This pipeline computes the correlation between significant arm-level copy number variations (cnvs) and selected clinical features.
Testing the association between copy number variation 59 arm-level events and 10 clinical features across 64 patients, 7 significant findings detected with Q value < 0.25.
-
5P GAIN MUTATION ANALYSIS cnv correlated to 'Time to Death'.
-
5Q GAIN MUTATION ANALYSIS cnv correlated to 'Time to Death'.
-
16P LOSS MUTATION ANALYSIS cnv correlated to 'Time to Death', 'NEOPLASM.DISEASESTAGE', and 'PATHOLOGY.N.STAGE'.
-
16Q LOSS MUTATION ANALYSIS cnv correlated to 'Time to Death' and 'PATHOLOGY.N.STAGE'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 59 arm-level events and 10 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 7 significant findings detected.
|
Clinical Features |
Time to Death |
AGE |
NEOPLASM DISEASESTAGE |
PATHOLOGY T STAGE |
PATHOLOGY N STAGE |
PATHOLOGY M STAGE |
GENDER |
KARNOFSKY PERFORMANCE SCORE |
NUMBERPACKYEARSSMOKED | YEAROFTOBACCOSMOKINGONSET | ||
| nCNV (%) | nWild-Type | logrank test | t-test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | t-test | t-test | t-test | |
| 16P LOSS MUTATION ANALYSIS | 4 (6%) | 60 |
2.71e-10 (1.33e-07) |
0.466 (1.00) |
0.000464 (0.226) |
0.00763 (1.00) |
3.36e-05 (0.0164) |
1 (1.00) |
||||
| 16Q LOSS MUTATION ANALYSIS | 5 (8%) | 59 |
2.35e-10 (1.15e-07) |
0.199 (1.00) |
0.00495 (1.00) |
0.0371 (1.00) |
0.000165 (0.0804) |
0.64 (1.00) |
||||
| 5P GAIN MUTATION ANALYSIS | 7 (11%) | 57 |
0.00042 (0.205) |
0.105 (1.00) |
0.000926 (0.446) |
0.278 (1.00) |
0.0874 (1.00) |
0.00903 (1.00) |
0.695 (1.00) |
|||
| 5Q GAIN MUTATION ANALYSIS | 7 (11%) | 57 |
0.00042 (0.205) |
0.105 (1.00) |
0.000926 (0.446) |
0.278 (1.00) |
0.0874 (1.00) |
0.00903 (1.00) |
0.695 (1.00) |
|||
| 3P GAIN MUTATION ANALYSIS | 8 (12%) | 56 |
0.00254 (1.00) |
0.0657 (1.00) |
