This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 71 focal events and 8 molecular subtypes across 103 patients, 20 significant findings detected with P value < 0.05 and Q value < 0.25.
-
amp_3q26.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_12p13.33 cnv correlated to 'CN_CNMF'.
-
amp_12p12.1 cnv correlated to 'CN_CNMF'.
-
del_3p14.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_9p21.3 cnv correlated to 'CN_CNMF', 'MIRSEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_16p13.3 cnv correlated to 'CN_CNMF'.
-
del_16q23.1 cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 71 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 20 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| amp 3q26 2 | 70 (68%) | 33 |
1.49e-10 (8.48e-08) |
4.31e-05 (0.024) |
0.00163 (0.867) |
8.57e-06 (0.00482) |
7.2e-06 (0.00405) |
3.2e-07 (0.000181) |
7.63e-05 (0.0424) |
3.2e-07 (0.000181) |
| del 3p14 3 | 71 (69%) | 32 |
0.000142 (0.0782) |
0.00167 (0.887) |
0.022 (1.00) |
0.000142 (0.0785) |
0.000222 (0.122) |
1.23e-05 (0.00691) |
0.00207 (1.00) |
1.23e-05 (0.00691) |
| del 9p21 3 | 78 (76%) | 25 |
2.04e-06 (0.00115) |
0.103 (1.00) |
0.045 (1.00) |
0.000969 (0.524) |
0.00774 (1.00) |
0.00011 (0.0609) |
0.000953 (0.517) |
0.00011 (0.0609) |
| del 16q23 1 | 45 (44%) | 58 |
2.5e-06 (0.00141) |
0.00039 (0.214) |
0.0014 (0.748) |
0.00083 (0.451) |
0.0017 (0.902) |
0.000737 (0.402) |
0.0154 (1.00) |
0.000737 (0.402) |
| amp 12p13 33 | 42 (41%) | 61 |
1.52e-05 (0.00848) |
0.0286 (1.00) |
0.0359 (1.00) |
0.00963 (1.00) |
0.0129 (1.00) |
0.00932 (1.00) |
0.115 (1.00) |
0.00932 (1.00) |
| amp 12p12 1 | 48 (47%) | 55 |
2.08e-05 (0.0116) |
0.136 (1.00) |
0.336 (1.00) |
0.0576 (1.00) |
0.0207 (1.00) |
0.0307 (1.00) |
0.0667 (1.00) |
0.0307 (1.00) |
| del 16p13 3 | 30 (29%) | 73 |
0.000135 (0.0747) |
0.00228 (1.00) |
0.0728 (1.00) |
0.023 (1.00) |
0.0669 (1.00) |
0.0621 (1.00) |
0.129 (1.00) |
0.0621 (1.00) |
| amp 1p34 2 | 20 (19%) | 83 |
0.228 (1.00) |
0.0908 (1.00) |
0.808 (1.00) |
0.883 (1.00) |
0.783 (1.00) |
0.587 (1.00) |
0.803 (1.00) |
0.587 (1.00) |
| amp 1q21 3 | 52 (50%) | 51 |
0.00401 (1.00) |
0.494 (1.00) |
1 (1.00) |
0.859 (1.00) |
0.93 (1.00) |
0.552 (1.00) |
0.854 (1.00) |
0.552 (1.00) |
| amp 2q14 2 | 34 (33%) | 69 |
0.545 (1.00) |
1 (1.00) |
0.581 (1.00) |
0.378 (1.00) |
0.422 (1.00) |
0.776 (1.00) |
0.446 (1.00) |
0.776 (1.00) |
| amp 5p15 33 | 59 (57%) | 44 |
0.00172 (0.911) |
0.0329 (1.00) |
