This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 47 focal events and 8 molecular subtypes across 150 patients, 42 significant findings detected with P value < 0.05 and Q value < 0.25.
-
amp_7q11.22 cnv correlated to 'CN_CNMF' and 'MRNASEQ_CNMF'.
-
amp_8q11.23 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
amp_8q23.3 cnv correlated to 'MRNASEQ_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
amp_16q21 cnv correlated to 'CN_CNMF'.
-
amp_22q11.21 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_6p22.3 cnv correlated to 'CN_CNMF'.
-
del_6q26 cnv correlated to 'METHLYATION_CNMF'.
-
del_9q33.3 cnv correlated to 'CN_CNMF'.
-
del_9q34.2 cnv correlated to 'CN_CNMF'.
-
del_10p15.3 cnv correlated to 'METHLYATION_CNMF' and 'MRNASEQ_CNMF'.
-
del_10q23.1 cnv correlated to 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_10q26.3 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', and 'MRNASEQ_CHIERARCHICAL'.
-
del_11p15.5 cnv correlated to 'MIRSEQ_CNMF' and 'MIRSEQ_CHIERARCHICAL'.
-
del_11q12.3 cnv correlated to 'CN_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_11q24.2 cnv correlated to 'CN_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
del_19q13.41 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', 'MIRSEQ_CHIERARCHICAL', 'MIRSEQ_MATURE_CNMF', and 'MIRSEQ_MATURE_CHIERARCHICAL'.
-
del_xp21.1 cnv correlated to 'METHLYATION_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 47 focal events and 8 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 42 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRSEQ MATURE CNMF |
MIRSEQ MATURE CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| del 19q13 41 | 58 (39%) | 92 |
1e-05 (0.00364) |
1e-05 (0.00364) |
1e-05 (0.00364) |
1e-05 (0.00364) |
1e-05 (0.00364) |
1e-05 (0.00364) |
1e-05 (0.00364) |
1e-05 (0.00364) |
| del 11q12 3 | 114 (76%) | 36 |
1e-05 (0.00364) |
0.186 (1.00) |
7e-05 (0.024) |
1e-05 (0.00364) |
3e-05 (0.0104) |
0.00029 (0.0969) |
0.00748 (1.00) |
0.00728 (1.00) |
| del 10q26 3 | 76 (51%) | 74 |
0.00047 (0.154) |
1e-05 (0.00364) |
1e-05 (0.00364) |
0.00015 (0.0507) |
0.588 (1.00) |
0.0347 (1.00) |
0.0384 (1.00) |
0.0111 (1.00) |
| amp 8q11 23 | 122 (81%) | 28 |
4e-05 (0.0139) |
0.00966 (1.00) |
1e-05 (0.00364) |
4e-05 (0.0139) |
0.0126 (1.00) |
0.0261 (1.00) |
0.0239 (1.00) |
0.0829 (1.00) |
| amp 22q11 21 | 57 (38%) | 93 |
1e-05 (0.00364) |
0.00719 (1.00) |
0.00013 (0.0441) |
1e-05 (0.00364) |
0.0702 (1.00) |
0.236 (1.00) |
0.331 (1.00) |
0.0443 (1.00) |
| del 10q23 1 | 71 (47%) | 79 |
0.00596 (1.00) |
1e-05 (0.00364) |
4e-05 (0.0139) |
0.00022 (0.0737) |
0.357 (1.00) |
0.0487 (1.00) |
0.0834 (1.00) |
0.0363 (1.00) |
| amp 7q11 22 | 120 (80%) | 30 |
0.0006 (0.196) |
0.0393 (1.00) |
5e-05 (0.0172) |
0.00229 (0.701) |
0.0159 (1.00) |
0.00368 (1.00) |
0.0153 (1.00) |
0.0467 (1.00) |
| amp 8q23 3 | 108 (72%) | 42 |
0.184 (1.00) |
0.00328 (0.977) |
0.00035 (0.116) |
0.00058 (0.19) |
0.00603 (1.00) |
0.0208 (1.00) |
0.0354 (1.00) |
0.143 (1.00) |
| del 10p15 3 | 84 (56%) | 66 |
0.00124 (0.391) |
0.00036 (0.119) |
0.0002 (0.0674) |
0.00113 (0.359) |
0.9 (1.00) |
0.0596 (1.00) |
0.234 (1.00) |
0.279 (1.00) |
| del 11p15 5 | 94 (63%) | 56 |
0.0251 (1.00) |
0.256 (1.00) |
0.003 (0.9) |
0.00402 (1.00) |
