | 0 | 11 | 0 | 1.87e-08 | 1 | 0.2 | 0.77 | 2.73e-07 | 0.0012 |
| 5 | TBP | TATA box binding protein | 321652 | 4 | 4 | 2 | 0 | 0 | 0 | 0 | 0 | 4 | 0 | 0.00023 | 0.000053 | 0.99 | 0.000096 | 4.07e-07 | 0.0014 |
| 6 | CSMD3 | CUB and Sushi multiple domains 3 | 3615081 | 18 | 18 | 18 | 3 | 0 | 2 | 7 | 8 | 1 | 0 | 1.78e-06 | 0.051 | 0.48 | 0.092 | 2.72e-06 | 0.0078 |
| 7 | RB1 | retinoblastoma 1 (including osteosarcoma) | 826059 | 9 | 9 | 9 | 0 | 0 | 1 | 3 | 0 | 5 | 0 | 7.71e-07 | 0.23 | 0.65 | 0.39 | 4.84e-06 | 0.011 |
| 8 | NF1 | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 2619095 | 15 | 14 | 15 | 0 | 1 | 1 | 1 | 3 | 9 | 0 | 1.67e-06 | 0.3 | 0.09 | 0.19 | 5.13e-06 | 0.011 |
| 9 | C9orf171 | 251675 | 5 | 5 | 5 | 0 | 2 | 0 | 2 | 0 | 1 | 0 | 0.000039 | 0.33 | 0.014 | 0.039 | 0.000022 | 0.042 | |
| 10 | GABRA6 | gamma-aminobutyric acid (GABA) A receptor, alpha 6 | 440417 | 6 | 6 | 6 | 1 | 1 | 3 | 1 | 1 | 0 | 0 | 0.000045 | 0.035 | 0.32 | 0.059 | 0.000037 | 0.063 |
| 11 | CDK12 | 1351667 | 9 | 9 | 9 | 0 | 0 | 0 | 1 | 3 | 5 | 0 | 0.000057 | 0.047 | 0.25 | 0.071 | 0.000054 | 0.084 | |
| 12 | FAT3 | FAT tumor suppressor homolog 3 (Drosophila) | 3706784 | 20 | 19 | 20 | 1 | 4 | 2 | 3 | 10 | 1 | 0 | 0.000012 | 0.74 | 0.28 | 0.62 | 0.000094 | 0.12 |
| 13 | ARHGEF9 | Cdc42 guanine nucleotide exchange factor (GEF) 9 | 494011 | 5 | 5 | 5 | 0 | 0 | 1 | 2 | 1 | 1 | 0 | 0.00041 | 0.015 | 0.33 | 0.018 | 0.000095 | 0.12 |
| 14 | EFEMP1 | EGF-containing fibulin-like extracellular matrix protein 1 | 475456 | 5 | 5 | 5 | 0 | 1 | 0 | 3 | 0 | 1 | 0 | 0.00036 | 0.25 | 0.012 | 0.028 | 0.00012 | 0.15 |
| 15 | CYP11B1 | cytochrome P450, family 11, subfamily B, polypeptide 1 | 456589 | 7 | 7 | 6 | 0 | 1 | 0 | 3 | 3 | 0 | 0 | 0.000019 | 0.45 | 0.8 | 0.65 | 0.00015 | 0.17 |
| 16 | LAIR1 | leukocyte-associated immunoglobulin-like receptor 1 | 267149 | 3 | 3 | 3 | 1 | 0 | 0 | 1 | 1 | 1 | 0 | 0.0033 | 0.036 | 0.015 | 0.0042 | 0.00017 | 0.17 |
| 17 | SLCO1C1 | solute carrier organic anion transporter family, member 1C1 | 717910 | 6 | 6 | 6 | 0 | 0 | 1 | 4 | 1 | 0 | 0 | 0.00013 | 0.047 | 0.72 | 0.1 | 0.00017 | 0.17 |
| 18 | CREBBP | CREB binding protein (Rubinstein-Taybi syndrome) | 1964764 | 7 | 7 | 7 | 1 | 0 | 1 | 0 | 3 | 3 | 0 | 0.0055 | 0.0044 | 0.071 | 0.0029 | 0.00019 | 0.18 |
| 19 | GAS2L1 | growth arrest-specific 2 like 1 | 197396 | 4 | 4 | 4 | 0 | 0 | 0 | 3 | 1 | 0 | 0 | 0.00017 | 0.065 | 0.4 | 0.1 | 0.00021 | 0.19 |
| 20 | DUSP19 | dual specificity phosphatase 19 | 211393 | 4 | 4 | 4 | 0 | 0 | 0 | 1 | 3 | 0 | 0 | 0.000040 | 0.63 | 0.47 | 0.7 | 0.00032 | 0.22 |
| 21 | CHST2 | carbohydrate (N-acetylglucosamine-6-O) sulfotransferase 2 | 172127 | 5 | 5 | 5 | 1 | 0 | 2 | 2 | 1 | 0 | 0 | 0.000028 | 1 | 0.97 | 1 | 0.00033 | 0.22 |
| 22 | USH2A | Usher syndrome 2A (autosomal recessive, mild) | 5014417 | 20 | 20 | 20 | 3 | 2 | 5 | 3 | 7 | 3 | 0 | 0.000074 | 0.42 | 0.2 | 0.39 | 0.00033 | 0.22 |
| 23 | BRCA2 | breast cancer 2, early onset | 2896444 | 11 | 11 | 11 | 0 | 1 | 0 | 0 | 1 | 8 | 1 | 0.000070 | 0.27 | 0.93 | 0.41 | 0.00033 | 0.22 |
| 24 | PPP1R3A | protein phosphatase 1, regulatory (inhibitor) subunit 3A | 1053877 | 8 | 8 | 8 | 2 | 1 | 1 | 2 | 4 | 0 | 0 | 0.000060 | 0.92 | 0.093 | 0.49 | 0.00034 | 0.22 |
| 25 | SLC35F5 | solute carrier family 35, member F5 | 489083 | 2 | 2 | 1 | 0 | 0 | 0 | 0 | 0 | 2 | 0 | 0.050 | 0.00062 | 0.098 | 0.00062 | 0.00036 | 0.22 |
| 26 | ATP5B | ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide | 467672 | 2 | 2 | 1 | 1 | 0 | 0 | 0 | 2 | 0 | 0 | 0.072 | 0.0018 | 0.22 | 0.00045 | 0.00037 | 0.22 |
| 27 | FAM171B | 744587 | 7 | 7 | 7 | 0 | 1 | 1 | 2 | 2 | 1 | 0 | 0.000084 | 0.22 | 0.64 | 0.4 | 0.00038 | 0.22 | |
| 28 | C6orf142 | chromosome 6 open reading frame 142 | 449405 | 5 | 5 | 5 | 3 | 0 | 2 | 1 | 1 | 1 | 0 | 0.00013 | 0.16 | 0.7 | 0.25 | 0.00038 | 0.22 |
| 29 | PDAP1 | PDGFA associated protein 1 | 170027 | 2 | 2 | 1 | 0 | 0 | 0 | 0 | 0 | 2 | 0 | 0.0036 | 0.0017 | 0.89 | 0.0095 | 0.00038 | 0.22 |
| 30 | MYO19 | myosin XIX | 675465 | 2 | 1 | 2 | 1 | 0 | 0 | 1 | 0 | 1 | 0 | 0.44 | 0.00019 | 0.19 | 0.000083 | 0.00041 | 0.22 |
| 31 | GLI2 | GLI-Kruppel family member GLI2 | 947026 | 10 | 9 | 10 | 0 | 2 | 3 | 3 | 1 | 1 | 0 | 0.000078 | 0.6 | 0.25 | 0.53 | 0.00046 | 0.22 |
| 32 | ODAM | odontogenic, ameloblast asssociated | 257049 | 3 | 3 | 3 | 0 | 0 | 0 | 1 | 2 | 0 | 0 | 0.0014 | 0.49 | 0.0057 | 0.03 | 0.00047 | 0.22 |
| 33 | COL5A3 | collagen, type V, alpha 3 | 1408232 | 7 | 7 | 7 | 1 | 0 | 3 | 3 | 0 | 1 | 0 | 0.0092 | 0.037 | 0.0041 | 0.0047 | 0.00047 | 0.22 |
| 34 | LCOR | ligand dependent nuclear receptor corepressor | 396634 | 4 | 4 | 4 | 0 | 0 | 1 | 1 | 0 | 2 | 0 | 0.0010 | 0.5 | 0.012 | 0.043 | 0.00048 | 0.22 |
| 35 | KCNJ12 | potassium inwardly-rectifying channel, subfamily J, member 12 | 281754 | 5 | 5 | 3 | 2 | 5 | 0 | 0 | 0 | 0 | 0 | 0.00022 | 0.14 | 0.21 | 0.2 | 0.00048 | 0.22 |
Note:
N - number of sequenced bases in this gene across the individual set.