0.0517 (1.00) |
0.00244 (1.00) |
0.0124 (1.00) |
0.00613 (1.00) |
0.0465 (1.00) |
0.00613 (1.00) |
| amp 6p21 1 | 35 (34%) | 68 |
0.00401 (1.00) |
0.525 (1.00) |
0.276 (1.00) |
0.287 (1.00) |
0.435 (1.00) |
0.46 (1.00) |
0.36 (1.00) |
0.46 (1.00) |
| amp 7p22 3 | 70 (68%) | 33 |
0.154 (1.00) |
0.59 (1.00) |
0.00149 (0.796) |
0.00883 (1.00) |
0.389 (1.00) |
0.394 (1.00) |
0.864 (1.00) |
0.394 (1.00) |
| amp 7p11 2 | 70 (68%) | 33 |
0.507 (1.00) |
0.435 (1.00) |
0.0427 (1.00) |
0.303 (1.00) |
0.103 (1.00) |
0.235 (1.00) |
0.287 (1.00) |
0.235 (1.00) |
| amp 7q21 2 | 66 (64%) | 37 |
0.0671 (1.00) |
0.146 (1.00) |
0.95 (1.00) |
1 (1.00) |
0.304 (1.00) |
0.418 (1.00) |
0.669 (1.00) |
0.418 (1.00) |
| amp 8p11 21 | 49 (48%) | 54 |
0.214 (1.00) |
0.776 (1.00) |
0.236 (1.00) |
1 (1.00) |
0.904 (1.00) |
0.681 (1.00) |
0.719 (1.00) |
0.681 (1.00) |
| amp 8q24 21 | 83 (81%) | 20 |
0.00211 (1.00) |
0.0129 (1.00) |
0.169 (1.00) |
0.035 (1.00) |
0.306 (1.00) |
0.0237 (1.00) |
0.024 (1.00) |
0.0237 (1.00) |
| amp 9p13 3 | 22 (21%) | 81 |
0.44 (1.00) |
0.312 (1.00) |
0.335 (1.00) |
0.593 (1.00) |
0.121 (1.00) |
0.599 (1.00) |
0.852 (1.00) |
0.599 (1.00) |
| amp 10p11 21 | 29 (28%) | 74 |
0.703 (1.00) |
0.18 (1.00) |
0.922 (1.00) |
0.862 (1.00) |
0.924 (1.00) |
0.802 (1.00) |
0.898 (1.00) |
0.802 (1.00) |
| amp 11p13 | 28 (27%) | 75 |
0.73 (1.00) |
0.863 (1.00) |
0.243 (1.00) |
0.72 (1.00) |
0.535 (1.00) |
0.463 (1.00) |
0.492 (1.00) |
0.463 (1.00) |
| amp 11q13 3 | 57 (55%) | 46 |
0.00214 (1.00) |
0.0507 (1.00) |
0.181 (1.00) |
0.0644 (1.00) |
0.104 (1.00) |
0.0208 (1.00) |
0.173 (1.00) |
0.0208 (1.00) |
| amp 13q22 1 | 36 (35%) | 67 |
0.0522 (1.00) |
0.232 (1.00) |
0.299 (1.00) |
0.104 (1.00) |
0.0311 (1.00) |
0.0644 (1.00) |
0.0842 (1.00) |
0.0644 (1.00) |
| amp 15q26 3 | 33 (32%) | 70 |
0.249 (1.00) |
0.0279 (1.00) |
0.299 (1.00) |
0.0888 (1.00) |
0.126 (1.00) |
0.0793 (1.00) |
0.124 (1.00) |
0.0793 (1.00) |
| amp 17q12 | 38 (37%) | 65 |
0.00142 (0.76) |
0.191 (1.00) |
0.339 (1.00) |
0.024 (1.00) |
0.0475 (1.00) |
0.101 (1.00) |
0.17 (1.00) |
0.101 (1.00) |
| amp 18p11 32 | 37 (36%) | 66 |
0.00477 (1.00) |
0.546 (1.00) |
0.778 (1.00) |
0.505 (1.00) |
0.662 (1.00) |
0.634 (1.00) |
0.462 (1.00) |
0.634 (1.00) |
| amp 18q11 2 | 31 (30%) | 72 |
0.522 (1.00) |
0.0375 (1.00) |
0.345 (1.00) |
0.149 (1.00) |
0.283 (1.00) |
0.208 (1.00) |
0.531 (1.00) |
0.208 (1.00) |
| amp 19p13 2 | 18 (17%) | 85 |
0.106 (1.00) |
0.204 (1.00) |
