This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 45 focal events and 8 molecular subtypes across 90 patients, 27 significant findings detected with P value < 0.05 and Q value < 0.25.
-
amp_1q22 cnv correlated to 'MRNASEQ_CHIERARCHICAL'.
-
amp_4q35.1 cnv correlated to 'CN_CNMF'.
-
amp_5p15.33 cnv correlated to 'CN_CNMF'.
-
amp_5q35.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
amp_9q31.3 cnv correlated to 'CN_CNMF'.
-
amp_12q14.1 cnv correlated to 'CN_CNMF'.
-
amp_14q11.2 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_MATURE_CNMF'.
-
amp_16p13.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
amp_16q22.1 cnv correlated to 'CN_CNMF'.
-
amp_16q24.2 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
amp_xp11.22 cnv correlated to 'CN_CNMF'.
-
amp_xq28 cnv correlated to 'CN_CNMF'.
-
del_9p23 cnv correlated to 'MRNASEQ_CNMF'.
-
del_9p21.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_11q14.1 cnv correlated to 'CN_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 45 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 27 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Chi-square test | Chi-square test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| amp 14q11 2 | 27 (30%) | 63 |
8.67e-05 (0.0301) |
0.018 (1.00) |
3.8e-05 (0.0134) |
5.99e-07 (0.000214) |
0.000295 (0.0995) |
0.00209 (0.67) |
0.000226 (0.0768) |
0.000753 (0.251) |
| amp 5q35 3 | 63 (70%) | 27 |
8.49e-08 (3.06e-05) |
0.0017 (0.55) |
2.2e-06 (0.000781) |
1.24e-05 (0.00437) |
0.00626 (1.00) |
0.311 (1.00) |
0.0241 (1.00) |
0.631 (1.00) |
| amp 16p13 3 | 50 (56%) | 40 |
1.34e-07 (4.78e-05) |
0.258 (1.00) |
0.000195 (0.0666) |
0.000585 (0.196) |
0.0744 (1.00) |
0.554 (1.00) |
0.0161 (1.00) |
0.841 (1.00) |
| amp 16q24 2 | 49 (54%) | 41 |
9.86e-08 (3.54e-05) |
0.0299 (1.00) |
0.000271 (0.0917) |
3.91e-05 (0.0137) |
0.0647 (1.00) |
0.249 (1.00) |
0.0292 (1.00) |
0.54 (1.00) |
| del 9p21 3 | 27 (30%) | 63 |
0.000657 (0.22) |
0.00316 (0.99) |
9.71e-06 (0.00344) |
0.00022 (0.0752) |
0.0169 (1.00) |
0.204 (1.00) |
0.0766 (1.00) |
0.213 (1.00) |
| amp 1q22 | 17 (19%) | 73 |
0.0111 (1.00) |
0.001 (0.329) |
0.00275 (0.875) |
8.89e-05 (0.0308) |
0.129 (1.00) |
0.27 (1.00) |
0.0765 (1.00) |
0.177 (1.00) |
| amp 4q35 1 | 29 (32%) | 61 |
4.62e-05 (0.0161) |
0.23 (1.00) |
0.000898 (0.297) |
0.0101 (1.00) |