n - number of (nonsilent) mutations in this gene across the individual set.
npat - number of patients (individuals) with at least one nonsilent mutation.
nsite - number of unique sites having a non-silent mutation.
nsil - number of silent mutations in this gene across the individual set.
n1 - number of nonsilent mutations of type: *CpG->T .
n2 - number of nonsilent mutations of type: *Cp(A/C/T)->T .
n3 - number of nonsilent mutations of type: C->(G/A) .
n4 - number of nonsilent mutations of type: A->mut .
n5 - number of nonsilent mutations of type: indel+null .
null - mutation category that includes nonsense, frameshift, splice-site mutations
p_classic = p-value for the observed amount of nonsilent mutations being elevated in this gene
p_ns_s = p-value for the observed nonsilent/silent ratio being elevated in this gene
p = p-value (overall)
q = q-value, False Discovery Rate (Benjamini-Hochberg procedure)
In this analysis, COSMIC is used as a filter to increase power by restricting the territory of each gene. Cosmic version: v48.
Table 4. Get Full Table Significantly mutated genes (COSMIC territory only). To access the database please go to: COSMIC. Number of significant genes found: 39. Number of genes displayed: 10
| rank | gene | description | n | cos | n_cos | N_cos | cos_ev | p | q |
|---|---|---|---|---|---|---|---|---|---|
| 1 | TP53 | tumor protein p53 | 279 | 823 | 278 | 260068 | 70907 | 0 | 0 |
| 2 | RB1 | retinoblastoma 1 (including osteosarcoma) | 9 | 267 | 8 | 84372 | 16 | 9.6e-12 | 2.1e-08 |
| 3 | NF1 | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 15 | 285 | 6 | 90060 | 8 | 2.7e-08 | 0.000039 |
| 4 | MYO3A | myosin IIIA | 7 | 14 | 3 | 4424 | 3 | 9.2e-08 | 0.0001 |
| 5 | KLK10 | kallikrein-related peptidase 10 | 2 | 2 | 2 | 632 | 2 | 6.9e-07 | 0.0006 |
| 6 | CDC27 | cell division cycle 27 homolog (S. cerevisiae) | 4 | 3 | 2 | 948 | 2 | 1.5e-06 | 0.0011 |
| 7 | ERBB2 | v-erb-b2 erythroblastic leukemia viral oncogene homolog 2, neuro/glioblastoma derived oncogene homolog (avian) | 4 | 41 | 3 | 12956 | 4 | 2.3e-06 | 0.0014 |
| 8 | MLL4 | 4 | 6 | 2 | 1896 | 2 | 6.2e-06 | 0.0034 | |
| 9 | NIPBL | Nipped-B homolog (Drosophila) | 5 | 7 | 2 | 2212 | 2 | 8.4e-06 | 0.0041 |
| 10 | EGFR | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 7 | 218 | 4 | 68888 | 53 | 1e-05 | 0.0044 |
Note:
n - number of (nonsilent) mutations in this gene across the individual set.
cos = number of unique mutated sites in this gene in COSMIC
n_cos = overlap between n and cos.
N_cos = number of individuals times cos.
cos_ev = total evidence: number of reports in COSMIC for mutations seen in this gene.
p = p-value for seeing the observed amount of overlap in this gene)
q = q-value, False Discovery Rate (Benjamini-Hochberg procedure)
There were no clustered mutations discovered.
Table 5. Get Full Table A Ranked List of Significantly Mutated Genesets. (Source: MSigDB GSEA Cannonical Pathway Set).Number of significant genesets found: 61. Number of genesets displayed: 10
| rank | geneset | description | genes | N_genes | mut_tally | N | n | npat | nsite | nsil | n1 | n2 | n3 | n4 | n5 | n6 | p | q |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HSA04010_MAPK_SIGNALING_PATHWAY | Genes involved in MAPK signaling pathway | ACVR1B, ACVR1C, AKT1, AKT2, AKT3, ARRB1, ARRB2, ATF2, ATF4, BDNF, BRAF, CACNA1A, CACNA1B, CACNA1C, CACNA1D, CACNA1E, CACNA1F, CACNA1G, CACNA1H, CACNA1I, CACNA1S, CACNA2D1, CACNA2D2, CACNA2D3, CACNA2D4, CACNB1, CACNB2, CACNB3, CACNB4, CACNG1, CACNG2, CACNG3, CACNG4, CACNG5, CACNG6, CACNG7, CACNG8, CASP3, CD14, CDC25B, CDC42, CHP, CHUK, CRK, CRKL, DAXX, DDIT3, DUSP1, DUSP10, DUSP14, DUSP16, DUSP2, DUSP3, DUSP4, DUSP5, DUSP6, DUSP7, DUSP8, DUSP9, ECSIT, EGF, EGFR, ELK1, ELK4, EVI1, FAS, FASLG, FGF1, FGF10, FGF11, FGF12, FGF13, FGF14, FGF16, FGF17, FGF18, FGF19, FGF2, FGF20, FGF21, FGF22, FGF23, FGF3, FGF4, FGF5, FGF6, FGF7, FGF8, FGF9, FGFR1, FGFR2, FGFR3, FGFR4, FLNA, FLNB, FLNC, FOS, GADD45A, GADD45B, GADD45G, GNA12, GNG12, GRB2, HRAS, IKBKB, IKBKG, IL1A, IL1B, IL1R1, IL1R2, JUN, JUND, KRAS, LOC653852, MAP2K1, MAP2K1IP1, MAP2K2, MAP2K3, MAP2K4, MAP2K5, MAP2K6, MAP2K7, MAP3K1, MAP3K10, MAP3K12, MAP3K13, MAP3K14, MAP3K2, MAP3K3, MAP3K4, MAP3K5, MAP3K6, MAP3K7, MAP3K7IP1, MAP3K7IP2, MAP3K8, MAP4K1, MAP4K2, MAP4K3, MAP4K4, MAPK1, MAPK10, MAPK11, MAPK12, MAPK13, MAPK14, MAPK3, MAPK7, MAPK8, MAPK8IP1, MAPK8IP2, MAPK8IP3, MAPK9, MAPKAPK2, MAPKAPK3, MAPKAPK5, MAPT, MAX, MEF2