This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 20 focal events and 6 molecular subtypes across 191 patients, 36 significant findings detected with P value < 0.05 and Q value < 0.25.
-
amp_1p33 cnv correlated to 'CN_CNMF'.
-
amp_11q23.3 cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
amp_19p13.2 cnv correlated to 'CN_CNMF'.
-
amp_21q22.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MIRSEQ_CNMF'.
-
del_3p13 cnv correlated to 'CN_CNMF'.
-
del_5q31.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_7p12.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_7q34 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_7q34 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_12p13.2 cnv correlated to 'CN_CNMF'.
-
del_12q21.33 cnv correlated to 'CN_CNMF'.
-
del_16q23.1 cnv correlated to 'CN_CNMF'.
-
del_17p13.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
del_17q11.2 cnv correlated to 'CN_CNMF'.
-
del_18p11.21 cnv correlated to 'CN_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 20 focal events and 6 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 36 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| del 5q31 2 | 18 (9%) | 173 |
1e-05 (0.00116) |
9e-05 (0.00909) |
0.00534 (0.406) |
0.00084 (0.0764) |
0.00077 (0.0716) |
0.0013 (0.112) |
| del 7q34 | 24 (13%) | 167 |
1e-05 (0.00116) |
6e-05 (0.00618) |
0.0215 (1.00) |
1e-05 (0.00116) |
0.0008 (0.0736) |
0.00055 (0.0533) |
| del 7q34 | 24 (13%) | 167 |
1e-05 (0.00116) |
6e-05 (0.00618) |
0.0216 (1.00) |
2e-05 (0.00216) |
0.00092 (0.0819) |
0.00056 (0.0538) |
| del 7p12 1 | 16 (8%) | 175 |
0.00019 (0.019) |
0.00062 (0.0589) |
0.00186 (0.154) |
5e-05 (0.0052) |
0.00614 (0.454) |
0.00639 (0.46) |
| del 17p13 2 | 15 (8%) | 176 |
1e-05 (0.00116) |
0.00085 (0.0765) |
0.0114 (0.767) |
0.0002 (0.0198) |
0.0865 (1.00) |
0.00116 (0.102) |
| amp 21q22 2 | 14 (7%) | 177 |
1e-05 (0.00116) |
0.00214 (0.175) |
0.00327 (0.262) |
0.0361 (1.00) |
0.00167 (0.14) |
0.0673 (1.00) |
| amp 11q23 3 | 17 (9%) | 174 |
1e-05 (0.00116) |
