This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 61 focal events and 8 molecular subtypes across 169 patients, 94 significant findings detected with P value < 0.05 and Q value < 0.25.
-
amp_1p32.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
amp_1q24.3 cnv correlated to 'CN_CNMF'.
-
amp_5p15.2 cnv correlated to 'CN_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
amp_6p21.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
amp_6q24.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_12q15 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
amp_17p11.2 cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
amp_17q24.3 cnv correlated to 'METHLYATION_CNMF'.
-
amp_19p13.2 cnv correlated to 'CN_CNMF'.
-
amp_19q12 cnv correlated to 'CN_CNMF'.
-
amp_20q13.33 cnv correlated to 'CN_CNMF'.
-
amp_21q21.1 cnv correlated to 'CN_CNMF'.
-
amp_xp21.2 cnv correlated to 'CN_CNMF'.
-
del_1p36.32 cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_1q44 cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
del_2p25.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_2q37.3 cnv correlated to 'CN_CNMF'.
-
del_2q37.3 cnv correlated to 'CN_CNMF'.
-
del_3p21.31 cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_3q29 cnv correlated to 'METHLYATION_CNMF'.
-
del_4q35.1 cnv correlated to 'CN_CNMF'.
-
del_6q14.1 cnv correlated to 'CN_CNMF'.
-
del_7q36.3 cnv correlated to 'CN_CNMF'.
-
del_8p23.3 cnv correlated to 'METHLYATION_CNMF'.
-
del_9q34.3 cnv correlated to 'CN_CNMF'.
-
del_10p15.3 cnv correlated to 'METHLYATION_CNMF'.
-
del_10q23.31 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_11p15.5 cnv correlated to 'CN_CNMF'.
-
del_11q24.3 cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
del_12p13.1 cnv correlated to 'METHLYATION_CNMF'.
-
del_13q14.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_14q24.1 cnv correlated to 'MRNASEQ_CHIERARCHICAL'.
-
del_15q11.2 cnv correlated to 'MRNASEQ_CNMF'.
-
del_16q12.1 cnv correlated to 'CN_CNMF'.
-
del_16q23.1 cnv correlated to 'CN_CNMF'.
-
del_17p13.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_17q11.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MIRSEQ_MATURE_CNMF'.
-
del_17q25.3 cnv correlated to 'CN_CNMF'.
-
del_19p13.3 cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
del_19q13.43 cnv correlated to 'CN_CNMF'.
-
del_22q13.31 cnv correlated to 'CN_CNMF'.
-
del_xq27.1 cnv correlated to 'CN_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 61 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 94 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| amp 6q24 3 | 53 (31%) | 116 |
0.00011 (0.0464) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
6e-05 (0.0259) |
1e-05 (0.00488) |
1e-05 (0.00488) |
0.00021 (0.0865) |
| amp 12q15 | 64 (38%) | 105 |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
| del 10q23 31 | 95 (56%) | 74 |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
0.00026 (0.106) |
5e-05 (0.0217) |
2e-05 (0.00894) |
0.00051 (0.203) |
| del 1p36 32 | 51 (30%) | 118 |
0.00844 (1.00) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
1e-05 (0.00488) |
| del 17p13 1 | 82 (49%) | 87 |
3e-05 (0.0132) |
0.0001 (0.0424) |
0.00013 (0.0546) |
0.00057 (0.226) |
0.0161 (1.00) |
0.00268 (0.93) |
0.00338 (1.00) |
0.00021 (0.0865) |
| amp 1p32 1 | 62 (37%) | 107 |
1e-05 (0.00488) |
1e-05 (0.00488) |
6e-05 (0.0259) |
3e-05 (0.0132) |
0.00544 (1.00) |
0.00338 (1.00) |
0.0156 (1.00) |
0.00358 (1.00) |
| amp 6p21 1 | 40 (24%) | 129 |
3e-05 (0.0132) |
1e-05 (0.00488) |
0.00061 (0.242) |
0.00081 (0.312) |
0.00316 (1.00) |
0.00013 (0.0546) |
0.00091 (0.344) |
0.00099 (0.373) |
| del 2p25 3 | 56 (33%) | 113 |
0.00017 (0.0707) |
0.00018 (0.0745) |
3e-05 (0.0132) |
0.0003 (0.121) |
0.00409 (1.00) |
0.00086 (0.328) |
0.0057 (1.00) |
0.00088 (0.334) |
| del 13q14 2 | 121 (72%) | 48 |
1e-05 (0.00488) |
1e-05 (0.00488) |
0.0001 (0.0424) |
1e-05 (0.00488) |
0.00264 (0.921) |
0.0019 (0.684) |
0.0022 (0.779) |
0.00481 (1.00) |
| del 3p21 31 | 37 (22%) | 132 |
0.03 (1.00) |
1e-05 (0.00488) |
0.00013 (0.0546) |
0.00303 (1.00) |
0.00671 (1.00) |
0.00017 (0.0707) |
0.0231 (1.00) |
0.00623 (1.00) |
| del 17q11 2 | 51 (30%) | 118 |
2e-05 (0.00894) |
1e-05 (0.00488) |
0.00064 (0.252) |
0.00328 (1.00) |
0.00107 (0.401) |
0.00114 (0.426) |
0.00044 (0.176) |
0.00104 (0.391) |
| amp 5p15 2 | 73 (43%) | 96 |
9e-05 (0.0382) |
0.0066 (1.00) |
0.0148 (1.00) |
0.00026 (0.106) |
0.868 (1.00) |
0.0536 (1.00) |
0.713 (1.00) |
0.808 (1.00) |
| amp 17p11 2 | 69 (41%) | 100 |
1e-05 (0.00488) |
7e-05 (0.03) |
0.155 (1.00) |
0.0112 (1.00) |
0.516 (1.00) |
0.117 (1.00) |
0.305 (1.00) |
0.659 (1.00) |
| del 1q44 | 53 (31%) | 116 |
1e-05 (0.00488) |
7e-05 (0.03) |
0.03 (1.00) |
0.0342 (1.00) |
0.0685 (1.00) |
0.0231 (1.00) |
0.0771 (1.00) |
0.134 (1.00) |
| del 11q24 3 | 75 (44%) | 94 |
1e-05 (0.00488) |
2e-05 (0.00894) |
0.0226 (1.00) |
0.0139 (1.00) |
0.00761 (1.00) |
0.0081 (1.00) |
0.00196 (0.704) |
0.0568 (1.00) |
| del 19p13 3 | 50 (30%) | 119 |
2e-05 (0.00894) |
0.00012 (0.0505) |
0.0054 (1.00) |
0.019 (1.00) |
0.0107 (1.00) |
0.00186 (0.671) |
0.00207 (0.737) |
0.00082 (0.314) |
| amp 1q24 3 | 71 (42%) | 98 |
2e-05 (0.00894) |
0.0017 (0.619) |
0.108 (1.00) |
0.0197 (1.00) |
0.312 (1.00) |
0.199 (1.00) |
0.224 (1.00) |
0.273 (1.00) |
| amp 17q24 3 | 48 (28%) | 121 |
0.00907 (1.00) |
0.00014 (0.0584) |
0.00508 (1.00) |
0.00422 (1.00) |
0.0557 (1.00) |
0.029 (1.00) |
0.101 (1.00) |
0.0142 (1.00) |
| amp 19p13 2 | 68 (40%) | 101 |
2e-05 (0.00894) |
0.0459 (1.00) |
0.105 (1.00) |
0.047 (1.00) |
0.819 (1.00) |
0.48 (1.00) |
0.758 (1.00) |
0.785 (1.00) |
| amp 19q12 | 61 (36%) | 108 |
1e-05 (0.00488) |
0.00083 (0.317) |
0.0157 (1.00) |
0.0142 (1.00) |
0.588 (1.00) |
0.00153 (0.56) |
0.382 (1.00) |
0.4 (1.00) |
| amp 20q13 33 | 68 (40%) | 101 |
8e-05 (0.0341) |
0.00337 (1.00) |
0.00174 (0.632) |
0.0108 (1.00) |
0.0256 (1.00) |
0.00286 (0.99) |
0.0159 (1.00) |
0.0314 (1.00) |
| amp 21q21 1 | 51 (30%) | 118 |
0.0002 (0.0826) |
0.00115 (0.429) |
0.134 (1.00) |
0.14 (1.00) |
0.224 (1.00) |
0.0109 (1.00) |
0.104 (1.00) |
0.177 (1.00) |
| amp xp21 2 | 60 (36%) | 109 |