0.0146 (1.00) |
0.46 (1.00) |
0.0327 (1.00) |
0.269 (1.00) |
0.135 (1.00) |
|||
| 3Q GAIN MUTATION ANALYSIS | 8 (12%) | 56 |
0.00254 (1.00) |
0.0657 (1.00) |
0.0146 (1.00) |
0.46 (1.00) |
0.0327 (1.00) |
0.269 (1.00) |
0.135 (1.00) |
|||
| 4P GAIN MUTATION ANALYSIS | 19 (30%) | 45 |
0.26 (1.00) |
0.575 (1.00) |
0.141 (1.00) |
0.941 (1.00) |
1 (1.00) |
0.127 (1.00) |
0.577 (1.00) |
0.863 (1.00) |
0.073 (1.00) |
0.0084 (1.00) |
| 4Q GAIN MUTATION ANALYSIS | 18 (28%) | 46 |
0.26 (1.00) |
0.902 (1.00) |
0.144 (1.00) |
1 (1.00) |
1 (1.00) |
0.127 (1.00) |
0.776 (1.00) |
0.863 (1.00) |
0.073 (1.00) |
0.0084 (1.00) |
| 7P GAIN MUTATION ANALYSIS | 23 (36%) | 41 |
0.0188 (1.00) |
0.264 (1.00) |
0.106 (1.00) |
0.547 (1.00) |
0.155 (1.00) |
0.107 (1.00) |
0.181 (1.00) |
0.863 (1.00) |
0.551 (1.00) |
0.0341 (1.00) |
| 7Q GAIN MUTATION ANALYSIS | 23 (36%) | 41 |
0.0188 (1.00) |
0.264 (1.00) |
0.106 (1.00) |
0.547 (1.00) |
0.155 (1.00) |
0.107 (1.00) |
0.181 (1.00) |
0.863 (1.00) |
0.551 (1.00) |
0.0341 (1.00) |
| 8P GAIN MUTATION ANALYSIS | 15 (23%) | 49 |
0.606 (1.00) |
0.202 (1.00) |
0.613 (1.00) |
0.661 (1.00) |
0.639 (1.00) |
0.482 (1.00) |
0.368 (1.00) |
0.863 (1.00) |
0.83 (1.00) |
|
| 8Q GAIN MUTATION ANALYSIS | 16 (25%) | 48 |
0.606 (1.00) |
0.211 (1.00) |
0.329 (1.00) |
0.671 (1.00) |
0.639 (1.00) |
0.0699 (1.00) |
0.243 (1.00) |
0.863 (1.00) |
0.83 (1.00) |
|
| 9P GAIN MUTATION ANALYSIS | 10 (16%) | 54 |
0.404 (1.00) |
0.958 (1.00) |
0.0852 (1.00) |
0.163 (1.00) |
0.306 (1.00) |
1 (1.00) |
1 (1.00) |
0.566 (1.00) |
||
| 9Q GAIN MUTATION ANALYSIS | 10 (16%) | 54 |
0.404 (1.00) |
0.958 (1.00) |
0.0852 (1.00) |
0.163 (1.00) |
0.306 (1.00) |
1 (1.00) |
1 (1.00) |
0.566 (1.00) |
||
| 11P GAIN MUTATION ANALYSIS | 14 (22%) | 50 |
0.575 (1.00) |
0.846 (1.00) |
0.608 (1.00) |
0.697 (1.00) |
0.617 (1.00) |
0.367 (1.00) |
1 (1.00) |
0.073 (1.00) |
