This pipeline performs Gene Set Enrichment Analysis (GSEA) using The Broad Institute GSEA tool with MSigDB - Class2: Canonical Pathways gene sets. For a given phenotype subtype, it shows what pathways are significantly enriched in each subtype by comparing gene expression profiles between subtypes. Here, the phenotype is mRNAseq_cNMF subtypes in KIRC-TP. This pipeline has the following features:
-
For each subtype, calculates enrichment scores (ES) using signal to noise (S2N) that checks similarity between subtypes in expression level then calculates p values through permutation test.
-
Lists pathways significantly enriched in each phenotype subtype and their enrichment scores (ES).
-
Lists top 20 core genes enriched in each significant gene set and their enrichment scores (ES).
-
Checks if the top core genes are up-regulated or down-regulated.
-
Checks if the top core genes are high expressed or low expressed.
-
Checks if the top core genes are significantly differently expressed genes.
Table 1. Get Full Table basic data info
| basic data info |
|---|
| Number of Gene Sets: 194 |
| Number of samples: 533 |
| Original number of Gene Sets: 404 |
| Maximum gene set size: 388 |
Table 2. Get Full Table pheno data info
| phenotype info |
|---|
| pheno.type: 1 - 3 :[ clus1 ] 116 |
| pheno.type: 2 - 3 :[ clus2 ] 223 |
| pheno.type: 3 - 3 :[ clus3 ] 194 |
For the expression subtypes of 18278 genes in 534 samples, GSEA found enriched gene sets in each cluster using 533 gene sets in MSigDB canonical pathways. Top enriched gene sets are listed as below.
-
clus1
-
Top enriched gene sets are BIOCARTA NO1 PATHWAY, BIOCARTA AGR PATHWAY, BIOCARTA RHO PATHWAY, BIOCARTA GPCR PATHWAY, BIOCARTA CREB PATHWAY, KEGG OXIDATIVE PHOSPHORYLATION, KEGG O GLYCAN BIOSYNTHESIS, KEGG INOSITOL PHOSPHATE METABOLISM, KEGG CARDIAC MUSCLE CONTRACTION, KEGG WNT SIGNALING PATHWAY
-
And common core enriched genes are PRKCA, CTNNB1, CXXC4, DAAM1, DKK1, FRAT1, FZD2, FZD3, FZD6, FZD7
-
clus2
-
Top enriched gene sets are BIOCARTA AT1R PATHWAY, BIOCARTA DEATH PATHWAY, BIOCARTA VEGF PATHWAY, KEGG GLYCOLYSIS GLUCONEOGENESIS, KEGG PENTOSE AND GLUCURONATE INTERCONVERSIONS, KEGG GALACTOSE METABOLISM, KEGG FATTY ACID METABOLISM, KEGG GLYCINE SERINE AND THREONINE METABOLISM, KEGG VALINE LEUCINE AND ISOLEUCINE DEGRADATION, KEGG LYSINE DEGRADATION
-
And common core enriched genes are EHHADH, ACADM, ALDH3A2, ACSL1, HMGCS2, SLC27A2, ABAT, ACAT1, ALDH6A1, ALDH7A1
-
clus3
-
Top enriched gene sets are BIOCARTA INFLAM PATHWAY, BIOCARTA GSK3 PATHWAY, BIOCARTA WNT PATHWAY, KEGG AMINO SUGAR AND NUCLEOTIDE SUGAR METABOLISM, KEGG DNA REPLICATION, KEGG BASE EXCISION REPAIR, KEGG HOMOLOGOUS RECOMBINATION, KEGG CHEMOKINE SIGNALING PATHWAY, KEGG CELL CYCLE, KEGG OOCYTE MEIOSIS
-
And common core enriched genes are CCL5, IL1B, IL6, IL18, PYCARD, CCNB1, CCNB2, CCNE1, CCNE2, CDK1
Table 3. Get Full Table This table shows top 10 pathways which are significantly enriched in cluster clus1. It displays only significant gene sets satisfying nom.p.val.threshold (-1), fwer.p.val.threshold (-1) , fdr.q.val.threshold (0.25) and the default table is sorted by Normalized Enrichment Score (NES). Further details on NES statistics, please visit The Broad GSEA website.