1e-05 (0.00488) |
0.00066 (0.259) |
0.0412 (1.00) |
0.137 (1.00) |
0.814 (1.00) |
0.0246 (1.00) |
0.501 (1.00) |
0.774 (1.00) |
| del 2q37 3 | 68 (40%) | 101 |
1e-05 (0.00488) |
0.0217 (1.00) |
0.107 (1.00) |
0.0397 (1.00) |
0.386 (1.00) |
0.4 (1.00) |
0.573 (1.00) |
0.578 (1.00) |
| del 2q37 3 | 70 (41%) | 99 |
1e-05 (0.00488) |
0.00731 (1.00) |
0.326 (1.00) |
0.0131 (1.00) |
0.805 (1.00) |
0.417 (1.00) |
0.956 (1.00) |
0.841 (1.00) |
| del 3q29 | 39 (23%) | 130 |
0.0442 (1.00) |
0.00034 (0.137) |
0.207 (1.00) |
0.55 (1.00) |
0.434 (1.00) |
0.0542 (1.00) |
0.241 (1.00) |
0.0755 (1.00) |
| del 4q35 1 | 63 (37%) | 106 |
2e-05 (0.00894) |
0.0255 (1.00) |
0.564 (1.00) |
0.615 (1.00) |
0.474 (1.00) |
0.459 (1.00) |
0.205 (1.00) |
0.977 (1.00) |
| del 6q14 1 | 49 (29%) | 120 |
0.00028 (0.114) |
0.0112 (1.00) |
0.308 (1.00) |
0.452 (1.00) |
0.946 (1.00) |
0.705 (1.00) |
0.848 (1.00) |
0.123 (1.00) |
| del 7q36 3 | 44 (26%) | 125 |
6e-05 (0.0259) |
0.077 (1.00) |
0.132 (1.00) |
0.448 (1.00) |
0.284 (1.00) |
0.308 (1.00) |
0.331 (1.00) |
0.461 (1.00) |
| del 8p23 3 | 52 (31%) | 117 |
0.0482 (1.00) |
1e-05 (0.00488) |
0.00723 (1.00) |
0.00948 (1.00) |
0.0007 (0.274) |
0.0196 (1.00) |
0.00432 (1.00) |
0.0121 (1.00) |
| del 9q34 3 | 46 (27%) | 123 |
0.00043 (0.172) |
0.13 (1.00) |
0.146 (1.00) |
0.199 (1.00) |
0.153 (1.00) |
0.0515 (1.00) |
0.0808 (1.00) |
0.0351 (1.00) |
| del 10p15 3 | 88 (52%) | 81 |
0.00311 (1.00) |
0.00036 (0.145) |
0.59 (1.00) |
0.0284 (1.00) |
0.792 (1.00) |
0.644 (1.00) |
0.818 (1.00) |
0.74 (1.00) |
| del 11p15 5 | 73 (43%) | 96 |
5e-05 (0.0217) |
0.0008 (0.309) |
0.927 (1.00) |
0.496 (1.00) |
0.522 (1.00) |
0.925 (1.00) |
0.553 (1.00) |
0.879 (1.00) |
| del 12p13 1 | 57 (34%) | 112 |
0.00131 (0.485) |
0.00045 (0.18) |
0.0203 (1.00) |
0.178 (1.00) |
0.0749 (1.00) |
0.0821 (1.00) |
0.14 (1.00) |
0.108 (1.00) |
| del 14q24 1 | 56 (33%) | 113 |
0.526 (1.00) |
0.0275 (1.00) |
0.75 (1.00) |
0.00022 (0.0902) |
0.165 (1.00) |
0.0315 (1.00) |
0.0501 (1.00) |
0.0799 (1.00) |
| del 15q11 2 | 32 (19%) | 137 |
0.19 (1.00) |
0.0384 (1.00) |
0.00022 (0.0902) |
0.00517 (1.00) |
0.0122 (1.00) |
0.0272 (1.00) |
0.0201 (1.00) |
0.0415 (1.00) |
| del 16q12 1 | 95 (56%) | 74 |
1e-05 (0.00488) |
0.00076 (0.295) |
0.0243 (1.00) |
0.02 (1.00) |
0.123 (1.00) |
0.0692 (1.00) |
0.0234 (1.00) |
0.101 (1.00) |
| del 16q23 1 | 94 (56%) | 75 |
4e-05 (0.0174) |
0.00069 (0.27) |
0.00834 (1.00) |
0.00681 (1.00) |
0.0222 (1.00) |
0.0329 (1.00) |
0.00304 (1.00) |
0.0103 (1.00) |
| del 17q25 3 | 45 (27%) | 124 |
0.00062 (0.245) |
0.00325 (1.00) |
0.0552 (1.00) |
0.0627 (1.00) |
0.514 (1.00) |
0.166 (1.00) |
0.473 (1.00) |
0.385 (1.00) |
| del 19q13 43 | 73 (43%) | 96 |
0.00042 (0.169) |
0.0126 (1.00) |
0.102 (1.00) |
0.185 (1.00) |
0.0714 (1.00) |
0.0272 (1.00) |
0.0669 (1.00) |
0.161 (1.00) |
| del 22q13 31 | 54 (32%) | 115 |
7e-05 (0.03) |
0.027 (1.00) |
0.891 (1.00) |
0.111 (1.00) |
0.441 (1.00) |
0.428 (1.00) |
0.675 (1.00) |
0.206 (1.00) |
| del xq27 1 | 82 (49%) | 87 |
3e-05 (0.0132) |
0.00175 (0.633) |
0.00072 (0.281) |
0.00226 (0.798) |
0.00262 (0.917) |
0.00266 (0.926) |
0.00212 (0.753) |
0.0013 (0.482) |
| amp 3p11 2 | 40 (24%) | 129 |
0.00967 (1.00) |
0.0356 (1.00) |
0.00642 (1.00) |
0.00198 (0.709) |
0.506 (1.00) |
0.0437 (1.00) |
0.274 (1.00) |
0.285 (1.00) |
| amp 7q31 1 | 49 (29%) | 120 |
0.00075 (0.292) |
0.0153 (1.00) |
0.0547 (1.00) |
0.206 (1.00) |
0.0384 (1.00) |
0.00876 (1.00) |
0.0196 (1.00) |
0.00246 (0.866) |
| amp 8q21 12 | 61 (36%) | 108 |
0.054 (1.00) |
0.0364 (1.00) |
0.236 (1.00) |
0.609 (1.00) |
1 (1.00) |
0.0453 (1.00) |
0.587 (1.00) |
0.956 (1.00) |
| amp 11q22 2 | 26 (15%) | 143 |
0.0549 (1.00) |
0.0579 (1.00) |
0.00579 (1.00) |
0.14 (1.00) |
0.136 (1.00) |
0.0178 (1.00) |
0.0857 (1.00) |
0.115 (1.00) |
| amp 12p13 32 | 32 (19%) | 137 |
0.0665 (1.00) |
0.4 (1.00) |
0.149 (1.00) |
0.314 (1.00) |
0.098 (1.00) |
0.173 (1.00) |
0.187 (1.00) |
0.244 (1.00) |
| amp 12p12 1 | 42 (25%) | 127 |
0.00137 (0.506) |
0.0245 (1.00) |
0.00947 (1.00) |
0.0779 (1.00) |
0.161 (1.00) |
0.00727 (1.00) |
0.144 (1.00) |
0.0912 (1.00) |
| amp 13q34 | 31 (18%) | 138 |
0.00149 (0.548) |
0.00081 (0.312) |
0.077 (1.00) |
0.103 (1.00) |
0.699 (1.00) |
0.17 (1.00) |
0.964 (1.00) |
0.607 (1.00) |
| amp xq21 2 | 25 (15%) | 144 |
0.00745 (1.00) |
0.0796 (1.00) |
0.6 (1.00) |
0.237 (1.00) |
0.425 (1.00) |
0.255 (1.00) |
0.559 (1.00) |
0.508 (1.00) |
| del 1p32 3 | 26 (15%) | 143 |
0.407 (1.00) |
0.0959 (1.00) |
0.989 (1.00) |
0.118 (1.00) |
0.617 (1.00) |
0.286 (1.00) |
0.555 (1.00) |
0.62 (1.00) |
| del 6p25 1 | 61 (36%) | 108 |
0.408 (1.00) |
0.0422 (1.00) |
0.0025 (0.877) |
0.0128 (1.00) |
0.0544 (1.00) |
0.00297 (1.00) |
0.0348 (1.00) |
0.0202 (1.00) |
| del 6q16 1 | 48 (28%) | 121 |
0.00442 (1.00) |
0.211 (1.00) |
0.241 (1.00) |
0.792 (1.00) |
0.865 (1.00) |
0.524 (1.00) |
0.476 (1.00) |
0.0583 (1.00) |
| del 9p23 | 58 (34%) | 111 |
0.00675 (1.00) |
0.315 (1.00) |
0.027 (1.00) |
0.272 (1.00) |
0.108 (1.00) |
0.0297 (1.00) |
0.0961 (1.00) |
0.00583 (1.00) |
| del 9p21 3 | 72 (43%) | 97 |
0.00129 (0.48) |
0.0382 (1.00) |
0.00151 (0.554) |
0.0212 (1.00) |
0.0293 (1.00) |
0.00076 (0.295) |
0.00206 (0.735) |
0.00426 (1.00) |
| del 12q12 | 34 (20%) | 135 |
0.0431 (1.00) |
0.0327 (1.00) |
0.83 (1.00) |
0.202 (1.00) |
0.171 (1.00) |
0.664 (1.00) |
0.597 (1.00) |
0.506 (1.00) |
| del 18q23 | 57 (34%) | 112 |
0.00952 (1.00) |
0.0329 (1.00) |
0.0178 (1.00) |
0.0798 (1.00) |
0.0352 (1.00) |
0.0835 (1.00) |
0.0291 (1.00) |
0.269 (1.00) |
| del 21q11 2 | 31 (18%) | 138 |
0.362 (1.00) |
0.408 (1.00) |
0.268 (1.00) |
0.328 (1.00) |
0.0151 (1.00) |
0.327 (1.00) |
0.102 (1.00) |
0.139 (1.00) |
| del 21q22 3 | 44 (26%) | 125 |
0.0698 (1.00) |
0.0371 (1.00) |
0.482 (1.00) |
0.575 (1.00) |
0.217 (1.00) |
0.553 (1.00) |
0.386 (1.00) |
0.722 (1.00) |
| del xp22 33 | 54 (32%) | 115 |
0.0181 (1.00) |
0.00158 (0.577) |
0.00971 (1.00) |
0.185 (1.00) |
0.119 (1.00) |
0.148 (1.00) |
0.175 (1.00) |
0.138 (1.00) |
| del xq21 1 | 80 (47%) | 89 |
0.035 (1.00) |
0.00087 (0.331) |
0.00391 (1.00) |
0.00499 (1.00) |
0.0137 (1.00) |
0.00637 (1.00) |