0.0579 (1.00) |
0.181 (1.00) |
0.155 (1.00) |
0.327 (1.00) |
| amp 5p15 33 | 65 (72%) | 25 |
5.21e-05 (0.0181) |
0.866 (1.00) |
0.0105 (1.00) |
0.0705 (1.00) |
0.0445 (1.00) |
0.82 (1.00) |
0.122 (1.00) |
0.599 (1.00) |
| amp 9q31 3 | 34 (38%) | 56 |
0.00017 (0.0584) |
0.0253 (1.00) |
0.0298 (1.00) |
0.00311 (0.976) |
0.0499 (1.00) |
0.1 (1.00) |
0.0379 (1.00) |
0.104 (1.00) |
| amp 12q14 1 | 70 (78%) | 20 |
0.000142 (0.0491) |
0.255 (1.00) |
0.024 (1.00) |
0.0792 (1.00) |
0.447 (1.00) |
0.556 (1.00) |
0.0752 (1.00) |
0.521 (1.00) |
| amp 16q22 1 | 56 (62%) | 34 |
9.31e-07 (0.000331) |
0.387 (1.00) |
0.0295 (1.00) |
0.0045 (1.00) |
0.0635 (1.00) |
0.786 (1.00) |
0.0363 (1.00) |
1 (1.00) |
| amp xp11 22 | 49 (54%) | 41 |
0.000189 (0.0648) |
0.902 (1.00) |
0.0341 (1.00) |
0.157 (1.00) |
0.717 (1.00) |
0.345 (1.00) |
0.853 (1.00) |
0.217 (1.00) |
| amp xq28 | 46 (51%) | 44 |
0.000345 (0.116) |
0.741 (1.00) |
0.00921 (1.00) |
0.126 (1.00) |
0.483 (1.00) |
0.204 (1.00) |
0.677 (1.00) |
0.273 (1.00) |
| del 9p23 | 24 (27%) | 66 |
0.00095 (0.314) |
0.00546 (1.00) |
3.54e-05 (0.0125) |
0.00146 (0.477) |
0.0069 (1.00) |
0.312 (1.00) |
0.0453 (1.00) |
0.347 (1.00) |
| del 11q14 1 | 25 (28%) | 65 |
0.000241 (0.0816) |
0.00593 (1.00) |
0.223 (1.00) |
0.343 (1.00) |
0.277 (1.00) |
0.371 (1.00) |
0.419 (1.00) |
0.585 (1.00) |
| amp 4p16 3 | 41 (46%) | 49 |
0.00282 (0.894) |
0.544 (1.00) |
0.0133 (1.00) |
0.0165 (1.00) |
0.485 (1.00) |
0.489 (1.00) |
0.555 (1.00) |
0.556 (1.00) |
| amp 6p21 31 | 19 (21%) | 71 |
0.11 (1.00) |
0.753 (1.00) |
0.0809 (1.00) |
1 (1.00) |
0.131 (1.00) |
0.225 (1.00) |
0.147 (1.00) |
0.177 (1.00) |
| amp 6q24 3 | 19 (21%) | 71 |
0.00689 (1.00) |
0.873 (1.00) |
0.0473 (1.00) |
0.87 (1.00) |
0.645 (1.00) |
0.737 (1.00) |
0.816 (1.00) |
0.903 (1.00) |
| amp 7p22 1 | 54 (60%) | 36 |
0.00894 (1.00) |
0.462 (1.00) |
0.00802 (1.00) |
0.0589 (1.00) |
0.534 (1.00) |
0.27 (1.00) |
0.424 (1.00) |
0.169 (1.00) |
| amp 17q25 3 | 18 (20%) | 72 |
0.124 (1.00) |
0.652 (1.00) |
0.126 (1.00) |
0.501 (1.00) |
0.398 (1.00) |
0.657 (1.00) |
0.54 (1.00) |
0.37 (1.00) |
| amp 19p13 12 | 58 (64%) | 32 |
0.00318 (0.992) |
0.0308 (1.00) |
0.00226 (0.725) |
0.00132 (0.431) |
0.325 (1.00) |
0.626 (1.00) |
0.241 (1.00) |
0.256 (1.00) |
| amp 19q12 | 56 (62%) | 34 |
0.00962 (1.00) |
0.0193 (1.00) |
0.00195 (0.63) |