| GeneSet(GS) | Size(#genes) | genes.ES.table | ES | NES | NOM.p.val | FDR.q.val | FWER.p.val | Tag.. | Gene.. | Signal | FDR..median. | glob.p.val |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BIOCARTA NO1 PATHWAY | 28 | genes.ES.table | 0.57 | 1.4 | 0.064 | 0.68 | 0.93 | 0.46 | 0.18 | 0.38 | 0.52 | 0.22 |
| BIOCARTA AGR PATHWAY | 34 | genes.ES.table | 0.58 | 1.7 | 0.0021 | 0.57 | 0.49 | 0.26 | 0.14 | 0.23 | 0 | 0.19 |
| BIOCARTA RHO PATHWAY | 31 | genes.ES.table | 0.43 | 1.6 | 0.024 | 0.79 | 0.78 | 0.19 | 0.16 | 0.16 | 0.5 | 0.26 |
| BIOCARTA GPCR PATHWAY | 32 | genes.ES.table | 0.48 | 1.5 | 0.061 | 0.56 | 0.86 | 0.47 | 0.26 | 0.35 | 0.37 | 0.18 |
| BIOCARTA CREB PATHWAY | 25 | genes.ES.table | 0.46 | 1.5 | 0.069 | 0.56 | 0.88 | 0.44 | 0.26 | 0.32 | 0.38 | 0.16 |
| KEGG OXIDATIVE PHOSPHORYLATION | 113 | genes.ES.table | 0.5 | 1.5 | 0.16 | 0.63 | 0.91 | 0.6 | 0.32 | 0.41 | 0.46 | 0.2 |
| KEGG O GLYCAN BIOSYNTHESIS | 28 | genes.ES.table | 0.48 | 1.3 | 0.086 | 0.73 | 0.97 | 0.46 | 0.19 | 0.38 | 0.61 | 0.24 |
| KEGG INOSITOL PHOSPHATE METABOLISM | 53 | genes.ES.table | 0.37 | 1.3 | 0.15 | 0.63 | 0.98 | 0.36 | 0.23 | 0.28 | 0.53 | 0.19 |
| KEGG CARDIAC MUSCLE CONTRACTION | 66 | genes.ES.table | 0.45 | 1.3 | 0.09 | 0.6 | 0.98 | 0.3 | 0.14 | 0.26 | 0.51 | 0.17 |
| KEGG WNT SIGNALING PATHWAY | 145 | genes.ES.table | 0.46 | 1.5 | 0.037 | 0.76 | 0.82 | 0.37 | 0.21 | 0.3 | 0.49 | 0.26 |
Table S1. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | ATP6V0A4 | ATP6V0A4 | ATP6V0A4 | 8 | 0.69 | 0.12 | YES |
| 2 | ATP6V1B1 | ATP6V1B1 | ATP6V1B1 | 40 | 0.59 | 0.22 | YES |
| 3 | CFTR | CFTR | CFTR | 54 | 0.58 | 0.32 | YES |
| 4 | ATP6V0D2 | ATP6V0D2 | ATP6V0D2 | 139 | 0.48 | 0.4 | YES |
| 5 | PLCG2 | PLCG2 | PLCG2 | 1071 | 0.23 | 0.38 | YES |
| 6 | KCNQ1 | KCNQ1 | KCNQ1 | 1101 | 0.22 | 0.42 | YES |
| 7 | ATP6V1C2 | ATP6V1C2 | ATP6V1C2 | 1102 | 0.22 | 0.46 | YES |
| 8 | ATP6V1G2 | ATP6V1G2 | ATP6V1G2 | 1115 | 0.22 | 0.5 | YES |
| 9 | SLC12A2 | SLC12A2 | SLC12A2 | 1997 | 0.14 | 0.47 | YES |
| 10 | ATP6V1A | ATP6V1A | ATP6V1A | 2299 | 0.12 | 0.48 | YES |
| 11 | ATP6V1D | ATP6V1D | ATP6V1D | 2454 | 0.11 | 0.49 | YES |
| 12 | ATP6V1H | ATP6V1H | ATP6V1H | 2724 | 0.1 | 0.49 | YES |
| 13 | PRKCA | PRKCA | PRKCA | 2922 | 0.092 | 0.5 | YES |
| 14 | ATP6V0B | ATP6V0B | ATP6V0B | 3032 | 0.088 | 0.5 | YES |