0.00452 (1.00) |
0.0166 (1.00) |
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S1. Gene #1: 'amp_1p32.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 1(1P32.1) MUTATED | 17 | 37 | 8 |
| AMP PEAK 1(1P32.1) WILD-TYPE | 29 | 29 | 49 |
Figure S1. Get High-res Image Gene #1: 'amp_1p32.1' versus Molecular Subtype #1: 'CN_CNMF'
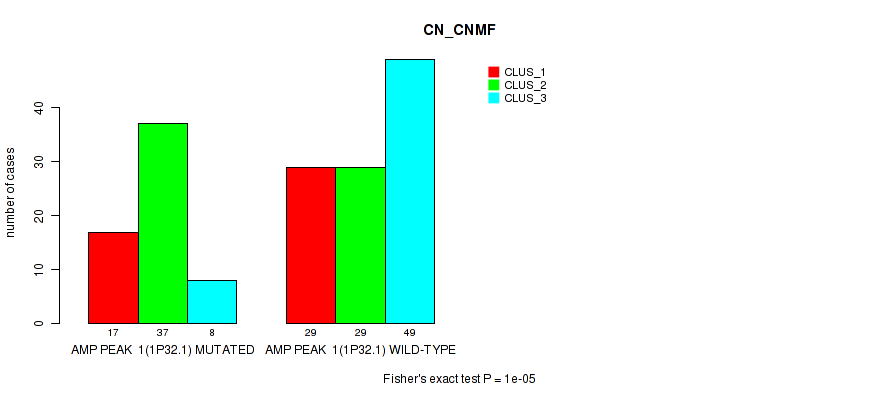
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S2. Gene #1: 'amp_1p32.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| AMP PEAK 1(1P32.1) MUTATED | 10 | 10 | 10 | 3 | 2 | 24 | 3 |
| AMP PEAK 1(1P32.1) WILD-TYPE | 20 | 18 | 32 | 18 | 14 | 4 | 1 |
Figure S2. Get High-res Image Gene #1: 'amp_1p32.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
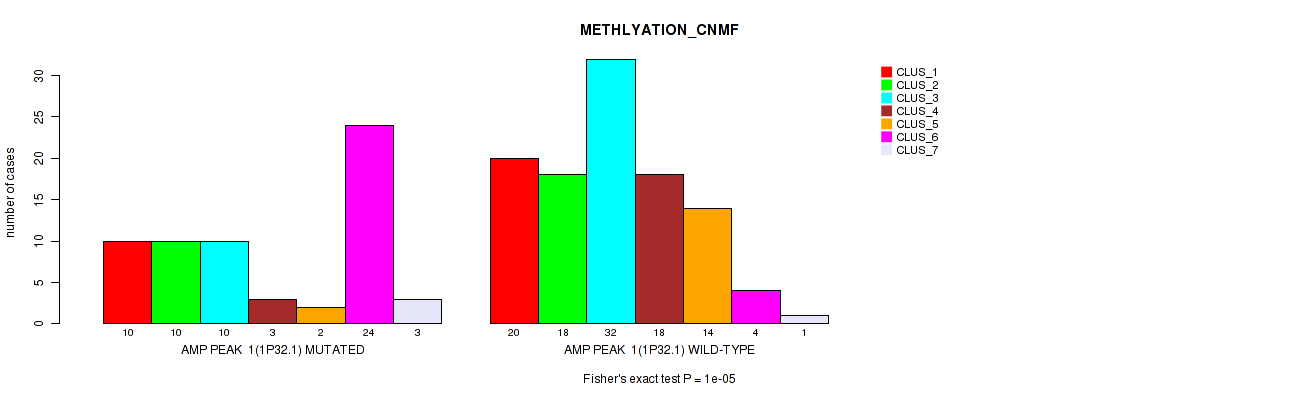
P value = 6e-05 (Fisher's exact test), Q value = 0.026
Table S3. Gene #1: 'amp_1p32.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| AMP PEAK 1(1P32.1) MUTATED | 27 | 15 | 6 | 13 |
| AMP PEAK 1(1P32.1) WILD-TYPE | 12 | 30 | 18 | 45 |
Figure S3. Get High-res Image Gene #1: 'amp_1p32.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
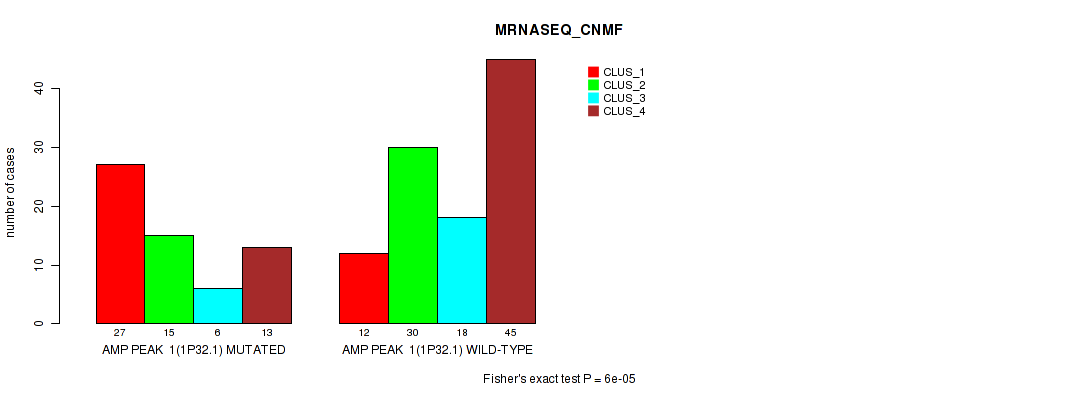
P value = 3e-05 (Fisher's exact test), Q value = 0.013
Table S4. Gene #1: 'amp_1p32.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 45 | 32 | 33 | 37 | 19 |
| AMP PEAK 1(1P32.1) MUTATED | 17 | 6 | 25 | 9 | 4 |
| AMP PEAK 1(1P32.1) WILD-TYPE | 28 | 26 | 8 | 28 | 15 |
Figure S4. Get High-res Image Gene #1: 'amp_1p32.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
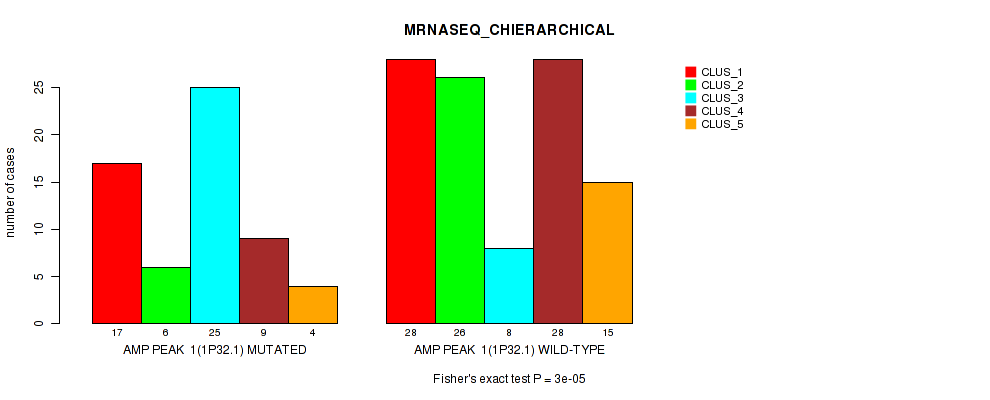
P value = 2e-05 (Fisher's exact test), Q value = 0.0089
Table S5. Gene #2: 'amp_1q24.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 2(1Q24.3) MUTATED | 22 | 39 | 10 |
| AMP PEAK 2(1Q24.3) WILD-TYPE | 24 | 27 | 47 |
Figure S5. Get High-res Image Gene #2: 'amp_1q24.3' versus Molecular Subtype #1: 'CN_CNMF'
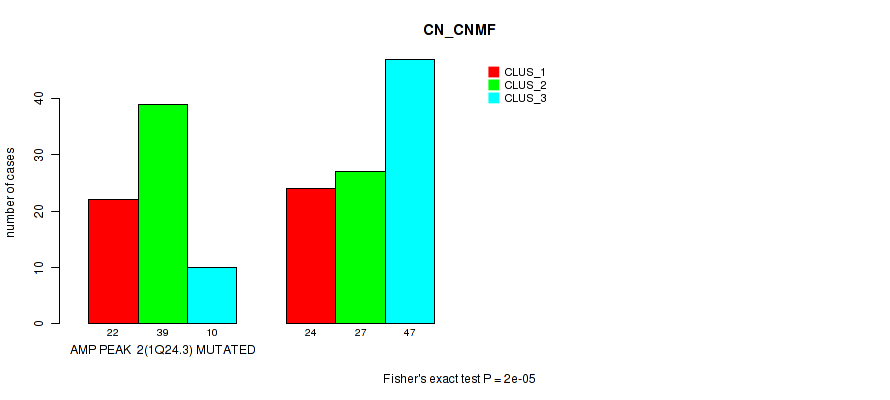
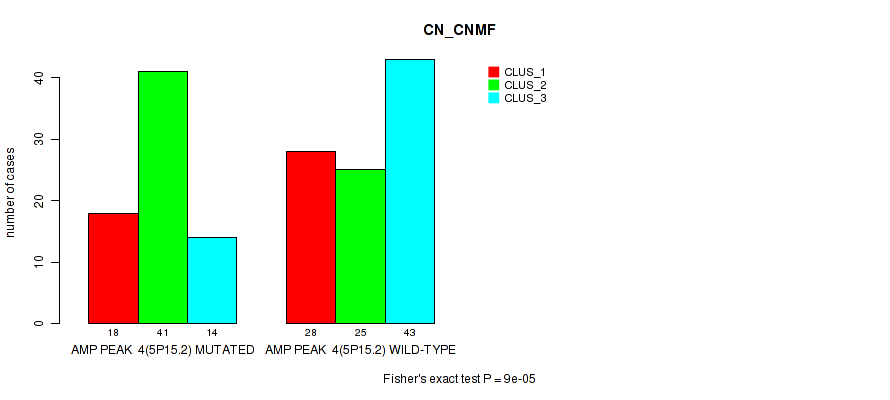
P value = 9e-05 (Fisher's exact test), Q value = 0.038
Table S6. Gene #4: 'amp_5p15.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 4(5P15.2) MUTATED | 18 | 41 | 14 |
| AMP PEAK 4(5P15.2) WILD-TYPE | 28 | 25 | 43 |
Figure S6. Get High-res Image Gene #4: 'amp_5p15.2' versus Molecular Subtype #1: 'CN_CNMF'
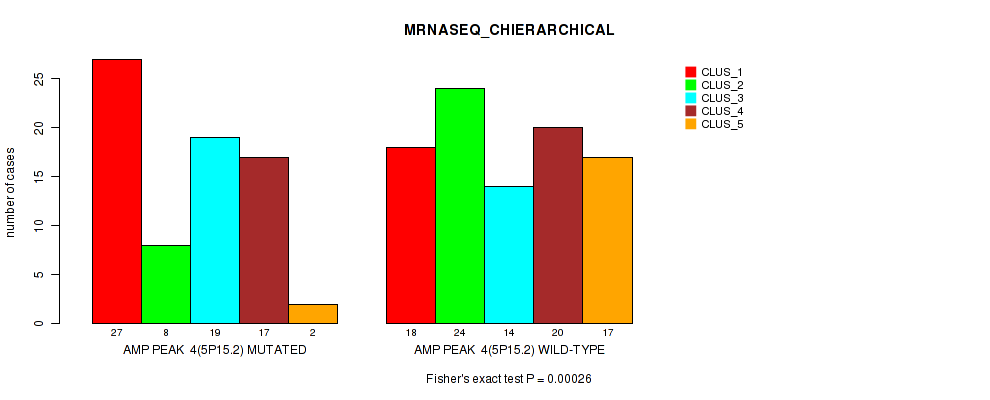
P value = 0.00026 (Fisher's exact test), Q value = 0.11
Table S7. Gene #4: 'amp_5p15.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 45 | 32 | 33 | 37 | 19 |
| AMP PEAK 4(5P15.2) MUTATED | 27 | 8 | 19 | 17 | 2 |
| AMP PEAK 4(5P15.2) WILD-TYPE | 18 | 24 | 14 | 20 | 17 |
Figure S7. Get High-res Image Gene #4: 'amp_5p15.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
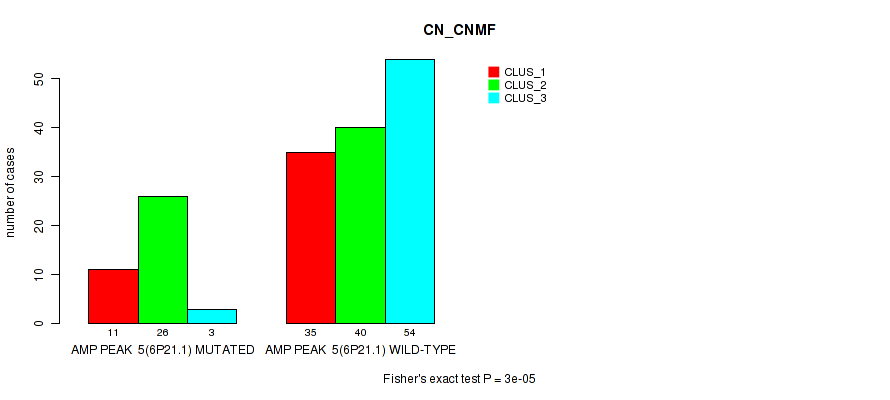
P value = 3e-05 (Fisher's exact test), Q value = 0.013
Table S8. Gene #5: 'amp_6p21.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 5(6P21.1) MUTATED | 11 | 26 | 3 |
| AMP PEAK 5(6P21.1) WILD-TYPE | 35 | 40 | 54 |
Figure S8. Get High-res Image Gene #5: 'amp_6p21.1' versus Molecular Subtype #1: 'CN_CNMF'
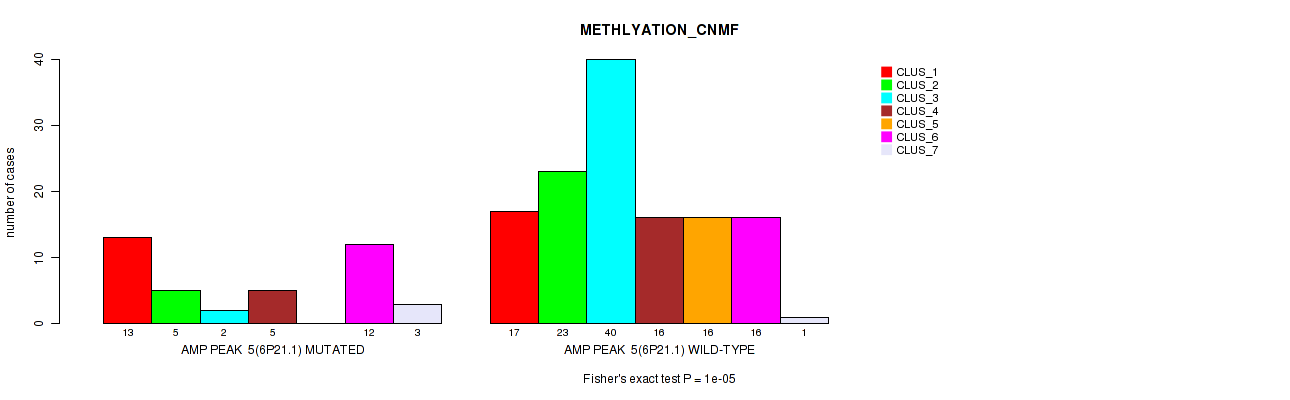
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S9. Gene #5: 'amp_6p21.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| AMP PEAK 5(6P21.1) MUTATED | 13 | 5 | 2 | 5 | 0 | 12 | 3 |
| AMP PEAK 5(6P21.1) WILD-TYPE | 17 | 23 | 40 | 16 | 16 | 16 | 1 |
Figure S9. Get High-res Image Gene #5: 'amp_6p21.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
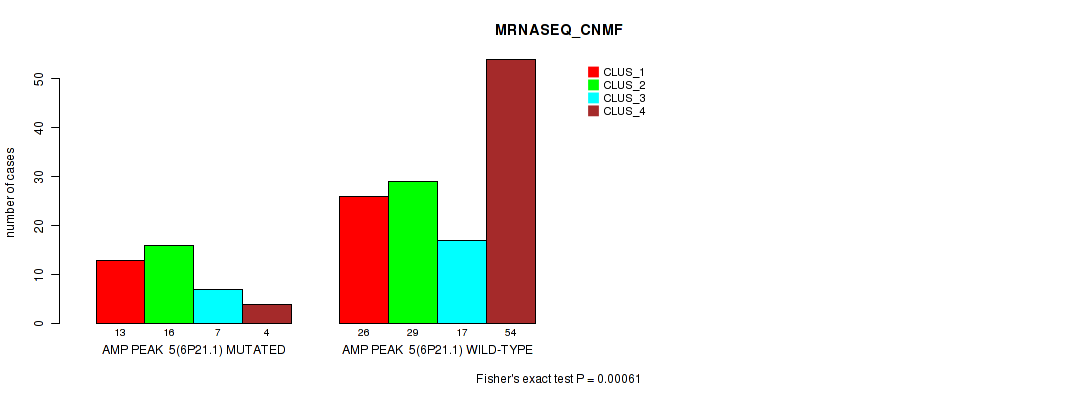
P value = 0.00061 (Fisher's exact test), Q value = 0.24
Table S10. Gene #5: 'amp_6p21.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| AMP PEAK 5(6P21.1) MUTATED | 13 | 16 | 7 | 4 |
| AMP PEAK 5(6P21.1) WILD-TYPE | 26 | 29 | 17 | 54 |
Figure S10. Get High-res Image Gene #5: 'amp_6p21.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
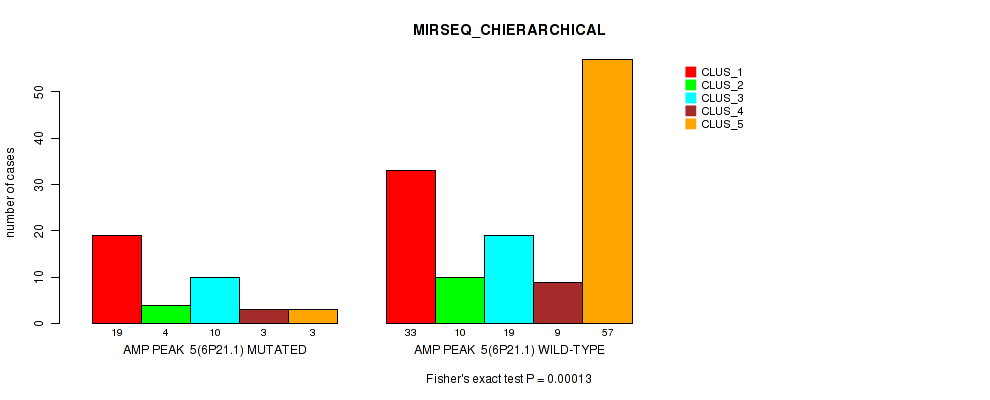
P value = 0.00013 (Fisher's exact test), Q value = 0.055
Table S11. Gene #5: 'amp_6p21.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 52 | 14 | 29 | 12 | 60 |
| AMP PEAK 5(6P21.1) MUTATED | 19 | 4 | 10 | 3 | 3 |
| AMP PEAK 5(6P21.1) WILD-TYPE | 33 | 10 | 19 | 9 | 57 |
Figure S11. Get High-res Image Gene #5: 'amp_6p21.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
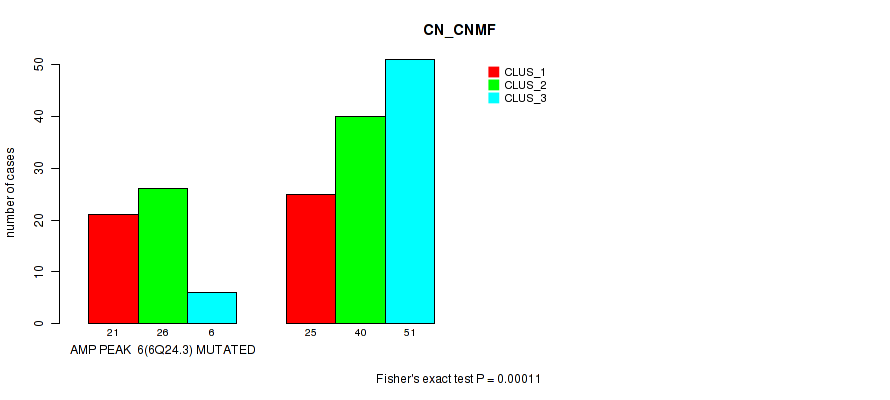
P value = 0.00011 (Fisher's exact test), Q value = 0.046
Table S12. Gene #6: 'amp_6q24.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 6(6Q24.3) MUTATED | 21 | 26 | 6 |
| AMP PEAK 6(6Q24.3) WILD-TYPE | 25 | 40 | 51 |
Figure S12. Get High-res Image Gene #6: 'amp_6q24.3' versus Molecular Subtype #1: 'CN_CNMF'
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S13. Gene #6: 'amp_6q24.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| AMP PEAK 6(6Q24.3) MUTATED | 21 | 12 | 3 | 3 | 1 | 12 | 1 |
| AMP PEAK 6(6Q24.3) WILD-TYPE | 9 | 16 | 39 | 18 | 15 | 16 | 3 |
Figure S13. Get High-res Image Gene #6: 'amp_6q24.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
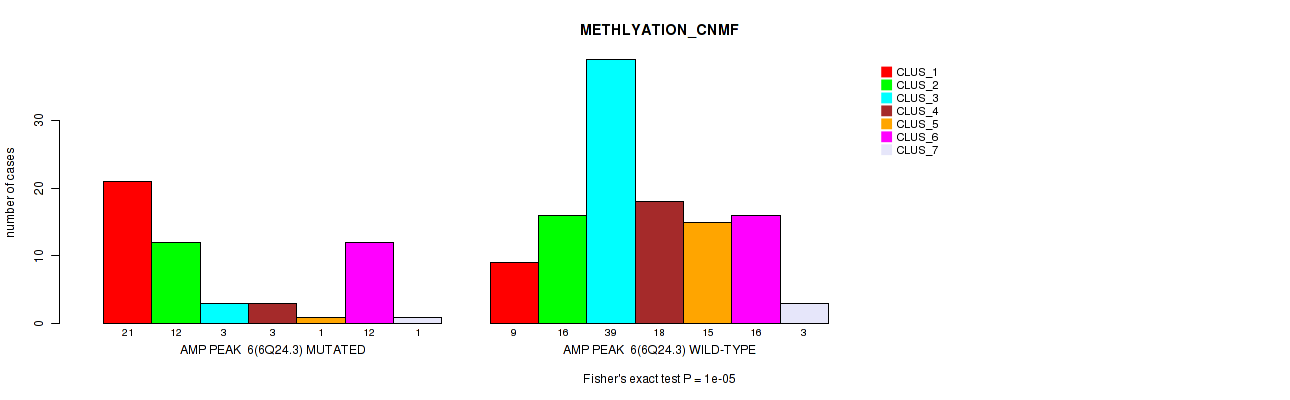
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S14. Gene #6: 'amp_6q24.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| AMP PEAK 6(6Q24.3) MUTATED | 15 | 28 | 5 | 4 |
| AMP PEAK 6(6Q24.3) WILD-TYPE | 24 | 17 | 19 | 54 |
Figure S14. Get High-res Image Gene #6: 'amp_6q24.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
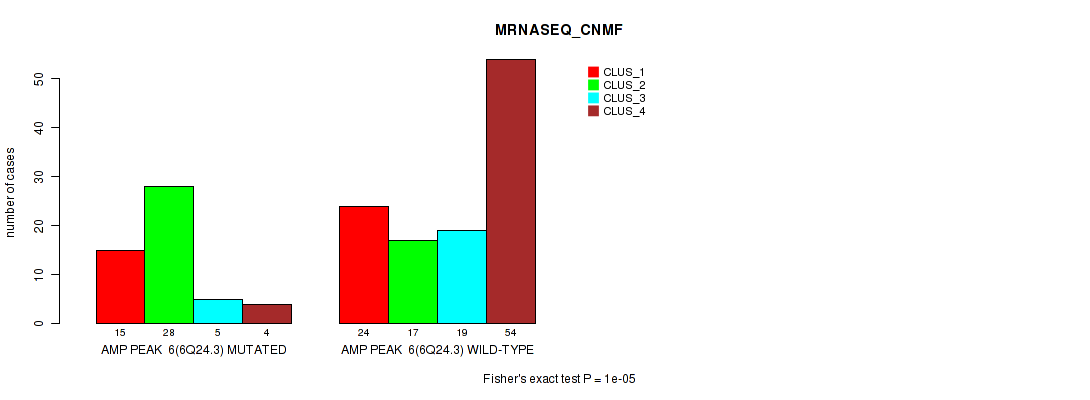
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S15. Gene #6: 'amp_6q24.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 45 | 32 | 33 | 37 | 19 |
| AMP PEAK 6(6Q24.3) MUTATED | 27 | 9 | 12 | 2 | 2 |
| AMP PEAK 6(6Q24.3) WILD-TYPE | 18 | 23 | 21 | 35 | 17 |
Figure S15. Get High-res Image Gene #6: 'amp_6q24.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
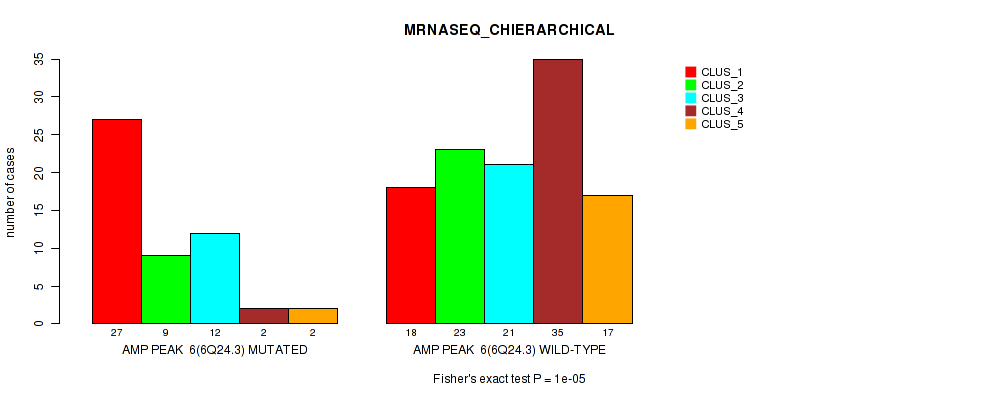
P value = 6e-05 (Fisher's exact test), Q value = 0.026
Table S16. Gene #6: 'amp_6q24.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 73 | 71 | 23 |
| AMP PEAK 6(6Q24.3) MUTATED | 32 | 10 | 11 |
| AMP PEAK 6(6Q24.3) WILD-TYPE | 41 | 61 | 12 |
Figure S16. Get High-res Image Gene #6: 'amp_6q24.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