C, MKNK1, MKNK2, MOS, MRAS, MYC, NF1, NFATC2, NFATC4, NFKB1, NFKB2, NGFB, NLK, NR4A1, NRAS, NTF3, NTF5, NTRK1, NTRK2, PAK1, PAK2, PDGFA, PDGFB, PDGFRA, PDGFRB, PLA2G10, PLA2G12A, PLA2G12B, PLA2G1B, PLA2G2A, PLA2G2D, PLA2G2E, PLA2G2F, PLA2G3, PLA2G4A, PLA2G5, PLA2G6, PPM1A, PPM1B, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PPP5C, PRKACA, PRKACB, PRKACG, PRKCA, PRKCB1, PRKCG, PRKX, PRKY, PTPN5, PTPN7, PTPRR, RAC1, RAC2, RAC3, RAF1, RAP1A, RAP1B, RAPGEF2, RASA1, RASA2, RASGRF1, RASGRF2, RASGRP1, RASGRP2, RASGRP3, RASGRP4, RPS6KA1, RPS6KA2, RPS6KA3, RPS6KA4, RPS6KA5, RPS6KA6, RRAS, RRAS2, SOS1, SOS2, SRF, STK3, STK4, STMN1, TAOK1, TAOK2, TAOK3, TGFB1, TGFB2, TGFB3, TGFBR1, TGFBR2, TNF, TNFRSF1A, TP53, TRAF2, TRAF6, ZAK | 232 | ACVR1B(1), ACVR1C(2), BRAF(2), CACNA1A(5), CACNA1B(1), CACNA1C(9), CACNA1D(6), CACNA1E(4), CACNA1F(4), CACNA1G(3), CACNA1S(5), CACNA2D1(4), CACNA2D2(1), CACNA2D3(1), CACNA2D4(1), CACNB1(1), CACNB2(1), CACNB4(1), CACNG3(1), CACNG6(2), CDC25B(2), CHUK(1), DAXX(1), DDIT3(2), DUSP1(2), DUSP7(1), DUSP9(2), ECSIT(1), EGFR(7), ELK1(2), FGF12(1), FGF2(1), FGF23(2), FGF5(1), FGF6(1), FGFR4(1), FLNA(1), FLNB(3), FLNC(3), FOS(1), GADD45A(1), GADD45B(1), GNA12(1), GRB2(2), IKBKB(1), KRAS(2), MAP2K3(1), MAP2K4(1), MAP2K6(1), MAP3K1(1), MAP3K10(2), MAP3K12(1), MAP3K13(2), MAP3K14(1), MAP3K2(2), MAP3K3(1), MAP3K4(2), MAP3K5(2), MAP3K6(1), MAP3K7(3), MAP4K1(1), MAP4K3(3), MAPK14(1), MAPK8(1), MAPK8IP1(1), MAPK9(1), MAPKAPK3(1), MAPKAPK5(1), MEF2C(1), MKNK1(1), MOS(1), MRAS(1), NF1(15), NFATC2(1), NFATC4(1), NFKB1(1), NLK(1), NRAS(2), NTF3(1), NTRK1(4), NTRK2(1), PAK2(1), PDGFRA(3), PDGFRB(3), PLA2G12A(1), PLA2G12B(1), PLA2G2F(1), PLA2G4A(1), PLA2G6(1), PPP3CC(1), PPP5C(1), PRKACG(1), PRKCA(1), PRKCG(1), PTPN5(3), PTPRR(1), RAPGEF2(1), RASA1(1), RASA2(2), RASGRF1(2), RASGRF2(4), RASGRP1(1), RASGRP2(1), RASGRP3(2), RPS6KA1(1), RPS6KA2(1), RPS6KA4(1), RPS6KA5(1), RRAS(1), SOS1(3), SOS2(1), SRF(1), STK4(1), TAOK1(1), TAOK3(1), TGFBR2(3), TNF(1), TNFRSF1A(1), TP53(279), TRAF2(1), TRAF6(2) | 116594517 | 496 | 293 | 358 | 53 | 79 | 59 | 109 | 115 | 134 | 0 | <1.00e-15 | <3.08e-14 |
| 2 | HSA04110_CELL_CYCLE | Genes involved in cell cycle | ABL1, ANAPC1, ANAPC10, ANAPC11, ANAPC2, ANAPC4, ANAPC5, ANAPC7, ATM, ATR, BUB1, BUB1B, BUB3, CCNA1, CCNA2, CCNB1, CCNB2, CCNB3, CCND1, CCND2, CCND3, CCNE1, CCNE2, CCNH, CDC14A, CDC14B, CDC16, CDC2, CDC20, CDC23, CDC25A, CDC25B, CDC25C, CDC26, CDC27, CDC45L, CDC6, CDC7, CDK2, CDK4, CDK6, CDK7, CDKN1A, CDKN1B, CDKN1C, CDKN2A, CDKN2B, CDKN2C, CDKN2D, CHEK1, CHEK2, CREBBP, CUL1, DBF4, E2F1, E2F2, E2F3, EP300, ESPL1, FZR1, GADD45A, GADD45B, GADD45G, GSK3B, hCG_1982709, HDAC1, HDAC2, LOC440917, LOC728919, MAD1L1, MAD2L1, MAD2L2, MCM2, MCM3, MCM4, MCM5, MCM6, MCM7, MDM2, ORC1L, ORC2L, ORC3L, ORC4L, ORC5L, ORC6L, PCNA, PKMYT1, PLK1, PRKDC, PTTG1, PTTG2, RB1, RBL1, RBL2, RBX1, SFN, SKP1, SKP2, SMAD2, SMAD3, SMAD4, SMC1A, SMC1B, TFDP1, TGFB1, TGFB2, TGFB3, TP53, WEE1, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ | 106 | ABL1(1), ANAPC1(1), ANAPC2(1), ANAPC4(1), ANAPC5(1), ATM(5), ATR(2), BUB1(1), BUB1B(2), BUB3(1), CCNA1(2), CCNA2(1), CCNB3(3), CCNH(1), CDC20(1), CDC25B(2), CDC27(4), CDC6(1), CDC7(2), CDK2(1), CDKN1A(1), CDKN1B(1), CDKN2C(2), CDKN2D(1), CHEK2(1), CREBBP(7), E2F2(1), EP300(1), GADD45A(1), GADD45B(1), GSK3B(1), HDAC1(1), MAD2L2(1), MCM2(2), MCM3(1), MCM4(2), MCM5(2), ORC1L(1), ORC4L(1), PRKDC(8), RB1(9), RBL1(1), RBL2(3), SKP2(2), SMC1A(5), SMC1B(3), TFDP1(2), TP53(279), YWHAB(1), YWHAE(2), YWHAG(2) | 59044916 | 380 | 287 | 244 | 28 | 60 | 44 | 71 | 84 | 121 | 0 | <1.00e-15 | <3.08e-14 |
| 3 | G1_TO_S_CELL_CYCLE_REACTOME | ATM, CCNA1, CCNB1, CCND1, CCND2, CCND3, CCNE1, CCNE2, CCNG2, CCNH, CDC25A, CDC45L, CDK2, CDK4, CDK7, CDKN1A, CDKN1B, CDKN1C, CDKN2A, CDKN2B, CDKN2C, CDKN2D, CREB3, CREB3L1, CREB3L3, CREB3L4, CREBL1, CREBL1, TNXB, E2F1, E2F2, E2F3, E2F4, E2F5, E2F6, FLJ14001, GADD45A, GBA2, MCM2, MCM3, MCM4, MCM5, MCM6, MCM7, MDM2, MNAT1, MYC, MYT1, NACA, NACA, FKSG17, ORC1L, ORC2L, ORC3L, ORC4L, ORC5L, ORC6L, PCNA, POLA2, POLE, POLE2, PRIM1, PRIM2A, RB1, RBL1, RPA1, RPA2, RPA3, TFDP1, TFDP2, TP53, WEE1 | 64 | ATM(5), CCNA1(2), CCNG2(1), CCNH(1), CDK2(1), CDKN1A(1), CDKN1B(1), CDKN2C(2), CDKN2D(1), CREB3L1(1), CREB3L4(1), E2F2(1), E2F5(1), GADD45A(1), GBA2(2), MCM2(2), MCM3(1), MCM4(2), MCM5(2), MNAT1(1), MYT1(2), NACA(4), ORC1L(1), ORC4L(1), POLE(2), POLE2(1), RB1(9), RBL1(1), RPA1(2), RPA2(1), TFDP1(2), TNXB(4), TP53(279) | 32487850 | 339 | 286 | 203 | 24 | 56 | 38 | 60 | 70 | 115 | 0 | <1.00e-15 | <3.08e-14 | |
