This pipeline computes the correlation between significantly recurrent gene mutations and molecular subtypes.
Testing the association between mutation status of 41 genes and 8 molecular subtypes across 323 patients, 20 significant findings detected with P value < 0.05 and Q value < 0.25.
-
BRAF mutation correlated to 'METHLYATION_CNMF', 'RPPA_CNMF', 'RPPA_CHIERARCHICAL', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
HRAS mutation correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'RPPA_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
NRAS mutation correlated to 'METHLYATION_CNMF', 'RPPA_CHIERARCHICAL', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
Table 1. Get Full Table Overview of the association between mutation status of 41 genes and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 20 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
RPPA CNMF |
RPPA CHIERARCHICAL |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
||
| nMutated (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| BRAF | 183 (57%) | 140 |
0.0194 (1.00) |
3.62e-38 (1.01e-35) |
9.4e-07 (0.000255) |
2.51e-11 (6.84e-09) |
3.88e-49 (1.09e-46) |
7.76e-49 (2.18e-46) |
2.35e-40 (6.57e-38) |
3.32e-43 (9.29e-41) |
| HRAS | 12 (4%) | 311 |
0.000222 (0.0586) |
1.48e-06 (0.000399) |
9.23e-05 (0.0245) |
0.00529 (1.00) |
1.71e-05 (0.00454) |
5.23e-06 (0.0014) |
1.69e-06 (0.000453) |
9.25e-06 (0.00247) |
| NRAS | 26 (8%) | 297 |
0.00692 (1.00) |
6.06e-14 (1.68e-11) |
1 (1.00) |
0.000583 (0.153) |
1.63e-12 (4.47e-10) |
1.11e-13 (3.08e-11) |
1.87e-12 (5.13e-10) |
3.3e-12 (9.02e-10) |
| EMG1 | 6 (2%) | 317 |
0.548 (1.00) |
1 (1.00) |
0.327 (1.00) |
0.0846 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
| PTTG1IP | 4 (1%) | 319 |
1 (1.00) |
1 (1.00) |
0.223 (1.00) |
0.451 (1.00) |
0.471 (1.00) |
0.396 (1.00) |
||