| 15 | ATP6V0A2 | ATP6V0A2 | ATP6V0A2 | 3047 | 0.088 | 0.52 | YES |
| 16 | TJP1 | TJP1 | TJP1 | 3400 | 0.076 | 0.51 | NO |
| 17 | ATP6V0E2 | ATP6V0E2 | ATP6V0E2 | 3670 | 0.068 | 0.51 | NO |
| 18 | ATP6V0D1 | ATP6V0D1 | ATP6V0D1 | 4097 | 0.059 | 0.5 | NO |
| 19 | ATP6V0C | ATP6V0C | ATP6V0C | 4127 | 0.058 | 0.5 | NO |
| 20 | ATP6V1G1 | ATP6V1G1 | ATP6V1G1 | 4243 | 0.056 | 0.51 | NO |
| 21 | ATP6AP1 | ATP6AP1 | ATP6AP1 | 4491 | 0.051 | 0.5 | NO |
| 22 | GNAS | GNAS | GNAS | 4530 | 0.05 | 0.51 | NO |
| 23 | ATP6V1E1 | ATP6V1E1 | ATP6V1E1 | 4740 | 0.046 | 0.51 | NO |
| 24 | PRKACB | PRKACB | PRKACB | 4823 | 0.045 | 0.51 | NO |
| 25 | TJP2 | TJP2 | TJP2 | 5652 | 0.032 | 0.47 | NO |
| 26 | PRKACA | PRKACA | PRKACA | 5827 | 0.03 | 0.46 | NO |
| 27 | ATP6V1F | ATP6V1F | ATP6V1F | 5830 | 0.03 | 0.47 | NO |
| 28 | ACTG1 | ACTG1 | ACTG1 | 6293 | 0.024 | 0.45 | NO |
| 29 | KDELR1 | KDELR1 | KDELR1 | 6742 | 0.019 | 0.43 | NO |
| 30 | ATP6V0E1 | ATP6V0E1 | ATP6V0E1 | 7328 | 0.012 | 0.4 | NO |
| 31 | ATP6V1C1 | ATP6V1C1 | ATP6V1C1 | 7633 | 0.0092 | 0.38 | NO |
| 32 | ACTB | ACTB | ACTB | 7964 | 0.0061 | 0.37 | NO |
| 33 | PRKX | PRKX | PRKX | 8539 | 0.0004 | 0.33 | NO |
| 34 | PLCG1 | PLCG1 | PLCG1 | 9144 | -0.0056 | 0.3 | NO |
| 35 | SEC61B | SEC61B | SEC61B | 9394 | -0.0081 | 0.29 | NO |
| 36 | ARF1 | ARF1 | ARF1 | 9606 | -0.01 | 0.28 | NO |
| 37 | MUC2 | MUC2 | MUC2 | 9991 | -0.014 | 0.26 | NO |
| 38 | KDELR2 | KDELR2 | KDELR2 | 10537 | -0.02 | 0.24 | NO |
| 39 | SEC61A1 | SEC61A1 | SEC61A1 | 10820 | -0.024 | 0.22 | NO |
| 40 | ERO1L | ERO1L | ERO1L | 11918 | -0.038 | 0.17 | NO |
| 41 | ADCY9 | ADCY9 | ADCY9 | 11963 | -0.039 | 0.17 | NO |
| 42 | ATP6V1B2 | ATP6V1B2 | ATP6V1B2 | 12650 | -0.051 | 0.14 | NO |
| 43 | PDIA4 | PDIA4 | PDIA4 | 13627 | -0.072 | 0.1 | NO |
| 44 | SEC61G | SEC61G | SEC61G | 13650 | -0.073 | 0.12 | NO |
| 45 | ATP6V0A1 | ATP6V0A1 | ATP6V0A1 | 13734 | -0.075 | 0.12 | NO |
| 46 | KDELR3 | KDELR3 | KDELR3 | 14435 | -0.096 | 0.1 | NO |
| 47 | SEC61A2 | SEC61A2 | SEC61A2 | 14609 | -0.1 | 0.11 | NO |
| 48 | TCIRG1 | TCIRG1 | TCIRG1 | 14615 | -0.1 | 0.13 | NO |
| 49 | ATP6V1E2 | ATP6V1E2 | ATP6V1E2 | 14777 | -0.11 | 0.14 | NO |
| 50 | PRKCG | PRKCG | PRKCG | 14786 | -0.11 | 0.16 | NO |
| 51 | PRKCB | PRKCB | PRKCB | 16685 | -0.2 | 0.086 | NO |