0.219 (1.00) |
0.272 (1.00) |
0.0508 (1.00) |
0.301 (1.00) |
0.0366 (1.00) |
0.301 (1.00) |
| amp 19q12 | 29 (28%) | 74 |
0.00104 (0.558) |
0.177 (1.00) |
0.427 (1.00) |
0.258 (1.00) |
0.283 (1.00) |
0.208 (1.00) |
0.0461 (1.00) |
0.208 (1.00) |
| del 1p36 13 | 33 (32%) | 70 |
0.0487 (1.00) |
0.0931 (1.00) |
0.316 (1.00) |
0.192 (1.00) |
0.374 (1.00) |
0.319 (1.00) |
0.67 (1.00) |
0.319 (1.00) |
| del 1p13 2 | 34 (33%) | 69 |
0.0461 (1.00) |
0.628 (1.00) |
0.255 (1.00) |
0.365 (1.00) |
0.836 (1.00) |
0.795 (1.00) |
0.688 (1.00) |
0.795 (1.00) |
| del 1q44 | 13 (13%) | 90 |
0.219 (1.00) |
0.824 (1.00) |
0.262 (1.00) |
0.594 (1.00) |
0.466 (1.00) |
0.391 (1.00) |
0.704 (1.00) |
0.391 (1.00) |
| del 2q22 1 | 32 (31%) | 71 |
0.0928 (1.00) |
0.0072 (1.00) |
0.395 (1.00) |
0.187 (1.00) |
0.0829 (1.00) |
0.0578 (1.00) |
0.0796 (1.00) |
0.0578 (1.00) |
| del 2q33 3 | 17 (17%) | 86 |
1 (1.00) |
0.0255 (1.00) |
0.261 (1.00) |
0.49 (1.00) |
0.128 (1.00) |
0.451 (1.00) |
0.31 (1.00) |
0.451 (1.00) |
| del 3p14 2 | 72 (70%) | 31 |
0.00174 (0.919) |
0.0174 (1.00) |
0.309 (1.00) |
0.00629 (1.00) |
0.0187 (1.00) |
0.00101 (0.544) |
0.0511 (1.00) |
0.00101 (0.544) |
| del 3q11 1 | 17 (17%) | 86 |
0.774 (1.00) |
0.887 (1.00) |
0.46 (1.00) |
0.523 (1.00) |
0.699 (1.00) |
0.583 (1.00) |
0.702 (1.00) |
0.583 (1.00) |
| del 3q26 31 | 8 (8%) | 95 |
0.0534 (1.00) |
0.0733 (1.00) |
0.068 (1.00) |
0.0205 (1.00) |
0.0443 (1.00) |
0.0191 (1.00) |
0.0843 (1.00) |
0.0191 (1.00) |
| del 4p16 3 | 53 (51%) | 50 |
0.702 (1.00) |
0.718 (1.00) |
0.362 (1.00) |
0.809 (1.00) |
0.193 (1.00) |
0.416 (1.00) |
0.267 (1.00) |
0.416 (1.00) |
| del 4p15 2 | 54 (52%) | 49 |
0.0185 (1.00) |
0.399 (1.00) |
0.17 (1.00) |
0.537 (1.00) |
0.144 (1.00) |
0.233 (1.00) |
0.664 (1.00) |
0.233 (1.00) |
| del 4q21 23 | 50 (49%) | 53 |
0.000746 (0.406) |
0.171 (1.00) |
0.0524 (1.00) |
0.489 (1.00) |
0.321 (1.00) |
0.694 (1.00) |
0.714 (1.00) |
0.694 (1.00) |
| del 4q22 1 | 53 (51%) | 50 |
0.000472 (0.259) |
0.248 (1.00) |
0.0333 (1.00) |
0.5 (1.00) |
0.319 (1.00) |
0.615 (1.00) |
0.664 (1.00) |
0.615 (1.00) |
| del 5q11 2 | 58 (56%) | 45 |
0.0717 (1.00) |
0.0283 (1.00) |
0.698 (1.00) |
0.698 (1.00) |
0.887 (1.00) |
0.684 (1.00) |
0.412 (1.00) |
0.684 (1.00) |
| del 6p25 3 | 37 (36%) | 66 |
0.775 (1.00) |
0.154 (1.00) |
0.661 (1.00) |
0.483 (1.00) |
0.301 (1.00) |
0.271 (1.00) |
0.161 (1.00) |
0.271 (1.00) |
| del 6q16 1 | 29 (28%) | 74 |
0.312 (1.00) |
0.6 (1.00) |
0.355 (1.00) |
