This pipeline computes the correlation between significantly recurrent gene mutations and molecular subtypes.
Testing the association between mutation status of 41 genes and 8 molecular subtypes across 323 patients, 20 significant findings detected with P value < 0.05 and Q value < 0.25.
-
BRAF mutation correlated to 'METHLYATION_CNMF', 'RPPA_CNMF', 'RPPA_CHIERARCHICAL', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
HRAS mutation correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'RPPA_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
NRAS mutation correlated to 'METHLYATION_CNMF', 'RPPA_CHIERARCHICAL', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
Table 1. Get Full Table Overview of the association between mutation status of 41 genes and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 20 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
RPPA CNMF |
RPPA CHIERARCHICAL |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
||
| nMutated (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| BRAF | 183 (57%) | 140 |
0.00338 (0.874) |
1.08e-37 (2.98e-35) |
3.6e-07 (9.6e-05) |
3.71e-13 (1.01e-10) |
4.03e-46 (1.12e-43) |
1.56e-39 (4.29e-37) |
1.34e-43 (3.7e-41) |
2.16e-45 (6e-43) |
| HRAS | 12 (4%) | 311 |
0.000107 (0.0279) |
2.48e-06 (0.000658) |
0.000224 (0.0582) |
0.00349 (0.9) |
1.37e-05 (0.00361) |
6.79e-06 (0.00179) |
1.49e-06 (0.000397) |
3.38e-07 (9.06e-05) |
| NRAS | 26 (8%) | 297 |
0.0098 (1.00) |
6.63e-14 (1.82e-11) |
1 (1.00) |
0.000209 (0.0545) |
4.7e-11 (1.26e-08) |
4.03e-11 (1.09e-08) |
1.64e-12 (4.43e-10) |
1.76e-13 (4.8e-11) |
| EMG1 | 6 (2%) | 317 |
0.557 (1.00) |
0.862 (1.00) |
0.267 (1.00) |
0.147 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.166 (1.00) |
| PTTG1IP | 4 (1%) | 319 |