0.00257 (0.819) |
0.178 (1.00) |
0.311 (1.00) |
0.125 (1.00) |
0.114 (1.00) |
| del 1p36 23 | 37 (41%) | 53 |
0.0361 (1.00) |
0.0156 (1.00) |
0.136 (1.00) |
0.0201 (1.00) |
0.924 (1.00) |
0.39 (1.00) |
0.693 (1.00) |
0.418 (1.00) |
| del 1q43 | 18 (20%) | 72 |
0.109 (1.00) |
0.0378 (1.00) |
0.588 (1.00) |
0.0875 (1.00) |
0.491 (1.00) |
0.248 (1.00) |
0.659 (1.00) |
0.307 (1.00) |
| del 2q22 1 | 18 (20%) | 72 |
0.00204 (0.657) |
0.00442 (1.00) |
0.237 (1.00) |
0.153 (1.00) |
0.319 (1.00) |
0.178 (1.00) |
0.393 (1.00) |
0.28 (1.00) |
| del 2q37 3 | 21 (23%) | 69 |
0.00343 (1.00) |
0.0313 (1.00) |
0.292 (1.00) |
0.199 (1.00) |
0.433 (1.00) |
0.463 (1.00) |
0.62 (1.00) |
0.631 (1.00) |
| del 3q13 31 | 23 (26%) | 67 |
0.0184 (1.00) |
0.264 (1.00) |
0.0139 (1.00) |
0.202 (1.00) |
0.275 (1.00) |
0.0978 (1.00) |
0.0757 (1.00) |
0.148 (1.00) |
| del 4q34 3 | 25 (28%) | 65 |
0.00986 (1.00) |
0.0441 (1.00) |
0.0227 (1.00) |
0.0296 (1.00) |
0.0349 (1.00) |
0.166 (1.00) |
0.118 (1.00) |
0.104 (1.00) |
| del 4q35 1 | 28 (31%) | 62 |
0.0469 (1.00) |
0.0331 (1.00) |
0.0974 (1.00) |
0.0697 (1.00) |
0.343 (1.00) |
0.643 (1.00) |
0.539 (1.00) |
0.532 (1.00) |
| del 6p24 3 | 20 (22%) | 70 |
0.00634 (1.00) |
0.9 (1.00) |
0.594 (1.00) |
0.687 (1.00) |
1 (1.00) |
0.737 (1.00) |
0.877 (1.00) |
0.903 (1.00) |
| del 6q26 | 21 (23%) | 69 |
0.0337 (1.00) |
0.881 (1.00) |
0.321 (1.00) |
0.322 (1.00) |
0.491 (1.00) |
0.662 (1.00) |
0.416 (1.00) |
0.524 (1.00) |
| del 7q32 3 | 9 (10%) | 81 |
0.232 (1.00) |
0.0305 (1.00) |
0.176 (1.00) |
0.0698 (1.00) |
0.895 (1.00) |
0.446 (1.00) |
1 (1.00) |
0.603 (1.00) |
| del 10q23 1 | 11 (12%) | 79 |
0.0379 (1.00) |
0.194 (1.00) |
0.237 (1.00) |
1 (1.00) |
1 (1.00) |
0.518 (1.00) |
0.891 (1.00) |
0.528 (1.00) |
| del 11p15 5 | 27 (30%) | 63 |
0.0015 (0.487) |
0.0189 (1.00) |
0.0917 (1.00) |
0.133 (1.00) |
0.123 (1.00) |
0.109 (1.00) |
0.199 (1.00) |
0.151 (1.00) |
| del 12q14 2 | 5 (6%) | 85 |
0.189 (1.00) |
0.246 (1.00) |
0.177 (1.00) |
0.505 (1.00) |
0.594 (1.00) |
1 (1.00) |
0.103 (1.00) |
1 (1.00) |
| del 13q14 2 | 42 (47%) | 48 |
0.00301 (0.95) |
0.0823 (1.00) |
0.333 (1.00) |
0.416 (1.00) |
1 (1.00) |
1 (1.00) |
0.903 (1.00) |
0.841 (1.00) |
| del 14q21 2 | 20 (22%) | 70 |
0.00729 (1.00) |
0.914 (1.00) |
0.308 (1.00) |
0.0992 (1.00) |
0.364 (1.00) |
0.357 (1.00) |
0.205 (1.00) |
0.363 (1.00) |