0.478 (1.00) |
0.93 (1.00) |
1 (1.00) |
0.847 (1.00) |
1 (1.00) |
| del 6q26 | 25 (24%) | 78 |
0.101 (1.00) |
0.384 (1.00) |
0.0741 (1.00) |
0.0489 (1.00) |
0.158 (1.00) |
0.0962 (1.00) |
0.312 (1.00) |
0.0962 (1.00) |
| del 7q31 1 | 30 (29%) | 73 |
0.76 (1.00) |
0.589 (1.00) |
0.318 (1.00) |
0.627 (1.00) |
0.9 (1.00) |
0.801 (1.00) |
0.668 (1.00) |
0.801 (1.00) |
| del 7q36 2 | 38 (37%) | 65 |
0.847 (1.00) |
0.6 (1.00) |
0.14 (1.00) |
0.901 (1.00) |
0.18 (1.00) |
0.756 (1.00) |
0.319 (1.00) |
0.756 (1.00) |
| del 8p23 2 | 46 (45%) | 57 |
0.00399 (1.00) |
0.522 (1.00) |
0.0402 (1.00) |
0.221 (1.00) |
0.185 (1.00) |
0.069 (1.00) |
0.131 (1.00) |
0.069 (1.00) |
| del 10p15 1 | 28 (27%) | 75 |
0.222 (1.00) |
0.565 (1.00) |
0.52 (1.00) |
0.168 (1.00) |
0.614 (1.00) |
0.577 (1.00) |
0.578 (1.00) |
0.577 (1.00) |
| del 10p11 21 | 23 (22%) | 80 |
0.00191 (1.00) |
0.0015 (0.797) |
0.109 (1.00) |
0.02 (1.00) |
0.017 (1.00) |
0.0206 (1.00) |
0.0413 (1.00) |
0.0206 (1.00) |
| del 10q22 3 | 33 (32%) | 70 |
0.00834 (1.00) |
0.00751 (1.00) |
0.203 (1.00) |
0.0307 (1.00) |
0.143 (1.00) |
0.0585 (1.00) |
0.151 (1.00) |
0.0585 (1.00) |
| del 10q23 31 | 41 (40%) | 62 |
0.0325 (1.00) |
0.145 (1.00) |
0.528 (1.00) |
0.753 (1.00) |
0.794 (1.00) |
0.836 (1.00) |
0.886 (1.00) |
0.836 (1.00) |
| del 11p15 4 | 39 (38%) | 64 |
0.361 (1.00) |
0.289 (1.00) |
0.0775 (1.00) |
0.107 (1.00) |
0.12 (1.00) |
0.0463 (1.00) |
0.0858 (1.00) |
0.0463 (1.00) |
| del 11q24 3 | 43 (42%) | 60 |
0.00301 (1.00) |
0.47 (1.00) |
0.665 (1.00) |
0.462 (1.00) |
0.592 (1.00) |
0.287 (1.00) |
0.187 (1.00) |
0.287 (1.00) |
| del 12q23 1 | 27 (26%) | 76 |
0.652 (1.00) |
0.497 (1.00) |
0.181 (1.00) |
0.108 (1.00) |
0.167 (1.00) |
0.208 (1.00) |
0.435 (1.00) |
0.208 (1.00) |
| del 13q14 2 | 46 (45%) | 57 |
0.000485 (0.265) |
0.00681 (1.00) |
0.0261 (1.00) |
0.0182 (1.00) |
0.0162 (1.00) |
0.00371 (1.00) |
0.00415 (1.00) |
0.00371 (1.00) |
| del 13q21 2 | 42 (41%) | 61 |
0.00142 (0.759) |
0.194 (1.00) |
0.322 (1.00) |
0.157 (1.00) |
0.12 (1.00) |
0.0463 (1.00) |
0.0204 (1.00) |
0.0463 (1.00) |
| del 15q11 2 | 33 (32%) | 70 |
0.218 (1.00) |
0.404 (1.00) |
0.784 (1.00) |
0.535 (1.00) |
0.665 (1.00) |
0.68 (1.00) |
0.462 (1.00) |
0.68 (1.00) |
| del 17p11 2 | 45 (44%) | 58 |
0.103 (1.00) |
0.0013 (0.699) |
0.00818 (1.00) |
0.00333 (1.00) |
0.00732 (1.00) |
0.00406 (1.00) |
0.0032 (1.00) |
0.00406 (1.00) |
| del 17q24 3 | 14 (14%) | 89 |
0.616 (1.00) |
0.658 (1.00) |
