This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 62 focal events and 8 molecular subtypes across 56 patients, one significant finding detected with P value < 0.05 and Q value < 0.25.
-
amp_19q12 cnv correlated to 'METHLYATION_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 62 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, one significant finding detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Chi-square test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| amp 19q12 | 45 (80%) | 11 |
0.000643 (0.318) |
0.00019 (0.0941) |
0.193 (1.00) |
0.0442 (1.00) |
0.642 (1.00) |
0.32 (1.00) |
0.443 (1.00) |
0.435 (1.00) |
| amp 1q22 | 42 (75%) | 14 |
0.896 (1.00) |
0.213 (1.00) |
0.475 (1.00) |
0.151 (1.00) |
0.179 (1.00) |
0.356 (1.00) |
0.241 (1.00) |
0.109 (1.00) |
| amp 2p14 | 28 (50%) | 28 |
0.506 (1.00) |
0.025 (1.00) |
0.00745 (1.00) |
0.0631 (1.00) |
0.463 (1.00) |
0.108 (1.00) |
0.359 (1.00) |
0.18 (1.00) |
| amp 2q13 | 29 (52%) | 27 |
0.358 (1.00) |
0.0165 (1.00) |
0.0477 (1.00) |
0.0706 (1.00) |
1 (1.00) |
0.459 (1.00) |
0.809 (1.00) |
0.64 (1.00) |
| amp 3p25 1 | 20 (36%) | 36 |
0.0286 (1.00) |
0.467 (1.00) |
0.289 (1.00) |
0.597 (1.00) |
0.219 (1.00) |
0.0122 (1.00) |
0.44 (1.00) |
0.0486 (1.00) |
| amp 3q26 2 | 37 (66%) | 19 |
0.00126 (0.621) |
0.0094 (1.00) |
0.0899 (1.00) |
0.0848 (1.00) |
0.342 (1.00) |
0.48 (1.00) |
0.704 (1.00) |
0.277 (1.00) |
| amp 4p16 3 | 25 (45%) | 31 |
0.026 (1.00) |
0.14 (1.00) |
0.38 (1.00) |
0.377 (1.00) |
0.955 (1.00) |
0.341 (1.00) |
0.854 (1.00) |
0.263 (1.00) |
| amp 5p13 2 | 26 (46%) | 30 |
0.0603 (1.00) |
0.548 (1.00) |
0.238 (1.00) |
0.475 (1.00) |
0.913 (1.00) |
0.446 (1.00) |
0.932 (1.00) |
1 (1.00) |
| amp 6p24 2 | 37 (66%) | 19 |
0.105 (1.00) |
0.967 (1.00) |
0.665 (1.00) |
0.429 (1.00) |
0.852 (1.00) |
0.264 (1.00) |
0.843 (1.00) |
0.51 (1.00) |
| amp 8p11 21 | 33 (59%) | 23 |
0.359 (1.00) |
0.6 (1.00) |
0.651 (1.00) |
0.198 (1.00) |
0.61 (1.00) |
0.765 (1.00) |
0.758 (1.00) |
1 (1.00) |
| amp 8q11 23 | 38 (68%) | 18 |
0.142 (1.00) |
0.207 (1.00) |
0.197 (1.00) |
0.122 (1.00) |
0.754 (1.00) |
1 (1.00) |
0.704 (1.00) |