| del 16p13 3 | 7 (8%) | 83 |
0.101 (1.00) |
0.463 (1.00) |
0.544 (1.00) |
1 (1.00) |
0.0241 (1.00) |
0.451 (1.00) |
0.045 (1.00) |
0.411 (1.00) |
| del 16q23 1 | 9 (10%) | 81 |
0.0408 (1.00) |
0.508 (1.00) |
0.017 (1.00) |
0.167 (1.00) |
0.141 (1.00) |
0.855 (1.00) |
0.299 (1.00) |
0.851 (1.00) |
| del 17q11 2 | 24 (27%) | 66 |
0.00343 (1.00) |
0.00644 (1.00) |
0.104 (1.00) |
0.175 (1.00) |
0.237 (1.00) |
0.279 (1.00) |
0.236 (1.00) |
0.183 (1.00) |
| del 17q21 31 | 26 (29%) | 64 |
0.00301 (0.95) |
0.00838 (1.00) |
0.0767 (1.00) |
0.0836 (1.00) |
0.035 (1.00) |
0.074 (1.00) |
0.0447 (1.00) |
0.0369 (1.00) |
| del 17q24 2 | 24 (27%) | 66 |
0.000867 (0.288) |
0.0113 (1.00) |
0.162 (1.00) |
0.0485 (1.00) |
0.148 (1.00) |
0.204 (1.00) |
0.172 (1.00) |
0.116 (1.00) |
| del 18q21 2 | 36 (40%) | 54 |
0.0757 (1.00) |
0.308 (1.00) |
0.811 (1.00) |
0.942 (1.00) |
0.471 (1.00) |
0.472 (1.00) |
0.732 (1.00) |
0.63 (1.00) |
| del 20p12 1 | 12 (13%) | 78 |
0.00118 (0.387) |
0.588 (1.00) |
0.0424 (1.00) |
0.0257 (1.00) |
0.00633 (1.00) |
0.547 (1.00) |
0.0143 (1.00) |
0.556 (1.00) |
| del 22q12 1 | 50 (56%) | 40 |
0.0391 (1.00) |
0.134 (1.00) |
0.734 (1.00) |
0.417 (1.00) |
0.629 (1.00) |
0.51 (1.00) |
0.471 (1.00) |
0.614 (1.00) |
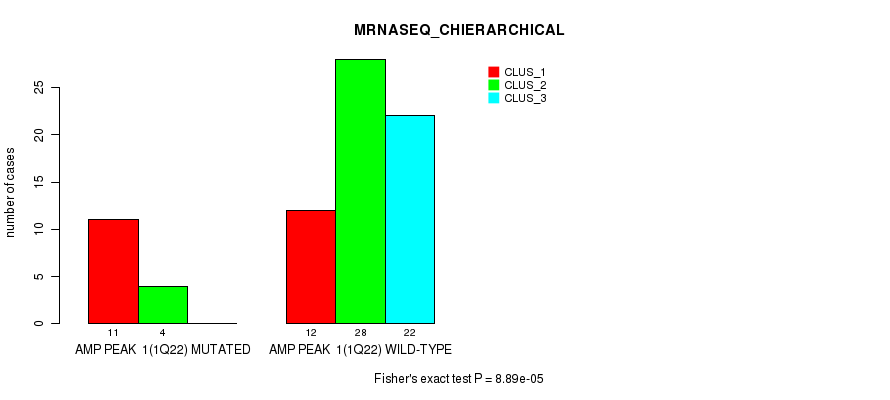
P value = 8.89e-05 (Fisher's exact test), Q value = 0.031
Table S1. Gene #1: 'amp_1q22' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 23 | 32 | 22 |
| AMP PEAK 1(1Q22) MUTATED | 11 | 4 | 0 |
| AMP PEAK 1(1Q22) WILD-TYPE | 12 | 28 | 22 |
Figure S1. Get High-res Image Gene #1: 'amp_1q22' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
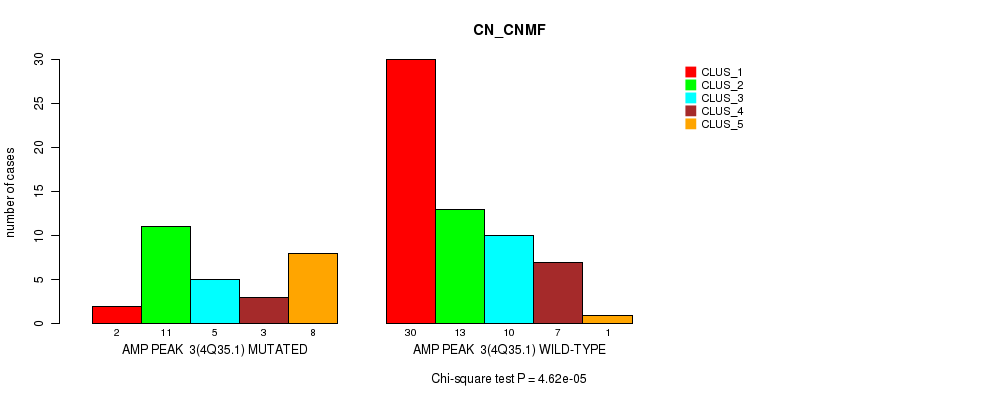
P value = 4.62e-05 (Chi-square test), Q value = 0.016
Table S2. Gene #3: 'amp_4q35.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 3(4Q35.1) MUTATED | 2 | 11 | 5 | 3 | 8 |
| AMP PEAK 3(4Q35.1) WILD-TYPE | 30 | 13 | 10 | 7 | 1 |
Figure S2. Get High-res Image Gene #3: 'amp_4q35.1' versus Molecular Subtype #1: 'CN_CNMF'
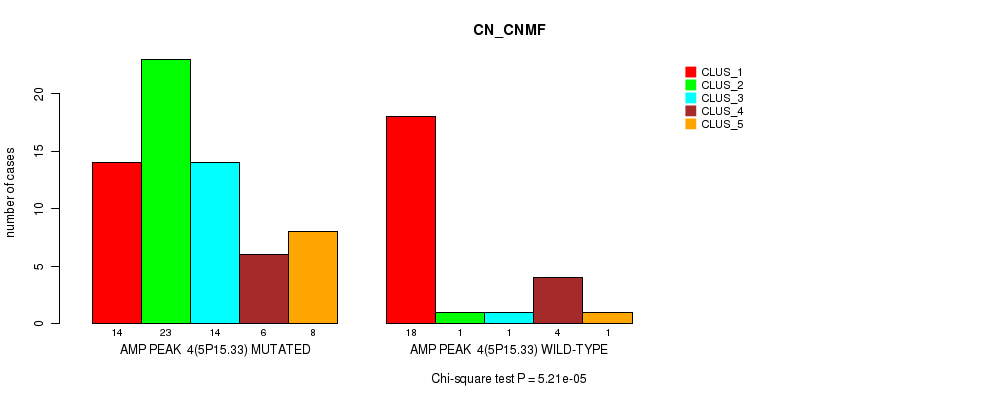
P value = 5.21e-05 (Chi-square test), Q value = 0.018
Table S3. Gene #4: 'amp_5p15.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 4(5P15.33) MUTATED | 14 | 23 | 14 | 6 | 8 |
| AMP PEAK 4(5P15.33) WILD-TYPE | 18 | 1 | 1 | 4 | 1 |
Figure S3. Get High-res Image Gene #4: 'amp_5p15.33' versus Molecular Subtype #1: 'CN_CNMF'