2e-05 (0.00216) |
0.373 (1.00) |
0.201 (1.00) |
0.746 (1.00) |
0.187 (1.00) |
| amp 1p33 | 7 (4%) | 184 |
0.00049 (0.048) |
0.0116 (0.767) |
0.125 (1.00) |
0.13 (1.00) |
0.00951 (0.656) |
0.0492 (1.00) |
| amp 19p13 2 | 6 (3%) | 185 |
0.0029 (0.235) |
0.555 (1.00) |
0.857 (1.00) |
0.632 (1.00) |
0.895 (1.00) |
1 (1.00) |
| del 3p13 | 8 (4%) | 183 |
0.00072 (0.0677) |
0.297 (1.00) |
0.0676 (1.00) |
0.236 (1.00) |
0.0365 (1.00) |
0.0478 (1.00) |
| del 12p13 2 | 10 (5%) | 181 |
2e-05 (0.00216) |
0.00519 (0.4) |
0.199 (1.00) |
0.0723 (1.00) |
0.0697 (1.00) |
0.00959 (0.656) |
| del 12q21 33 | 3 (2%) | 188 |
0.00135 (0.115) |
0.304 (1.00) |
0.187 (1.00) |
0.587 (1.00) |
0.00843 (0.59) |
0.006 (0.45) |
| del 16q23 1 | 9 (5%) | 182 |
0.00121 (0.105) |
0.149 (1.00) |
0.33 (1.00) |
0.494 (1.00) |
0.043 (1.00) |
0.0657 (1.00) |
| del 17q11 2 | 13 (7%) | 178 |
1e-05 (0.00116) |
0.0224 (1.00) |
0.156 (1.00) |
0.00661 (0.469) |
0.136 (1.00) |
0.0035 (0.273) |
| del 18p11 21 | 9 (5%) | 182 |
2e-05 (0.00216) |
0.0178 (1.00) |
0.382 (1.00) |
0.335 (1.00) |
0.213 (1.00) |
0.0311 (1.00) |
| amp 1q43 | 7 (4%) | 184 |
0.00338 (0.267) |
0.0124 (0.808) |
0.377 (1.00) |
0.0364 (1.00) |
0.137 (1.00) |
0.015 (0.944) |
| amp 13q31 3 | 7 (4%) | 184 |
0.0807 (1.00) |
0.0124 (0.808) |
0.067 (1.00) |
0.0364 (1.00) |
0.737 (1.00) |
0.0566 (1.00) |
| amp 20q11 21 | 3 (2%) | 188 |
0.00618 (0.454) |
0.267 (1.00) |
||||
| del 9q21 32 | 5 (3%) | 186 |
0.236 (1.00) |
0.516 (1.00) |
0.176 (1.00) |
0.602 (1.00) |
0.182 (1.00) |
0.105 (1.00) |
| del 20q13 13 | 4 (2%) | 187 |
0.0288 (1.00) |
0.148 (1.00) |
0.787 (1.00) |
0.588 (1.00) |
0.126 (1.00) |
0.411 (1.00) |
P value = 0.00049 (Fisher's exact test), Q value = 0.048
Table S1. Gene #1: 'amp_1p33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| AMP PEAK 1(1P33) MUTATED | 1 | 3 | 3 | 0 |
| AMP PEAK 1(1P33) WILD-TYPE | 147 | 11 | 24 | 2 |
Figure S1. Get High-res Image Gene #1: 'amp_1p33' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 0.0012