0.0084 (1.00) |
|
| 11Q GAIN MUTATION ANALYSIS | 15 (23%) | 49 |
0.575 (1.00) |
0.866 (1.00) |
0.279 (1.00) |
0.708 (1.00) |
0.617 (1.00) |
0.0529 (1.00) |
0.765 (1.00) |
0.073 (1.00) |
0.0084 (1.00) |
|
| 12P GAIN MUTATION ANALYSIS | 17 (27%) | 47 |
0.714 (1.00) |
0.0319 (1.00) |
0.385 (1.00) |
0.881 (1.00) |
1 (1.00) |
0.0939 (1.00) |
0.397 (1.00) |
0.863 (1.00) |
0.83 (1.00) |
|
| 12Q GAIN MUTATION ANALYSIS | 17 (27%) | 47 |
0.167 (1.00) |
0.0956 (1.00) |
0.138 (1.00) |
0.938 (1.00) |
0.315 (1.00) |
0.0939 (1.00) |
0.397 (1.00) |
0.863 (1.00) |
0.473 (1.00) |
0.0084 (1.00) |
| 14Q GAIN MUTATION ANALYSIS | 17 (27%) | 47 |
0.806 (1.00) |
0.276 (1.00) |
0.256 (1.00) |
0.356 (1.00) |
0.651 (1.00) |
0.671 (1.00) |
0.778 (1.00) |
0.863 (1.00) |
0.473 (1.00) |
0.0084 (1.00) |
| 15Q GAIN MUTATION ANALYSIS | 18 (28%) | 46 |
0.344 (1.00) |
0.821 (1.00) |
0.743 (1.00) |
0.735 (1.00) |
1 (1.00) |
0.695 (1.00) |
1 (1.00) |
0.863 (1.00) |
0.368 (1.00) |
0.161 (1.00) |
| 16P GAIN MUTATION ANALYSIS | 18 (28%) | 46 |
0.501 (1.00) |
0.693 (1.00) |
0.495 (1.00) |
0.444 (1.00) |
0.641 (1.00) |
0.695 (1.00) |
1 (1.00) |
0.863 (1.00) |
0.073 (1.00) |
0.0084 (1.00) |
| 16Q GAIN MUTATION ANALYSIS | 18 (28%) | 46 |
0.501 (1.00) |
0.693 (1.00) |
0.495 (1.00) |
0.444 (1.00) |
0.641 (1.00) |
0.695 (1.00) |
1 (1.00) |
0.863 (1.00) |
0.073 (1.00) |
0.0084 (1.00) |
| 18P GAIN MUTATION ANALYSIS | 14 (22%) | 50 |
0.591 (1.00) |
0.427 (1.00) |
0.399 (1.00) |
0.648 (1.00) |
0.313 (1.00) |
0.337 (1.00) |
0.537 (1.00) |
0.745 (1.00) |
||
| 18Q GAIN MUTATION ANALYSIS | 13 (20%) | 51 |
0.699 (1.00) |
0.755 (1.00) |
0.497 (1.00) |
0.339 (1.00) |
0.313 (1.00) |
0.337 (1.00) |
0.544 (1.00) |
0.745 (1.00) |
||
| 19P GAIN MUTATION ANALYSIS | 17 (27%) | 47 |
0.000914 (0.442) |
0.0575 (1.00) |
0.0158 (1.00) |
0.527 (1.00) |
0.00246 (1.00) |
0.0699 (1.00) |
0.397 (1.00) |