0.0002 (0.0674) |
0.00037 (0.122) |
0.00125 (0.392) |
0.0163 (1.00) |
| del 11q24 2 | 122 (81%) | 28 |
7e-05 (0.024) |
0.725 (1.00) |
0.00897 (1.00) |
0.00068 (0.22) |
0.00094 (0.301) |
0.0379 (1.00) |
0.019 (1.00) |
0.0431 (1.00) |
| amp 16q21 | 21 (14%) | 129 |
8e-05 (0.0272) |
0.373 (1.00) |
0.00282 (0.854) |
0.0076 (1.00) |
0.0346 (1.00) |
0.00285 (0.861) |
0.0011 (0.351) |
0.0407 (1.00) |
| del 6p22 3 | 24 (16%) | 126 |
7e-05 (0.024) |
0.0157 (1.00) |
0.00386 (1.00) |
0.0126 (1.00) |
0.322 (1.00) |
0.0777 (1.00) |
0.116 (1.00) |
0.224 (1.00) |
| del 6q26 | 26 (17%) | 124 |
0.0324 (1.00) |
0.00075 (0.242) |
0.00592 (1.00) |
0.00135 (0.421) |
0.468 (1.00) |
0.29 (1.00) |
0.246 (1.00) |
0.459 (1.00) |
| del 9q33 3 | 60 (40%) | 90 |
0.00037 (0.122) |
0.394 (1.00) |
0.118 (1.00) |
0.0156 (1.00) |
0.32 (1.00) |
0.00252 (0.769) |
0.0262 (1.00) |
0.0538 (1.00) |
| del 9q34 2 | 60 (40%) | 90 |
0.00029 (0.0969) |
0.276 (1.00) |
0.0954 (1.00) |
0.00396 (1.00) |
0.509 (1.00) |
0.0125 (1.00) |
0.0754 (1.00) |
0.182 (1.00) |
| del xp21 1 | 28 (19%) | 122 |
0.0585 (1.00) |
0.00067 (0.218) |
0.0453 (1.00) |
0.0523 (1.00) |
0.357 (1.00) |
0.245 (1.00) |
0.082 (1.00) |
0.158 (1.00) |
| amp 1q21 2 | 69 (46%) | 81 |
0.0778 (1.00) |
0.718 (1.00) |
0.87 (1.00) |
0.00085 (0.274) |
0.0178 (1.00) |
0.445 (1.00) |
0.673 (1.00) |
0.194 (1.00) |
| amp 2p24 1 | 62 (41%) | 88 |
0.00407 (1.00) |
0.00087 (0.279) |
0.0191 (1.00) |
0.0441 (1.00) |
0.188 (1.00) |
0.19 (1.00) |
0.202 (1.00) |
0.316 (1.00) |
| amp 3q27 3 | 49 (33%) | 101 |
0.0776 (1.00) |
0.102 (1.00) |
0.00254 (0.772) |
0.00771 (1.00) |
0.152 (1.00) |
0.183 (1.00) |
0.259 (1.00) |
0.233 (1.00) |
| amp 4q12 | 20 (13%) | 130 |
0.00849 (1.00) |
0.661 (1.00) |
0.393 (1.00) |
0.394 (1.00) |
0.863 (1.00) |
0.69 (1.00) |
0.129 (1.00) |
0.429 (1.00) |
| amp 7p22 1 | 126 (84%) | 24 |
0.0863 (1.00) |
0.0779 (1.00) |
0.0289 (1.00) |
0.103 (1.00) |
0.042 (1.00) |
0.0362 (1.00) |
0.191 (1.00) |
0.177 (1.00) |
| amp 12p13 33 | 145 (97%) | 5 |
0.053 (1.00) |
0.102 (1.00) |
0.0826 (1.00) |
0.684 (1.00) |
||||
| amp 12p12 1 | 147 (98%) | 3 |
0.0549 (1.00) |
0.109 (1.00) |
0.444 (1.00) |
0.558 (1.00) |
||||
| amp 12p11 21 | 144 (96%) | 6 |
0.108 (1.00) |
0.0391 (1.00) |
0.036 (1.00) |
0.497 (1.00) |
0.11 (1.00) |
0.469 (1.00) |
0.188 (1.00) |
0.236 (1.00) |
| amp 12q15 | 100 (67%) | 50 |
0.527 (1.00) |
0.93 (1.00) |
0.813 (1.00) |
0.734 (1.00) |
0.325 (1.00) |
0.564 (1.00) |
0.837 (1.00) |
0.628 (1.00) |
| amp 20q11 21 | 51 (34%) | 99 |
0.0048 (1.00) |
0.00292 (0.879) |
0.00301 (0.9) |
0.0342 (1.00) |
0.0121 (1.00) |
0.0487 (1.00) |
0.0304 (1.00) |
0.0279 (1.00) |
| del 1p36 31 | 34 (23%) | 116 |
0.282 (1.00) |
0.733 (1.00) |
0.00228 (0.7) |
0.0507 (1.00) |
0.115 (1.00) |
0.0778 (1.00) |
0.127 (1.00) |
0.33 (1.00) |
| del 1p31 1 | 27 (18%) | 123 |
0.00709 (1.00) |
0.601 (1.00) |
0.247 (1.00) |
0.723 (1.00) |
0.396 (1.00) |
0.42 (1.00) |
0.498 (1.00) |
0.866 (1.00) |
| del 1q42 12 | 9 (6%) | 141 |
0.0386 (1.00) |
0.52 (1.00) |
0.562 (1.00) |
0.195 (1.00) |
0.133 (1.00) |
0.188 (1.00) |
0.114 (1.00) |
0.0605 (1.00) |
| del 2q37 3 | 15 (10%) | 135 |
0.457 (1.00) |
0.0408 (1.00) |
0.0576 (1.00) |
0.173 (1.00) |
0.676 (1.00) |
0.0283 (1.00) |
0.117 (1.00) |
0.215 (1.00) |
| del 3p26 2 | 41 (27%) | 109 |
