This pipeline computes the correlation between significant copy number variation (cnv focal) genes and molecular subtypes.
Testing the association between copy number variation 20 focal events and 6 molecular subtypes across 191 patients, 69 significant findings detected with P value < 0.05 and Q value < 0.25.
-
amp_1p33 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
amp_1q43 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
amp_11q23.3 cnv correlated to 'CN_CNMF' and 'METHLYATION_CNMF'.
-
amp_13q31.3 cnv correlated to 'METHLYATION_CNMF' and 'MRNASEQ_CHIERARCHICAL'.
-
amp_19p13.2 cnv correlated to 'CN_CNMF'.
-
amp_20q11.21 cnv correlated to 'CN_CNMF'.
-
amp_21q22.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CNMF'.
-
del_3p13 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_5q31.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_7p12.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_7q34 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_7q34 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_12p13.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_12q21.33 cnv correlated to 'CN_CNMF', 'MIRSEQ_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_16q23.1 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MIRSEQ_CNMF'.
-
del_17p13.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
del_17q11.2 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', 'MRNASEQ_CHIERARCHICAL', and 'MIRSEQ_CHIERARCHICAL'.
-
del_18p11.21 cnv correlated to 'CN_CNMF', 'METHLYATION_CNMF', and 'MIRSEQ_CHIERARCHICAL'.
-
del_20q13.13 cnv correlated to 'CN_CNMF'.
Table 1. Get Full Table Overview of the association between significant copy number variation of 20 focal events and 6 molecular subtypes. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 69 significant findings detected.
|
Clinical Features |
CN CNMF |
METHLYATION CNMF |
MRNASEQ CNMF |
MRNASEQ CHIERARCHICAL |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
||
| nCNV (%) | nWild-Type | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| del 5q31 2 | 18 (9%) | 173 |
1e-05 (0.000109) |
1e-05 (0.000109) |
0.0055 (0.0143) |
0.00061 (0.00318) |
0.00102 (0.00437) |
0.00129 (0.00516) |
| del 7p12 1 | 16 (8%) | 175 |
0.00025 (0.00176) |
0.00034 (0.00215) |
0.0017 (0.00607) |
2e-05 (2e-04) |
0.00595 (0.0152) |
0.0076 (0.0179) |
| del 7q34 | 24 (13%) | 167 |
1e-05 (0.000109) |
0.00011 (0.000943) |
0.0228 (0.0464) |
1e-05 (0.000109) |
0.00079 (0.00365) |
0.00061 (0.00318) |
| del 7q34 | 24 (13%) | 167 |
1e-05 (0.000109) |
0.00012 (0.00096) |
0.0222 (0.046) |
1e-05 (0.000109) |
0.00084 (0.00373) |
0.00072 (0.00346) |
| amp 21q22 2 | 14 (7%) | 177 |
1e-05 (0.000109) |
0.00054 (0.00309) |
0.00325 (0.00929) |
0.037 (0.0683) |
0.00176 (0.00607) |
0.0675 (0.107) |
| del 17p13 2 | 15 (8%) | 176 |
1e-05 (0.000109) |
0.00033 (0.00215) |
0.0114 (0.0248) |
0.00018 (0.00135) |
0.0867 (0.132) |
0.00124 (0.00513) |
| amp 1p33 | 7 (4%) | 184 |
0.00052 (0.00309) |
0.00194 (0.00647) |
0.124 (0.182) |
0.13 (0.188) |
0.0097 (0.022) |
0.0485 (0.0844) |
| amp 1q43 | 7 (4%) | 184 |
0.00317 (0.00928) |
0.00218 (0.00688) |
0.373 (0.443) |
0.0362 (0.0683) |
0.138 (0.195) |
0.0146 (0.0313) |
| del 3p13 | 8 (4%) | 183 |
0.00065 (0.00325) |
0.0469 (0.084) |
0.066 (0.107) |
0.234 (0.293) |
0.0367 (0.0683) |
0.0485 (0.0844) |
| del 17q11 2 | 13 (7%) | 178 |
1e-05 (0.000109) |
0.00493 (0.0131) |
0.157 (0.218) |
0.00717 (0.0172) |
0.138 (0.195) |
0.0038 (0.0106) |
| del 12p13 2 | 10 (5%) | 181 |
4e-05 (0.000369) |
0.00151 (0.00566) |
0.198 (0.261) |
0.073 (0.114) |
0.0679 (0.107) |
0.01 (0.0223) |
| del 12q21 33 | 3 (2%) | 188 |
0.00135 (0.00523) |
0.703 (0.767) |
0.185 (0.247) |
0.586 (0.662) |
0.0085 (0.0196) |
0.00682 (0.0167) |
| del 16q23 1 | 9 (5%) | 182 |
0.00177 (0.00607) |
0.0222 (0.046) |
0.332 (0.406) |
0.494 (0.564) |
0.0437 (0.0795) |
0.0651 (0.107) |
| del 18p11 21 | 9 (5%) | 182 |
1e-05 (0.000109) |
