This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 45 focal events and 8 molecular subtypes across 90 patients, 31 significant findings detected with P value < 0.05 and Q value < 0.25.
-
amp_1q22 cnv correlated to 'METHLYATION_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
amp_4q35.1 cnv correlated to 'CN_CNMF' and 'MIRSEQ_CHIERARCHICAL'.
-
amp_5p15.33 cnv correlated to 'CN_CNMF'.
-
amp_5q35.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
amp_9q31.3 cnv correlated to 'CN_CNMF'.
-
amp_12q14.1 cnv correlated to 'CN_CNMF'.
-
amp_14q11.2 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_MATURE_CNMF'.
-
amp_16p13.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
amp_16q22.1 cnv correlated to 'CN_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
amp_16q24.2 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
amp_19p13.12 cnv correlated to 'MRNASEQ_CHIERARCHICAL'.
-
amp_xp11.22 cnv correlated to 'CN_CNMF'.
-
amp_xq28 cnv correlated to 'CN_CNMF'.
-
del_9p23 cnv correlated to 'MRNASEQ_CNMF'.
-
del_9p21.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_11q14.1 cnv correlated to 'CN_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 45 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 31 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| amp 14q11 2 | 27 (30%) | 63 |
0.00011 (0.0378) |
0.0121 (1.00) |
5e-05 (0.0173) |
2e-05 (0.00702) |
0.00043 (0.144) |
0.00693 (1.00) |
0.00021 (0.0712) |
0.0054 (1.00) |
| amp 5q35 3 | 63 (70%) | 27 |
1e-05 (0.0036) |
0.00139 (0.446) |
1e-05 (0.0036) |
1e-05 (0.0036) |
0.00652 (1.00) |
0.0466 (1.00) |
0.0258 (1.00) |
0.161 (1.00) |
| amp 16p13 3 | 50 (56%) | 40 |
1e-05 (0.0036) |
0.261 (1.00) |
0.00016 (0.0547) |
1e-05 (0.0036) |
0.0778 (1.00) |
0.00915 (1.00) |
0.0156 (1.00) |
0.0469 (1.00) |
| amp 16q24 2 | 49 (54%) | 41 |
1e-05 (0.0036) |
0.0267 (1.00) |
0.00026 (0.0876) |
1e-05 (0.0036) |
0.0635 (1.00) |
0.00283 (0.866) |
0.0288 (1.00) |
0.0497 (1.00) |
| del 9p21 3 | 27 (30%) | 63 |
0.0005 (0.166) |
0.00364 (1.00) |
2e-05 (0.00702) |
0.00053 (0.176) |
0.0175 (1.00) |
0.181 (1.00) |
0.0727 (1.00) |
0.153 (1.00) |
| amp 1q22 | 17 (19%) | 73 |
0.0101 (1.00) |
0.00053 (0.176) |
0.00264 (0.816) |
0.00018 (0.0614) |
0.116 (1.00) |
0.157 (1.00) |
0.0763 (1.00) |
0.0965 (1.00) |
| amp 4q35 1 | 29 (32%) | 61 |
2e-05 (0.00702) |
0.196 (1.00) |
0.00078 (0.256) |
0.00081 (0.265) |
0.0567 (1.00) |
0.00075 (0.247) |
0.178 (1.00) |
0.00258 (0.802) |
| amp 16q22 1 | 56 (62%) | 34 |
1e-05 (0.0036) |
0.37 (1.00) |
0.0279 (1.00) |
0.00024 (0.0811) |
0.0662 (1.00) |