| RPTN | 8 (2%) | 315 |
0.805 (1.00) |
0.442 (1.00) |
0.265 (1.00) |
0.801 (1.00) |
0.0267 (1.00) |
0.663 (1.00) |
0.74 (1.00) |
0.672 (1.00) |
| EIF1AX | 5 (2%) | 318 |
0.731 (1.00) |
0.0278 (1.00) |
0.444 (1.00) |
0.132 (1.00) |
0.0654 (1.00) |
0.0257 (1.00) |
0.0634 (1.00) |
0.199 (1.00) |
| CCDC15 | 5 (2%) | 318 |
0.731 (1.00) |
0.115 (1.00) |
1 (1.00) |
0.381 (1.00) |
0.211 (1.00) |
0.327 (1.00) |
0.627 (1.00) |
0.108 (1.00) |
| ZNF845 | 6 (2%) | 317 |
0.42 (1.00) |
0.637 (1.00) |
0.639 (1.00) |
0.125 (1.00) |
0.943 (1.00) |
0.552 (1.00) |
0.875 (1.00) |
0.334 (1.00) |
| ZNF878 | 4 (1%) | 319 |
1 (1.00) |
0.487 (1.00) |
0.0475 (1.00) |
0.561 (1.00) |
0.694 (1.00) |
1 (1.00) |
||
| TG | 16 (5%) | 307 |
1 (1.00) |
0.0205 (1.00) |
0.34 (1.00) |
0.492 (1.00) |
0.145 (1.00) |
0.0643 (1.00) |
0.0453 (1.00) |
0.021 (1.00) |
| PRB2 | 6 (2%) | 317 |
1 (1.00) |
0.637 (1.00) |
0.0801 (1.00) |
0.851 (1.00) |
0.415 (1.00) |
0.623 (1.00) |
0.529 (1.00) |
0.518 (1.00) |
| R3HDM2 | 4 (1%) | 319 |
0.0124 (1.00) |
0.063 (1.00) |
0.0598 (1.00) |
0.0654 (1.00) |
0.021 (1.00) |
0.00694 (1.00) |
||
| IL32 | 3 (1%) | 320 |
1 (1.00) |
0.742 (1.00) |
1 (1.00) |
1 (1.00) |
0.209 (1.00) |
1 (1.00) |
||
| TMCO2 | 3 (1%) | 320 |
1 (1.00) |
0.416 (1.00) |
0.778 (1.00) |
0.2 (1.00) |
0.5 (1.00) |
0.214 (1.00) |
0.418 (1.00) |
0.778 (1.00) |
| PPTC7 | 3 (1%) | 320 |
0.0791 (1.00) |
0.0627 (1.00) |
0.174 (1.00) |
0.0688 (1.00) |
0.312 (1.00) |
0.292 (1.00) |
||
| MUC7 | 5 (2%) | 318 |
0.731 (1.00) |
0.372 (1.00) |
0.683 (1.00) |
0.26 (1.00) |
0.806 (1.00) |
0.908 (1.00) |
0.869 (1.00) |
0.631 (1.00) |
| LYPD3 | 3 (1%) | 320 |
0.547 (1.00) |
0.416 (1.00) |
0.5 (1.00) |
0.427 (1.00) |
0.653 (1.00) |
0.778 (1.00) |
||
| TMEM90B | 3 (1%) | 320 |
0.135 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
||
| ZNF799 | 5 (2%) | 318 |
0.465 (1.00) |