| 4 | HSA04310_WNT_SIGNALING_PATHWAY | Genes involved in Wnt signaling pathway | APC, APC2, AXIN1, AXIN2, BTRC, CACYBP, CAMK2A, CAMK2B, CAMK2D, CAMK2G, CCND1, CCND2, CCND3, CER1, CHD8, CHP, CREBBP, CSNK1A1, CSNK1A1L, CSNK1E, CSNK2A1, CSNK2A2, CSNK2B, CTBP1, CTBP2, CTNNB1, CTNNBIP1, CUL1, CXXC4, DAAM1, DAAM2, DKK1, DKK2, DKK4, DVL1, DVL2, DVL3, EP300, FBXW11, FOSL1, FRAT1, FRAT2, FZD1, FZD10, FZD2, FZD3, FZD4, FZD5, FZD6, FZD7, FZD8, FZD9, GSK3B, JUN, LEF1, LOC652788, LRP5, LRP6, MAP3K7, MAPK10, MAPK8, MAPK9, MMP7, MYC, NFAT5, NFATC1, NFATC2, NFATC3, NFATC4, NKD1, NKD2, NLK, PLCB1, PLCB2, PLCB3, PLCB4, PORCN, PPARD, PPP2CA, PPP2CB, PPP2R1A, PPP2R1B, PPP2R2A, PPP2R2B, PPP2R2C, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PRICKLE1, PRICKLE2, PRKACA, PRKACB, PRKACG, PRKCA, PRKCB1, PRKCG, PRKX, PRKY, PSEN1, RAC1, RAC2, RAC3, RBX1, RHOA, ROCK1, ROCK2, RUVBL1, SENP2, SFRP1, SFRP2, SFRP4, SFRP5, SIAH1, SKP1, SMAD2, SMAD3, SMAD4, SOX17, TBL1X, TBL1XR1, TBL1Y, TCF7, TCF7L1, TCF7L2, TP53, VANGL1, VANGL2, WIF1, WNT1, WNT10A, WNT10B, WNT11, WNT16, WNT2, WNT2B, WNT3, WNT3A, WNT4, WNT5A, WNT5B, WNT6, WNT7A, WNT7B, WNT8A, WNT8B, WNT9A, WNT9B | 138 | APC(8), AXIN2(1), BTRC(2), CAMK2B(2), CAMK2D(1), CAMK2G(1), CHD8(5), CREBBP(7), CSNK1A1L(1), CSNK1E(1), CSNK2A1(1), CSNK2B(1), CTBP1(1), CTBP2(1), CTNNB1(2), CXXC4(1), DAAM1(2), DAAM2(2), DKK2(2), DVL3(1), EP300(1), FBXW11(1), FOSL1(1), FZD5(2), FZD7(2), GSK3B(1), LRP5(3), LRP6(1), MAP3K7(3), MAPK8(1), MAPK9(1), MMP7(1), NFAT5(1), NFATC1(1), NFATC2(1), NFATC4(1), NLK(1), PLCB1(4), PLCB2(4), PLCB3(2), PLCB4(1), PORCN(2), PPARD(2), PPP2CA(2), PPP2R1A(4), PPP2R2A(1), PPP2R2B(1), PPP3CC(1), PRICKLE1(3), PRICKLE2(2), PRKACG(1), PRKCA(1), PRKCG(1), RHOA(2), SFRP2(1), SFRP5(1), TBL1X(1), TBL1XR1(1), TCF7L2(2), TP53(279), VANGL2(1), WIF1(3), WNT11(2), WNT16(3), WNT2(1), WNT2B(1), WNT4(1), WNT6(2), WNT7A(2), WNT7B(2), WNT8A(1), WNT9A(2), WNT9B(1) | 65706417 | 407 | 284 | 270 | 37 | 65 | 52 | 80 | 90 | 120 | 0 | <1.00e-15 | <3.08e-14 |
| 5 | CELL_CYCLE_KEGG | ABL1, ASK, ATM, BUB1, BUB1B, BUB3, CCNA1, CCNA2, CCNB1, CCNB2, CCNB3, CCND2, CCND3, CCNE1, CCNE2, CCNH, CDAN1, CDC14A, CDC14B, CDC14B, CDC14C, CDC2, CDC20, CDC25A, CDC25B, CDC25C, CDC45L, CDC6, CDC7, CDH1, CDK2, CDK4, CDKN1A, CDKN2A, CHEK1, CHEK2, DTX4, E2F1, E2F2, E2F3, E2F4, E2F5, E2F6, EP300, ESPL1, FLJ14001, GADD45A, GSK3B, HDAC1, HDAC2, HDAC3, HDAC4, HDAC5, HDAC6, HDAC7A, HDAC8, MAD1L1, MAD2L1, MAD2L2, MCM2, MCM3, MCM4, MCM5, MCM6, MCM7, MDM2, MPEG1, MPL, ORC1L, ORC2L, ORC3L, ORC4L, ORC5L, ORC6L, PCNA, PLK1, PRKDC, PTPRA, PTTG1, PTTG2, PTTG3, RB1, RBL1, SKP2, SMAD4, SMC1L1, TBC1D8, TFDP1, TGFB1, TP53, WEE1 | 82 | ABL1(1), ATM(5), BUB1(1), BUB1B(2), BUB3(1), CCNA1(2), CCNA2(1), CCNB3(3), CCNH(1), CDAN1(2), CDC20(1), CDC25B(2), CDC6(1), CDC7(2), CDH1(2), CDK2(1), CDKN1A(1), CHEK2(1), E2F2(1), E2F5(1), EP300(1), GADD45A(1), GSK3B(1), HDAC1(1), HDAC3(2), HDAC4(2), HDAC5(1), HDAC6(2), MAD2L2(1), MCM2(2), MCM3(1), MCM4(2), MCM5(2), MPEG1(1), ORC1L(1), ORC4L(1), PRKDC(8), RB1(9), RBL1(1), SKP2(2), TBC1D8(3), TFDP1(2), TP53(279) | 48736010 | 358 | 283 | 222 | 27 | 56 | 42 | 65 | 80 | 115 | 0 | <1.00e-15 | <3.08e-14 | |
| 6 | HSA04210_APOPTOSIS | Genes involved in apoptosis | AIFM1, AKT1, AKT2, AKT3, APAF1, ATM, BAD, BAX, BCL2, BCL2L1, BID, BIRC2, BIRC3, BIRC4, CAPN1, CAPN2, CASP10, CASP3, CASP6, CASP7, CASP8, CASP9, CFLAR, CHP, CHUK, CSF2RB, CYCS, DFFA, DFFB, ENDOG, FADD, FAS, FASLG, IKBKB, IKBKG, IL1A, IL1B, IL1R1, IL1RAP, IL3, IL3RA, IRAK1, IRAK2, IRAK3, IRAK4, MAP3K14, MYD88, NFKB1, NFKB2, NFKBIA, NGFB, NTRK1, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PPP3CA, PPP3CB, PPP3CC, PPP3R1, PPP3R2, PRKACA, PRKACB, PRKACG, PRKAR1A, PRKAR1B, PRKAR2A, PRKAR2B, RELA, RIPK1, TNF, TNFRSF10A, TNFRSF10B, TNFRSF10C, TNFRSF10D, TNFRSF1A, TNFSF10, TP53, TRADD, TRAF2 | 78 | AIFM1(2), APAF1(2), ATM(5), BID(1), BIRC3(2), CASP6(1), CASP7(1), CASP9(1), CHUK(1), CSF2RB(3), FADD(1), IKBKB(1), IRAK1(2), IRAK2(1), IRAK3(2), IRAK4(1), MAP3K14(1), NFKB1(1), NTRK1(4), PIK3CA(2), PIK3CB(1), PIK3CG(3), PIK3R1(1), PIK3R2(2), PIK3R5(1), PPP3CC(1), PRKACG(1), PRKAR2A(1), PRKAR2B(1), RELA(1), TNF(1), TNFRSF10A(2), TNFRSF10D(1), TNFRSF1A(1), TNFSF10(1), TP53(279), TRAF2(1) | 35255671 | 334 | 282 | 198 | 25 | 53 | 42 | 59 | 75 | 105 | 0 | <1.00e-15 | <3.08e-14 |