0.00391 (0.0107) |
0.38 (0.447) |
0.335 (0.406) |
0.214 (0.273) |
0.0323 (0.0635) |
| amp 11q23 3 | 17 (9%) | 174 |
1e-05 (0.000109) |
0.00236 (0.00726) |
0.37 (0.443) |
0.2 (0.261) |
0.745 (0.799) |
0.182 (0.247) |
| amp 13q31 3 | 7 (4%) | 184 |
0.0802 (0.123) |
0.00211 (0.00684) |
0.0666 (0.107) |
0.0366 (0.0683) |
0.733 (0.793) |
0.0564 (0.0961) |
| amp 19p13 2 | 6 (3%) | 185 |
0.00279 (0.00837) |
0.25 (0.31) |
0.857 (0.902) |
0.632 (0.696) |
0.894 (0.933) |
1 (1.00) |
| amp 20q11 21 | 3 (2%) | 188 |
0.00654 (0.0163) |
0.214 (0.273) |
||||
| del 20q13 13 | 4 (2%) | 187 |
0.0299 (0.0597) |
0.0568 (0.0961) |
0.786 (0.835) |
0.59 (0.662) |
0.124 (0.182) |
0.414 (0.482) |
| del 9q21 32 | 5 (3%) | 186 |
0.233 (0.293) |
0.467 (0.539) |
0.178 (0.245) |
0.601 (0.668) |
0.184 (0.247) |
0.104 (0.157) |
P value = 0.00052 (Fisher's exact test), Q value = 0.0031
Table S1. Gene #1: 'amp_1p33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| AMP PEAK 1(1P33) MUTATED | 1 | 3 | 3 | 0 |
| AMP PEAK 1(1P33) WILD-TYPE | 147 | 11 | 24 | 2 |
Figure S1. Get High-res Image Gene #1: 'amp_1p33' versus Molecular Subtype #1: 'CN_CNMF'
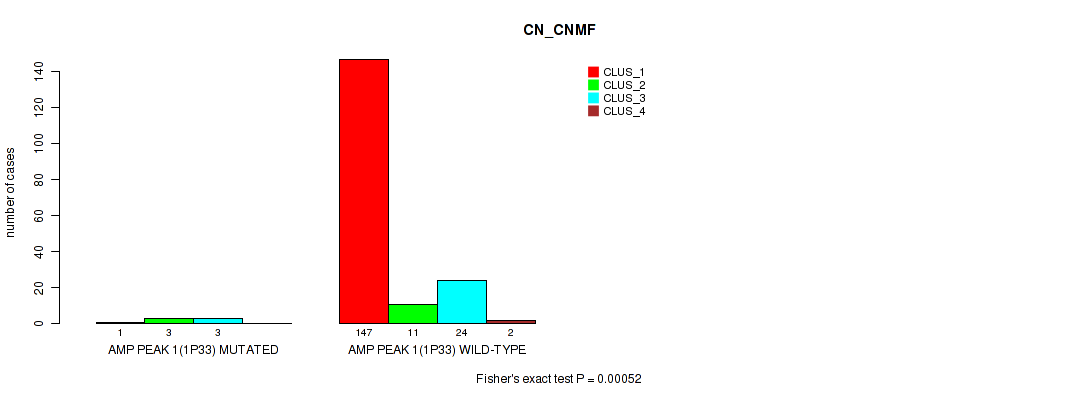
P value = 0.00194 (Fisher's exact test), Q value = 0.0065
Table S2. Gene #1: 'amp_1p33' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| AMP PEAK 1(1P33) MUTATED | 0 | 7 | 0 | 0 |
| AMP PEAK 1(1P33) WILD-TYPE | 33 | 54 | 47 | 44 |
Figure S2. Get High-res Image Gene #1: 'amp_1p33' versus Molecular Subtype #2: 'METHLYATION_CNMF'
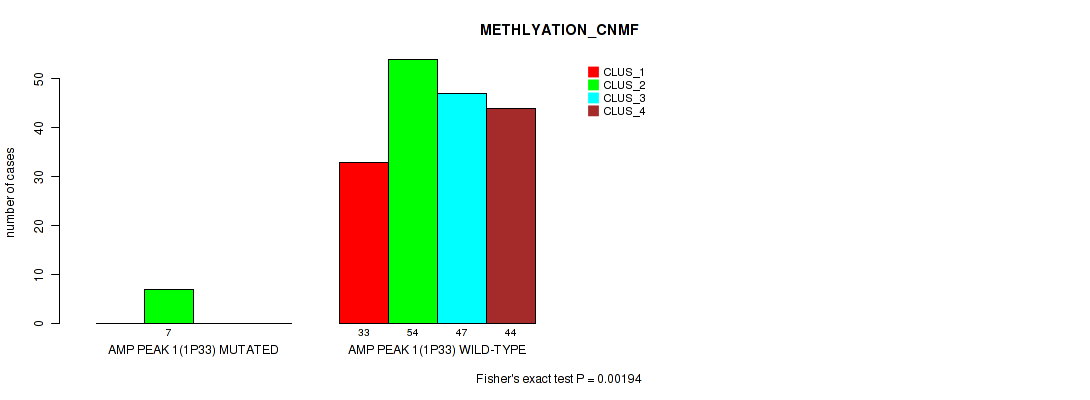
P value = 0.0097 (Fisher's exact test), Q value = 0.022
Table S3. Gene #1: 'amp_1p33' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| AMP PEAK 1(1P33) MUTATED | 0 | 3 | 0 | 4 |
| AMP PEAK 1(1P33) WILD-TYPE | 58 | 34 | 41 | 39 |
Figure S3. Get High-res Image Gene #1: 'amp_1p33' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
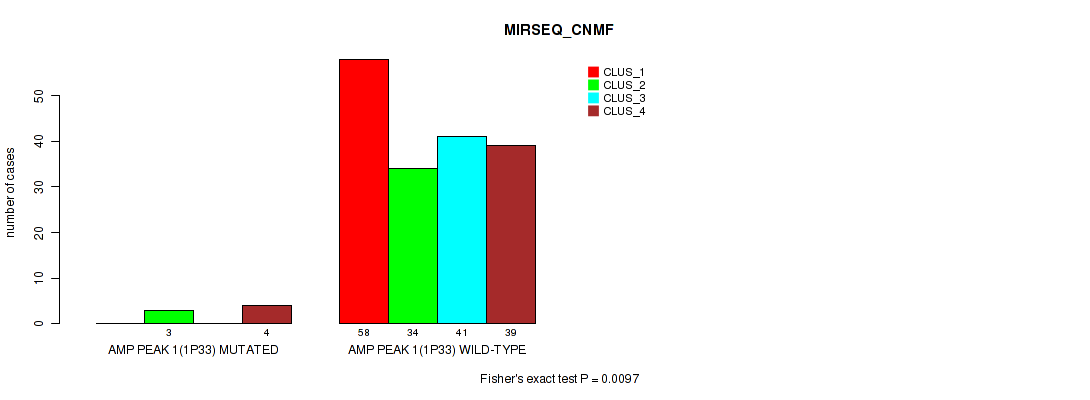
P value = 0.0485 (Fisher's exact test), Q value = 0.084
Table S4. Gene #1: 'amp_1p33' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| AMP PEAK 1(1P33) MUTATED | 0 | 3 | 4 |
| AMP PEAK 1(1P33) WILD-TYPE | 67 | 31 | 74 |
Figure S4. Get High-res Image Gene #1: 'amp_1p33' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
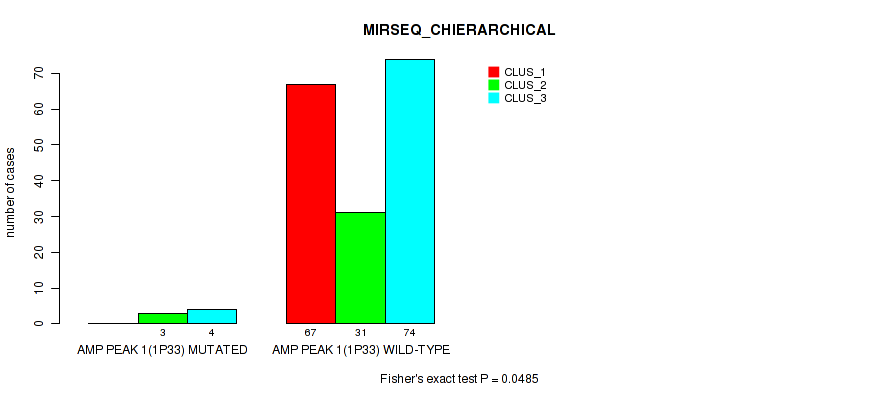
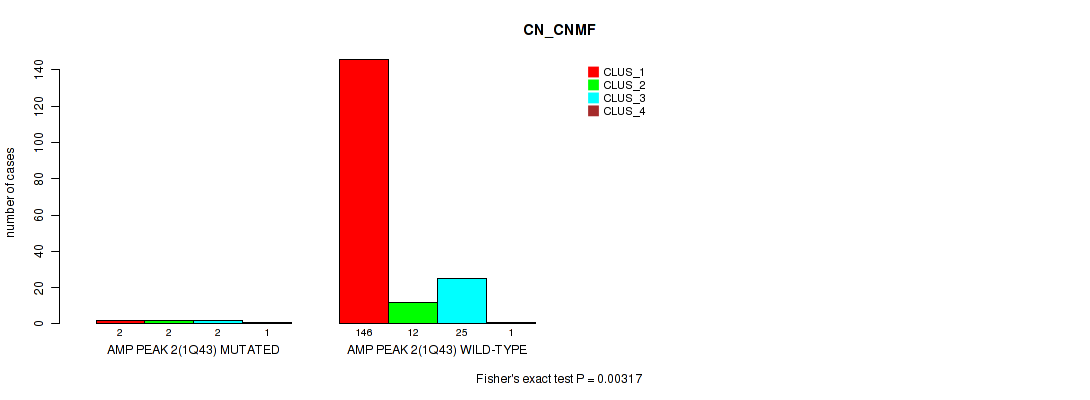
P value = 0.00317 (Fisher's exact test), Q value = 0.0093
Table S5. Gene #2: 'amp_1q43' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| AMP PEAK 2(1Q43) MUTATED | 2 | 2 | 2 | 1 |
| AMP PEAK 2(1Q43) WILD-TYPE | 146 | 12 | 25 | 1 |
Figure S5. Get High-res Image Gene #2: 'amp_1q43' versus Molecular Subtype #1: 'CN_CNMF'
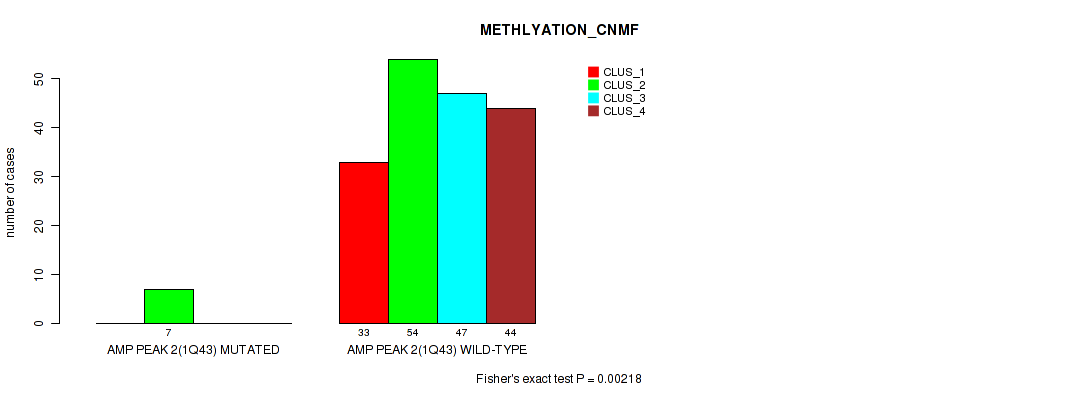
P value = 0.00218 (Fisher's exact test), Q value = 0.0069
Table S6. Gene #2: 'amp_1q43' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| AMP PEAK 2(1Q43) MUTATED | 0 | 7 | 0 | 0 |
| AMP PEAK 2(1Q43) WILD-TYPE | 33 | 54 | 47 | 44 |
Figure S6. Get High-res Image Gene #2: 'amp_1q43' versus Molecular Subtype #2: 'METHLYATION_CNMF'
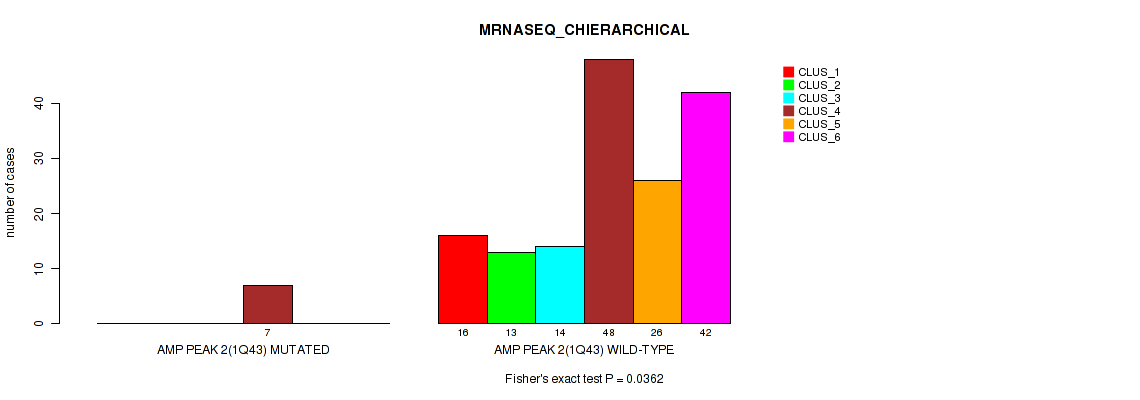
P value = 0.0362 (Fisher's exact test), Q value = 0.068
Table S7. Gene #2: 'amp_1q43' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| AMP PEAK 2(1Q43) MUTATED | 0 | 0 | 0 | 7 | 0 | 0 |
| AMP PEAK 2(1Q43) WILD-TYPE | 16 | 13 | 14 | 48 | 26 | 42 |
Figure S7. Get High-res Image Gene #2: 'amp_1q43' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
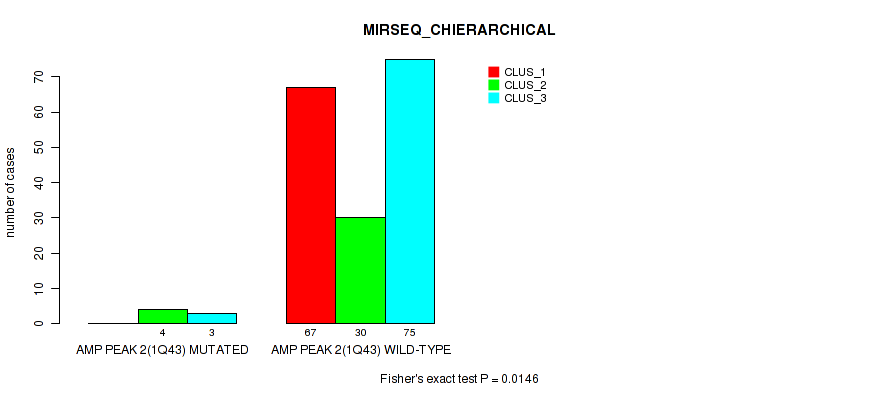
P value = 0.0146 (Fisher's exact test), Q value = 0.031
Table S8. Gene #2: 'amp_1q43' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| AMP PEAK 2(1Q43) MUTATED | 0 | 4 | 3 |
| AMP PEAK 2(1Q43) WILD-TYPE | 67 | 30 | 75 |
Figure S8. Get High-res Image Gene #2: 'amp_1q43' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
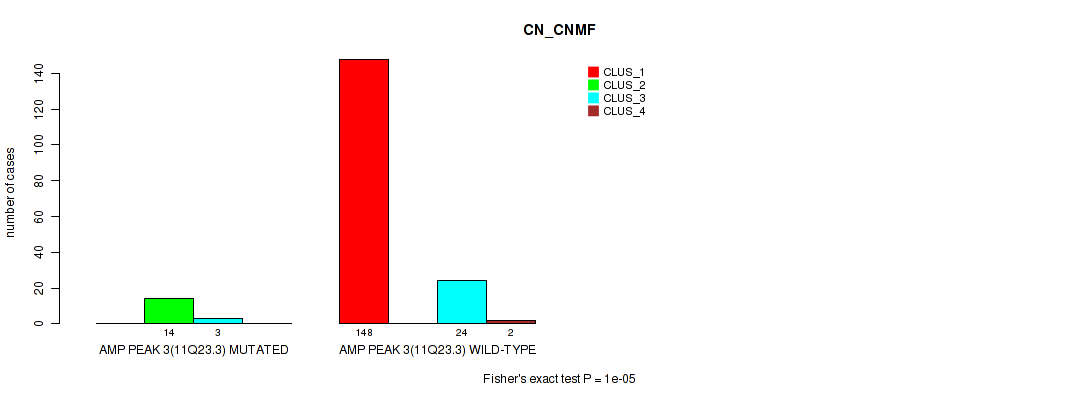
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S9. Gene #3: 'amp_11q23.3' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| AMP PEAK 3(11Q23.3) MUTATED | 0 | 14 | 3 | 0 |