Table S2. Gene #3: 'amp_11q23.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| AMP PEAK 3(11Q23.3) MUTATED | 0 | 14 | 3 | 0 |
| AMP PEAK 3(11Q23.3) WILD-TYPE | 148 | 0 | 24 | 2 |
Figure S2. Get High-res Image Gene #3: 'amp_11q23.3' versus Molecular Subtype #1: 'CN_CNMF'

P value = 2e-05 (Fisher's exact test), Q value = 0.0022
Table S3. Gene #3: 'amp_11q23.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 46 | 44 | 61 | 14 | 20 |
| AMP PEAK 3(11Q23.3) MUTATED | 0 | 0 | 12 | 4 | 1 |
| AMP PEAK 3(11Q23.3) WILD-TYPE | 46 | 44 | 49 | 10 | 19 |
Figure S3. Get High-res Image Gene #3: 'amp_11q23.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 0.0029 (Fisher's exact test), Q value = 0.23
Table S4. Gene #5: 'amp_19p13.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| AMP PEAK 5(19P13.2) MUTATED | 1 | 1 | 4 | 0 |
| AMP PEAK 5(19P13.2) WILD-TYPE | 147 | 13 | 23 | 2 |
Figure S4. Get High-res Image Gene #5: 'amp_19p13.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 0.0012
Table S5. Gene #7: 'amp_21q22.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| AMP PEAK 7(21Q22.2) MUTATED | 0 | 0 | 14 | 0 |
| AMP PEAK 7(21Q22.2) WILD-TYPE | 148 | 14 | 13 | 2 |
Figure S5. Get High-res Image Gene #7: 'amp_21q22.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00214 (Fisher's exact test), Q value = 0.18
Table S6. Gene #7: 'amp_21q22.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 46 | 44 | 61 | 14 | 20 |
| AMP PEAK 7(21Q22.2) MUTATED | 1 | 0 | 11 | 0 | 1 |
| AMP PEAK 7(21Q22.2) WILD-TYPE | 45 | 44 | 50 | 14 | 19 |
Figure S6. Get High-res Image Gene #7: 'amp_21q22.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'

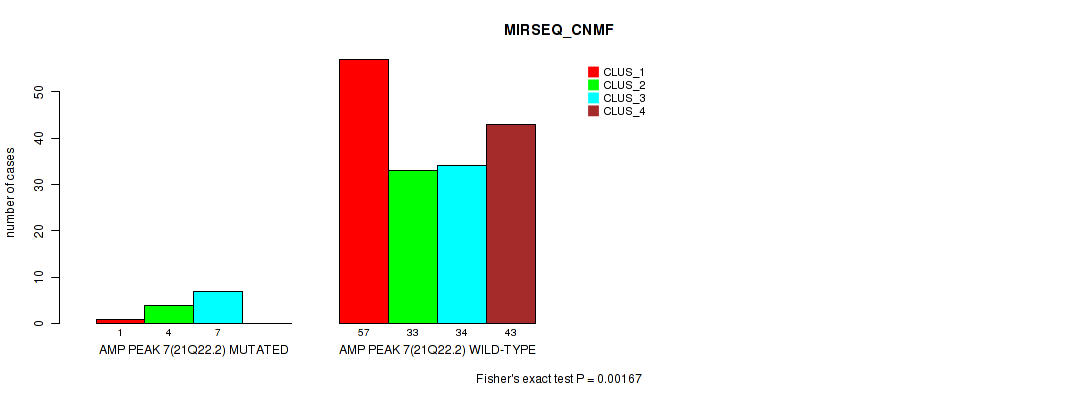
P value = 0.00167 (Fisher's exact test), Q value = 0.14
Table S7. Gene #7: 'amp_21q22.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| AMP PEAK 7(21Q22.2) MUTATED | 1 | 4 | 7 | 0 |
| AMP PEAK 7(21Q22.2) WILD-TYPE | 57 | 33 | 34 | 43 |
Figure S7. Get High-res Image Gene #7: 'amp_21q22.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
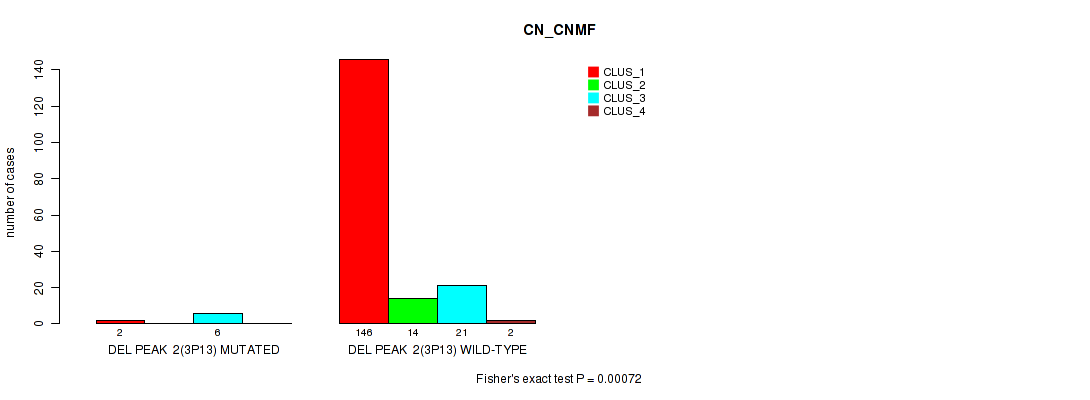