0.551 (1.00) |
0.0341 (1.00) |
|
| 19Q GAIN MUTATION ANALYSIS | 16 (25%) | 48 |
0.0137 (1.00) |
0.0274 (1.00) |
0.107 (1.00) |
0.719 (1.00) |
0.027 (1.00) |
0.0366 (1.00) |
0.561 (1.00) |
0.368 (1.00) |
0.161 (1.00) |
|
| 20P GAIN MUTATION ANALYSIS | 18 (28%) | 46 |
0.353 (1.00) |
0.261 (1.00) |
0.428 (1.00) |
0.941 (1.00) |
0.35 (1.00) |
0.695 (1.00) |
0.776 (1.00) |
0.863 (1.00) |
0.473 (1.00) |
0.0084 (1.00) |
| 20Q GAIN MUTATION ANALYSIS | 18 (28%) | 46 |
0.288 (1.00) |
0.219 (1.00) |
0.144 (1.00) |
1 (1.00) |
0.33 (1.00) |
0.695 (1.00) |
0.776 (1.00) |
0.863 (1.00) |
0.073 (1.00) |
0.0084 (1.00) |
| 21Q GAIN MUTATION ANALYSIS | 3 (5%) | 61 |
0.604 (1.00) |
0.853 (1.00) |
1 (1.00) |
1 (1.00) |
0.555 (1.00) |
|||||
| 22Q GAIN MUTATION ANALYSIS | 17 (27%) | 47 |
0.0287 (1.00) |
0.733 (1.00) |
0.521 (1.00) |
0.527 (1.00) |
0.166 (1.00) |
0.34 (1.00) |
0.397 (1.00) |
0.551 (1.00) |
0.0341 (1.00) |
|
| XQ GAIN MUTATION ANALYSIS | 10 (16%) | 54 |
0.335 (1.00) |
0.159 (1.00) |
0.191 (1.00) |
0.756 (1.00) |
1 (1.00) |
0.167 (1.00) |
0.292 (1.00) |
|||
| 1P LOSS MUTATION ANALYSIS | 47 (73%) | 17 |
0.447 (1.00) |
0.0592 (1.00) |
0.0376 (1.00) |
0.148 (1.00) |
1 (1.00) |
0.0671 (1.00) |
0.155 (1.00) |
0.374 (1.00) |
0.661 (1.00) |
0.308 (1.00) |
| 1Q LOSS MUTATION ANALYSIS | 47 (73%) | 17 |
0.816 (1.00) |
0.063 (1.00) |
0.0461 (1.00) |
0.095 (1.00) |
1 (1.00) |
0.0964 (1.00) |
0.155 (1.00) |
0.374 (1.00) |
0.164 (1.00) |
0.308 (1.00) |
| 2P LOSS MUTATION ANALYSIS | 42 (66%) | 22 |
0.488 (1.00) |
0.0564 (1.00) |
0.241 (1.00) |
0.197 (1.00) |
1 (1.00) |
0.372 (1.00) |
0.188 (1.00) |
0.374 (1.00) |
0.164 (1.00) |
0.308 (1.00) |
| 2Q LOSS MUTATION ANALYSIS | 42 (66%) | 22 |
0.488 (1.00) |
0.0564 (1.00) |
0.241 (1.00) |
0.197 (1.00) |
1 (1.00) |
0.372 (1.00) |
0.188 (1.00) |
0.374 (1.00) |
0.164 (1.00) |
0.308 (1.00) |
| 3P LOSS MUTATION ANALYSIS | 7 (11%) | 57 |