1 (1.00) |
1 (1.00) |
0.242 (1.00) |
0.252 (1.00) |
0.471 (1.00) |
0.0379 (1.00) |
||
| RPTN | 8 (2%) | 315 |
1 (1.00) |
0.688 (1.00) |
0.327 (1.00) |
0.485 (1.00) |
0.0249 (1.00) |
0.415 (1.00) |
0.741 (1.00) |
1 (1.00) |
| EIF1AX | 5 (2%) | 318 |
1 (1.00) |
0.028 (1.00) |
0.444 (1.00) |
0.115 (1.00) |
0.0621 (1.00) |
0.0109 (1.00) |
0.0616 (1.00) |
0.234 (1.00) |
| CCDC15 | 5 (2%) | 318 |
1 (1.00) |
0.17 (1.00) |
1 (1.00) |
0.778 (1.00) |
0.132 (1.00) |
0.136 (1.00) |
0.628 (1.00) |
0.527 (1.00) |
| ZNF845 | 6 (2%) | 317 |
0.557 (1.00) |
0.615 (1.00) |
0.64 (1.00) |
0.35 (1.00) |
0.943 (1.00) |
0.166 (1.00) |
0.38 (1.00) |
0.327 (1.00) |
| ZNF878 | 4 (1%) | 319 |
1 (1.00) |
0.456 (1.00) |
0.0454 (1.00) |
0.833 (1.00) |
0.694 (1.00) |
1 (1.00) |
||
| TG | 16 (5%) | 307 |
1 (1.00) |
0.0255 (1.00) |
0.286 (1.00) |
0.273 (1.00) |
0.406 (1.00) |
0.293 (1.00) |
0.0426 (1.00) |
0.0244 (1.00) |
| PRB2 | 6 (2%) | 317 |
1 (1.00) |
0.615 (1.00) |
0.0809 (1.00) |
1 (1.00) |
0.474 (1.00) |
0.723 (1.00) |
0.525 (1.00) |
0.594 (1.00) |
| R3HDM2 | 4 (1%) | 319 |
0.0134 (1.00) |
0.0495 (1.00) |
0.0699 (1.00) |
0.0388 (1.00) |
0.0202 (1.00) |
0.00788 (1.00) |
||
| IL32 | 3 (1%) | 320 |
1 (1.00) |
0.39 (1.00) |
1 (1.00) |
0.78 (1.00) |
0.206 (1.00) |
0.641 (1.00) |
||
| TMCO2 | 3 (1%) | 320 |
1 (1.00) |
0.545 (1.00) |
0.778 (1.00) |
0.185 (1.00) |
0.492 (1.00) |
0.303 (1.00) |
0.413 (1.00) |
0.779 (1.00) |
| PPTC7 | 3 (1%) | 320 |
0.079 (1.00) |
0.0835 (1.00) |
0.172 (1.00) |
0.0916 (1.00) |
0.306 (1.00) |
0.395 (1.00) |
||
| MUC7 | 5 (2%) | 318 |
1 (1.00) |
0.274 (1.00) |
0.684 (1.00) |
0.778 (1.00) |
0.805 (1.00) |
0.848 (1.00) |
0.453 (1.00) |
0.632 (1.00) |
| LYPD3 | 3 (1%) | 320 |
0.55 (1.00) |
0.545 (1.00) |
0.492 (1.00) |
0.303 (1.00) |
0.65 (1.00) |