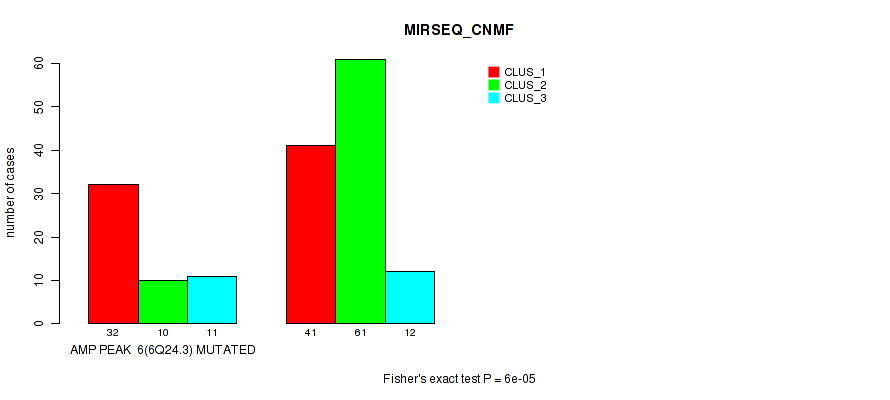
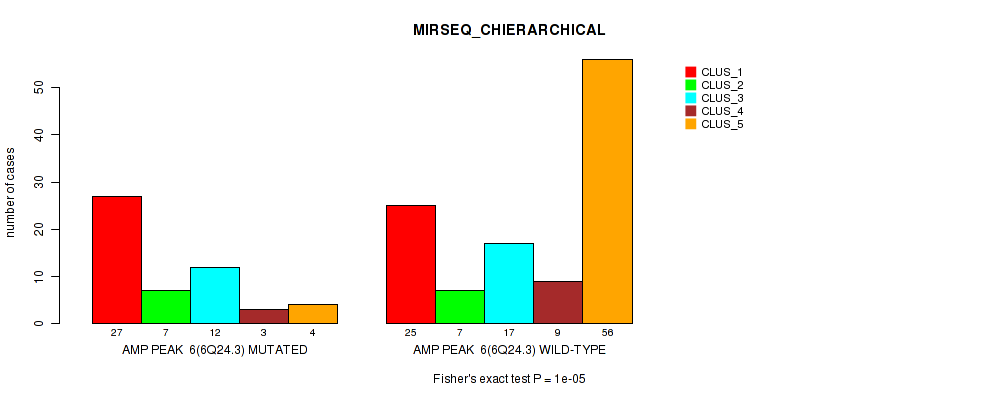
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S17. Gene #6: 'amp_6q24.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 52 | 14 | 29 | 12 | 60 |
| AMP PEAK 6(6Q24.3) MUTATED | 27 | 7 | 12 | 3 | 4 |
| AMP PEAK 6(6Q24.3) WILD-TYPE | 25 | 7 | 17 | 9 | 56 |
Figure S17. Get High-res Image Gene #6: 'amp_6q24.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
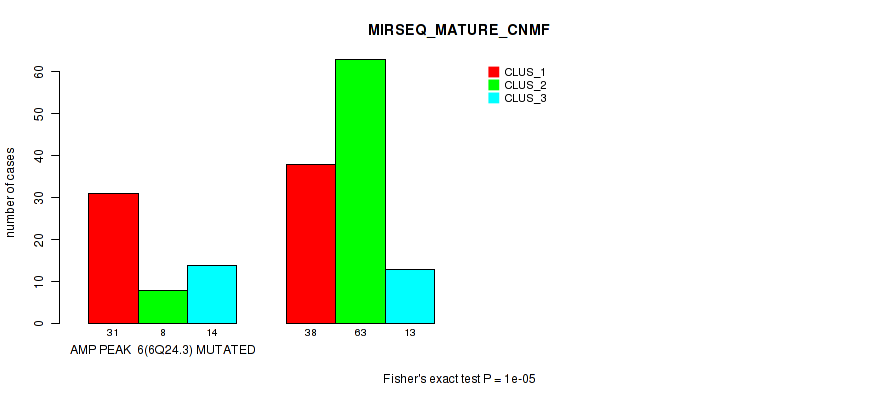
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S18. Gene #6: 'amp_6q24.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 69 | 71 | 27 |
| AMP PEAK 6(6Q24.3) MUTATED | 31 | 8 | 14 |
| AMP PEAK 6(6Q24.3) WILD-TYPE | 38 | 63 | 13 |
Figure S18. Get High-res Image Gene #6: 'amp_6q24.3' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
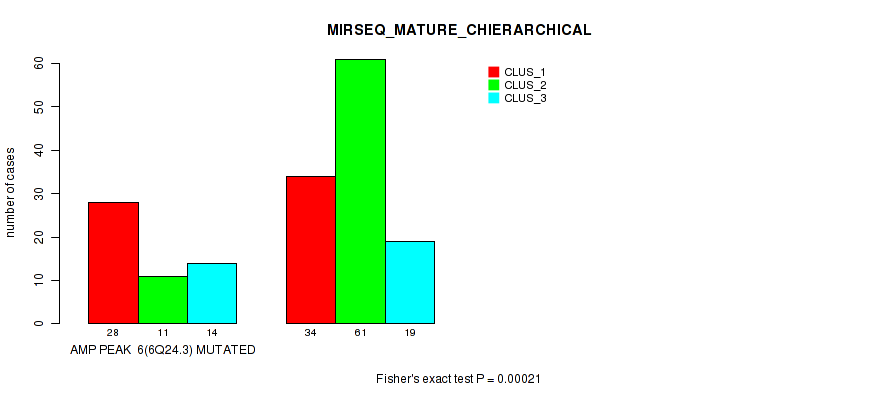
P value = 0.00021 (Fisher's exact test), Q value = 0.087
Table S19. Gene #6: 'amp_6q24.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 62 | 72 | 33 |
| AMP PEAK 6(6Q24.3) MUTATED | 28 | 11 | 14 |
| AMP PEAK 6(6Q24.3) WILD-TYPE | 34 | 61 | 19 |
Figure S19. Get High-res Image Gene #6: 'amp_6q24.3' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
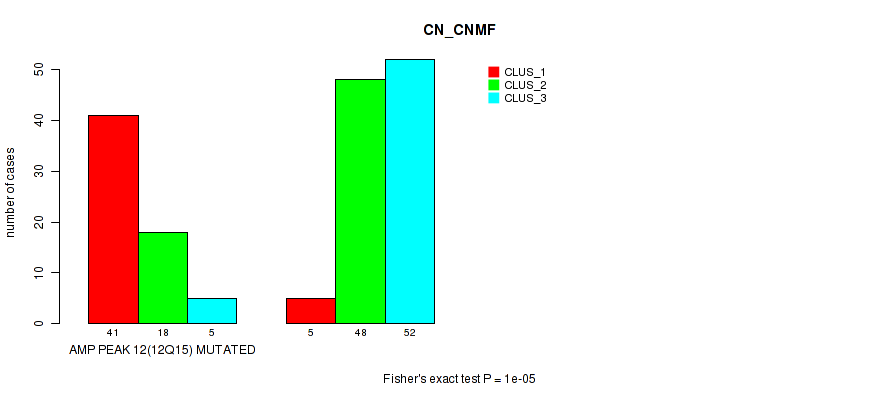
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S20. Gene #12: 'amp_12q15' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 12(12Q15) MUTATED | 41 | 18 | 5 |
| AMP PEAK 12(12Q15) WILD-TYPE | 5 | 48 | 52 |
Figure S20. Get High-res Image Gene #12: 'amp_12q15' versus Molecular Subtype #1: 'CN_CNMF'
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S21. Gene #12: 'amp_12q15' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| AMP PEAK 12(12Q15) MUTATED | 22 | 17 | 1 | 9 | 2 | 11 | 2 |
| AMP PEAK 12(12Q15) WILD-TYPE | 8 | 11 | 41 | 12 | 14 | 17 | 2 |
Figure S21. Get High-res Image Gene #12: 'amp_12q15' versus Molecular Subtype #2: 'METHLYATION_CNMF'
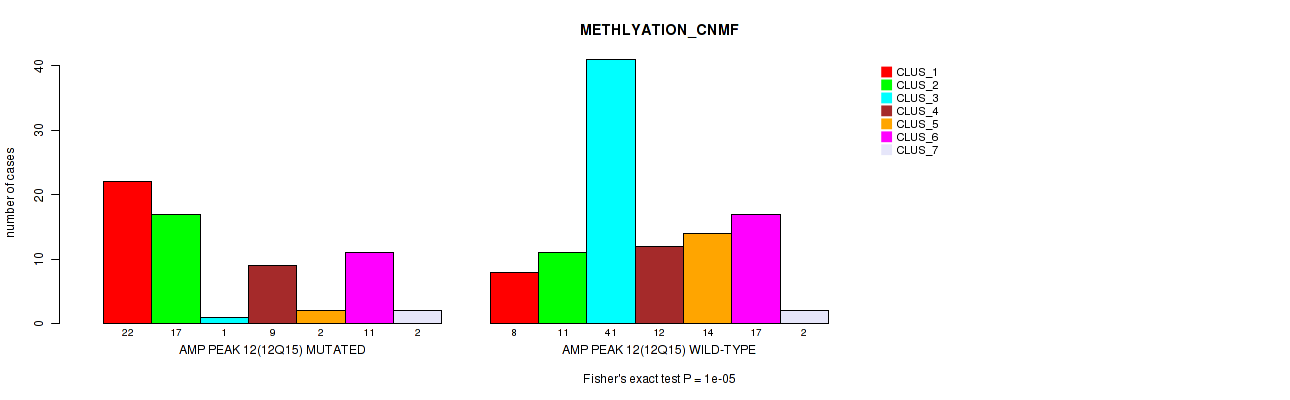
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S22. Gene #12: 'amp_12q15' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| AMP PEAK 12(12Q15) MUTATED | 17 | 32 | 12 | 2 |
| AMP PEAK 12(12Q15) WILD-TYPE | 22 | 13 | 12 | 56 |
Figure S22. Get High-res Image Gene #12: 'amp_12q15' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
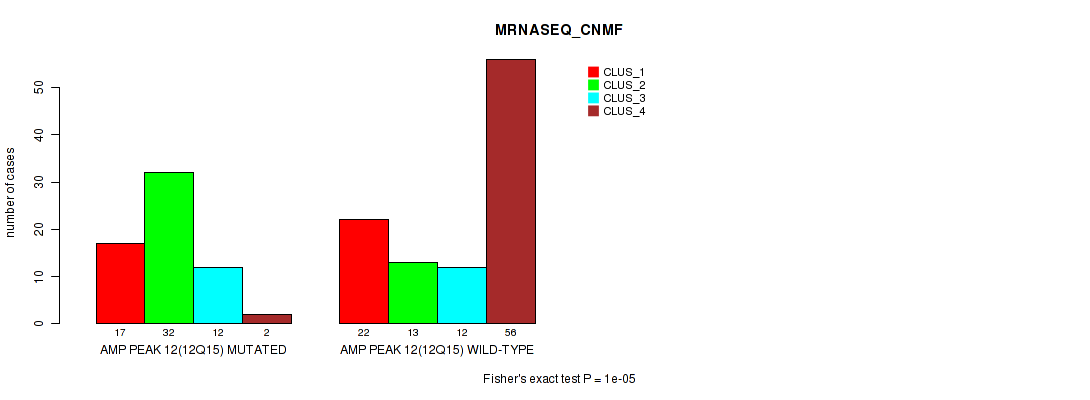
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S23. Gene #12: 'amp_12q15' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 45 | 32 | 33 | 37 | 19 |
| AMP PEAK 12(12Q15) MUTATED | 30 | 17 | 14 | 1 | 1 |
| AMP PEAK 12(12Q15) WILD-TYPE | 15 | 15 | 19 | 36 | 18 |
Figure S23. Get High-res Image Gene #12: 'amp_12q15' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
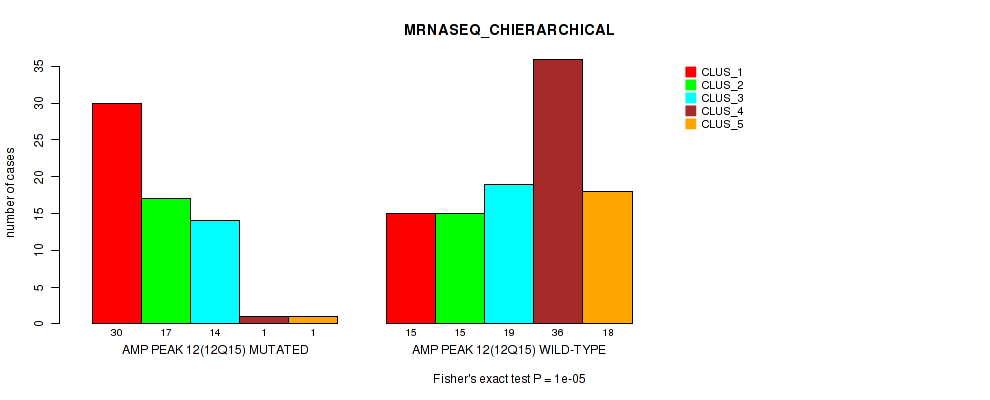
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S24. Gene #12: 'amp_12q15' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 73 | 71 | 23 |
| AMP PEAK 12(12Q15) MUTATED | 41 | 9 | 12 |
| AMP PEAK 12(12Q15) WILD-TYPE | 32 | 62 | 11 |
Figure S24. Get High-res Image Gene #12: 'amp_12q15' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
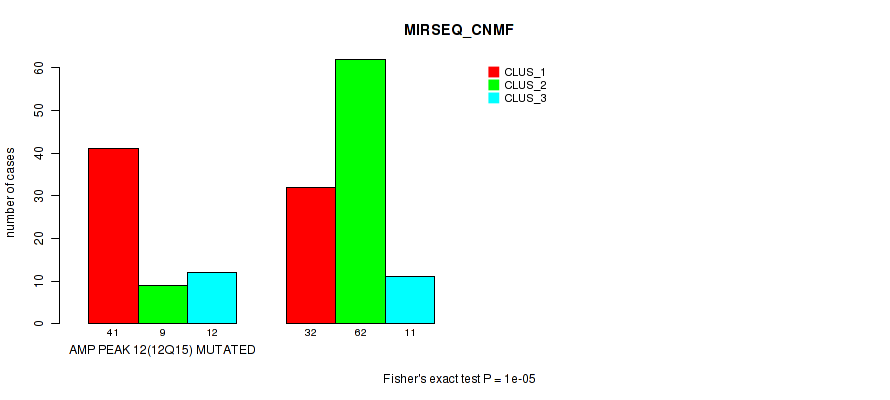
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S25. Gene #12: 'amp_12q15' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 52 | 14 | 29 | 12 | 60 |
| AMP PEAK 12(12Q15) MUTATED | 36 | 8 | 14 | 3 | 1 |
| AMP PEAK 12(12Q15) WILD-TYPE | 16 | 6 | 15 | 9 | 59 |
Figure S25. Get High-res Image Gene #12: 'amp_12q15' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
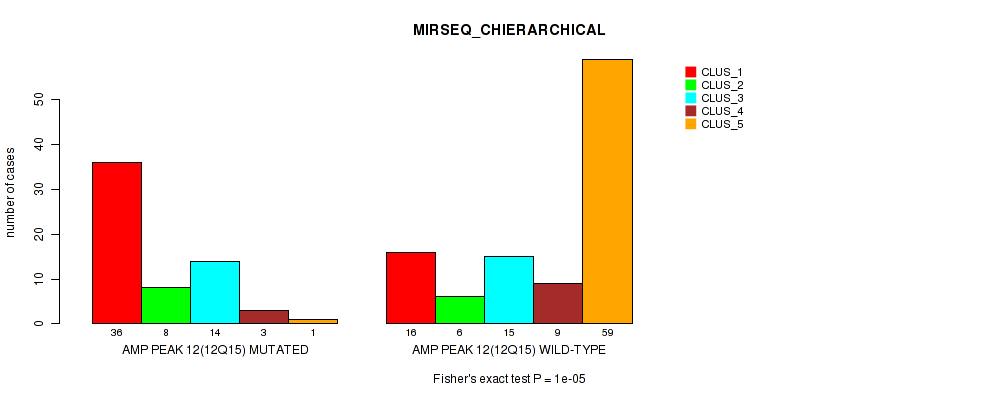
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S26. Gene #12: 'amp_12q15' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 69 | 71 | 27 |
| AMP PEAK 12(12Q15) MUTATED | 43 | 7 | 12 |
| AMP PEAK 12(12Q15) WILD-TYPE | 26 | 64 | 15 |
Figure S26. Get High-res Image Gene #12: 'amp_12q15' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
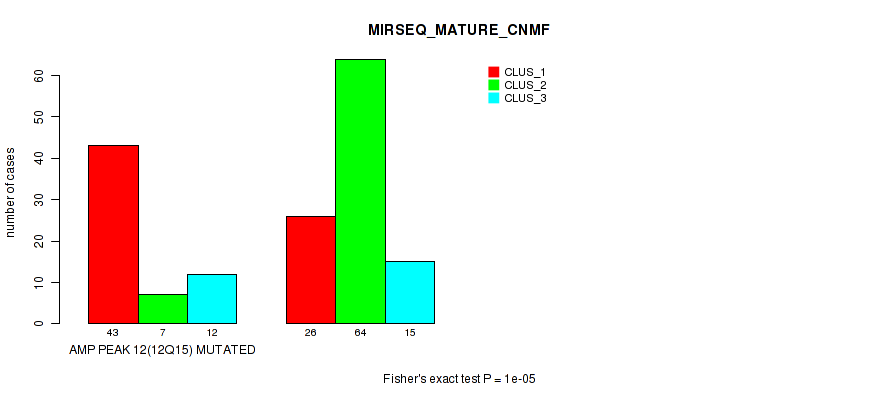
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S27. Gene #12: 'amp_12q15' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 62 | 72 | 33 |
| AMP PEAK 12(12Q15) MUTATED | 37 | 9 | 16 |
| AMP PEAK 12(12Q15) WILD-TYPE | 25 | 63 | 17 |
Figure S27. Get High-res Image Gene #12: 'amp_12q15' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
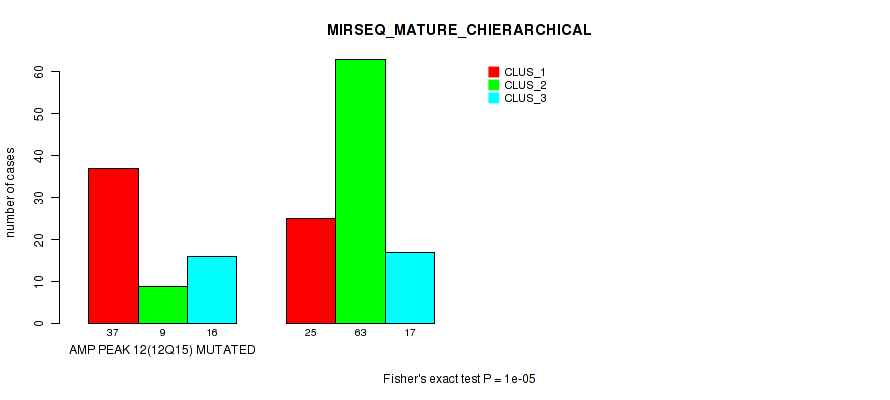
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S28. Gene #14: 'amp_17p11.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 14(17P11.2) MUTATED | 9 | 46 | 14 |
| AMP PEAK 14(17P11.2) WILD-TYPE | 37 | 20 | 43 |
Figure S28. Get High-res Image Gene #14: 'amp_17p11.2' versus Molecular Subtype #1: 'CN_CNMF'
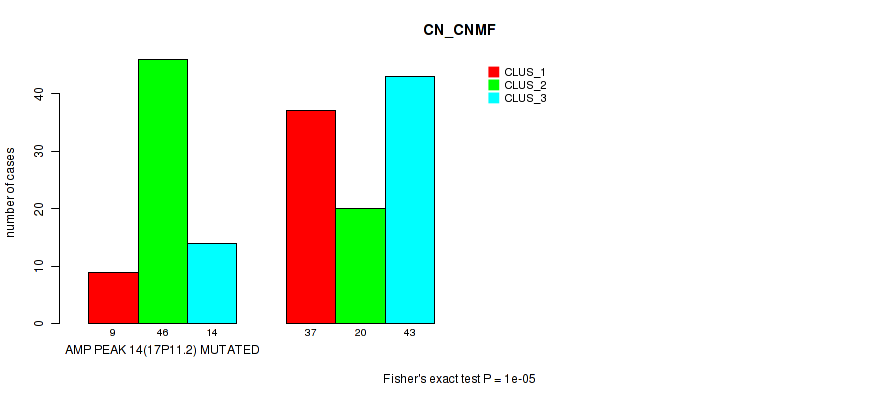
P value = 7e-05 (Fisher's exact test), Q value = 0.03
Table S29. Gene #14: 'amp_17p11.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| AMP PEAK 14(17P11.2) MUTATED | 14 | 6 | 21 | 6 | 1 | 20 | 1 |
| AMP PEAK 14(17P11.2) WILD-TYPE | 16 | 22 | 21 | 15 | 15 | 8 | 3 |
Figure S29. Get High-res Image Gene #14: 'amp_17p11.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
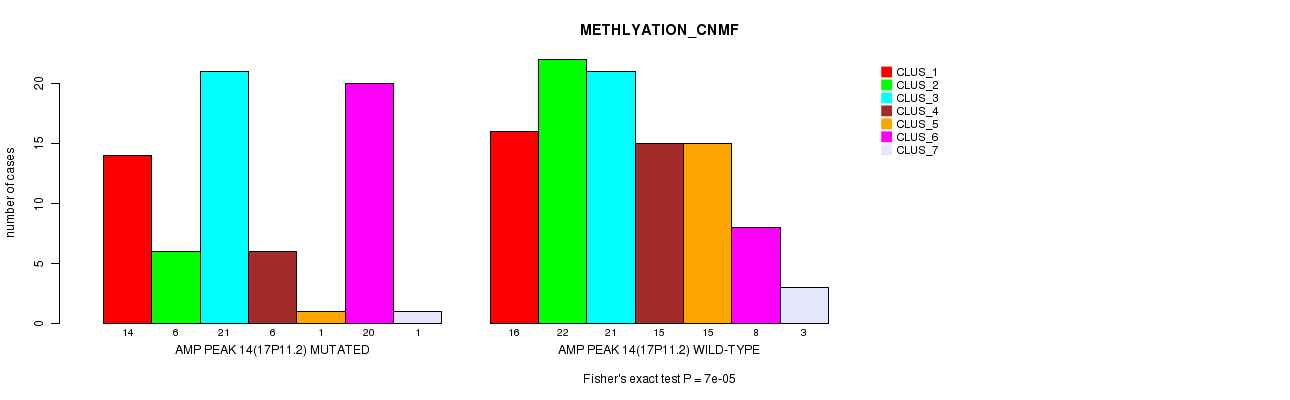
P value = 0.00014 (Fisher's exact test), Q value = 0.058
Table S30. Gene #15: 'amp_17q24.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| AMP PEAK 15(17Q24.3) MUTATED | 11 | 6 | 4 | 8 | 2 | 13 | 4 |
| AMP PEAK 15(17Q24.3) WILD-TYPE | 19 | 22 | 38 | 13 | 14 | 15 | 0 |
Figure S30. Get High-res Image Gene #15: 'amp_17q24.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
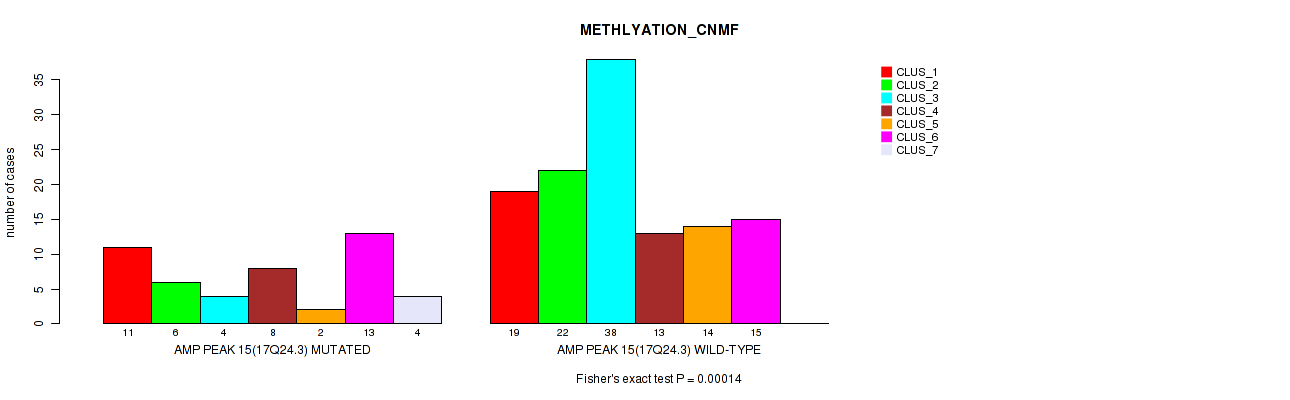
P value = 2e-05 (Fisher's exact test), Q value = 0.0089
Table S31. Gene #16: 'amp_19p13.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 16(19P13.2) MUTATED | 17 | 40 | 11 |
| AMP PEAK 16(19P13.2) WILD-TYPE | 29 | 26 | 46 |
Figure S31. Get High-res Image Gene #16: 'amp_19p13.2' versus Molecular Subtype #1: 'CN_CNMF'
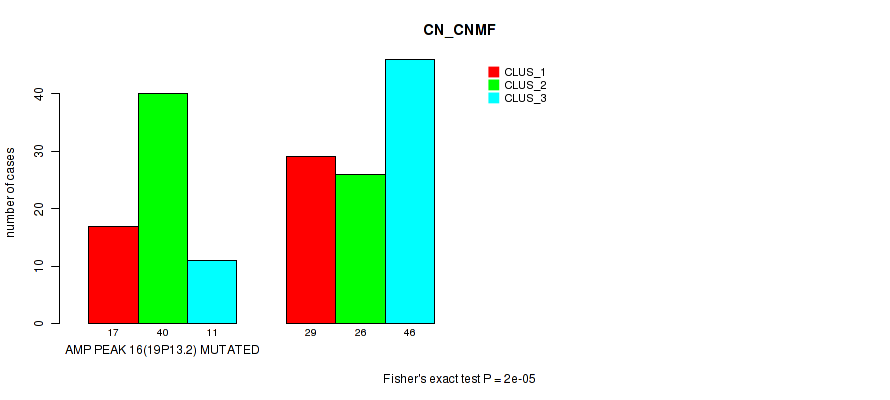
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S32. Gene #17: 'amp_19q12' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 17(19Q12) MUTATED | 14 | 39 | 8 |