0.353 (1.00) |
0.195 (1.00) |
0.218 (1.00) |
0.083 (1.00) |
1 (1.00) |
1 (1.00) |
0.417 (1.00) |
|||
| 3Q LOSS MUTATION ANALYSIS | 6 (9%) | 58 |
0.409 (1.00) |
0.37 (1.00) |
0.337 (1.00) |
0.175 (1.00) |
1 (1.00) |
0.199 (1.00) |
||||
| 5P LOSS MUTATION ANALYSIS | 6 (9%) | 58 |
0.617 (1.00) |
0.000736 (0.357) |
0.166 (1.00) |
0.15 (1.00) |
0.387 (1.00) |
0.455 (1.00) |
0.671 (1.00) |
|||
| 5Q LOSS MUTATION ANALYSIS | 6 (9%) | 58 |
0.617 (1.00) |
0.000736 (0.357) |
0.166 (1.00) |
0.15 (1.00) |
0.387 (1.00) |
0.455 (1.00) |
0.671 (1.00) |
|||
| 6P LOSS MUTATION ANALYSIS | 46 (72%) | 18 |
0.178 (1.00) |
0.0745 (1.00) |
0.146 (1.00) |
0.0887 (1.00) |
1 (1.00) |
0.21 (1.00) |
0.0972 (1.00) |
0.374 (1.00) |
0.164 (1.00) |
0.308 (1.00) |
| 6Q LOSS MUTATION ANALYSIS | 47 (73%) | 17 |
0.178 (1.00) |
0.0843 (1.00) |
0.149 (1.00) |
0.095 (1.00) |
0.571 (1.00) |
0.21 (1.00) |
0.155 (1.00) |
0.374 (1.00) |
0.164 (1.00) |
0.308 (1.00) |
| 8P LOSS MUTATION ANALYSIS | 10 (16%) | 54 |
0.233 (1.00) |
0.888 (1.00) |
0.582 (1.00) |
0.316 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
|||
| 8Q LOSS MUTATION ANALYSIS | 9 (14%) | 55 |
0.271 (1.00) |
0.767 (1.00) |
0.684 (1.00) |
0.435 (1.00) |
1 (1.00) |
1 (1.00) |
0.728 (1.00) |
|||
| 9P LOSS MUTATION ANALYSIS | 6 (9%) | 58 |
0.0342 (1.00) |
0.16 (1.00) |
0.581 (1.00) |
0.424 (1.00) |
0.387 (1.00) |
1 (1.00) |
0.391 (1.00) |
|||
| 9Q LOSS MUTATION ANALYSIS | 7 (11%) | 57 |
0.0882 (1.00) |
0.341 (1.00) |
0.29 (1.00) |
0.237 (1.00) |
0.461 (1.00) |
1 (1.00) |
0.695 (1.00) |
|||
| 10P LOSS MUTATION ANALYSIS | 45 (70%) | 19 |
0.192 (1.00) |
0.432 (1.00) |
0.422 (1.00) |
0.313 (1.00) |
1 (1.00) |
0.0233 (1.00) |
0.577 (1.00) |
0.107 (1.00) |
||
| 10Q LOSS MUTATION ANALYSIS | 45 (70%) | 19 |
0.192 (1.00) |
0.432 (1.00) |
0.422 (1.00) |
0.313 (1.00) |
1 (1.00) |
0.0233 (1.00) |
0.577 (1.00) |