P value = 8.49e-08 (Chi-square test), Q value = 3.1e-05
Table S4. Gene #5: 'amp_5q35.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 5(5Q35.3) MUTATED | 10 | 24 | 14 | 7 | 8 |
| AMP PEAK 5(5Q35.3) WILD-TYPE | 22 | 0 | 1 | 3 | 1 |
Figure S4. Get High-res Image Gene #5: 'amp_5q35.3' versus Molecular Subtype #1: 'CN_CNMF'

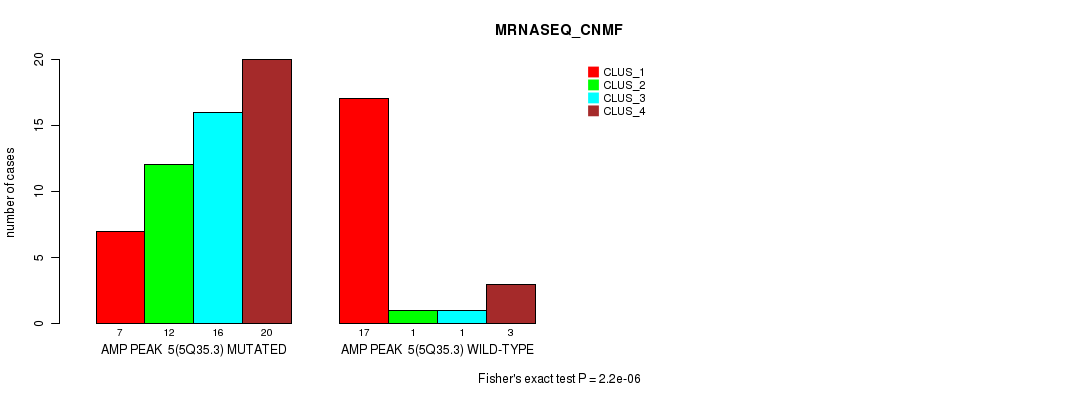
P value = 2.2e-06 (Fisher's exact test), Q value = 0.00078
Table S5. Gene #5: 'amp_5q35.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| AMP PEAK 5(5Q35.3) MUTATED | 7 | 12 | 16 | 20 |
| AMP PEAK 5(5Q35.3) WILD-TYPE | 17 | 1 | 1 | 3 |
Figure S5. Get High-res Image Gene #5: 'amp_5q35.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
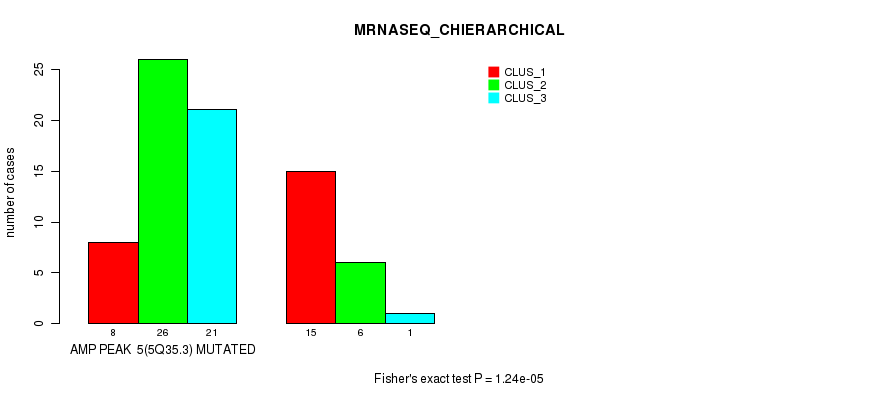
P value = 1.24e-05 (Fisher's exact test), Q value = 0.0044
Table S6. Gene #5: 'amp_5q35.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 23 | 32 | 22 |
| AMP PEAK 5(5Q35.3) MUTATED | 8 | 26 | 21 |
| AMP PEAK 5(5Q35.3) WILD-TYPE | 15 | 6 | 1 |
Figure S6. Get High-res Image Gene #5: 'amp_5q35.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
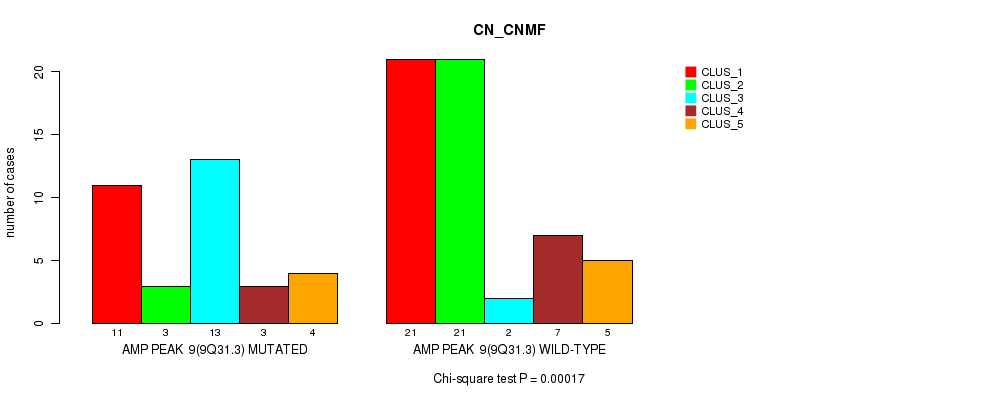
P value = 0.00017 (Chi-square test), Q value = 0.058
Table S7. Gene #9: 'amp_9q31.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 9(9Q31.3) MUTATED | 11 | 3 | 13 | 3 | 4 |
| AMP PEAK 9(9Q31.3) WILD-TYPE | 21 | 21 | 2 | 7 | 5 |
Figure S7. Get High-res Image Gene #9: 'amp_9q31.3' versus Molecular Subtype #1: 'CN_CNMF'
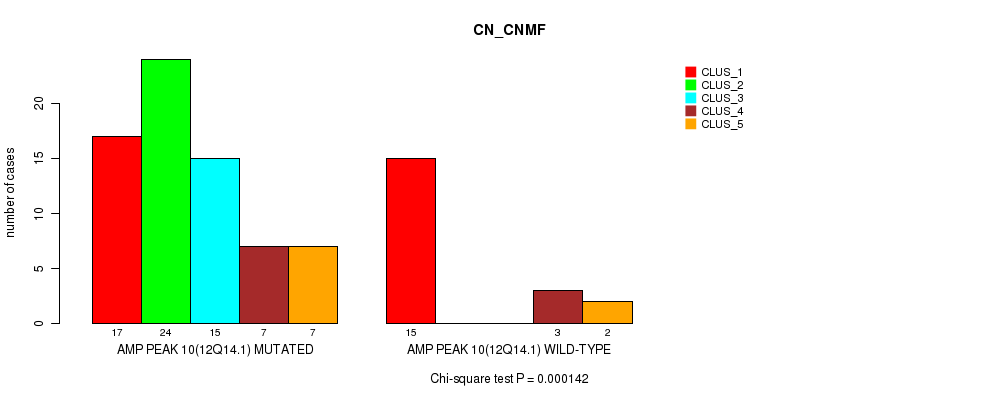
P value = 0.000142 (Chi-square test), Q value = 0.049
Table S8. Gene #10: 'amp_12q14.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 10(12Q14.1) MUTATED | 17 | 24 | 15 | 7 | 7 |
| AMP PEAK 10(12Q14.1) WILD-TYPE | 15 | 0 | 0 | 3 | 2 |
Figure S8. Get High-res Image Gene #10: 'amp_12q14.1' versus Molecular Subtype #1: 'CN_CNMF'
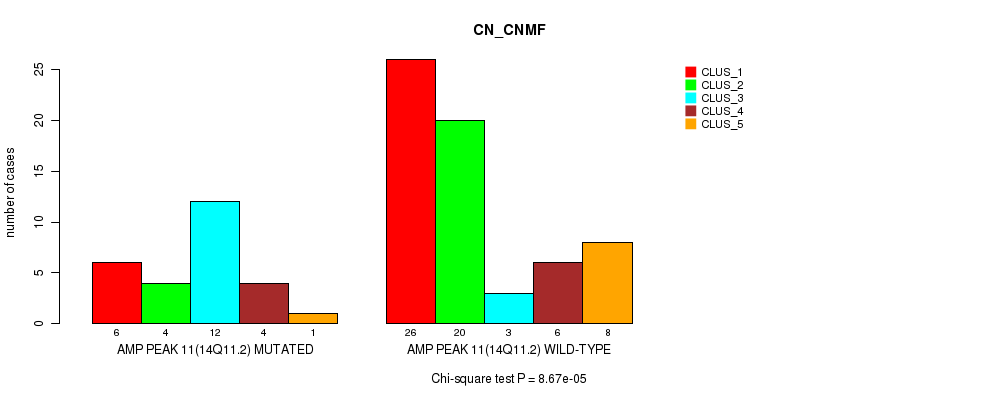
P value = 8.67e-05 (Chi-square test), Q value = 0.03
Table S9. Gene #11: 'amp_14q11.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 11(14Q11.2) MUTATED | 6 | 4 | 12 | 4 | 1 |
| AMP PEAK 11(14Q11.2) WILD-TYPE | 26 | 20 | 3 | 6 | 8 |
Figure S9. Get High-res Image Gene #11: 'amp_14q11.2' versus Molecular Subtype #1: 'CN_CNMF'
P value = 3.8e-05 (Fisher's exact test), Q value = 0.013