| AMP PEAK 17(19Q12) WILD-TYPE | 32 | 27 | 49 |
Figure S32. Get High-res Image Gene #17: 'amp_19q12' versus Molecular Subtype #1: 'CN_CNMF'
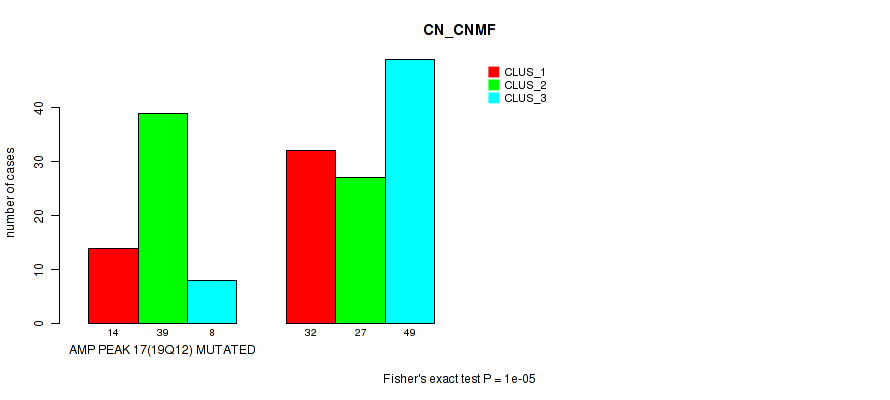
P value = 8e-05 (Fisher's exact test), Q value = 0.034
Table S33. Gene #18: 'amp_20q13.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 18(20Q13.33) MUTATED | 19 | 38 | 11 |
| AMP PEAK 18(20Q13.33) WILD-TYPE | 27 | 28 | 46 |
Figure S33. Get High-res Image Gene #18: 'amp_20q13.33' versus Molecular Subtype #1: 'CN_CNMF'
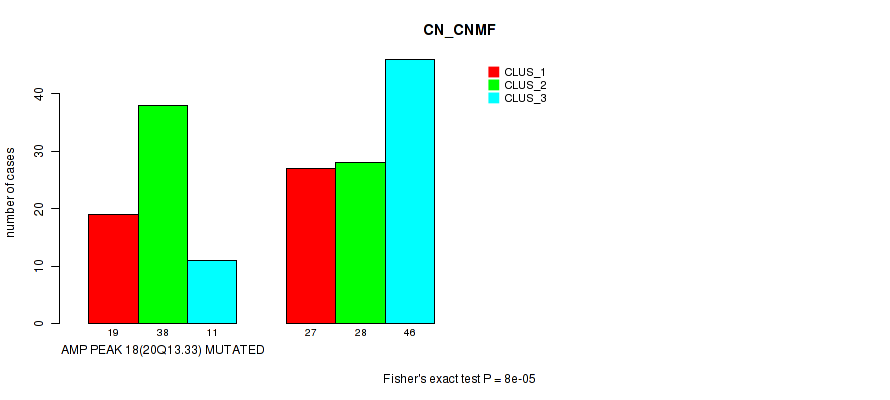
P value = 2e-04 (Fisher's exact test), Q value = 0.083
Table S34. Gene #19: 'amp_21q21.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 19(21Q21.1) MUTATED | 9 | 32 | 10 |
| AMP PEAK 19(21Q21.1) WILD-TYPE | 37 | 34 | 47 |
Figure S34. Get High-res Image Gene #19: 'amp_21q21.1' versus Molecular Subtype #1: 'CN_CNMF'
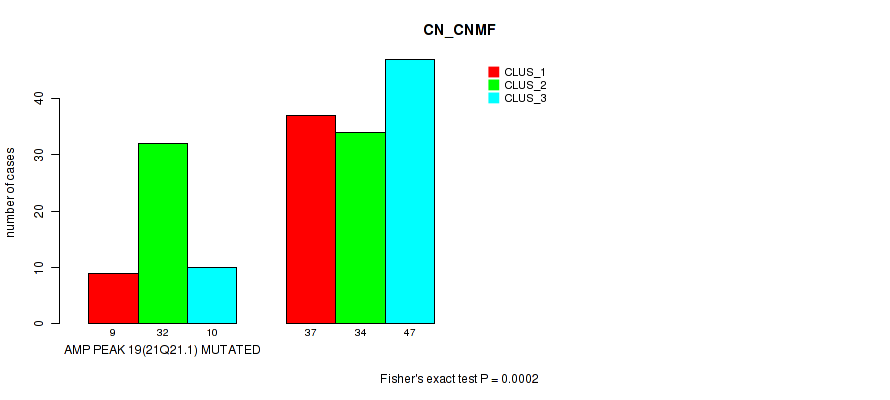
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S35. Gene #20: 'amp_xp21.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| AMP PEAK 20(XP21.2) MUTATED | 11 | 39 | 10 |
| AMP PEAK 20(XP21.2) WILD-TYPE | 35 | 27 | 47 |
Figure S35. Get High-res Image Gene #20: 'amp_xp21.2' versus Molecular Subtype #1: 'CN_CNMF'
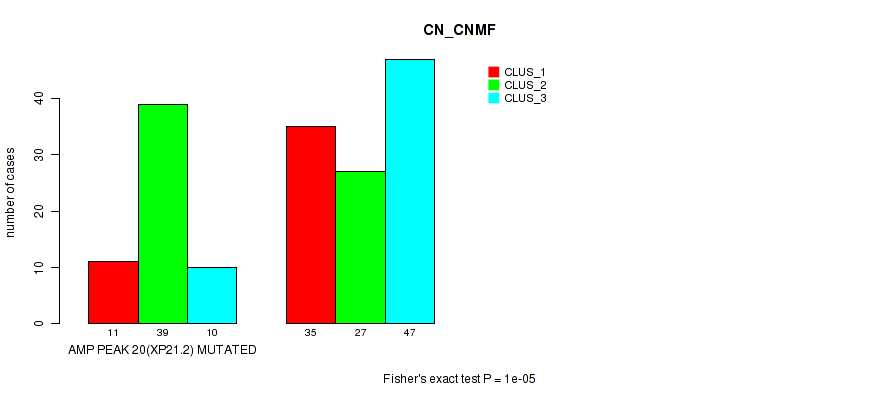
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S36. Gene #22: 'del_1p36.32' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 1(1P36.32) MUTATED | 7 | 1 | 27 | 5 | 6 | 3 | 2 |
| DEL PEAK 1(1P36.32) WILD-TYPE | 23 | 27 | 15 | 16 | 10 | 25 | 2 |
Figure S36. Get High-res Image Gene #22: 'del_1p36.32' versus Molecular Subtype #2: 'METHLYATION_CNMF'
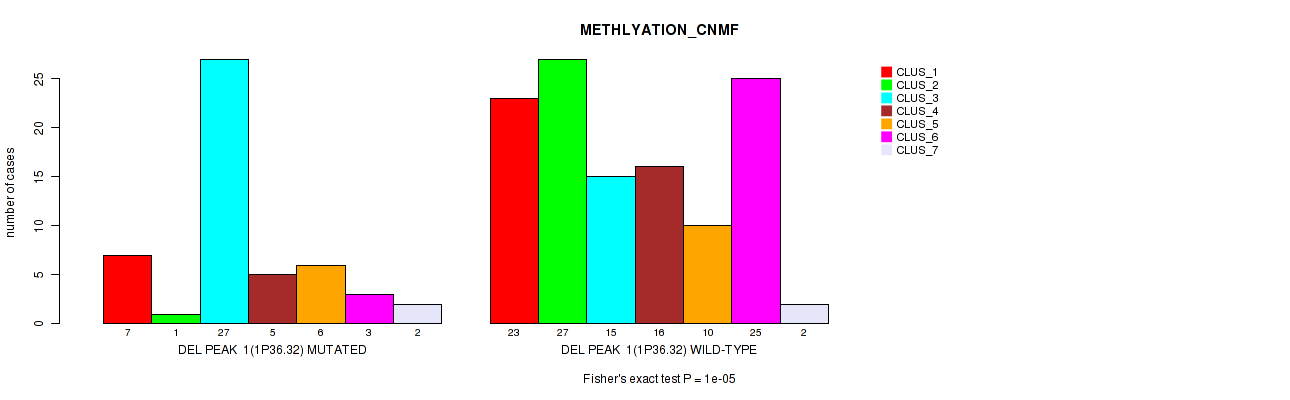
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S37. Gene #22: 'del_1p36.32' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| DEL PEAK 1(1P36.32) MUTATED | 5 | 7 | 4 | 34 |
| DEL PEAK 1(1P36.32) WILD-TYPE | 34 | 38 | 20 | 24 |
Figure S37. Get High-res Image Gene #22: 'del_1p36.32' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
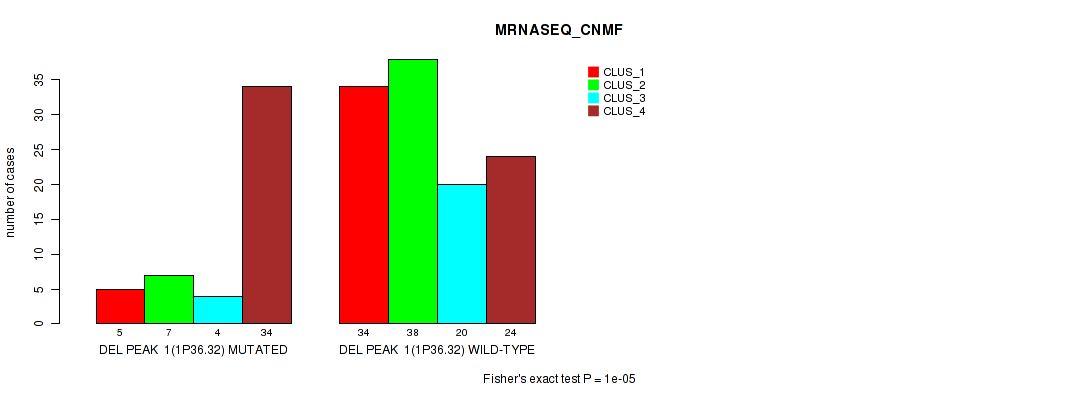
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S38. Gene #22: 'del_1p36.32' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 45 | 32 | 33 | 37 | 19 |
| DEL PEAK 1(1P36.32) MUTATED | 7 | 5 | 3 | 23 | 12 |
| DEL PEAK 1(1P36.32) WILD-TYPE | 38 | 27 | 30 | 14 | 7 |
Figure S38. Get High-res Image Gene #22: 'del_1p36.32' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
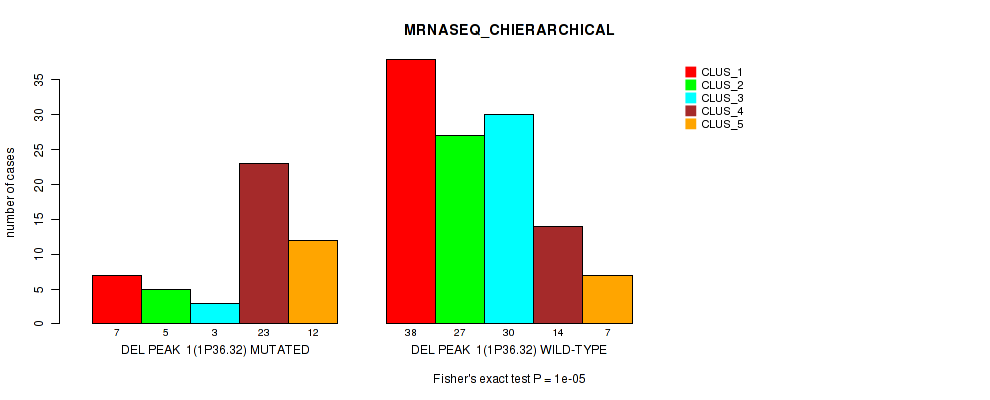
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S39. Gene #22: 'del_1p36.32' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 73 | 71 | 23 |
| DEL PEAK 1(1P36.32) MUTATED | 10 | 37 | 4 |
| DEL PEAK 1(1P36.32) WILD-TYPE | 63 | 34 | 19 |
Figure S39. Get High-res Image Gene #22: 'del_1p36.32' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
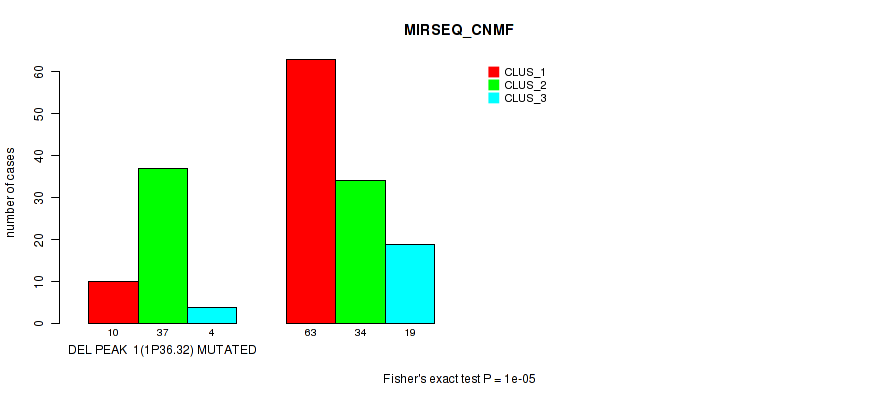
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S40. Gene #22: 'del_1p36.32' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 52 | 14 | 29 | 12 | 60 |
| DEL PEAK 1(1P36.32) MUTATED | 7 | 3 | 4 | 1 | 36 |
| DEL PEAK 1(1P36.32) WILD-TYPE | 45 | 11 | 25 | 11 | 24 |
Figure S40. Get High-res Image Gene #22: 'del_1p36.32' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
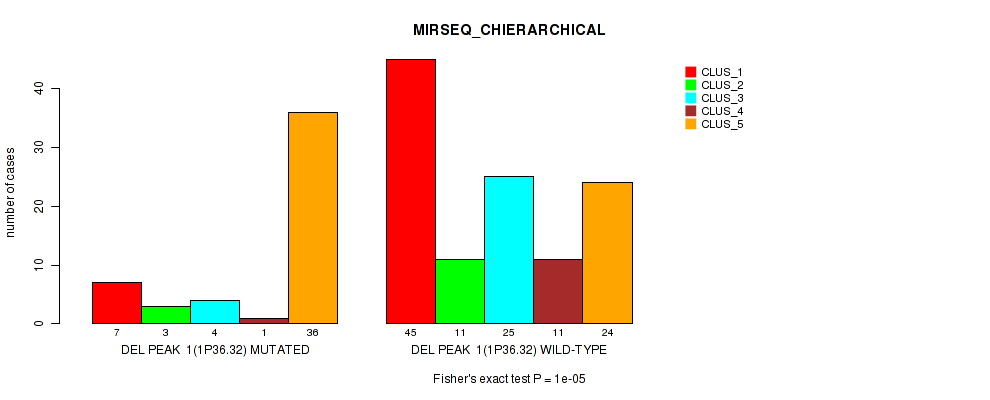
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S41. Gene #22: 'del_1p36.32' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 69 | 71 | 27 |
| DEL PEAK 1(1P36.32) MUTATED | 9 | 38 | 4 |
| DEL PEAK 1(1P36.32) WILD-TYPE | 60 | 33 | 23 |
Figure S41. Get High-res Image Gene #22: 'del_1p36.32' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
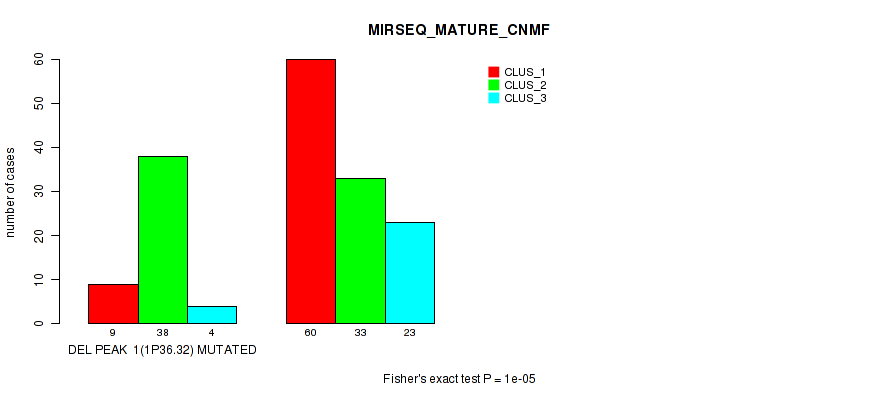
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S42. Gene #22: 'del_1p36.32' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 62 | 72 | 33 |
| DEL PEAK 1(1P36.32) MUTATED | 9 | 38 | 4 |
| DEL PEAK 1(1P36.32) WILD-TYPE | 53 | 34 | 29 |
Figure S42. Get High-res Image Gene #22: 'del_1p36.32' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
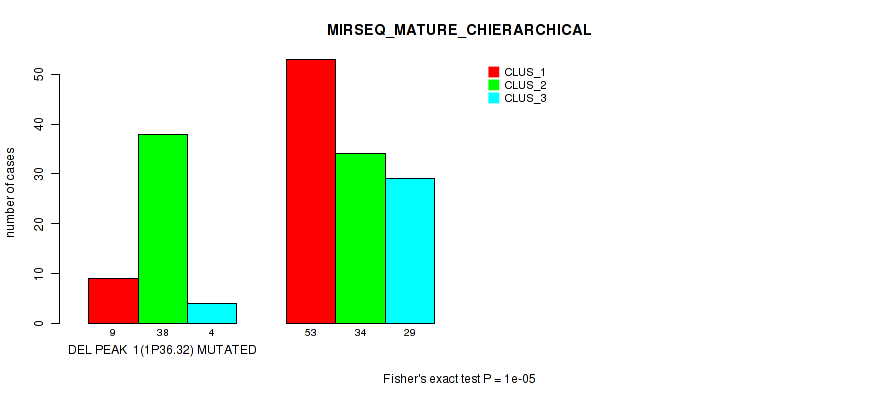
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S43. Gene #24: 'del_1q44' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 3(1Q44) MUTATED | 16 | 33 | 4 |
| DEL PEAK 3(1Q44) WILD-TYPE | 30 | 33 | 53 |
Figure S43. Get High-res Image Gene #24: 'del_1q44' versus Molecular Subtype #1: 'CN_CNMF'
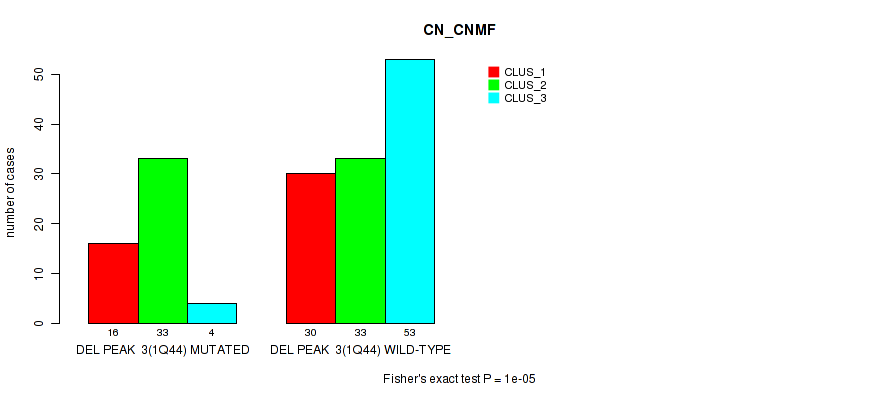
P value = 7e-05 (Fisher's exact test), Q value = 0.03
Table S44. Gene #24: 'del_1q44' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 3(1Q44) MUTATED | 18 | 5 | 8 | 3 | 2 | 15 | 2 |
| DEL PEAK 3(1Q44) WILD-TYPE | 12 | 23 | 34 | 18 | 14 | 13 | 2 |
Figure S44. Get High-res Image Gene #24: 'del_1q44' versus Molecular Subtype #2: 'METHLYATION_CNMF'
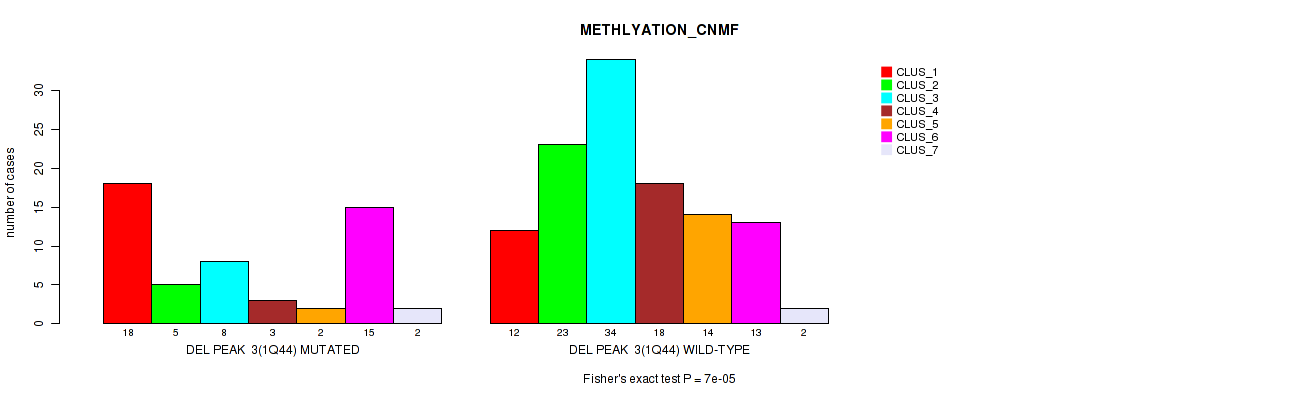
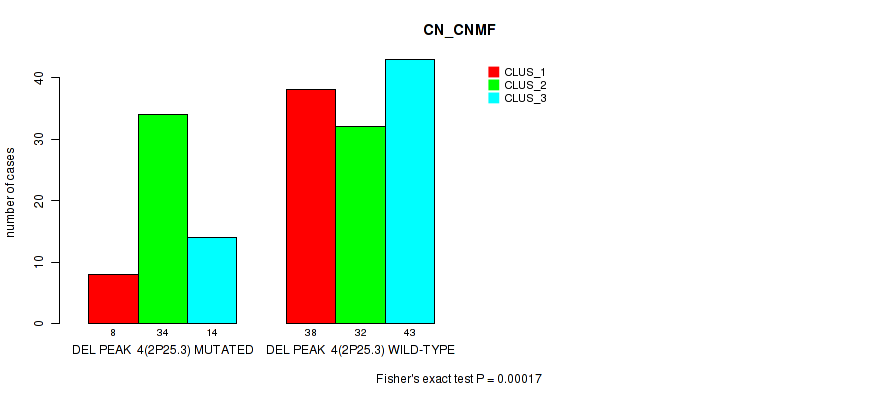
P value = 0.00017 (Fisher's exact test), Q value = 0.071
Table S45. Gene #25: 'del_2p25.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 4(2P25.3) MUTATED | 8 | 34 | 14 |
| DEL PEAK 4(2P25.3) WILD-TYPE | 38 | 32 | 43 |
Figure S45. Get High-res Image Gene #25: 'del_2p25.3' versus Molecular Subtype #1: 'CN_CNMF'
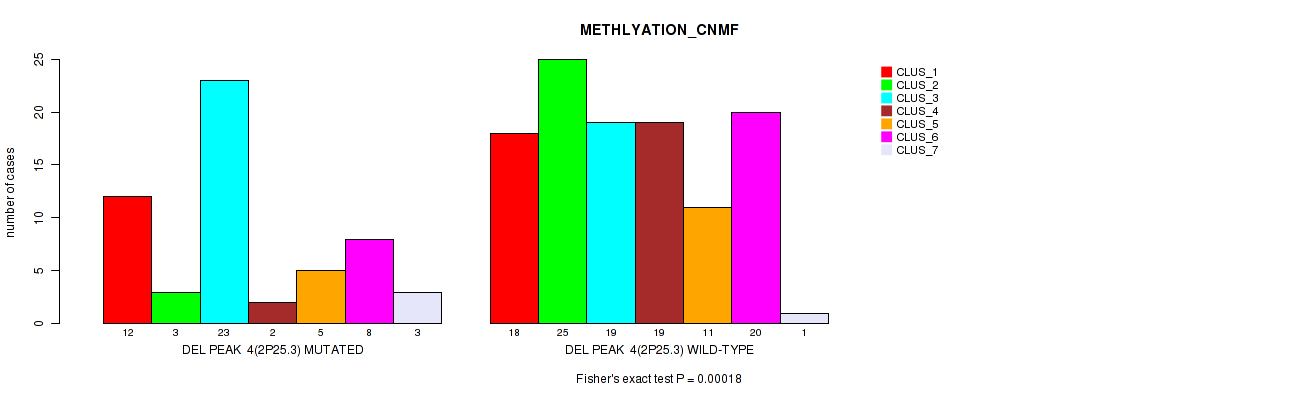
P value = 0.00018 (Fisher's exact test), Q value = 0.075
Table S46. Gene #25: 'del_2p25.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 4(2P25.3) MUTATED | 12 | 3 | 23 | 2 | 5 | 8 | 3 |
| DEL PEAK 4(2P25.3) WILD-TYPE | 18 | 25 | 19 | 19 | 11 | 20 | 1 |
Figure S46. Get High-res Image Gene #25: 'del_2p25.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
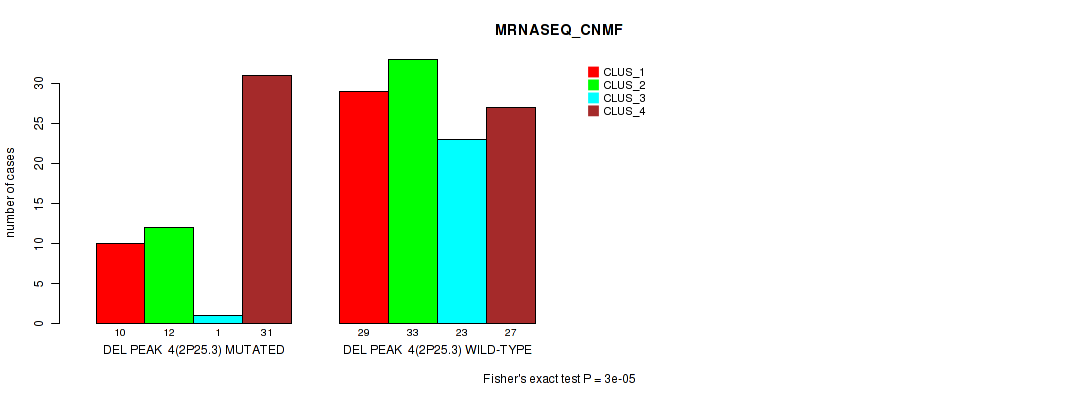
P value = 3e-05 (Fisher's exact test), Q value = 0.013
Table S47. Gene #25: 'del_2p25.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| DEL PEAK 4(2P25.3) MUTATED | 10 | 12 | 1 | 31 |
| DEL PEAK 4(2P25.3) WILD-TYPE | 29 | 33 | 23 | 27 |
Figure S47. Get High-res Image Gene #25: 'del_2p25.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
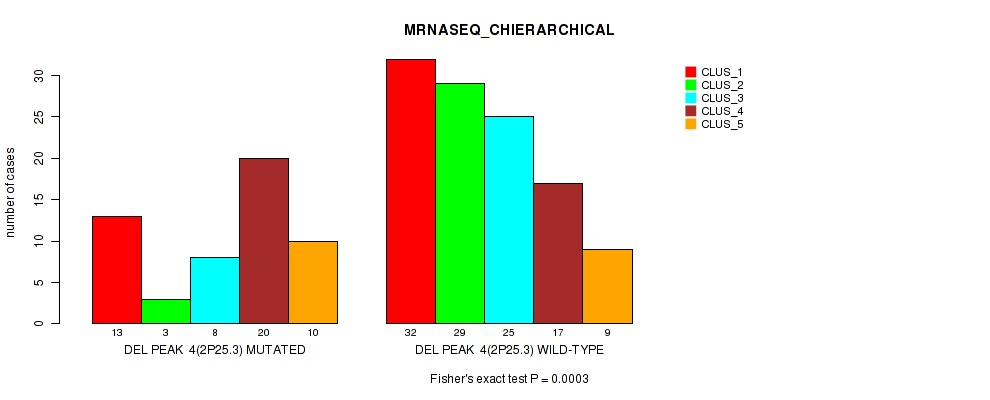
P value = 3e-04 (Fisher's exact test), Q value = 0.12
Table S48. Gene #25: 'del_2p25.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 45 | 32 | 33 | 37 | 19 |