| 7 | TELPATHWAY | Telomerase is a ribonucleotide protein that adds telomeric repeats to the 3' ends of chromosomes. | AKT1, BCL2, EGFR, G22P1, HSPCA, IGF1R, KRAS2, MYC, POLR2A, PPP2CA, PRKCA, RB1, TEP1, TERF1, TERT, TNKS, TP53, XRCC5 | 15 | EGFR(7), IGF1R(1), PPP2CA(2), PRKCA(1), RB1(9), TEP1(5), TERF1(1), TERT(1), TNKS(1), TP53(279) | 12360237 | 307 | 281 | 171 | 10 | 50 | 33 | 49 | 65 | 110 | 0 | <1.00e-15 | <3.08e-14 |
| 8 | APOPTOSIS_GENMAPP | APAF1, BAK1, BCL2L7P1, BAX, BCL2, BCL2L1, BID, BIRC2, BIRC3, BIRC4, CASP2, CASP3, CASP6, CASP7, CASP8, CASP9, CYCS, FADD, FAS, FASLG, GZMB, IKBKG, JUN, MAP2K4, MAP3K1, MAP3K14, MAPK10, MCL1, MDM2, MYC, NFKB1, NFKBIA, PARP1, PRF1, RELA, RIPK1, TNF, TNFRSF1A, TNFRSF1B, TNFSF10, TP53, TRADD, TRAF1, TRAF2 | 40 | APAF1(2), BAK1(1), BID(1), BIRC3(2), CASP2(1), CASP6(1), CASP7(1), CASP9(1), FADD(1), MAP2K4(1), MAP3K1(1), MAP3K14(1), NFKB1(1), PRF1(2), RELA(1), TNF(1), TNFRSF1A(1), TNFSF10(1), TP53(279), TRAF1(4), TRAF2(1) | 16033981 | 305 | 280 | 169 | 10 | 50 | 36 | 48 | 67 | 104 | 0 | <1.00e-15 | <3.08e-14 | |
| 9 | G1PATHWAY | CDK4/6-cyclin D and CDK2-cyclin E phosphorylate Rb, which allows the transcription of genes needed for the G1/S cell cycle transition. | ABL1, ATM, ATR, CCNA1, CCND1, CCNE1, CDC2, CDC25A, CDK2, CDK4, CDK6, CDKN1A, CDKN1B, CDKN2A, CDKN2B, DHFR, E2F1, GSK3B, HDAC1, MADH3, MADH4, RB1, SKP2, TFDP1, TGFB1, TGFB2, TGFB3, TP53 | 25 | ABL1(1), ATM(5), ATR(2), CCNA1(2), CDK2(1), CDKN1A(1), CDKN1B(1), GSK3B(1), HDAC1(1), RB1(9), SKP2(2), TFDP1(2), TP53(279) | 13302114 | 307 | 280 | 171 | 8 | 48 | 37 | 47 | 66 | 109 | 0 | <1.00e-15 | <3.08e-14 |
| 10 | G2PATHWAY | Activated Cdc2-cyclin B kinase regulates the G2/M transition; DNA damage stimulates the DNA-PK/ATM/ATR kinases, which inactivate Cdc2. | ATM, ATR, BRCA1, CCNB1, CDC2, CDC25A, CDC25B, CDC25C, CDC34, CDKN1A, CDKN2D, CHEK1, CHEK2, EP300, GADD45A, MDM2, MYT1, PLK, PRKDC, RPS6KA1, TP53, WEE1, YWHAH, YWHAQ | 21 | ATM(5), ATR(2), BRCA1(12), CDC25B(2), CDKN1A(1), CDKN2D(1), CHEK2(1), EP300(1), GADD45A(1), MYT1(2), PRKDC(8), RPS6KA1(1), TP53(279) | 18900608 | 316 | 280 | 180 | 7 | 49 | 34 | 46 | 71 | 116 | 0 | <1.00e-15 | <3.08e-14 |
Table 6. Get Full Table A Ranked List of Significantly Mutated Genesets (Excluding Significantly Mutated Genes). Number of significant genesets found: 13. Number of genesets displayed: 10
| rank | geneset | description | genes | N_genes | mut_tally | N | n | npat | nsite | nsil | n1 | n2 | n3 | n4 | n5 | n6 | p | q |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HSA04080_NEUROACTIVE_LIGAND_RECEPTOR_INTERACTION | Genes involved in neuroactive ligand-receptor interaction | ADCYAP1R1, ADORA1, ADORA2A, ADORA2B, ADORA3, ADRA1A, ADRA1B, ADRA2A, ADRA2B, ADRA2C, ADRB1, ADRB2, ADRB3, AGTR1, AGTR2, AGTRL1, AVPR1A, AVPR1B, AVPR2, BDKRB1, BDKRB2, BRS3, C3AR1, C5AR1, CALCR, CALCRL, CCKAR, CCKBR, CGA, CHRM1, CHRM2, CHRM3, CHRM4, CHRM5, CNR1, CNR2, CRHR1, CRHR2, CTSG, CYSLTR1, CYSLTR2, DRD1, DRD2, DRD3, DRD4, DRD5, EDG1, EDG2, EDG3, EDG4, EDG5, EDG6, EDG7, EDG8, EDNRA, EDNRB, F2, F2R, F2RL1, F2RL2, F2RL3, FPR1, FPRL1, FPRL2, FSHB, FSHR, GABBR1, GABBR2, GABRA1, GABRA2, GABRA3, GABRA4, GABRA5, GABRA6, GABRB1, GABRB2, GABRB3, GABRD, GABRE, GABRG1, GABRG2, GABRG3, GABRP, GABRQ, GABRR1, GABRR2, GALR1, GALR2, GALR3, GCGR, GH1, GH2, GHR, GHRHR, GHSR, GIPR, GLP1R, GLP2R, GLRA1, GLRA2, GLRA3, GLRB, GNRHR, GPR156, GPR23, GPR35, GPR50, GPR63, GPR83, GRIA1, GRIA2, GRIA3, GRIA4, GRID1, GRID2, GRIK1, GRIK2, GRIK3, GRIK4, GRIK5, GRIN1, GRIN2A, GRIN2B, GRIN2C, GRIN2D, GRIN3A, GRIN3B, GRM1, GRM2, GRM3, GRM4, GRM5, GRM6, GRM7, GRM8, GRPR, GZMA, HCRTR1, HCRTR2, HRH1, HRH2, HRH3, HRH4, HTR1A, HTR1B, HTR1D, HTR1E, HTR1F, HTR2A, HTR2B, HTR2C, HTR4, HTR5A, HTR6, HTR7, KISS1R, LEP, LEPR, LHB, LHCGR, LTB4R, LTB4R2, MAS1, MC1R, MC2R, MC3R, MC4R, MC5R, MCHR1, MCHR2, MLNR, MTNR1A, MTNR1B, NMBR, NMUR1, NMUR2, NPBWR1, NPBWR2, NPFFR1, NPFFR2, NPY1R, NPY2R, NPY5R, NR3C1, NTSR1, NTSR2, OPRD1, OPRK1, OPRL1, OPRM1, OXTR, P2RX1, P2RX2, P2RX3, P2RX4, P2RX5, P2RX7, P2RXL1, P2RY1, P2RY10, P2RY11, P2RY13, P2RY14, P2RY2, P2RY4, P2RY5, P2RY6, P2RY8, PARD3, PPYR1, PRL, PRLHR, PRLR, PRSS1, PRSS2, PRSS3, PTAFR, PTGDR, PTGER1, PTGER2, PTGER3, PTGER4, PTGFR, PTGIR, PTH2R, PTHR1, RXFP1, RXFP2, SCTR, SSTR1, SSTR2, SSTR3, SSTR4, SSTR5, TAAR1, TAAR2, TAAR5, TAAR6, TAAR8, TAAR9, TACR1, TACR2, TACR3, TBXA2R, THRA, THRB, TRHR, TRPV1, TSHB, TSHR, TSPO, UTS2R, VIPR1, VIPR2 | 220 | ADCYAP1R1(1), ADORA2B(2), ADORA3(2), ADRA1A(1), ADRA2B(3), ADRB2(1), AGTR2(2), AVPR1B(3), AVPR2(1), BDKRB1(4), BDKRB2(3), BRS3(1), C5AR1(2), CALCR(3), CALCRL(1), CCKBR(1), CGA(1), CHRM2(4), CHRM3(3), CHRM5(1), CTSG(1), CYSLTR1(1), DRD1(2), DRD3(1), EDNRA(1), F2(3), F2R(2), F2RL2(1), FPR1(1), FSHB(1), FSHR(4), GABBR1(2), GABBR2(2), GABRA1(2), GABRA2(2), GABRA3(1), GABRA4(1), GABRA5(1), GABRB1(2), GABRB3(1), GABRE(3), GABRG1(2), GABRG2(2), GABRQ(2), GALR1(1), GH2(1), GHR(1), GHRHR(1), GHSR(1), GLP2R(1), GLRA1(1), GLRA2(1), GLRA3(4), GPR156(1), GPR35(1), GPR50(2), GRIA1(1), GRIA2(2), GRIA3(2), GRIA4(3), GRID1(3), GRID2(2), GRIK1(1), GRIK2(2), GRIK3(2), GRIK5(2), GRIN2A(3), GRIN2B(3), GRIN2C(1), GRIN2D(3), GRIN3A(3), GRM1(4), GRM3(1), GRM4(1), GRM5(2), GRM6(1), GRM7(4), GRM8(3), GRPR(1), GZMA(2), HCRTR1(1), HCRTR2(1), HRH1(1), HRH4(2), HTR1A(2), HTR1E(2), HTR1F(3), HTR2A(3), HTR2C(4), HTR4(2), HTR5A(2), HTR6(1), LEPR(1), LHCGR(3), MAS1(1), MC2R(3), MC3R(2), MC4R(1), MC5R(2), MCHR2(2), MLNR(1), MTNR1A(1), NMBR(1), NMUR2(2), NPBWR1(1), NPFFR2(2), NR3C1(2), OPRD1(2), OPRK1(1), P2RX1(1), P2RX5(1), P2RY10(2), P2RY13(4), P2RY2(2), P2RY6(1), PARD3(2), PPYR1(2), PRLHR(1), PRLR(2), PRSS2(1), PRSS3(1), PTGDR(1), PTGFR(3), PTH2R(1), RXFP2(1), SCTR(1), SSTR1(1), TAAR6(2), TACR1(1), TACR2(2), TACR3(1), TRPV1(1), TSHR(4), VIPR2(1) | 92059511 | 241 | 147 | 240 | 74 | 46 | 26 | 84 | 64 | 21 | 0 | 9.8e-08 | 6e-05 |
| 2 | HSA04530_TIGHT_JUNCTION | Genes involved in tight junction | ACTB, ACTG1, ACTN1, ACTN2, ACTN3, ACTN4, AKT1, AKT2, AKT3, AMOTL1, ASH1L, CASK, CDC42, CDK4, CGN, CLDN1, CLDN10, CLDN11, CLDN14, CLDN15, CLDN16, CLDN17, CLDN18, CLDN19, CLDN2, CLDN20, CLDN22, CLDN23, CLDN3, CLDN4, CLDN5, CLDN6, CLDN7, CLDN8, CLDN9, CRB3, CSDA, CSNK2A1, CSNK2A2, CSNK2B, CTNNA1, CTNNA2, CTNNA3, CTNNB1, CTTN, EPB41, EPB41L1, EPB41L2, EPB41L3, EXOC3, EXOC4, F11R, GNAI1, GNAI2, GNAI3, HCLS1, HRAS, IGSF5, INADL, JAM2, JAM3, KRAS, LLGL1, LLGL2, MAGI1, MAGI2, MAGI3, MLLT4, MPDZ, MPP5, MRAS, MRCL3, MRLC2, MYH1, MYH10, MYH11, MYH13, MYH14, MYH15, MYH2, MYH3, MYH4, MYH6, MYH7, MYH7B, MYH8, MYH9, MYL2, MYL5, MYL7, MYL8P, MYL9, MYLC2PL, MYLPF, NRAS, OCLN, PARD3, PARD6A, PARD6B, PARD6G, PPM1J, PPP2CA, PPP2CB, PPP2R1A, PPP2R1B, PPP2R2A, PPP2R2B, PPP2R2C, PPP2R3A, PPP2R3B, PPP2R4, PRKCA, PRKCB1, PRKCD, PRKCE, PRKCG, PRKCH, PRKCI, PRKCQ, PRKCZ, PTEN, RAB13, RAB3B, RHOA, RRAS, RRAS2, SPTAN1, SRC, SYMPK, TJAP1, TJP1, TJP2, TJP3, VAPA, YES1, ZAK | 127 | ACTG1(2), ACTN2(3), ACTN3(2), AMOTL1(1), ASH1L(3), CASK(2), CGN(2), CLDN10(1), CLDN11(2), CLDN15(1), CLDN16(1), CLDN17(2), CLDN18(2), CLDN2(1), CLDN6(1), CLDN9(1), CSDA(1), CSNK2A1(1), CSNK2B(1), CTNNA1(2), CTNNA3(1), CTNNB1(2), CTTN(1), EPB41(2), EPB41L3(5), EXOC3(1), EXOC4(4), F11R(2), GNAI2(1), INADL(2), JAM3(1), KRAS(2), LLGL2(3), MAGI1(2), MAGI2(4), MAGI3(3), MLLT4(2), MPDZ(4), MPP5(1), MRAS(1), MYH1(9), MYH10(3), MYH11(7), MYH13(6), MYH14(2), MYH15(5), MYH2(5), MYH3(4), MYH4(9), MYH7(3), MYH7B(5), MYH8(3), MYH9(3), MYL2(1), NRAS(2), PARD3(2), PARD6A(1), PPP2CA(2), PPP2R1A(4), PPP2R2A(1), PPP2R2B(1), PPP2R3A(2), PPP2R4(1), PRKCA(1), PRKCE(1), PRKCG(1), PRKCH(1), PRKCI(2), PRKCQ(3), PRKCZ(2), PTEN(2), RAB3B(1), RHOA(2), RRAS(1), SPTAN1(5), TJAP1(1), TJP1(3), TJP3(1), YES1(1) | 82023148 | 185 | 138 | 183 | 38 | 29 | 15 | 64 | 50 | 27 | 0 | 1.6e-06 | 0.00049 |
| 3 | GPCRDB_CLASS_A_RHODOPSIN_LIKE | ADORA1, ADORA2A, ADORA2B, ADORA3, ADRA1A, ADRA1B, ADRA1D, ADRA2A, ADRA2C, ADRB1, ADRB2, ADRB3, AGTR1, AGTR2, AGTRL1, AVPR1A, AVPR1B, AVPR2, BDKRB1, BDKRB2, BLR1, BRS3, C3AR1, C5R1, CCBP2, CCKAR, CCKBR, CCR1, CCR10, CCR2, CCR3, CCR4, CCR5, CCR6, CCR7, CCR8, CCR9, CCRL1, CCRL2, CHML, CHRM1, CHRM2, CHRM3, CHRM4, CHRM5, CMKLR1, CMKOR1, CNR1, CNR2, CX3CR1, CXCR3, CXCR4, DRD1, DRD2, DRD3, DRD4, DRD5, EDNRA, EDNRB, ELA3A, F2R, F2RL1, F2RL2, F2RL3, FPR1, FPRL1, FPRL2, FSHR, GALR1, GALR2, GALR3, GALT, GHSR, GNB2L1, GPR10, GPR147, GPR17, GPR173, GPR174, GPR23, GPR24, GPR27, GPR3, GPR30, GPR35, GPR37, GPR37L1, GPR4, GPR44, GPR50, GPR6, GPR63, GPR74, GPR77, GPR83, GPR85, GPR87, GPR92, GRPR, HCRTR1, HCRTR2, HRH1, HRH2, HRH3, HTR1A, HTR1B, HTR1D, HTR1E, HTR1F, HTR2A, HTR2B, HTR2C, HTR4, HTR5A, HTR6, HTR7, HTR7, LOC93164, IL8RA, IL8RB, LHCGR, LTB4R, MAS1, MC1R, MC3R, MC4R, MC5R, MLNR, MTNR1A, MTNR1B, NMBR, NMUR1, NMUR2, NPY1R, NPY2R, NPY5R, NPY6R, NTSR1, NTSR2, OPN1SW, OPN3, OPRD1, OPRK1, OPRL1, OPRM1, OR10A5, OR11A1, OR12D3, OR1C1, OR1F1, OR1Q1, OR2H1, OR5V1, OR5V1, OR12D3, OR7A5, OR7C1, OR8B8, OXTR, P2RY1, P2RY10, P2RY11, P2RY12, P2RY13, P2RY14, P2RY2, P2RY5, P2RY6, PPYR1, PTAFR, PTGDR, PTGER1, PTGER2, PTGER4, PTGFR, PTGIR, Rgr, RGR, RHO, RRH, SSTR1, SSTR2, SSTR3, SSTR4, SUCNR1, TBXA2R, TRHR | 151 | ADORA2B(2), ADORA3(2), ADRA1A(1), ADRB2(1), AGTR2(2), AVPR1B(3), AVPR2(1), BDKRB1(4), BDKRB2(3), BRS3(1), CCKBR(1), CCR3(2), CCR6(1), CCR8(1), CCR9(1), CHML(3), CHRM2(4), CHRM3(3), CHRM5(1), CXCR3(2), DRD1(2), DRD3(1), EDNRA(1), F2R(2), F2RL2(1), FPR1(1), FSHR(4), GALR1(1), GHSR(1), GPR173(1), GPR174(1), GPR35(1), GPR37(2), GPR37L1(1), GPR4(2), GPR50(2), GPR77(1), GRPR(1), HCRTR1(1), HCRTR2(1), HRH1(1), HTR1A(2), HTR1E(2), HTR1F(3), HTR2A(3), HTR2C(4), HTR4(2), HTR5A(2), HTR6(1), LHCGR(3), MAS1(1), MC3R(2), MC4R(1), MC5R(2), MLNR(1), MTNR1A(1), NMBR(1), NMUR2(2), OPN1SW(2), OPRD1(2), OPRK1(1), OR10A5(1), OR11A1(1), OR12D3(2), OR5V1(3), OR7C1(2), OR8B8(1), P2RY10(2), P2RY12(1), P2RY13(4), P2RY2(2), P2RY6(1), PPYR1(2), PTGDR(1), PTGFR(3), RRH(1), SSTR1(1), SUCNR1(1) | 48728343 | 135 | 103 | 135 | 30 | 22 | 12 | 52 | 38 | 11 | 0 | 0.000061 | 0.013 | |
| 4 | HSA04810_REGULATION_OF_ACTIN_CYTOSKELETON | Genes involved in regulation of actin cytoskeleton | ABI2, ACTN1, ACTN2, ACTN3, ACTN4, APC, APC2, ARAF, ARHGEF1, ARHGEF12, ARHGEF4, ARHGEF6, ARHGEF7, ARPC1A, ARPC1B, ARPC2, ARPC3, ARPC4, ARPC5, ARPC5L, BAIAP2, BCAR1, BDKRB1, BDKRB2, BRAF, C3orf10, CD14, CDC42, CFL1, CFL2, CHRM1, CHRM2, CHRM3, CHRM4, CHRM5, CRK, CRKL, CSK, CYFIP1, CYFIP2, DIAPH1, DIAPH2, DIAPH3, DOCK1, EGF, EGFR, EZR, F2, F2R, FGD1, FGD3, FGF1, FGF10, FGF11, FGF12, FGF13, FGF14, FGF16, FGF17, FGF18, FGF19, FGF2, FGF20, FGF21, FGF22, FGF23, FGF3, FGF4, FGF5, FGF6, FGF7, FGF8, FGF9, FGFR1, FGFR2, FGFR3, FGFR4, FN1, GIT1, GNA12, GNA13, GNG12, GRLF1, GSN, HRAS, INS, IQGAP1, IQGAP2, IQGAP3, ITGA1, ITGA10, ITGA11, ITGA2, ITGA2B, ITGA3, ITGA4, ITGA5, ITGA6, ITGA7, ITGA8, ITGA9, ITGAD, ITGAE, ITGAL, ITGAM, ITGAV, ITGAX, ITGB1, ITGB2, ITGB3, ITGB4, ITGB5, ITGB6, ITGB7, ITGB8, KRAS, LIMK1, LIMK2, LOC200025, LOC645126, LOC653888, MAP2K1, MAP2K2, MAPK1, MAPK3, MLCK, MOS, MRAS, MRCL3, MRLC2, MSN, MYH10, MYH14, MYH9, MYL2, MYL5, MYL7, MYL8P, MYL9, MYLC2PL, MYLK, MYLK2, MYLPF, NCKAP1, NCKAP1L, NRAS, PAK1, PAK2, PAK3, PAK4, PAK6, PAK7, PDGFA, PDGFB, PDGFRA, PDGFRB, PFN1, PFN2, PFN3, PFN4, PIK3CA, PIK3CB, PIK3CD, PIK3CG, PIK3R1, PIK3R2, PIK3R3, PIK3R5, PIP4K2A, PIP4K2B, PIP4K2C, PIP5K1A, PIP5K1B, PIP5K1C, PIP5K3, PPP1CA, PPP1CB, PPP1CC, PPP1R12A, PPP1R12B, PTK2, PXN, RAC1, RAC2, RAC3, RAF1, RDX, RHOA, ROCK1, ROCK2, RRAS, RRAS2, SCIN, SLC9A1, SOS1, SOS2, SSH1, SSH2, SSH3, TIAM1, TIAM2, TMSB4X, TMSB4Y, TMSL3, VAV1, VAV2, VAV3, VCL, WAS, WASF1, WASF2, WASL | 192 | ACTN2(3), ACTN3(2), APC(8), ARHGEF1(3), ARHGEF12(1), ARHGEF4(2), ARHGEF7(2), ARPC1A(1), ARPC3(1), BDKRB1(4), BDKRB2(3), BRAF(2), CHRM2(4), CHRM3(3), CHRM5(1), CSK(1), CYFIP1(5), CYFIP2(2), DIAPH2(4), DIAPH3(2), DOCK1(4), EGFR(7), F2(3), F2R(2), FGD1(3), FGD3(1), FGF12(1), FGF2(1), FGF23(2), FGF5(1), FGF6(1), FGFR4(1), FN1(1), GNA12(1), GRLF1(6), GSN(1), IQGAP1(2), IQGAP2(6), IQGAP3(2), ITGA1(1), ITGA10(2), ITGA2B(4), ITGA3(2), ITGA4(3), ITGA5(1), ITGA6(1), ITGA8(3), ITGAD(1), ITGAE(1), ITGAL(3), ITGAV(2), ITGAX(1), ITGB2(2), ITGB6(1), ITGB7(4), KRAS(2), LIMK1(1), LIMK2(1), MOS(1), MRAS(1), MSN(2), MYH10(3), MYH14(2), MYH9(3), MYL2(1), MYLK(6), MYLK2(1), NCKAP1(2), NCKAP1L(3), NRAS(2), PAK2(1), PAK3(5), PAK6(2), PAK7(1), PDGFRA(3), PDGFRB(3), PFN2(1), PFN4(1), PIK3CA(2), PIK3CB(1), PIK3CG(3), PIK3R1(1), PIK3R2(2), PIK3R5(1), PIP4K2A(1), PIP4K2B(1), PIP5K1A(1), PIP5K1C(1), PPP1R12A(2), PPP1R12B(4), PTK2(1), PXN(5), RHOA(2), RRAS(1), SCIN(2), SOS1(3), SOS2(1), SSH1(4), SSH2(1), SSH3(3), TIAM1(1), TIAM2(3), VAV1(2), VAV2(4), VAV3(3), VCL(2), WAS(1), WASF2(1) | 116072989 | 240 | 159 | 237 | 64 | 31 | 29 | 88 | 64 | 28 | 0 | 0.0002 | 0.021 |
| 5 | AMIPATHWAY | Endogenous anti-thrombosis pathways are overwhelmed in plaque-narrowed blood vessels, resulting in potentially lethal myocardial infarction. | ADCY1, CD3D, CD3E, CD3G, CD3Z, CD4, CREBBP, CSK, GNAS, GNB1, GNGT1, HLA-DRA, HLA-DRB1, LCK, PRKACB, PRKACG, PRKAR1A, PRKAR1B, PRKAR2A, PRKAR2B, PTPRC, TRA@, TRB@, ZAP70 | 21 | ADCY1(4), CD3E(2), CD4(1), CREBBP(7), CSK(1), GNAS(3), HLA-DRA(1), HLA-DRB1(1), LCK(1), PRKACG(1), PRKAR2A(1), PRKAR2B(1), PTPRC(3), ZAP70(1) | 9488232 | 28 | 28 | 27 | 3 | 5 | 3 | 6 | 6 | 8 | 0 | 0.0002 | 0.021 |