1 (1.00) |
0.505 (1.00) |
1 (1.00) |
0.598 (1.00) |
0.908 (1.00) |
0.453 (1.00) |
1 (1.00) |
| PPM1D | 5 (2%) | 318 |
1 (1.00) |
1 (1.00) |
0.778 (1.00) |
0.125 (1.00) |
0.695 (1.00) |
0.212 (1.00) |
0.869 (1.00) |
0.328 (1.00) |
| MAP3K3 | 4 (1%) | 319 |
0.061 (1.00) |
0.224 (1.00) |
0.459 (1.00) |
0.0846 (1.00) |
0.117 (1.00) |
0.313 (1.00) |
0.471 (1.00) |
0.557 (1.00) |
| TROAP | 3 (1%) | 320 |
1 (1.00) |
0.742 (1.00) |
0.778 (1.00) |
0.535 (1.00) |
0.751 (1.00) |
0.85 (1.00) |
0.533 (1.00) |
1 (1.00) |
| SYNPO2L | 3 (1%) | 320 |
0.547 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
||
| ATAD2 | 4 (1%) | 319 |
1 (1.00) |
0.823 (1.00) |
0.813 (1.00) |
0.561 (1.00) |
0.694 (1.00) |
1 (1.00) |
||
| KRAS | 3 (1%) | 320 |
1 (1.00) |
0.0627 (1.00) |
0.174 (1.00) |
0.0974 (1.00) |
0.0661 (1.00) |
0.0243 (1.00) |
||
| SLC5A11 | 3 (1%) | 320 |
0.547 (1.00) |
0.742 (1.00) |
0.657 (1.00) |
0.365 (1.00) |
0.312 (1.00) |
0.292 (1.00) |
||
| PRG4 | 4 (1%) | 319 |
0.23 (1.00) |
0.104 (1.00) |
0.0143 (1.00) |
0.028 (1.00) |
0.471 (1.00) |
0.557 (1.00) |
||
| SCUBE2 | 3 (1%) | 320 |
1 (1.00) |
0.281 (1.00) |
0.32 (1.00) |
0.259 (1.00) |
0.653 (1.00) |
0.643 (1.00) |
||
| ZNF479 | 3 (1%) | 320 |
1 (1.00) |
0.742 (1.00) |
1 (1.00) |
1 (1.00) |
0.533 (1.00) |
1 (1.00) |
||
| SREBF2 | 3 (1%) | 320 |
1 (1.00) |
0.742 (1.00) |
0.577 (1.00) |
0.214 (1.00) |
0.418 (1.00) |
0.778 (1.00) |
||
| SLC26A11 | 3 (1%) | 320 |
0.0791 (1.00) |
1 (1.00) |
0.375 (1.00) |
1 (1.00) |
0.776 (1.00) |
0.398 (1.00) |
||
| TSC22D1 | 3 (1%) | 320 |
1 (1.00) |
1 (1.00) |
0.751 (1.00) |
0.85 (1.00) |
1 (1.00) |
0.512 (1.00) |
||
| ANKRD30A | 5 (2%) | 318 |
0.731 (1.00) |
0.462 (1.00) |
0.75 (1.00) |
0.514 (1.00) |
0.743 (1.00) |
0.744 (1.00) |
||
| FAM155A | 4 (1%) | 319 |
0.144 (1.00) |
0.156 (1.00) |
0.778 (1.00) |
0.535 (1.00) |
0.0361 (1.00) |