0.0229 (1.00) |
0.0386 (1.00) |
0.402 (1.00) |
| amp 5p15 33 | 65 (72%) | 25 |
1e-05 (0.0036) |
0.882 (1.00) |
0.00991 (1.00) |
0.00086 (0.28) |
0.0443 (1.00) |
0.181 (1.00) |
0.112 (1.00) |
0.0593 (1.00) |
| amp 9q31 3 | 34 (38%) | 56 |
0.00012 (0.0412) |
0.0202 (1.00) |
0.0275 (1.00) |
0.0194 (1.00) |
0.0543 (1.00) |
0.199 (1.00) |
0.0411 (1.00) |
0.158 (1.00) |
| amp 12q14 1 | 70 (78%) | 20 |
3e-05 (0.0104) |
0.283 (1.00) |
0.0222 (1.00) |
0.00191 (0.605) |
0.46 (1.00) |
0.489 (1.00) |
0.0769 (1.00) |
0.196 (1.00) |
| amp 19p13 12 | 58 (64%) | 32 |
0.00104 (0.337) |
0.0349 (1.00) |
0.002 (0.63) |
0.00045 (0.15) |
0.324 (1.00) |
0.481 (1.00) |
0.271 (1.00) |
0.54 (1.00) |
| amp xp11 22 | 49 (54%) | 41 |
9e-05 (0.031) |
0.92 (1.00) |
0.0331 (1.00) |
0.0966 (1.00) |
0.761 (1.00) |
0.145 (1.00) |
0.875 (1.00) |
0.00862 (1.00) |
| amp xq28 | 46 (51%) | 44 |
0.00018 (0.0614) |
0.756 (1.00) |
0.00874 (1.00) |
0.0118 (1.00) |
0.513 (1.00) |
0.263 (1.00) |
0.733 (1.00) |
0.0433 (1.00) |
| del 9p23 | 24 (27%) | 66 |
0.00077 (0.253) |
0.00804 (1.00) |
4e-05 (0.0139) |
0.00152 (0.485) |
0.00792 (1.00) |
0.172 (1.00) |
0.0494 (1.00) |
0.182 (1.00) |
| del 11q14 1 | 25 (28%) | 65 |
0.00035 (0.118) |
0.00685 (1.00) |
0.217 (1.00) |
0.35 (1.00) |
0.279 (1.00) |
0.741 (1.00) |
0.443 (1.00) |
0.871 (1.00) |
| amp 4p16 3 | 41 (46%) | 49 |
0.00226 (0.707) |
0.547 (1.00) |
0.0131 (1.00) |
0.00562 (1.00) |
0.489 (1.00) |
0.0462 (1.00) |
0.561 (1.00) |
0.387 (1.00) |
| amp 6p21 31 | 19 (21%) | 71 |
0.126 (1.00) |
0.798 (1.00) |
0.0813 (1.00) |
1 (1.00) |
0.139 (1.00) |
0.0268 (1.00) |
0.147 (1.00) |
0.00576 (1.00) |
| amp 6q24 3 | 19 (21%) | 71 |
0.0088 (1.00) |
0.91 (1.00) |
0.0475 (1.00) |
0.935 (1.00) |
0.668 (1.00) |
0.23 (1.00) |
0.816 (1.00) |
0.0563 (1.00) |
| amp 7p22 1 | 54 (60%) | 36 |
0.00808 (1.00) |
0.484 (1.00) |
0.00772 (1.00) |
0.0333 (1.00) |
0.572 (1.00) |
0.492 (1.00) |
0.435 (1.00) |
0.696 (1.00) |
| amp 17q25 3 | 18 (20%) | 72 |
0.106 (1.00) |
0.689 (1.00) |
0.127 (1.00) |
0.671 (1.00) |
0.397 (1.00) |
0.692 (1.00) |
0.539 (1.00) |
0.915 (1.00) |
| amp 19q12 | 56 (62%) | 34 |
0.0029 (0.884) |
0.02 (1.00) |
0.00215 (0.675) |
0.00097 (0.315) |
0.196 (1.00) |
0.252 (1.00) |
0.128 (1.00) |
0.407 (1.00) |
| del 1p36 23 | 37 (41%) | 53 |
0.0301 (1.00) |
0.0116 (1.00) |
0.139 (1.00) |
0.00968 (1.00) |
0.955 (1.00) |
0.686 (1.00) |
0.668 (1.00) |
0.734 (1.00) |
| del 1q43 | 18 (20%) | 72 |
0.11 (1.00) |
0.0451 (1.00) |
0.59 (1.00) |
0.139 (1.00) |
0.526 (1.00) |
0.55 (1.00) |
0.66 (1.00) |
0.538 (1.00) |
| del 2q22 1 | 18 (20%) | 72 |