| AMP PEAK 3(11Q23.3) WILD-TYPE | 148 | 0 | 24 | 2 |
Figure S9. Get High-res Image Gene #3: 'amp_11q23.3' versus Molecular Subtype #1: 'CN_CNMF'
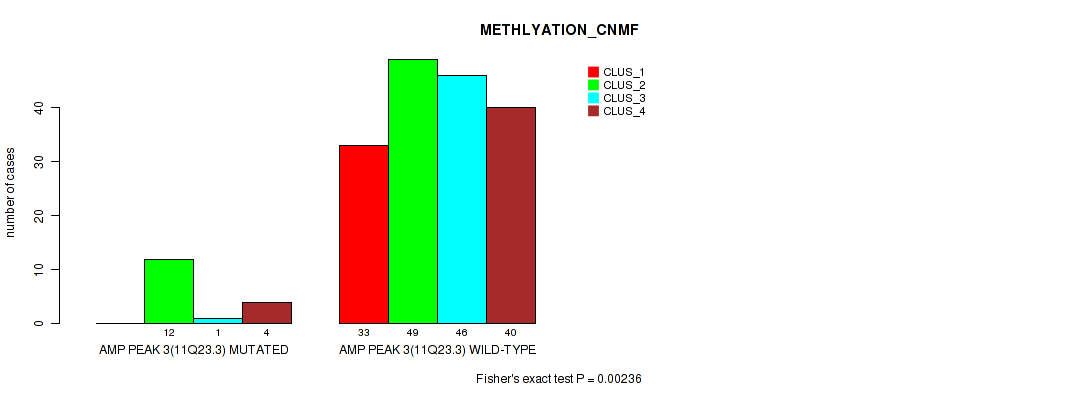
P value = 0.00236 (Fisher's exact test), Q value = 0.0073
Table S10. Gene #3: 'amp_11q23.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| AMP PEAK 3(11Q23.3) MUTATED | 0 | 12 | 1 | 4 |
| AMP PEAK 3(11Q23.3) WILD-TYPE | 33 | 49 | 46 | 40 |
Figure S10. Get High-res Image Gene #3: 'amp_11q23.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
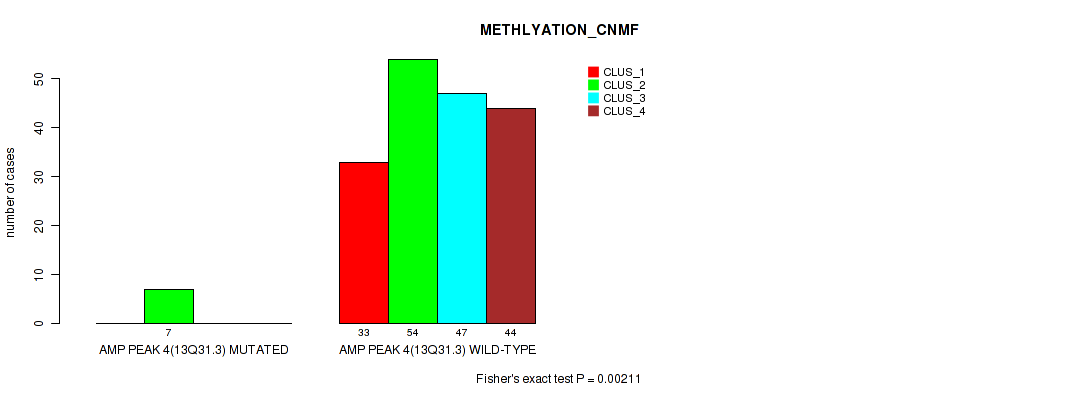
P value = 0.00211 (Fisher's exact test), Q value = 0.0068
Table S11. Gene #4: 'amp_13q31.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| AMP PEAK 4(13Q31.3) MUTATED | 0 | 7 | 0 | 0 |
| AMP PEAK 4(13Q31.3) WILD-TYPE | 33 | 54 | 47 | 44 |
Figure S11. Get High-res Image Gene #4: 'amp_13q31.3' versus Molecular Subtype #2: 'METHLYATION_CNMF'
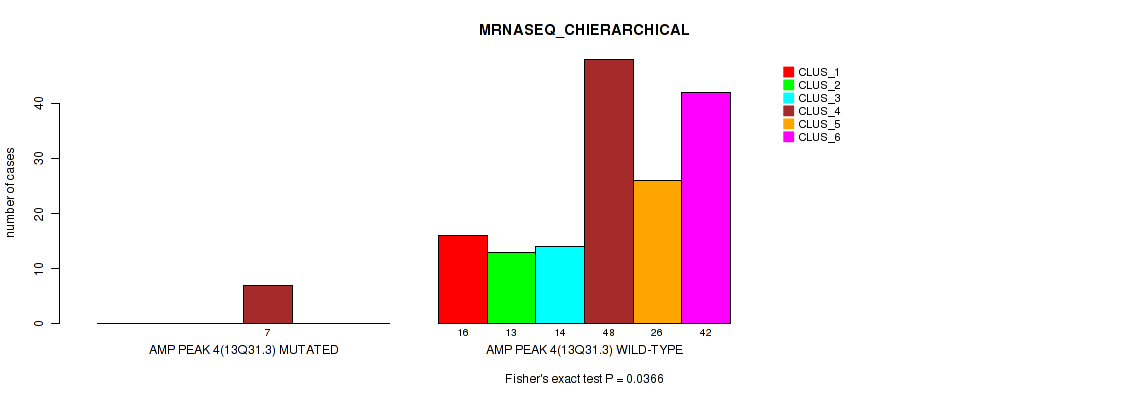
P value = 0.0366 (Fisher's exact test), Q value = 0.068
Table S12. Gene #4: 'amp_13q31.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| AMP PEAK 4(13Q31.3) MUTATED | 0 | 0 | 0 | 7 | 0 | 0 |
| AMP PEAK 4(13Q31.3) WILD-TYPE | 16 | 13 | 14 | 48 | 26 | 42 |
Figure S12. Get High-res Image Gene #4: 'amp_13q31.3' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
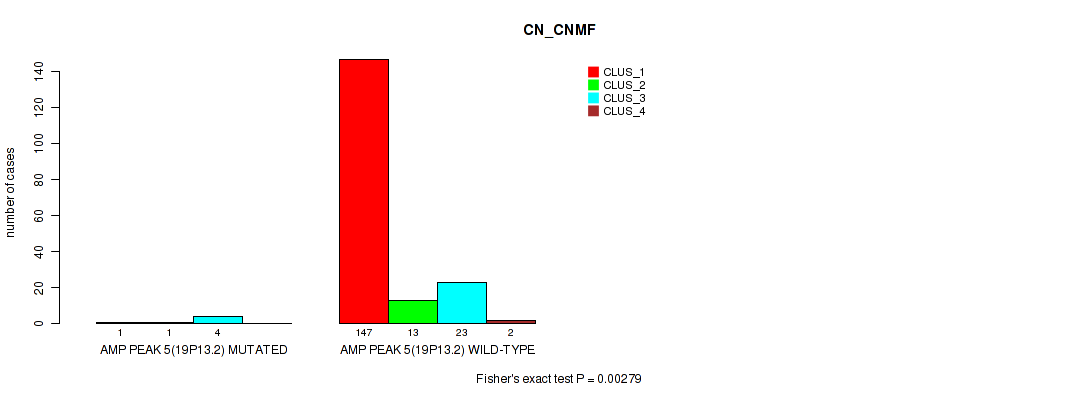
P value = 0.00279 (Fisher's exact test), Q value = 0.0084
Table S13. Gene #5: 'amp_19p13.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| AMP PEAK 5(19P13.2) MUTATED | 1 | 1 | 4 | 0 |
| AMP PEAK 5(19P13.2) WILD-TYPE | 147 | 13 | 23 | 2 |
Figure S13. Get High-res Image Gene #5: 'amp_19p13.2' versus Molecular Subtype #1: 'CN_CNMF'
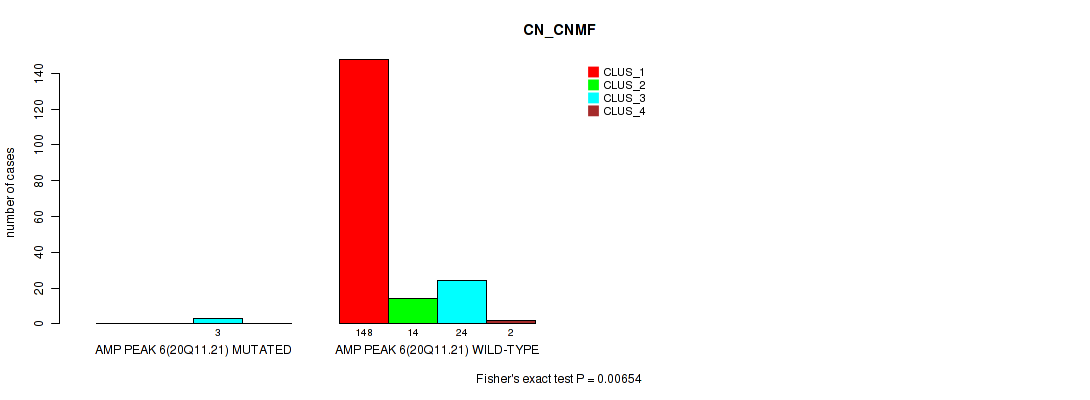
P value = 0.00654 (Fisher's exact test), Q value = 0.016
Table S14. Gene #6: 'amp_20q11.21' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| AMP PEAK 6(20Q11.21) MUTATED | 0 | 0 | 3 | 0 |
| AMP PEAK 6(20Q11.21) WILD-TYPE | 148 | 14 | 24 | 2 |
Figure S14. Get High-res Image Gene #6: 'amp_20q11.21' versus Molecular Subtype #1: 'CN_CNMF'
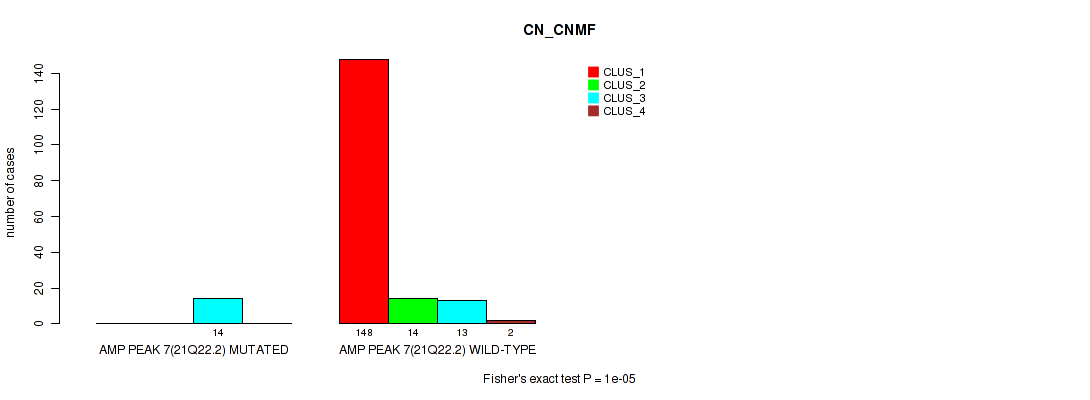
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S15. Gene #7: 'amp_21q22.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| AMP PEAK 7(21Q22.2) MUTATED | 0 | 0 | 14 | 0 |
| AMP PEAK 7(21Q22.2) WILD-TYPE | 148 | 14 | 13 | 2 |
Figure S15. Get High-res Image Gene #7: 'amp_21q22.2' versus Molecular Subtype #1: 'CN_CNMF'
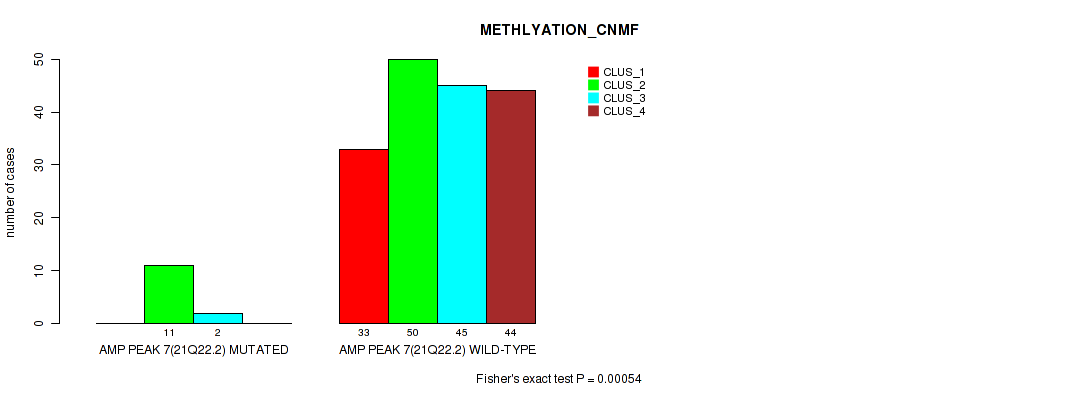
P value = 0.00054 (Fisher's exact test), Q value = 0.0031
Table S16. Gene #7: 'amp_21q22.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| AMP PEAK 7(21Q22.2) MUTATED | 0 | 11 | 2 | 0 |
| AMP PEAK 7(21Q22.2) WILD-TYPE | 33 | 50 | 45 | 44 |
Figure S16. Get High-res Image Gene #7: 'amp_21q22.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
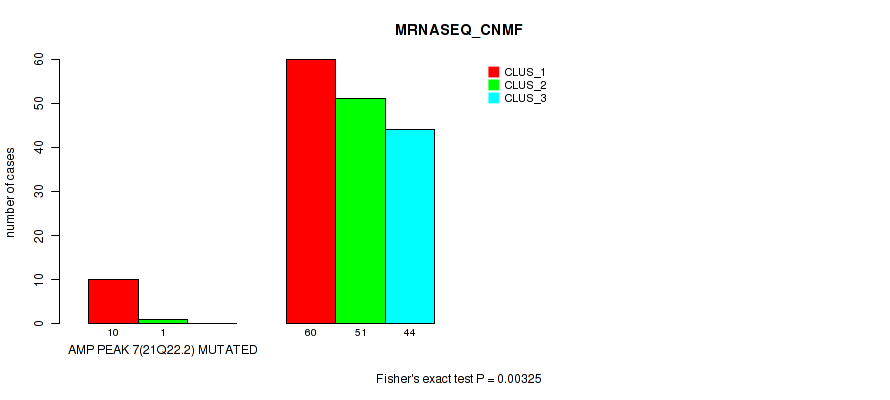
P value = 0.00325 (Fisher's exact test), Q value = 0.0093
Table S17. Gene #7: 'amp_21q22.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 52 | 44 |
| AMP PEAK 7(21Q22.2) MUTATED | 10 | 1 | 0 |
| AMP PEAK 7(21Q22.2) WILD-TYPE | 60 | 51 | 44 |
Figure S17. Get High-res Image Gene #7: 'amp_21q22.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
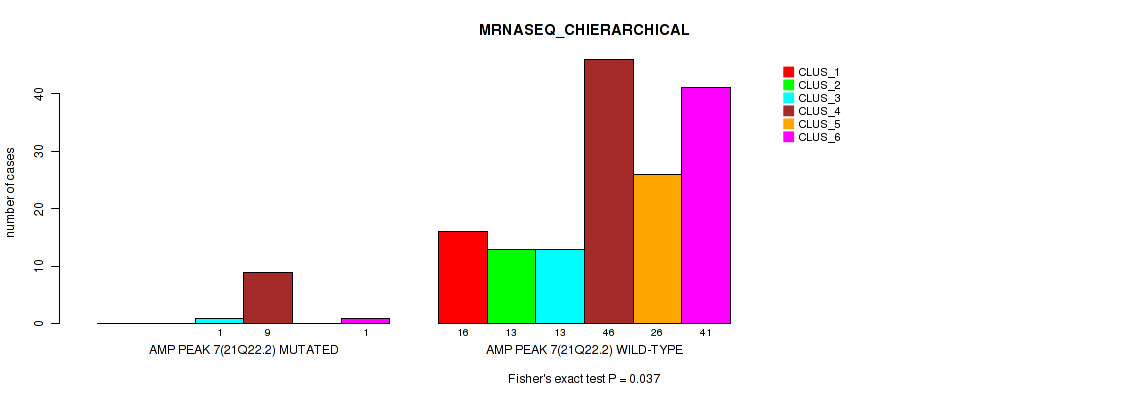
P value = 0.037 (Fisher's exact test), Q value = 0.068
Table S18. Gene #7: 'amp_21q22.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| AMP PEAK 7(21Q22.2) MUTATED | 0 | 0 | 1 | 9 | 0 | 1 |
| AMP PEAK 7(21Q22.2) WILD-TYPE | 16 | 13 | 13 | 46 | 26 | 41 |
Figure S18. Get High-res Image Gene #7: 'amp_21q22.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
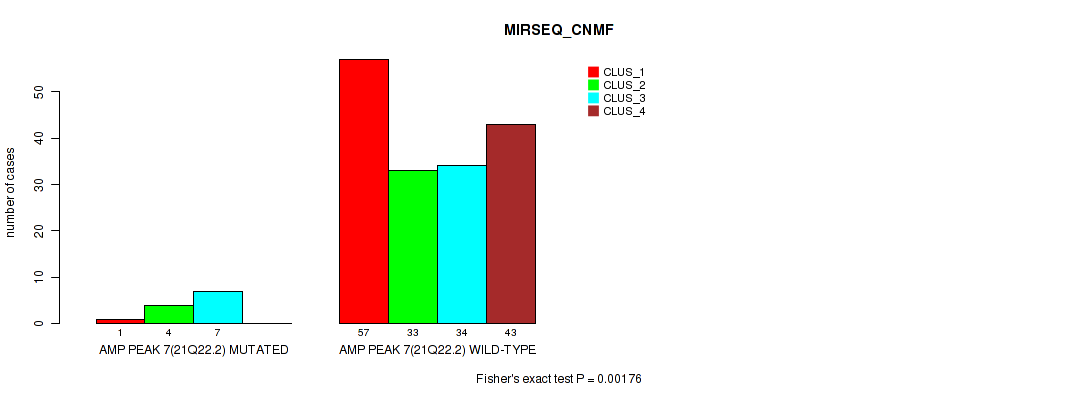
P value = 0.00176 (Fisher's exact test), Q value = 0.0061
Table S19. Gene #7: 'amp_21q22.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| AMP PEAK 7(21Q22.2) MUTATED | 1 | 4 | 7 | 0 |