1 (1.00) |
||
| SLA | 3 (1%) | 320 |
0.547 (1.00) |
0.0627 (1.00) |
0.286 (1.00) |
0.285 (1.00) |
0.174 (1.00) |
0.0974 (1.00) |
0.0661 (1.00) |
0.0243 (1.00) |
| ZFHX3 | 10 (3%) | 313 |
0.171 (1.00) |
0.616 (1.00) |
0.48 (1.00) |
0.632 (1.00) |
0.583 (1.00) |
0.459 (1.00) |
0.617 (1.00) |
0.721 (1.00) |
| ARMCX3 | 3 (1%) | 320 |
0.547 (1.00) |
0.742 (1.00) |
0.657 (1.00) |
0.132 (1.00) |
0.533 (1.00) |
0.2 (1.00) |
||
| SLC25A45 | 3 (1%) | 320 |
0.135 (1.00) |
1 (1.00) |
0.209 (1.00) |
0.292 (1.00) |
||||
| COL5A3 | 6 (2%) | 317 |
0.752 (1.00) |
0.317 (1.00) |
0.778 (1.00) |
0.758 (1.00) |
0.0513 (1.00) |
0.518 (1.00) |
0.875 (1.00) |
0.596 (1.00) |
| CDC27 | 3 (1%) | 320 |
1 (1.00) |
1 (1.00) |
0.375 (1.00) |
0.17 (1.00) |
0.776 (1.00) |
0.398 (1.00) |
P value = 3.62e-38 (Fisher's exact test), Q value = 1e-35
Table S1. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 105 | 52 | 166 |
| BRAF MUTATED | 10 | 30 | 143 |
| BRAF WILD-TYPE | 95 | 22 | 23 |
Figure S1. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'

P value = 9.4e-07 (Fisher's exact test), Q value = 0.00025
Table S2. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 62 | 76 | 79 |
| BRAF MUTATED | 19 | 57 | 42 |
| BRAF WILD-TYPE | 43 | 19 | 37 |
Figure S2. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
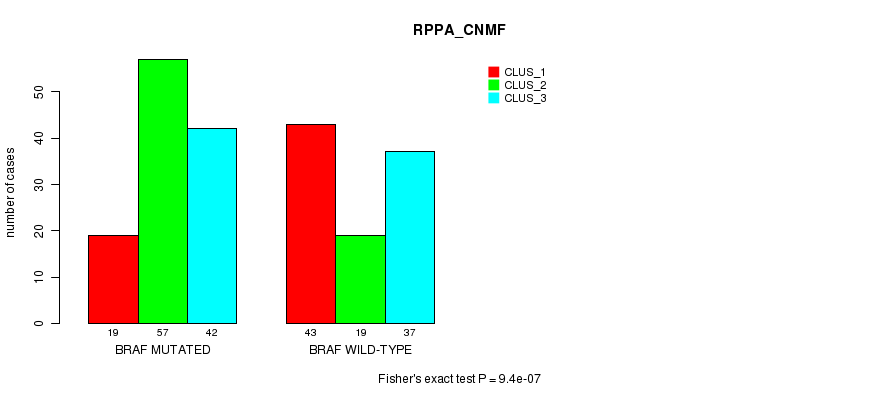
P value = 2.51e-11 (Fisher's exact test), Q value = 6.8e-09
Table S3. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 96 | 32 | 89 |
| BRAF MUTATED | 30 | 16 | 72 |
| BRAF WILD-TYPE | 66 | 16 | 17 |
Figure S3. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'
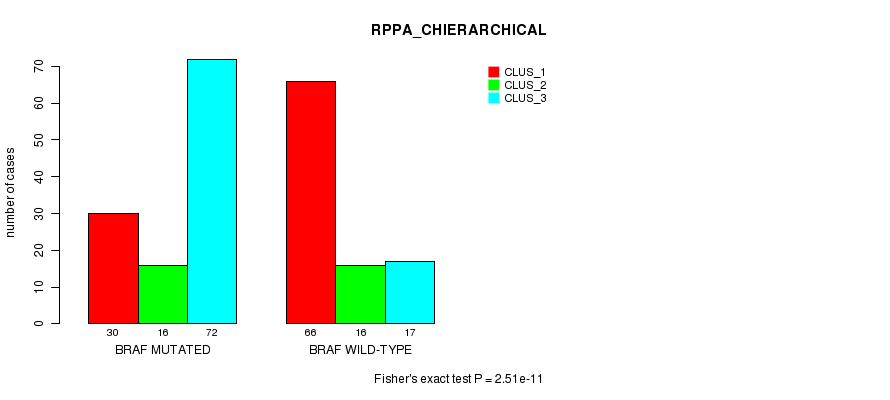
P value = 3.88e-49 (Fisher's exact test), Q value = 1.1e-46
Table S4. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 106 | 39 | 74 | 96 |
| BRAF MUTATED | 5 | 22 | 60 | 91 |