0.00278 (0.853) |
0.00316 (0.961) |
0.239 (1.00) |
0.0903 (1.00) |
0.314 (1.00) |
0.665 (1.00) |
0.386 (1.00) |
0.702 (1.00) |
| del 2q37 3 | 21 (23%) | 69 |
0.00273 (0.841) |
0.0261 (1.00) |
0.296 (1.00) |
0.0916 (1.00) |
0.481 (1.00) |
0.525 (1.00) |
0.662 (1.00) |
0.967 (1.00) |
| del 3q13 31 | 23 (26%) | 67 |
0.0233 (1.00) |
0.304 (1.00) |
0.0132 (1.00) |
0.0877 (1.00) |
0.27 (1.00) |
0.372 (1.00) |
0.0839 (1.00) |
0.076 (1.00) |
| del 4q34 3 | 25 (28%) | 65 |
0.00497 (1.00) |
0.0643 (1.00) |
0.0216 (1.00) |
0.0379 (1.00) |
0.038 (1.00) |
0.0421 (1.00) |
0.116 (1.00) |
0.06 (1.00) |
| del 4q35 1 | 28 (31%) | 62 |
0.0408 (1.00) |
0.0397 (1.00) |
0.0959 (1.00) |
0.0774 (1.00) |
0.32 (1.00) |
0.139 (1.00) |
0.557 (1.00) |
0.172 (1.00) |
| del 6p24 3 | 20 (22%) | 70 |
0.00335 (1.00) |
0.92 (1.00) |
0.623 (1.00) |
0.482 (1.00) |
1 (1.00) |
0.101 (1.00) |
0.878 (1.00) |
0.0822 (1.00) |
| del 6q26 | 21 (23%) | 69 |
0.0185 (1.00) |
0.862 (1.00) |
0.322 (1.00) |
0.307 (1.00) |
0.505 (1.00) |
0.345 (1.00) |
0.419 (1.00) |
0.0284 (1.00) |
| del 7q32 3 | 9 (10%) | 81 |
0.153 (1.00) |
0.0284 (1.00) |
0.175 (1.00) |
0.167 (1.00) |
0.895 (1.00) |
0.737 (1.00) |
1 (1.00) |
0.863 (1.00) |
| del 10q23 1 | 11 (12%) | 79 |
0.0434 (1.00) |
0.259 (1.00) |
0.233 (1.00) |
0.749 (1.00) |
1 (1.00) |
0.969 (1.00) |
0.889 (1.00) |
0.775 (1.00) |
| del 11p15 5 | 27 (30%) | 63 |
0.00192 (0.607) |
0.0196 (1.00) |
0.0883 (1.00) |
0.0333 (1.00) |
0.125 (1.00) |
0.597 (1.00) |
0.224 (1.00) |
0.736 (1.00) |
| del 12q14 2 | 5 (6%) | 85 |
0.23 (1.00) |
0.221 (1.00) |
0.177 (1.00) |
0.00107 (0.346) |
0.596 (1.00) |
0.305 (1.00) |
0.104 (1.00) |
0.00146 (0.467) |
| del 13q14 2 | 42 (47%) | 48 |
0.00262 (0.812) |
0.0874 (1.00) |
0.333 (1.00) |
0.557 (1.00) |
1 (1.00) |
0.531 (1.00) |
0.914 (1.00) |
0.629 (1.00) |
| del 14q21 2 | 20 (22%) | 70 |
0.0112 (1.00) |
0.924 (1.00) |
0.303 (1.00) |
0.285 (1.00) |
0.368 (1.00) |
0.736 (1.00) |
0.205 (1.00) |
0.788 (1.00) |
| del 16p13 3 | 7 (8%) | 83 |
0.119 (1.00) |
0.558 (1.00) |
0.541 (1.00) |
0.00806 (1.00) |
0.0237 (1.00) |
0.6 (1.00) |
0.0457 (1.00) |
0.585 (1.00) |
| del 16q23 1 | 9 (10%) | 81 |
0.0427 (1.00) |
0.658 (1.00) |
0.0169 (1.00) |
0.0783 (1.00) |
0.141 (1.00) |
0.591 (1.00) |
0.298 (1.00) |
0.363 (1.00) |
| del 17q11 2 | 24 (27%) | 66 |
0.00367 (1.00) |
0.00544 (1.00) |
0.104 (1.00) |
0.106 (1.00) |
0.26 (1.00) |
0.0185 (1.00) |
0.235 (1.00) |
0.0623 (1.00) |
| del 17q21 31 | 26 (29%) | 64 |
0.00244 (0.761) |
0.00923 (1.00) |
0.0782 (1.00) |
0.0472 (1.00) |
0.0346 (1.00) |
0.00479 (1.00) |
0.046 (1.00) |