| AMP PEAK 7(21Q22.2) WILD-TYPE | 57 | 33 | 34 | 43 |
Figure S19. Get High-res Image Gene #7: 'amp_21q22.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
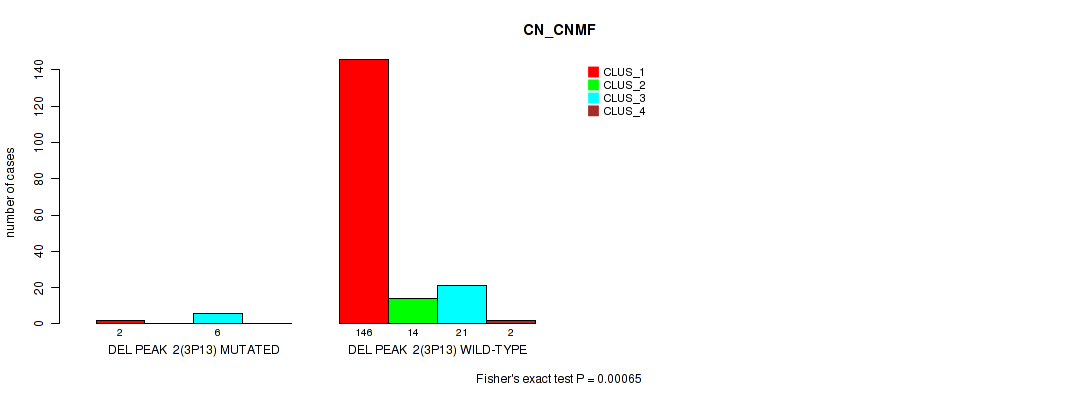
P value = 0.00065 (Fisher's exact test), Q value = 0.0032
Table S20. Gene #8: 'del_3p13' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 2(3P13) MUTATED | 2 | 0 | 6 | 0 |
| DEL PEAK 2(3P13) WILD-TYPE | 146 | 14 | 21 | 2 |
Figure S20. Get High-res Image Gene #8: 'del_3p13' versus Molecular Subtype #1: 'CN_CNMF'
P value = 0.0469 (Fisher's exact test), Q value = 0.084
Table S21. Gene #8: 'del_3p13' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| DEL PEAK 2(3P13) MUTATED | 0 | 6 | 2 | 0 |
| DEL PEAK 2(3P13) WILD-TYPE | 33 | 55 | 45 | 44 |
Figure S21. Get High-res Image Gene #8: 'del_3p13' versus Molecular Subtype #2: 'METHLYATION_CNMF'
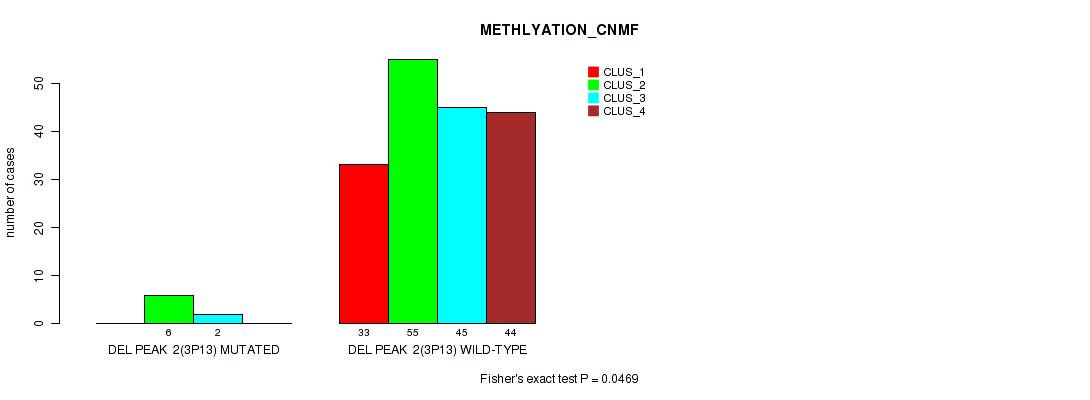
P value = 0.0367 (Fisher's exact test), Q value = 0.068
Table S22. Gene #8: 'del_3p13' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| DEL PEAK 2(3P13) MUTATED | 0 | 4 | 2 | 1 |
| DEL PEAK 2(3P13) WILD-TYPE | 58 | 33 | 39 | 42 |
Figure S22. Get High-res Image Gene #8: 'del_3p13' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
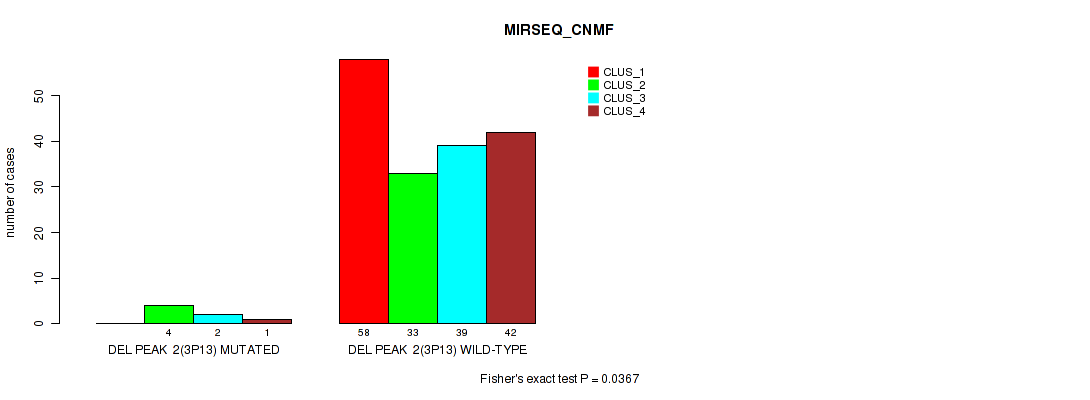
P value = 0.0485 (Fisher's exact test), Q value = 0.084
Table S23. Gene #8: 'del_3p13' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 2(3P13) MUTATED | 0 | 3 | 4 |
| DEL PEAK 2(3P13) WILD-TYPE | 67 | 31 | 74 |
Figure S23. Get High-res Image Gene #8: 'del_3p13' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
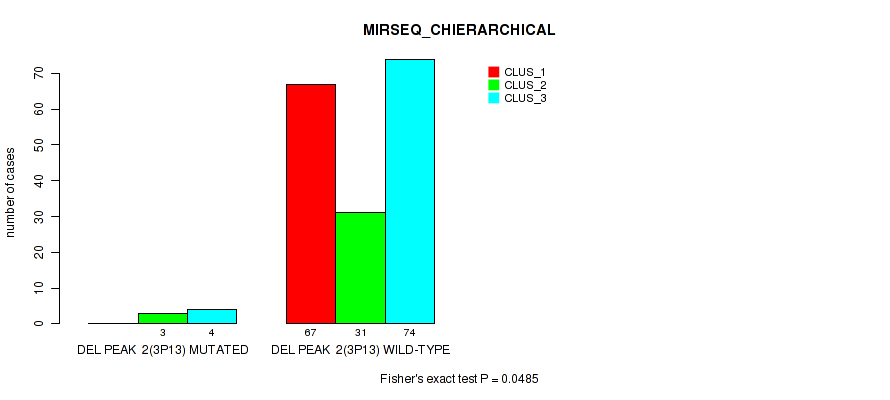
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S24. Gene #9: 'del_5q31.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 3(5Q31.2) MUTATED | 1 | 1 | 15 | 1 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 147 | 13 | 12 | 1 |
Figure S24. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #1: 'CN_CNMF'
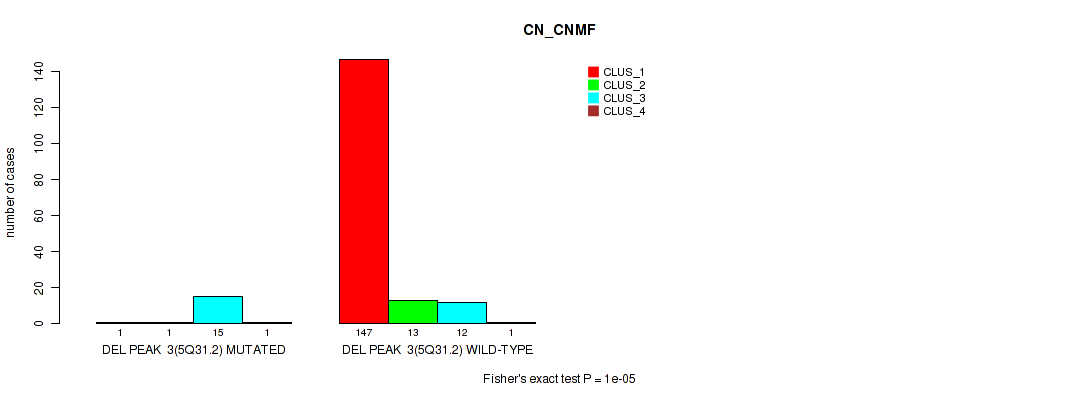
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S25. Gene #9: 'del_5q31.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| DEL PEAK 3(5Q31.2) MUTATED | 0 | 15 | 3 | 0 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 33 | 46 | 44 | 44 |
Figure S25. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
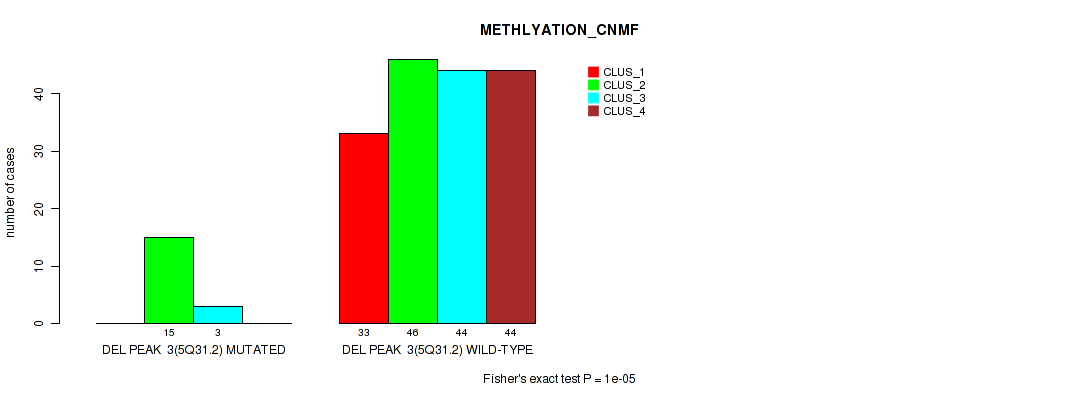
P value = 0.0055 (Fisher's exact test), Q value = 0.014
Table S26. Gene #9: 'del_5q31.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 52 | 44 |
| DEL PEAK 3(5Q31.2) MUTATED | 13 | 2 | 1 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 57 | 50 | 43 |
Figure S26. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
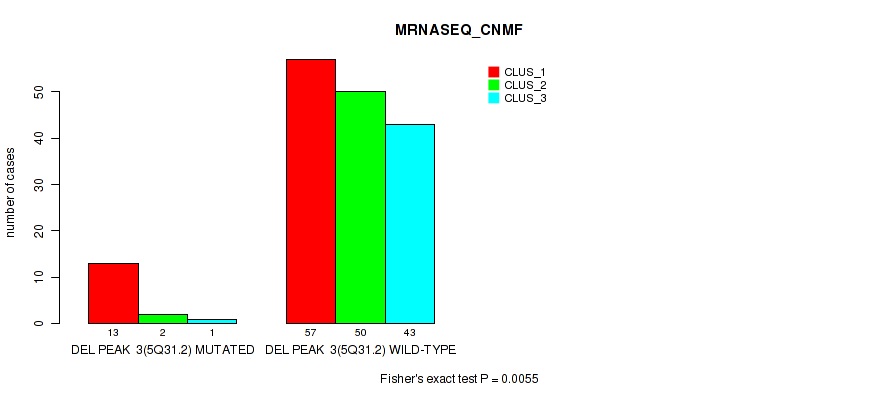
P value = 0.00061 (Fisher's exact test), Q value = 0.0032
Table S27. Gene #9: 'del_5q31.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 3(5Q31.2) MUTATED | 0 | 0 | 0 | 14 | 0 | 2 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 16 | 13 | 14 | 41 | 26 | 40 |
Figure S27. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
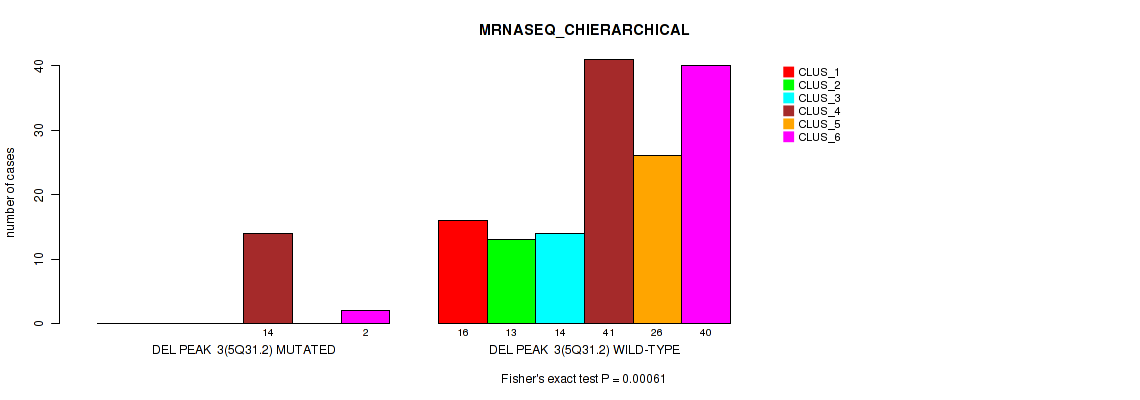
P value = 0.00102 (Fisher's exact test), Q value = 0.0044
Table S28. Gene #9: 'del_5q31.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| DEL PEAK 3(5Q31.2) MUTATED | 1 | 10 | 3 | 3 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 57 | 27 | 38 | 40 |
Figure S28. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
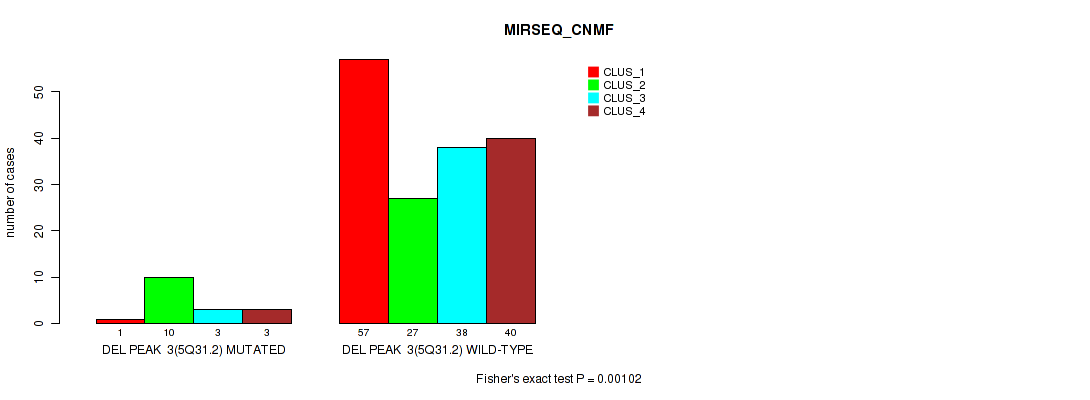
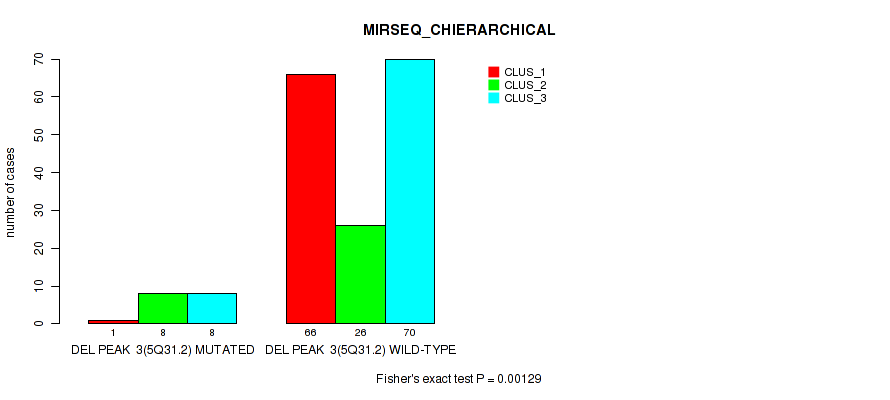
P value = 0.00129 (Fisher's exact test), Q value = 0.0052
Table S29. Gene #9: 'del_5q31.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 3(5Q31.2) MUTATED | 1 | 8 | 8 |
| DEL PEAK 3(5Q31.2) WILD-TYPE | 66 | 26 | 70 |
Figure S29. Get High-res Image Gene #9: 'del_5q31.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
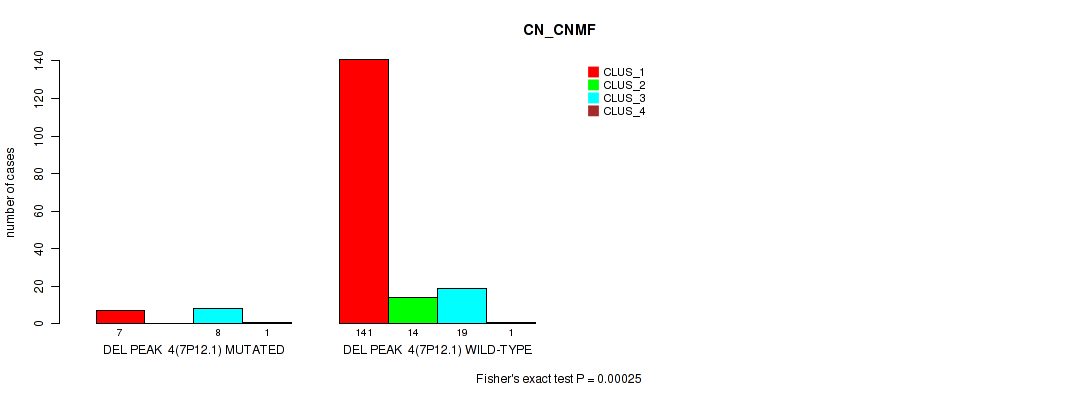
P value = 0.00025 (Fisher's exact test), Q value = 0.0018