0.107 (1.00) |
||
| 11P LOSS MUTATION ANALYSIS | 7 (11%) | 57 |
0.391 (1.00) |
0.0226 (1.00) |
0.0877 (1.00) |
0.0502 (1.00) |
0.461 (1.00) |
1 (1.00) |
0.695 (1.00) |
|||
| 11Q LOSS MUTATION ANALYSIS | 7 (11%) | 57 |
0.391 (1.00) |
0.0226 (1.00) |
0.0877 (1.00) |
0.0502 (1.00) |
0.461 (1.00) |
1 (1.00) |
0.695 (1.00) |
|||
| 13Q LOSS MUTATION ANALYSIS | 40 (62%) | 24 |
0.064 (1.00) |
0.01 (1.00) |
0.341 (1.00) |
0.244 (1.00) |
0.153 (1.00) |
0.628 (1.00) |
0.112 (1.00) |
0.81 (1.00) |
0.0556 (1.00) |
1 (1.00) |
| 17P LOSS MUTATION ANALYSIS | 46 (72%) | 18 |
0.764 (1.00) |
0.0449 (1.00) |
0.0305 (1.00) |
0.0228 (1.00) |
1 (1.00) |
0.178 (1.00) |
0.0972 (1.00) |
0.374 (1.00) |
||
| 17Q LOSS MUTATION ANALYSIS | 46 (72%) | 18 |
0.764 (1.00) |
0.0449 (1.00) |
0.0305 (1.00) |
0.0228 (1.00) |
1 (1.00) |
0.178 (1.00) |
0.0972 (1.00) |
0.374 (1.00) |
||
| 18P LOSS MUTATION ANALYSIS | 6 (9%) | 58 |
0.685 (1.00) |
0.97 (1.00) |
0.269 (1.00) |
0.582 (1.00) |
0.125 (1.00) |
0.638 (1.00) |
0.199 (1.00) |
|||
| 18Q LOSS MUTATION ANALYSIS | 8 (12%) | 56 |
0.18 (1.00) |
0.595 (1.00) |
0.0854 (1.00) |
0.572 (1.00) |
0.166 (1.00) |
0.638 (1.00) |
0.245 (1.00) |
|||
| 20P LOSS MUTATION ANALYSIS | 4 (6%) | 60 |
0.442 (1.00) |
0.172 (1.00) |
0.162 (1.00) |
0.11 (1.00) |
0.304 (1.00) |
1 (1.00) |
0.64 (1.00) |
|||
| 20Q LOSS MUTATION ANALYSIS | 3 (5%) | 61 |
0.313 (1.00) |
0.423 (1.00) |
0.0432 (1.00) |
0.0274 (1.00) |
0.304 (1.00) |
1 (1.00) |
1 (1.00) |
|||
| 21Q LOSS MUTATION ANALYSIS | 32 (50%) | 32 |
0.0913 (1.00) |
0.897 (1.00) |
0.717 (1.00) |
0.546 (1.00) |
0.0634 (1.00) |
0.1 (1.00) |
1 (1.00) |
0.278 (1.00) |
0.28 (1.00) |
0.101 (1.00) |
| 22Q LOSS MUTATION ANALYSIS | 8 (12%) | 56 |
0.18 (1.00) |
0.542 (1.00) |
0.12 (1.00) |
0.716 (1.00) |
0.166 (1.00) |
0.342 (1.00) |
0.048 (1.00) |
|||
| XQ LOSS MUTATION ANALYSIS | 40 (62%) | 24 |
0.31 (1.00) |