0.779 (1.00) |
| amp 8q24 21 | 42 (75%) | 14 |
0.0399 (1.00) |
0.722 (1.00) |
0.189 (1.00) |
0.299 (1.00) |
0.0408 (1.00) |
0.0339 (1.00) |
0.0176 (1.00) |
0.00819 (1.00) |
| amp 8q24 21 | 40 (71%) | 16 |
0.00686 (1.00) |
0.328 (1.00) |
0.0762 (1.00) |
0.103 (1.00) |
0.0485 (1.00) |
0.0435 (1.00) |
0.0177 (1.00) |
0.00961 (1.00) |
| amp 10q22 2 | 26 (46%) | 30 |
0.232 (1.00) |
0.47 (1.00) |
0.961 (1.00) |
1 (1.00) |
0.696 (1.00) |
0.306 (1.00) |
0.609 (1.00) |
0.513 (1.00) |
| amp 11q13 1 | 19 (34%) | 37 |
0.44 (1.00) |
0.606 (1.00) |
0.332 (1.00) |
0.36 (1.00) |
0.714 (1.00) |
0.678 (1.00) |
0.655 (1.00) |
1 (1.00) |
| amp 12q12 | 22 (39%) | 34 |
0.15 (1.00) |
0.0932 (1.00) |
0.255 (1.00) |
0.421 (1.00) |
0.866 (1.00) |
0.83 (1.00) |
0.667 (1.00) |
1 (1.00) |
| amp 12q15 | 17 (30%) | 39 |
0.424 (1.00) |
0.365 (1.00) |
0.254 (1.00) |
0.774 (1.00) |
0.802 (1.00) |
0.509 (1.00) |
0.526 (1.00) |
0.723 (1.00) |
| amp 13q31 3 | 25 (45%) | 31 |
0.396 (1.00) |
0.415 (1.00) |
0.134 (1.00) |
0.0788 (1.00) |
0.0853 (1.00) |
0.00214 (1.00) |
0.0311 (1.00) |
0.0069 (1.00) |
| amp 16p11 2 | 20 (36%) | 36 |
0.00479 (1.00) |
0.281 (1.00) |
0.663 (1.00) |
0.138 (1.00) |
0.736 (1.00) |
0.546 (1.00) |
0.379 (1.00) |
0.545 (1.00) |
| amp 17q12 | 27 (48%) | 29 |
0.926 (1.00) |
0.177 (1.00) |
0.313 (1.00) |
0.433 (1.00) |
0.346 (1.00) |
0.839 (1.00) |
0.581 (1.00) |
0.565 (1.00) |
| amp 17q25 1 | 32 (57%) | 24 |
0.626 (1.00) |
0.0129 (1.00) |
0.605 (1.00) |
0.269 (1.00) |
0.666 (1.00) |
0.375 (1.00) |
0.475 (1.00) |
0.494 (1.00) |
| amp 19p13 2 | 19 (34%) | 37 |
0.338 (1.00) |
0.248 (1.00) |
0.797 (1.00) |
0.753 (1.00) |
0.331 (1.00) |
0.264 (1.00) |
0.22 (1.00) |
0.116 (1.00) |
| amp 20q11 21 | 48 (86%) | 8 |
0.0145 (1.00) |
0.186 (1.00) |
0.545 (1.00) |
0.128 (1.00) |
0.161 (1.00) |
0.0338 (1.00) |
0.13 (1.00) |
0.111 (1.00) |
| amp 20q11 21 | 48 (86%) | 8 |
0.155 (1.00) |
0.263 (1.00) |
0.195 (1.00) |
0.258 (1.00) |
0.179 (1.00) |
0.261 (1.00) |
0.148 (1.00) |
0.369 (1.00) |
| amp xp11 21 | 28 (50%) | 28 |
0.248 (1.00) |
0.543 (1.00) |
0.0481 (1.00) |
0.0719 (1.00) |
0.0287 (1.00) |
0.00689 (1.00) |
0.0789 (1.00) |
0.015 (1.00) |