Table S30. Gene #10: 'del_7p12.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 4(7P12.1) MUTATED | 7 | 0 | 8 | 1 |
| DEL PEAK 4(7P12.1) WILD-TYPE | 141 | 14 | 19 | 1 |
Figure S30. Get High-res Image Gene #10: 'del_7p12.1' versus Molecular Subtype #1: 'CN_CNMF'
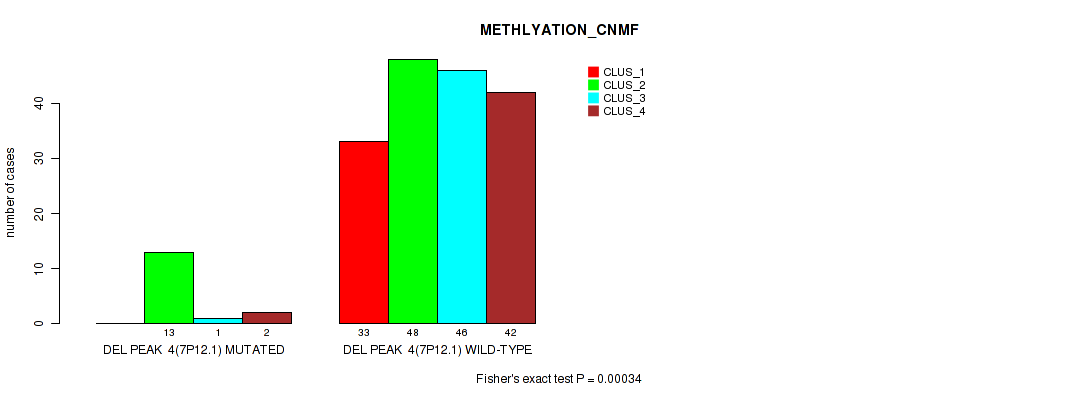
P value = 0.00034 (Fisher's exact test), Q value = 0.0021
Table S31. Gene #10: 'del_7p12.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| DEL PEAK 4(7P12.1) MUTATED | 0 | 13 | 1 | 2 |
| DEL PEAK 4(7P12.1) WILD-TYPE | 33 | 48 | 46 | 42 |
Figure S31. Get High-res Image Gene #10: 'del_7p12.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
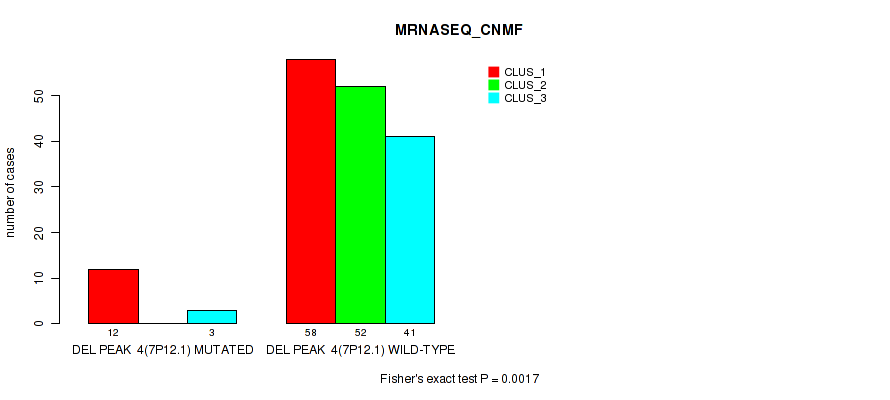
P value = 0.0017 (Fisher's exact test), Q value = 0.0061
Table S32. Gene #10: 'del_7p12.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 52 | 44 |
| DEL PEAK 4(7P12.1) MUTATED | 12 | 0 | 3 |
| DEL PEAK 4(7P12.1) WILD-TYPE | 58 | 52 | 41 |
Figure S32. Get High-res Image Gene #10: 'del_7p12.1' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
P value = 2e-05 (Fisher's exact test), Q value = 2e-04
Table S33. Gene #10: 'del_7p12.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 4(7P12.1) MUTATED | 0 | 0 | 0 | 15 | 0 | 0 |
| DEL PEAK 4(7P12.1) WILD-TYPE | 16 | 13 | 14 | 40 | 26 | 42 |
Figure S33. Get High-res Image Gene #10: 'del_7p12.1' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
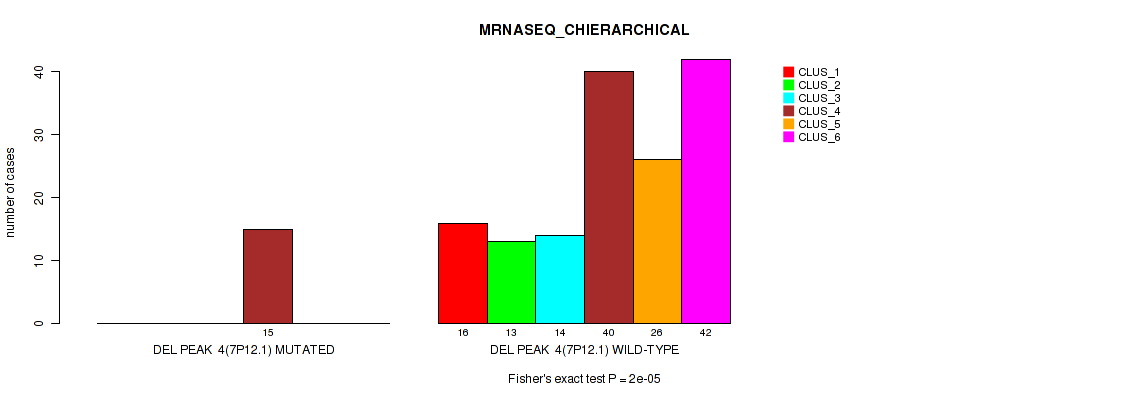
P value = 0.00595 (Fisher's exact test), Q value = 0.015
Table S34. Gene #10: 'del_7p12.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| DEL PEAK 4(7P12.1) MUTATED | 1 | 8 | 2 | 5 |
| DEL PEAK 4(7P12.1) WILD-TYPE | 57 | 29 | 39 | 38 |
Figure S34. Get High-res Image Gene #10: 'del_7p12.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
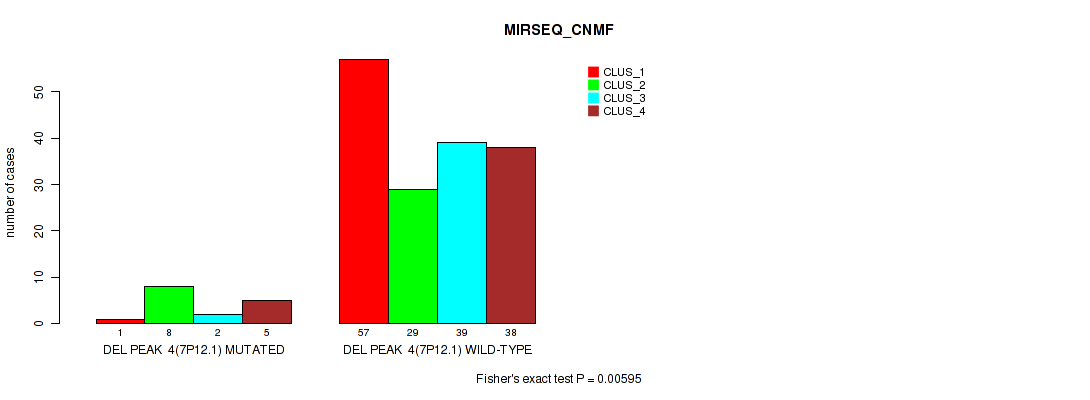
P value = 0.0076 (Fisher's exact test), Q value = 0.018
Table S35. Gene #10: 'del_7p12.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 4(7P12.1) MUTATED | 1 | 6 | 9 |
| DEL PEAK 4(7P12.1) WILD-TYPE | 66 | 28 | 69 |
Figure S35. Get High-res Image Gene #10: 'del_7p12.1' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
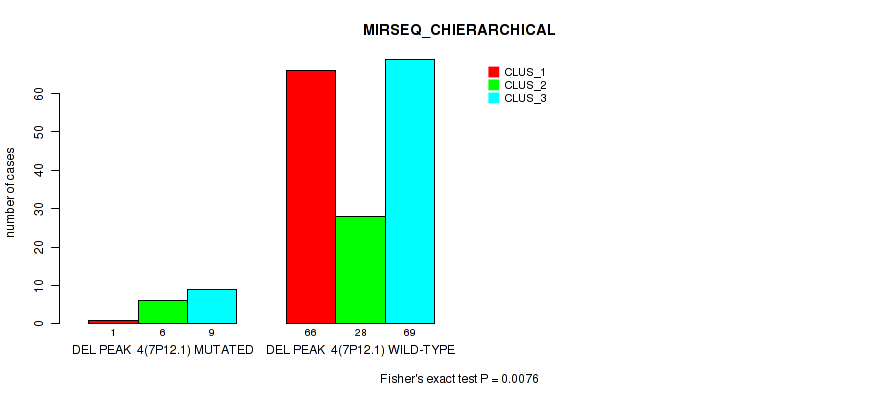
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S36. Gene #11: 'del_7q34' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 5(7Q34) MUTATED | 9 | 2 | 11 | 2 |
| DEL PEAK 5(7Q34) WILD-TYPE | 139 | 12 | 16 | 0 |
Figure S36. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #1: 'CN_CNMF'
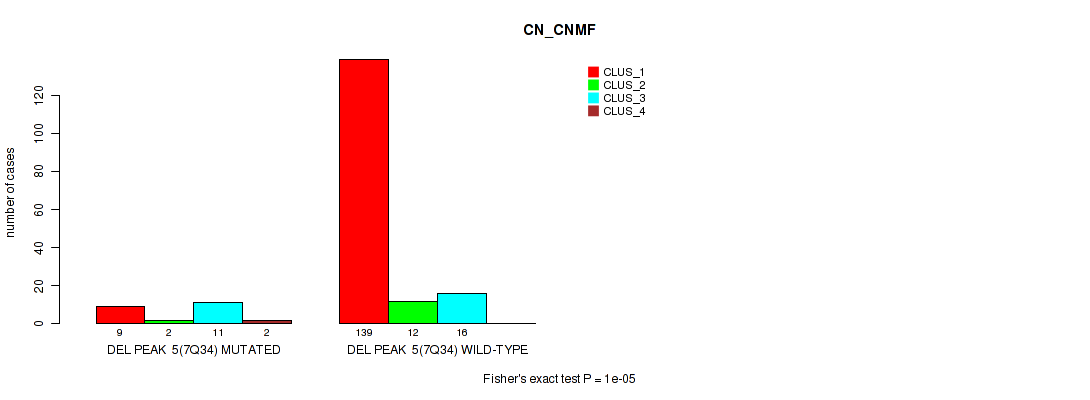
P value = 0.00011 (Fisher's exact test), Q value = 0.00094
Table S37. Gene #11: 'del_7q34' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| DEL PEAK 5(7Q34) MUTATED | 1 | 18 | 2 | 3 |
| DEL PEAK 5(7Q34) WILD-TYPE | 32 | 43 | 45 | 41 |
Figure S37. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #2: 'METHLYATION_CNMF'
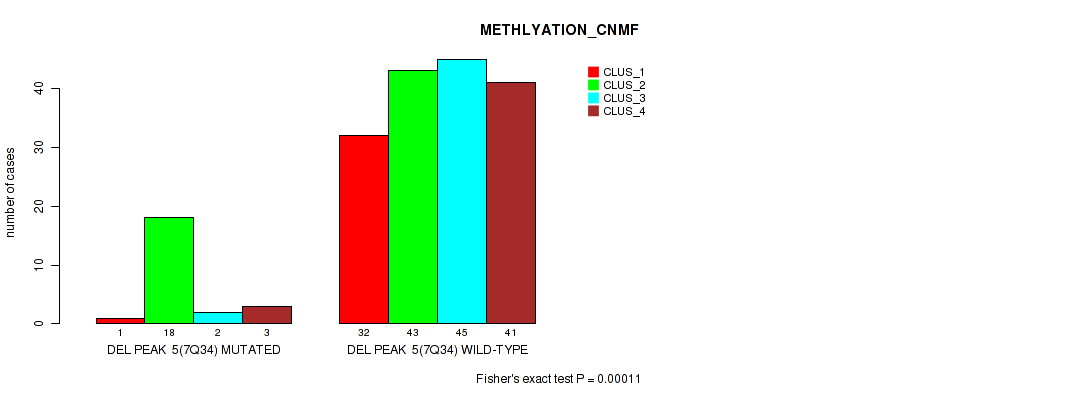
P value = 0.0228 (Fisher's exact test), Q value = 0.046
Table S38. Gene #11: 'del_7q34' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 52 | 44 |
| DEL PEAK 5(7Q34) MUTATED | 14 | 2 | 4 |
| DEL PEAK 5(7Q34) WILD-TYPE | 56 | 50 | 40 |
Figure S38. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
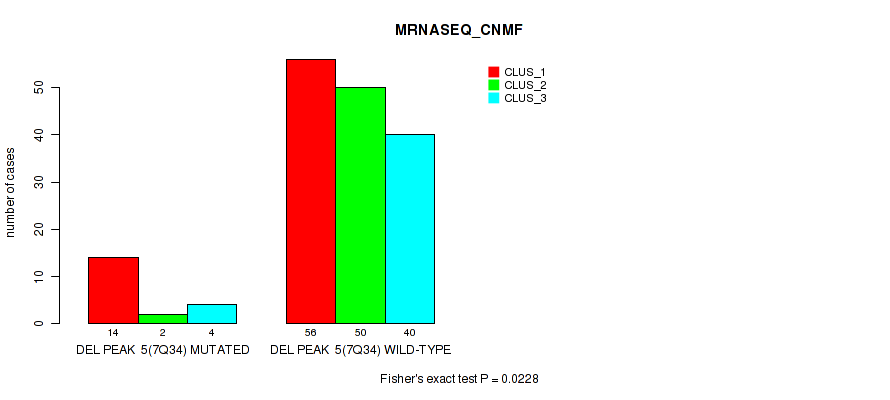
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S39. Gene #11: 'del_7q34' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 5(7Q34) MUTATED | 1 | 1 | 0 | 18 | 0 | 0 |
| DEL PEAK 5(7Q34) WILD-TYPE | 15 | 12 | 14 | 37 | 26 | 42 |
Figure S39. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
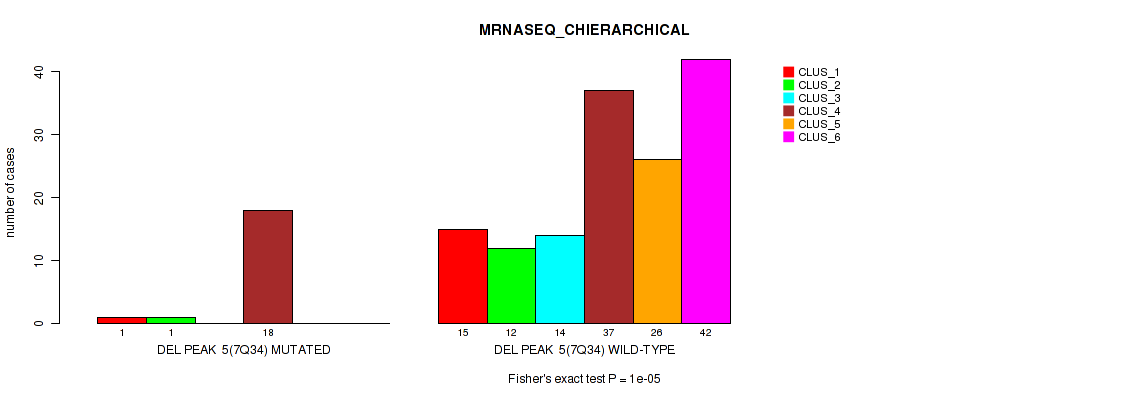
P value = 0.00079 (Fisher's exact test), Q value = 0.0036
Table S40. Gene #11: 'del_7q34' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| DEL PEAK 5(7Q34) MUTATED | 2 | 12 | 5 | 4 |
| DEL PEAK 5(7Q34) WILD-TYPE | 56 | 25 | 36 | 39 |
Figure S40. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
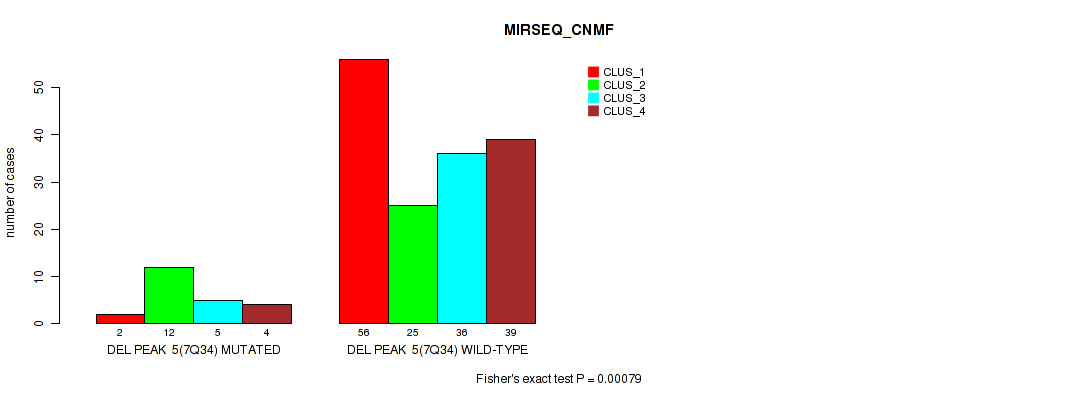