Figure S1. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA NO1 PATHWAY.
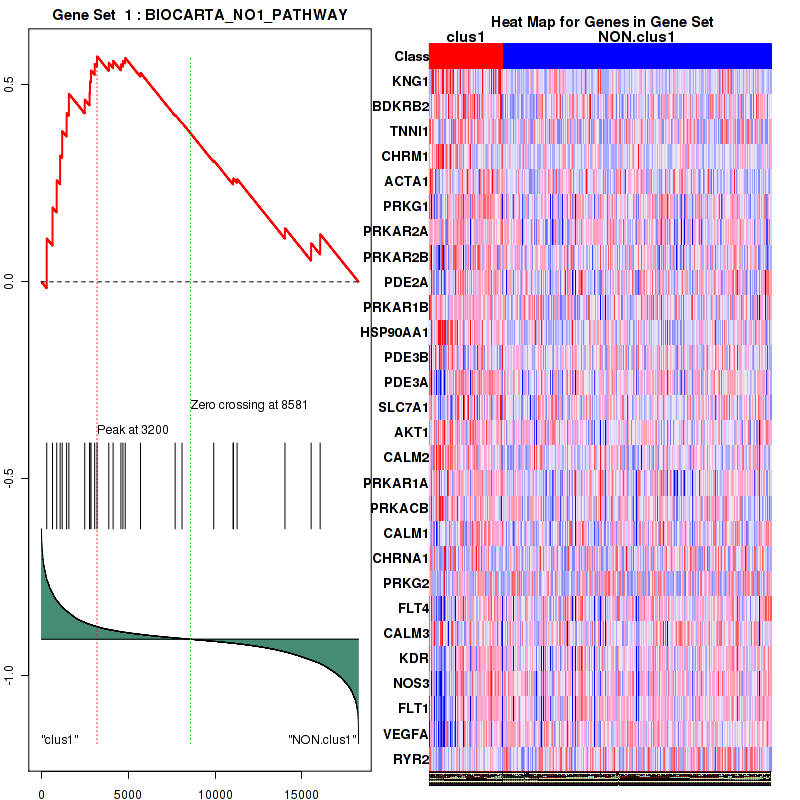
Figure S2. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA NO1 PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
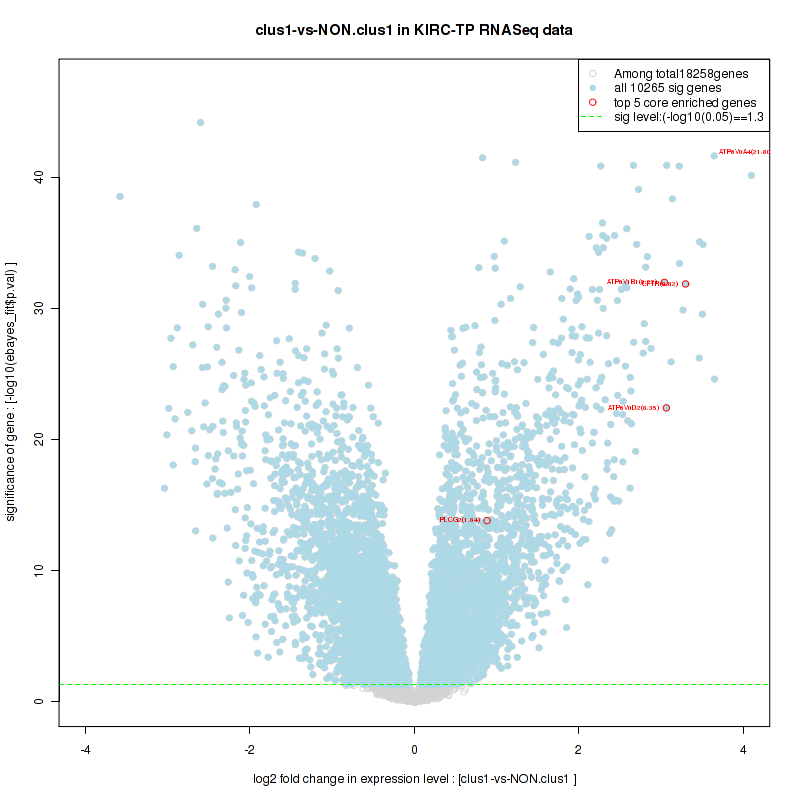
Table S2. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | LAMA2 | LAMA2 | LAMA2 | 32 | 0.6 | 0.17 | YES |
| 2 | PAK6 | PAK6 | PAK6 | 48 | 0.58 | 0.33 | YES |
| 3 | PAK7 | PAK7 | PAK7 | 401 | 0.36 | 0.41 | YES |
| 4 | CHRM1 | CHRM1 | CHRM1 | 1076 | 0.23 | 0.44 | YES |
| 5 | ACTA1 | ACTA1 | ACTA1 | 1193 | 0.21 | 0.49 | YES |
| 6 | NRG2 | NRG2 | NRG2 | 1221 | 0.21 | 0.55 | YES |
| 7 | DMD | DMD | DMD | 1942 | 0.14 | 0.55 | YES |
| 8 | DAG1 | DAG1 | DAG1 | 2222 | 0.12 | 0.56 | YES |
| 9 | LAMA3 | LAMA3 | LAMA3 | 2552 | 0.11 | 0.58 | YES |
| 10 | MAPK8 | MAPK8 | MAPK8 | 4785 | 0.046 | 0.47 | NO |
| 11 | PTK2 | PTK2 | PTK2 | 5172 | 0.039 | 0.46 | NO |
| 12 | PAK4 | PAK4 | PAK4 | 5807 | 0.03 | 0.43 | NO |
| 13 | CDC42 | CDC42 | CDC42 | 6780 | 0.018 | 0.38 | NO |
| 14 | UTRN | UTRN | UTRN | 6948 | 0.017 | 0.38 | NO |
| 15 | ITGB1 | ITGB1 | ITGB1 | 7161 | 0.014 | 0.37 | NO |
| 16 | CHRNA1 | CHRNA1 | CHRNA1 | 7700 | 0.0087 | 0.34 | NO |
| 17 | RAC1 | RAC1 | RAC1 | 7856 | 0.0071 | 0.34 | NO |
| 18 | CTTN | CTTN | CTTN | 8870 | -0.003 | 0.28 | NO |
| 19 | MAPK3 | MAPK3 | MAPK3 | 9071 | -0.005 | 0.27 | NO |
| 20 | PXN | PXN | PXN | 9426 | -0.0084 | 0.26 | NO |
| 21 | ITGA1 | ITGA1 | ITGA1 | 9512 | -0.0093 | 0.25 | NO |
| 22 | PAK2 | PAK2 | PAK2 | 9603 | -0.01 | 0.25 | NO |
| 23 | DVL1 | DVL1 | DVL1 | 9928 | -0.013 | 0.24 | NO |
| 24 | LAMA4 | LAMA4 | LAMA4 | 10475 | -0.019 | 0.21 | NO |
| 25 | MAPK1 | MAPK1 | MAPK1 | 11822 | -0.037 | 0.15 | NO |
| 26 | SP1 | SP1 | SP1 | 12235 | -0.043 | 0.14 | NO |
| 27 | SRC | SRC | SRC | 12472 | -0.048 | 0.14 | NO |
| 28 | PAK1 | PAK1 | PAK1 | 12783 | -0.054 | 0.14 | NO |
| 29 | PAK3 | PAK3 | PAK3 | 13640 | -0.073 | 0.11 | NO |
| 30 | EGFR | EGFR | EGFR | 13768 | -0.076 | 0.12 | NO |
| 31 | JUN | JUN | JUN | 13855 | -0.079 | 0.14 | NO |
| 32 | NRG1 | NRG1 | NRG1 | 14797 | -0.11 | 0.12 | NO |
| 33 | RAPSN | RAPSN | RAPSN | 15023 | -0.12 | 0.14 | NO |
| 34 | GIT2 | GIT2 | GIT2 | 15488 | -0.14 | 0.15 | NO |
Figure S3. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA AGR PATHWAY.