0.0931 (1.00) |
| del 17q24 2 | 24 (27%) | 66 |
0.00111 (0.357) |
0.0135 (1.00) |
0.16 (1.00) |
0.0404 (1.00) |
0.157 (1.00) |
0.0272 (1.00) |
0.158 (1.00) |
0.317 (1.00) |
| del 18q21 2 | 36 (40%) | 54 |
0.0752 (1.00) |
0.321 (1.00) |
0.825 (1.00) |
1 (1.00) |
0.494 (1.00) |
0.915 (1.00) |
0.761 (1.00) |
0.0982 (1.00) |
| del 20p12 1 | 12 (13%) | 78 |
0.00188 (0.598) |
0.642 (1.00) |
0.0429 (1.00) |
0.0262 (1.00) |
0.00631 (1.00) |
0.516 (1.00) |
0.0144 (1.00) |
0.582 (1.00) |
| del 22q12 1 | 50 (56%) | 40 |
0.0397 (1.00) |
0.139 (1.00) |
0.746 (1.00) |
0.183 (1.00) |
0.637 (1.00) |
0.88 (1.00) |
0.512 (1.00) |
0.814 (1.00) |
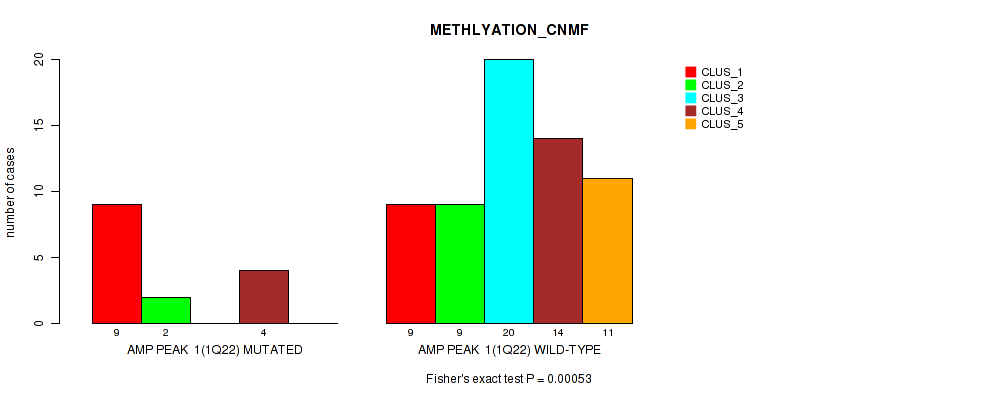
P value = 0.00053 (Fisher's exact test), Q value = 0.18
Table S1. Gene #1: 'amp_1q22' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 18 | 11 | 20 | 18 | 11 |
| AMP PEAK 1(1Q22) MUTATED | 9 | 2 | 0 | 4 | 0 |
| AMP PEAK 1(1Q22) WILD-TYPE | 9 | 9 | 20 | 14 | 11 |
Figure S1. Get High-res Image Gene #1: 'amp_1q22' versus Molecular Subtype #2: 'METHLYATION_CNMF'
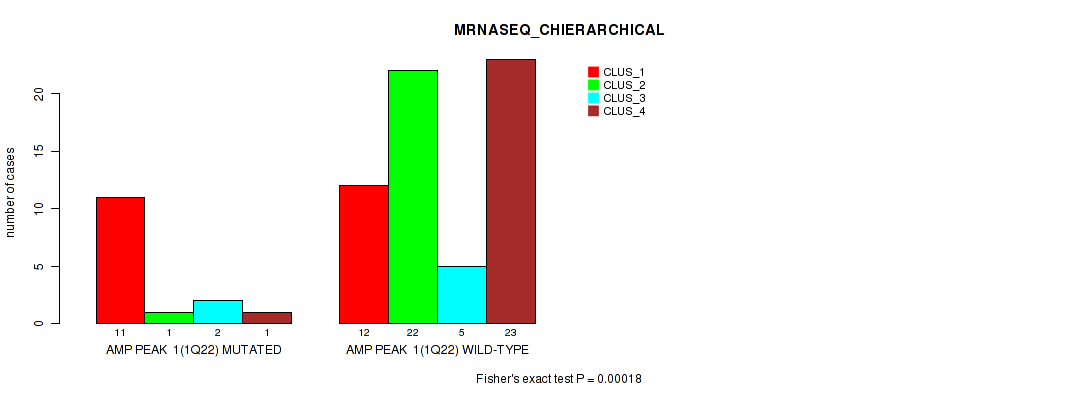
P value = 0.00018 (Fisher's exact test), Q value = 0.061
Table S2. Gene #1: 'amp_1q22' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 23 | 7 | 24 |
| AMP PEAK 1(1Q22) MUTATED | 11 | 1 | 2 | 1 |
| AMP PEAK 1(1Q22) WILD-TYPE | 12 | 22 | 5 | 23 |
Figure S2. Get High-res Image Gene #1: 'amp_1q22' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
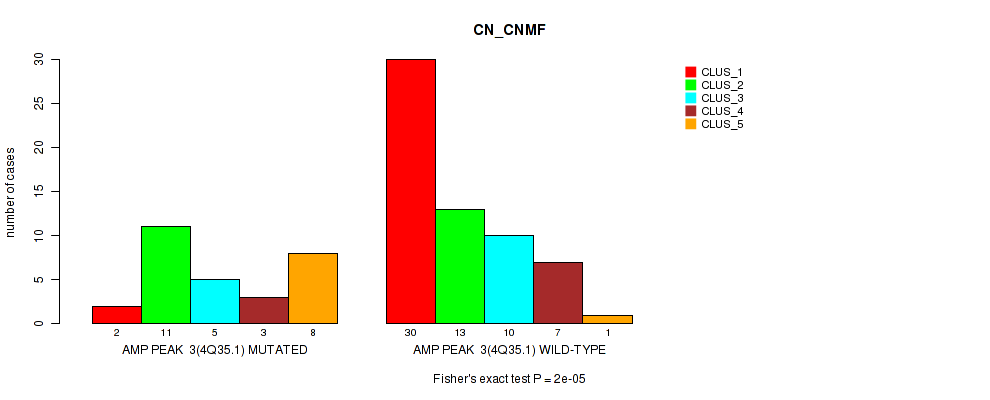
P value = 2e-05 (Fisher's exact test), Q value = 0.007
Table S3. Gene #3: 'amp_4q35.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 3(4Q35.1) MUTATED | 2 | 11 | 5 | 3 | 8 |
| AMP PEAK 3(4Q35.1) WILD-TYPE | 30 | 13 | 10 | 7 | 1 |
Figure S3. Get High-res Image Gene #3: 'amp_4q35.1' versus Molecular Subtype #1: 'CN_CNMF'
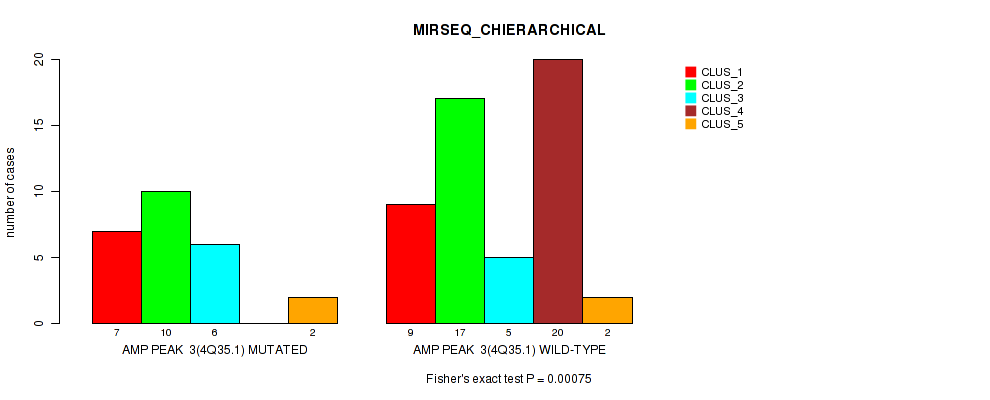
P value = 0.00075 (Fisher's exact test), Q value = 0.25
Table S4. Gene #3: 'amp_4q35.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 16 | 27 | 11 | 20 | 4 |
| AMP PEAK 3(4Q35.1) MUTATED | 7 | 10 | 6 | 0 | 2 |
| AMP PEAK 3(4Q35.1) WILD-TYPE | 9 | 17 | 5 | 20 | 2 |
Figure S4. Get High-res Image Gene #3: 'amp_4q35.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S5. Gene #4: 'amp_5p15.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 4(5P15.33) MUTATED | 14 | 23 | 14 | 6 | 8 |
| AMP PEAK 4(5P15.33) WILD-TYPE | 18 | 1 | 1 | 4 | 1 |
Figure S5. Get High-res Image Gene #4: 'amp_5p15.33' versus Molecular Subtype #1: 'CN_CNMF'
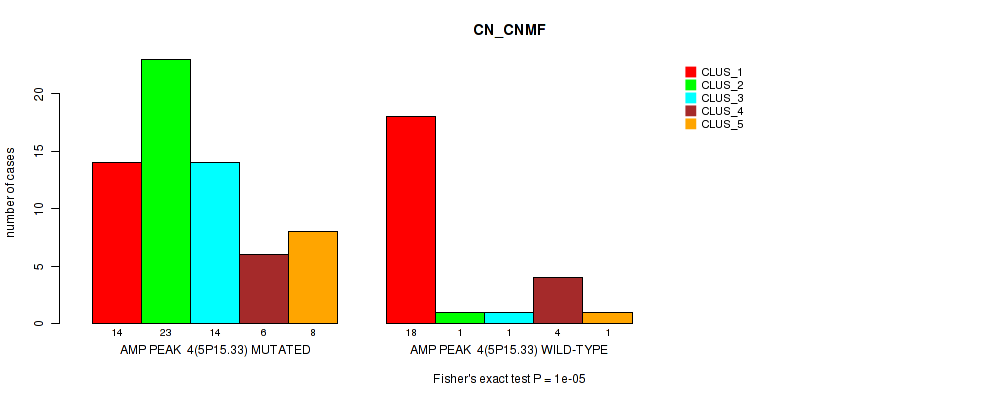