P value = 0.00061 (Fisher's exact test), Q value = 0.0032
Table S41. Gene #11: 'del_7q34' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 5(7Q34) MUTATED | 2 | 10 | 11 |
| DEL PEAK 5(7Q34) WILD-TYPE | 65 | 24 | 67 |
Figure S41. Get High-res Image Gene #11: 'del_7q34' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
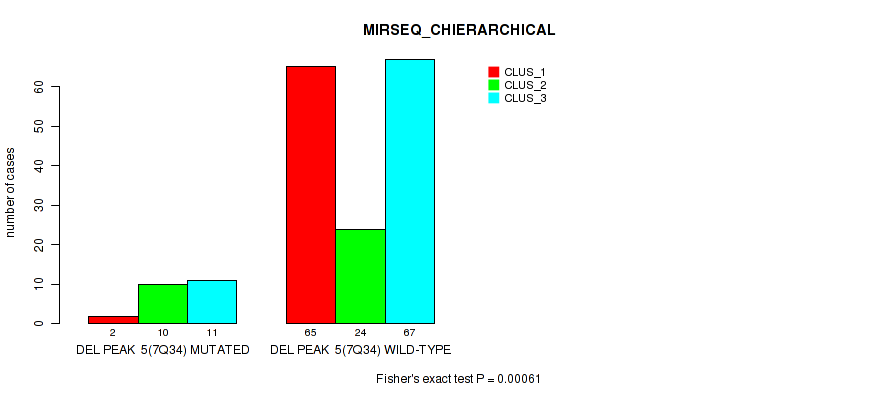
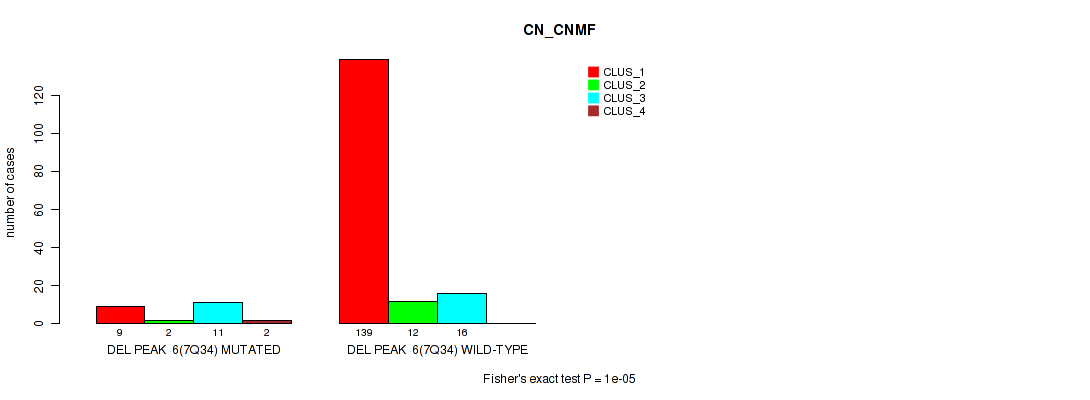
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S42. Gene #12: 'del_7q34' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 6(7Q34) MUTATED | 9 | 2 | 11 | 2 |
| DEL PEAK 6(7Q34) WILD-TYPE | 139 | 12 | 16 | 0 |
Figure S42. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #1: 'CN_CNMF'
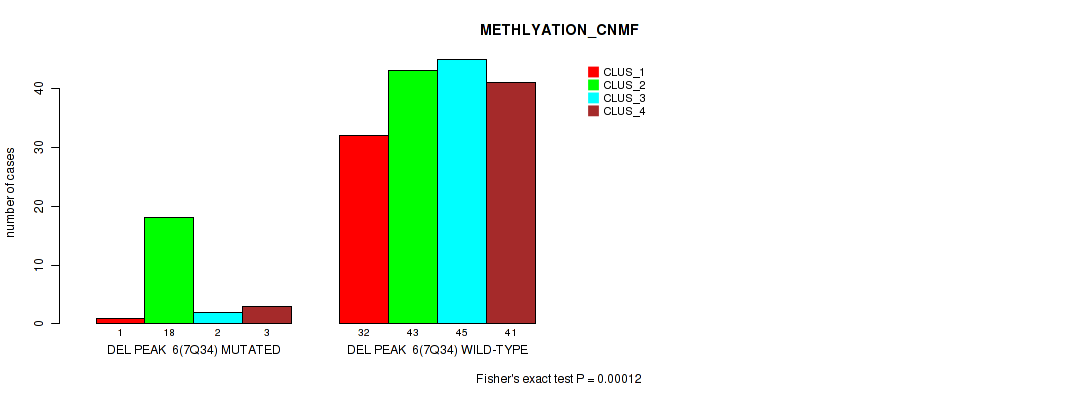
P value = 0.00012 (Fisher's exact test), Q value = 0.00096
Table S43. Gene #12: 'del_7q34' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| DEL PEAK 6(7Q34) MUTATED | 1 | 18 | 2 | 3 |
| DEL PEAK 6(7Q34) WILD-TYPE | 32 | 43 | 45 | 41 |
Figure S43. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #2: 'METHLYATION_CNMF'
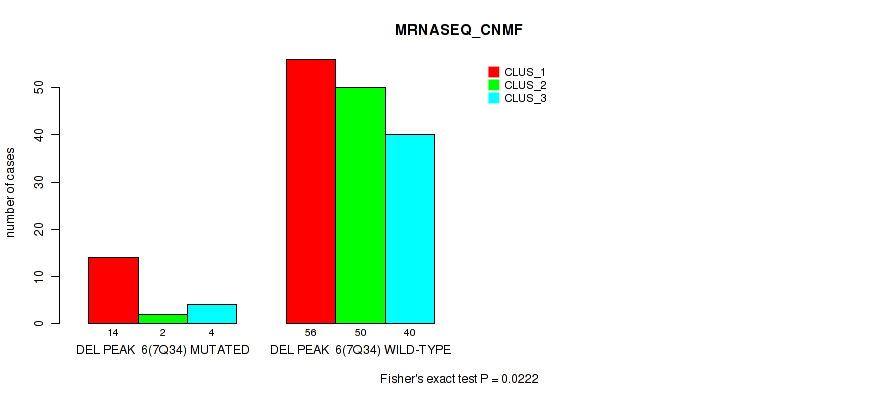
P value = 0.0222 (Fisher's exact test), Q value = 0.046
Table S44. Gene #12: 'del_7q34' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 52 | 44 |
| DEL PEAK 6(7Q34) MUTATED | 14 | 2 | 4 |
| DEL PEAK 6(7Q34) WILD-TYPE | 56 | 50 | 40 |
Figure S44. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S45. Gene #12: 'del_7q34' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 6(7Q34) MUTATED | 1 | 1 | 0 | 18 | 0 | 0 |
| DEL PEAK 6(7Q34) WILD-TYPE | 15 | 12 | 14 | 37 | 26 | 42 |
Figure S45. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
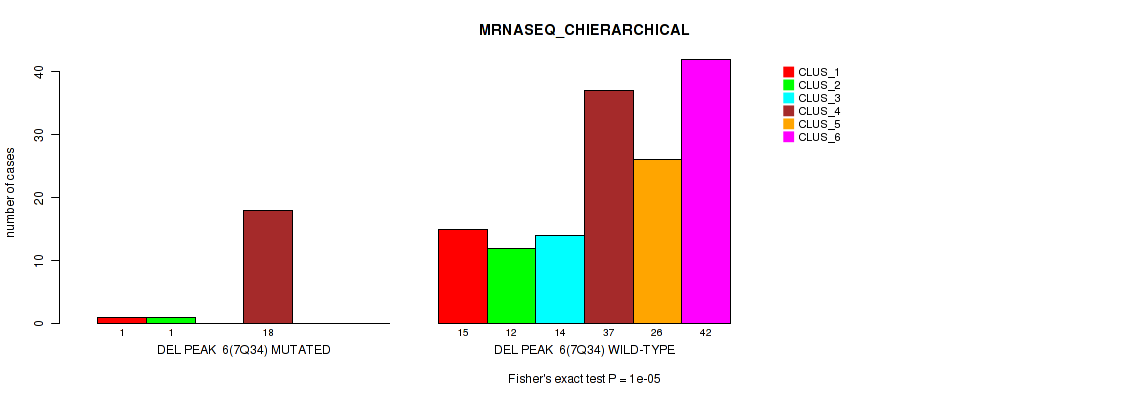
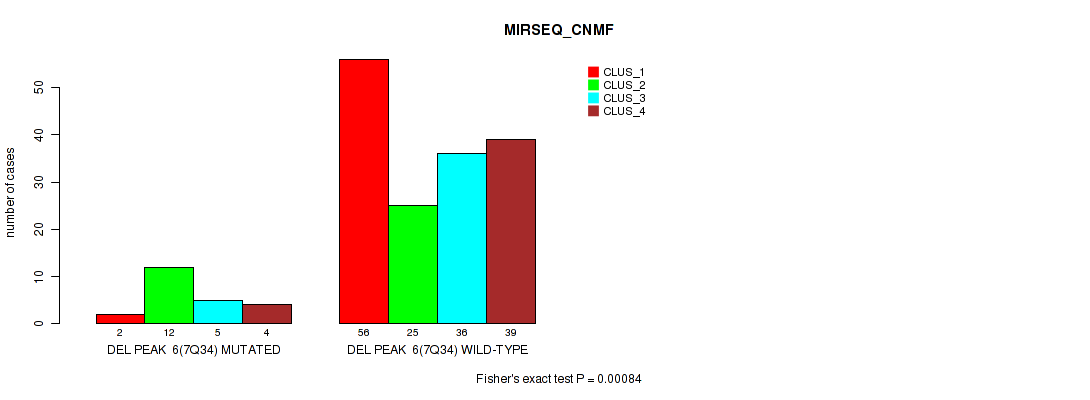
P value = 0.00084 (Fisher's exact test), Q value = 0.0037
Table S46. Gene #12: 'del_7q34' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| DEL PEAK 6(7Q34) MUTATED | 2 | 12 | 5 | 4 |
| DEL PEAK 6(7Q34) WILD-TYPE | 56 | 25 | 36 | 39 |
Figure S46. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
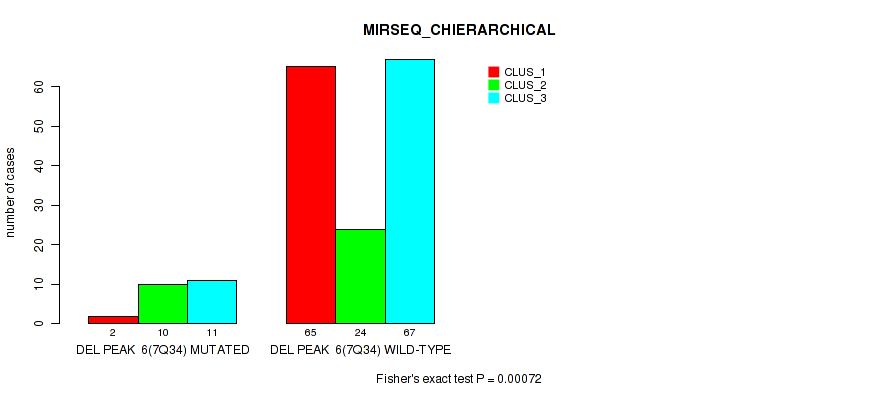
P value = 0.00072 (Fisher's exact test), Q value = 0.0035
Table S47. Gene #12: 'del_7q34' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 6(7Q34) MUTATED | 2 | 10 | 11 |
| DEL PEAK 6(7Q34) WILD-TYPE | 65 | 24 | 67 |
Figure S47. Get High-res Image Gene #12: 'del_7q34' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
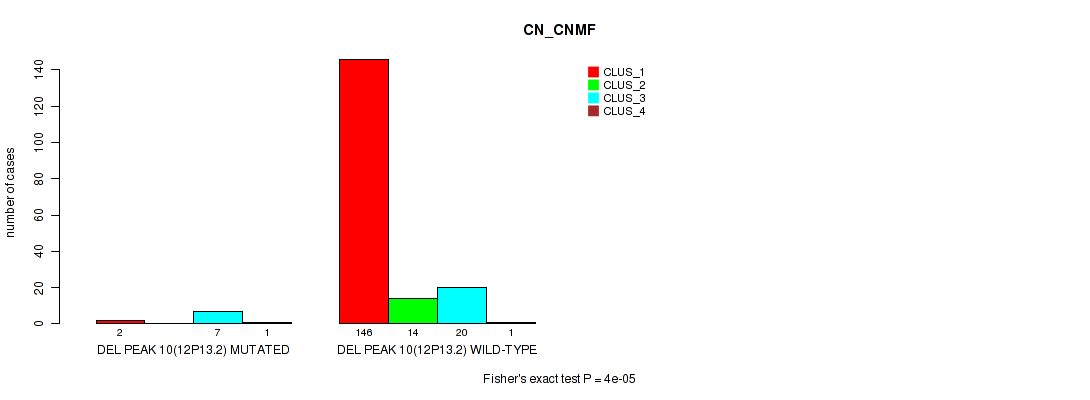
P value = 4e-05 (Fisher's exact test), Q value = 0.00037
Table S48. Gene #14: 'del_12p13.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 10(12P13.2) MUTATED | 2 | 0 | 7 | 1 |
| DEL PEAK 10(12P13.2) WILD-TYPE | 146 | 14 | 20 | 1 |
Figure S48. Get High-res Image Gene #14: 'del_12p13.2' versus Molecular Subtype #1: 'CN_CNMF'
P value = 0.00151 (Fisher's exact test), Q value = 0.0057
Table S49. Gene #14: 'del_12p13.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| DEL PEAK 10(12P13.2) MUTATED | 0 | 9 | 1 | 0 |
| DEL PEAK 10(12P13.2) WILD-TYPE | 33 | 52 | 46 | 44 |
Figure S49. Get High-res Image Gene #14: 'del_12p13.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
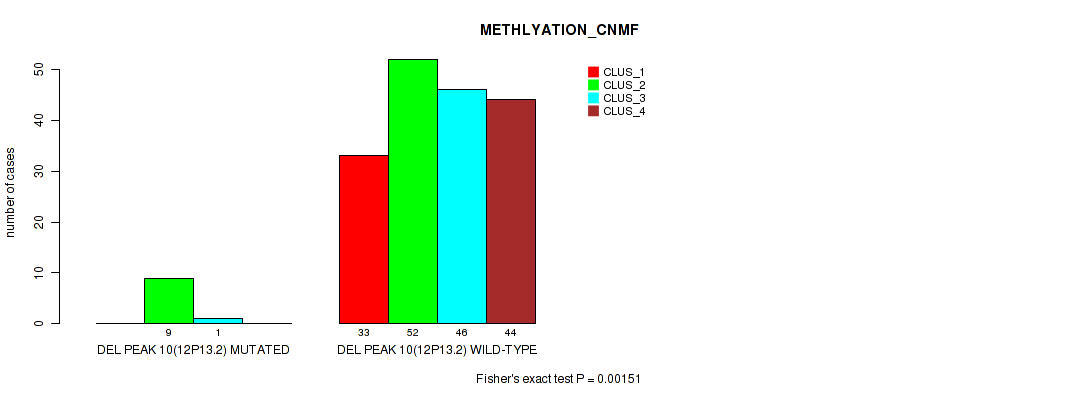
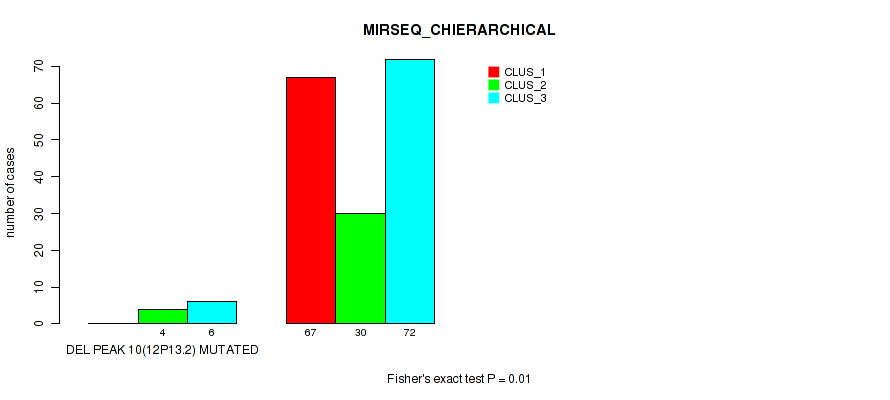
P value = 0.01 (Fisher's exact test), Q value = 0.022
Table S50. Gene #14: 'del_12p13.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 10(12P13.2) MUTATED | 0 | 4 | 6 |
| DEL PEAK 10(12P13.2) WILD-TYPE | 67 | 30 | 72 |
Figure S50. Get High-res Image Gene #14: 'del_12p13.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
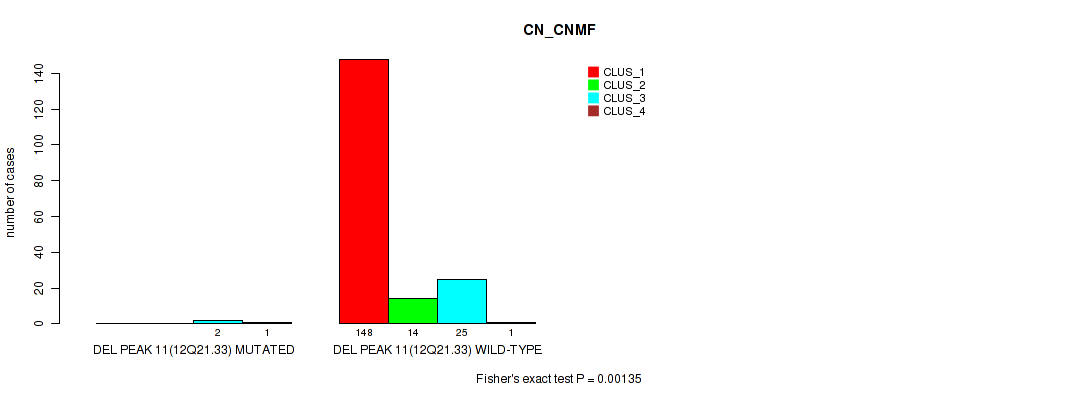
P value = 0.00135 (Fisher's exact test), Q value = 0.0052
Table S51. Gene #15: 'del_12q21.33' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 11(12Q21.33) MUTATED | 0 | 0 | 2 | 1 |
| DEL PEAK 11(12Q21.33) WILD-TYPE | 148 | 14 | 25 | 1 |
Figure S51. Get High-res Image Gene #15: 'del_12q21.33' versus Molecular Subtype #1: 'CN_CNMF'