0.779 (1.00) |
||
| TMEM90B | 3 (1%) | 320 |
0.136 (1.00) |
1 (1.00) |
1 (1.00) |
0.78 (1.00) |
1 (1.00) |
1 (1.00) |
||
| ZNF799 | 5 (2%) | 318 |
0.473 (1.00) |
0.834 (1.00) |
0.51 (1.00) |
1 (1.00) |
0.592 (1.00) |
0.848 (1.00) |
0.453 (1.00) |
1 (1.00) |
| PPM1D | 5 (2%) | 318 |
1 (1.00) |
1 (1.00) |
0.778 (1.00) |
0.35 (1.00) |
0.693 (1.00) |
0.848 (1.00) |
0.868 (1.00) |
0.281 (1.00) |
| MAP3K3 | 4 (1%) | 319 |
0.381 (1.00) |
0.276 (1.00) |
0.393 (1.00) |
0.147 (1.00) |
0.274 (1.00) |
0.184 (1.00) |
0.471 (1.00) |
0.391 (1.00) |
| TROAP | 3 (1%) | 320 |
1 (1.00) |
0.39 (1.00) |
0.778 (1.00) |
1 (1.00) |
0.746 (1.00) |
1 (1.00) |
0.306 (1.00) |
0.395 (1.00) |
| SYNPO2L | 3 (1%) | 320 |
0.55 (1.00) |
1 (1.00) |
1 (1.00) |
0.78 (1.00) |
1 (1.00) |
1 (1.00) |
||
| ATAD2 | 4 (1%) | 319 |
1 (1.00) |
0.612 (1.00) |
0.808 (1.00) |
1 (1.00) |
0.694 (1.00) |
1 (1.00) |
||
| KRAS | 3 (1%) | 320 |
1 (1.00) |
0.0835 (1.00) |
0.172 (1.00) |
0.0916 (1.00) |
0.0642 (1.00) |
0.0267 (1.00) |
||
| SLC5A11 | 3 (1%) | 320 |
0.55 (1.00) |
0.716 (1.00) |
0.306 (1.00) |
0.395 (1.00) |
||||
| PRG4 | 4 (1%) | 319 |
0.232 (1.00) |
0.362 (1.00) |
0.0128 (1.00) |
0.142 (1.00) |
0.471 (1.00) |
0.558 (1.00) |
||
| SCUBE2 | 3 (1%) | 320 |
0.246 (1.00) |
0.314 (1.00) |
0.303 (1.00) |
0.65 (1.00) |
0.511 (1.00) |
|||
| ZNF479 | 3 (1%) | 320 |
1 (1.00) |
0.39 (1.00) |
1 (1.00) |
0.78 (1.00) |
1 (1.00) |
1 (1.00) |
||
| SREBF2 | 3 (1%) | 320 |
1 (1.00) |
0.39 (1.00) |
0.568 (1.00) |
1 (1.00) |
0.413 (1.00) |
0.779 (1.00) |
||
| SLC26A11 | 3 (1%) | 320 |
0.079 (1.00) |
1 (1.00) |
0.368 (1.00) |
0.78 (1.00) |
0.777 (1.00) |
0.292 (1.00) |
||
| TSC22D1 | 3 (1%) | 320 |
1 (1.00) |
1 (1.00) |
0.746 (1.00) |
1 (1.00) |
1 (1.00) |
0.641 (1.00) |
||
| ANKRD30A | 5 (2%) | 318 |
1 (1.00) |