P value = 0.00072 (Fisher's exact test), Q value = 0.068
Table S8. Gene #8: 'del_3p13' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 2(3P13) MUTATED | 2 | 0 | 6 | 0 |
| DEL PEAK 2(3P13) WILD-TYPE | 146 | 14 | 21 | 2 |
Figure S8. Get High-res Image Gene #8: 'del_3p13' versus Molecular Subtype #1: 'CN_CNMF'
P value = 1e-05 (Fisher's exact test), Q value = 0.0012
Table S9. Gene #9: 'del_5q31.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 3(5Q31.2) MUTATED | 1 | 1 | 15 | 1 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 147 | 13 | 12 | 1 |
Figure S9. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 9e-05 (Fisher's exact test), Q value = 0.0091
Table S10. Gene #9: 'del_5q31.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 46 | 44 | 61 | 14 | 20 |
| DEL PEAK 3(5Q31.2) MUTATED | 1 | 0 | 15 | 0 | 2 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 45 | 44 | 46 | 14 | 18 |
Figure S10. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'

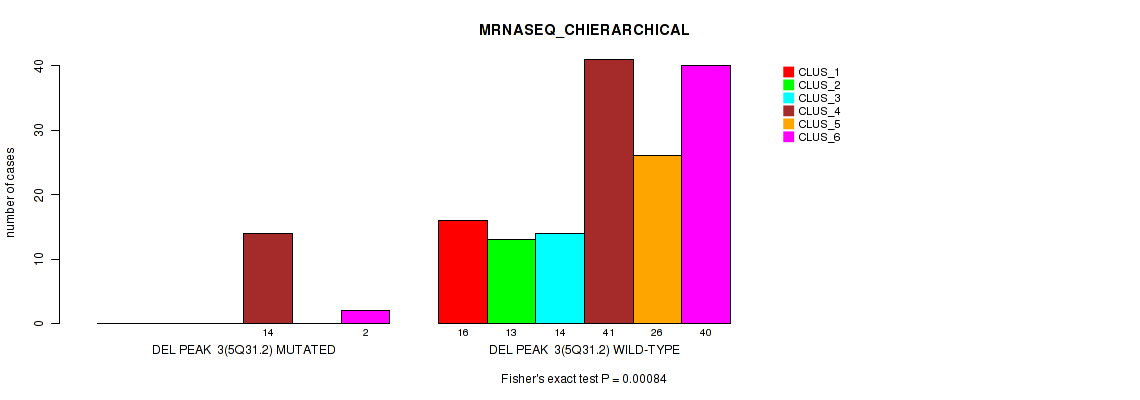
P value = 0.00084 (Fisher's exact test), Q value = 0.076
Table S11. Gene #9: 'del_5q31.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 3(5Q31.2) MUTATED | 0 | 0 | 0 | 14 | 0 | 2 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 16 | 13 | 14 | 41 | 26 | 40 |
Figure S11. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
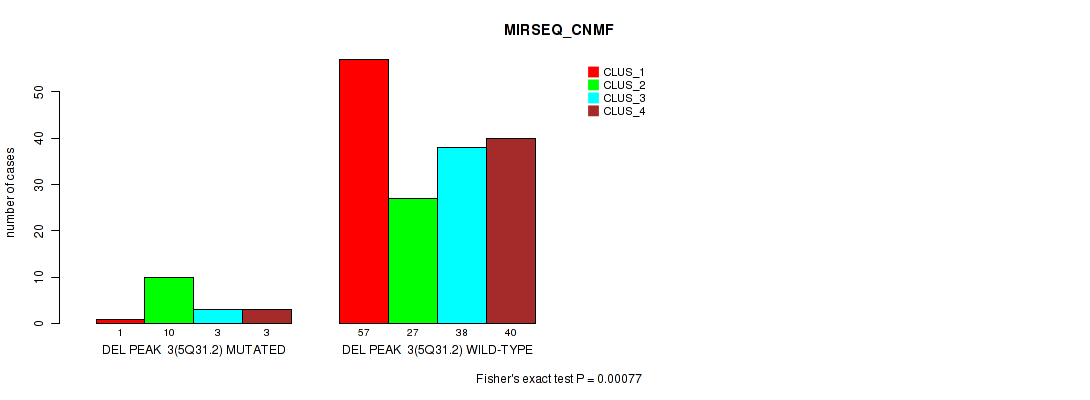
P value = 0.00077 (Fisher's exact test), Q value = 0.072
Table S12. Gene #9: 'del_5q31.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| DEL PEAK 3(5Q31.2) MUTATED | 1 | 10 | 3 | 3 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 57 | 27 | 38 | 40 |
Figure S12. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
P value = 0.0013 (Fisher's exact test), Q value = 0.11
Table S13. Gene #9: 'del_5q31.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 3(5Q31.2) MUTATED | 1 | 8 | 8 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 66 | 26 | 70 |
Figure S13. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 0.00019 (Fisher's exact test), Q value = 0.019