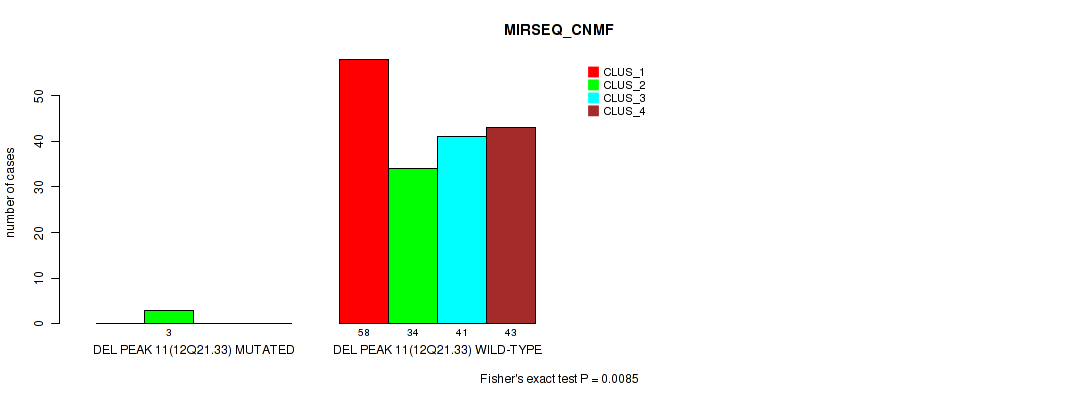
P value = 0.0085 (Fisher's exact test), Q value = 0.02
Table S52. Gene #15: 'del_12q21.33' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| DEL PEAK 11(12Q21.33) MUTATED | 0 | 3 | 0 | 0 |
| DEL PEAK 11(12Q21.33) WILD-TYPE | 58 | 34 | 41 | 43 |
Figure S52. Get High-res Image Gene #15: 'del_12q21.33' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
P value = 0.00682 (Fisher's exact test), Q value = 0.017
Table S53. Gene #15: 'del_12q21.33' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 11(12Q21.33) MUTATED | 0 | 3 | 0 |
| DEL PEAK 11(12Q21.33) WILD-TYPE | 67 | 31 | 78 |
Figure S53. Get High-res Image Gene #15: 'del_12q21.33' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
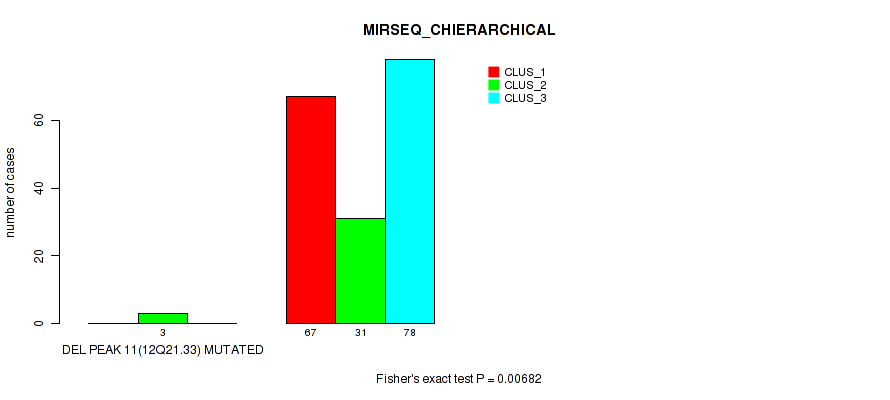
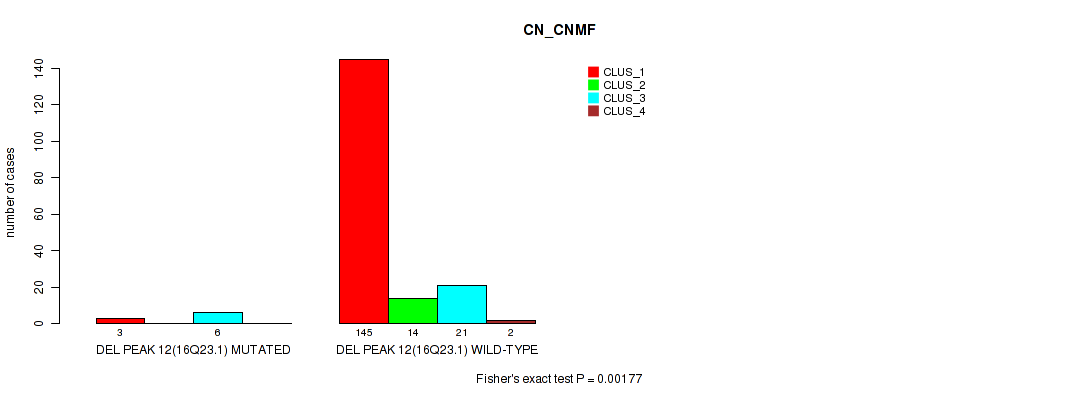
P value = 0.00177 (Fisher's exact test), Q value = 0.0061
Table S54. Gene #16: 'del_16q23.1' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 12(16Q23.1) MUTATED | 3 | 0 | 6 | 0 |
| DEL PEAK 12(16Q23.1) WILD-TYPE | 145 | 14 | 21 | 2 |
Figure S54. Get High-res Image Gene #16: 'del_16q23.1' versus Molecular Subtype #1: 'CN_CNMF'
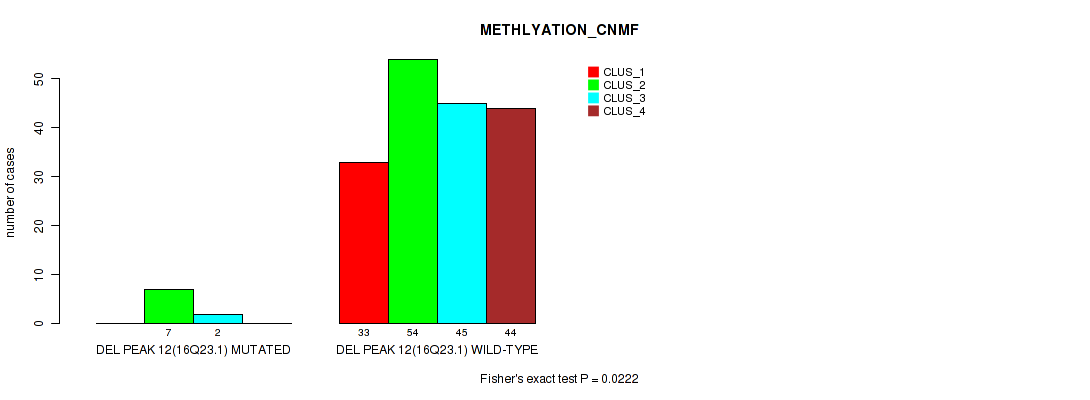
P value = 0.0222 (Fisher's exact test), Q value = 0.046
Table S55. Gene #16: 'del_16q23.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| DEL PEAK 12(16Q23.1) MUTATED | 0 | 7 | 2 | 0 |
| DEL PEAK 12(16Q23.1) WILD-TYPE | 33 | 54 | 45 | 44 |
Figure S55. Get High-res Image Gene #16: 'del_16q23.1' versus Molecular Subtype #2: 'METHLYATION_CNMF'
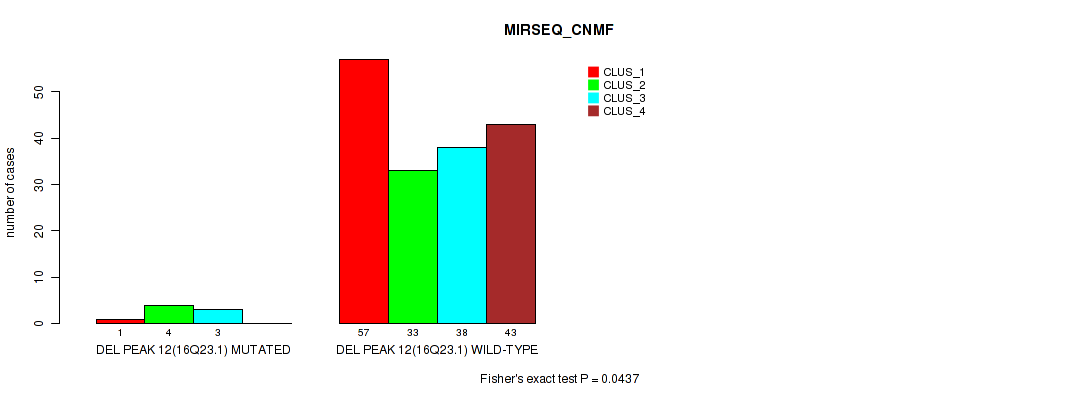
P value = 0.0437 (Fisher's exact test), Q value = 0.079
Table S56. Gene #16: 'del_16q23.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 58 | 37 | 41 | 43 |
| DEL PEAK 12(16Q23.1) MUTATED | 1 | 4 | 3 | 0 |
| DEL PEAK 12(16Q23.1) WILD-TYPE | 57 | 33 | 38 | 43 |
Figure S56. Get High-res Image Gene #16: 'del_16q23.1' versus Molecular Subtype #5: 'MIRSEQ_CNMF'
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S57. Gene #17: 'del_17p13.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 13(17P13.2) MUTATED | 0 | 0 | 15 | 0 |
| DEL PEAK 13(17P13.2) WILD-TYPE | 148 | 14 | 12 | 2 |
Figure S57. Get High-res Image Gene #17: 'del_17p13.2' versus Molecular Subtype #1: 'CN_CNMF'
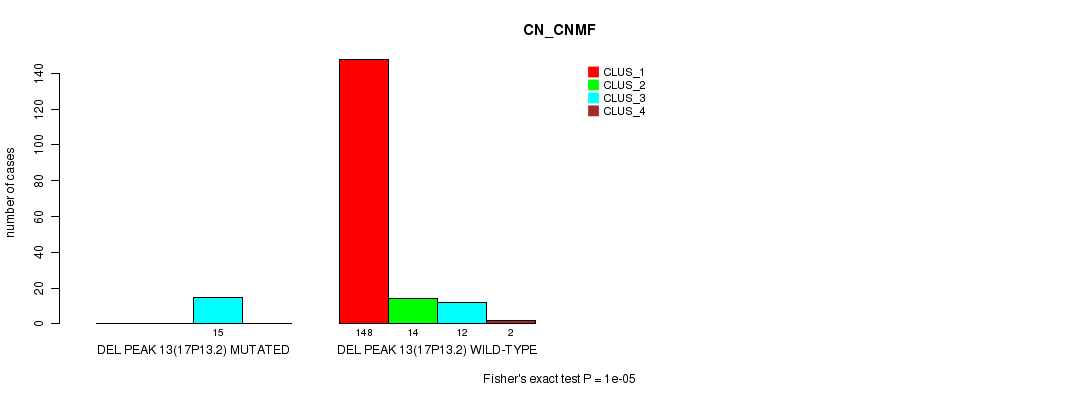
P value = 0.00033 (Fisher's exact test), Q value = 0.0021
Table S58. Gene #17: 'del_17p13.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| DEL PEAK 13(17P13.2) MUTATED | 0 | 12 | 3 | 0 |
| DEL PEAK 13(17P13.2) WILD-TYPE | 33 | 49 | 44 | 44 |
Figure S58. Get High-res Image Gene #17: 'del_17p13.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
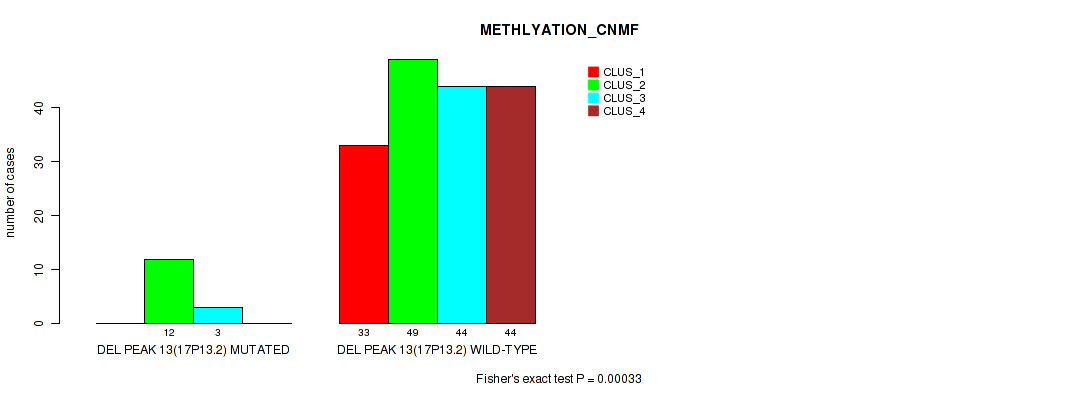
P value = 0.0114 (Fisher's exact test), Q value = 0.025
Table S59. Gene #17: 'del_17p13.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 70 | 52 | 44 |
| DEL PEAK 13(17P13.2) MUTATED | 12 | 2 | 1 |
| DEL PEAK 13(17P13.2) WILD-TYPE | 58 | 50 | 43 |
Figure S59. Get High-res Image Gene #17: 'del_17p13.2' versus Molecular Subtype #3: 'MRNASEQ_CNMF'
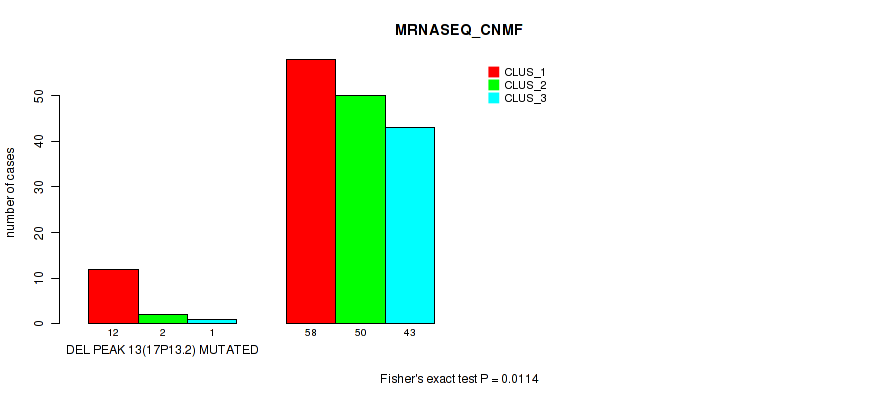
P value = 0.00018 (Fisher's exact test), Q value = 0.0013
Table S60. Gene #17: 'del_17p13.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 13(17P13.2) MUTATED | 0 | 0 | 0 | 14 | 0 | 1 |
| DEL PEAK 13(17P13.2) WILD-TYPE | 16 | 13 | 14 | 41 | 26 | 41 |
Figure S60. Get High-res Image Gene #17: 'del_17p13.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
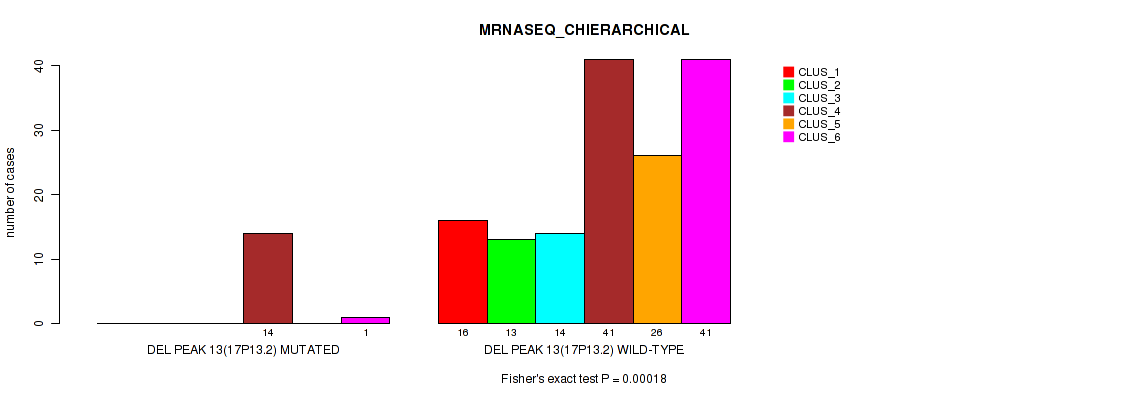
P value = 0.00124 (Fisher's exact test), Q value = 0.0051
Table S61. Gene #17: 'del_17p13.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 13(17P13.2) MUTATED | 0 | 5 | 10 |
| DEL PEAK 13(17P13.2) WILD-TYPE | 67 | 29 | 68 |
Figure S61. Get High-res Image Gene #17: 'del_17p13.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
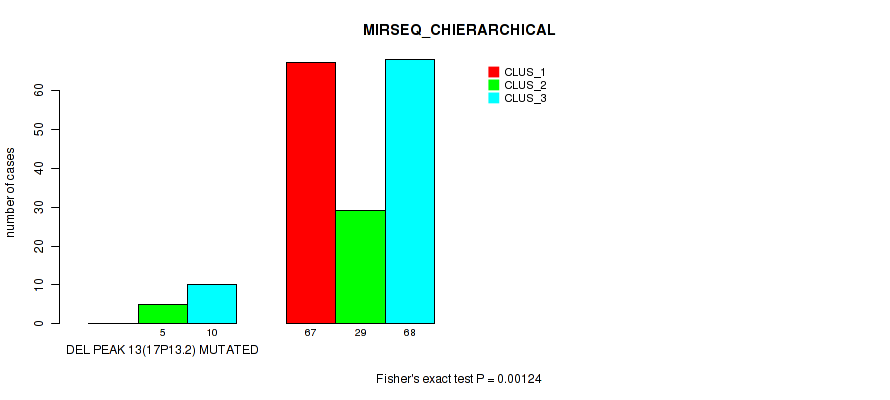