| DEL PEAK 4(2P25.3) MUTATED | 13 | 3 | 8 | 20 | 10 |
| DEL PEAK 4(2P25.3) WILD-TYPE | 32 | 29 | 25 | 17 | 9 |
Figure S48. Get High-res Image Gene #25: 'del_2p25.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S49. Gene #26: 'del_2q37.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 5(2Q37.3) MUTATED | 13 | 46 | 9 |
| DEL PEAK 5(2Q37.3) WILD-TYPE | 33 | 20 | 48 |
Figure S49. Get High-res Image Gene #26: 'del_2q37.3' versus Molecular Subtype #1: 'CN_CNMF'
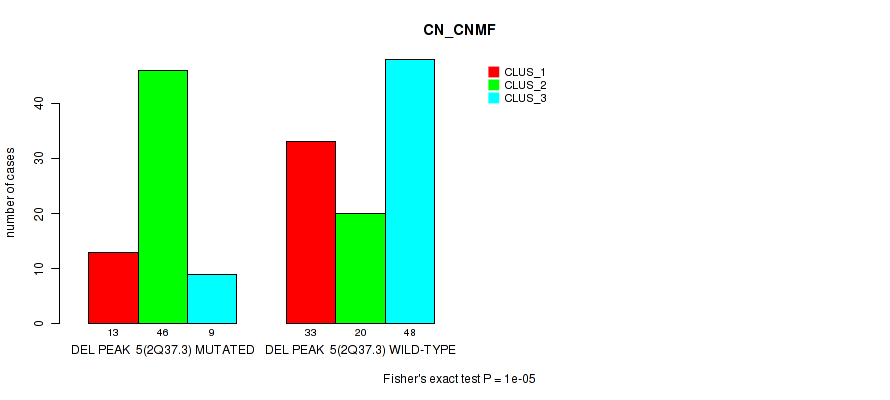
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S50. Gene #27: 'del_2q37.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 6(2Q37.3) MUTATED | 14 | 47 | 9 |
| DEL PEAK 6(2Q37.3) WILD-TYPE | 32 | 19 | 48 |
Figure S50. Get High-res Image Gene #27: 'del_2q37.3' versus Molecular Subtype #1: 'CN_CNMF'
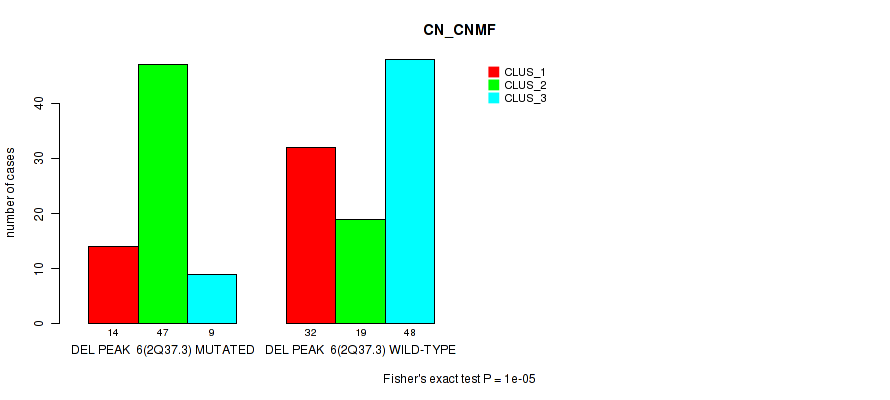
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S51. Gene #28: 'del_3p21.31' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 7(3P21.31) MUTATED | 16 | 3 | 1 | 7 | 2 | 8 | 0 |
| DEL PEAK 7(3P21.31) WILD-TYPE | 14 | 25 | 41 | 14 | 14 | 20 | 4 |
Figure S51. Get High-res Image Gene #28: 'del_3p21.31' versus Molecular Subtype #2: 'METHLYATION_CNMF'
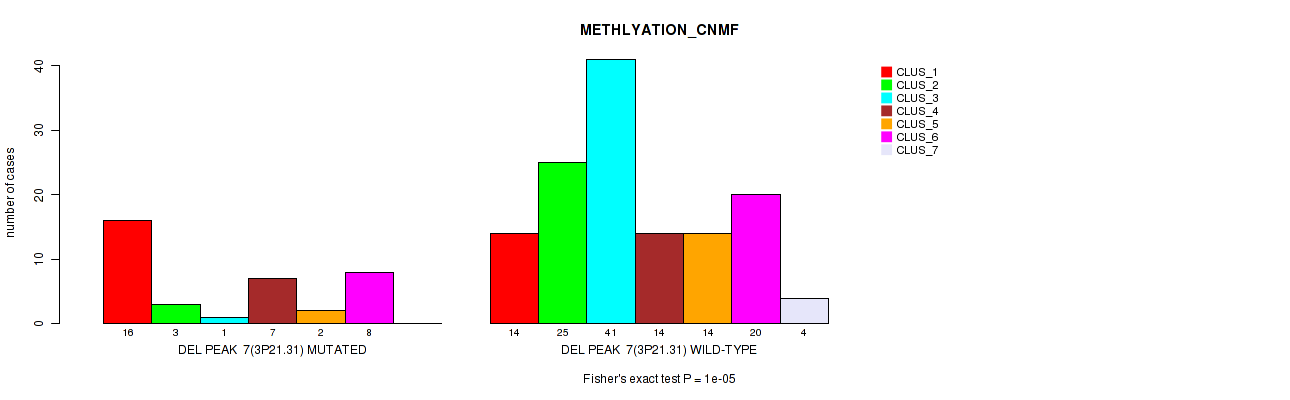
P value = 0.00013 (Fisher's exact test), Q value = 0.055
Table S52. Gene #28: 'del_3p21.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| DEL PEAK 7(3P21.31) MUTATED | 9 | 16 | 9 | 3 |
| DEL PEAK 7(3P21.31) WILD-TYPE | 30 | 29 | 15 | 55 |
Figure S52. Get High-res Image Gene #28: 'del_3p21.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
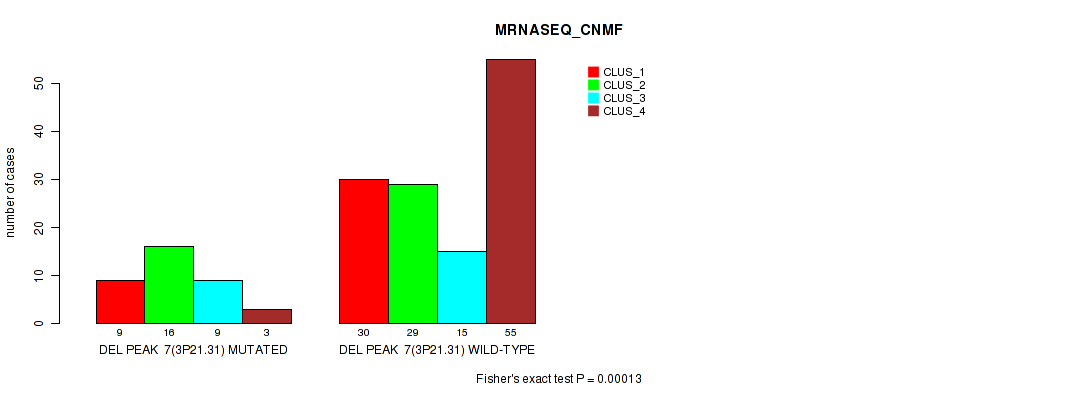
P value = 0.00017 (Fisher's exact test), Q value = 0.071
Table S53. Gene #28: 'del_3p21.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 52 | 14 | 29 | 12 | 60 |
| DEL PEAK 7(3P21.31) MUTATED | 19 | 6 | 4 | 4 | 4 |
| DEL PEAK 7(3P21.31) WILD-TYPE | 33 | 8 | 25 | 8 | 56 |
Figure S53. Get High-res Image Gene #28: 'del_3p21.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
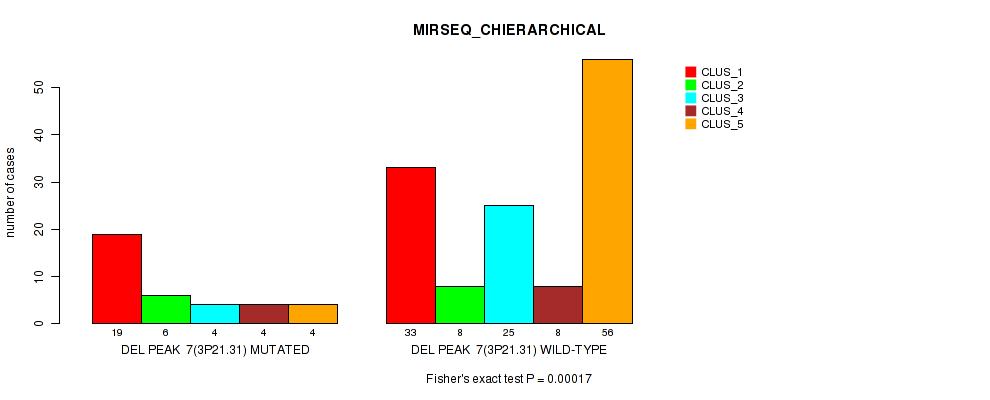
P value = 0.00034 (Fisher's exact test), Q value = 0.14
Table S54. Gene #29: 'del_3q29' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 8(3Q29) MUTATED | 13 | 2 | 7 | 5 | 0 | 9 | 3 |
| DEL PEAK 8(3Q29) WILD-TYPE | 17 | 26 | 35 | 16 | 16 | 19 | 1 |
Figure S54. Get High-res Image Gene #29: 'del_3q29' versus Molecular Subtype #2: 'METHLYATION_CNMF'
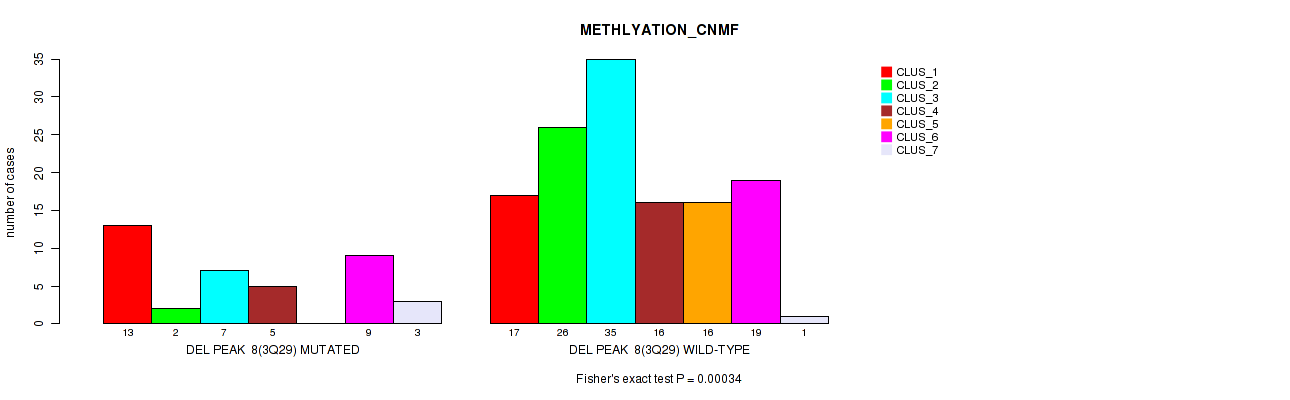
P value = 2e-05 (Fisher's exact test), Q value = 0.0089
Table S55. Gene #30: 'del_4q35.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 9(4Q35.1) MUTATED | 15 | 39 | 9 |
| DEL PEAK 9(4Q35.1) WILD-TYPE | 31 | 27 | 48 |
Figure S55. Get High-res Image Gene #30: 'del_4q35.1' versus Molecular Subtype #1: 'CN_CNMF'
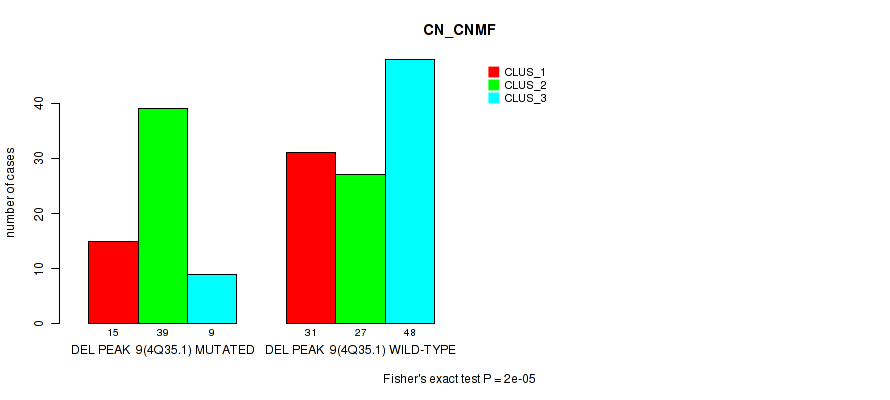
P value = 0.00028 (Fisher's exact test), Q value = 0.11
Table S56. Gene #32: 'del_6q14.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 11(6Q14.1) MUTATED | 8 | 31 | 10 |
| DEL PEAK 11(6Q14.1) WILD-TYPE | 38 | 35 | 47 |
Figure S56. Get High-res Image Gene #32: 'del_6q14.1' versus Molecular Subtype #1: 'CN_CNMF'
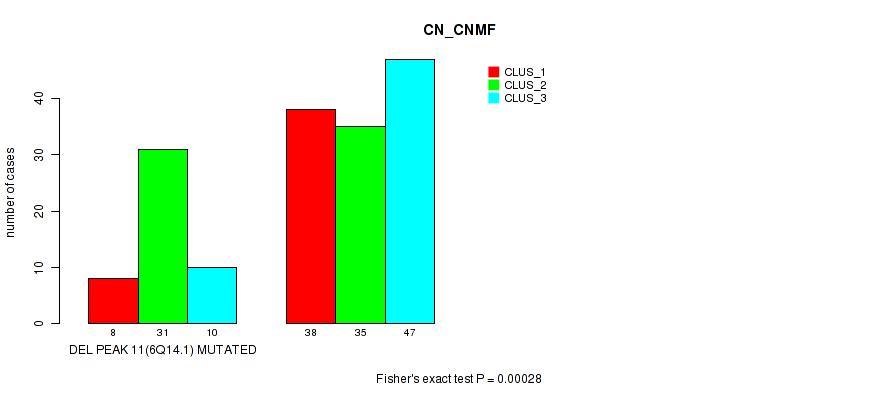
P value = 6e-05 (Fisher's exact test), Q value = 0.026
Table S57. Gene #34: 'del_7q36.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 13(7Q36.3) MUTATED | 14 | 26 | 4 |
| DEL PEAK 13(7Q36.3) WILD-TYPE | 32 | 40 | 53 |
Figure S57. Get High-res Image Gene #34: 'del_7q36.3' versus Molecular Subtype #1: 'CN_CNMF'
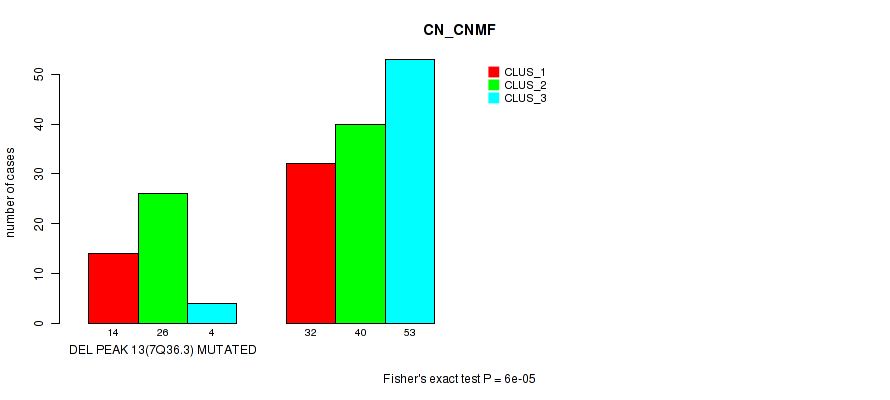
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S58. Gene #35: 'del_8p23.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 14(8P23.3) MUTATED | 14 | 5 | 4 | 9 | 2 | 15 | 3 |
| DEL PEAK 14(8P23.3) WILD-TYPE | 16 | 23 | 38 | 12 | 14 | 13 | 1 |
Figure S58. Get High-res Image Gene #35: 'del_8p23.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
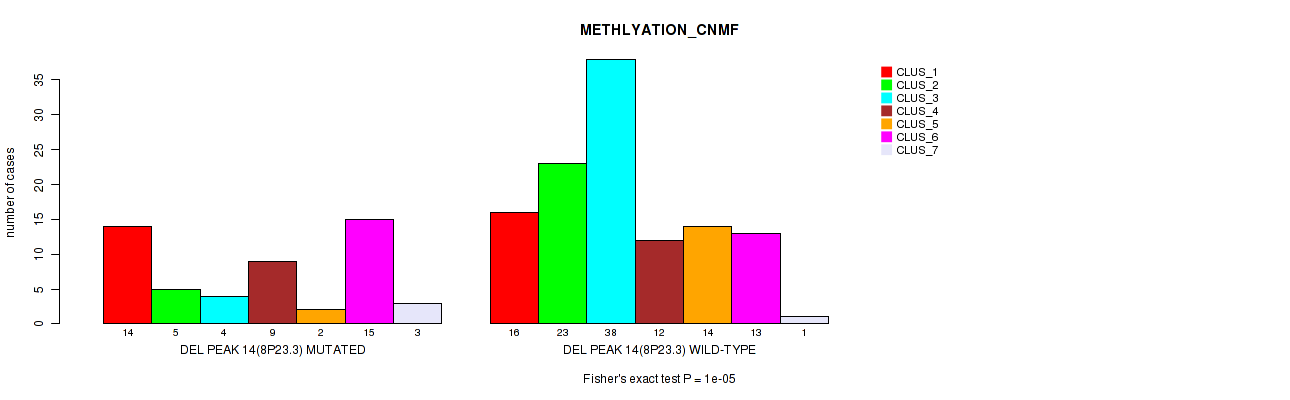
P value = 0.00043 (Fisher's exact test), Q value = 0.17
Table S59. Gene #38: 'del_9q34.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 17(9Q34.3) MUTATED | 7 | 29 | 10 |
| DEL PEAK 17(9Q34.3) WILD-TYPE | 39 | 37 | 47 |
Figure S59. Get High-res Image Gene #38: 'del_9q34.3' versus Molecular Subtype #1: 'CN_CNMF'
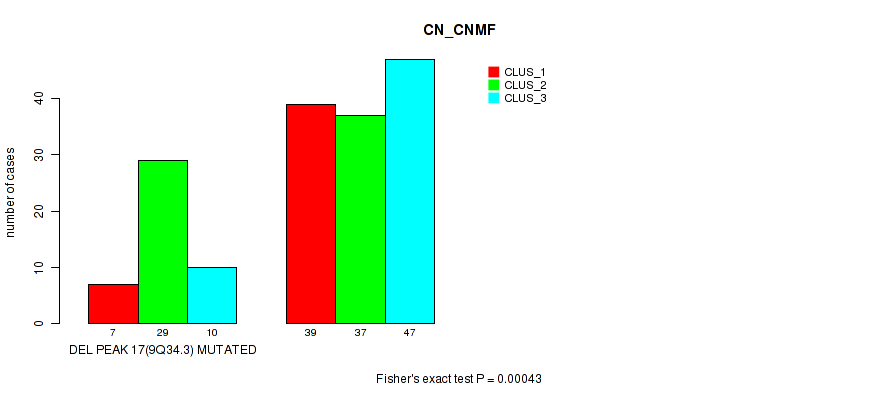
P value = 0.00036 (Fisher's exact test), Q value = 0.15
Table S60. Gene #39: 'del_10p15.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 18(10P15.3) MUTATED | 20 | 6 | 28 | 10 | 4 | 18 | 2 |
| DEL PEAK 18(10P15.3) WILD-TYPE | 10 | 22 | 14 | 11 | 12 | 10 | 2 |
Figure S60. Get High-res Image Gene #39: 'del_10p15.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
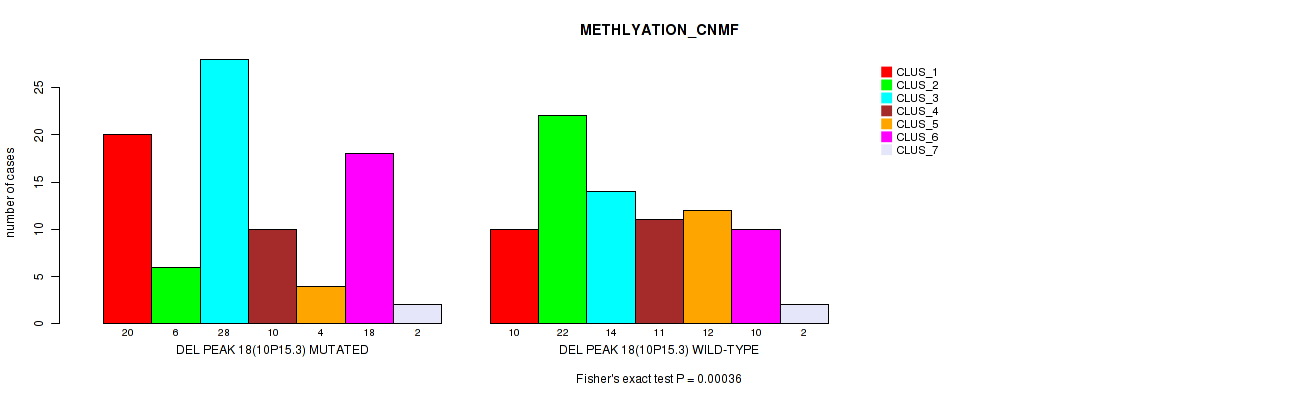
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S61. Gene #40: 'del_10q23.31' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 19(10Q23.31) MUTATED | 8 | 54 | 33 |
| DEL PEAK 19(10Q23.31) WILD-TYPE | 38 | 12 | 24 |
Figure S61. Get High-res Image Gene #40: 'del_10q23.31' versus Molecular Subtype #1: 'CN_CNMF'
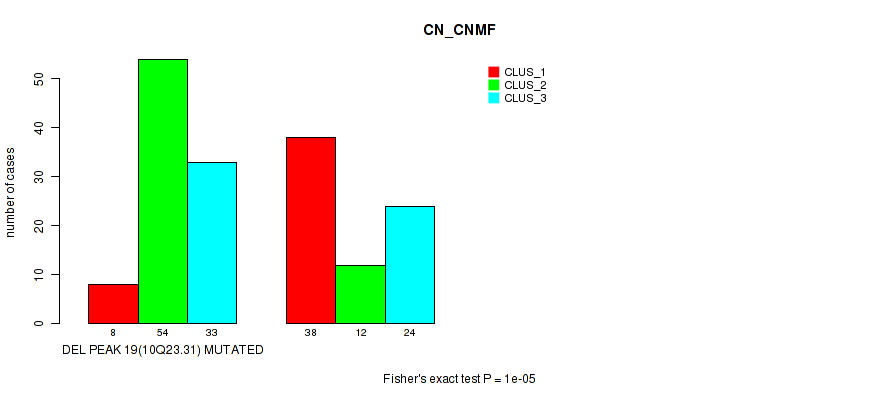
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S62. Gene #40: 'del_10q23.31' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 19(10Q23.31) MUTATED | 17 | 4 | 39 | 5 | 6 | 21 | 3 |
| DEL PEAK 19(10Q23.31) WILD-TYPE | 13 | 24 | 3 | 16 | 10 | 7 | 1 |
Figure S62. Get High-res Image Gene #40: 'del_10q23.31' versus Molecular Subtype #2: 'METHLYATION_CNMF'
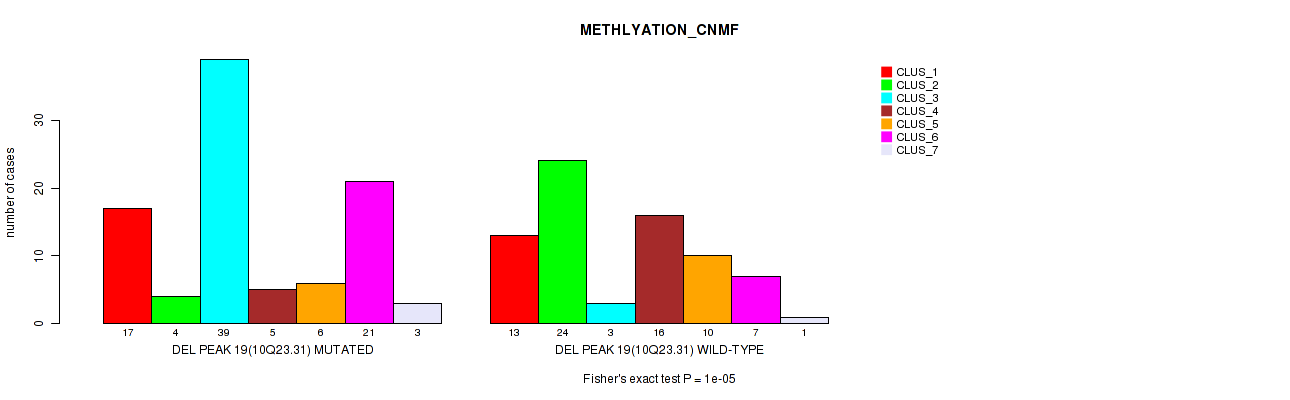
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S63. Gene #40: 'del_10q23.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| DEL PEAK 19(10Q23.31) MUTATED | 24 | 15 | 6 | 48 |
| DEL PEAK 19(10Q23.31) WILD-TYPE | 15 | 30 | 18 | 10 |
Figure S63. Get High-res Image Gene #40: 'del_10q23.31' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
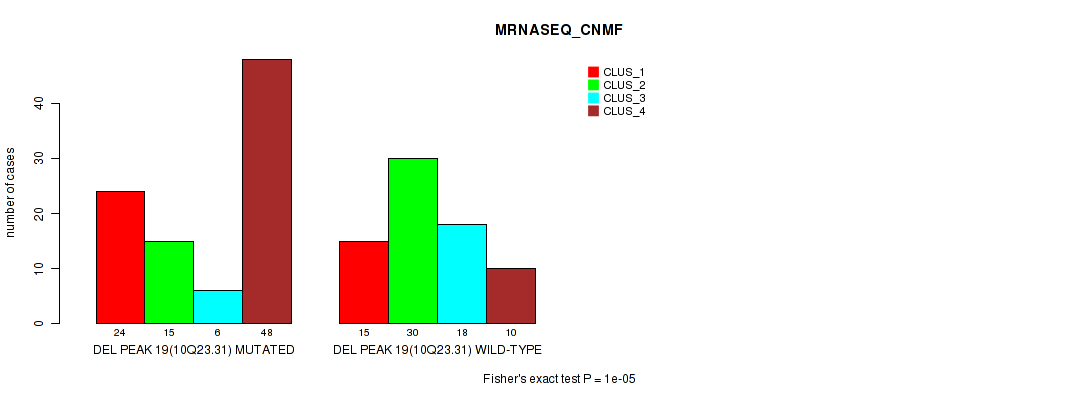
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S64. Gene #40: 'del_10q23.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 45 | 32 | 33 | 37 | 19 |
| DEL PEAK 19(10Q23.31) MUTATED | 18 | 8 | 20 | 36 | 11 |