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S6. Gene #5: 'amp_5q35.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 5(5Q35.3) MUTATED | 10 | 24 | 14 | 7 | 8 |
| AMP PEAK 5(5Q35.3) WILD-TYPE | 22 | 0 | 1 | 3 | 1 |
Figure S6. Get High-res Image Gene #5: 'amp_5q35.3' versus Molecular Subtype #1: 'CN_CNMF'
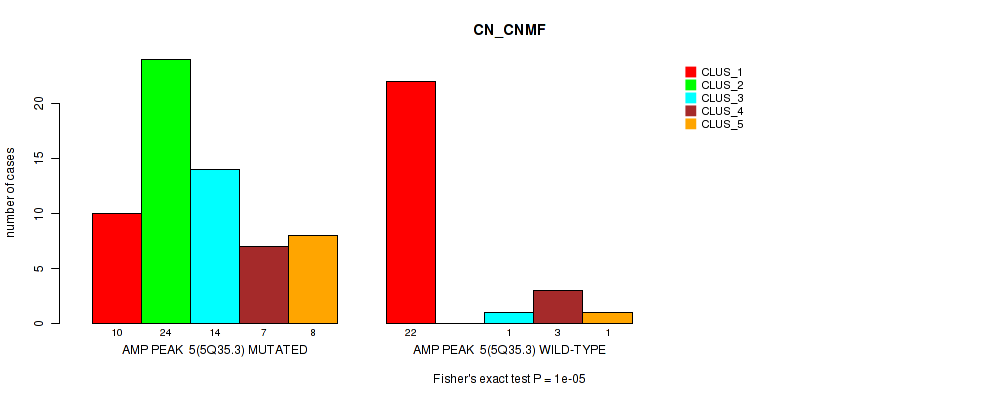
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S7. Gene #5: 'amp_5q35.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| AMP PEAK 5(5Q35.3) MUTATED | 7 | 12 | 16 | 20 |
| AMP PEAK 5(5Q35.3) WILD-TYPE | 17 | 1 | 1 | 3 |
Figure S7. Get High-res Image Gene #5: 'amp_5q35.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
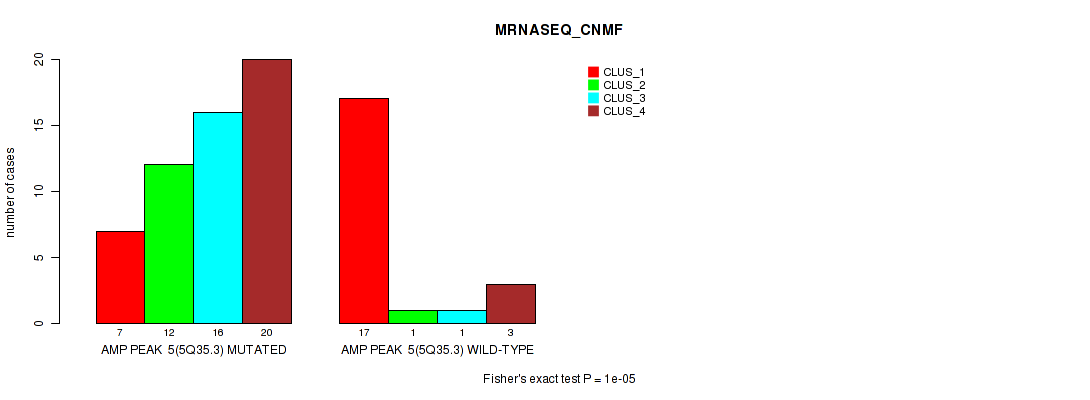
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S8. Gene #5: 'amp_5q35.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 23 | 7 | 24 |
| AMP PEAK 5(5Q35.3) MUTATED | 8 | 22 | 2 | 23 |
| AMP PEAK 5(5Q35.3) WILD-TYPE | 15 | 1 | 5 | 1 |
Figure S8. Get High-res Image Gene #5: 'amp_5q35.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
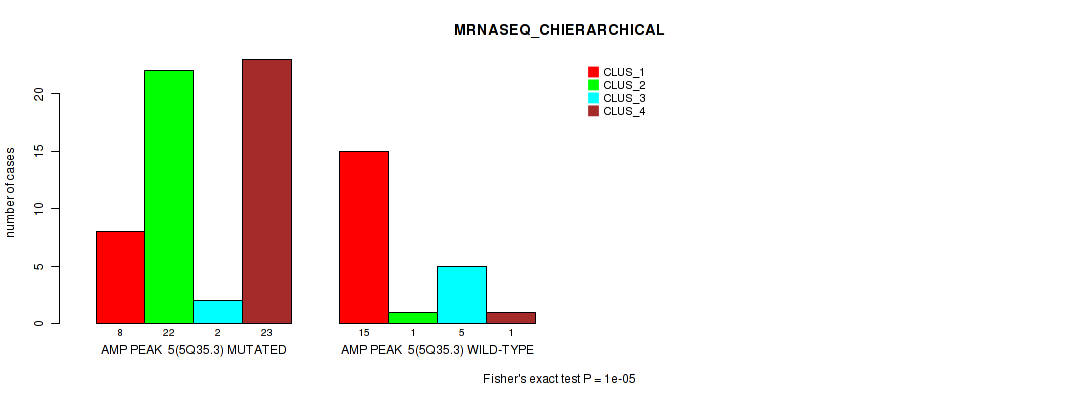
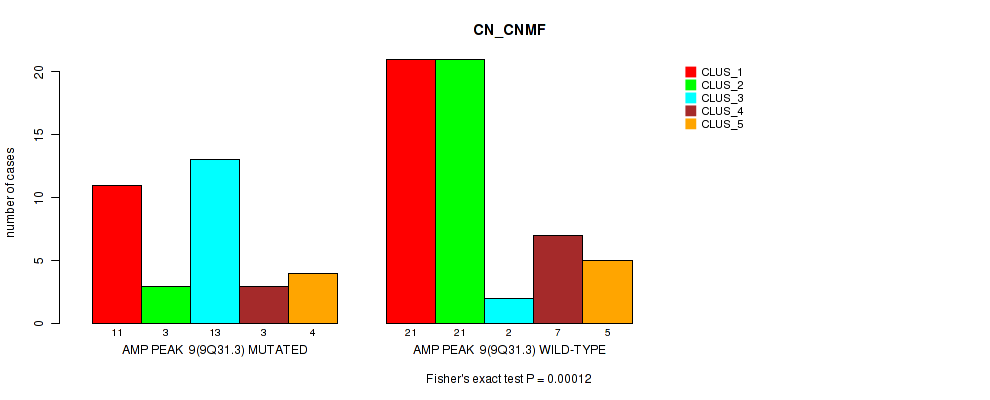
P value = 0.00012 (Fisher's exact test), Q value = 0.041
Table S9. Gene #9: 'amp_9q31.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 9(9Q31.3) MUTATED | 11 | 3 | 13 | 3 | 4 |
| AMP PEAK 9(9Q31.3) WILD-TYPE | 21 | 21 | 2 | 7 | 5 |
Figure S9. Get High-res Image Gene #9: 'amp_9q31.3' versus Molecular Subtype #1: 'CN_CNMF'
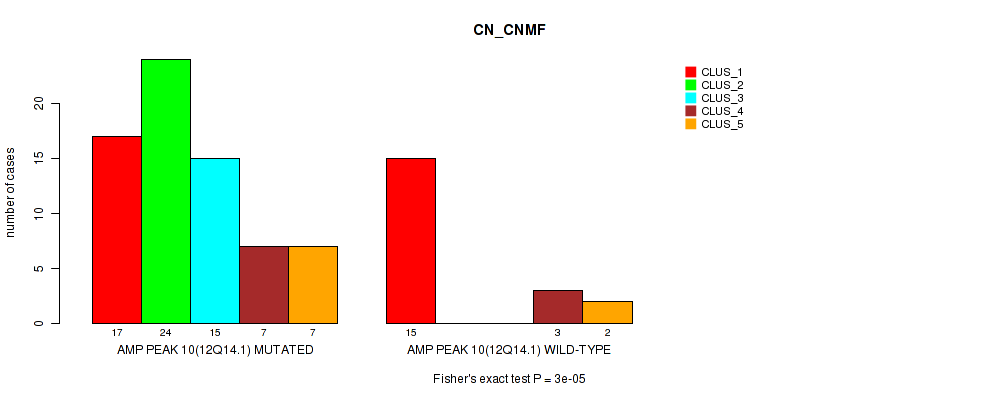
P value = 3e-05 (Fisher's exact test), Q value = 0.01
Table S10. Gene #10: 'amp_12q14.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 10(12Q14.1) MUTATED | 17 | 24 | 15 | 7 | 7 |
| AMP PEAK 10(12Q14.1) WILD-TYPE | 15 | 0 | 0 | 3 | 2 |
Figure S10. Get High-res Image Gene #10: 'amp_12q14.1' versus Molecular Subtype #1: 'CN_CNMF'
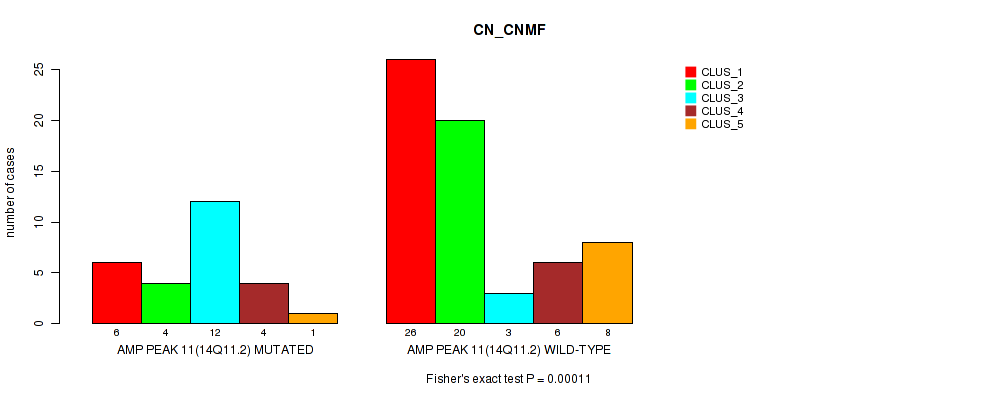
P value = 0.00011 (Fisher's exact test), Q value = 0.038
Table S11. Gene #11: 'amp_14q11.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 11(14Q11.2) MUTATED | 6 | 4 | 12 | 4 | 1 |
| AMP PEAK 11(14Q11.2) WILD-TYPE | 26 | 20 | 3 | 6 | 8 |
Figure S11. Get High-res Image Gene #11: 'amp_14q11.2' versus Molecular Subtype #1: 'CN_CNMF'
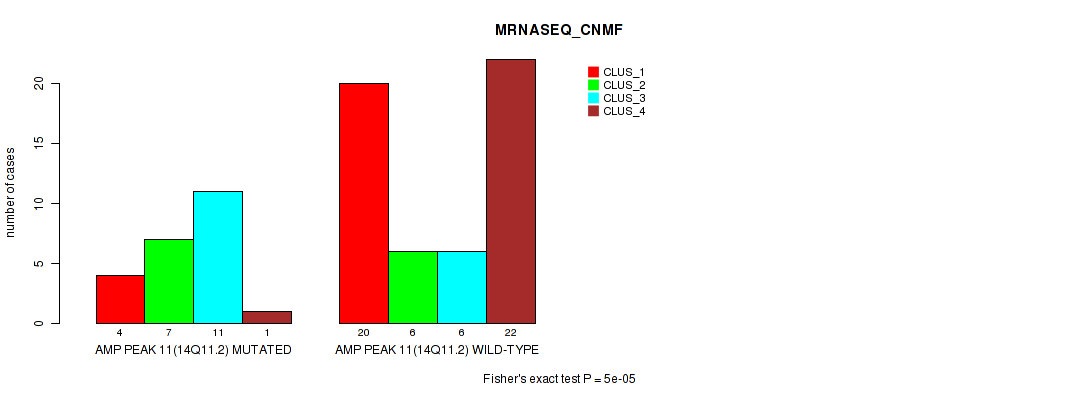
P value = 5e-05 (Fisher's exact test), Q value = 0.017
Table S12. Gene #11: 'amp_14q11.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| AMP PEAK 11(14Q11.2) MUTATED | 4 | 7 | 11 | 1 |
| AMP PEAK 11(14Q11.2) WILD-TYPE | 20 | 6 | 6 | 22 |
Figure S12. Get High-res Image Gene #11: 'amp_14q11.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