0.43 (1.00) |
0.749 (1.00) |
0.507 (1.00) |
0.742 (1.00) |
0.865 (1.00) |
||
| FAM155A | 4 (1%) | 319 |
0.144 (1.00) |
0.612 (1.00) |
0.778 (1.00) |
1 (1.00) |
0.332 (1.00) |
0.832 (1.00) |
||
| SLA | 3 (1%) | 320 |
0.55 (1.00) |
0.0835 (1.00) |
0.29 (1.00) |
0.35 (1.00) |
0.172 (1.00) |
0.0916 (1.00) |
0.0642 (1.00) |
0.0267 (1.00) |
| ZFHX3 | 10 (3%) | 313 |
0.178 (1.00) |
0.737 (1.00) |
0.483 (1.00) |
0.533 (1.00) |
0.794 (1.00) |
0.437 (1.00) |
0.615 (1.00) |
0.727 (1.00) |
| ARMCX3 | 3 (1%) | 320 |
0.55 (1.00) |
0.39 (1.00) |
0.646 (1.00) |
0.213 (1.00) |
0.528 (1.00) |
0.199 (1.00) |
||
| SLC25A45 | 3 (1%) | 320 |
0.136 (1.00) |
1 (1.00) |
0.206 (1.00) |
0.641 (1.00) |
||||
| COL5A3 | 6 (2%) | 317 |
0.758 (1.00) |
1 (1.00) |
0.778 (1.00) |
1 (1.00) |
0.044 (1.00) |
0.753 (1.00) |
0.876 (1.00) |
0.519 (1.00) |
| CDC27 | 3 (1%) | 320 |
1 (1.00) |
1 (1.00) |
0.368 (1.00) |
0.0446 (1.00) |
0.777 (1.00) |
1 (1.00) |
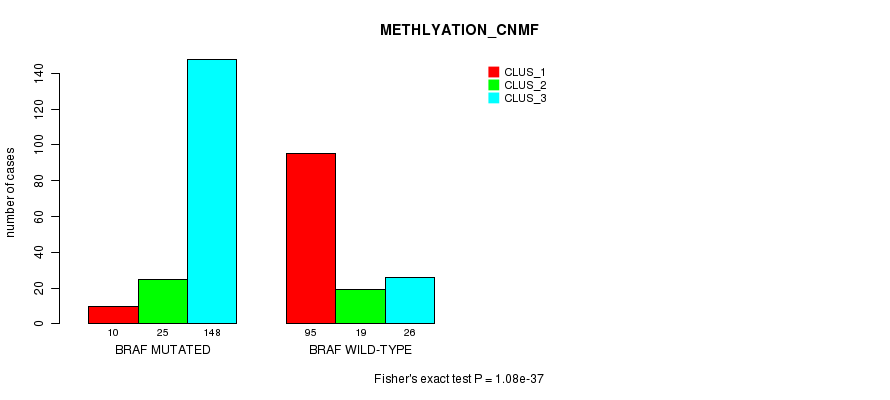
P value = 1.08e-37 (Fisher's exact test), Q value = 3e-35
Table S1. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 105 | 44 | 174 |
| BRAF MUTATED | 10 | 25 | 148 |
| BRAF WILD-TYPE | 95 | 19 | 26 |
Figure S1. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
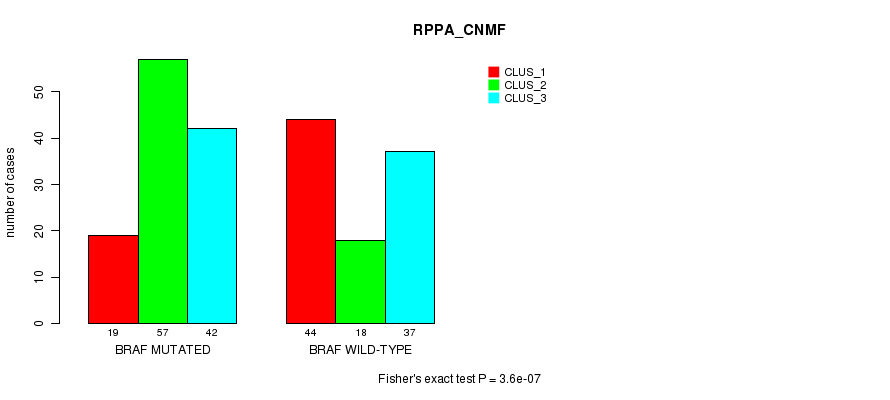
P value = 3.6e-07 (Fisher's exact test), Q value = 9.6e-05
Table S2. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 63 | 75 | 79 |
| BRAF MUTATED | 19 | 57 | 42 |
| BRAF WILD-TYPE | 44 | 18 | 37 |
Figure S2. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
P value = 3.71e-13 (Fisher's exact test), Q value = 1e-10
Table S3. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 95 | 95 |
| BRAF MUTATED | 11 | 78 | 29 |
| BRAF WILD-TYPE | 16 | 17 | 66 |