0.823 (1.00) |
0.282 (1.00) |
0.794 (1.00) |
0.737 (1.00) |
0.34 (1.00) |
0.737 (1.00) |
| del 17q25 3 | 25 (24%) | 78 |
0.24 (1.00) |
0.924 (1.00) |
0.171 (1.00) |
0.577 (1.00) |
0.535 (1.00) |
0.76 (1.00) |
0.266 (1.00) |
0.76 (1.00) |
| del 18q12 2 | 55 (53%) | 48 |
0.307 (1.00) |
0.136 (1.00) |
0.111 (1.00) |
0.0667 (1.00) |
0.0873 (1.00) |
0.0743 (1.00) |
0.0472 (1.00) |
0.0743 (1.00) |
| del 18q21 2 | 61 (59%) | 42 |
0.635 (1.00) |
0.0866 (1.00) |
0.289 (1.00) |
0.109 (1.00) |
0.358 (1.00) |
0.181 (1.00) |
0.135 (1.00) |
0.181 (1.00) |
| del 19p13 3 | 46 (45%) | 57 |
0.0491 (1.00) |
0.927 (1.00) |
0.705 (1.00) |
0.815 (1.00) |
0.202 (1.00) |
0.79 (1.00) |
0.441 (1.00) |
0.79 (1.00) |
| del 19q13 31 | 28 (27%) | 75 |
0.0572 (1.00) |
0.043 (1.00) |
0.327 (1.00) |
0.183 (1.00) |
0.0861 (1.00) |
0.137 (1.00) |
0.398 (1.00) |
0.137 (1.00) |
| del 20p12 1 | 19 (18%) | 84 |
0.0746 (1.00) |
0.835 (1.00) |
0.579 (1.00) |
0.317 (1.00) |
0.158 (1.00) |
0.426 (1.00) |
0.214 (1.00) |
0.426 (1.00) |
| del 21q11 2 | 61 (59%) | 42 |
0.676 (1.00) |
0.396 (1.00) |
0.308 (1.00) |
0.0172 (1.00) |
0.567 (1.00) |
0.228 (1.00) |
0.472 (1.00) |
0.228 (1.00) |
| del 21q22 12 | 57 (55%) | 46 |
0.459 (1.00) |
0.0583 (1.00) |
0.0167 (1.00) |
0.00991 (1.00) |
0.129 (1.00) |
0.0516 (1.00) |
0.0735 (1.00) |
0.0516 (1.00) |
| del 22q12 3 | 38 (37%) | 65 |
0.0133 (1.00) |
0.00361 (1.00) |
0.0346 (1.00) |
0.062 (1.00) |
0.123 (1.00) |
0.0656 (1.00) |
0.163 (1.00) |
0.0656 (1.00) |
| del xp21 1 | 40 (39%) | 63 |
0.00522 (1.00) |
0.0371 (1.00) |
0.0469 (1.00) |
0.112 (1.00) |
0.684 (1.00) |
0.375 (1.00) |
0.769 (1.00) |
0.375 (1.00) |
| del xp11 3 | 42 (41%) | 61 |
0.00181 (0.953) |
0.0267 (1.00) |
0.0611 (1.00) |
0.00833 (1.00) |
0.105 (1.00) |
0.0353 (1.00) |
0.213 (1.00) |
0.0353 (1.00) |
| del xq21 33 | 23 (22%) | 80 |
0.591 (1.00) |
0.595 (1.00) |
0.516 (1.00) |
0.755 (1.00) |
0.794 (1.00) |
0.724 (1.00) |
1 (1.00) |
0.724 (1.00) |
P value = 1.49e-10 (Fisher's exact test), Q value = 8.5e-08
Table S1. Gene #4: 'amp_3q26.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 38 | 39 | 26 |
| AMP PEAK 4(3Q26.2) MUTATED | 13 | 39 | 18 |
| AMP PEAK 4(3Q26.2) WILD-TYPE | 25 | 0 | 8 |
Figure S1. Get High-res Image Gene #4: 'amp_3q26.2' versus Molecular Subtype #1: 'CN_CNMF'
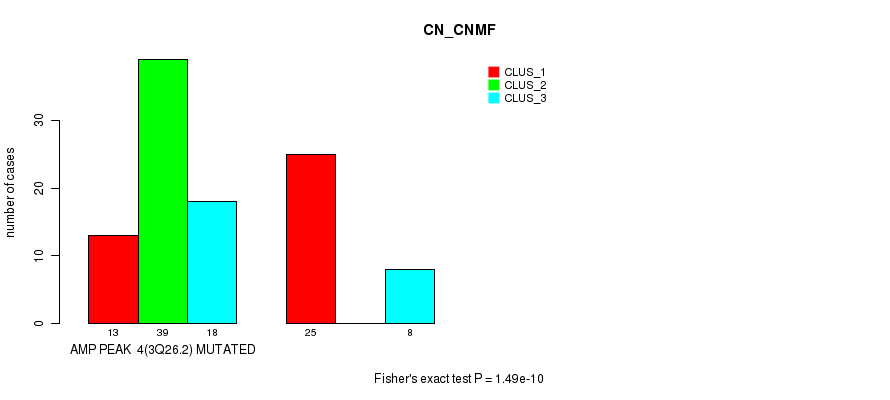
P value = 4.31e-05 (Fisher's exact test), Q value = 0.024
Table S2. Gene #4: 'amp_3q26.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 43 | 20 |