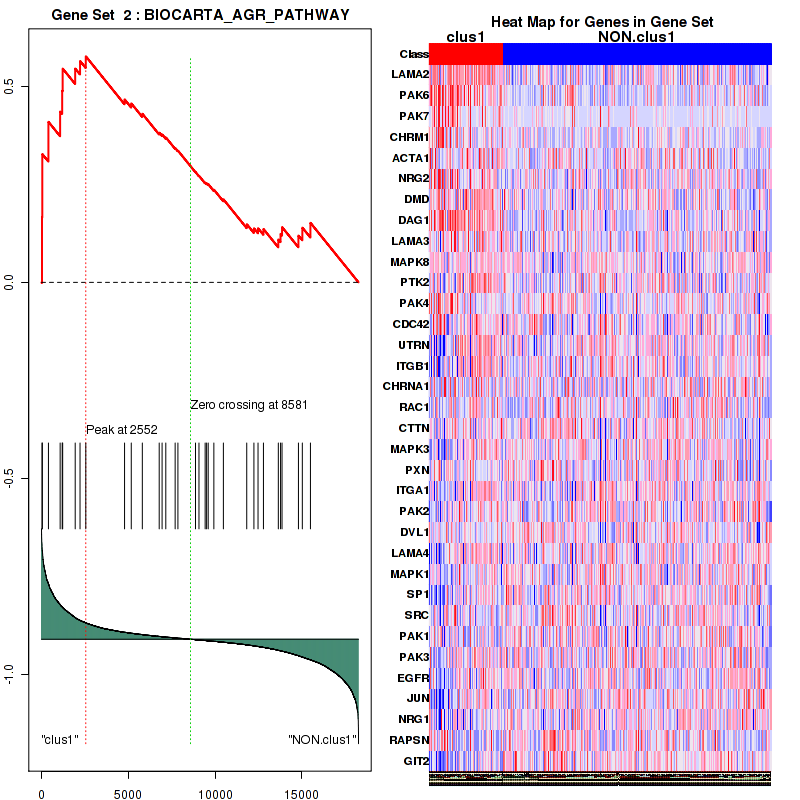
Figure S4. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA AGR PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
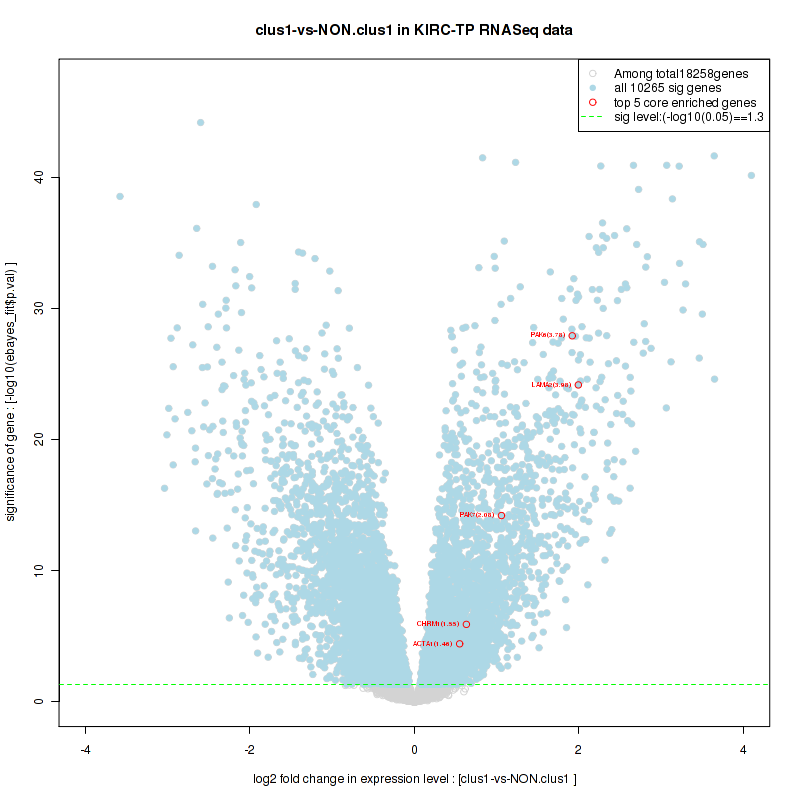
Table S3. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | SFRP1 | SFRP1 | SFRP1 | 82 | 0.54 | 0.057 | YES |
| 2 | PYGO1 | PYGO1 | PYGO1 | 205 | 0.45 | 0.1 | YES |
| 3 | NKD1 | NKD1 | NKD1 | 294 | 0.4 | 0.14 | YES |
| 4 | WNT4 | WNT4 | WNT4 | 374 | 0.37 | 0.18 | YES |
| 5 | WNT7A | WNT7A | WNT7A | 473 | 0.34 | 0.22 | YES |
| 6 | WNT11 | WNT11 | WNT11 | 479 | 0.34 | 0.25 | YES |
| 7 | WNT2 | WNT2 | WNT2 | 529 | 0.33 | 0.29 | YES |
| 8 | TLE2 | TLE2 | TLE2 | 622 | 0.3 | 0.32 | YES |
| 9 | FZD7 | FZD7 | FZD7 | 869 | 0.26 | 0.33 | YES |
| 10 | WNT7B | WNT7B | WNT7B | 881 | 0.26 | 0.36 | YES |
| 11 | DAAM1 | DAAM1 | DAAM1 | 901 | 0.25 | 0.39 | YES |
| 12 | FZD2 | FZD2 | FZD2 | 1235 | 0.21 | 0.4 | YES |
| 13 | WISP1 | WISP1 | WISP1 | 1346 | 0.2 | 0.41 | YES |
| 14 | WNT5A | WNT5A | WNT5A | 1750 | 0.16 | 0.41 | YES |
| 15 | CXXC4 | CXXC4 | CXXC4 | 1853 | 0.15 | 0.42 | YES |
| 16 | FZD3 | FZD3 | FZD3 | 1867 | 0.15 | 0.44 | YES |
| 17 | HPRT1 | HPRT1 | HPRT1 | 2067 | 0.13 | 0.44 | YES |
| 18 | PORCN | PORCN | PORCN | 2124 | 0.13 | 0.45 | YES |
| 19 | TCF7L1 | TCF7L1 | TCF7L1 | 2223 | 0.12 | 0.46 | YES |
| 20 | TLE1 | TLE1 | TLE1 | 2416 | 0.11 | 0.46 | YES |
| 21 | WNT6 | WNT6 | WNT6 | 2428 | 0.11 | 0.48 | YES |
| 22 | FRAT1 | FRAT1 | FRAT1 | 2550 | 0.11 | 0.48 | YES |
| 23 | LEF1 | LEF1 | LEF1 | 2713 | 0.1 | 0.48 | YES |
| 24 | WIF1 | WIF1 | WIF1 | 3046 | 0.088 | 0.48 | YES |
| 25 | WNT5B | WNT5B | WNT5B | 3265 | 0.08 | 0.47 | YES |
| 26 | LRP6 | LRP6 | LRP6 | 3351 | 0.078 | 0.48 | YES |