P value = 2e-05 (Fisher's exact test), Q value = 0.007
Table S13. Gene #11: 'amp_14q11.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 23 | 7 | 24 |
| AMP PEAK 11(14Q11.2) MUTATED | 5 | 16 | 0 | 2 |
| AMP PEAK 11(14Q11.2) WILD-TYPE | 18 | 7 | 7 | 22 |
Figure S13. Get High-res Image Gene #11: 'amp_14q11.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
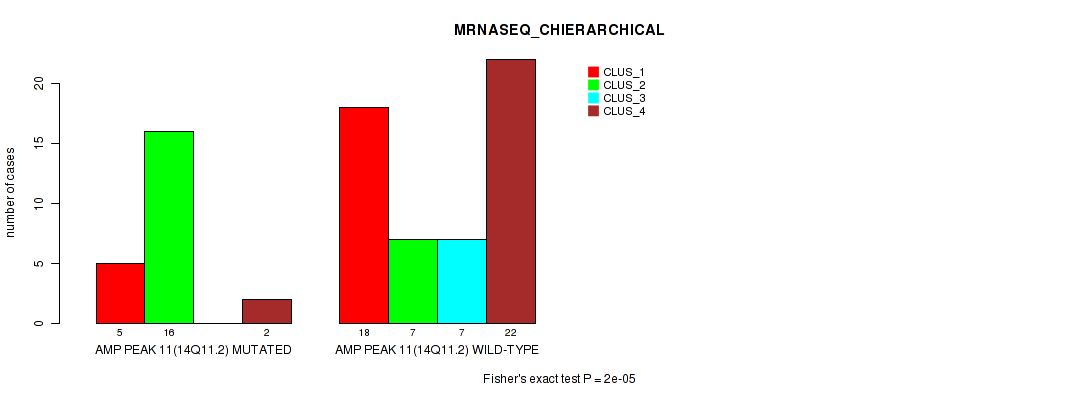
P value = 0.00043 (Fisher's exact test), Q value = 0.14
Table S14. Gene #11: 'amp_14q11.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 36 | 16 | 26 |
| AMP PEAK 11(14Q11.2) MUTATED | 19 | 3 | 2 |
| AMP PEAK 11(14Q11.2) WILD-TYPE | 17 | 13 | 24 |
Figure S14. Get High-res Image Gene #11: 'amp_14q11.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
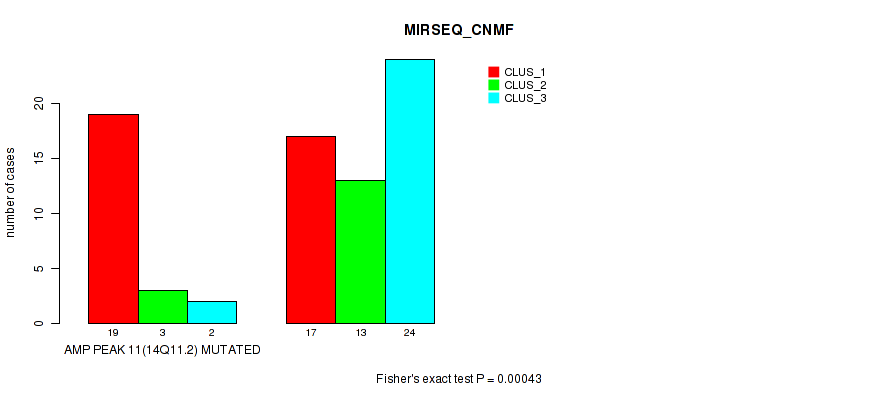
P value = 0.00021 (Fisher's exact test), Q value = 0.071
Table S15. Gene #11: 'amp_14q11.2' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 37 | 18 | 23 |
| AMP PEAK 11(14Q11.2) MUTATED | 19 | 4 | 1 |
| AMP PEAK 11(14Q11.2) WILD-TYPE | 18 | 14 | 22 |
Figure S15. Get High-res Image Gene #11: 'amp_14q11.2' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
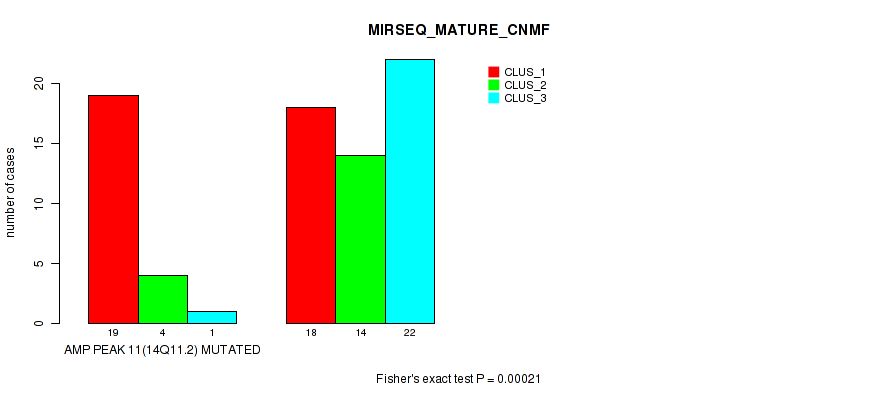
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S16. Gene #12: 'amp_16p13.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 12(16P13.3) MUTATED | 6 | 24 | 10 | 5 | 5 |
| AMP PEAK 12(16P13.3) WILD-TYPE | 26 | 0 | 5 | 5 | 4 |
Figure S16. Get High-res Image Gene #12: 'amp_16p13.3' versus Molecular Subtype #1: 'CN_CNMF'
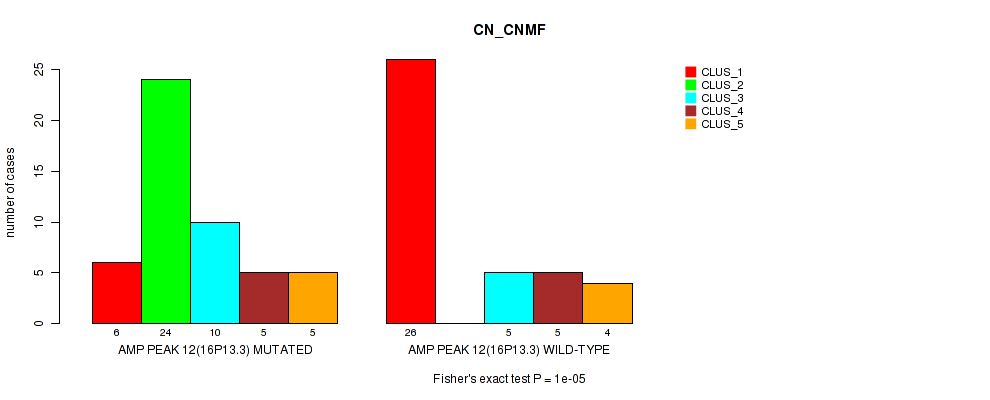
P value = 0.00016 (Fisher's exact test), Q value = 0.055
Table S17. Gene #12: 'amp_16p13.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| AMP PEAK 12(16P13.3) MUTATED | 5 | 9 | 14 | 16 |
| AMP PEAK 12(16P13.3) WILD-TYPE | 19 | 4 | 3 | 7 |
Figure S17. Get High-res Image Gene #12: 'amp_16p13.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
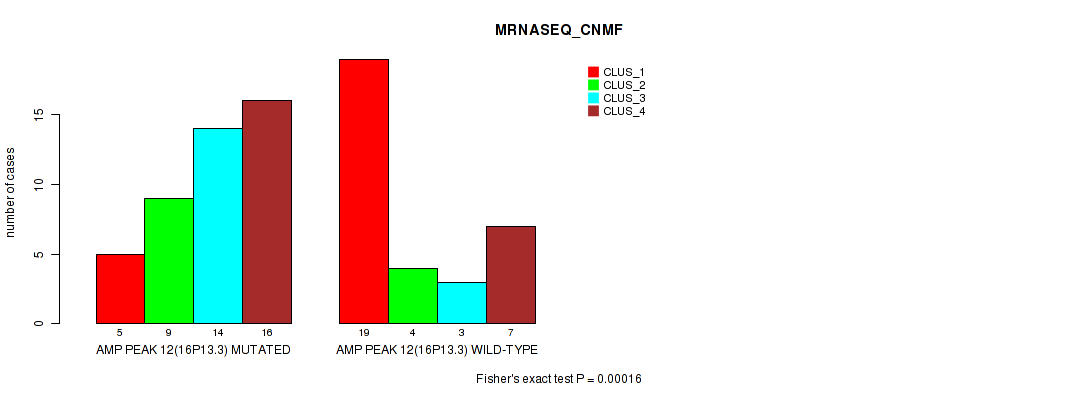
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S18. Gene #12: 'amp_16p13.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 23 | 7 | 24 |
| AMP PEAK 12(16P13.3) MUTATED | 6 | 19 | 0 | 19 |
| AMP PEAK 12(16P13.3) WILD-TYPE | 17 | 4 | 7 | 5 |
Figure S18. Get High-res Image Gene #12: 'amp_16p13.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
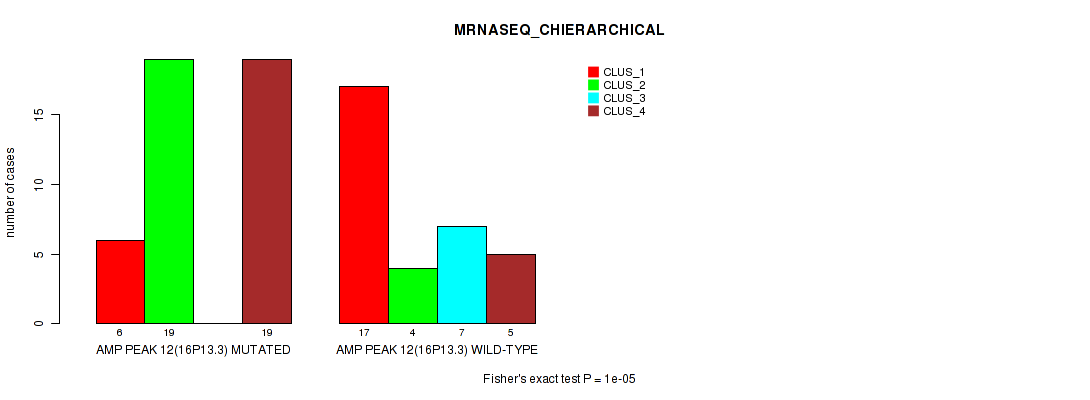
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S19. Gene #13: 'amp_16q22.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 13(16Q22.1) MUTATED | 11 | 24 | 13 | 3 | 5 |