| 6 | CSKPATHWAY | Csk inhibits T-cell activation by phosphorylating Lck; Csk is regulated by cAMP-dependent kinases and is opposed by the T-cell activator CD45. | ADCY1, CD3D, CD3E, CD3G, CD3Z, CD4, CREBBP, CSK, GNAS, GNB1, GNGT1, HLA-DRA, HLA-DRB1, LCK, PRKACB, PRKACG, PRKAR1A, PRKAR1B, PRKAR2A, PRKAR2B, PTPRC, TRA@, TRB@, ZAP70 | 21 | ADCY1(4), CD3E(2), CD4(1), CREBBP(7), CSK(1), GNAS(3), HLA-DRA(1), HLA-DRB1(1), LCK(1), PRKACG(1), PRKAR2A(1), PRKAR2B(1), PTPRC(3), ZAP70(1) | 9488232 | 28 | 28 | 27 | 3 | 5 | 3 | 6 | 6 | 8 | 0 | 0.0002 | 0.021 |
| 7 | GPCRDB_OTHER | ADORA3, ALG6, C5R1, CCKBR, CCR2, CCR3, CCR5, CELSR1, CELSR2, CELSR3, CHRM2, CHRM3, CIDEB, CXCR3, DRD4, EBI2, EDG1, EDNRA, ELA3A, EMR2, EMR3, F2R, FSHR, FY, GHRHR, GNRHR, GPR, GPR116, GPR132, GPR133, GPR135, GPR143, GPR145, GPR17, GPR18, GPR55, GPR56, GPR61, GPR73L1, GPR77, GPR84, GPR88, GRCA, GRM1, GRPR, HRH4, IL8RA, IL8RB, LGR6, LGR7, LPHN2, LPHN3, LTB4R2, MASS1, NTSR1, OR2A9P, OR2M4, OR5E1P, OR7E19P, OR7E47P, OR7E37P, OR7E18P, OR7E35P, LOC441453, OR8G1, LOC442754, OR8G2, P2RY11, P2RY13, PTGFR, RLN3R1, SMO, SSTR2, TAAR5, TSHR, VN1R1 | 50 | ADORA3(2), CCKBR(1), CCR3(2), CELSR1(1), CELSR2(4), CELSR3(6), CHRM2(4), CHRM3(3), CXCR3(2), EDNRA(1), EMR2(1), EMR3(5), F2R(2), FSHR(4), GHRHR(1), GPR116(1), GPR132(2), GPR133(2), GPR55(1), GPR77(1), GRM1(4), GRPR(1), HRH4(2), LGR6(3), LPHN2(1), LPHN3(4), OR8G2(1), P2RY13(4), PTGFR(3), TSHR(4) | 25452491 | 73 | 60 | 73 | 20 | 11 | 10 | 27 | 19 | 6 | 0 | 0.0005 | 0.039 | |
| 8 | FXRPATHWAY | The nuclear receptor transcription factors FXR and LXR are activated by cholesterol metabolites and regulate cholesterol homeostasis. | FABP6, LDLR, NR0B2, NR1H3, NR1H4, RXRA | 6 | LDLR(3), NR1H3(4), NR1H4(3), RXRA(2) | 2264708 | 12 | 12 | 12 | 0 | 2 | 2 | 3 | 2 | 3 | 0 | 0.0005 | 0.039 |
| 9 | HSA04916_MELANOGENESIS | Genes involved in melanogenesis | ADCY1, ADCY2, ADCY3, ADCY4, ADCY5, ADCY6, ADCY7, ADCY8, ADCY9, ASIP, CALM1, CALM2, CALM3, CALML3, CALML6, CAMK2A, CAMK2B, CAMK2D, CAMK2G, CREB1, CREB3, CREB3L1, CREB3L2, CREB3L3, CREB3L4, CREBBP, CTNNB1, DCT, DVL1, DVL2, DVL3, EDN1, EDNRB, EP300, FZD1, FZD10, FZD2, FZD3, FZD4, FZD5, FZD6, FZD7, FZD8, FZD9, GNAI1, GNAI2, GNAI3, GNAO1, GNAQ, GNAS, GSK3B, HRAS, KIT, KITLG, KRAS, LEF1, LOC652788, MAP2K1, MAP2K2, MAPK1, MAPK3, MC1R, MITF, NRAS, PLCB1, PLCB2, PLCB3, PLCB4, POMC, PRKACA, PRKACB, PRKACG, PRKCA, PRKCB1, PRKCG, PRKX, PRKY, RAF1, TCF7, TCF7L1, TCF7L2, TYR, TYRP1, WNT1, WNT10A, WNT10B, WNT11, WNT16, WNT2, WNT2B, WNT3, WNT3A, WNT4, WNT5A, WNT5B, WNT6, WNT7A, WNT7B, WNT8A, WNT8B, WNT9A, WNT9B | 96 | ADCY1(4), ADCY2(6), ADCY3(2), ADCY4(1), ADCY5(3), ADCY7(1), ADCY9(5), CALM3(1), CAMK2B(2), CAMK2D(1), CAMK2G(1), CREB3L1(1), CREB3L4(1), CREBBP(7), CTNNB1(2), DCT(2), DVL3(1), EP300(1), FZD5(2), FZD7(2), GNAI2(1), GNAS(3), GSK3B(1), KIT(7), KITLG(1), KRAS(2), MITF(2), NRAS(2), PLCB1(4), PLCB2(4), PLCB3(2), PLCB4(1), POMC(1), PRKACG(1), PRKCA(1), PRKCG(1), TCF7L2(2), TYR(1), WNT11(2), WNT16(3), WNT2(1), WNT2B(1), WNT4(1), WNT6(2), WNT7A(2), WNT7B(2), WNT8A(1), WNT9A(2), WNT9B(1) | 43571057 | 101 | 86 | 96 | 28 | 21 | 14 | 30 | 20 | 16 | 0 | 0.00073 | 0.05 |
| 10 | HSA04340_HEDGEHOG_SIGNALING_PATHWAY | Genes involved in Hedgehog signaling pathway | BMP2, BMP4, BMP5, BMP6, BMP7, BMP8A, BMP8B, BTRC, CSNK1A1, CSNK1A1L, CSNK1D, CSNK1E, CSNK1G1, CSNK1G2, CSNK1G3, DHH, FBXW11, GAS1, GLI1, GLI2, GLI3, GSK3B, HHIP, IHH, LRP2, PRKACA, PRKACB, PRKACG, PRKX, PRKY, PTCH1, PTCH2, RAB23, SHH, SMO, STK36, SUFU, WNT1, WNT10A, WNT10B, WNT11, WNT16, WNT2, WNT2B, WNT3, WNT3A, WNT4, WNT5A, WNT5B, WNT6, WNT7A, WNT7B, WNT8A, WNT8B, WNT9A, WNT9B, ZIC2 | 54 | BMP2(1), BMP5(1), BMP6(1), BTRC(2), CSNK1A1L(1), CSNK1E(1), FBXW11(1), GLI1(5), GLI2(10), GLI3(2), GSK3B(1), HHIP(2), LRP2(16), PRKACG(1), PTCH1(6), PTCH2(1), STK36(1), WNT11(2), WNT16(3), WNT2(1), WNT2B(1), WNT4(1), WNT6(2), WNT7A(2), WNT7B(2), WNT8A(1), WNT9A(2), WNT9B(1) | 24713567 | 71 | 61 | 70 | 10 | 10 | 14 | 26 | 15 | 6 | 0 | 0.00095 | 0.057 |
In brief, we tabulate the number of mutations and the number of covered bases for each gene. The counts are broken down by mutation context category: four context categories that are discovered by MutSig, and one for indel and 'null' mutations, which include indels, nonsense mutations, splice-site mutations, and non-stop (read-through) mutations. For each gene, we calculate the probability of seeing the observed constellation of mutations, i.e. the product P1 x P2 x ... x Pm, or a more extreme one, given the background mutation rates calculated across the dataset. [1]
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.