| DEL PEAK 19(10Q23.31) WILD-TYPE | 27 | 24 | 13 | 1 | 8 |
Figure S64. Get High-res Image Gene #40: 'del_10q23.31' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
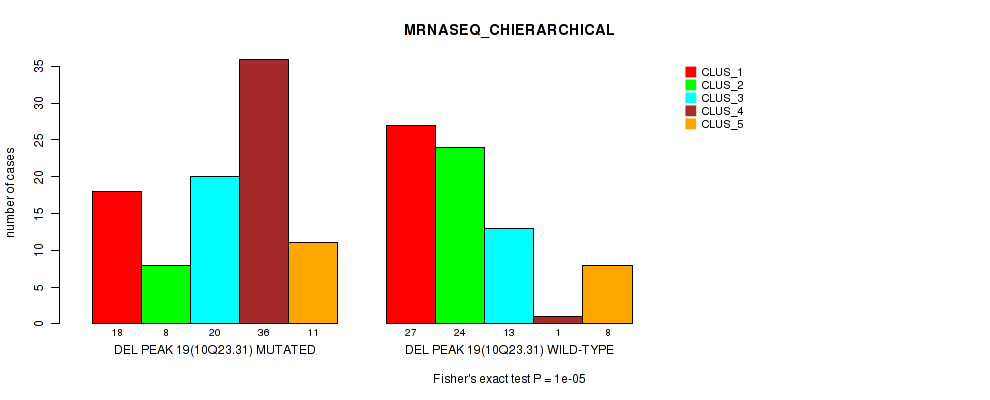
P value = 0.00026 (Fisher's exact test), Q value = 0.11
Table S65. Gene #40: 'del_10q23.31' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 73 | 71 | 23 |
| DEL PEAK 19(10Q23.31) MUTATED | 29 | 52 | 13 |
| DEL PEAK 19(10Q23.31) WILD-TYPE | 44 | 19 | 10 |
Figure S65. Get High-res Image Gene #40: 'del_10q23.31' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
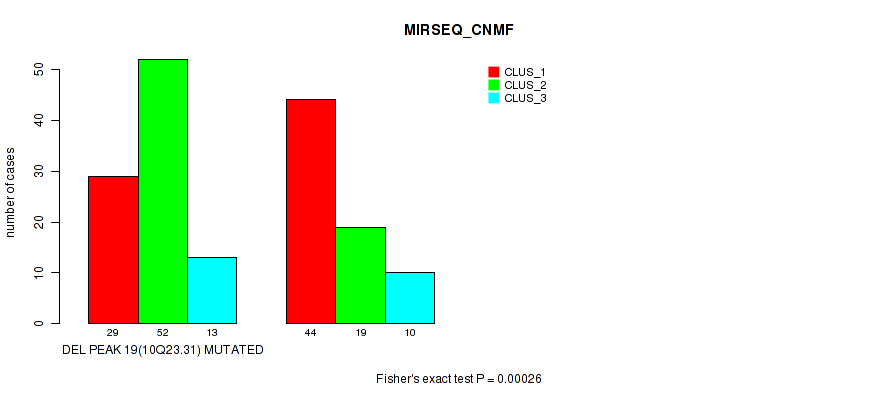
P value = 5e-05 (Fisher's exact test), Q value = 0.022
Table S66. Gene #40: 'del_10q23.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 52 | 14 | 29 | 12 | 60 |
| DEL PEAK 19(10Q23.31) MUTATED | 24 | 6 | 14 | 2 | 48 |
| DEL PEAK 19(10Q23.31) WILD-TYPE | 28 | 8 | 15 | 10 | 12 |
Figure S66. Get High-res Image Gene #40: 'del_10q23.31' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
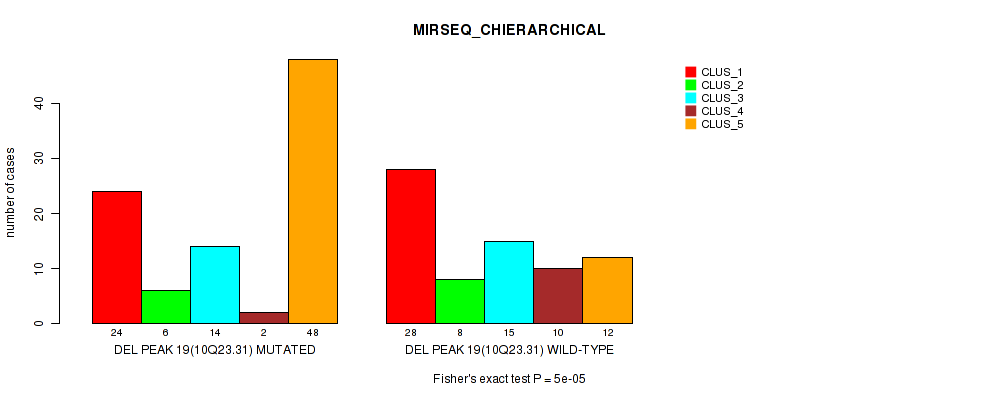
P value = 2e-05 (Fisher's exact test), Q value = 0.0089
Table S67. Gene #40: 'del_10q23.31' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 69 | 71 | 27 |
| DEL PEAK 19(10Q23.31) MUTATED | 26 | 54 | 14 |
| DEL PEAK 19(10Q23.31) WILD-TYPE | 43 | 17 | 13 |
Figure S67. Get High-res Image Gene #40: 'del_10q23.31' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
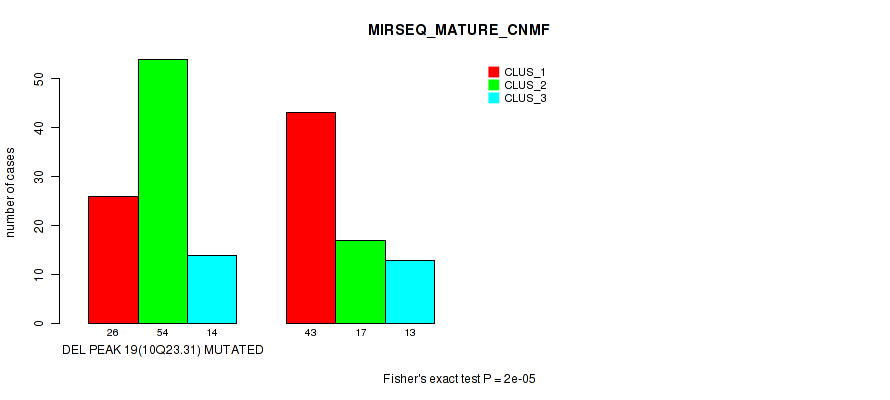
P value = 0.00051 (Fisher's exact test), Q value = 0.2
Table S68. Gene #40: 'del_10q23.31' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 62 | 72 | 33 |
| DEL PEAK 19(10Q23.31) MUTATED | 24 | 52 | 18 |
| DEL PEAK 19(10Q23.31) WILD-TYPE | 38 | 20 | 15 |
Figure S68. Get High-res Image Gene #40: 'del_10q23.31' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
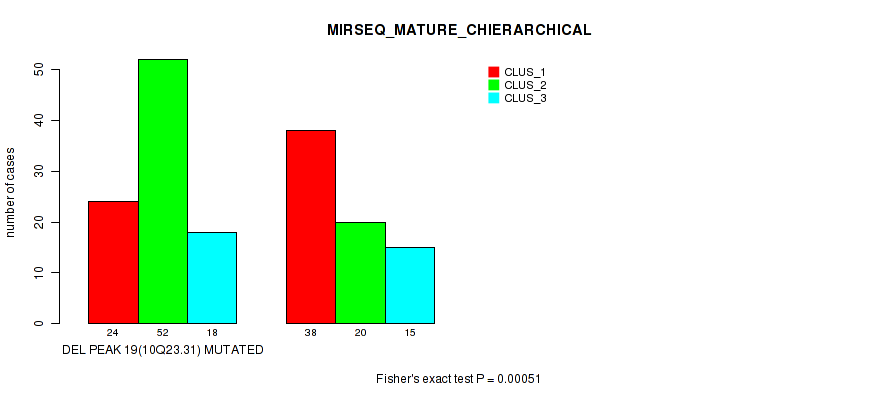
P value = 5e-05 (Fisher's exact test), Q value = 0.022
Table S69. Gene #41: 'del_11p15.5' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 20(11P15.5) MUTATED | 21 | 40 | 12 |
| DEL PEAK 20(11P15.5) WILD-TYPE | 25 | 26 | 45 |
Figure S69. Get High-res Image Gene #41: 'del_11p15.5' versus Molecular Subtype #1: 'CN_CNMF'
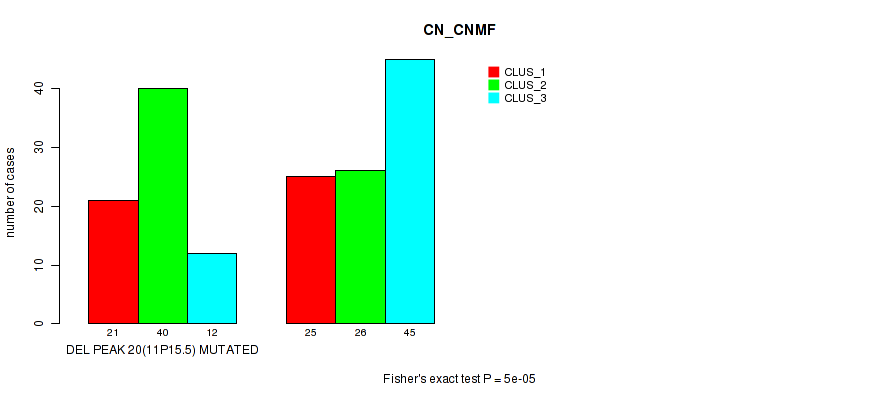
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S70. Gene #42: 'del_11q24.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 21(11Q24.3) MUTATED | 27 | 41 | 7 |
| DEL PEAK 21(11Q24.3) WILD-TYPE | 19 | 25 | 50 |
Figure S70. Get High-res Image Gene #42: 'del_11q24.3' versus Molecular Subtype #1: 'CN_CNMF'
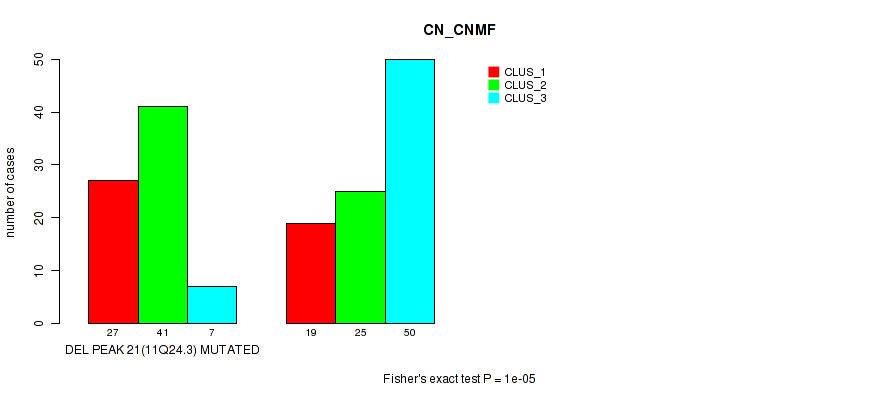
P value = 2e-05 (Fisher's exact test), Q value = 0.0089
Table S71. Gene #42: 'del_11q24.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 21(11Q24.3) MUTATED | 24 | 9 | 13 | 8 | 2 | 17 | 2 |
| DEL PEAK 21(11Q24.3) WILD-TYPE | 6 | 19 | 29 | 13 | 14 | 11 | 2 |
Figure S71. Get High-res Image Gene #42: 'del_11q24.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
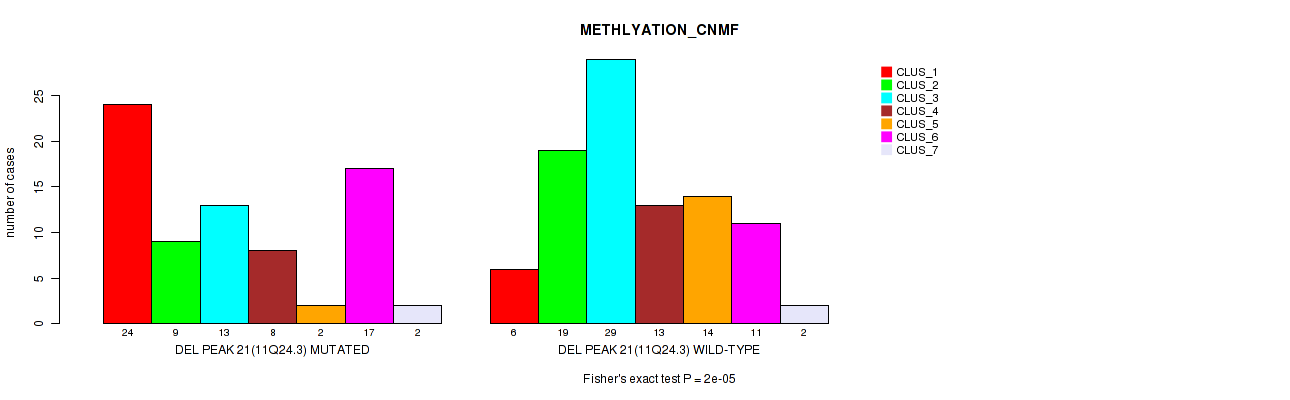
P value = 0.00045 (Fisher's exact test), Q value = 0.18
Table S72. Gene #43: 'del_12p13.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 22(12P13.1) MUTATED | 18 | 5 | 13 | 3 | 2 | 15 | 1 |
| DEL PEAK 22(12P13.1) WILD-TYPE | 12 | 23 | 29 | 18 | 14 | 13 | 3 |
Figure S72. Get High-res Image Gene #43: 'del_12p13.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
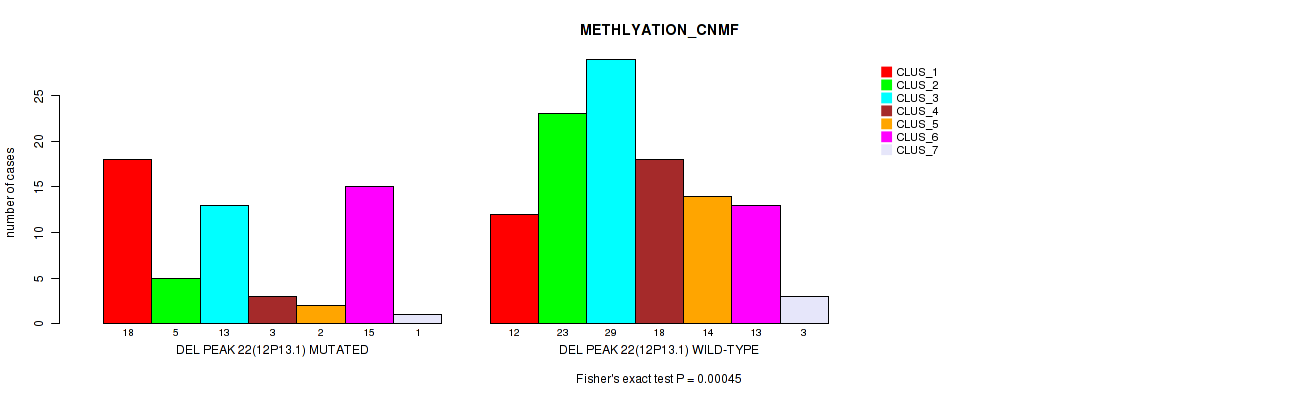
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S73. Gene #45: 'del_13q14.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 24(13Q14.2) MUTATED | 23 | 63 | 35 |
| DEL PEAK 24(13Q14.2) WILD-TYPE | 23 | 3 | 22 |
Figure S73. Get High-res Image Gene #45: 'del_13q14.2' versus Molecular Subtype #1: 'CN_CNMF'
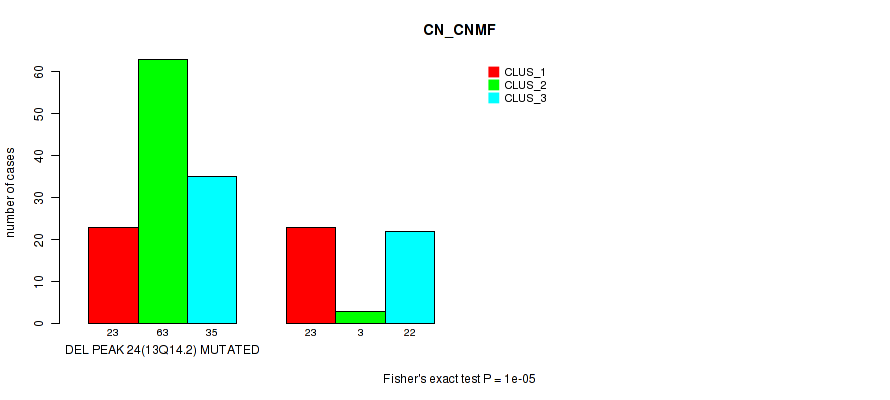
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S74. Gene #45: 'del_13q14.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 24(13Q14.2) MUTATED | 24 | 11 | 41 | 9 | 7 | 25 | 4 |
| DEL PEAK 24(13Q14.2) WILD-TYPE | 6 | 17 | 1 | 12 | 9 | 3 | 0 |
Figure S74. Get High-res Image Gene #45: 'del_13q14.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
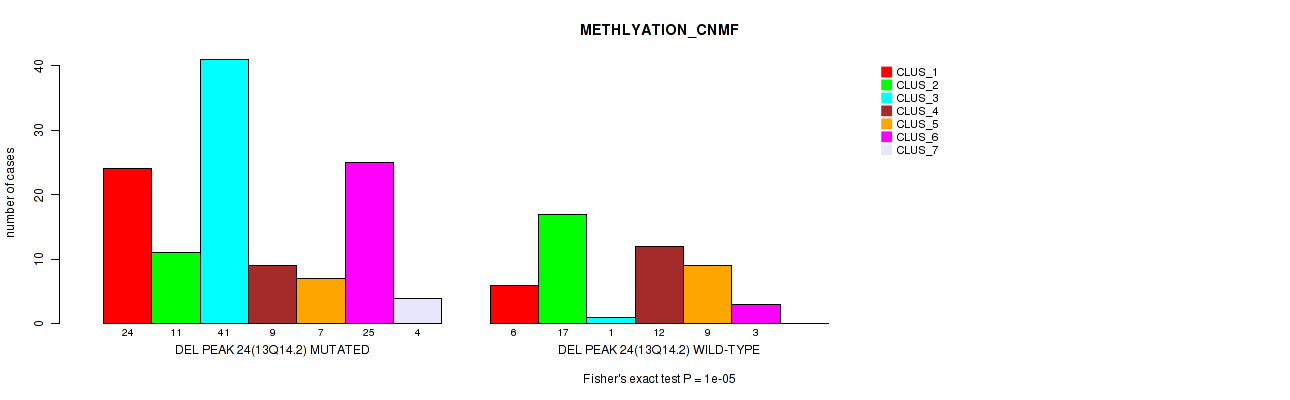
P value = 1e-04 (Fisher's exact test), Q value = 0.042
Table S75. Gene #45: 'del_13q14.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| DEL PEAK 24(13Q14.2) MUTATED | 31 | 24 | 12 | 51 |
| DEL PEAK 24(13Q14.2) WILD-TYPE | 8 | 21 | 12 | 7 |
Figure S75. Get High-res Image Gene #45: 'del_13q14.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
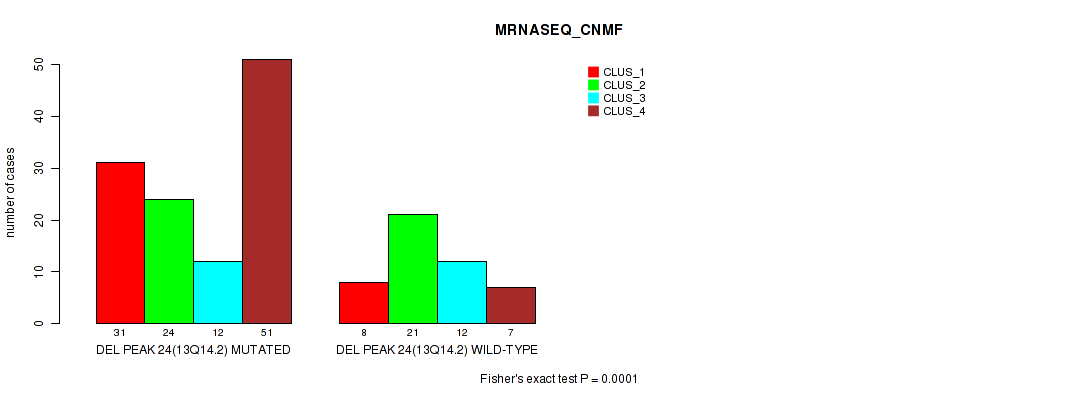
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S76. Gene #45: 'del_13q14.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 45 | 32 | 33 | 37 | 19 |
| DEL PEAK 24(13Q14.2) MUTATED | 27 | 13 | 26 | 36 | 16 |
| DEL PEAK 24(13Q14.2) WILD-TYPE | 18 | 19 | 7 | 1 | 3 |
Figure S76. Get High-res Image Gene #45: 'del_13q14.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
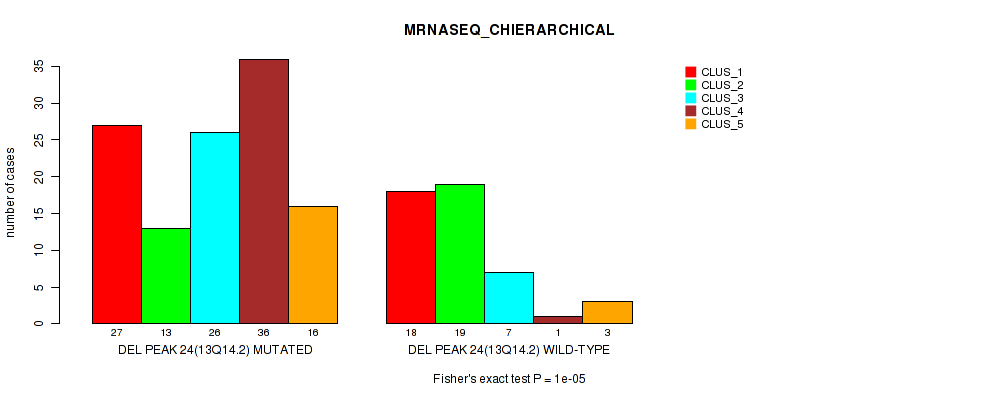
P value = 0.00022 (Fisher's exact test), Q value = 0.09
Table S77. Gene #46: 'del_14q24.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 45 | 32 | 33 | 37 | 19 |
| DEL PEAK 25(14Q24.1) MUTATED | 17 | 7 | 9 | 8 | 15 |
| DEL PEAK 25(14Q24.1) WILD-TYPE | 28 | 25 | 24 | 29 | 4 |
Figure S77. Get High-res Image Gene #46: 'del_14q24.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
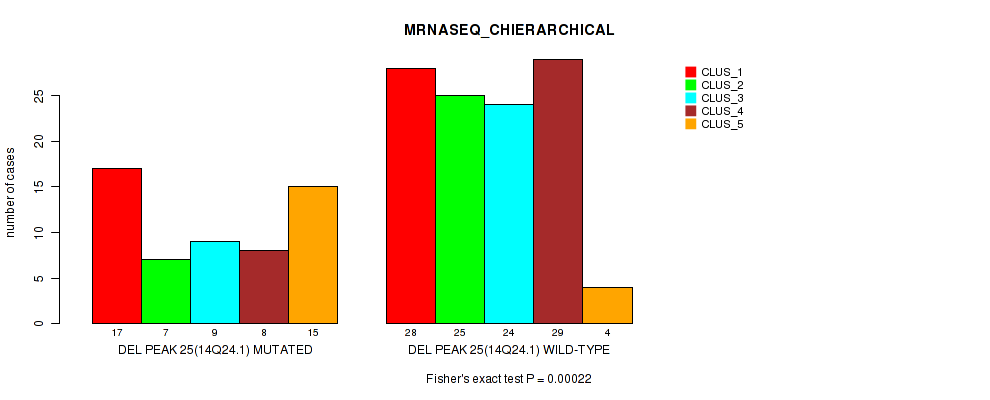
P value = 0.00022 (Fisher's exact test), Q value = 0.09
Table S78. Gene #47: 'del_15q11.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| DEL PEAK 26(15Q11.2) MUTATED | 17 | 6 | 3 | 5 |
| DEL PEAK 26(15Q11.2) WILD-TYPE | 22 | 39 | 21 | 53 |
Figure S78. Get High-res Image Gene #47: 'del_15q11.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
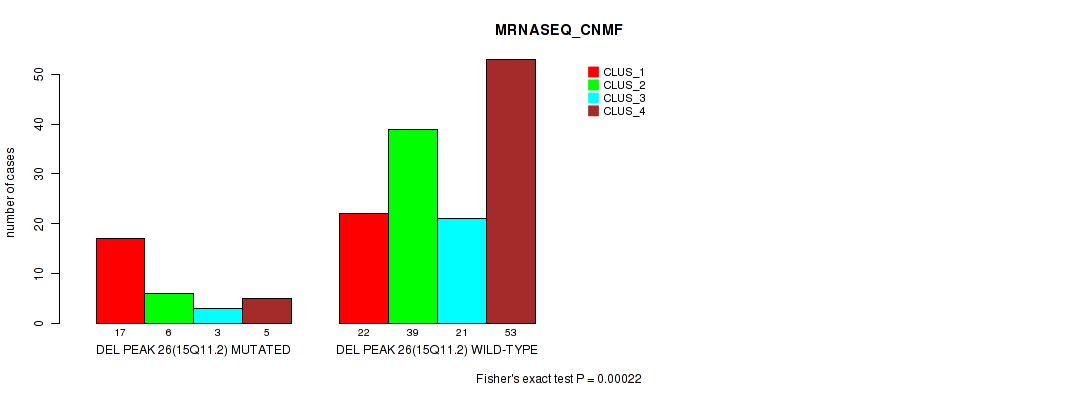
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S79. Gene #48: 'del_16q12.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 27(16Q12.1) MUTATED | 13 | 52 | 30 |
| DEL PEAK 27(16Q12.1) WILD-TYPE | 33 | 14 | 27 |
Figure S79. Get High-res Image Gene #48: 'del_16q12.1' versus Molecular Subtype #1: 'CN_CNMF'