Table S14. Gene #10: 'del_7p12.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 4(7P12.1) MUTATED | 7 | 0 | 8 | 1 |
| DEL PEAK 4(7P12.1) WILD-TYPE | 141 | 14 | 19 | 1 |
Figure S14. Get High-res Image Gene #10: 'del_7p12.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00062 (Fisher's exact test), Q value = 0.059
Table S15. Gene #10: 'del_7p12.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 46 | 44 | 61 | 14 | 20 |
| DEL PEAK 4(7P12.1) MUTATED | 1 | 0 | 13 | 1 | 1 |
| DEL PEAK 4(7P12.1) WILD-TYPE | 45 | 44 | 48 | 13 | 19 |
Figure S15. Get High-res Image Gene #10: 'del_7p12.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
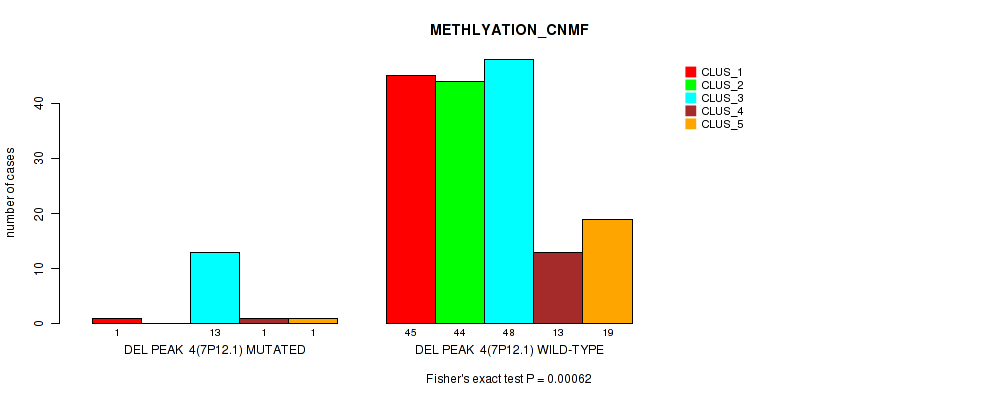
P value = 0.00186 (Fisher's exact test), Q value = 0.15
Table S16. Gene #10: 'del_7p12.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 52 | 44 |
| DEL PEAK 4(7P12.1) MUTATED | 12 | 0 | 3 |
| DEL PEAK 4(7P12.1) WILD-TYPE | 58 | 52 | 41 |
Figure S16. Get High-res Image Gene #10: 'del_7p12.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'

P value = 5e-05 (Fisher's exact test), Q value = 0.0052
Table S17. Gene #10: 'del_7p12.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 4(7P12.1) MUTATED | 0 | 0 | 0 | 15 | 0 | 0 |
| DEL PEAK 4(7P12.1) WILD-TYPE | 16 | 13 | 14 | 40 | 26 | 42 |
Figure S17. Get High-res Image Gene #10: 'del_7p12.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
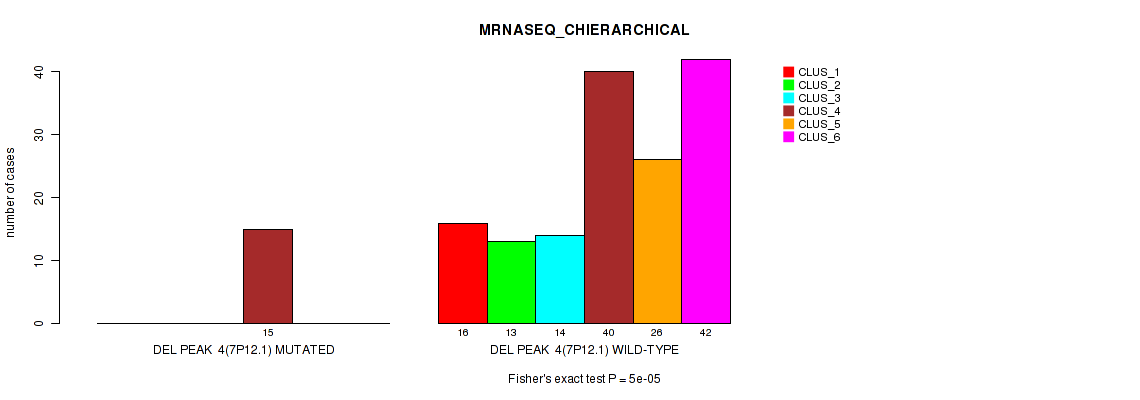
P value = 1e-05 (Fisher's exact test), Q value = 0.0012
Table S18. Gene #11: 'del_7q34' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 5(7Q34) MUTATED | 9 | 2 | 11 | 2 |
| DEL PEAK 5(7Q34) WILD-TYPE | 139 | 12 | 16 | 0 |
Figure S18. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #1: 'CN_CNMF'

P value = 6e-05 (Fisher's exact test), Q value = 0.0062
Table S19. Gene #11: 'del_7q34' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 46 | 44 | 61 | 14 | 20 |
| DEL PEAK 5(7Q34) MUTATED | 1 | 1 | 18 | 2 | 2 |
| DEL PEAK 5(7Q34) WILD-TYPE | 45 | 43 | 43 | 12 | 18 |