Table S10. Gene #11: 'amp_14q11.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| AMP PEAK 11(14Q11.2) MUTATED | 4 | 7 | 11 | 1 |
| AMP PEAK 11(14Q11.2) WILD-TYPE | 20 | 6 | 6 | 22 |
Figure S10. Get High-res Image Gene #11: 'amp_14q11.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
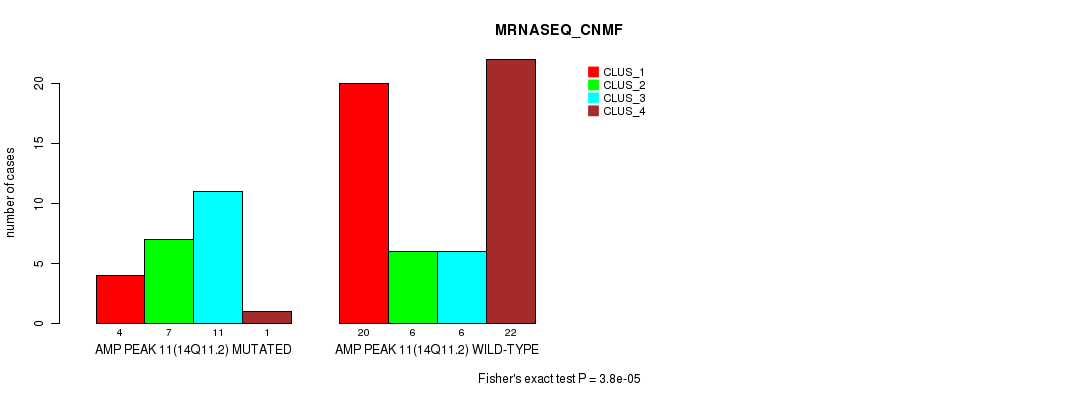
P value = 5.99e-07 (Fisher's exact test), Q value = 0.00021
Table S11. Gene #11: 'amp_14q11.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 23 | 32 | 22 |
| AMP PEAK 11(14Q11.2) MUTATED | 5 | 2 | 16 |
| AMP PEAK 11(14Q11.2) WILD-TYPE | 18 | 30 | 6 |
Figure S11. Get High-res Image Gene #11: 'amp_14q11.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.000295 (Fisher's exact test), Q value = 0.1
Table S12. Gene #11: 'amp_14q11.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 36 | 16 | 26 |
| AMP PEAK 11(14Q11.2) MUTATED | 19 | 3 | 2 |
| AMP PEAK 11(14Q11.2) WILD-TYPE | 17 | 13 | 24 |
Figure S12. Get High-res Image Gene #11: 'amp_14q11.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
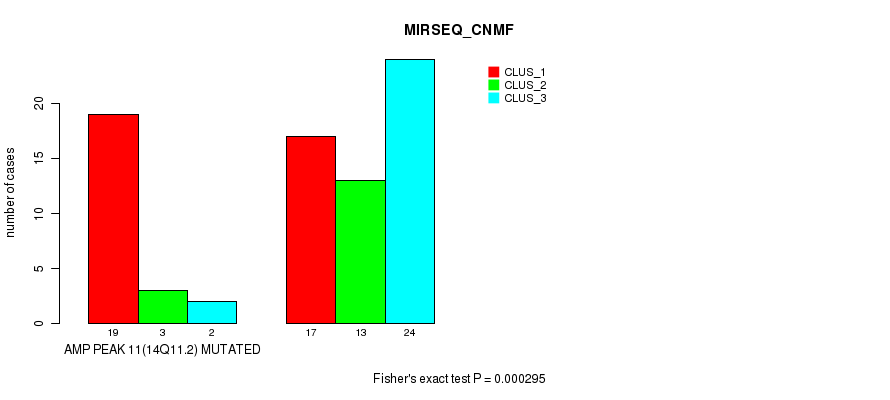
P value = 0.000226 (Fisher's exact test), Q value = 0.077
Table S13. Gene #11: 'amp_14q11.2' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 18 | 23 |
| AMP PEAK 11(14Q11.2) MUTATED | 19 | 4 | 1 |
| AMP PEAK 11(14Q11.2) WILD-TYPE | 18 | 14 | 22 |
Figure S13. Get High-res Image Gene #11: 'amp_14q11.2' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
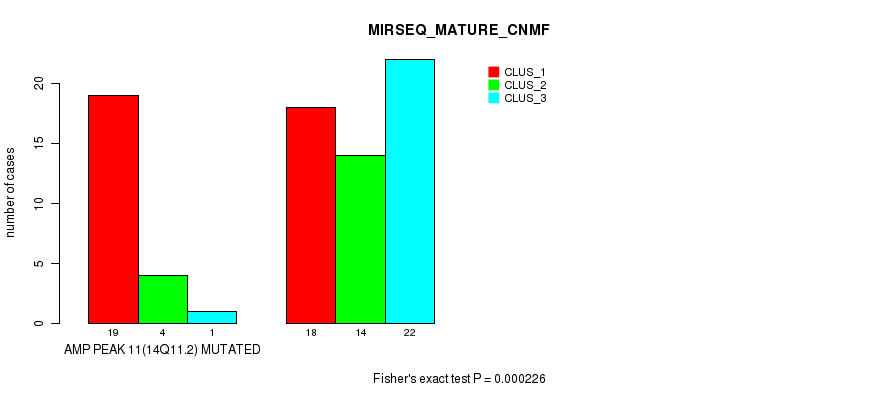
P value = 1.34e-07 (Chi-square test), Q value = 4.8e-05
Table S14. Gene #12: 'amp_16p13.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 12(16P13.3) MUTATED | 6 | 24 | 10 | 5 | 5 |
| AMP PEAK 12(16P13.3) WILD-TYPE | 26 | 0 | 5 | 5 | 4 |
Figure S14. Get High-res Image Gene #12: 'amp_16p13.3' versus Molecular Subtype #1: 'CN_CNMF'
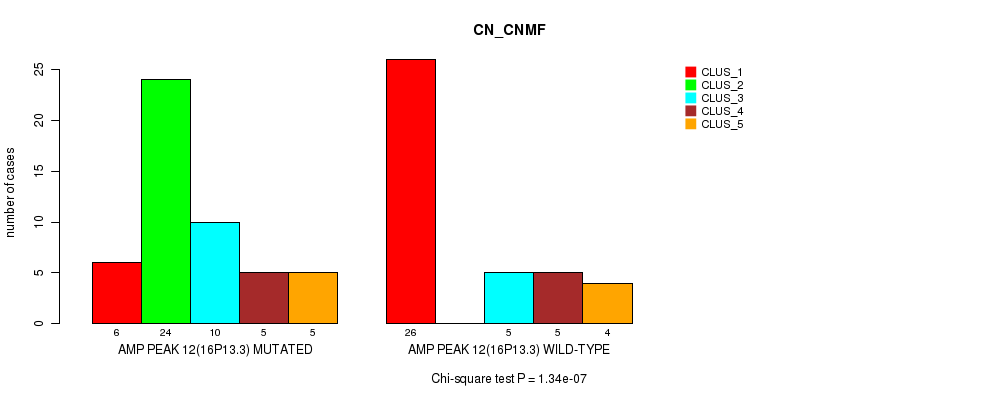
P value = 0.000195 (Fisher's exact test), Q value = 0.067
Table S15. Gene #12: 'amp_16p13.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| AMP PEAK 12(16P13.3) MUTATED | 5 | 9 | 14 | 16 |
| AMP PEAK 12(16P13.3) WILD-TYPE | 19 | 4 | 3 | 7 |
Figure S15. Get High-res Image Gene #12: 'amp_16p13.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
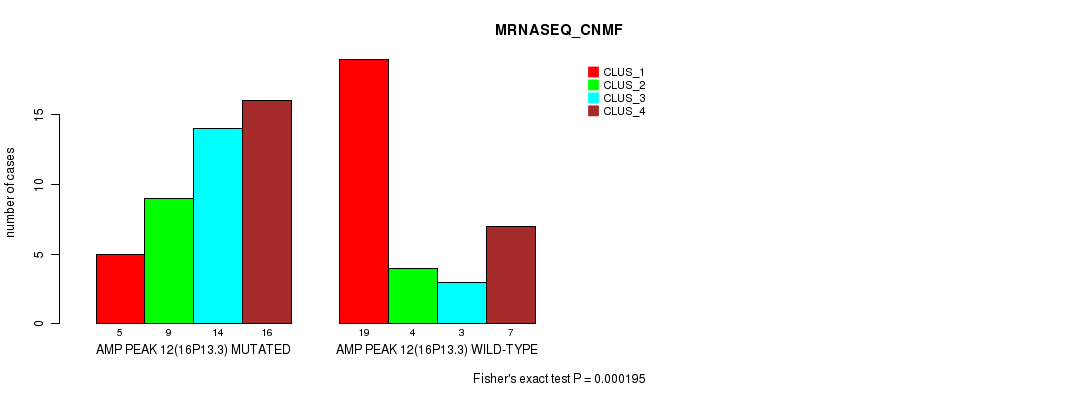
P value = 0.000585 (Fisher's exact test), Q value = 0.2
Table S16. Gene #12: 'amp_16p13.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 23 | 32 | 22 |