| AMP PEAK 4(3Q26.2) MUTATED | 19 | 39 | 12 |
| AMP PEAK 4(3Q26.2) WILD-TYPE | 21 | 4 | 8 |
Figure S2. Get High-res Image Gene #4: 'amp_3q26.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
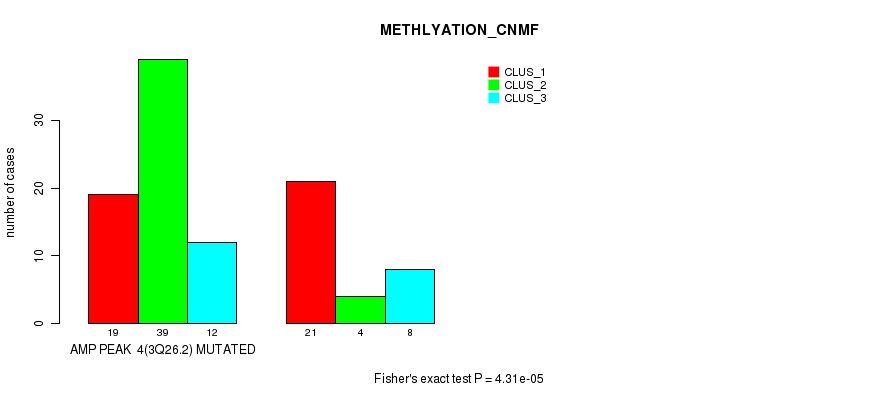
P value = 8.57e-06 (Fisher's exact test), Q value = 0.0048
Table S3. Gene #4: 'amp_3q26.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 4 | 39 |
| AMP PEAK 4(3Q26.2) MUTATED | 12 | 3 | 37 |
| AMP PEAK 4(3Q26.2) WILD-TYPE | 15 | 1 | 2 |
Figure S3. Get High-res Image Gene #4: 'amp_3q26.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 7.2e-06 (Fisher's exact test), Q value = 0.0041
Table S4. Gene #4: 'amp_3q26.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 30 | 14 | 28 |
| AMP PEAK 4(3Q26.2) MUTATED | 29 | 12 | 12 |
| AMP PEAK 4(3Q26.2) WILD-TYPE | 1 | 2 | 16 |
Figure S4. Get High-res Image Gene #4: 'amp_3q26.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
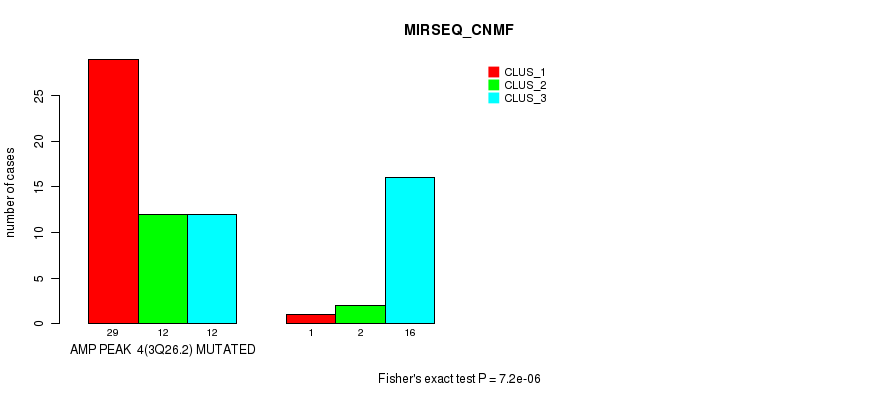
P value = 3.2e-07 (Fisher's exact test), Q value = 0.00018
Table S5. Gene #4: 'amp_3q26.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 1 | 28 | 43 |
| AMP PEAK 4(3Q26.2) MUTATED | 0 | 12 | 41 |
| AMP PEAK 4(3Q26.2) WILD-TYPE | 1 | 16 | 2 |
Figure S5. Get High-res Image Gene #4: 'amp_3q26.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
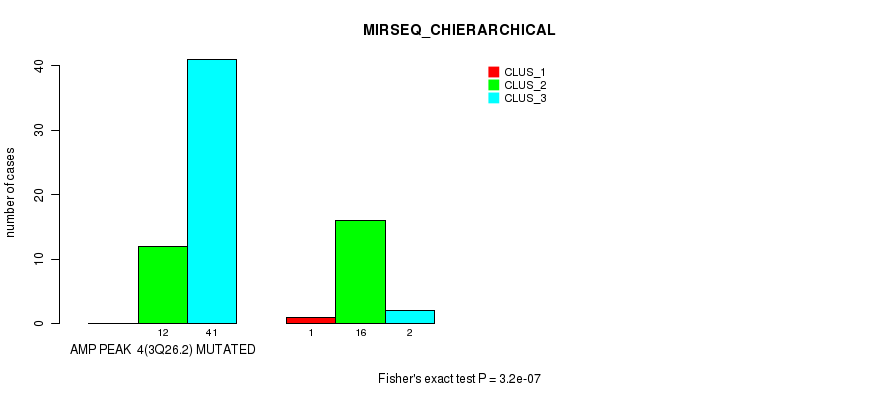
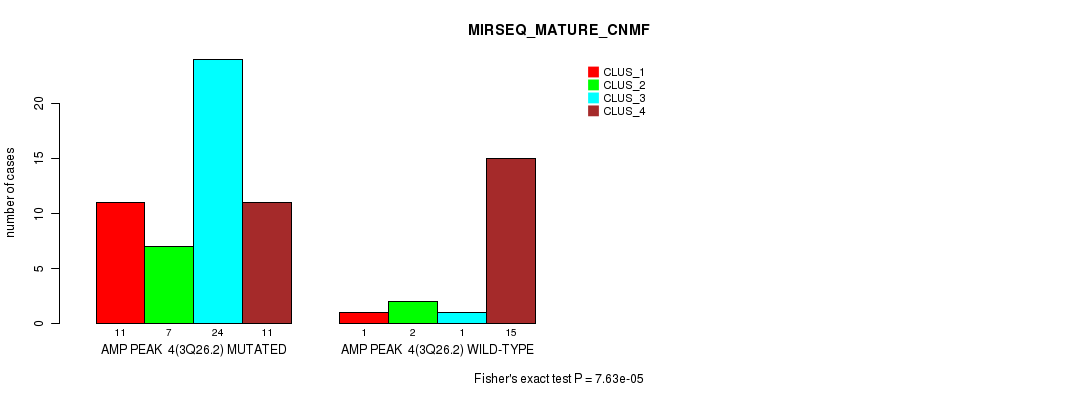
P value = 7.63e-05 (Fisher's exact test), Q value = 0.042
Table S6. Gene #4: 'amp_3q26.2' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 9 | 25 | 26 |
| AMP PEAK 4(3Q26.2) MUTATED | 11 | 7 | 24 | 11 |
| AMP PEAK 4(3Q26.2) WILD-TYPE | 1 | 2 | 1 | 15 |
Figure S6. Get High-res Image Gene #4: 'amp_3q26.2' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
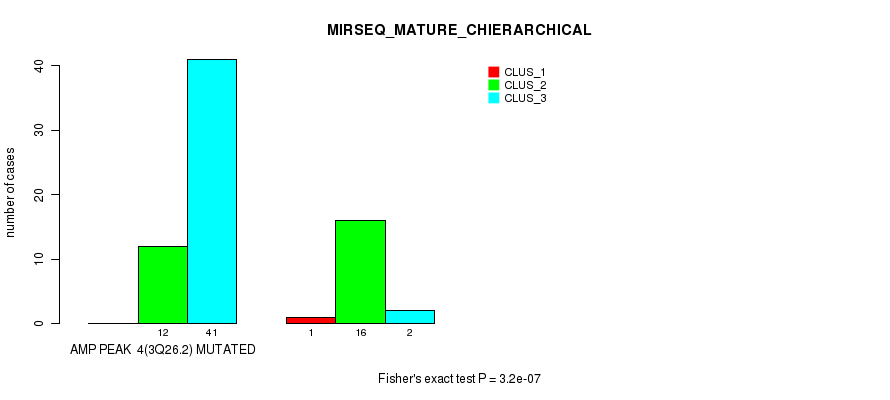
P value = 3.2e-07 (Fisher's exact test), Q value = 0.00018
Table S7. Gene #4: 'amp_3q26.2' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 1 | 28 | 43 |
| AMP PEAK 4(3Q26.2) MUTATED | 0 | 12 | 41 |
| AMP PEAK 4(3Q26.2) WILD-TYPE | 1 | 16 | 2 |
Figure S7. Get High-res Image Gene #4: 'amp_3q26.2' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
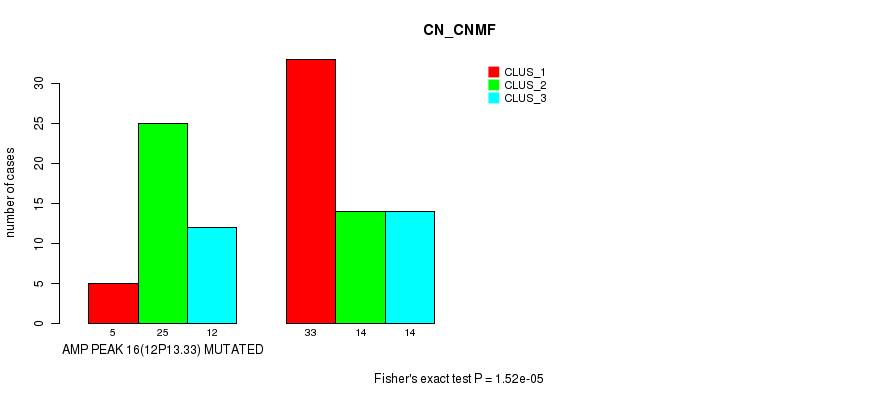
P value = 1.52e-05 (Fisher's exact test), Q value = 0.0085
Table S8. Gene #16: 'amp_12p13.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 38 | 39 | 26 |
| AMP PEAK 16(12P13.33) MUTATED | 5 | 25 | 12 |
| AMP PEAK 16(12P13.33) WILD-TYPE | 33 | 14 | 14 |