Figure S19. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 0.0012
Table S20. Gene #11: 'del_7q34' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 5(7Q34) MUTATED | 1 | 1 | 0 | 18 | 0 | 0 |
| DEL PEAK 5(7Q34) WILD-TYPE | 15 | 12 | 14 | 37 | 26 | 42 |
Figure S20. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
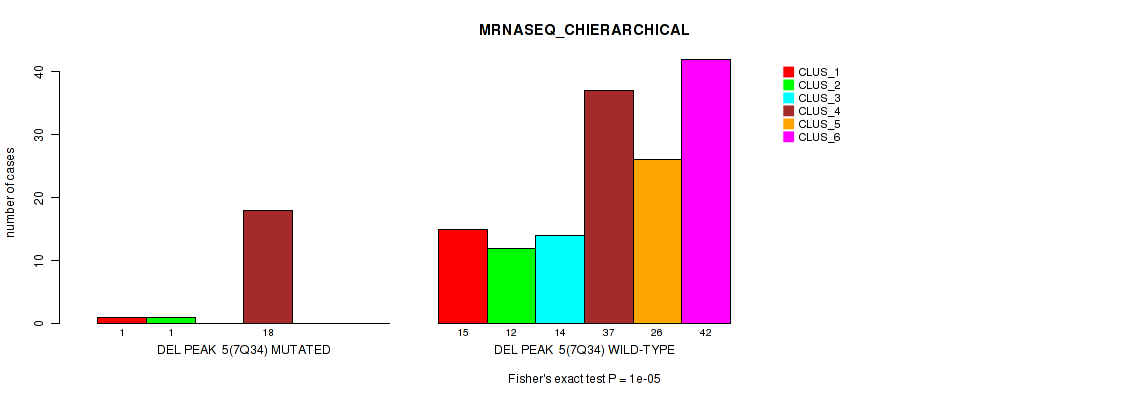
P value = 8e-04 (Fisher's exact test), Q value = 0.074
Table S21. Gene #11: 'del_7q34' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| DEL PEAK 5(7Q34) MUTATED | 2 | 12 | 5 | 4 |
| DEL PEAK 5(7Q34) WILD-TYPE | 56 | 25 | 36 | 39 |
Figure S21. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
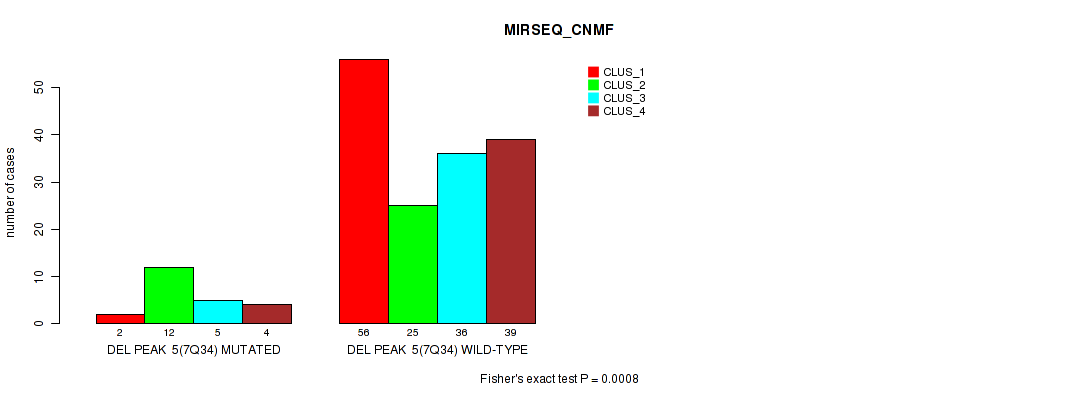
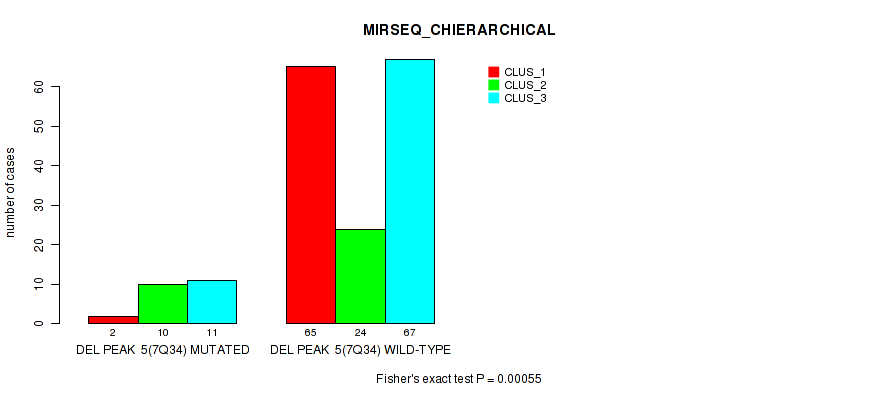
P value = 0.00055 (Fisher's exact test), Q value = 0.053
Table S22. Gene #11: 'del_7q34' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 5(7Q34) MUTATED | 2 | 10 | 11 |
| DEL PEAK 5(7Q34) WILD-TYPE | 65 | 24 | 67 |
Figure S22. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
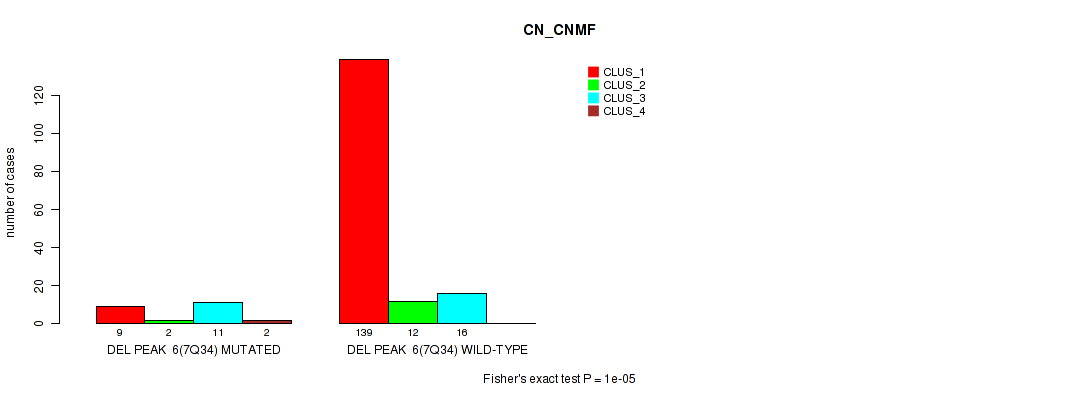
P value = 1e-05 (Fisher's exact test), Q value = 0.0012
Table S23. Gene #12: 'del_7q34' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 6(7Q34) MUTATED | 9 | 2 | 11 | 2 |
| DEL PEAK 6(7Q34) WILD-TYPE | 139 | 12 | 16 | 0 |
Figure S23. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #1: 'CN_CNMF'
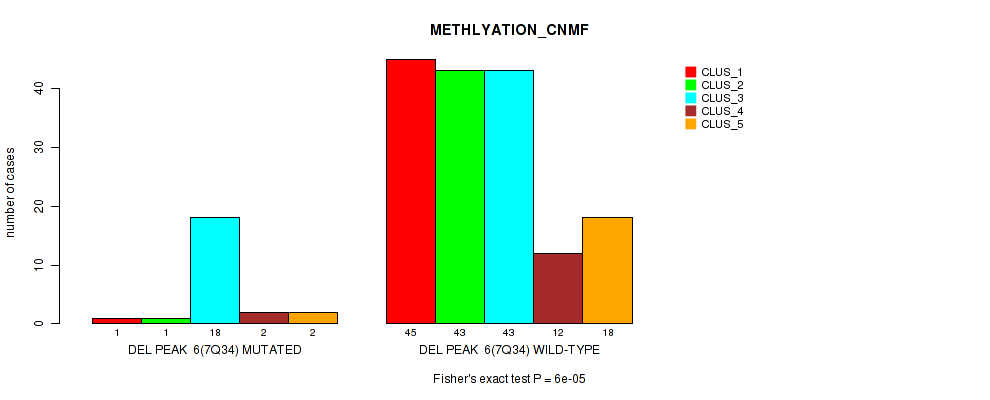
P value = 6e-05 (Fisher's exact test), Q value = 0.0062
Table S24. Gene #12: 'del_7q34' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 46 | 44 | 61 | 14 | 20 |
| DEL PEAK 6(7Q34) MUTATED | 1 | 1 | 18 | 2 | 2 |
| DEL PEAK 6(7Q34) WILD-TYPE | 45 | 43 | 43 | 12 | 18 |
Figure S24. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #2: 'METHLYATION_CNMF'
P value = 2e-05 (Fisher's exact test), Q value = 0.0022
Table S25. Gene #12: 'del_7q34' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 6(7Q34) MUTATED | 1 | 1 | 0 | 18 | 0 | 0 |
| DEL PEAK 6(7Q34) WILD-TYPE | 15 | 12 | 14 | 37 | 26 | 42 |