0.76 (1.00) |
0.0207 (1.00) |
0.00133 (0.416) |
0.00382 (1.00) |
0.0497 (1.00) |
0.0743 (1.00) |
0.0907 (1.00) |
0.0391 (1.00) |
| del 4q22 2 | 110 (73%) | 40 |
0.0211 (1.00) |
0.0321 (1.00) |
0.00897 (1.00) |
0.00144 (0.448) |
0.116 (1.00) |
0.108 (1.00) |
0.624 (1.00) |
0.506 (1.00) |
| del 5p15 2 | 101 (67%) | 49 |
0.187 (1.00) |
0.489 (1.00) |
0.214 (1.00) |
0.457 (1.00) |
0.685 (1.00) |
0.222 (1.00) |
0.34 (1.00) |
0.368 (1.00) |
| del 5p14 3 | 103 (69%) | 47 |
0.264 (1.00) |
0.318 (1.00) |
0.18 (1.00) |
0.294 (1.00) |
0.528 (1.00) |
0.123 (1.00) |
0.237 (1.00) |
0.304 (1.00) |
| del 5q12 1 | 105 (70%) | 45 |
0.0895 (1.00) |
0.104 (1.00) |
0.0302 (1.00) |
0.00745 (1.00) |
0.501 (1.00) |
0.0489 (1.00) |
0.1 (1.00) |
0.155 (1.00) |
| del 7q11 22 | 4 (3%) | 146 |
0.315 (1.00) |
0.00722 (1.00) |
0.00916 (1.00) |
0.0301 (1.00) |
||||
| del 7q36 1 | 15 (10%) | 135 |
0.0509 (1.00) |
0.889 (1.00) |
0.201 (1.00) |
0.00989 (1.00) |
0.455 (1.00) |
0.724 (1.00) |
0.0837 (1.00) |
0.18 (1.00) |
| del 9p24 3 | 70 (47%) | 80 |
0.0909 (1.00) |
0.896 (1.00) |
1 (1.00) |
0.805 (1.00) |
0.284 (1.00) |
0.768 (1.00) |
0.32 (1.00) |
0.374 (1.00) |
| del 9p23 | 66 (44%) | 84 |
0.0383 (1.00) |
0.915 (1.00) |
0.908 (1.00) |
0.523 (1.00) |
0.325 (1.00) |
0.804 (1.00) |
0.293 (1.00) |
0.41 (1.00) |
| del 11p12 | 99 (66%) | 51 |
0.0556 (1.00) |
0.111 (1.00) |
0.00163 (0.504) |
0.00523 (1.00) |
0.00338 (1.00) |
0.00373 (1.00) |
0.0296 (1.00) |
0.0483 (1.00) |
| del 13q34 | 113 (75%) | 37 |
0.0441 (1.00) |
0.554 (1.00) |
0.0828 (1.00) |
0.00191 (0.588) |
0.418 (1.00) |
0.276 (1.00) |
0.00349 (1.00) |
0.0274 (1.00) |
| del 16q23 1 | 66 (44%) | 84 |
0.00118 (0.373) |
0.234 (1.00) |
0.357 (1.00) |
0.393 (1.00) |
0.292 (1.00) |
0.437 (1.00) |
0.162 (1.00) |
0.251 (1.00) |
| del 18p11 32 | 118 (79%) | 32 |
0.699 (1.00) |
0.132 (1.00) |
0.0503 (1.00) |
0.247 (1.00) |
0.124 (1.00) |
0.098 (1.00) |
0.0844 (1.00) |
0.108 (1.00) |
| del 18q22 2 | 125 (83%) | 25 |
0.0586 (1.00) |
0.143 (1.00) |
0.109 (1.00) |
0.133 (1.00) |
0.155 (1.00) |
0.2 (1.00) |
0.0207 (1.00) |
0.00596 (1.00) |
| del 20p12 1 | 29 (19%) | 121 |
0.335 (1.00) |
0.336 (1.00) |
0.31 (1.00) |
0.095 (1.00) |
0.492 (1.00) |
0.187 (1.00) |
0.111 (1.00) |
0.0741 (1.00) |
| del xq27 3 | 30 (20%) | 120 |
0.00113 (0.359) |
0.00157 (0.487) |
0.0942 (1.00) |
0.0627 (1.00) |
0.591 (1.00) |
0.383 (1.00) |
0.101 (1.00) |
0.604 (1.00) |
P value = 6e-04 (Fisher's exact test), Q value = 0.2
Table S1. Gene #6: 'amp_7q11.22' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| AMP PEAK 6(7Q11.22) MUTATED | 35 | 43 | 42 |
| AMP PEAK 6(7Q11.22) WILD-TYPE | 13 | 1 | 16 |
Figure S1. Get High-res Image Gene #6: 'amp_7q11.22' versus Molecular Subtype #1: 'CN_CNMF'
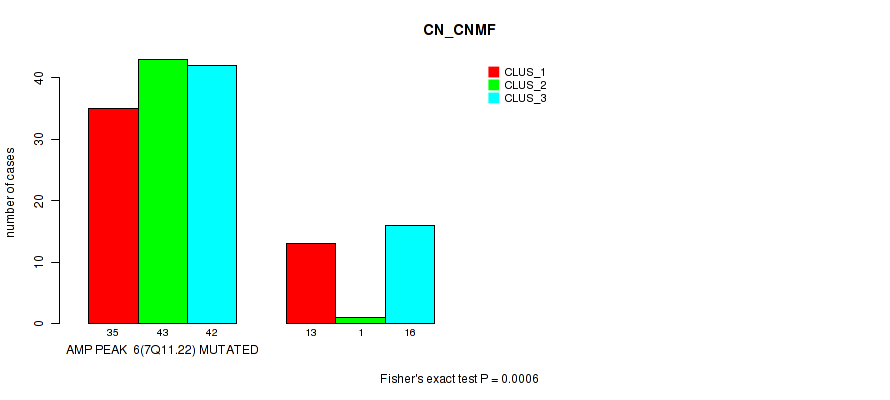
P value = 5e-05 (Fisher's exact test), Q value = 0.017
Table S2. Gene #6: 'amp_7q11.22' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 40 | 40 |
| AMP PEAK 6(7Q11.22) MUTATED | 63 | 35 | 22 |
| AMP PEAK 6(7Q11.22) WILD-TYPE | 7 | 5 | 18 |
Figure S2. Get High-res Image Gene #6: 'amp_7q11.22' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
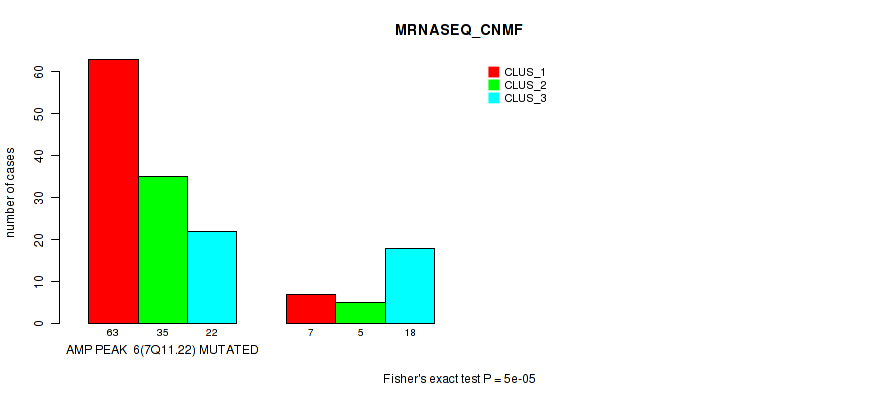
P value = 4e-05 (Fisher's exact test), Q value = 0.014