| AMP PEAK 13(16Q22.1) WILD-TYPE | 21 | 0 | 2 | 7 | 4 |
Figure S19. Get High-res Image Gene #13: 'amp_16q22.1' versus Molecular Subtype #1: 'CN_CNMF'
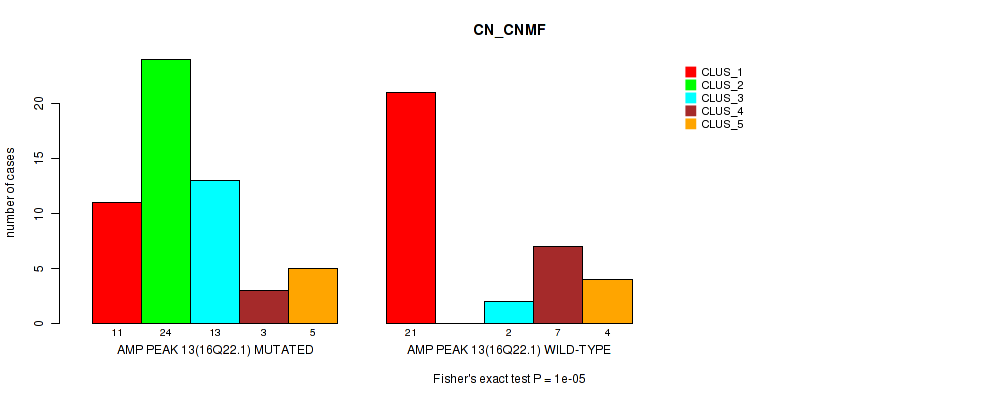
P value = 0.00024 (Fisher's exact test), Q value = 0.081
Table S20. Gene #13: 'amp_16q22.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 23 | 7 | 24 |
| AMP PEAK 13(16Q22.1) MUTATED | 9 | 20 | 1 | 18 |
| AMP PEAK 13(16Q22.1) WILD-TYPE | 14 | 3 | 6 | 6 |
Figure S20. Get High-res Image Gene #13: 'amp_16q22.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
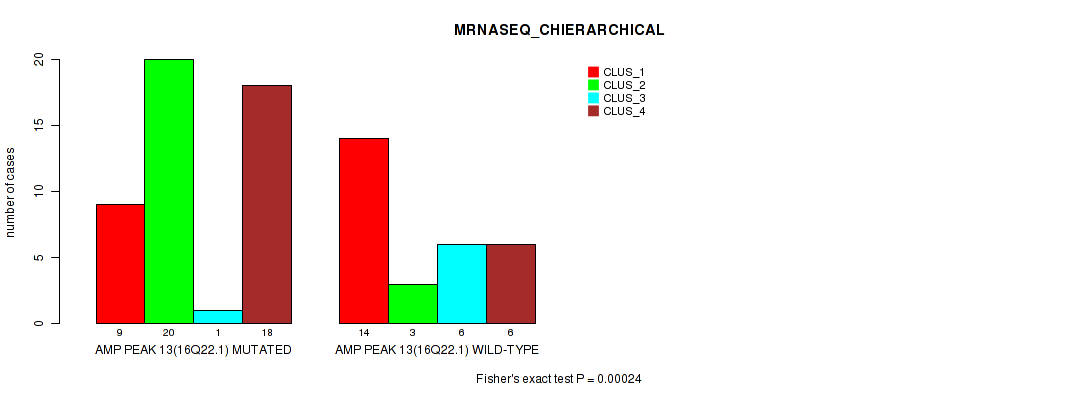
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S21. Gene #14: 'amp_16q24.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 14(16Q24.2) MUTATED | 6 | 24 | 10 | 4 | 5 |
| AMP PEAK 14(16Q24.2) WILD-TYPE | 26 | 0 | 5 | 6 | 4 |
Figure S21. Get High-res Image Gene #14: 'amp_16q24.2' versus Molecular Subtype #1: 'CN_CNMF'
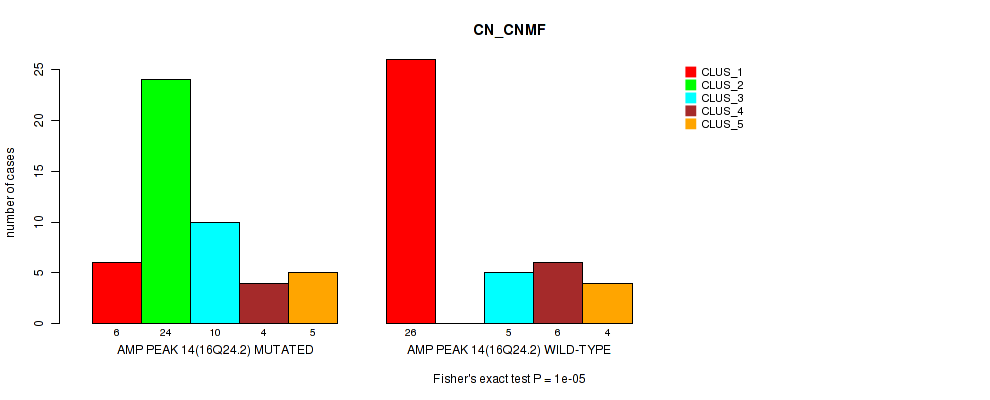
P value = 0.00026 (Fisher's exact test), Q value = 0.088
Table S22. Gene #14: 'amp_16q24.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| AMP PEAK 14(16Q24.2) MUTATED | 5 | 10 | 13 | 16 |
| AMP PEAK 14(16Q24.2) WILD-TYPE | 19 | 3 | 4 | 7 |
Figure S22. Get High-res Image Gene #14: 'amp_16q24.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
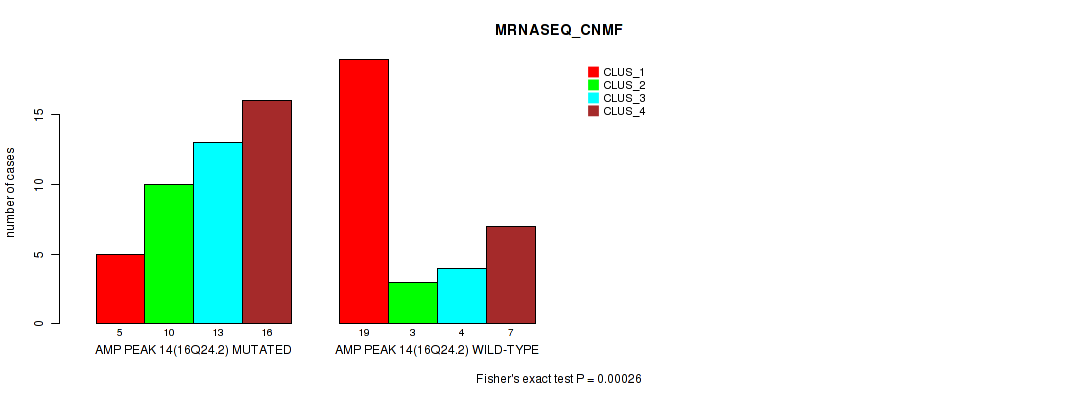
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S23. Gene #14: 'amp_16q24.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 23 | 7 | 24 |
| AMP PEAK 14(16Q24.2) MUTATED | 5 | 20 | 1 | 18 |
| AMP PEAK 14(16Q24.2) WILD-TYPE | 18 | 3 | 6 | 6 |
Figure S23. Get High-res Image Gene #14: 'amp_16q24.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
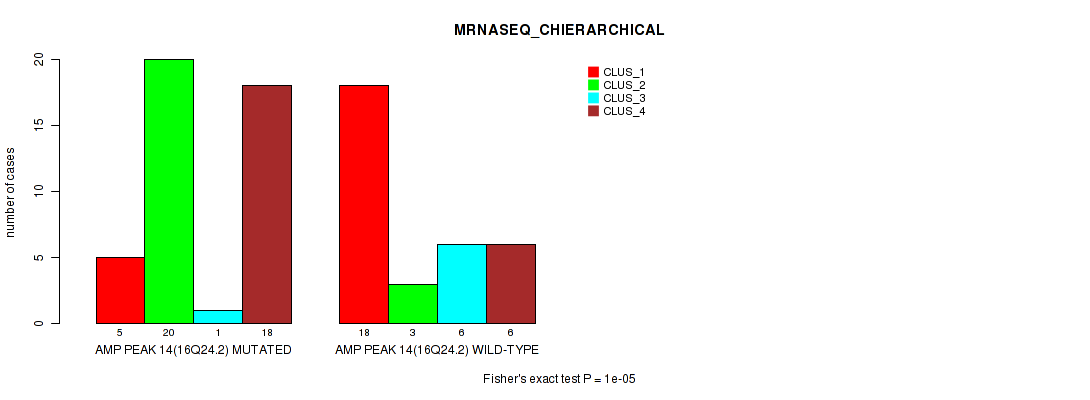
P value = 0.00045 (Fisher's exact test), Q value = 0.15
Table S24. Gene #16: 'amp_19p13.12' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 23 | 7 | 24 |
| AMP PEAK 16(19P13.12) MUTATED | 15 | 21 | 1 | 12 |
| AMP PEAK 16(19P13.12) WILD-TYPE | 8 | 2 | 6 | 12 |
Figure S24. Get High-res Image Gene #16: 'amp_19p13.12' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
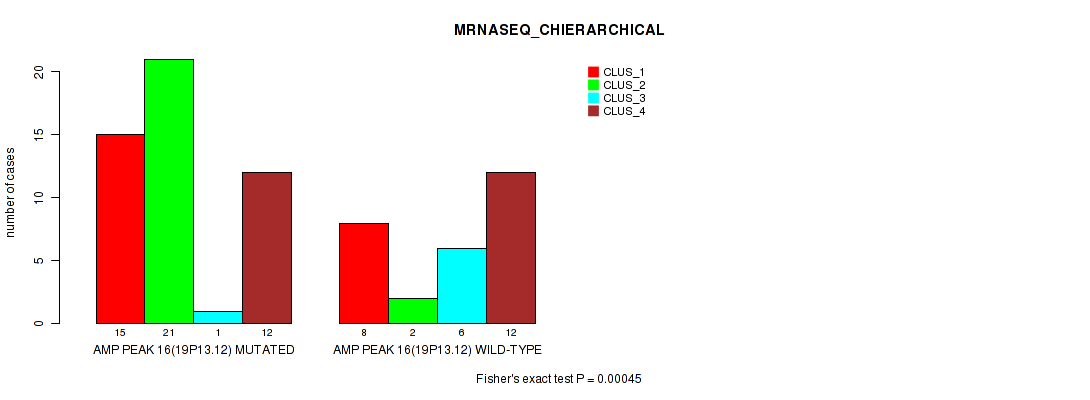
P value = 9e-05 (Fisher's exact test), Q value = 0.031
Table S25. Gene #18: 'amp_xp11.22' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 18(XP11.22) MUTATED | 10 | 21 | 11 | 4 | 3 |
| AMP PEAK 18(XP11.22) WILD-TYPE | 22 | 3 | 4 | 6 | 6 |
Figure S25. Get High-res Image Gene #18: 'amp_xp11.22' versus Molecular Subtype #1: 'CN_CNMF'
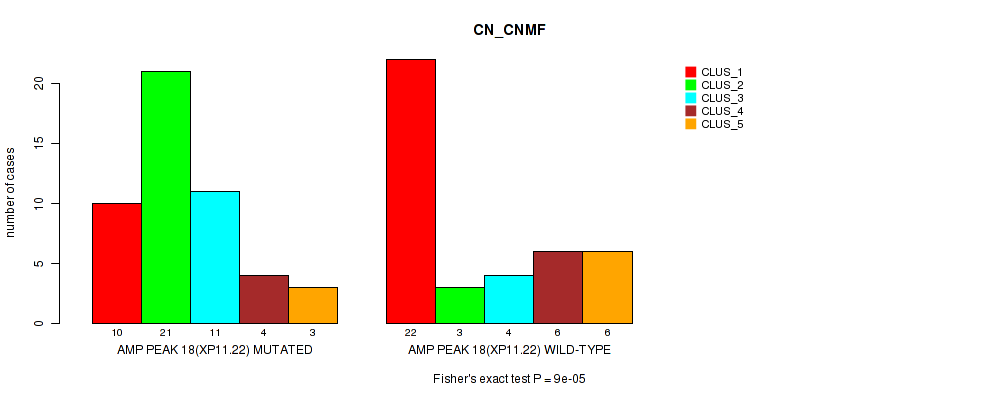