0.35 (1.00) |
0.233 (1.00) |
0.244 (1.00) |
1 (1.00) |
0.141 (1.00) |
0.291 (1.00) |
0.0791 (1.00) |
0.0398 (1.00) |
0.277 (1.00) |
P value = 0.00042 (logrank test), Q value = 0.21
Table S1. Gene #5: '5P GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 63 | 7 | 0.6 - 151.9 (63.9) |
| 5P GAIN MUTATED | 7 | 3 | 0.6 - 137.1 (28.1) |
| 5P GAIN WILD-TYPE | 56 | 4 | 2.5 - 151.9 (65.9) |
Figure S1. Get High-res Image Gene #5: '5P GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
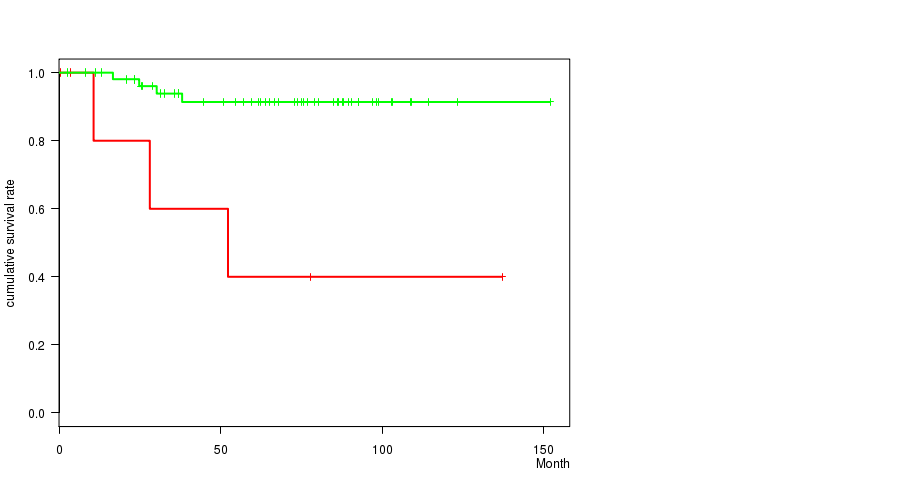
P value = 0.00042 (logrank test), Q value = 0.21
Table S2. Gene #6: '5Q GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 63 | 7 | 0.6 - 151.9 (63.9) |
| 5Q GAIN MUTATED | 7 | 3 | 0.6 - 137.1 (28.1) |
| 5Q GAIN WILD-TYPE | 56 | 4 | 2.5 - 151.9 (65.9) |
Figure S2. Get High-res Image Gene #6: '5Q GAIN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
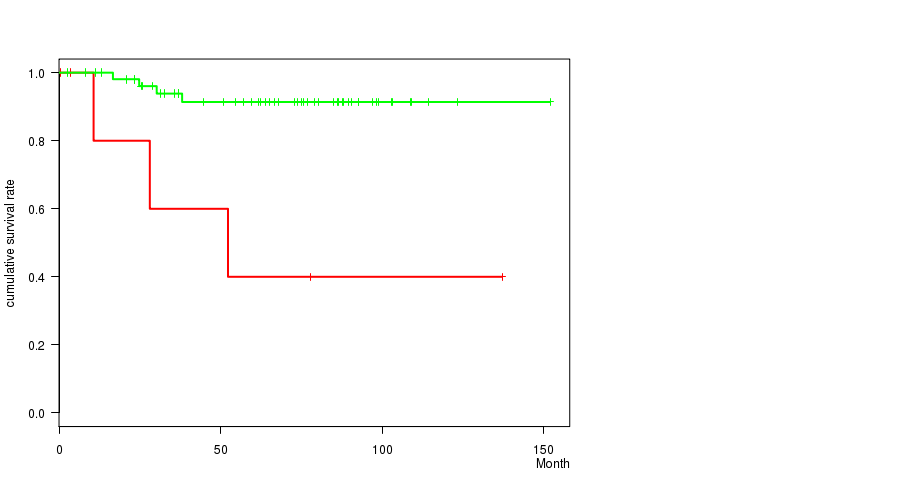
P value = 2.71e-10 (logrank test), Q value = 1.3e-07
Table S3. Gene #49: '16P LOSS MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 63 | 7 | 0.6 - 151.9 (63.9) |
| 16P LOSS MUTATED | 4 | 3 | 0.6 - 30.2 (22.4) |
| 16P LOSS WILD-TYPE | 59 | 4 | 2.5 - 151.9 (66.5) |
Figure S3. Get High-res Image Gene #49: '16P LOSS MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
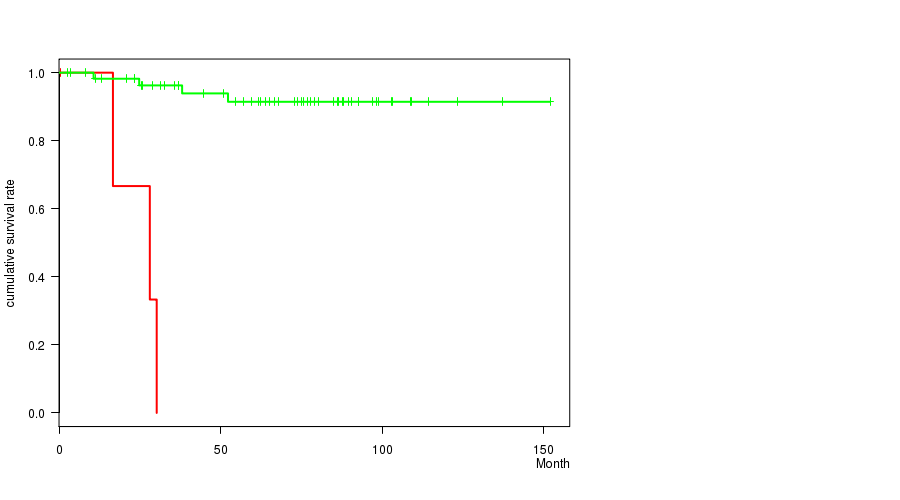
P value = 0.000464 (Fisher's exact test), Q value = 0.23
Table S4. Gene #49: '16P LOSS MUTATION STATUS' versus Clinical Feature #3: 'NEOPLASM.DISEASESTAGE'
| nPatients | STAGE I | STAGE II | STAGE III | STAGE IV |
|---|---|---|---|---|
| ALL | 21 | 23 | 14 | 6 |
| 16P LOSS MUTATED | 0 | 0 | 1 | 3 |
| 16P LOSS WILD-TYPE | 21 | 23 | 13 | 3 |
Figure S4. Get High-res Image Gene #49: '16P LOSS MUTATION STATUS' versus Clinical Feature #3: 'NEOPLASM.DISEASESTAGE'
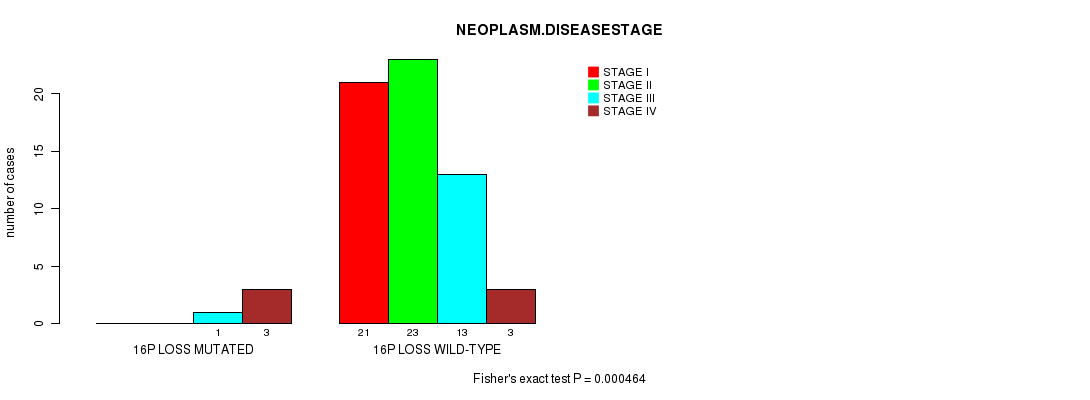
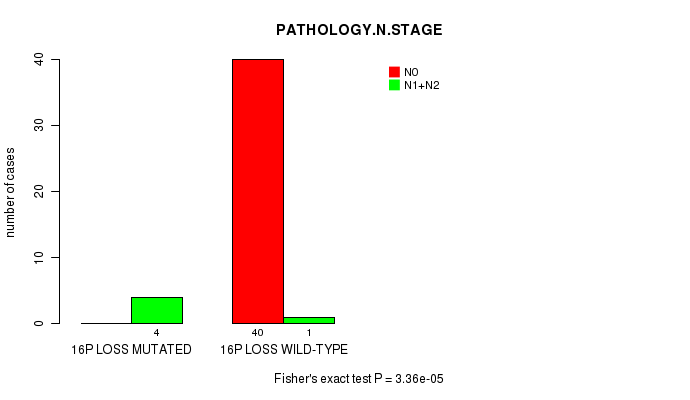