Table S3. Gene #7: 'amp_8q11.23' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| AMP PEAK 7(8Q11.23) MUTATED | 35 | 44 | 43 |
| AMP PEAK 7(8Q11.23) WILD-TYPE | 13 | 0 | 15 |
Figure S3. Get High-res Image Gene #7: 'amp_8q11.23' versus Molecular Subtype #1: 'CN_CNMF'
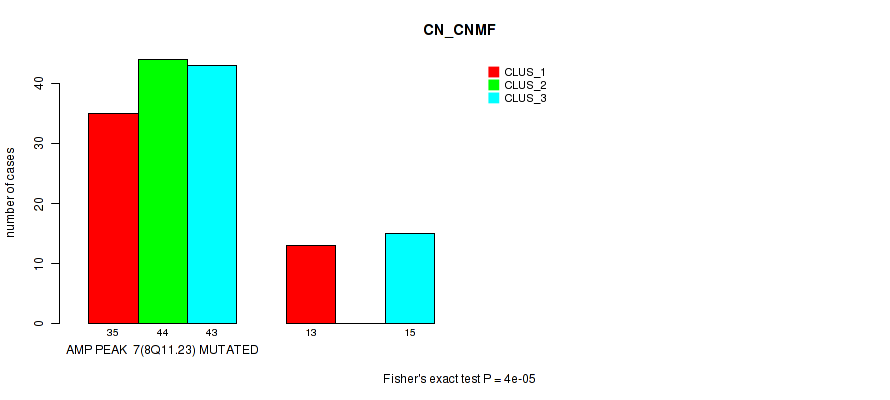
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S4. Gene #7: 'amp_8q11.23' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 40 | 40 |
| AMP PEAK 7(8Q11.23) MUTATED | 67 | 31 | 24 |
| AMP PEAK 7(8Q11.23) WILD-TYPE | 3 | 9 | 16 |
Figure S4. Get High-res Image Gene #7: 'amp_8q11.23' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
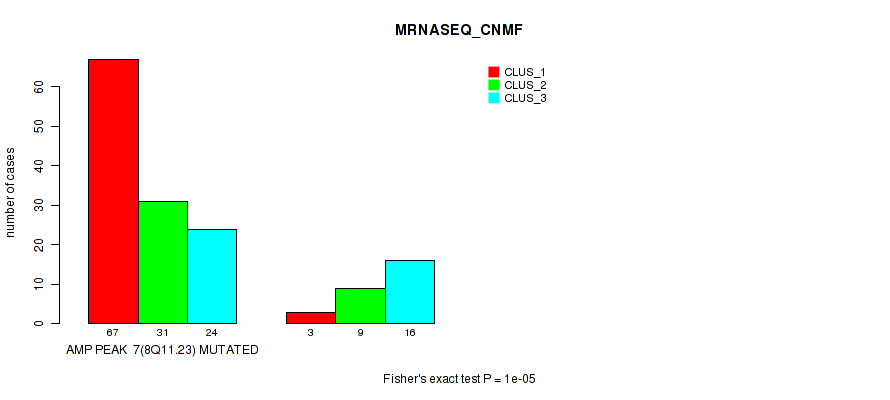
P value = 4e-05 (Fisher's exact test), Q value = 0.014
Table S5. Gene #7: 'amp_8q11.23' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 43 | 27 | 12 | 24 | 8 | 23 | 13 |
| AMP PEAK 7(8Q11.23) MUTATED | 40 | 27 | 8 | 18 | 6 | 19 | 4 |
| AMP PEAK 7(8Q11.23) WILD-TYPE | 3 | 0 | 4 | 6 | 2 | 4 | 9 |
Figure S5. Get High-res Image Gene #7: 'amp_8q11.23' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
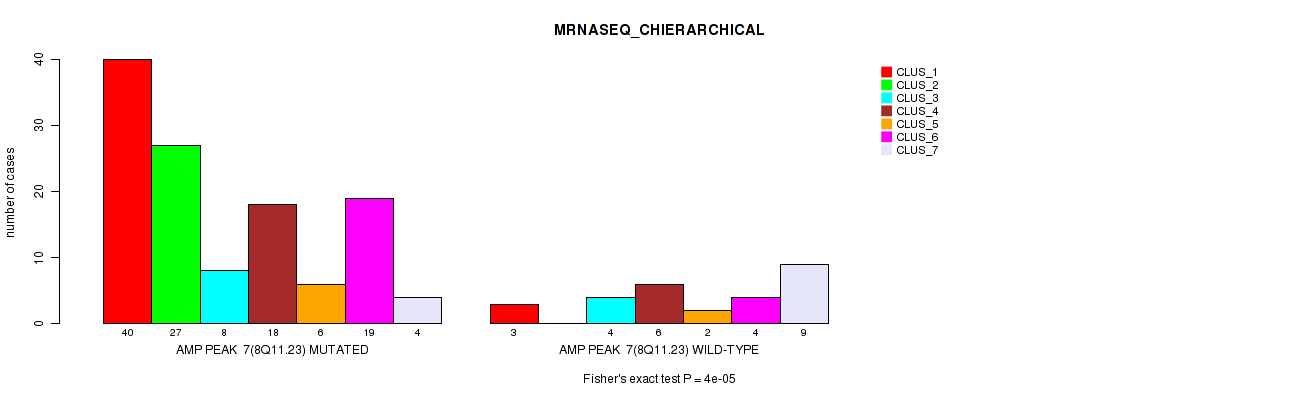
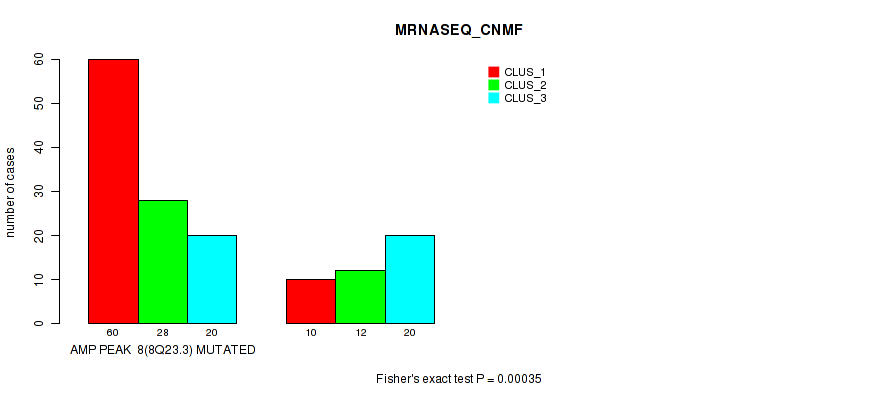
P value = 0.00035 (Fisher's exact test), Q value = 0.12
Table S6. Gene #8: 'amp_8q23.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 40 | 40 |
| AMP PEAK 8(8Q23.3) MUTATED | 60 | 28 | 20 |
| AMP PEAK 8(8Q23.3) WILD-TYPE | 10 | 12 | 20 |
Figure S6. Get High-res Image Gene #8: 'amp_8q23.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
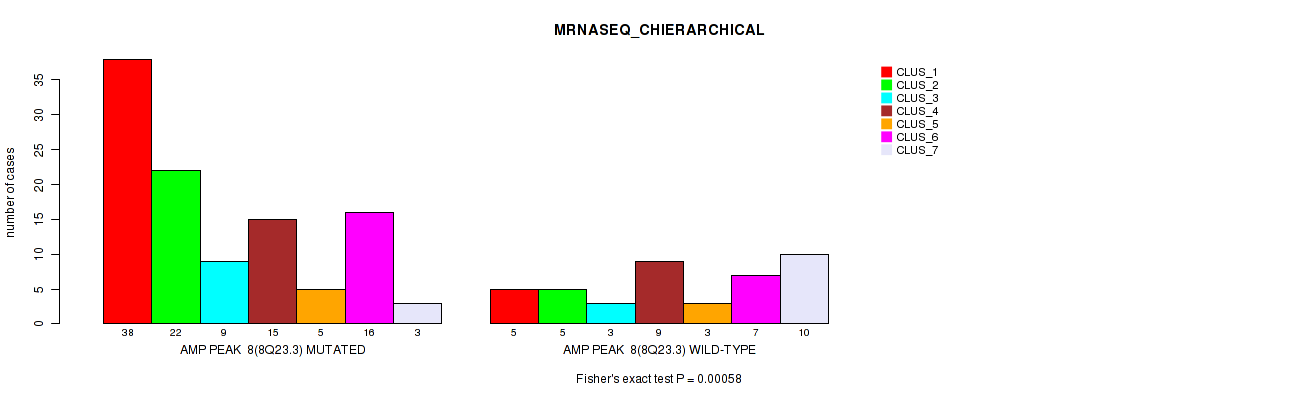
P value = 0.00058 (Fisher's exact test), Q value = 0.19
Table S7. Gene #8: 'amp_8q23.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 43 | 27 | 12 | 24 | 8 | 23 | 13 |
| AMP PEAK 8(8Q23.3) MUTATED | 38 | 22 | 9 | 15 | 5 | 16 | 3 |
| AMP PEAK 8(8Q23.3) WILD-TYPE | 5 | 5 | 3 | 9 | 3 | 7 | 10 |
Figure S7. Get High-res Image Gene #8: 'amp_8q23.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
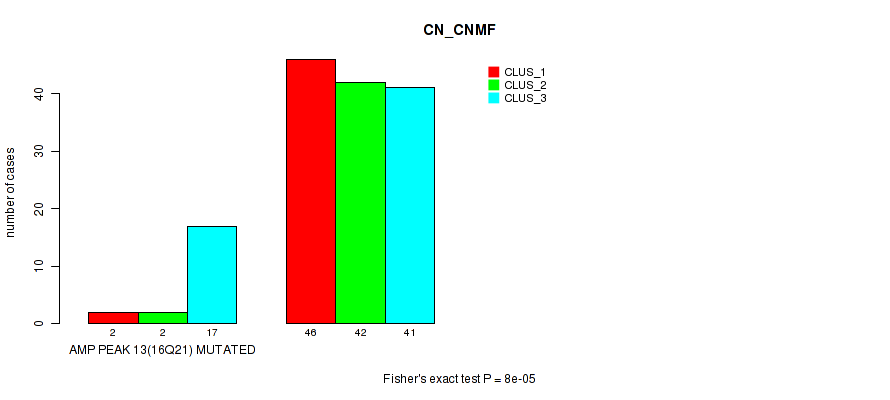
P value = 8e-05 (Fisher's exact test), Q value = 0.027
Table S8. Gene #13: 'amp_16q21' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| AMP PEAK 13(16Q21) MUTATED | 2 | 2 | 17 |
| AMP PEAK 13(16Q21) WILD-TYPE | 46 | 42 | 41 |
Figure S8. Get High-res Image Gene #13: 'amp_16q21' versus Molecular Subtype #1: 'CN_CNMF'
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S9. Gene #15: 'amp_22q11.21' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| AMP PEAK 15(22Q11.21) MUTATED | 13 | 30 | 14 |
| AMP PEAK 15(22Q11.21) WILD-TYPE | 35 | 14 | 44 |
Figure S9. Get High-res Image Gene #15: 'amp_22q11.21' versus Molecular Subtype #1: 'CN_CNMF'
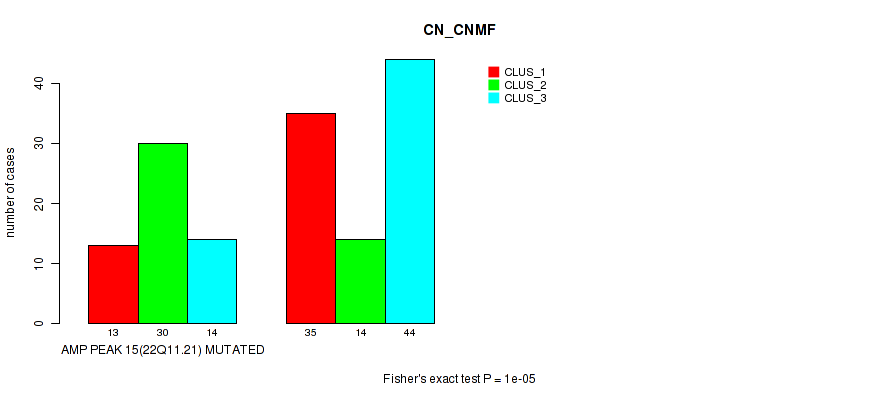
P value = 0.00013 (Fisher's exact test), Q value = 0.044
Table S10. Gene #15: 'amp_22q11.21' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 40 | 40 |
| AMP PEAK 15(22Q11.21) MUTATED | 39 | 10 | 8 |
| AMP PEAK 15(22Q11.21) WILD-TYPE | 31 | 30 | 32 |
Figure S10. Get High-res Image Gene #15: 'amp_22q11.21' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
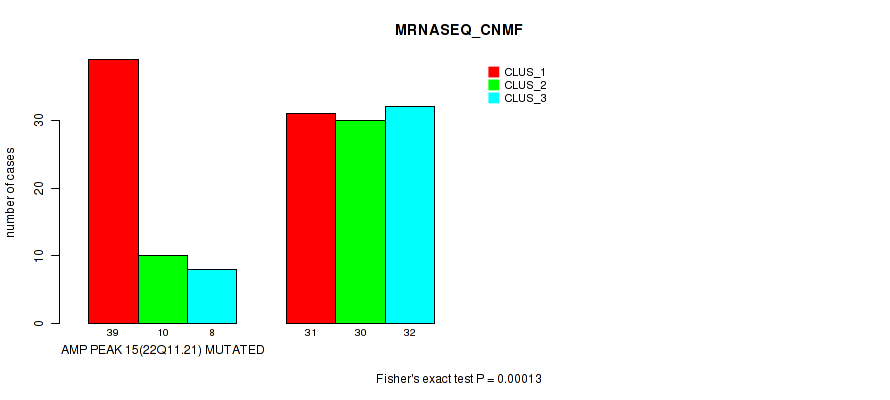
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S11. Gene #15: 'amp_22q11.21' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 43 | 27 | 12 | 24 | 8 | 23 | 13 |
| AMP PEAK 15(22Q11.21) MUTATED | 16 | 23 | 4 | 5 | 1 | 5 | 3 |