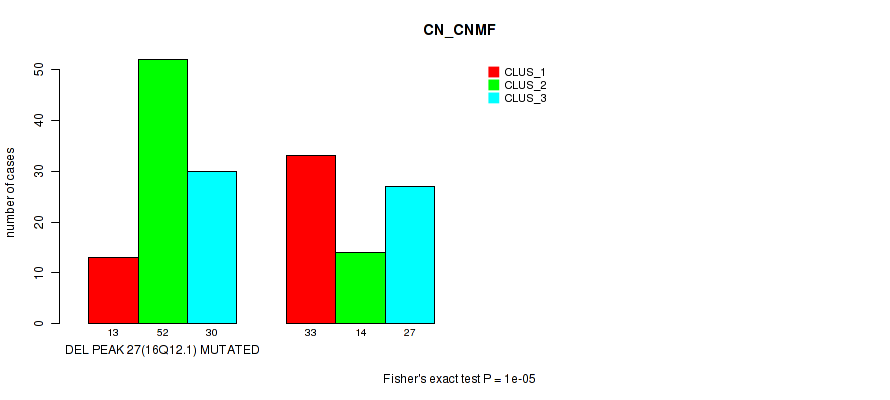
P value = 4e-05 (Fisher's exact test), Q value = 0.017
Table S80. Gene #49: 'del_16q23.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 28(16Q23.1) MUTATED | 16 | 50 | 28 |
| DEL PEAK 28(16Q23.1) WILD-TYPE | 30 | 16 | 29 |
Figure S80. Get High-res Image Gene #49: 'del_16q23.1' versus Molecular Subtype #1: 'CN_CNMF'
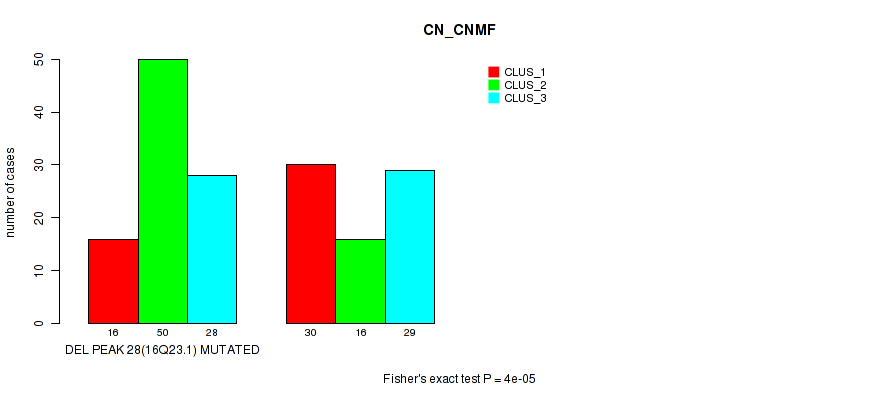
P value = 3e-05 (Fisher's exact test), Q value = 0.013
Table S81. Gene #50: 'del_17p13.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 29(17P13.1) MUTATED | 9 | 41 | 32 |
| DEL PEAK 29(17P13.1) WILD-TYPE | 37 | 25 | 25 |
Figure S81. Get High-res Image Gene #50: 'del_17p13.1' versus Molecular Subtype #1: 'CN_CNMF'
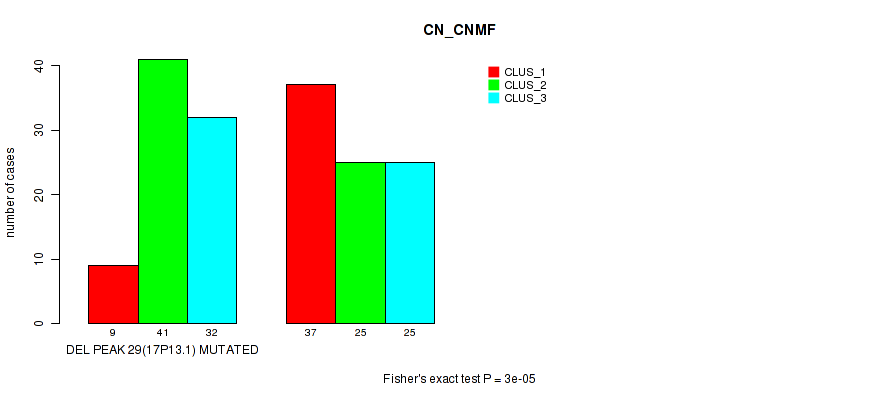
P value = 1e-04 (Fisher's exact test), Q value = 0.042
Table S82. Gene #50: 'del_17p13.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 29(17P13.1) MUTATED | 9 | 7 | 33 | 10 | 6 | 16 | 1 |
| DEL PEAK 29(17P13.1) WILD-TYPE | 21 | 21 | 9 | 11 | 10 | 12 | 3 |
Figure S82. Get High-res Image Gene #50: 'del_17p13.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
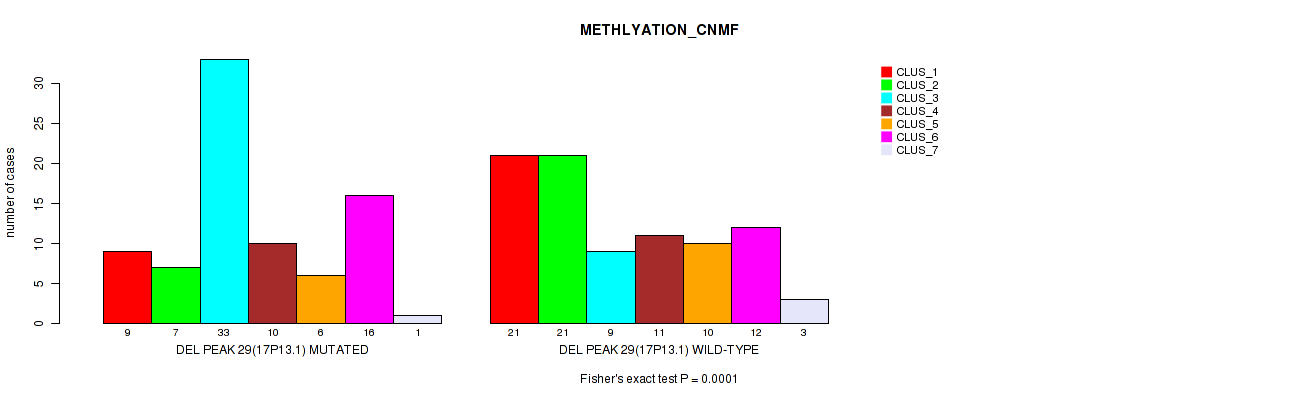
P value = 0.00013 (Fisher's exact test), Q value = 0.055
Table S83. Gene #50: 'del_17p13.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 39 | 45 | 24 | 58 |
| DEL PEAK 29(17P13.1) MUTATED | 21 | 11 | 9 | 39 |
| DEL PEAK 29(17P13.1) WILD-TYPE | 18 | 34 | 15 | 19 |
Figure S83. Get High-res Image Gene #50: 'del_17p13.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
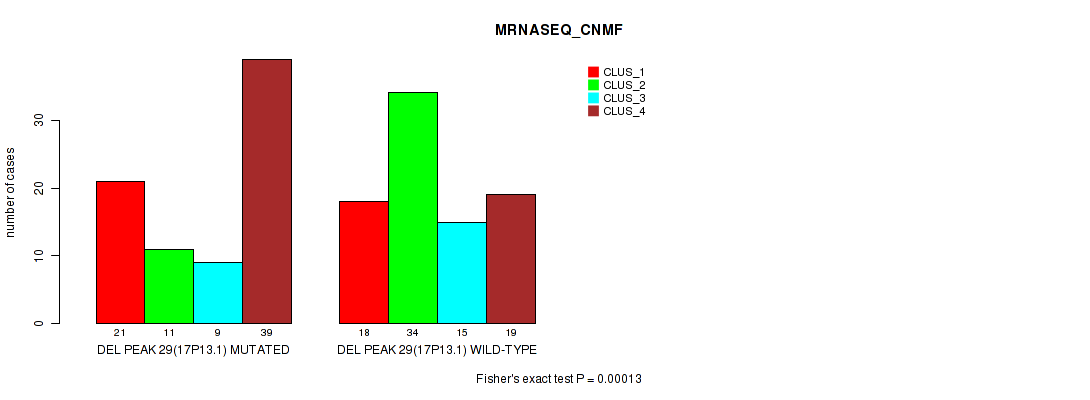
P value = 0.00057 (Fisher's exact test), Q value = 0.23
Table S84. Gene #50: 'del_17p13.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 45 | 32 | 33 | 37 | 19 |
| DEL PEAK 29(17P13.1) MUTATED | 15 | 12 | 16 | 29 | 8 |
| DEL PEAK 29(17P13.1) WILD-TYPE | 30 | 20 | 17 | 8 | 11 |
Figure S84. Get High-res Image Gene #50: 'del_17p13.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
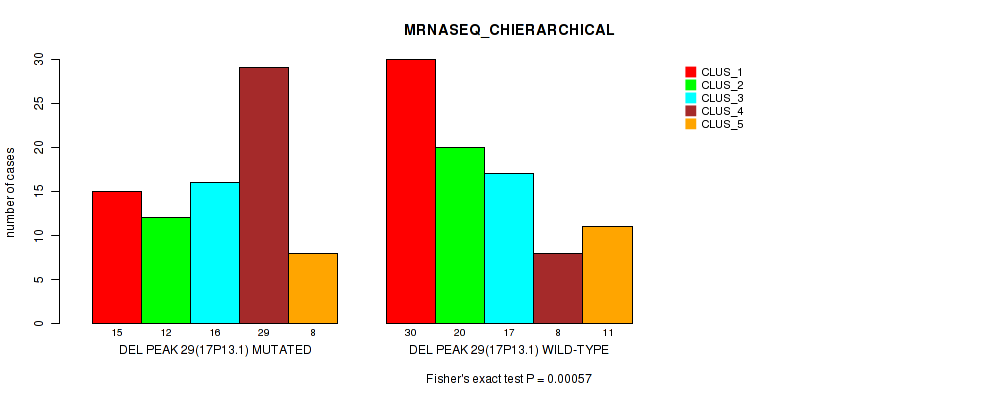
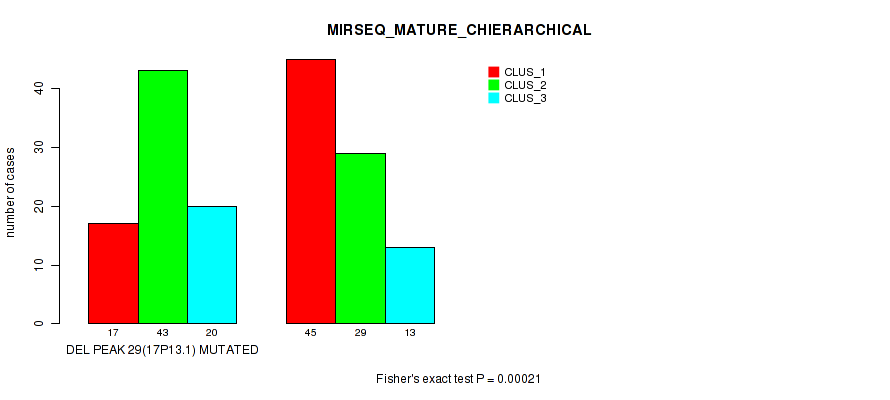
P value = 0.00021 (Fisher's exact test), Q value = 0.087
Table S85. Gene #50: 'del_17p13.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 62 | 72 | 33 |
| DEL PEAK 29(17P13.1) MUTATED | 17 | 43 | 20 |
| DEL PEAK 29(17P13.1) WILD-TYPE | 45 | 29 | 13 |
Figure S85. Get High-res Image Gene #50: 'del_17p13.1' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
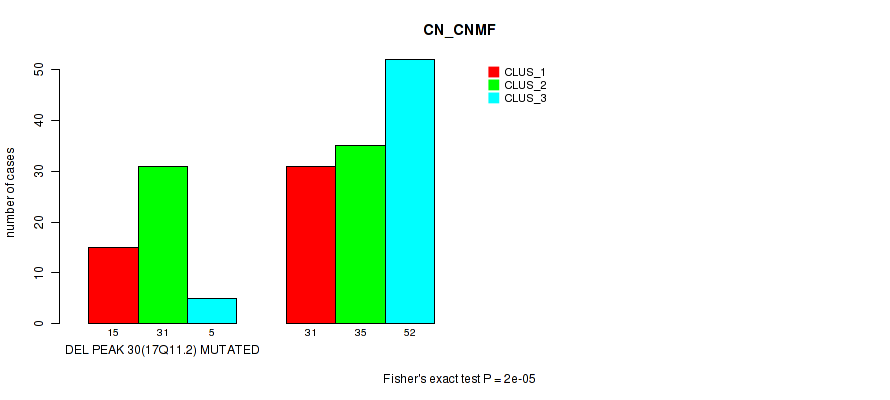
P value = 2e-05 (Fisher's exact test), Q value = 0.0089
Table S86. Gene #51: 'del_17q11.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 30(17Q11.2) MUTATED | 15 | 31 | 5 |
| DEL PEAK 30(17Q11.2) WILD-TYPE | 31 | 35 | 52 |
Figure S86. Get High-res Image Gene #51: 'del_17q11.2' versus Molecular Subtype #1: 'CN_CNMF'
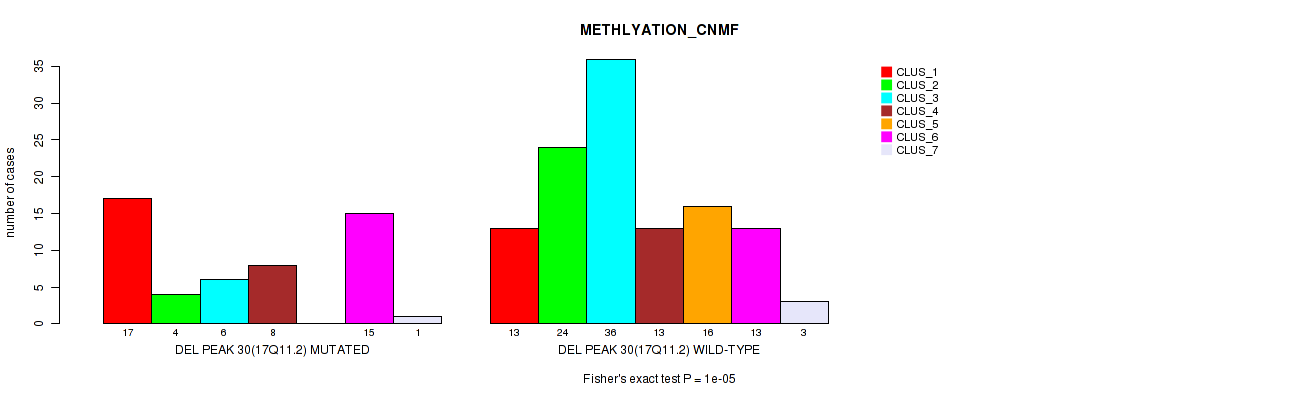
P value = 1e-05 (Fisher's exact test), Q value = 0.0049
Table S87. Gene #51: 'del_17q11.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 30(17Q11.2) MUTATED | 17 | 4 | 6 | 8 | 0 | 15 | 1 |
| DEL PEAK 30(17Q11.2) WILD-TYPE | 13 | 24 | 36 | 13 | 16 | 13 | 3 |
Figure S87. Get High-res Image Gene #51: 'del_17q11.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
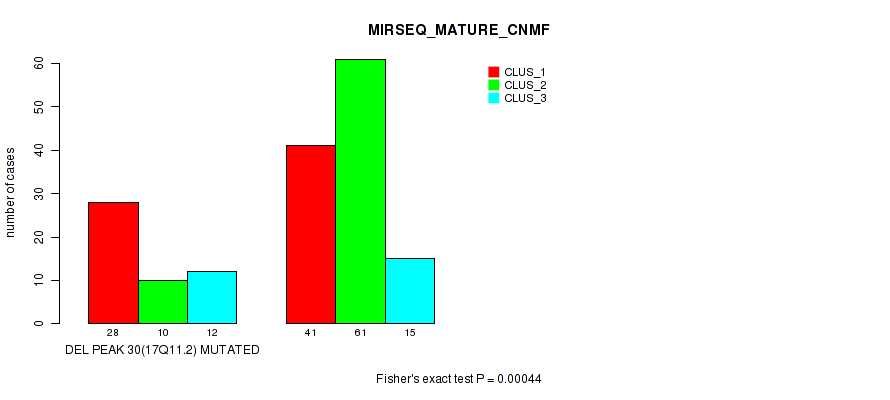
P value = 0.00044 (Fisher's exact test), Q value = 0.18
Table S88. Gene #51: 'del_17q11.2' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 69 | 71 | 27 |
| DEL PEAK 30(17Q11.2) MUTATED | 28 | 10 | 12 |
| DEL PEAK 30(17Q11.2) WILD-TYPE | 41 | 61 | 15 |
Figure S88. Get High-res Image Gene #51: 'del_17q11.2' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
P value = 0.00062 (Fisher's exact test), Q value = 0.24
Table S89. Gene #52: 'del_17q25.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 31(17Q25.3) MUTATED | 10 | 28 | 7 |
| DEL PEAK 31(17Q25.3) WILD-TYPE | 36 | 38 | 50 |
Figure S89. Get High-res Image Gene #52: 'del_17q25.3' versus Molecular Subtype #1: 'CN_CNMF'
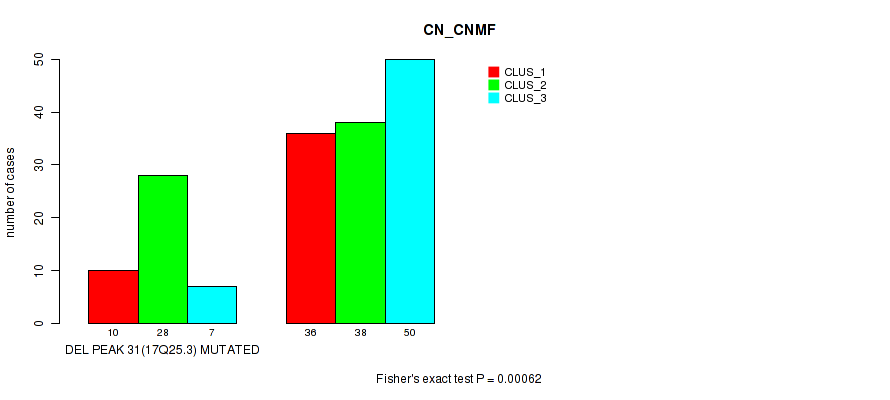
P value = 2e-05 (Fisher's exact test), Q value = 0.0089
Table S90. Gene #54: 'del_19p13.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 33(19P13.3) MUTATED | 6 | 32 | 12 |
| DEL PEAK 33(19P13.3) WILD-TYPE | 40 | 34 | 45 |
Figure S90. Get High-res Image Gene #54: 'del_19p13.3' versus Molecular Subtype #1: 'CN_CNMF'
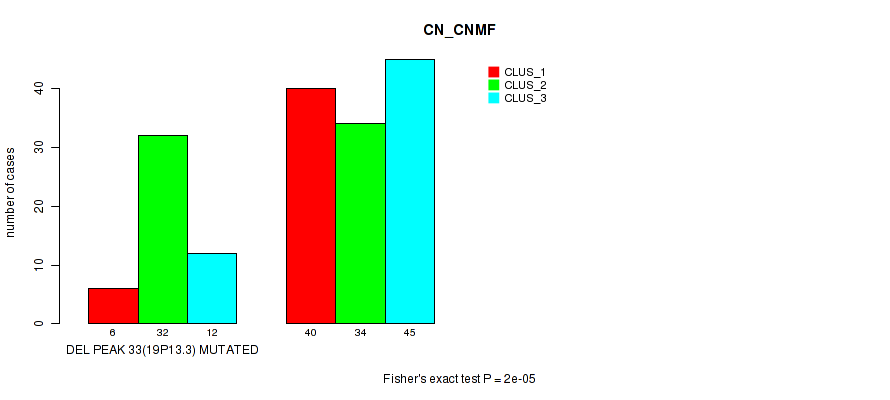
P value = 0.00012 (Fisher's exact test), Q value = 0.051
Table S91. Gene #54: 'del_19p13.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 30 | 28 | 42 | 21 | 16 | 28 | 4 |
| DEL PEAK 33(19P13.3) MUTATED | 5 | 1 | 17 | 5 | 5 | 14 | 3 |
| DEL PEAK 33(19P13.3) WILD-TYPE | 25 | 27 | 25 | 16 | 11 | 14 | 1 |
Figure S91. Get High-res Image Gene #54: 'del_19p13.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
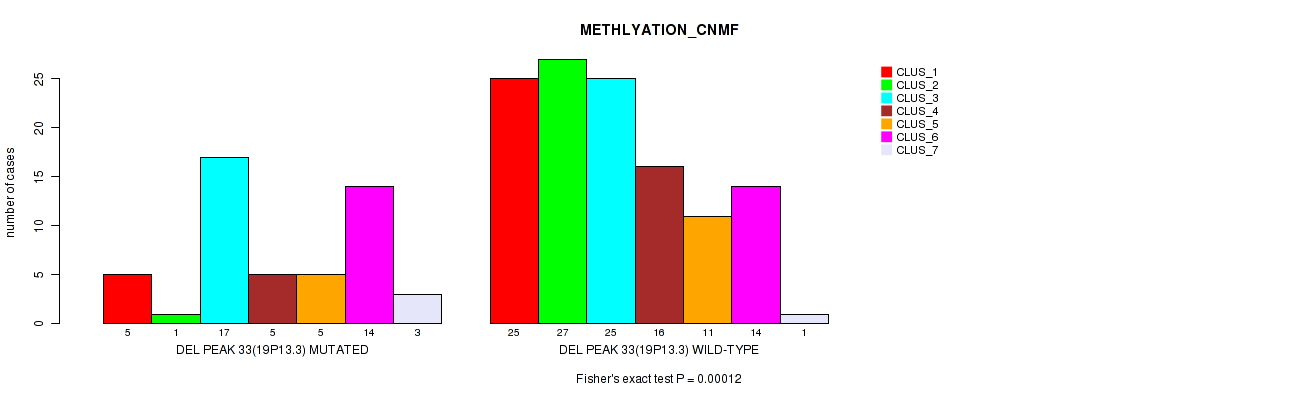
P value = 0.00042 (Fisher's exact test), Q value = 0.17
Table S92. Gene #55: 'del_19q13.43' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 34(19Q13.43) MUTATED | 11 | 40 | 22 |
| DEL PEAK 34(19Q13.43) WILD-TYPE | 35 | 26 | 35 |
Figure S92. Get High-res Image Gene #55: 'del_19q13.43' versus Molecular Subtype #1: 'CN_CNMF'
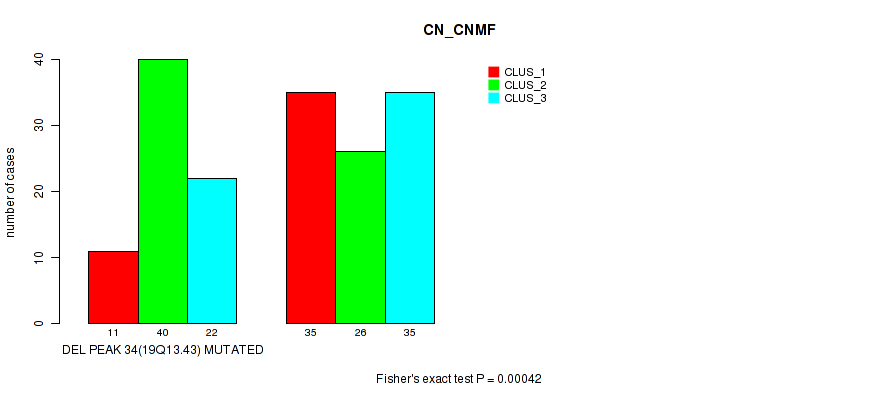
P value = 7e-05 (Fisher's exact test), Q value = 0.03
Table S93. Gene #58: 'del_22q13.31' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 37(22Q13.31) MUTATED | 15 | 32 | 7 |
| DEL PEAK 37(22Q13.31) WILD-TYPE | 31 | 34 | 50 |
Figure S93. Get High-res Image Gene #58: 'del_22q13.31' versus Molecular Subtype #1: 'CN_CNMF'
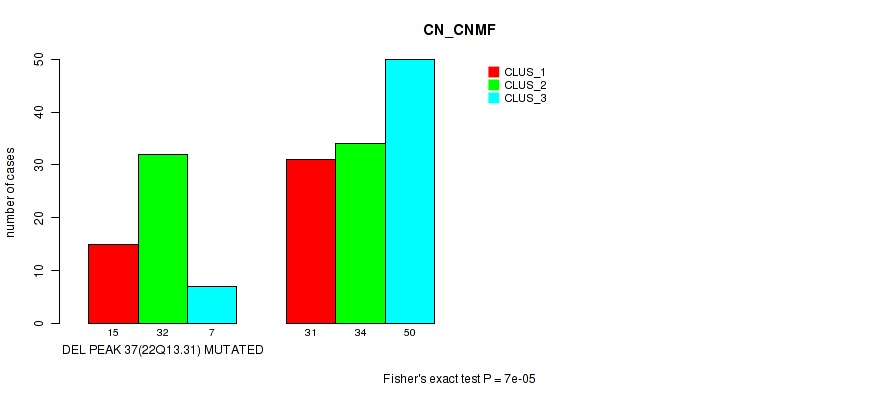
P value = 3e-05 (Fisher's exact test), Q value = 0.013
Table S94. Gene #61: 'del_xq27.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 46 | 66 | 57 |
| DEL PEAK 40(XQ27.1) MUTATED | 13 | 46 | 23 |
| DEL PEAK 40(XQ27.1) WILD-TYPE | 33 | 20 | 34 |
Figure S94. Get High-res Image Gene #61: 'del_xq27.1' versus Molecular Subtype #1: 'CN_CNMF'
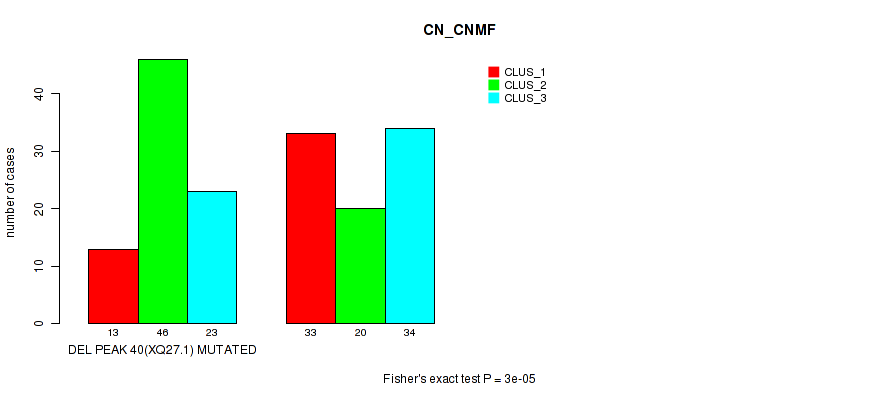
-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtype file = SARC-TP.transferedmergedcluster.txt
-
Number of patients = 169
-
Number of significantly focal cnvs = 61
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.