| del 1p36 21 | 19 (34%) | 37 |
0.0832 (1.00) |
0.379 (1.00) |
0.337 (1.00) |
0.467 (1.00) |
0.136 (1.00) |
0.00802 (1.00) |
0.123 (1.00) |
0.0083 (1.00) |
| del 2q22 1 | 11 (20%) | 45 |
0.502 (1.00) |
0.507 (1.00) |
0.527 (1.00) |
0.372 (1.00) |
0.552 (1.00) |
0.211 (1.00) |
0.282 (1.00) |
0.142 (1.00) |
| del 3p21 1 | 37 (66%) | 19 |
0.431 (1.00) |
0.418 (1.00) |
0.467 (1.00) |
0.399 (1.00) |
0.504 (1.00) |
1 (1.00) |
0.497 (1.00) |
0.624 (1.00) |
| del 3p14 1 | 30 (54%) | 26 |
1 (1.00) |
0.637 (1.00) |
0.173 (1.00) |
0.359 (1.00) |
0.089 (1.00) |
0.775 (1.00) |
0.207 (1.00) |
0.455 (1.00) |
| del 3q13 31 | 19 (34%) | 37 |
0.112 (1.00) |
0.118 (1.00) |
0.923 (1.00) |
1 (1.00) |
0.94 (1.00) |
0.387 (1.00) |
0.672 (1.00) |
0.383 (1.00) |
| del 4q22 1 | 37 (66%) | 19 |
0.0619 (1.00) |
0.0486 (1.00) |
0.0253 (1.00) |
0.0442 (1.00) |
0.014 (1.00) |
0.299 (1.00) |
0.0288 (1.00) |
0.0817 (1.00) |
| del 4q34 3 | 37 (66%) | 19 |
0.00379 (1.00) |
0.0185 (1.00) |
0.0112 (1.00) |
0.0442 (1.00) |
0.00412 (1.00) |
0.621 (1.00) |
0.00956 (1.00) |
0.3 (1.00) |
| del 5q11 2 | 26 (46%) | 30 |
0.133 (1.00) |
0.0971 (1.00) |
0.597 (1.00) |
0.129 (1.00) |
0.541 (1.00) |
0.446 (1.00) |
0.0816 (1.00) |
1 (1.00) |
| del 6q26 | 10 (18%) | 46 |
0.528 (1.00) |
0.519 (1.00) |
0.122 (1.00) |
0.751 (1.00) |
0.329 (1.00) |
0.196 (1.00) |
0.174 (1.00) |
0.252 (1.00) |
| del 7q11 22 | 12 (21%) | 44 |
0.262 (1.00) |
0.44 (1.00) |
0.76 (1.00) |
0.813 (1.00) |
0.861 (1.00) |
0.684 (1.00) |
0.808 (1.00) |
1 (1.00) |
| del 7q36 2 | 20 (36%) | 36 |
0.156 (1.00) |
0.171 (1.00) |
0.483 (1.00) |
0.473 (1.00) |
0.372 (1.00) |
0.057 (1.00) |
0.528 (1.00) |
0.175 (1.00) |
| del 8p21 3 | 32 (57%) | 24 |
0.000664 (0.328) |
0.455 (1.00) |
0.251 (1.00) |
0.286 (1.00) |
0.789 (1.00) |
0.375 (1.00) |
0.421 (1.00) |
0.825 (1.00) |
| del 9p23 | 34 (61%) | 22 |
0.0919 (1.00) |
0.135 (1.00) |
0.619 (1.00) |
0.624 (1.00) |
0.482 (1.00) |
0.146 (1.00) |
0.839 (1.00) |
0.591 (1.00) |
| del 9q21 13 | 39 (70%) | 17 |
0.112 (1.00) |
0.306 (1.00) |
0.211 (1.00) |
0.382 (1.00) |
0.467 (1.00) |
0.513 (1.00) |
0.654 (1.00) |
0.532 (1.00) |
| del 9q33 3 | 38 (68%) | 18 |