| AMP PEAK 15(22Q11.21) WILD-TYPE | 27 | 4 | 8 | 19 | 7 | 18 | 10 |
Figure S11. Get High-res Image Gene #15: 'amp_22q11.21' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
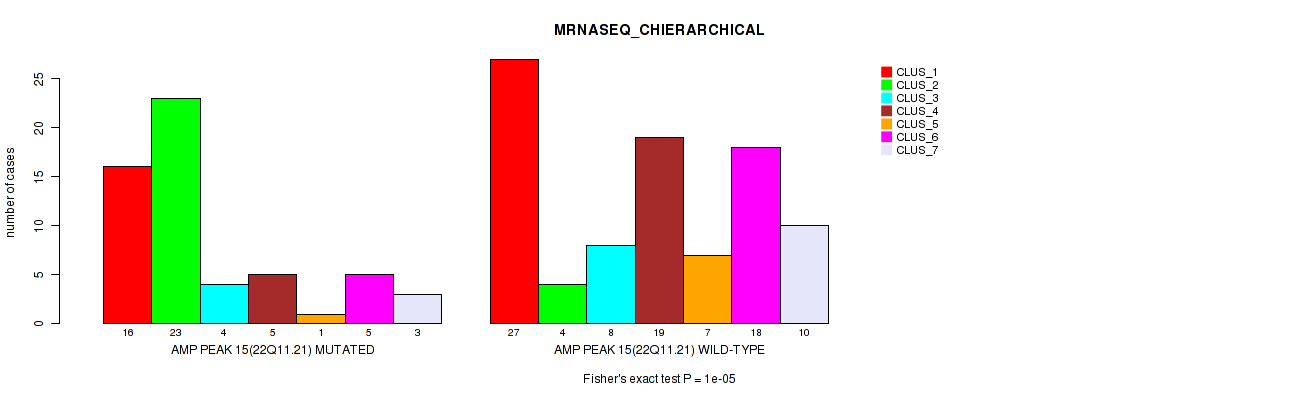
P value = 7e-05 (Fisher's exact test), Q value = 0.024
Table S12. Gene #25: 'del_6p22.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| DEL PEAK 10(6P22.3) MUTATED | 2 | 3 | 19 |
| DEL PEAK 10(6P22.3) WILD-TYPE | 46 | 41 | 39 |
Figure S12. Get High-res Image Gene #25: 'del_6p22.3' versus Molecular Subtype #1: 'CN_CNMF'
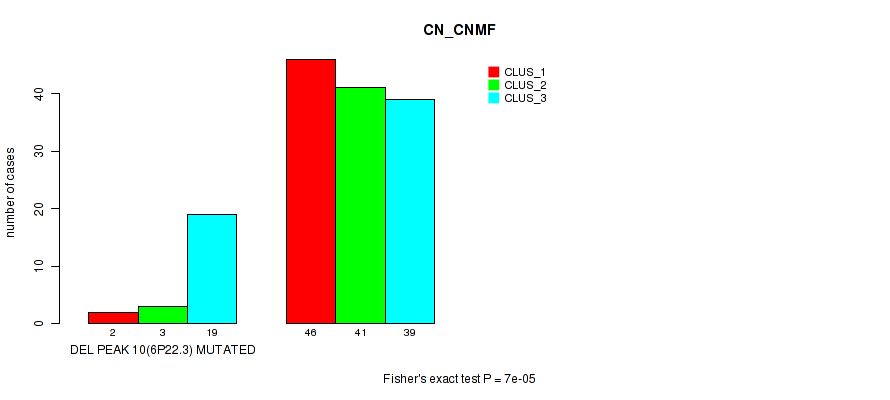
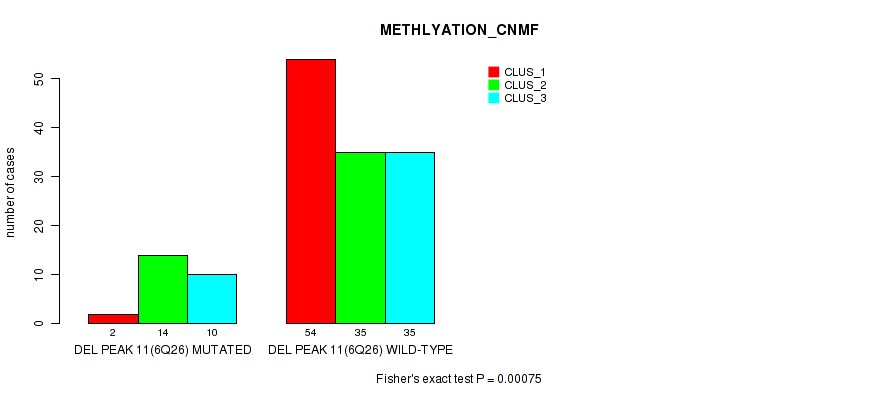
P value = 0.00075 (Fisher's exact test), Q value = 0.24
Table S13. Gene #26: 'del_6q26' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 56 | 49 | 45 |
| DEL PEAK 11(6Q26) MUTATED | 2 | 14 | 10 |
| DEL PEAK 11(6Q26) WILD-TYPE | 54 | 35 | 35 |
Figure S13. Get High-res Image Gene #26: 'del_6q26' versus Molecular Subtype #2: 'METHLYATION_CNMF'
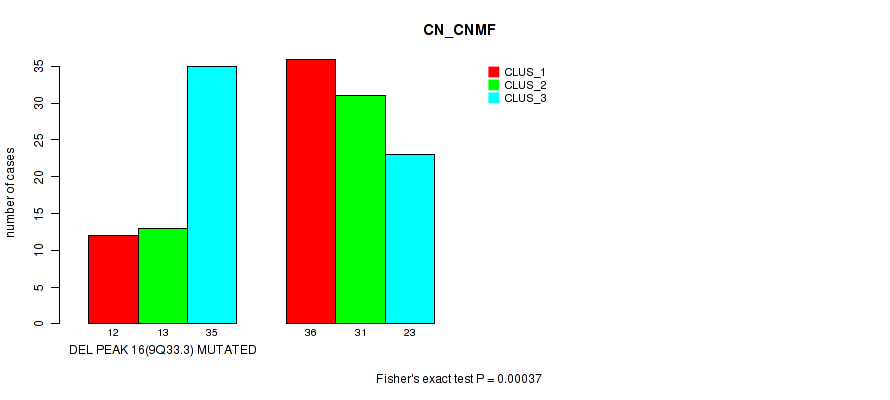
P value = 0.00037 (Fisher's exact test), Q value = 0.12
Table S14. Gene #31: 'del_9q33.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| DEL PEAK 16(9Q33.3) MUTATED | 12 | 13 | 35 |
| DEL PEAK 16(9Q33.3) WILD-TYPE | 36 | 31 | 23 |
Figure S14. Get High-res Image Gene #31: 'del_9q33.3' versus Molecular Subtype #1: 'CN_CNMF'
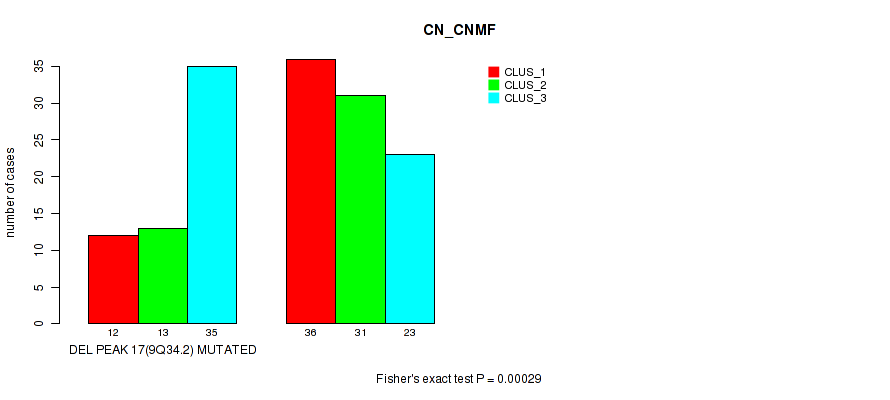
P value = 0.00029 (Fisher's exact test), Q value = 0.097
Table S15. Gene #32: 'del_9q34.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| DEL PEAK 17(9Q34.2) MUTATED | 12 | 13 | 35 |
| DEL PEAK 17(9Q34.2) WILD-TYPE | 36 | 31 | 23 |
Figure S15. Get High-res Image Gene #32: 'del_9q34.2' versus Molecular Subtype #1: 'CN_CNMF'
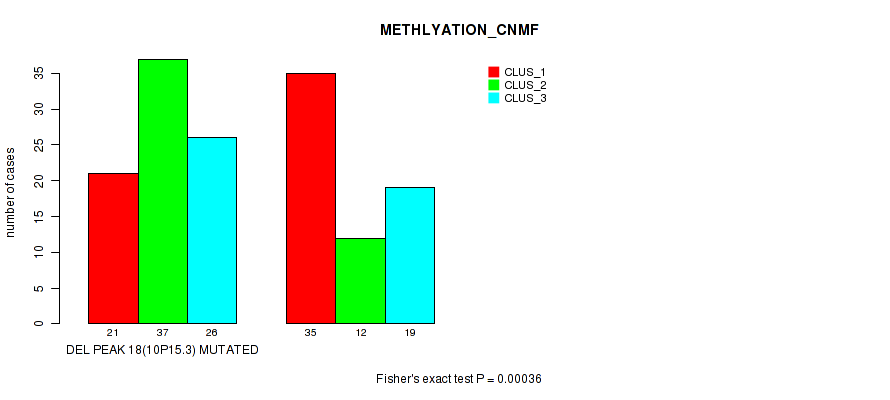
P value = 0.00036 (Fisher's exact test), Q value = 0.12
Table S16. Gene #33: 'del_10p15.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 56 | 49 | 45 |
| DEL PEAK 18(10P15.3) MUTATED | 21 | 37 | 26 |
| DEL PEAK 18(10P15.3) WILD-TYPE | 35 | 12 | 19 |
Figure S16. Get High-res Image Gene #33: 'del_10p15.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
P value = 2e-04 (Fisher's exact test), Q value = 0.067
Table S17. Gene #33: 'del_10p15.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 40 | 40 |
| DEL PEAK 18(10P15.3) MUTATED | 28 | 32 | 24 |
| DEL PEAK 18(10P15.3) WILD-TYPE | 42 | 8 | 16 |
Figure S17. Get High-res Image Gene #33: 'del_10p15.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
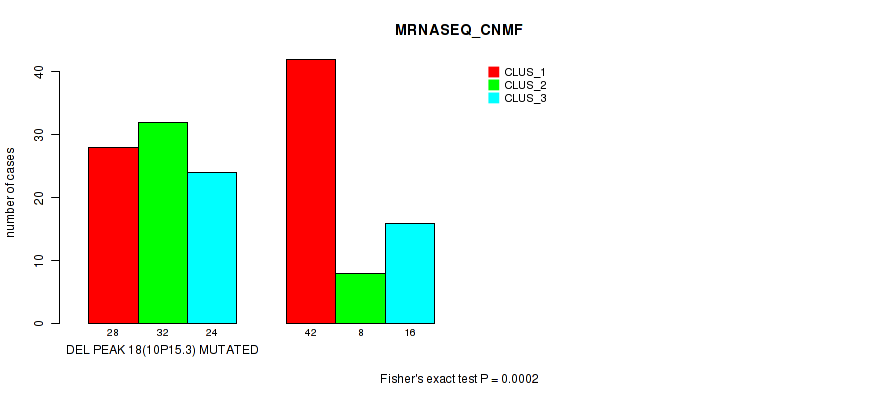
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S18. Gene #34: 'del_10q23.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 56 | 49 | 45 |
| DEL PEAK 19(10Q23.1) MUTATED | 15 | 37 | 19 |
| DEL PEAK 19(10Q23.1) WILD-TYPE | 41 | 12 | 26 |
Figure S18. Get High-res Image Gene #34: 'del_10q23.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
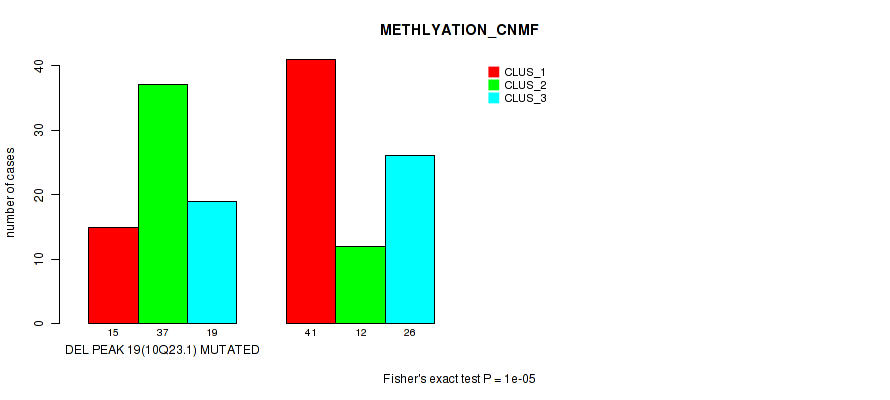
P value = 4e-05 (Fisher's exact test), Q value = 0.014
Table S19. Gene #34: 'del_10q23.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 40 | 40 |
| DEL PEAK 19(10Q23.1) MUTATED | 22 | 31 | 18 |
| DEL PEAK 19(10Q23.1) WILD-TYPE | 48 | 9 | 22 |
Figure S19. Get High-res Image Gene #34: 'del_10q23.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