P value = 0.00018 (Fisher's exact test), Q value = 0.061
Table S26. Gene #19: 'amp_xq28' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| AMP PEAK 19(XQ28) MUTATED | 8 | 20 | 10 | 4 | 4 |
| AMP PEAK 19(XQ28) WILD-TYPE | 24 | 4 | 5 | 6 | 5 |
Figure S26. Get High-res Image Gene #19: 'amp_xq28' versus Molecular Subtype #1: 'CN_CNMF'
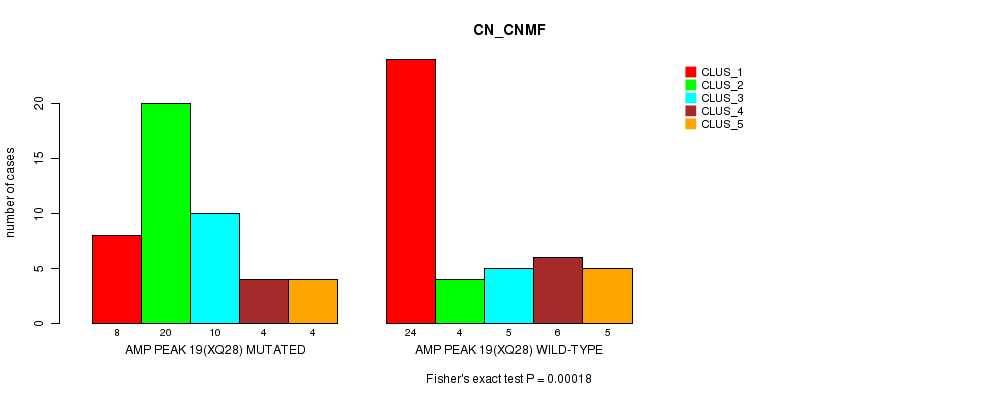
P value = 4e-05 (Fisher's exact test), Q value = 0.014
Table S27. Gene #30: 'del_9p23' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| DEL PEAK 11(9P23) MUTATED | 15 | 1 | 4 | 1 |
| DEL PEAK 11(9P23) WILD-TYPE | 9 | 12 | 13 | 22 |
Figure S27. Get High-res Image Gene #30: 'del_9p23' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
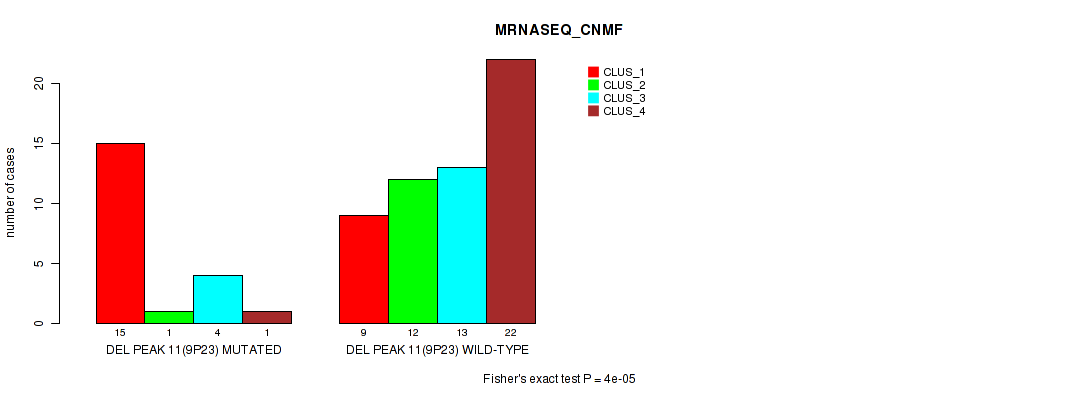
P value = 5e-04 (Fisher's exact test), Q value = 0.17
Table S28. Gene #31: 'del_9p21.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| DEL PEAK 12(9P21.3) MUTATED | 17 | 3 | 2 | 5 | 0 |
| DEL PEAK 12(9P21.3) WILD-TYPE | 15 | 21 | 13 | 5 | 9 |
Figure S28. Get High-res Image Gene #31: 'del_9p21.3' versus Molecular Subtype #1: 'CN_CNMF'
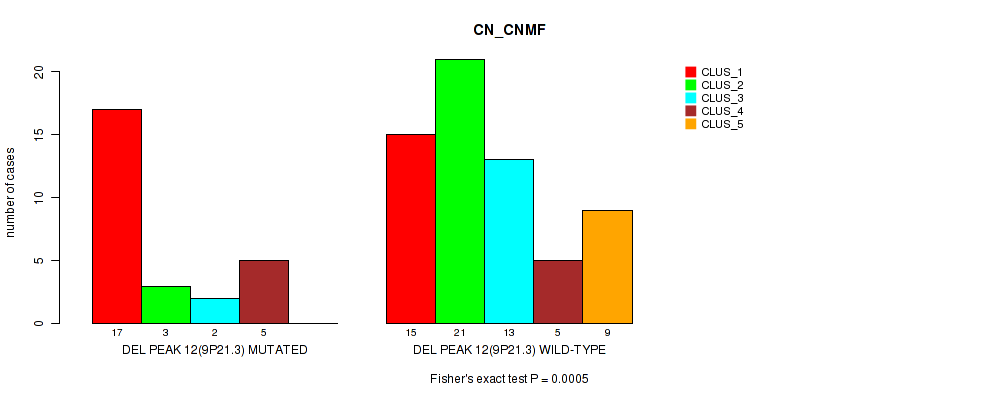
P value = 2e-05 (Fisher's exact test), Q value = 0.007
Table S29. Gene #31: 'del_9p21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 24 | 13 | 17 | 23 |
| DEL PEAK 12(9P21.3) MUTATED | 17 | 1 | 4 | 2 |
| DEL PEAK 12(9P21.3) WILD-TYPE | 7 | 12 | 13 | 21 |
Figure S29. Get High-res Image Gene #31: 'del_9p21.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
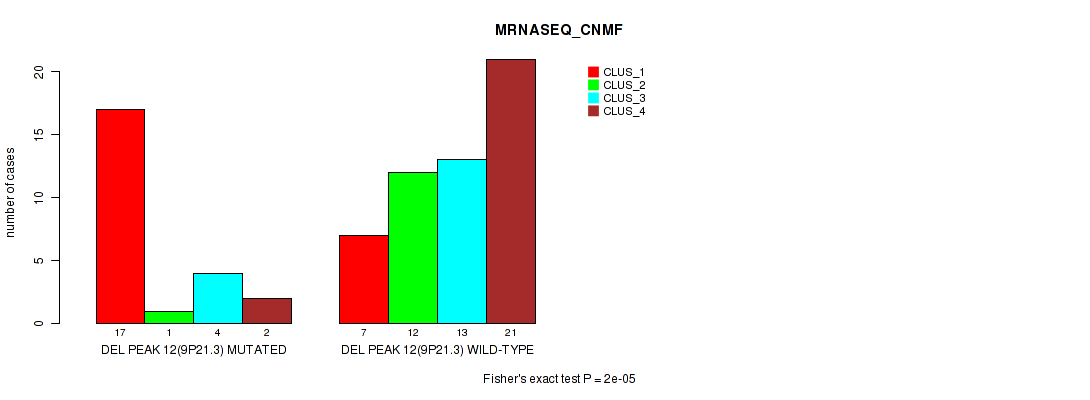
P value = 0.00053 (Fisher's exact test), Q value = 0.18
Table S30. Gene #31: 'del_9p21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 23 | 23 | 7 | 24 |
| DEL PEAK 12(9P21.3) MUTATED | 15 | 4 | 2 | 3 |
| DEL PEAK 12(9P21.3) WILD-TYPE | 8 | 19 | 5 | 21 |
Figure S30. Get High-res Image Gene #31: 'del_9p21.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
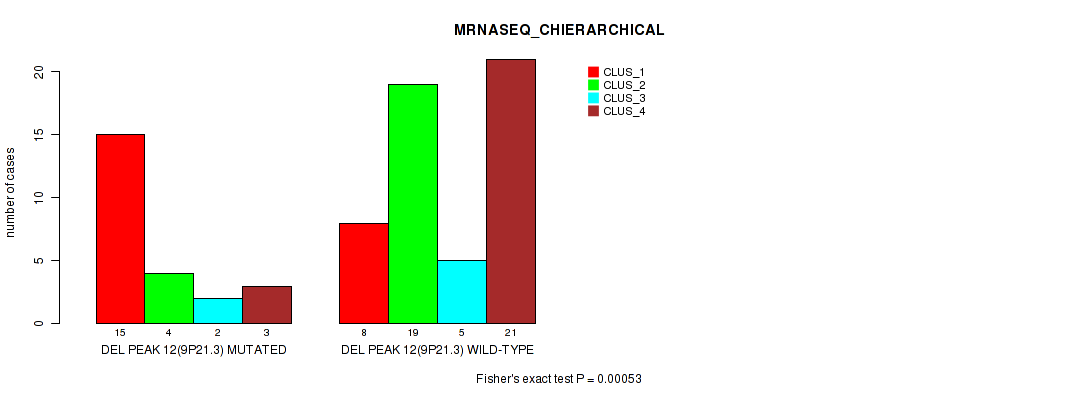
P value = 0.00035 (Fisher's exact test), Q value = 0.12
Table S31. Gene #34: 'del_11q14.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 32 | 24 | 15 | 10 | 9 |
| DEL PEAK 15(11Q14.1) MUTATED | 6 | 3 | 3 | 8 | 5 |
| DEL PEAK 15(11Q14.1) WILD-TYPE | 26 | 21 | 12 | 2 | 4 |
Figure S31. Get High-res Image Gene #34: 'del_11q14.1' versus Molecular Subtype #1: 'CN_CNMF'
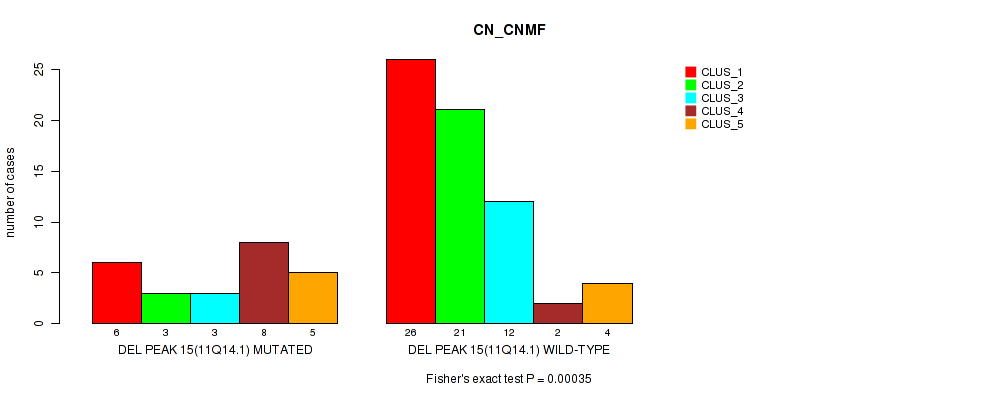
-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtype file = ACC-TP.transferedmergedcluster.txt
-
Number of patients = 90
-
Number of significantly focal cnvs = 45
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.