0.442 (1.00) |
0.11 (1.00) |
0.088 (1.00) |
0.294 (1.00) |
1 (1.00) |
0.733 (1.00) |
0.785 (1.00) |
1 (1.00) |
| del 10q21 1 | 18 (32%) | 38 |
0.505 (1.00) |
0.392 (1.00) |
0.601 (1.00) |
0.538 (1.00) |
0.449 (1.00) |
0.299 (1.00) |
0.302 (1.00) |
0.3 (1.00) |
| del 10q23 31 | 20 (36%) | 36 |
0.143 (1.00) |
0.126 (1.00) |
0.119 (1.00) |
0.191 (1.00) |
0.736 (1.00) |
1 (1.00) |
0.581 (1.00) |
0.666 (1.00) |
| del 11p15 5 | 25 (45%) | 31 |
0.192 (1.00) |
0.739 (1.00) |
0.884 (1.00) |
0.951 (1.00) |
0.696 (1.00) |
0.761 (1.00) |
0.235 (1.00) |
0.513 (1.00) |
| del 11q14 1 | 26 (46%) | 30 |
0.616 (1.00) |
0.202 (1.00) |
1 (1.00) |
1 (1.00) |
0.259 (1.00) |
0.186 (1.00) |
0.251 (1.00) |
0.309 (1.00) |
| del 11q24 2 | 27 (48%) | 29 |
0.58 (1.00) |
0.0343 (1.00) |
0.134 (1.00) |
0.696 (1.00) |
0.246 (1.00) |
0.286 (1.00) |
0.236 (1.00) |
0.424 (1.00) |
| del 12q23 1 | 20 (36%) | 36 |
0.64 (1.00) |
0.878 (1.00) |
0.663 (1.00) |
0.454 (1.00) |
0.922 (1.00) |
0.0524 (1.00) |
0.912 (1.00) |
0.308 (1.00) |
| del 12q24 31 | 20 (36%) | 36 |
0.774 (1.00) |
0.826 (1.00) |
0.153 (1.00) |
0.492 (1.00) |
0.577 (1.00) |
0.151 (1.00) |
0.152 (1.00) |
0.308 (1.00) |
| del 13q12 11 | 30 (54%) | 26 |
0.428 (1.00) |
0.0184 (1.00) |
0.366 (1.00) |
0.314 (1.00) |
0.696 (1.00) |
0.112 (1.00) |
0.478 (1.00) |
0.211 (1.00) |
| del 13q14 2 | 32 (57%) | 24 |
0.575 (1.00) |
0.000796 (0.392) |
0.0135 (1.00) |
0.0347 (1.00) |
0.457 (1.00) |
0.124 (1.00) |
0.134 (1.00) |
0.0293 (1.00) |
| del 14q21 1 | 28 (50%) | 28 |
0.732 (1.00) |
0.0539 (1.00) |
0.0608 (1.00) |
0.117 (1.00) |
0.373 (1.00) |
0.405 (1.00) |
0.342 (1.00) |
0.542 (1.00) |
| del 15q15 1 | 43 (77%) | 13 |
0.0403 (1.00) |
0.0367 (1.00) |
0.348 (1.00) |
0.378 (1.00) |
0.112 (1.00) |
1 (1.00) |
0.0481 (1.00) |
0.757 (1.00) |
| del 16p13 3 | 33 (59%) | 23 |
0.175 (1.00) |
0.527 (1.00) |
0.562 (1.00) |
0.429 (1.00) |
0.339 (1.00) |
1 (1.00) |
0.585 (1.00) |
1 (1.00) |
| del 16q23 1 | 39 (70%) | 17 |
0.152 (1.00) |
0.396 (1.00) |
0.395 (1.00) |
0.305 (1.00) |
0.86 (1.00) |
1 (1.00) |
0.451 (1.00) |
0.652 (1.00) |
| del 17p13 1 | 42 (75%) | 14 |
0.0629 (1.00) |
0.0586 (1.00) |