Figure S3. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'

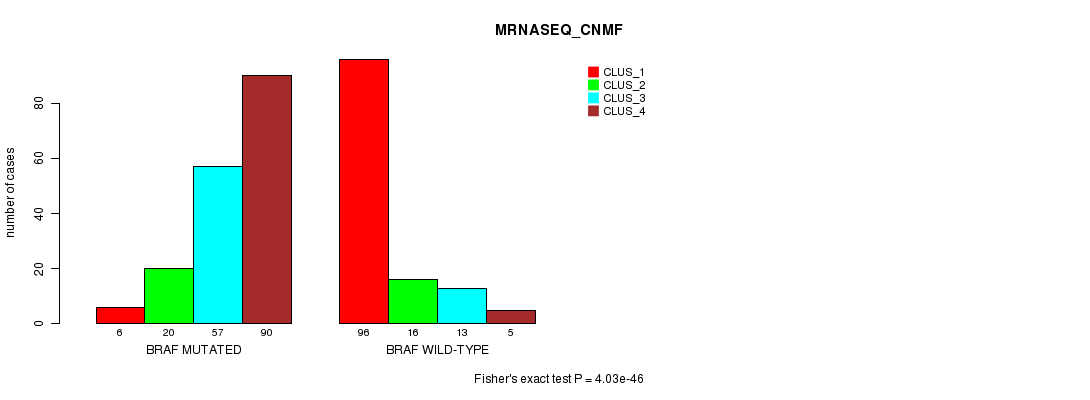
P value = 4.03e-46 (Fisher's exact test), Q value = 1.1e-43
Table S4. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 102 | 36 | 70 | 95 |
| BRAF MUTATED | 6 | 20 | 57 | 90 |
| BRAF WILD-TYPE | 96 | 16 | 13 | 5 |
Figure S4. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
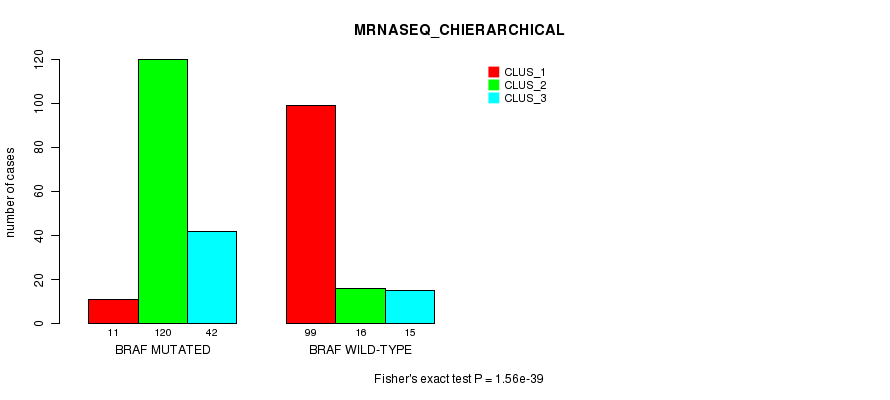
P value = 1.56e-39 (Fisher's exact test), Q value = 4.3e-37
Table S5. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 110 | 136 | 57 |
| BRAF MUTATED | 11 | 120 | 42 |
| BRAF WILD-TYPE | 99 | 16 | 15 |
Figure S5. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
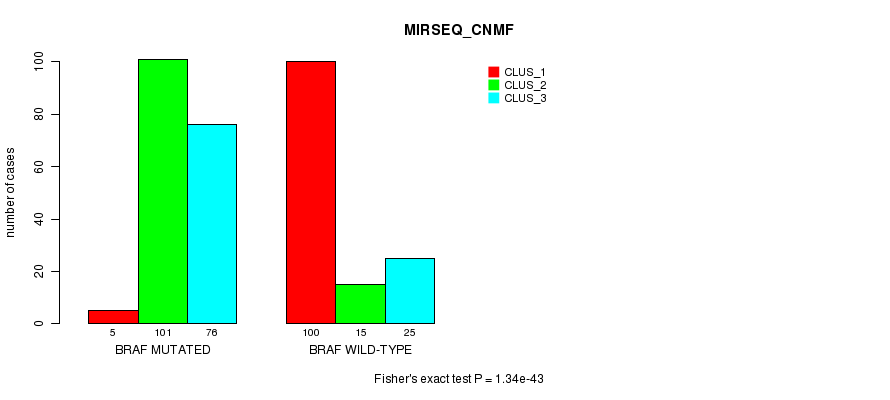
P value = 1.34e-43 (Fisher's exact test), Q value = 3.7e-41
Table S6. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 105 | 116 | 101 |
| BRAF MUTATED | 5 | 101 | 76 |
| BRAF WILD-TYPE | 100 | 15 | 25 |
Figure S6. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
P value = 2.16e-45 (Fisher's exact test), Q value = 6e-43
Table S7. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 97 | 122 | 103 |
| BRAF MUTATED | 1 | 97 | 84 |
| BRAF WILD-TYPE | 96 | 25 | 19 |
Figure S7. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'

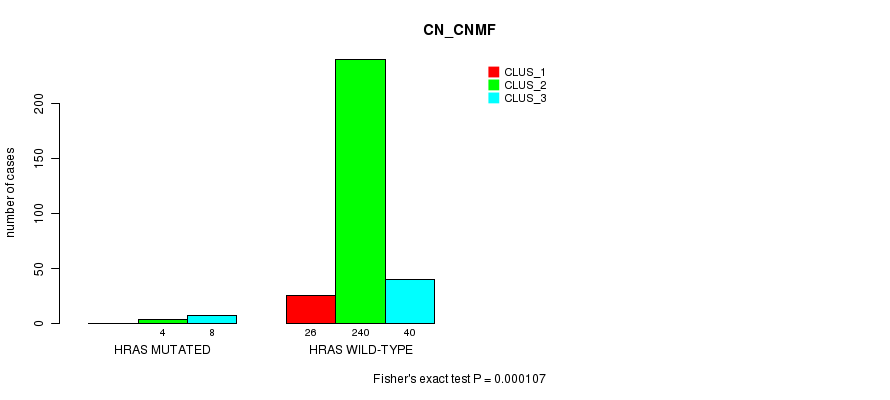
P value = 0.000107 (Fisher's exact test), Q value = 0.028
Table S8. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 26 | 244 | 48 |
| HRAS MUTATED | 0 | 4 | 8 |
| HRAS WILD-TYPE | 26 | 240 | 40 |
Figure S8. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
P value = 2.48e-06 (Fisher's exact test), Q value = 0.00066
Table S9. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 105 | 44 | 174 |
| HRAS MUTATED | 12 | 0 | 0 |
| HRAS WILD-TYPE | 93 | 44 | 174 |