| 27 | FZD6 | FZD6 | FZD6 | 3366 | 0.077 | 0.48 | YES |
| 28 | BCL9 | BCL9 | BCL9 | 3373 | 0.077 | 0.49 | YES |
| 29 | WNT9A | WNT9A | WNT9A | 3451 | 0.074 | 0.5 | YES |
| 30 | DKK1 | DKK1 | DKK1 | 3532 | 0.072 | 0.5 | YES |
| 31 | CTNNB1 | CTNNB1 | CTNNB1 | 3624 | 0.069 | 0.5 | YES |
| 32 | TCF7 | TCF7 | TCF7 | 3722 | 0.067 | 0.51 | YES |
| 33 | GSK3B | GSK3B | GSK3B | 3846 | 0.065 | 0.51 | YES |
| 34 | FBXW2 | FBXW2 | FBXW2 | 4267 | 0.055 | 0.49 | NO |
| 35 | FZD8 | FZD8 | FZD8 | 4475 | 0.051 | 0.49 | NO |
| 36 | CTNNBIP1 | CTNNBIP1 | CTNNBIP1 | 4946 | 0.043 | 0.46 | NO |
| 37 | FOXN1 | FOXN1 | FOXN1 | 5052 | 0.041 | 0.46 | NO |
| 38 | AES | AES | AES | 5328 | 0.037 | 0.45 | NO |
| 39 | FBXW11 | FBXW11 | FBXW11 | 5534 | 0.034 | 0.45 | NO |
| 40 | CSNK2A1 | CSNK2A1 | CSNK2A1 | 6036 | 0.027 | 0.42 | NO |
| 41 | SOX17 | SOX17 | SOX17 | 6841 | 0.018 | 0.38 | NO |
| 42 | BTRC | BTRC | BTRC | 7082 | 0.015 | 0.37 | NO |
| 43 | RHOU | RHOU | RHOU | 7219 | 0.014 | 0.36 | NO |
| 44 | KREMEN1 | KREMEN1 | KREMEN1 | 7243 | 0.013 | 0.36 | NO |
| 45 | EP300 | EP300 | EP300 | 7375 | 0.012 | 0.36 | NO |
| 46 | PPP2R1A | PPP2R1A | PPP2R1A | 7410 | 0.012 | 0.36 | NO |
| 47 | DVL2 | DVL2 | DVL2 | 7756 | 0.008 | 0.34 | NO |
| 48 | GSK3A | GSK3A | GSK3A | 7769 | 0.008 | 0.34 | NO |
| 49 | LRP5 | LRP5 | LRP5 | 7838 | 0.0072 | 0.34 | NO |
| 50 | PPP2CA | PPP2CA | PPP2CA | 7844 | 0.0072 | 0.34 | NO |
| 51 | ACTB | ACTB | ACTB | 7964 | 0.0061 | 0.33 | NO |
| 52 | CSNK1G1 | CSNK1G1 | CSNK1G1 | 7990 | 0.0059 | 0.33 | NO |
| 53 | RPL13A | RPL13A | RPL13A | 8052 | 0.0054 | 0.33 | NO |
| 54 | SENP2 | SENP2 | SENP2 | 8266 | 0.0032 | 0.32 | NO |
| 55 | CSNK1D | CSNK1D | CSNK1D | 8383 | 0.0019 | 0.31 | NO |
| 56 | APC | APC | APC | 8501 | 0.00067 | 0.3 | NO |