0.0581 (1.00) |
0.113 (1.00) |
0.132 (1.00) |
0.443 (1.00) |
0.06 (1.00) |
0.393 (1.00) |
| del 17q21 32 | 19 (34%) | 37 |
0.899 (1.00) |
0.753 (1.00) |
0.182 (1.00) |
0.0301 (1.00) |
0.215 (1.00) |
0.745 (1.00) |
0.197 (1.00) |
0.548 (1.00) |
| del 18q22 2 | 26 (46%) | 30 |
0.12 (1.00) |
0.341 (1.00) |
0.472 (1.00) |
0.314 (1.00) |
0.0231 (1.00) |
0.0153 (1.00) |
0.0887 (1.00) |
0.00746 (1.00) |
| del 19p13 3 | 47 (84%) | 9 |
0.14 (1.00) |
0.0279 (1.00) |
0.0984 (1.00) |
0.164 (1.00) |
0.0149 (1.00) |
0.0338 (1.00) |
0.00828 (1.00) |
0.111 (1.00) |
| del 19q13 33 | 17 (30%) | 39 |
0.0356 (1.00) |
0.00903 (1.00) |
0.000748 (0.369) |
0.0107 (1.00) |
0.227 (1.00) |
0.451 (1.00) |
0.526 (1.00) |
0.457 (1.00) |
| del 20p12 1 | 9 (16%) | 47 |
1 (1.00) |
0.671 (1.00) |
0.391 (1.00) |
0.284 (1.00) |
0.22 (1.00) |
0.403 (1.00) |
0.317 (1.00) |
0.292 (1.00) |
| del 22q13 31 | 40 (71%) | 16 |
0.563 (1.00) |
0.4 (1.00) |
0.893 (1.00) |
0.951 (1.00) |
0.548 (1.00) |
0.388 (1.00) |
0.479 (1.00) |
0.384 (1.00) |
| del xp21 1 | 18 (32%) | 38 |
0.556 (1.00) |
0.383 (1.00) |
0.82 (1.00) |
0.814 (1.00) |
0.36 (1.00) |
0.784 (1.00) |
0.734 (1.00) |
0.779 (1.00) |
| del xq25 | 23 (41%) | 33 |
0.515 (1.00) |
0.297 (1.00) |
0.112 (1.00) |
0.306 (1.00) |
0.692 (1.00) |
1 (1.00) |
0.537 (1.00) |
1 (1.00) |
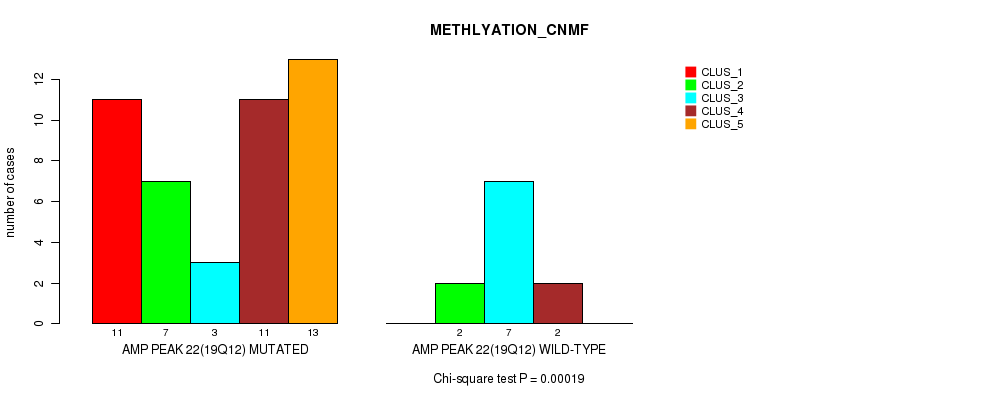
P value = 0.00019 (Chi-square test), Q value = 0.094
Table S1. Gene #22: 'amp_19q12' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 11 | 9 | 10 | 13 | 13 |
| AMP PEAK 22(19Q12) MUTATED | 11 | 7 | 3 | 11 | 13 |
| AMP PEAK 22(19Q12) WILD-TYPE | 0 | 2 | 7 | 2 | 0 |
Figure S1. Get High-res Image Gene #22: 'amp_19q12' versus Molecular Subtype #2: 'METHLYATION_CNMF'
-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtype file = UCS-TP.transferedmergedcluster.txt
-
Number of patients = 56
-
Number of significantly focal cnvs = 62
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multi-class clinical features (nominal or ordinal), Chi-square tests (Greenwood and Nikulin 1996) were used to estimate the P values using the 'chisq.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.