Figure S8. Get High-res Image Gene #16: 'amp_12p13.33' versus Molecular Subtype #1: 'CN_CNMF'
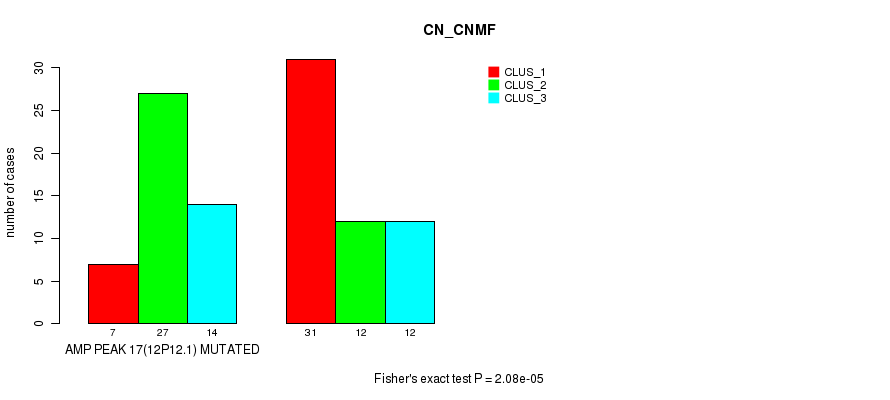
P value = 2.08e-05 (Fisher's exact test), Q value = 0.012
Table S9. Gene #17: 'amp_12p12.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 38 | 39 | 26 |
| AMP PEAK 17(12P12.1) MUTATED | 7 | 27 | 14 |
| AMP PEAK 17(12P12.1) WILD-TYPE | 31 | 12 | 12 |
Figure S9. Get High-res Image Gene #17: 'amp_12p12.1' versus Molecular Subtype #1: 'CN_CNMF'
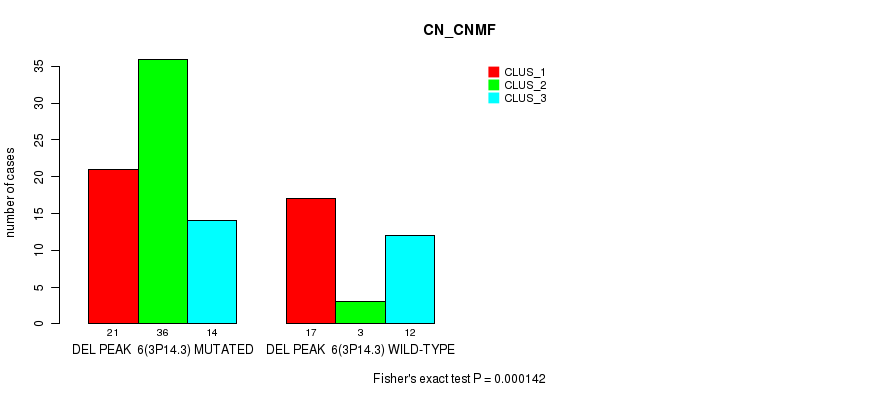
P value = 0.000142 (Fisher's exact test), Q value = 0.078
Table S10. Gene #30: 'del_3p14.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 38 | 39 | 26 |
| DEL PEAK 6(3P14.3) MUTATED | 21 | 36 | 14 |
| DEL PEAK 6(3P14.3) WILD-TYPE | 17 | 3 | 12 |
Figure S10. Get High-res Image Gene #30: 'del_3p14.3' versus Molecular Subtype #1: 'CN_CNMF'
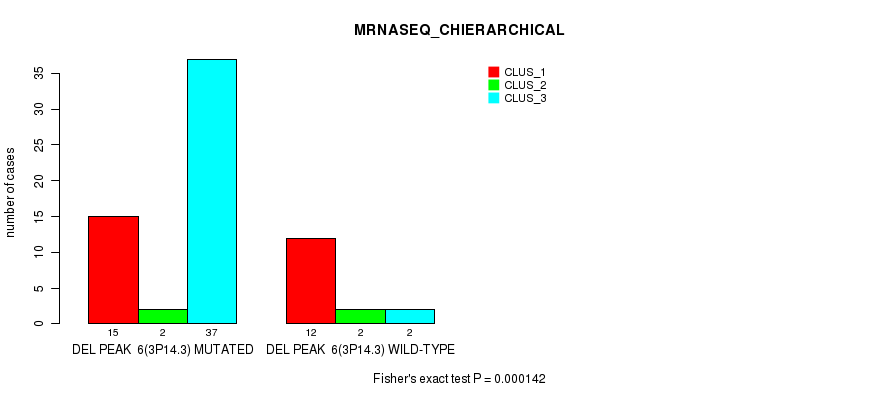
P value = 0.000142 (Fisher's exact test), Q value = 0.078
Table S11. Gene #30: 'del_3p14.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 27 | 4 | 39 |
| DEL PEAK 6(3P14.3) MUTATED | 15 | 2 | 37 |
| DEL PEAK 6(3P14.3) WILD-TYPE | 12 | 2 | 2 |
Figure S11. Get High-res Image Gene #30: 'del_3p14.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
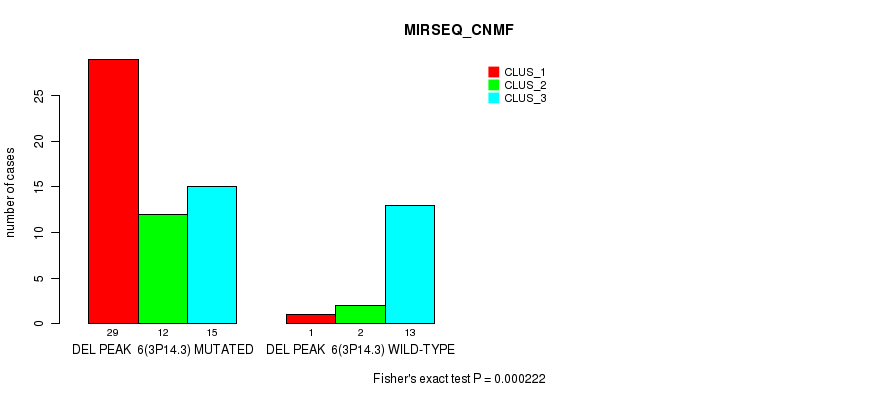
P value = 0.000222 (Fisher's exact test), Q value = 0.12
Table S12. Gene #30: 'del_3p14.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 30 | 14 | 28 |
| DEL PEAK 6(3P14.3) MUTATED | 29 | 12 | 15 |
| DEL PEAK 6(3P14.3) WILD-TYPE | 1 | 2 | 13 |
Figure S12. Get High-res Image Gene #30: 'del_3p14.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
P value = 1.23e-05 (Fisher's exact test), Q value = 0.0069
Table S13. Gene #30: 'del_3p14.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 1 | 28 | 43 |
| DEL PEAK 6(3P14.3) MUTATED | 0 | 15 | 41 |
| DEL PEAK 6(3P14.3) WILD-TYPE | 1 | 13 | 2 |
Figure S13. Get High-res Image Gene #30: 'del_3p14.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

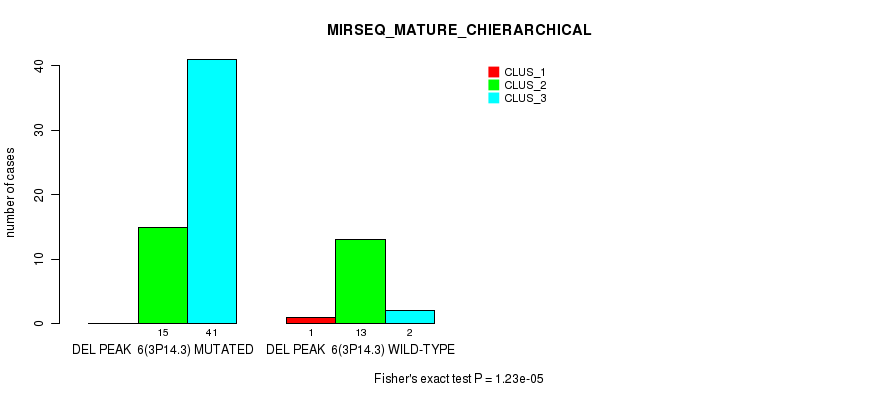
P value = 1.23e-05 (Fisher's exact test), Q value = 0.0069
Table S14. Gene #30: 'del_3p14.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 1 | 28 | 43 |
| DEL PEAK 6(3P14.3) MUTATED | 0 | 15 | 41 |
| DEL PEAK 6(3P14.3) WILD-TYPE | 1 | 13 | 2 |
Figure S14. Get High-res Image Gene #30: 'del_3p14.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
P value = 2.04e-06 (Fisher's exact test), Q value = 0.0012