Figure S25. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00092 (Fisher's exact test), Q value = 0.082
Table S26. Gene #12: 'del_7q34' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| DEL PEAK 6(7Q34) MUTATED | 2 | 12 | 5 | 4 |
| DEL PEAK 6(7Q34) WILD-TYPE | 56 | 25 | 36 | 39 |
Figure S26. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #5: 'MIRSEQ_CNMF'

P value = 0.00056 (Fisher's exact test), Q value = 0.054
Table S27. Gene #12: 'del_7q34' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 6(7Q34) MUTATED | 2 | 10 | 11 |
| DEL PEAK 6(7Q34) WILD-TYPE | 65 | 24 | 67 |
Figure S27. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 2e-05 (Fisher's exact test), Q value = 0.0022
Table S28. Gene #14: 'del_12p13.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 10(12P13.2) MUTATED | 2 | 0 | 7 | 1 |
| DEL PEAK 10(12P13.2) WILD-TYPE | 146 | 14 | 20 | 1 |
Figure S28. Get High-res Image Gene #14: 'del_12p13.2' versus Molecular Subtype #1: 'CN_CNMF'

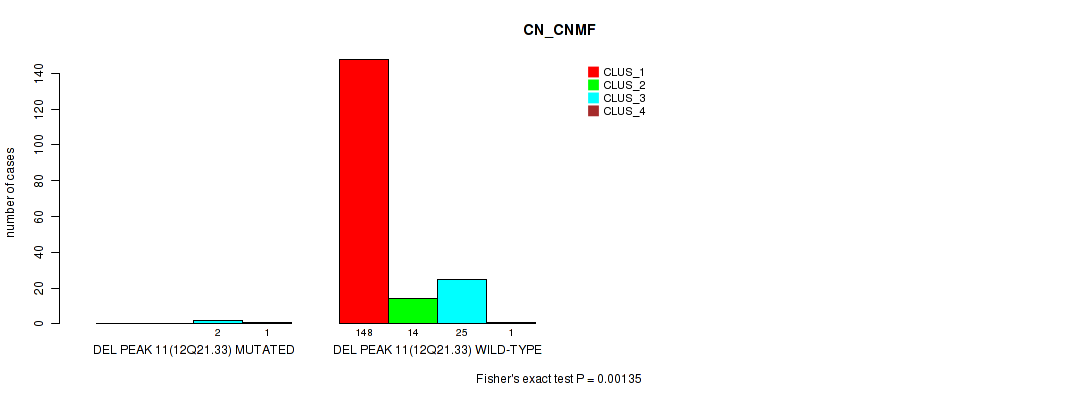
P value = 0.00135 (Fisher's exact test), Q value = 0.11
Table S29. Gene #15: 'del_12q21.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 11(12Q21.33) MUTATED | 0 | 0 | 2 | 1 |
| DEL PEAK 11(12Q21.33) WILD-TYPE | 148 | 14 | 25 | 1 |
Figure S29. Get High-res Image Gene #15: 'del_12q21.33' versus Molecular Subtype #1: 'CN_CNMF'
P value = 0.00121 (Fisher's exact test), Q value = 0.11
Table S30. Gene #16: 'del_16q23.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 12(16Q23.1) MUTATED | 3 | 0 | 6 | 0 |
| DEL PEAK 12(16Q23.1) WILD-TYPE | 145 | 14 | 21 | 2 |
Figure S30. Get High-res Image Gene #16: 'del_16q23.1' versus Molecular Subtype #1: 'CN_CNMF'

P value = 1e-05 (Fisher's exact test), Q value = 0.0012
Table S31. Gene #17: 'del_17p13.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 13(17P13.2) MUTATED | 0 | 0 | 15 | 0 |
| DEL PEAK 13(17P13.2) WILD-TYPE | 148 | 14 | 12 | 2 |
Figure S31. Get High-res Image Gene #17: 'del_17p13.2' versus Molecular Subtype #1: 'CN_CNMF'

P value = 0.00085 (Fisher's exact test), Q value = 0.076