| 57 | CTBP1 | CTBP1 | CTBP1 | 8673 | -0.001 | 0.29 | NO |
| 58 | AXIN1 | AXIN1 | AXIN1 | 9215 | -0.0063 | 0.26 | NO |
| 59 | CTBP2 | CTBP2 | CTBP2 | 9452 | -0.0087 | 0.25 | NO |
| 60 | CCND3 | CCND3 | CCND3 | 9559 | -0.0098 | 0.25 | NO |
| 61 | CCND2 | CCND2 | CCND2 | 9809 | -0.012 | 0.24 | NO |
| 62 | FRZB | FRZB | FRZB | 9810 | -0.012 | 0.24 | NO |
| 63 | NLK | NLK | NLK | 9827 | -0.012 | 0.24 | NO |
| 64 | DVL1 | DVL1 | DVL1 | 9928 | -0.013 | 0.23 | NO |
| 65 | FBXW4 | FBXW4 | FBXW4 | 10095 | -0.015 | 0.22 | NO |
| 66 | CSNK1A1 | CSNK1A1 | CSNK1A1 | 10525 | -0.02 | 0.2 | NO |
| 67 | B2M | B2M | B2M | 11404 | -0.031 | 0.16 | NO |
| 68 | SFRP4 | SFRP4 | SFRP4 | 11774 | -0.036 | 0.14 | NO |
| 69 | FZD4 | FZD4 | FZD4 | 12617 | -0.05 | 0.1 | NO |
| 70 | FOSL1 | FOSL1 | FOSL1 | 13077 | -0.059 | 0.084 | NO |
| 71 | GAPDH | GAPDH | GAPDH | 13153 | -0.061 | 0.087 | NO |
| 72 | WNT16 | WNT16 | WNT16 | 13175 | -0.062 | 0.093 | NO |
| 73 | WNT2B | WNT2B | WNT2B | 13310 | -0.064 | 0.093 | NO |
| 74 | JUN | JUN | JUN | 13855 | -0.079 | 0.072 | NO |
| 75 | DIXDC1 | DIXDC1 | DIXDC1 | 13935 | -0.081 | 0.077 | NO |
| 76 | WNT10A | WNT10A | WNT10A | 13945 | -0.081 | 0.086 | NO |
| 77 | CCND1 | CCND1 | CCND1 | 14009 | -0.083 | 0.092 | NO |
| 78 | SLC9A3R1 | SLC9A3R1 | SLC9A3R1 | 15432 | -0.13 | 0.029 | NO |
| 79 | FZD1 | FZD1 | FZD1 | 15552 | -0.14 | 0.038 | NO |
| 80 | PITX2 | PITX2 | PITX2 | 15717 | -0.14 | 0.046 | NO |
| 81 | MYC | MYC | MYC | 15790 | -0.15 | 0.059 | NO |
| 82 | WNT3 | WNT3 | WNT3 | 15824 | -0.15 | 0.074 | NO |
| 83 | FZD5 | FZD5 | FZD5 | 16822 | -0.21 | 0.044 | NO |
| 84 | WNT1 | WNT1 | WNT1 | 17674 | -0.31 | 0.032 | NO |
Figure S5. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA RHO PATHWAY.