Table S15. Gene #45: 'del_9p21.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 38 | 39 | 26 |
| DEL PEAK 21(9P21.3) MUTATED | 22 | 39 | 17 |
| DEL PEAK 21(9P21.3) WILD-TYPE | 16 | 0 | 9 |
Figure S15. Get High-res Image Gene #45: 'del_9p21.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00011 (Fisher's exact test), Q value = 0.061
Table S16. Gene #45: 'del_9p21.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 1 | 28 | 43 |
| DEL PEAK 21(9P21.3) MUTATED | 0 | 19 | 42 |
| DEL PEAK 21(9P21.3) WILD-TYPE | 1 | 9 | 1 |
Figure S16. Get High-res Image Gene #45: 'del_9p21.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
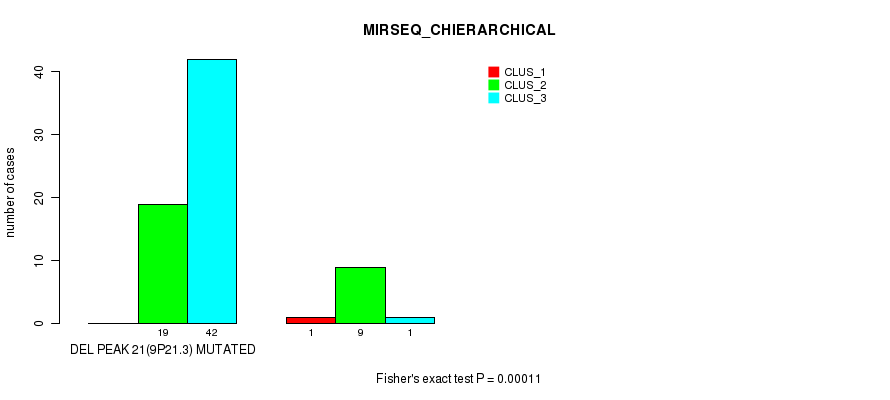
P value = 0.00011 (Fisher's exact test), Q value = 0.061
Table S17. Gene #45: 'del_9p21.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 1 | 28 | 43 |
| DEL PEAK 21(9P21.3) MUTATED | 0 | 19 | 42 |
| DEL PEAK 21(9P21.3) WILD-TYPE | 1 | 9 | 1 |
Figure S17. Get High-res Image Gene #45: 'del_9p21.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
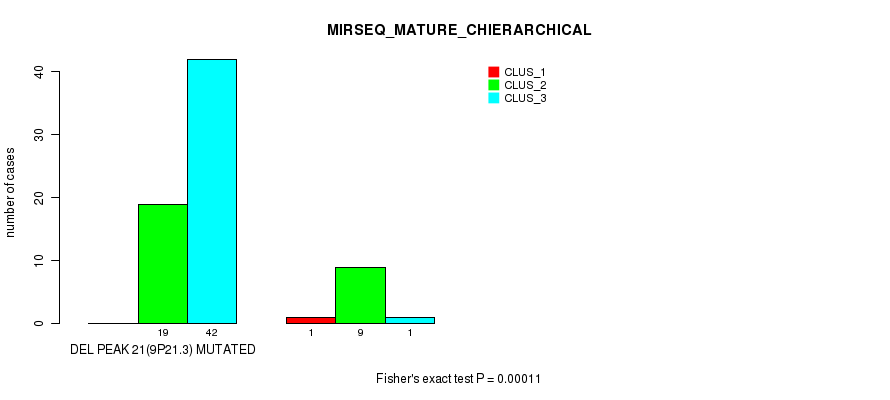
P value = 0.000135 (Fisher's exact test), Q value = 0.075
Table S18. Gene #56: 'del_16p13.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 38 | 39 | 26 |
| DEL PEAK 32(16P13.3) MUTATED | 13 | 3 | 14 |
| DEL PEAK 32(16P13.3) WILD-TYPE | 25 | 36 | 12 |
Figure S18. Get High-res Image Gene #56: 'del_16p13.3' versus Molecular Subtype #1: 'CN_CNMF'
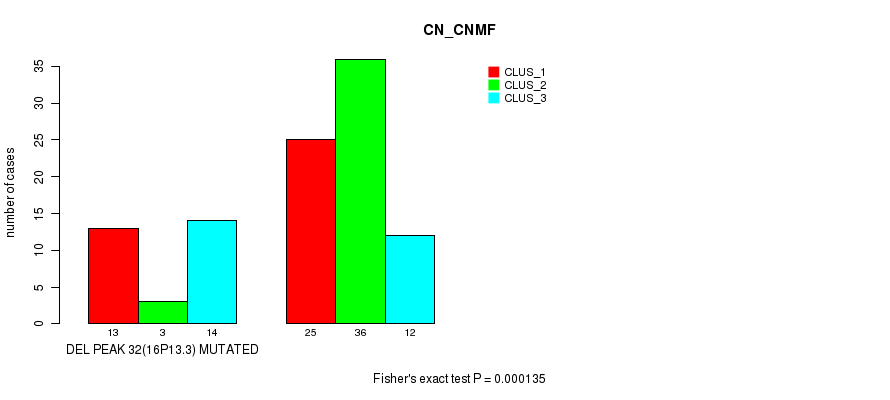
P value = 2.5e-06 (Fisher's exact test), Q value = 0.0014
Table S19. Gene #57: 'del_16q23.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 38 | 39 | 26 |
| DEL PEAK 33(16Q23.1) MUTATED | 14 | 9 | 22 |
| DEL PEAK 33(16Q23.1) WILD-TYPE | 24 | 30 | 4 |
Figure S19. Get High-res Image Gene #57: 'del_16q23.1' versus Molecular Subtype #1: 'CN_CNMF'
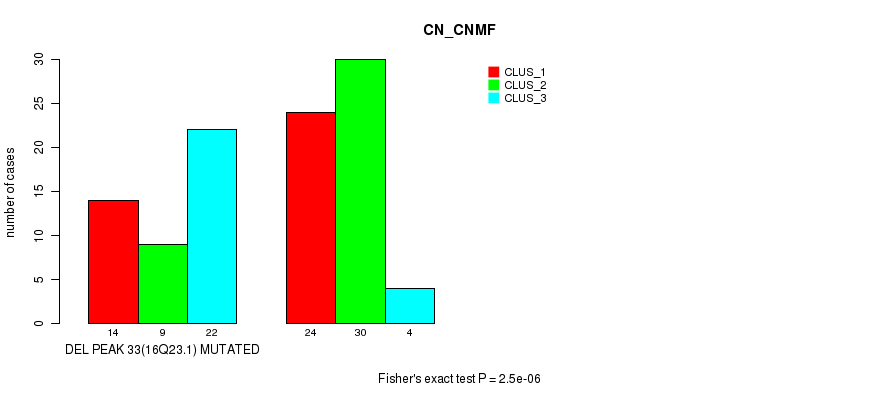
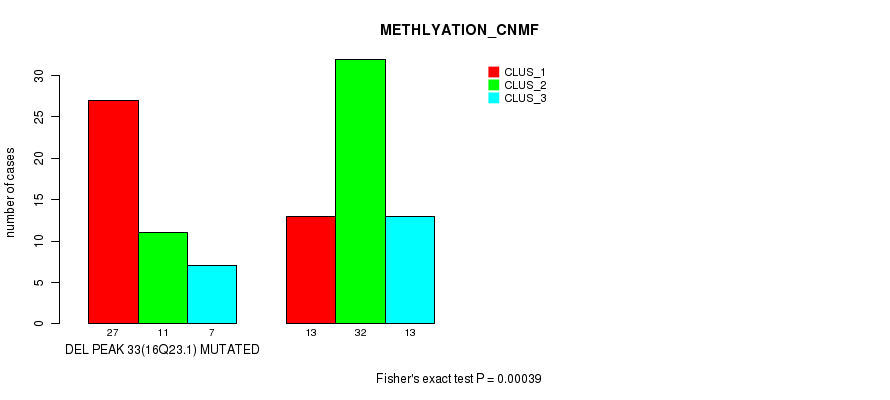
P value = 0.00039 (Fisher's exact test), Q value = 0.21
Table S20. Gene #57: 'del_16q23.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 40 | 43 | 20 |
| DEL PEAK 33(16Q23.1) MUTATED | 27 | 11 | 7 |
| DEL PEAK 33(16Q23.1) WILD-TYPE | 13 | 32 | 13 |
Figure S20. Get High-res Image Gene #57: 'del_16q23.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtype file = ESCA-TP.transferedmergedcluster.txt
-
Number of patients = 103
-
Number of significantly focal cnvs = 71
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.