Figure S9. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'

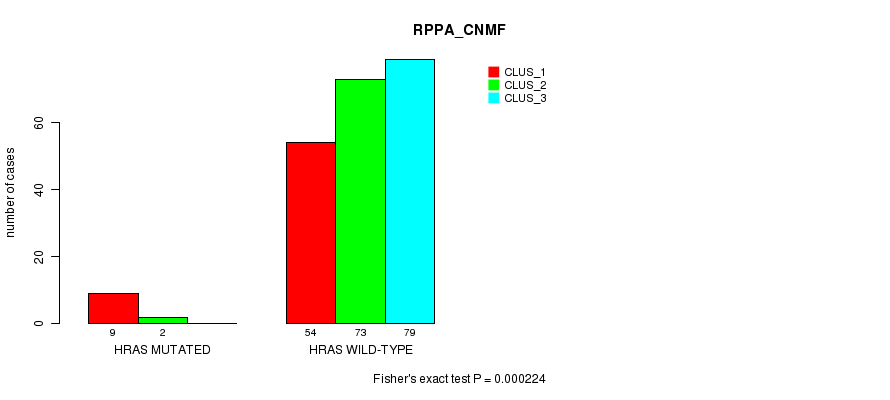
P value = 0.000224 (Fisher's exact test), Q value = 0.058
Table S10. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 63 | 75 | 79 |
| HRAS MUTATED | 9 | 2 | 0 |
| HRAS WILD-TYPE | 54 | 73 | 79 |
Figure S10. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
P value = 1.37e-05 (Fisher's exact test), Q value = 0.0036
Table S11. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 102 | 36 | 70 | 95 |
| HRAS MUTATED | 12 | 0 | 0 | 0 |
| HRAS WILD-TYPE | 90 | 36 | 70 | 95 |
Figure S11. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'

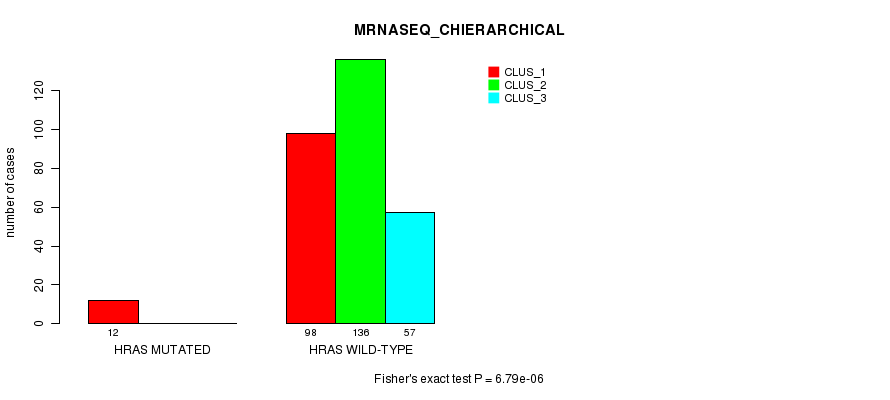
P value = 6.79e-06 (Fisher's exact test), Q value = 0.0018
Table S12. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 110 | 136 | 57 |
| HRAS MUTATED | 12 | 0 | 0 |
| HRAS WILD-TYPE | 98 | 136 | 57 |
Figure S12. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
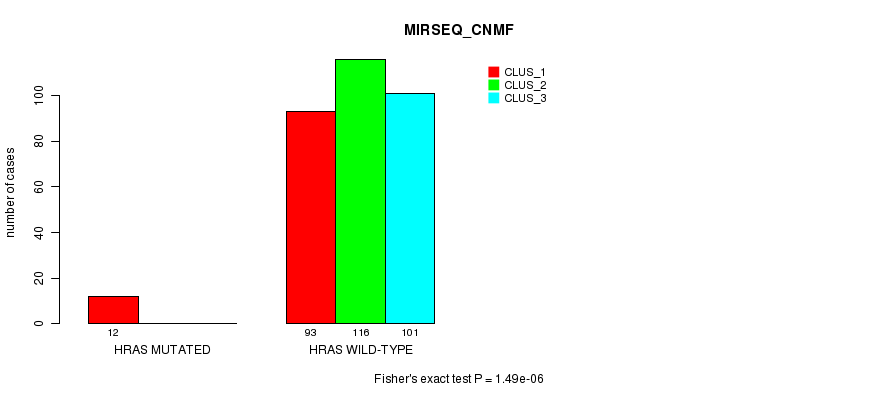
P value = 1.49e-06 (Fisher's exact test), Q value = 4e-04
Table S13. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 105 | 116 | 101 |
| HRAS MUTATED | 12 | 0 | 0 |
| HRAS WILD-TYPE | 93 | 116 | 101 |
Figure S13. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
P value = 3.38e-07 (Fisher's exact test), Q value = 9.1e-05
Table S14. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 97 | 122 | 103 |
| HRAS MUTATED | 12 | 0 | 0 |
| HRAS WILD-TYPE | 85 | 122 | 103 |
Figure S14. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'

P value = 6.63e-14 (Fisher's exact test), Q value = 1.8e-11
Table S15. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 105 | 44 | 174 |
| NRAS MUTATED | 26 | 0 | 0 |
| NRAS WILD-TYPE | 79 | 44 | 174 |