| AMP PEAK 12(16P13.3) MUTATED | 6 | 20 | 18 |
| AMP PEAK 12(16P13.3) WILD-TYPE | 17 | 12 | 4 |
Figure S16. Get High-res Image Gene #12: 'amp_16p13.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
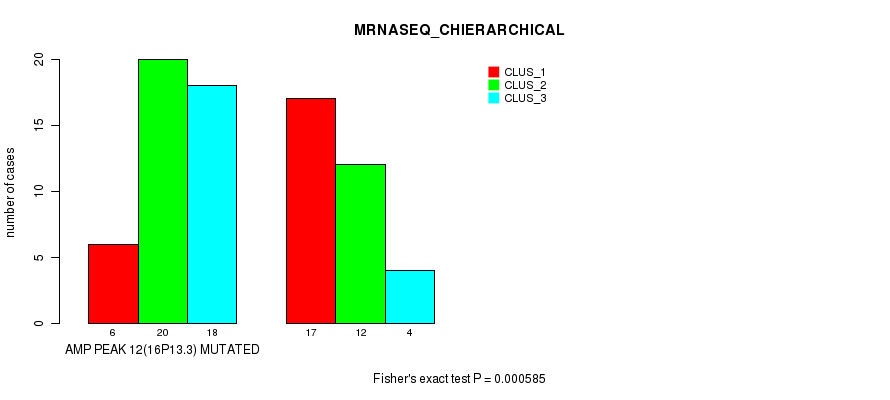
P value = 9.31e-07 (Chi-square test), Q value = 0.00033
Table S17. Gene #13: 'amp_16q22.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 13(16Q22.1) MUTATED | 11 | 24 | 13 | 3 | 5 |
| AMP PEAK 13(16Q22.1) WILD-TYPE | 21 | 0 | 2 | 7 | 4 |
Figure S17. Get High-res Image Gene #13: 'amp_16q22.1' versus Molecular Subtype #1: 'CN_CNMF'
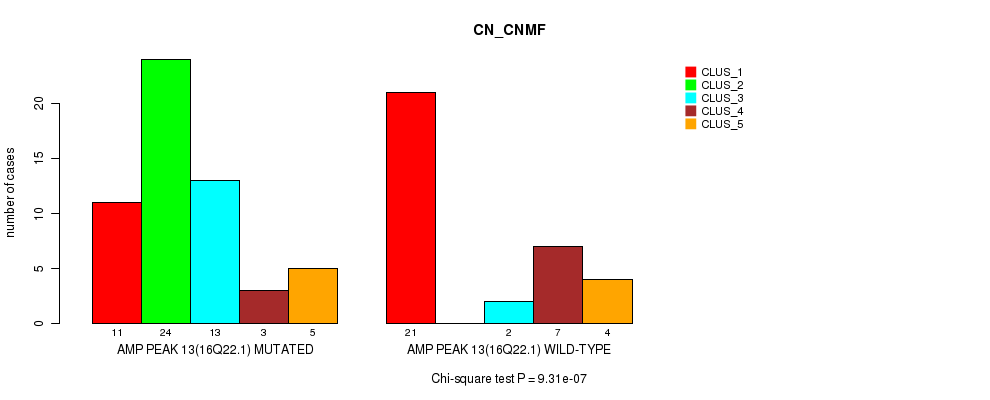
P value = 9.86e-08 (Chi-square test), Q value = 3.5e-05
Table S18. Gene #14: 'amp_16q24.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 14(16Q24.2) MUTATED | 6 | 24 | 10 | 4 | 5 |
| AMP PEAK 14(16Q24.2) WILD-TYPE | 26 | 0 | 5 | 6 | 4 |
Figure S18. Get High-res Image Gene #14: 'amp_16q24.2' versus Molecular Subtype #1: 'CN_CNMF'
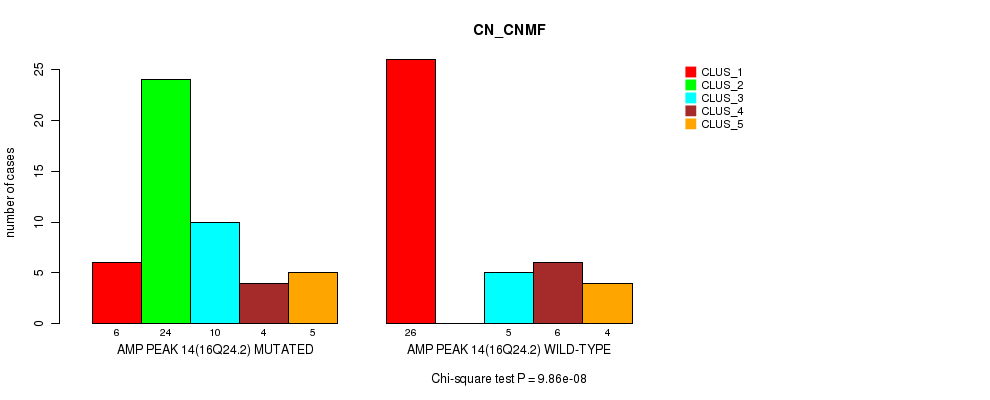
P value = 0.000271 (Fisher's exact test), Q value = 0.092
Table S19. Gene #14: 'amp_16q24.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| AMP PEAK 14(16Q24.2) MUTATED | 5 | 10 | 13 | 16 |
| AMP PEAK 14(16Q24.2) WILD-TYPE | 19 | 3 | 4 | 7 |
Figure S19. Get High-res Image Gene #14: 'amp_16q24.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
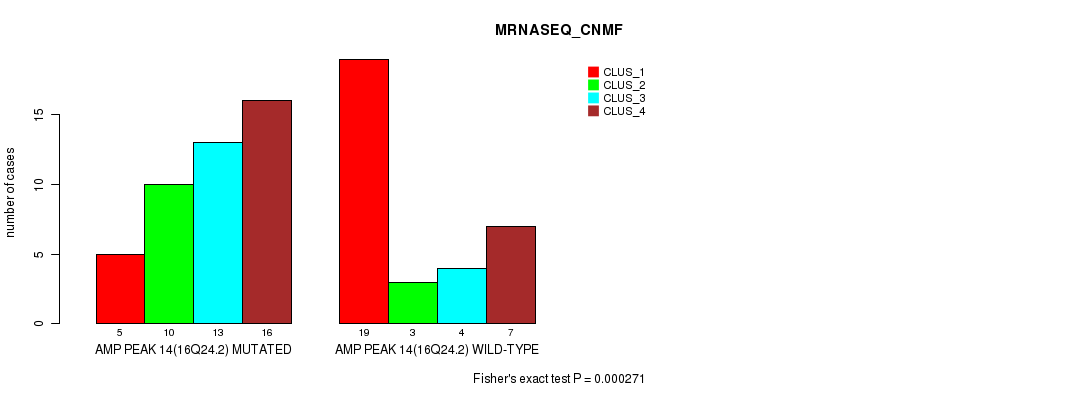
P value = 3.91e-05 (Fisher's exact test), Q value = 0.014
Table S20. Gene #14: 'amp_16q24.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 23 | 32 | 22 |
| AMP PEAK 14(16Q24.2) MUTATED | 5 | 20 | 19 |
| AMP PEAK 14(16Q24.2) WILD-TYPE | 18 | 12 | 3 |
Figure S20. Get High-res Image Gene #14: 'amp_16q24.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
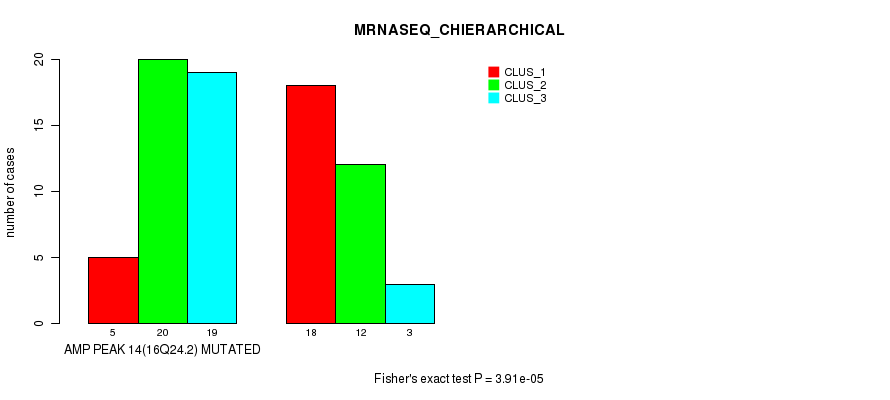
P value = 0.000189 (Chi-square test), Q value = 0.065
Table S21. Gene #18: 'amp_xp11.22' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 18(XP11.22) MUTATED | 10 | 21 | 11 | 4 | 3 |
| AMP PEAK 18(XP11.22) WILD-TYPE | 22 | 3 | 4 | 6 | 6 |
Figure S21. Get High-res Image Gene #18: 'amp_xp11.22' versus Molecular Subtype #1: 'CN_CNMF'
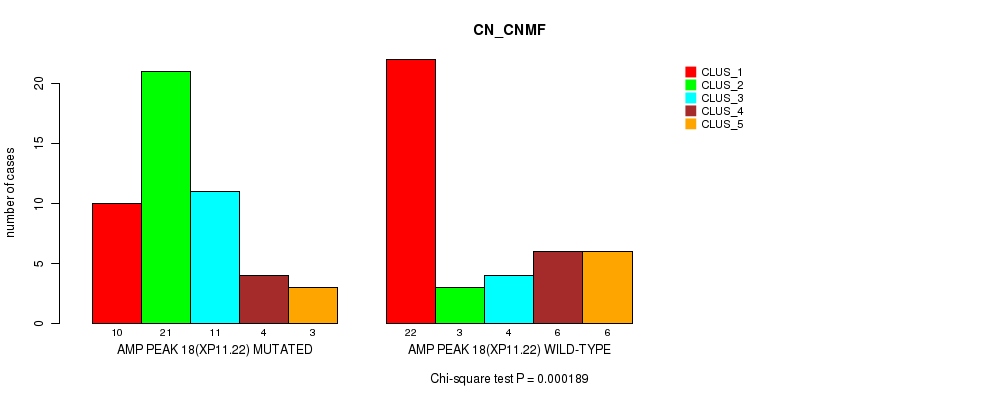
P value = 0.000345 (Chi-square test), Q value = 0.12
Table S22. Gene #19: 'amp_xq28' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 19(XQ28) MUTATED | 8 | 20 | 10 | 4 | 4 |