| BRAF WILD-TYPE | 101 | 17 | 14 | 5 |
Figure S4. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'

P value = 7.76e-49 (Fisher's exact test), Q value = 2.2e-46
Table S5. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 97 | 125 | 29 | 64 |
| BRAF MUTATED | 1 | 112 | 17 | 48 |
| BRAF WILD-TYPE | 96 | 13 | 12 | 16 |
Figure S5. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
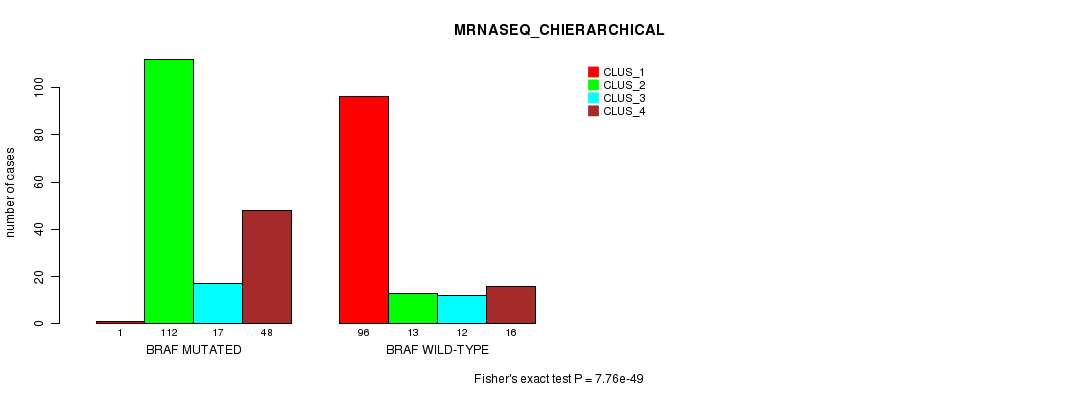
P value = 2.35e-40 (Fisher's exact test), Q value = 6.6e-38
Table S6. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 106 | 114 | 102 |
| BRAF MUTATED | 7 | 98 | 77 |
| BRAF WILD-TYPE | 99 | 16 | 25 |
Figure S6. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
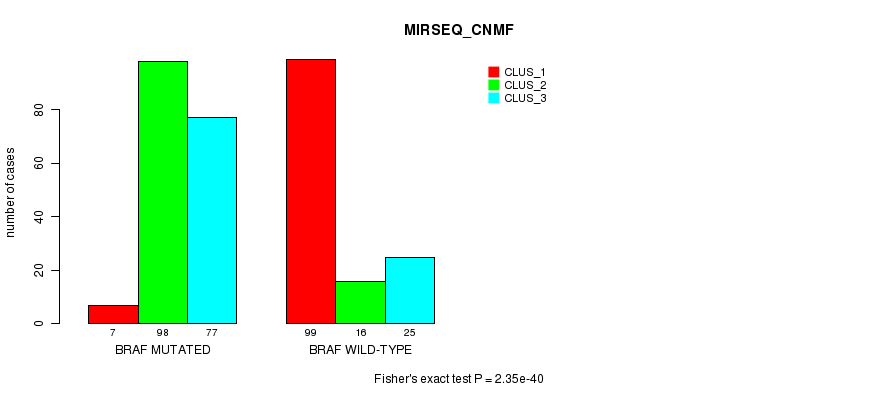
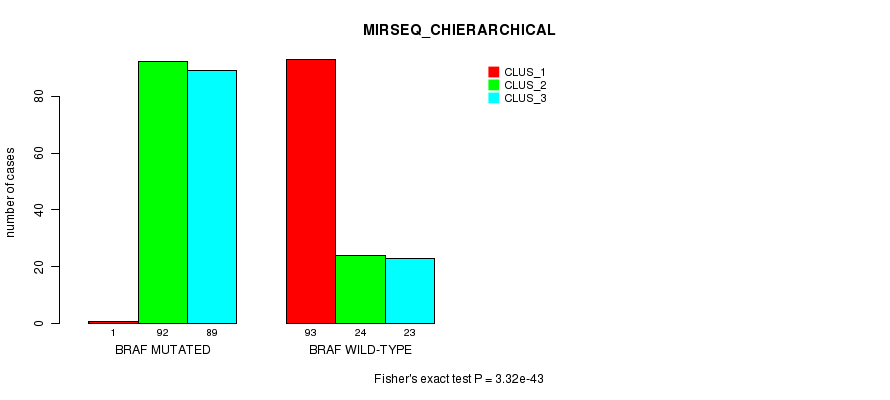
P value = 3.32e-43 (Fisher's exact test), Q value = 9.3e-41
Table S7. Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 116 | 112 |
| BRAF MUTATED | 1 | 92 | 89 |
| BRAF WILD-TYPE | 93 | 24 | 23 |
Figure S7. Get High-res Image Gene #1: 'BRAF MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
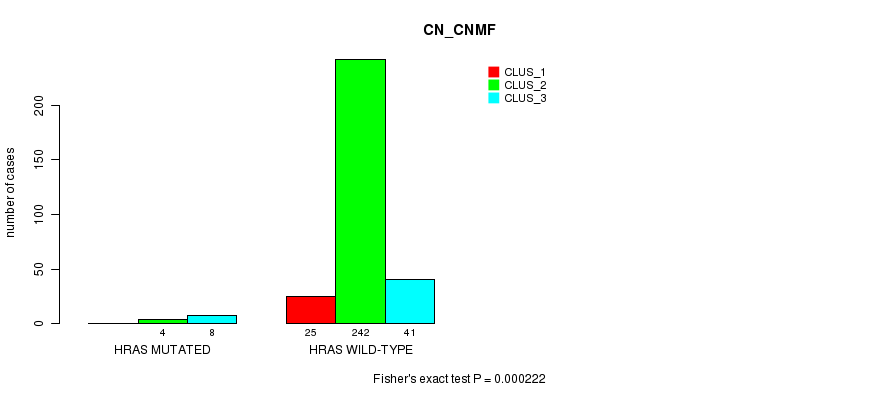
P value = 0.000222 (Fisher's exact test), Q value = 0.059
Table S8. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 25 | 246 | 49 |
| HRAS MUTATED | 0 | 4 | 8 |
| HRAS WILD-TYPE | 25 | 242 | 41 |
Figure S8. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #1: 'CN_CNMF'
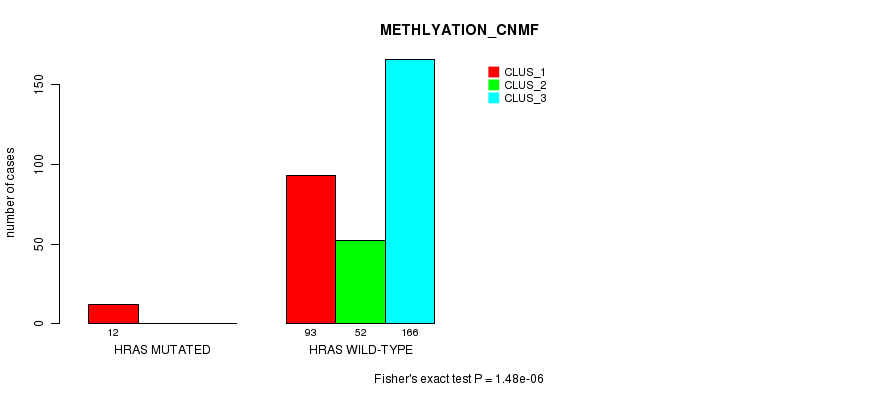
P value = 1.48e-06 (Fisher's exact test), Q value = 4e-04
Table S9. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 105 | 52 | 166 |
| HRAS MUTATED | 12 | 0 | 0 |
| HRAS WILD-TYPE | 93 | 52 | 166 |
Figure S9. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
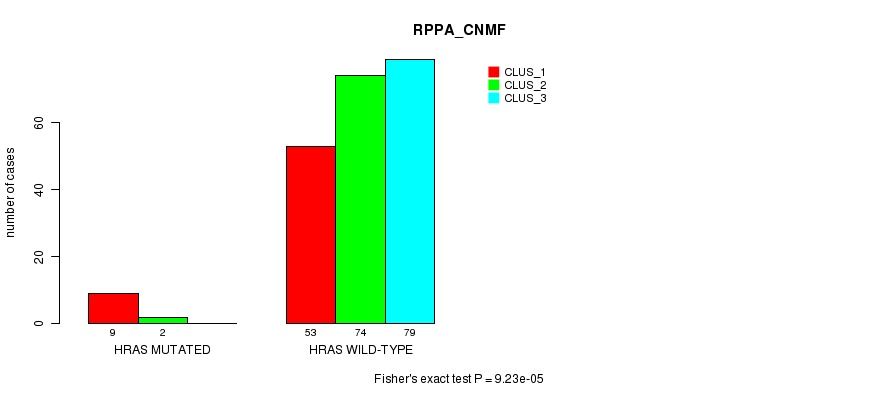
P value = 9.23e-05 (Fisher's exact test), Q value = 0.024
Table S10. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 62 | 76 | 79 |