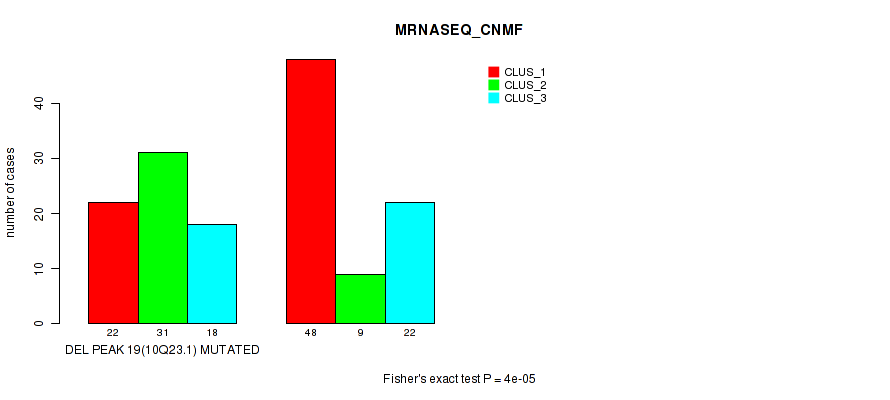
P value = 0.00022 (Fisher's exact test), Q value = 0.074
Table S20. Gene #34: 'del_10q23.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 43 | 27 | 12 | 24 | 8 | 23 | 13 |
| DEL PEAK 19(10Q23.1) MUTATED | 14 | 8 | 9 | 12 | 5 | 19 | 4 |
| DEL PEAK 19(10Q23.1) WILD-TYPE | 29 | 19 | 3 | 12 | 3 | 4 | 9 |
Figure S20. Get High-res Image Gene #34: 'del_10q23.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
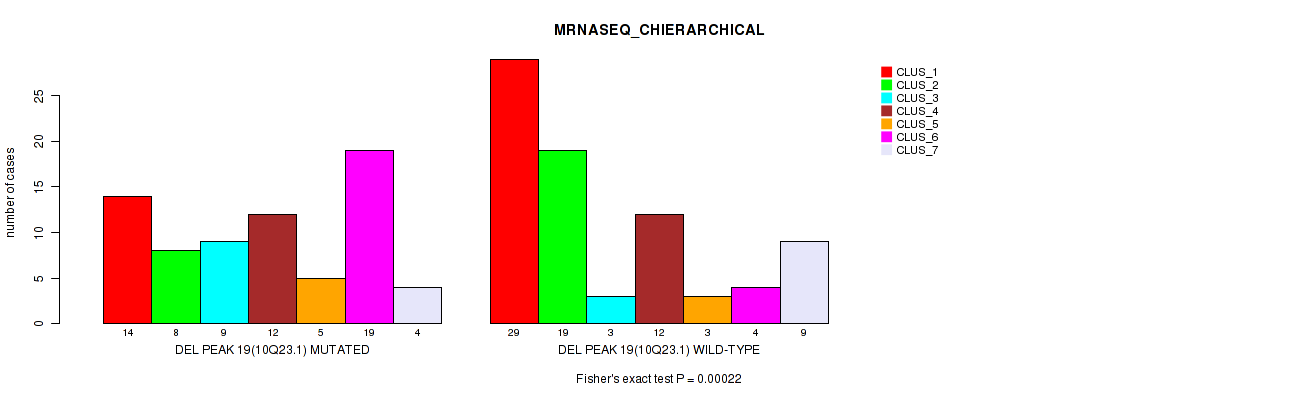
P value = 0.00047 (Fisher's exact test), Q value = 0.15
Table S21. Gene #35: 'del_10q26.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| DEL PEAK 20(10Q26.3) MUTATED | 18 | 17 | 41 |
| DEL PEAK 20(10Q26.3) WILD-TYPE | 30 | 27 | 17 |
Figure S21. Get High-res Image Gene #35: 'del_10q26.3' versus Molecular Subtype #1: 'CN_CNMF'
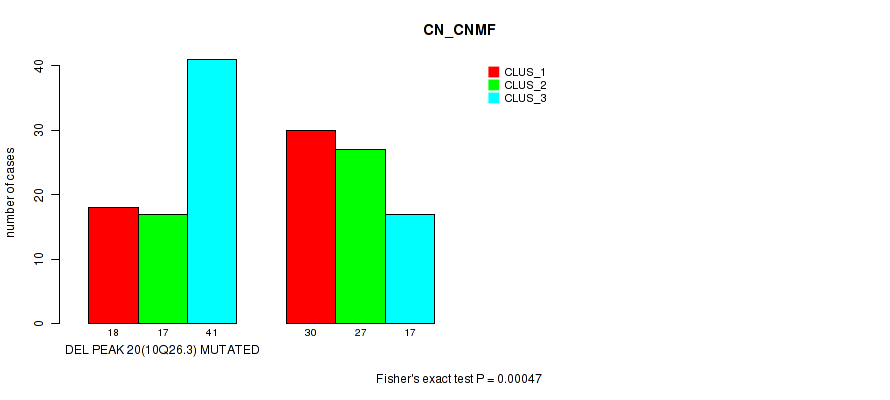
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S22. Gene #35: 'del_10q26.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 56 | 49 | 45 |
| DEL PEAK 20(10Q26.3) MUTATED | 16 | 38 | 22 |
| DEL PEAK 20(10Q26.3) WILD-TYPE | 40 | 11 | 23 |
Figure S22. Get High-res Image Gene #35: 'del_10q26.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
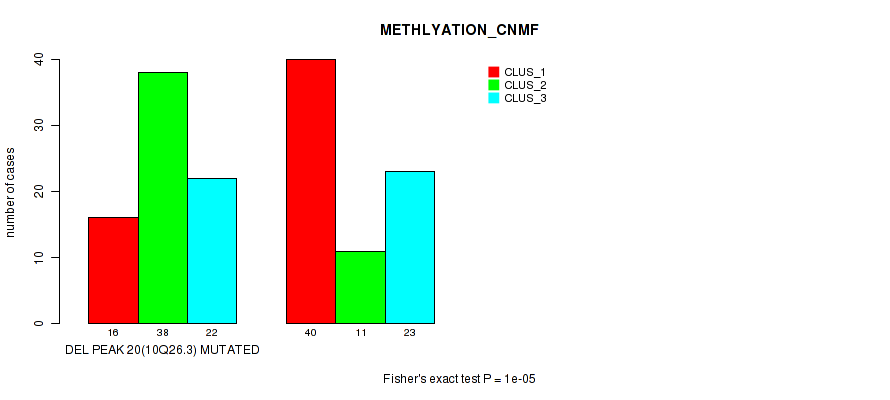
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S23. Gene #35: 'del_10q26.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 40 | 40 |
| DEL PEAK 20(10Q26.3) MUTATED | 24 | 33 | 19 |
| DEL PEAK 20(10Q26.3) WILD-TYPE | 46 | 7 | 21 |
Figure S23. Get High-res Image Gene #35: 'del_10q26.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
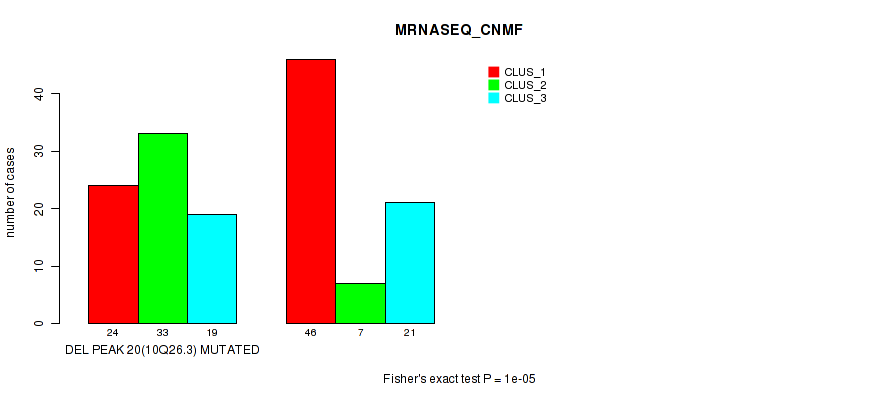
P value = 0.00015 (Fisher's exact test), Q value = 0.051
Table S24. Gene #35: 'del_10q26.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 43 | 27 | 12 | 24 | 8 | 23 | 13 |
| DEL PEAK 20(10Q26.3) MUTATED | 16 | 8 | 10 | 13 | 6 | 19 | 4 |
| DEL PEAK 20(10Q26.3) WILD-TYPE | 27 | 19 | 2 | 11 | 2 | 4 | 9 |
Figure S24. Get High-res Image Gene #35: 'del_10q26.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
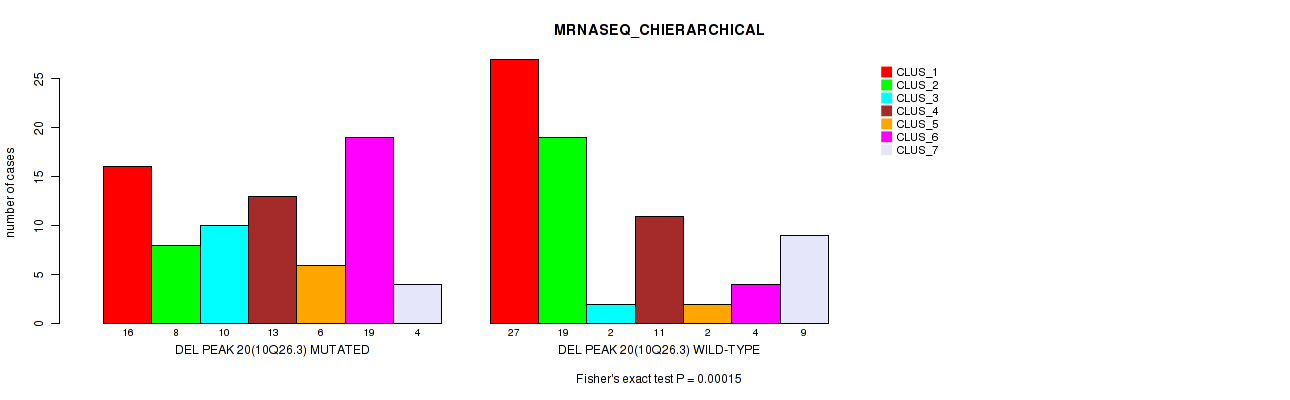
P value = 2e-04 (Fisher's exact test), Q value = 0.067
Table S25. Gene #36: 'del_11p15.5' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 21 | 16 | 25 |
| DEL PEAK 21(11P15.5) MUTATED | 10 | 15 | 4 | 22 |
| DEL PEAK 21(11P15.5) WILD-TYPE | 2 | 6 | 12 | 3 |
Figure S25. Get High-res Image Gene #36: 'del_11p15.5' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
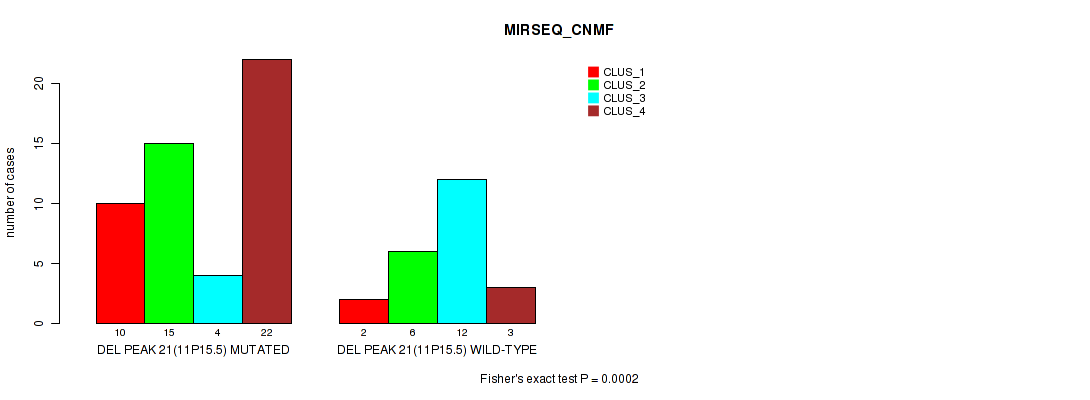
P value = 0.00037 (Fisher's exact test), Q value = 0.12
Table S26. Gene #36: 'del_11p15.5' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 44 | 15 | 15 |
| DEL PEAK 21(11P15.5) MUTATED | 36 | 11 | 4 |
| DEL PEAK 21(11P15.5) WILD-TYPE | 8 | 4 | 11 |
Figure S26. Get High-res Image Gene #36: 'del_11p15.5' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
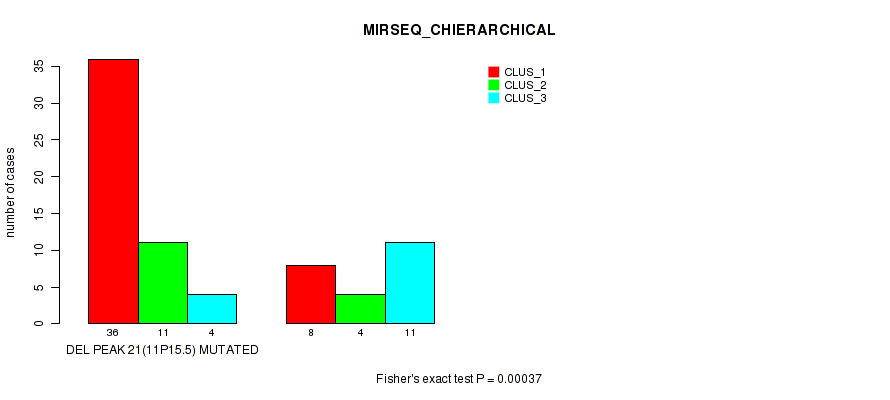
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S27. Gene #38: 'del_11q12.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| DEL PEAK 23(11Q12.3) MUTATED | 30 | 43 | 41 |
| DEL PEAK 23(11Q12.3) WILD-TYPE | 18 | 1 | 17 |
Figure S27. Get High-res Image Gene #38: 'del_11q12.3' versus Molecular Subtype #1: 'CN_CNMF'