| AMP PEAK 19(XQ28) WILD-TYPE | 24 | 4 | 5 | 6 | 5 |
Figure S22. Get High-res Image Gene #19: 'amp_xq28' versus Molecular Subtype #1: 'CN_CNMF'
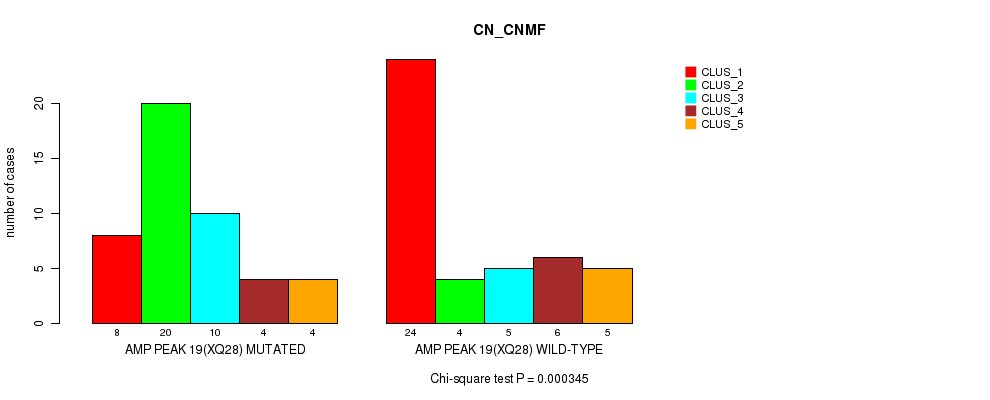
P value = 3.54e-05 (Fisher's exact test), Q value = 0.012
Table S23. Gene #30: 'del_9p23' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| DEL PEAK 11(9P23) MUTATED | 15 | 1 | 4 | 1 |
| DEL PEAK 11(9P23) WILD-TYPE | 9 | 12 | 13 | 22 |
Figure S23. Get High-res Image Gene #30: 'del_9p23' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
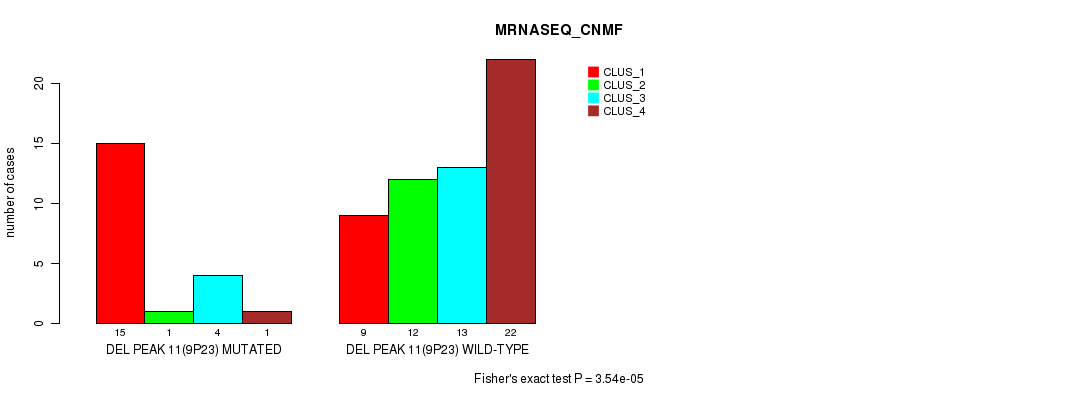
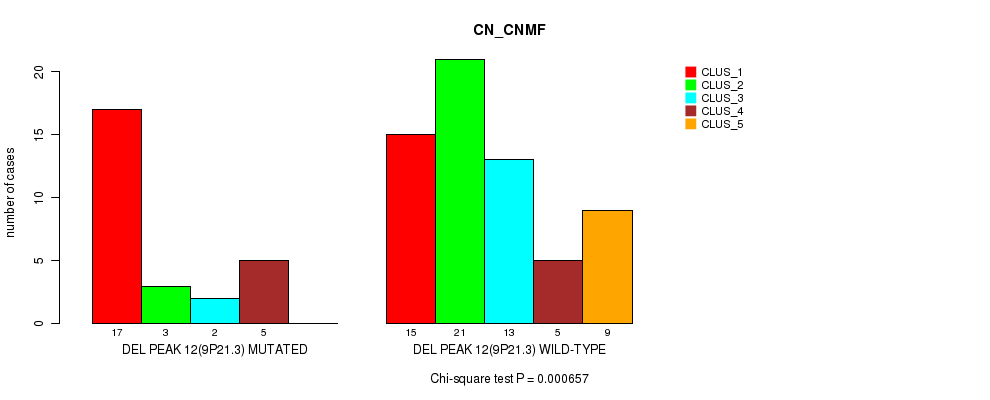
P value = 0.000657 (Chi-square test), Q value = 0.22
Table S24. Gene #31: 'del_9p21.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| DEL PEAK 12(9P21.3) MUTATED | 17 | 3 | 2 | 5 | 0 |
| DEL PEAK 12(9P21.3) WILD-TYPE | 15 | 21 | 13 | 5 | 9 |
Figure S24. Get High-res Image Gene #31: 'del_9p21.3' versus Molecular Subtype #1: 'CN_CNMF'
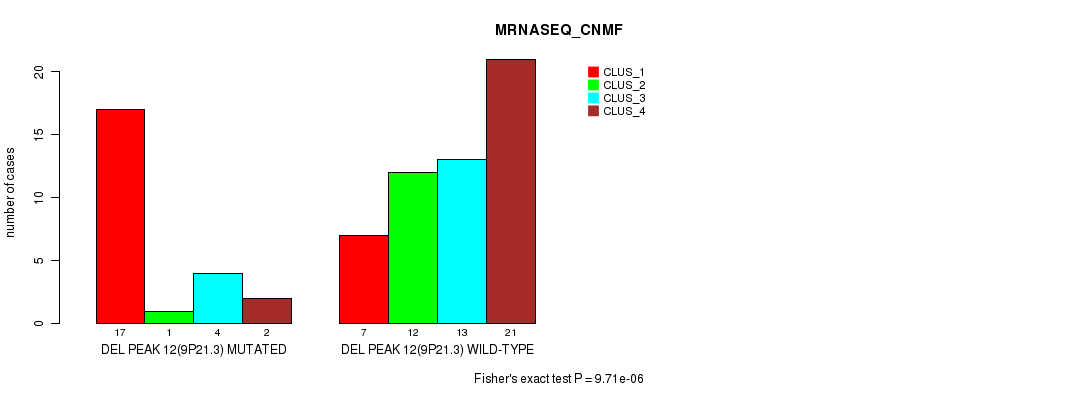
P value = 9.71e-06 (Fisher's exact test), Q value = 0.0034
Table S25. Gene #31: 'del_9p21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| DEL PEAK 12(9P21.3) MUTATED | 17 | 1 | 4 | 2 |
| DEL PEAK 12(9P21.3) WILD-TYPE | 7 | 12 | 13 | 21 |
Figure S25. Get High-res Image Gene #31: 'del_9p21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
P value = 0.00022 (Fisher's exact test), Q value = 0.075
Table S26. Gene #31: 'del_9p21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 23 | 32 | 22 |
| DEL PEAK 12(9P21.3) MUTATED | 15 | 6 | 3 |
| DEL PEAK 12(9P21.3) WILD-TYPE | 8 | 26 | 19 |
Figure S26. Get High-res Image Gene #31: 'del_9p21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

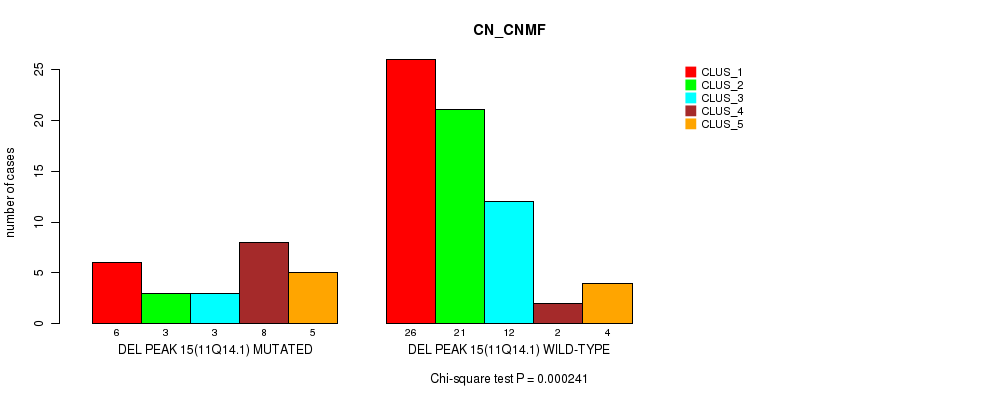
P value = 0.000241 (Chi-square test), Q value = 0.082
Table S27. Gene #34: 'del_11q14.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| DEL PEAK 15(11Q14.1) MUTATED | 6 | 3 | 3 | 8 | 5 |
| DEL PEAK 15(11Q14.1) WILD-TYPE | 26 | 21 | 12 | 2 | 4 |
Figure S27. Get High-res Image Gene #34: 'del_11q14.1' versus Molecular Subtype #1: 'CN_CNMF'
-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtype file = ACC-TP.transferedmergedcluster.txt
-
Number of patients = 90
-
Number of significantly focal cnvs = 45
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.