Figure S15. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'

P value = 0.000209 (Fisher's exact test), Q value = 0.054
Table S16. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 95 | 95 |
| NRAS MUTATED | 5 | 0 | 9 |
| NRAS WILD-TYPE | 22 | 95 | 86 |
Figure S16. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'

P value = 4.7e-11 (Fisher's exact test), Q value = 1.3e-08
Table S17. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 102 | 36 | 70 | 95 |
| NRAS MUTATED | 23 | 0 | 0 | 0 |
| NRAS WILD-TYPE | 79 | 36 | 70 | 95 |
Figure S17. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'

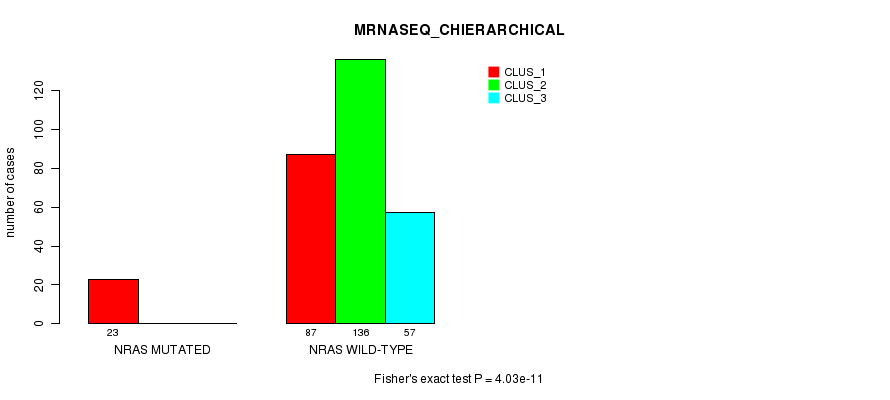
P value = 4.03e-11 (Fisher's exact test), Q value = 1.1e-08
Table S18. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 110 | 136 | 57 |
| NRAS MUTATED | 23 | 0 | 0 |
| NRAS WILD-TYPE | 87 | 136 | 57 |
Figure S18. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
P value = 1.64e-12 (Fisher's exact test), Q value = 4.4e-10
Table S19. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 105 | 116 | 101 |
| NRAS MUTATED | 25 | 0 | 1 |
| NRAS WILD-TYPE | 80 | 116 | 100 |
Figure S19. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'

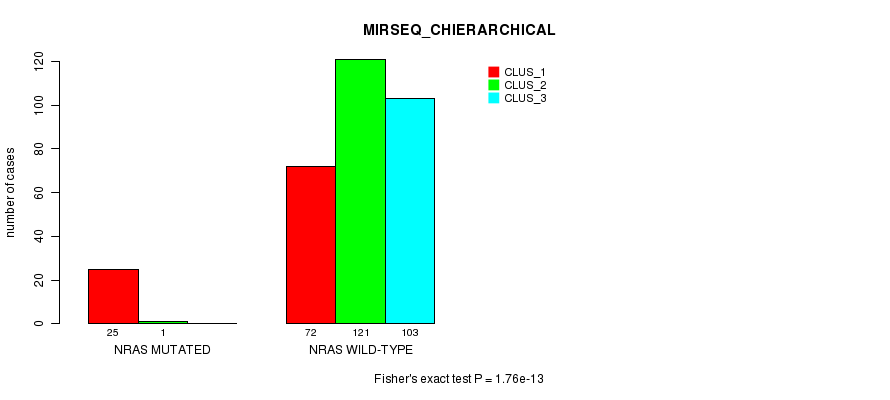
P value = 1.76e-13 (Fisher's exact test), Q value = 4.8e-11
Table S20. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 97 | 122 | 103 |
| NRAS MUTATED | 25 | 1 | 0 |
| NRAS WILD-TYPE | 72 | 121 | 103 |
Figure S20. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
-
Mutation data file = THCA-TP.mutsig.cluster.txt
-
Molecular subtypes file = THCA-TP.transferedmergedcluster.txt
-
Number of patients = 323
-
Number of significantly mutated genes = 41
-
Number of Molecular subtypes = 8
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.