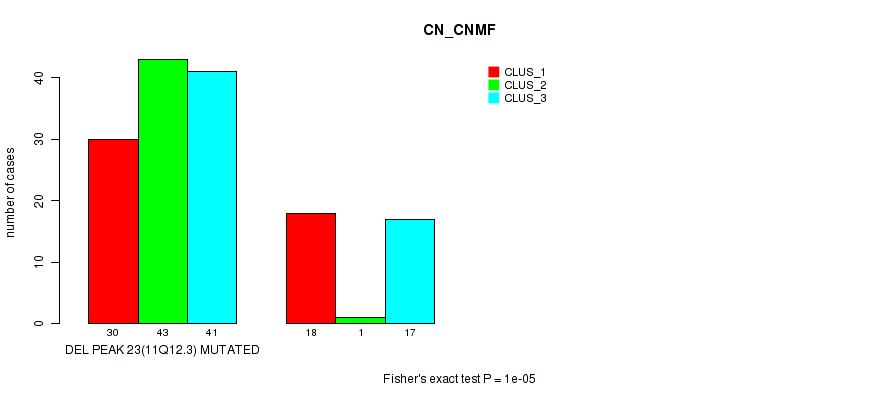
P value = 7e-05 (Fisher's exact test), Q value = 0.024
Table S28. Gene #38: 'del_11q12.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 40 | 40 |
| DEL PEAK 23(11Q12.3) MUTATED | 62 | 32 | 20 |
| DEL PEAK 23(11Q12.3) WILD-TYPE | 8 | 8 | 20 |
Figure S28. Get High-res Image Gene #38: 'del_11q12.3' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
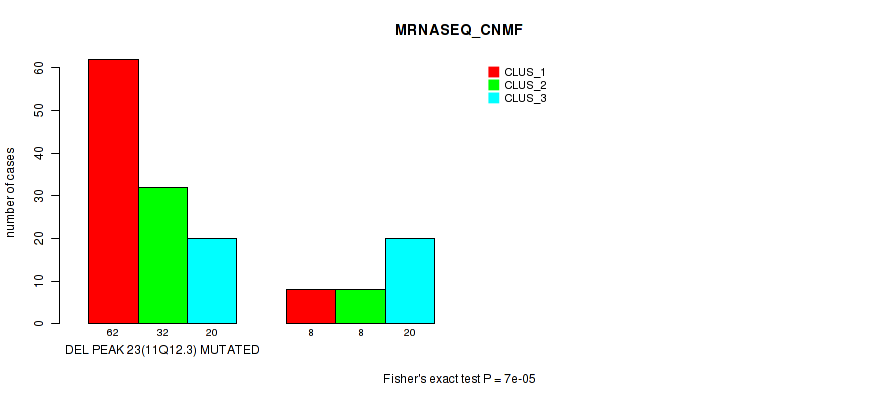
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S29. Gene #38: 'del_11q12.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 43 | 27 | 12 | 24 | 8 | 23 | 13 |
| DEL PEAK 23(11Q12.3) MUTATED | 35 | 27 | 9 | 14 | 6 | 20 | 3 |
| DEL PEAK 23(11Q12.3) WILD-TYPE | 8 | 0 | 3 | 10 | 2 | 3 | 10 |
Figure S29. Get High-res Image Gene #38: 'del_11q12.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
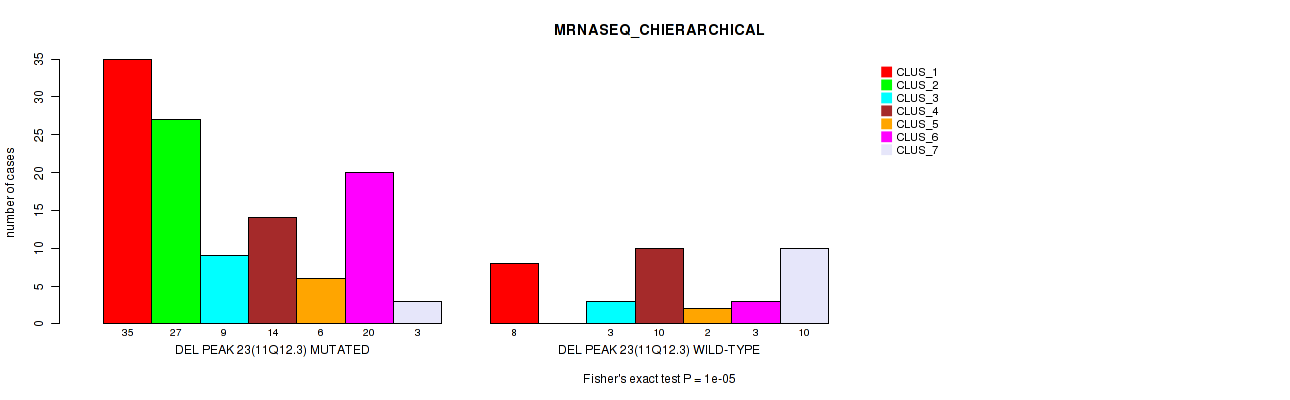
P value = 3e-05 (Fisher's exact test), Q value = 0.01
Table S30. Gene #38: 'del_11q12.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 21 | 16 | 25 |
| DEL PEAK 23(11Q12.3) MUTATED | 9 | 20 | 6 | 24 |
| DEL PEAK 23(11Q12.3) WILD-TYPE | 3 | 1 | 10 | 1 |
Figure S30. Get High-res Image Gene #38: 'del_11q12.3' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
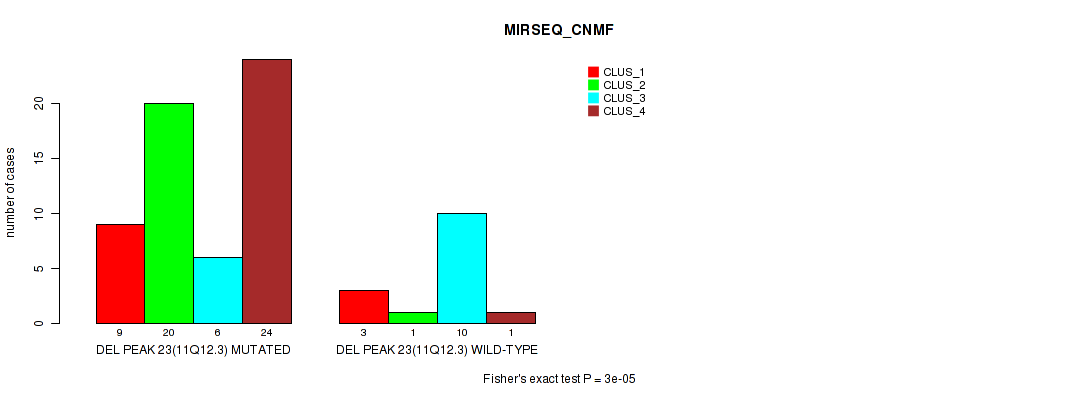
P value = 0.00029 (Fisher's exact test), Q value = 0.097
Table S31. Gene #38: 'del_11q12.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 44 | 15 | 15 |
| DEL PEAK 23(11Q12.3) MUTATED | 39 | 14 | 6 |
| DEL PEAK 23(11Q12.3) WILD-TYPE | 5 | 1 | 9 |
Figure S31. Get High-res Image Gene #38: 'del_11q12.3' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
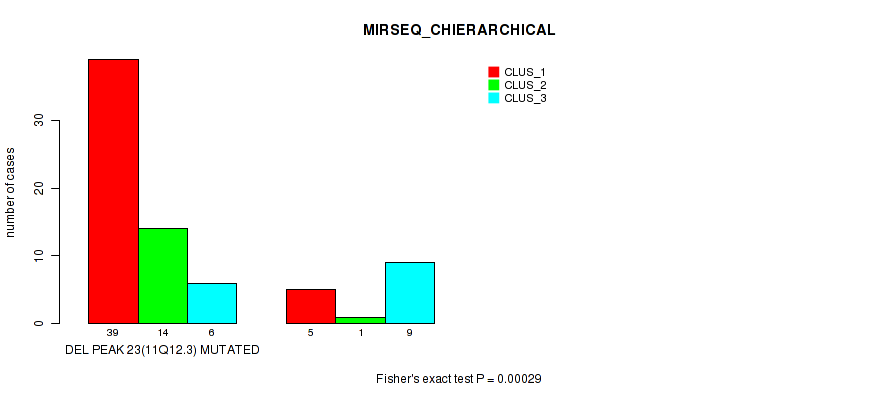
P value = 7e-05 (Fisher's exact test), Q value = 0.024
Table S32. Gene #39: 'del_11q24.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| DEL PEAK 24(11Q24.2) MUTATED | 33 | 44 | 45 |
| DEL PEAK 24(11Q24.2) WILD-TYPE | 15 | 0 | 13 |
Figure S32. Get High-res Image Gene #39: 'del_11q24.2' versus Molecular Subtype #1: 'CN_CNMF'
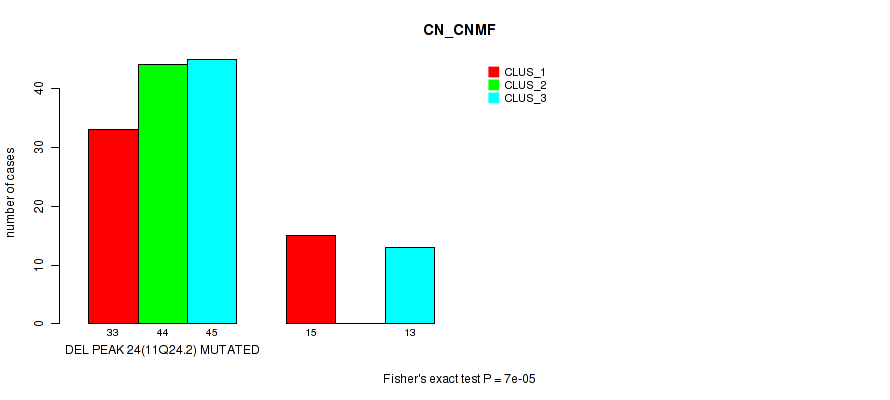
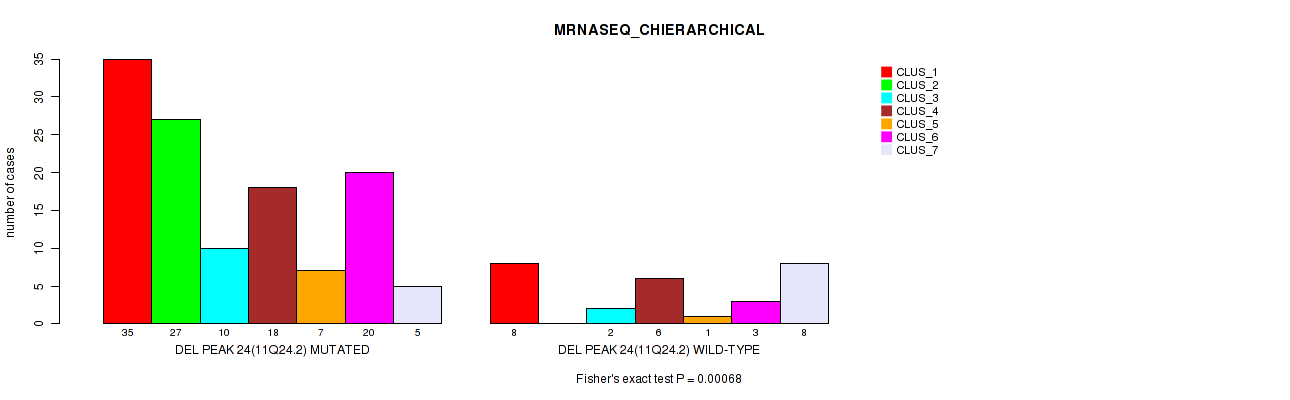
P value = 0.00068 (Fisher's exact test), Q value = 0.22
Table S33. Gene #39: 'del_11q24.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 43 | 27 | 12 | 24 | 8 | 23 | 13 |
| DEL PEAK 24(11Q24.2) MUTATED | 35 | 27 | 10 | 18 | 7 | 20 | 5 |
| DEL PEAK 24(11Q24.2) WILD-TYPE | 8 | 0 | 2 | 6 | 1 | 3 | 8 |
Figure S33. Get High-res Image Gene #39: 'del_11q24.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
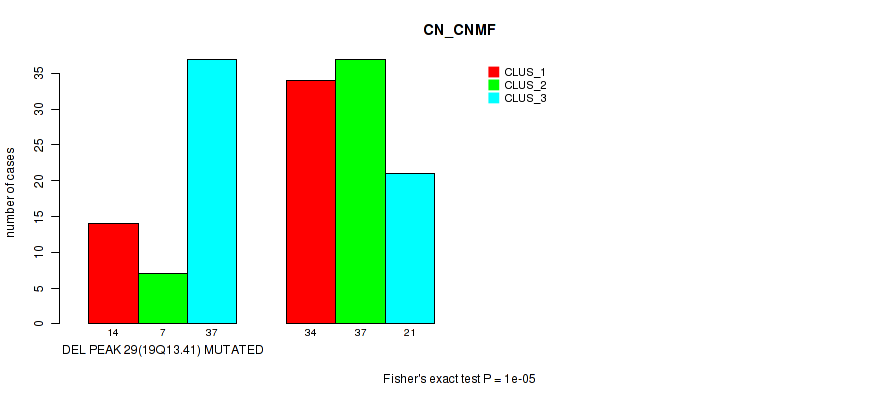
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S34. Gene #44: 'del_19q13.41' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 48 | 44 | 58 |
| DEL PEAK 29(19Q13.41) MUTATED | 14 | 7 | 37 |
| DEL PEAK 29(19Q13.41) WILD-TYPE | 34 | 37 | 21 |
Figure S34. Get High-res Image Gene #44: 'del_19q13.41' versus Molecular Subtype #1: 'CN_CNMF'
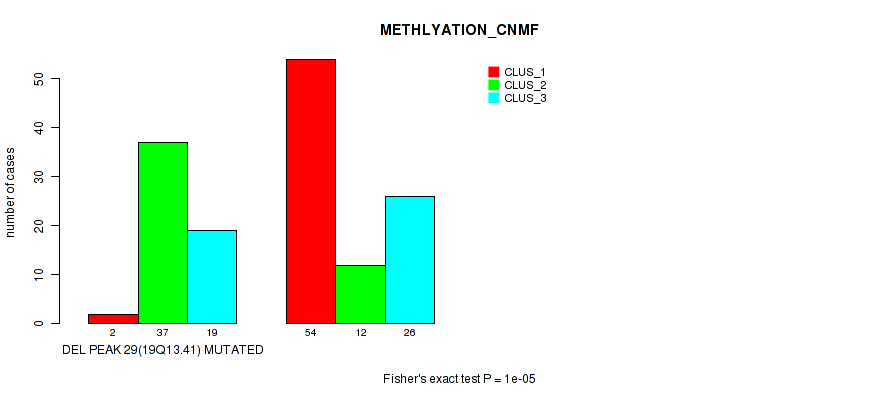
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S35. Gene #44: 'del_19q13.41' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 56 | 49 | 45 |
| DEL PEAK 29(19Q13.41) MUTATED | 2 | 37 | 19 |
| DEL PEAK 29(19Q13.41) WILD-TYPE | 54 | 12 | 26 |
Figure S35. Get High-res Image Gene #44: 'del_19q13.41' versus Molecular Subtype #2: 'METHLYATION_CNMF'