| HRAS MUTATED | 9 | 2 | 0 |
| HRAS WILD-TYPE | 53 | 74 | 79 |
Figure S10. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #3: 'RPPA_CNMF'
P value = 1.71e-05 (Fisher's exact test), Q value = 0.0045
Table S11. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 106 | 39 | 74 | 96 |
| HRAS MUTATED | 12 | 0 | 0 | 0 |
| HRAS WILD-TYPE | 94 | 39 | 74 | 96 |
Figure S11. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'

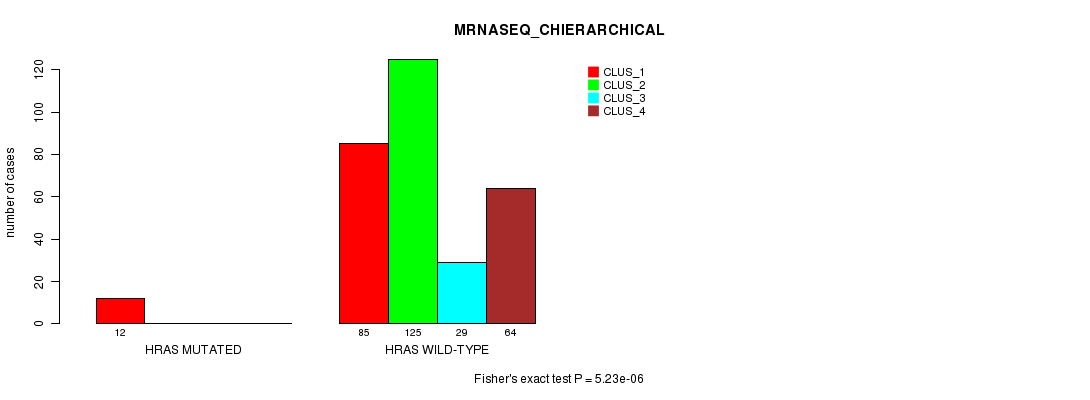
P value = 5.23e-06 (Fisher's exact test), Q value = 0.0014
Table S12. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 97 | 125 | 29 | 64 |
| HRAS MUTATED | 12 | 0 | 0 | 0 |
| HRAS WILD-TYPE | 85 | 125 | 29 | 64 |
Figure S12. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
P value = 1.69e-06 (Fisher's exact test), Q value = 0.00045
Table S13. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 106 | 114 | 102 |
| HRAS MUTATED | 12 | 0 | 0 |
| HRAS WILD-TYPE | 94 | 114 | 102 |
Figure S13. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'

P value = 9.25e-06 (Fisher's exact test), Q value = 0.0025
Table S14. Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 116 | 112 |
| HRAS MUTATED | 11 | 0 | 1 |
| HRAS WILD-TYPE | 83 | 116 | 111 |
Figure S14. Get High-res Image Gene #3: 'HRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'

P value = 6.06e-14 (Fisher's exact test), Q value = 1.7e-11
Table S15. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 105 | 52 | 166 |
| NRAS MUTATED | 26 | 0 | 0 |
| NRAS WILD-TYPE | 79 | 52 | 166 |
Figure S15. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #2: 'METHLYATION_CNMF'

P value = 0.000583 (Fisher's exact test), Q value = 0.15
Table S16. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 96 | 32 | 89 |
| NRAS MUTATED | 9 | 5 | 0 |
| NRAS WILD-TYPE | 87 | 27 | 89 |