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S62. Gene #18: 'del_17q11.2' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 14(17Q11.2) MUTATED | 2 | 2 | 9 | 0 |
| DEL PEAK 14(17Q11.2) WILD-TYPE | 146 | 12 | 18 | 2 |
Figure S62. Get High-res Image Gene #18: 'del_17q11.2' versus Molecular Subtype #1: 'CN_CNMF'
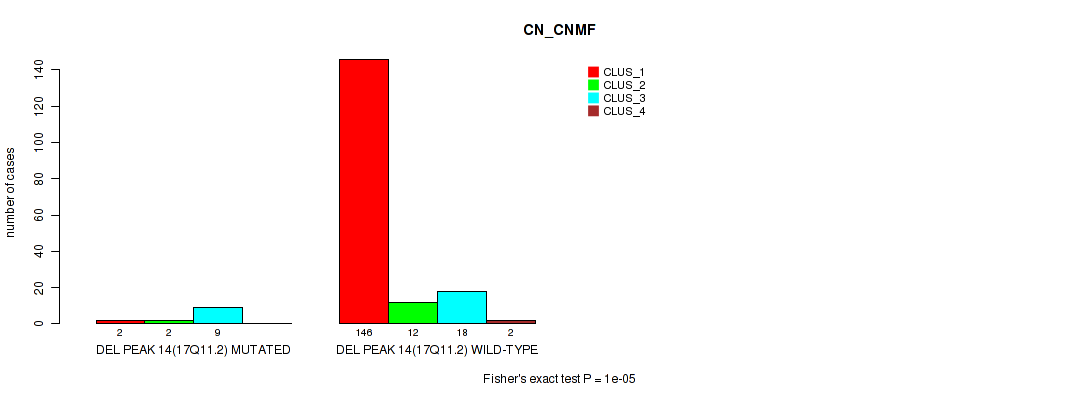
P value = 0.00493 (Fisher's exact test), Q value = 0.013
Table S63. Gene #18: 'del_17q11.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| DEL PEAK 14(17Q11.2) MUTATED | 0 | 9 | 4 | 0 |
| DEL PEAK 14(17Q11.2) WILD-TYPE | 33 | 52 | 43 | 44 |
Figure S63. Get High-res Image Gene #18: 'del_17q11.2' versus Molecular Subtype #2: 'METHLYATION_CNMF'
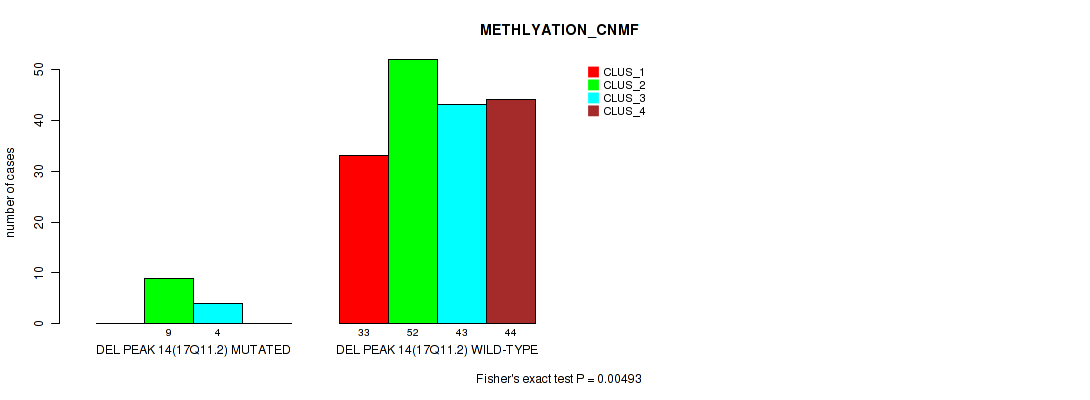
P value = 0.00717 (Fisher's exact test), Q value = 0.017
Table S64. Gene #18: 'del_17q11.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 | CLUS_5 | CLUS_6 |
|---|---|---|---|---|---|---|
| ALL | 16 | 13 | 14 | 55 | 26 | 42 |
| DEL PEAK 14(17Q11.2) MUTATED | 0 | 1 | 0 | 11 | 0 | 1 |
| DEL PEAK 14(17Q11.2) WILD-TYPE | 16 | 12 | 14 | 44 | 26 | 41 |
Figure S64. Get High-res Image Gene #18: 'del_17q11.2' versus Molecular Subtype #4: 'MRNASEQ_CHIERARCHICAL'
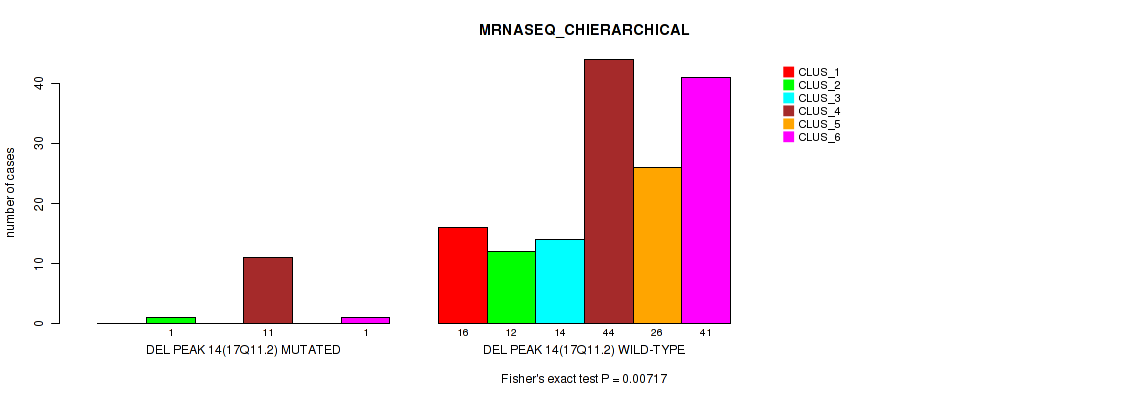
P value = 0.0038 (Fisher's exact test), Q value = 0.011
Table S65. Gene #18: 'del_17q11.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 14(17Q11.2) MUTATED | 0 | 4 | 9 |
| DEL PEAK 14(17Q11.2) WILD-TYPE | 67 | 30 | 69 |
Figure S65. Get High-res Image Gene #18: 'del_17q11.2' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
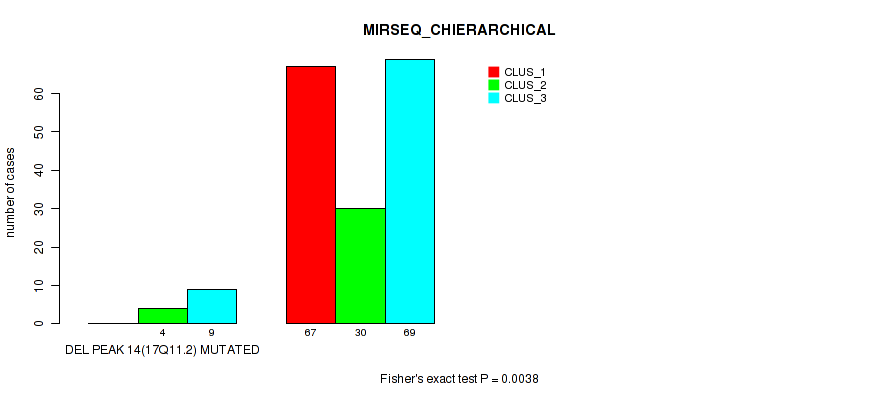
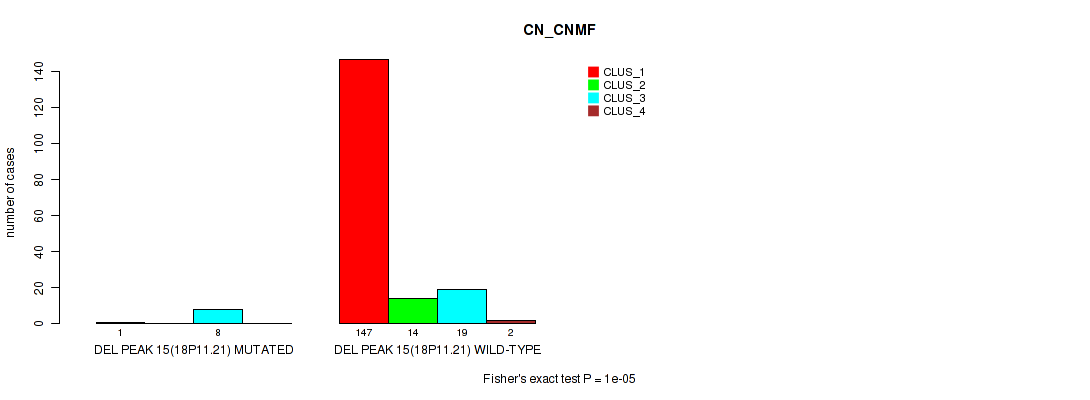
P value = 1e-05 (Fisher's exact test), Q value = 0.00011
Table S66. Gene #19: 'del_18p11.21' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 15(18P11.21) MUTATED | 1 | 0 | 8 | 0 |
| DEL PEAK 15(18P11.21) WILD-TYPE | 147 | 14 | 19 | 2 |
Figure S66. Get High-res Image Gene #19: 'del_18p11.21' versus Molecular Subtype #1: 'CN_CNMF'
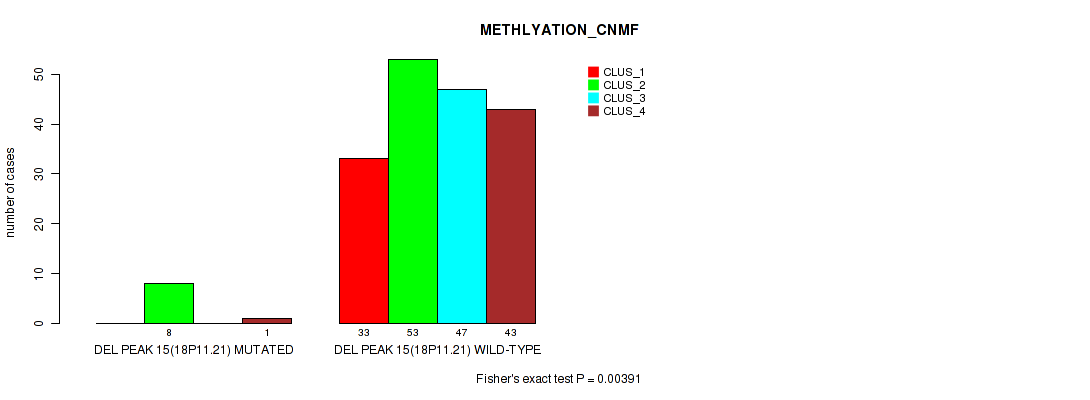
P value = 0.00391 (Fisher's exact test), Q value = 0.011
Table S67. Gene #19: 'del_18p11.21' versus Molecular Subtype #2: 'METHLYATION_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 33 | 61 | 47 | 44 |
| DEL PEAK 15(18P11.21) MUTATED | 0 | 8 | 0 | 1 |
| DEL PEAK 15(18P11.21) WILD-TYPE | 33 | 53 | 47 | 43 |
Figure S67. Get High-res Image Gene #19: 'del_18p11.21' versus Molecular Subtype #2: 'METHLYATION_CNMF'
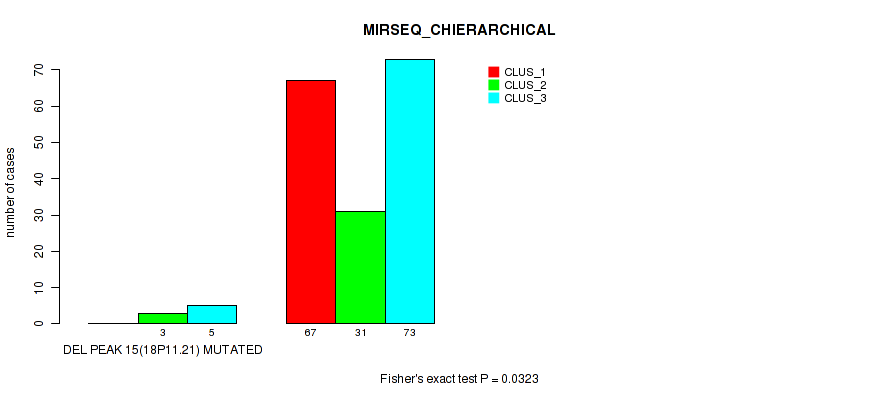
P value = 0.0323 (Fisher's exact test), Q value = 0.063
Table S68. Gene #19: 'del_18p11.21' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 |
|---|---|---|---|
| ALL | 67 | 34 | 78 |
| DEL PEAK 15(18P11.21) MUTATED | 0 | 3 | 5 |
| DEL PEAK 15(18P11.21) WILD-TYPE | 67 | 31 | 73 |
Figure S68. Get High-res Image Gene #19: 'del_18p11.21' versus Molecular Subtype #6: 'MIRSEQ_CHIERARCHICAL'
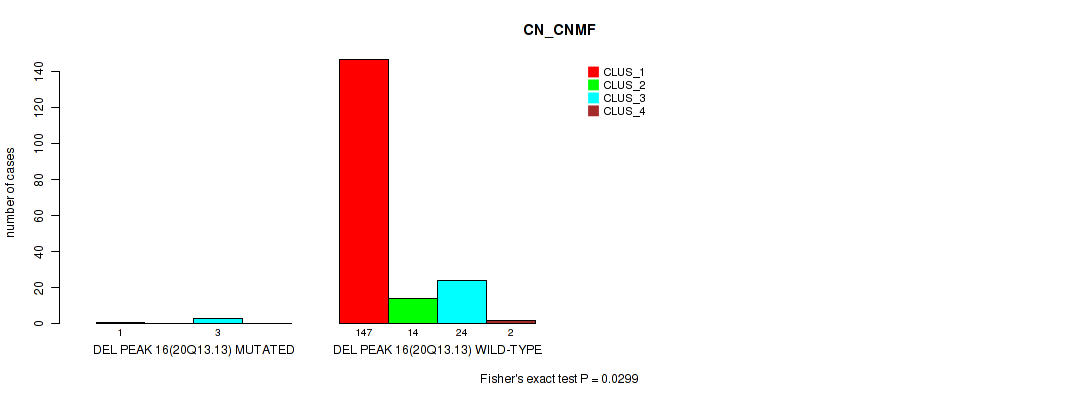
P value = 0.0299 (Fisher's exact test), Q value = 0.06
Table S69. Gene #20: 'del_20q13.13' versus Molecular Subtype #1: 'CN_CNMF'
| nPatients | CLUS_1 | CLUS_2 | CLUS_3 | CLUS_4 |
|---|---|---|---|---|
| ALL | 148 | 14 | 27 | 2 |
| DEL PEAK 16(20Q13.13) MUTATED | 1 | 0 | 3 | 0 |
| DEL PEAK 16(20Q13.13) WILD-TYPE | 147 | 14 | 24 | 2 |
Figure S69. Get High-res Image Gene #20: 'del_20q13.13' versus Molecular Subtype #1: 'CN_CNMF'
-
Copy number data file = all_lesions.txt from GISTIC pipeline
-
Processed Copy number data file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_Correlate_Genomic_Events_Preprocess/LAML-TB/15087095/transformed.cor.cli.txt
-
Molecular subtype file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_mergedClustering/LAML-TB/15098833/LAML-TB.transferedmergedcluster.txt
-
Number of patients = 191
-
Number of significantly focal cnvs = 20
-
Number of molecular subtypes = 6
-
Exclude genes that fewer than K tumors have alterations, K = 3
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.