P value = 3.36e-05 (Fisher's exact test), Q value = 0.016
Table S5. Gene #49: '16P LOSS MUTATION STATUS' versus Clinical Feature #5: 'PATHOLOGY.N.STAGE'
| nPatients | N0 | N1+N2 |
|---|---|---|
| ALL | 40 | 5 |
| 16P LOSS MUTATED | 0 | 4 |
| 16P LOSS WILD-TYPE | 40 | 1 |
Figure S5. Get High-res Image Gene #49: '16P LOSS MUTATION STATUS' versus Clinical Feature #5: 'PATHOLOGY.N.STAGE'
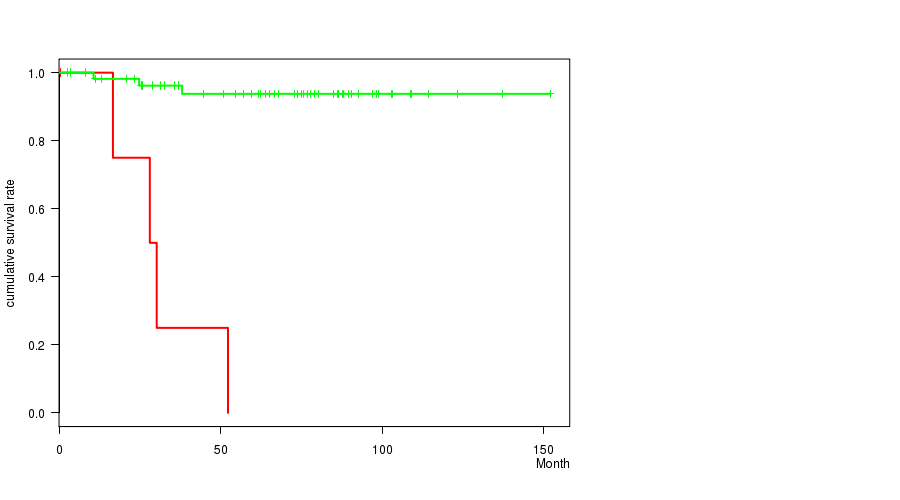
P value = 2.35e-10 (logrank test), Q value = 1.2e-07
Table S6. Gene #50: '16Q LOSS MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 63 | 7 | 0.6 - 151.9 (63.9) |
| 16Q LOSS MUTATED | 5 | 4 | 0.6 - 52.3 (28.1) |
| 16Q LOSS WILD-TYPE | 58 | 3 | 2.5 - 151.9 (67.2) |
Figure S6. Get High-res Image Gene #50: '16Q LOSS MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
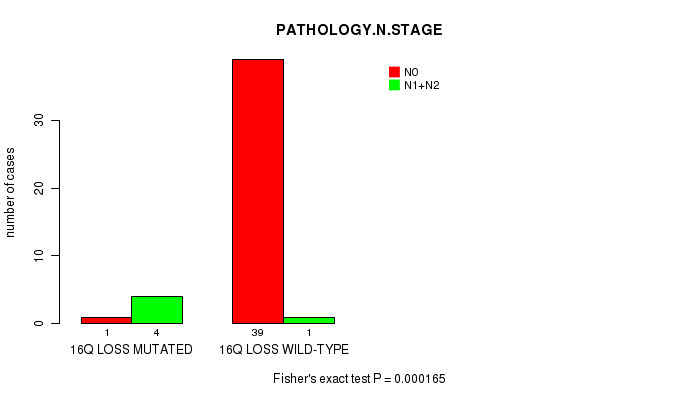
P value = 0.000165 (Fisher's exact test), Q value = 0.08
Table S7. Gene #50: '16Q LOSS MUTATION STATUS' versus Clinical Feature #5: 'PATHOLOGY.N.STAGE'
| nPatients | N0 | N1+N2 |
|---|---|---|
| ALL | 40 | 5 |
| 16Q LOSS MUTATED | 1 | 4 |
| 16Q LOSS WILD-TYPE | 39 | 1 |
Figure S7. Get High-res Image Gene #50: '16Q LOSS MUTATION STATUS' versus Clinical Feature #5: 'PATHOLOGY.N.STAGE'
-
Copy number data file = transformed.cor.cli.txt
-
Clinical data file = KICH-TP.clin.merged.picked.txt
-
Number of patients = 64
-
Number of significantly arm-level cnvs = 59
-
Number of selected clinical features = 10
-
Exclude regions that fewer than K tumors have mutations, K = 3
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For continuous numerical clinical features, two-tailed Student's t test with unequal variance (Lehmann and Romano 2005) was applied to compare the clinical values between tumors with and without gene mutations using 't.test' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.