Figure S16. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #4: 'RPPA_CHIERARCHICAL'

P value = 1.63e-12 (Fisher's exact test), Q value = 4.5e-10
Table S17. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 106 | 39 | 74 | 96 |
| NRAS MUTATED | 26 | 0 | 0 | 0 |
| NRAS WILD-TYPE | 80 | 39 | 74 | 96 |
Figure S17. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #5: 'MRNASEQ_CNMF'

P value = 1.11e-13 (Fisher's exact test), Q value = 3.1e-11
Table S18. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 97 | 125 | 29 | 64 |
| NRAS MUTATED | 26 | 0 | 0 | 0 |
| NRAS WILD-TYPE | 71 | 125 | 29 | 64 |
Figure S18. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #6: 'MRNASEQ_CHIERARCHICAL'

P value = 1.87e-12 (Fisher's exact test), Q value = 5.1e-10
Table S19. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 106 | 114 | 102 |
| NRAS MUTATED | 25 | 0 | 1 |
| NRAS WILD-TYPE | 81 | 114 | 101 |
Figure S19. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #7: 'MIRSEQ_CNMF'

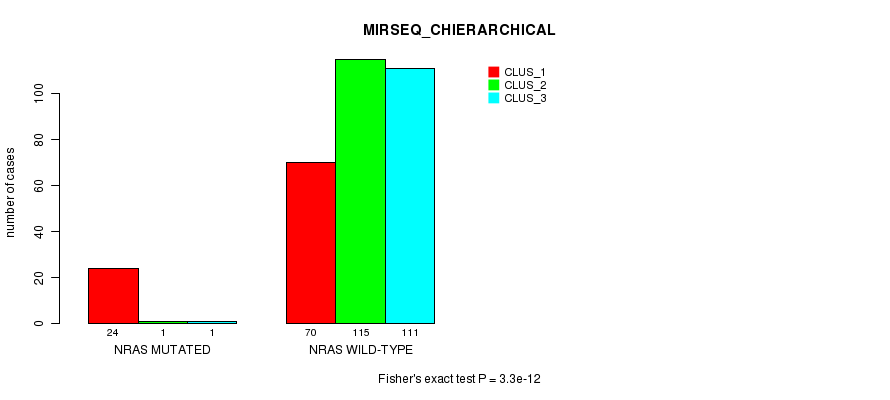
P value = 3.3e-12 (Fisher's exact test), Q value = 9e-10
Table S20. Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 94 | 116 | 112 |
| NRAS MUTATED | 24 | 1 | 1 |
| NRAS WILD-TYPE | 70 | 115 | 111 |
Figure S20. Get High-res Image Gene #4: 'NRAS MUTATION STATUS' versus Clinical Feature #8: 'MIRSEQ_CHIERARCHICAL'
-
Mutation data file = THCA-TP.mutsig.cluster.txt
-
Molecular subtypes file = THCA-TP.transferedmergedcluster.txt
-
Number of patients = 323
-
Number of significantly mutated genes = 41
-
Number of Molecular subtypes = 8
-
Exclude genes that fewer than K tumors have mutations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
This is an experimental feature. The full results of the analysis summarized in this report can be downloaded from the TCGA Data Coordination Center.