Table S32. Gene #17: 'del_17p13.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 |
|---|---|---|---|---|---|
| ALL | 46 | 44 | 61 | 14 | 20 |
| DEL PEAK 13(17P13.2) MUTATED | 0 | 1 | 12 | 0 | 2 |
| DEL PEAK 13(17P13.2) WILD-TYPE | 46 | 43 | 49 | 14 | 18 |
Figure S32. Get High-res Image Gene #17: 'del_17p13.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'

P value = 2e-04 (Fisher's exact test), Q value = 0.02
Table S33. Gene #17: 'del_17p13.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 13(17P13.2) MUTATED | 0 | 0 | 0 | 14 | 0 | 1 |
| DEL PEAK 13(17P13.2) WILD-TYPE | 16 | 13 | 14 | 41 | 26 | 41 |
Figure S33. Get High-res Image Gene #17: 'del_17p13.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'

P value = 0.00116 (Fisher's exact test), Q value = 0.1
Table S34. Gene #17: 'del_17p13.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 13(17P13.2) MUTATED | 0 | 5 | 10 |
| DEL PEAK 13(17P13.2) WILD-TYPE | 67 | 29 | 68 |
Figure S34. Get High-res Image Gene #17: 'del_17p13.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'

P value = 1e-05 (Fisher's exact test), Q value = 0.0012
Table S35. Gene #18: 'del_17q11.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 14(17Q11.2) MUTATED | 2 | 2 | 9 | 0 |
| DEL PEAK 14(17Q11.2) WILD-TYPE | 146 | 12 | 18 | 2 |
Figure S35. Get High-res Image Gene #18: 'del_17q11.2' versus Molecular Subtype #1: 'CN_CNMF'

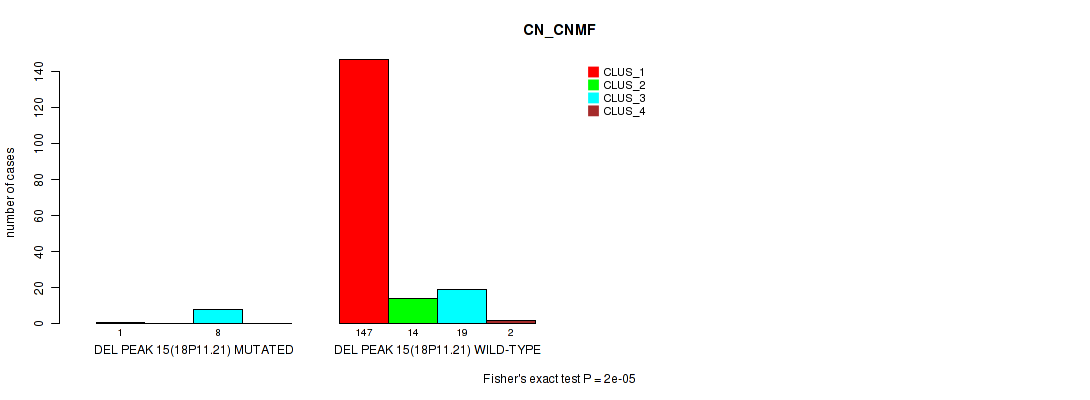
P value = 2e-05 (Fisher's exact test), Q value = 0.0022
Table S36. Gene #19: 'del_18p11.21' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 15(18P11.21) MUTATED | 1 | 0 | 8 | 0 |
| DEL PEAK 15(18P11.21) WILD-TYPE | 147 | 14 | 19 | 2 |
Figure S36. Get High-res Image Gene #19: 'del_18p11.21' versus Molecular Subtype #1: 'CN_CNMF'
-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtype file = LAML-TB.transferedmergedcluster.txt
-
Number of patients = 191
-
Number of significantly focal cnvs = 20
-
Number of molecular subtypes = 6
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.