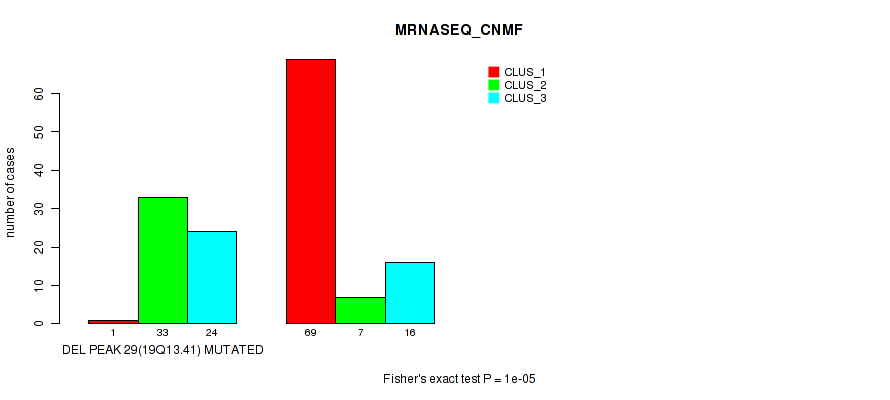
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S36. Gene #44: 'del_19q13.41' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 40 | 40 |
| DEL PEAK 29(19Q13.41) MUTATED | 1 | 33 | 24 |
| DEL PEAK 29(19Q13.41) WILD-TYPE | 69 | 7 | 16 |
Figure S36. Get High-res Image Gene #44: 'del_19q13.41' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
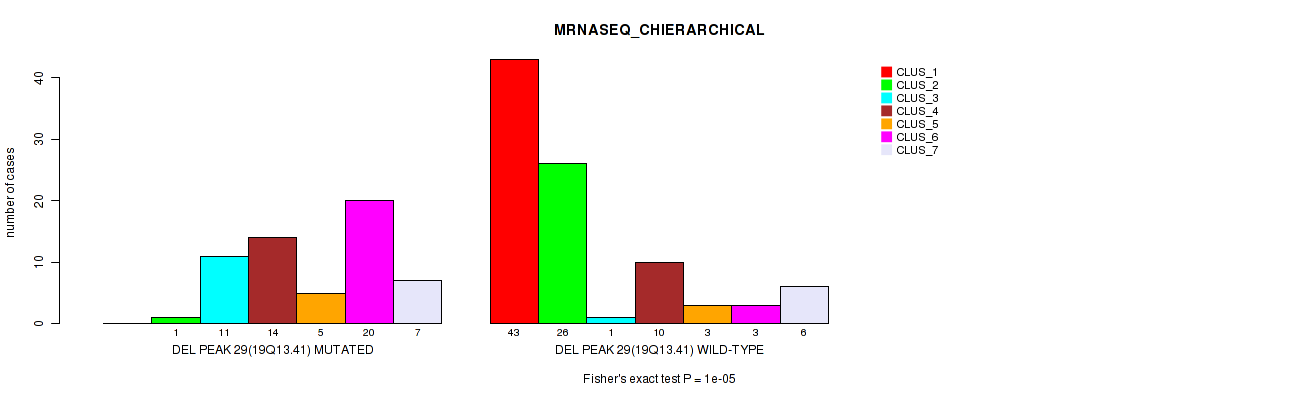
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S37. Gene #44: 'del_19q13.41' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 43 | 27 | 12 | 24 | 8 | 23 | 13 |
| DEL PEAK 29(19Q13.41) MUTATED | 0 | 1 | 11 | 14 | 5 | 20 | 7 |
| DEL PEAK 29(19Q13.41) WILD-TYPE | 43 | 26 | 1 | 10 | 3 | 3 | 6 |
Figure S37. Get High-res Image Gene #44: 'del_19q13.41' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
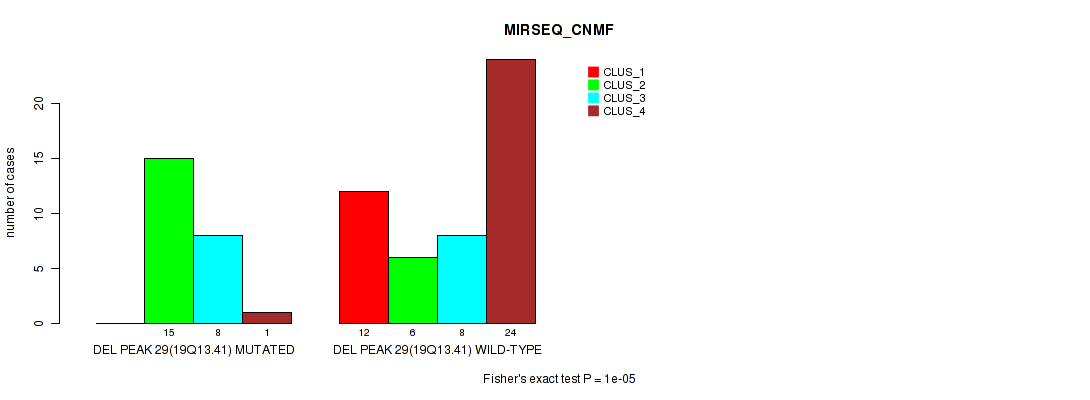
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S38. Gene #44: 'del_19q13.41' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 12 | 21 | 16 | 25 |
| DEL PEAK 29(19Q13.41) MUTATED | 0 | 15 | 8 | 1 |
| DEL PEAK 29(19Q13.41) WILD-TYPE | 12 | 6 | 8 | 24 |
Figure S38. Get High-res Image Gene #44: 'del_19q13.41' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
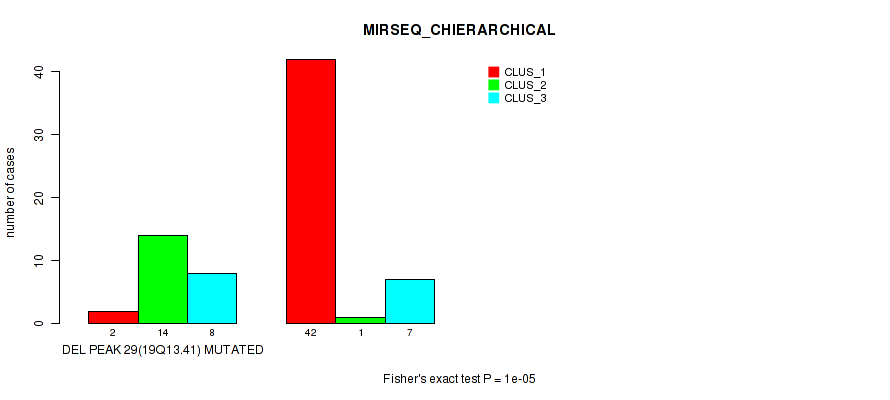
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S39. Gene #44: 'del_19q13.41' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 44 | 15 | 15 |
| DEL PEAK 29(19Q13.41) MUTATED | 2 | 14 | 8 |
| DEL PEAK 29(19Q13.41) WILD-TYPE | 42 | 1 | 7 |
Figure S39. Get High-res Image Gene #44: 'del_19q13.41' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
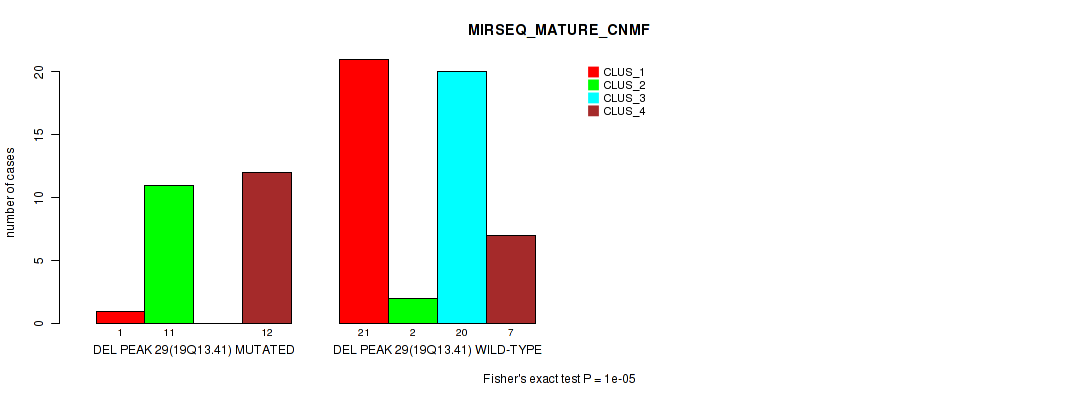
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S40. Gene #44: 'del_19q13.41' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 22 | 13 | 20 | 19 |
| DEL PEAK 29(19Q13.41) MUTATED | 1 | 11 | 0 | 12 |
| DEL PEAK 29(19Q13.41) WILD-TYPE | 21 | 2 | 20 | 7 |
Figure S40. Get High-res Image Gene #44: 'del_19q13.41' versus Molecular Subtype #7: 'MIRSEQ_MATURE_CNMF'
P value = 1e-05 (Fisher's exact test), Q value = 0.0036
Table S41. Gene #44: 'del_19q13.41' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 | CLUS_7 |
|---|---|---|---|---|---|---|---|
| ALL | 11 | 8 | 15 | 11 | 5 | 13 | 11 |
| DEL PEAK 29(19Q13.41) MUTATED | 0 | 0 | 14 | 6 | 3 | 1 | 0 |
| DEL PEAK 29(19Q13.41) WILD-TYPE | 11 | 8 | 1 | 5 | 2 | 12 | 11 |
Figure S41. Get High-res Image Gene #44: 'del_19q13.41' versus Molecular Subtype #8: 'MIRSEQ_MATURE_CHIERARCHICAL'
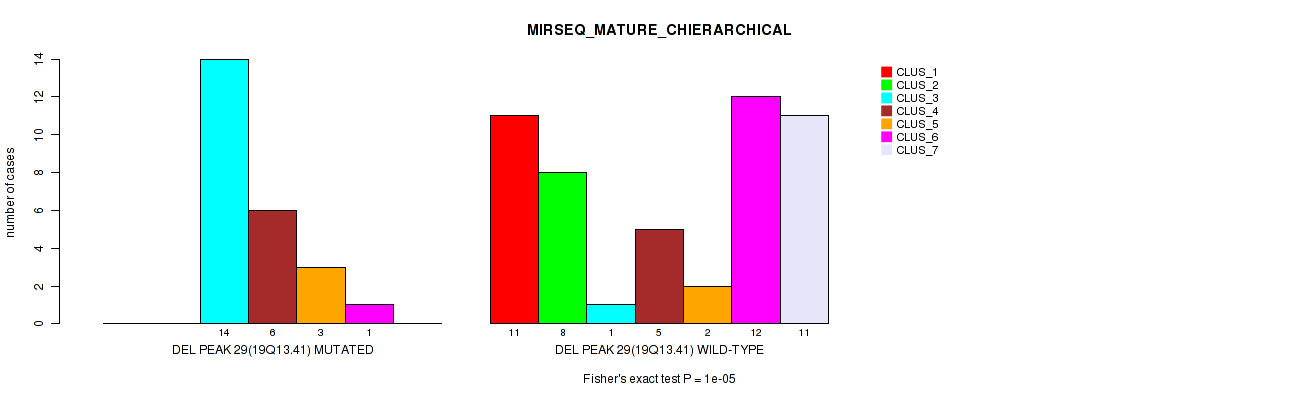
P value = 0.00067 (Fisher's exact test), Q value = 0.22
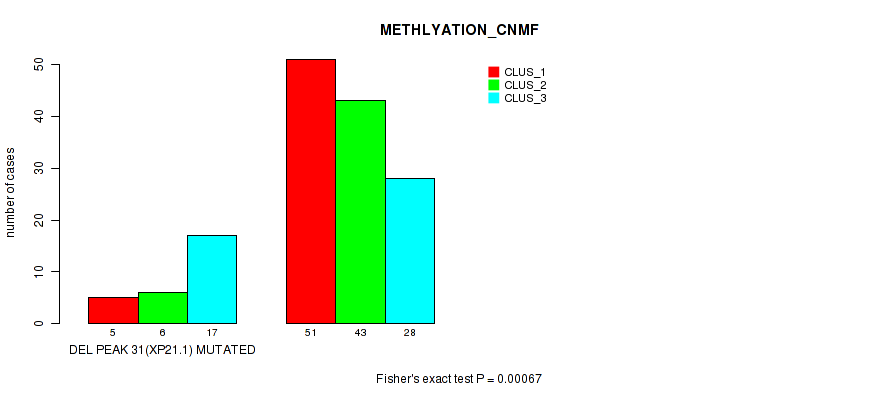
Table S42. Gene #46: 'del_xp21.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 56 | 49 | 45 |
| DEL PEAK 31(XP21.1) MUTATED | 5 | 6 | 17 |
| DEL PEAK 31(XP21.1) WILD-TYPE | 51 | 43 | 28 |
Figure S42. Get High-res Image Gene #46: 'del_xp21.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
-
Copy number data file = transformed.cor.cli.txt
-
Molecular subtype file = TGCT-TP.transferedmergedcluster.txt
-
Number of patients = 150
-
Number of significantly focal cnvs = 47
-
Number of molecular subtypes = 8
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.