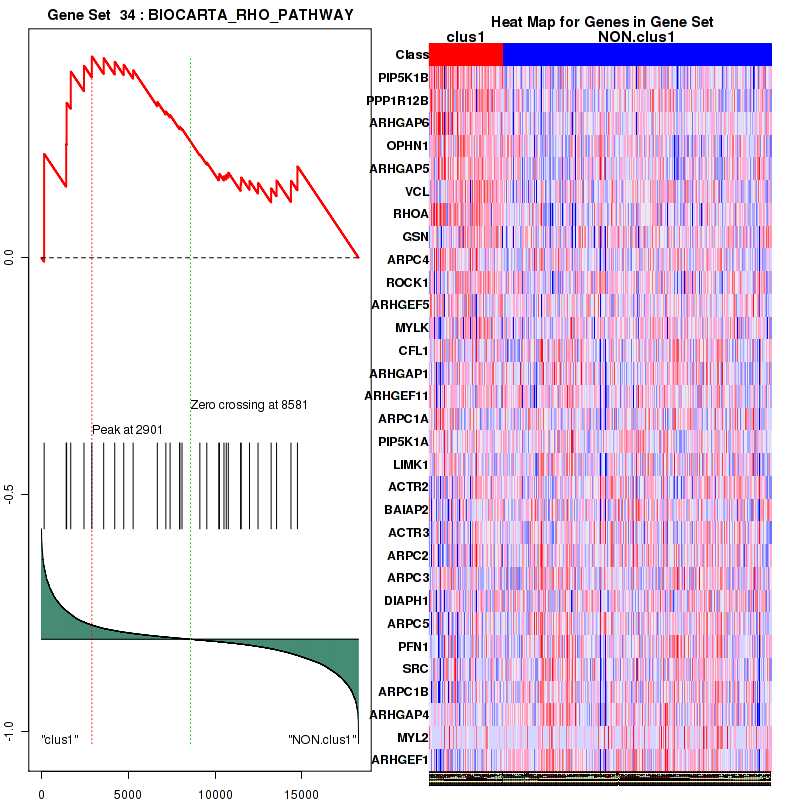
Figure S6. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA RHO PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
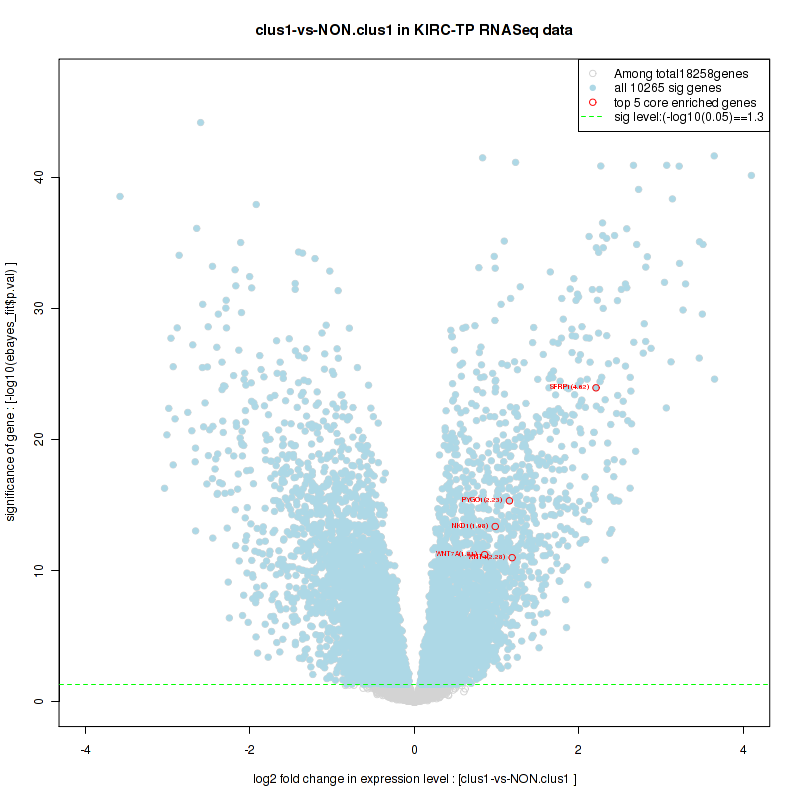
Table S4. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | PIP5K1B | PIP5K1B | PIP5K1B | 153 | 0.48 | 0.22 | YES |
| 2 | PPP1R12B | PPP1R12B | PPP1R12B | 1424 | 0.19 | 0.24 | YES |
| 3 | ARHGAP6 | ARHGAP6 | ARHGAP6 | 1448 | 0.19 | 0.33 | YES |
| 4 | OPHN1 | OPHN1 | OPHN1 | 1687 | 0.16 | 0.39 | YES |
| 5 | ARHGAP5 | ARHGAP5 | ARHGAP5 | 2449 | 0.11 | 0.4 | YES |
| 6 | VCL | VCL | VCL | 2901 | 0.093 | 0.42 | YES |
| 7 | RHOA | RHOA | RHOA | 3584 | 0.07 | 0.42 | NO |
| 8 | GSN | GSN | GSN | 4218 | 0.056 | 0.41 | NO |
| 9 | ARPC4 | ARPC4 | ARPC4 | 4736 | 0.046 | 0.41 | NO |
| 10 | ROCK1 | ROCK1 | ROCK1 | 5280 | 0.038 | 0.4 | NO |
| 11 | ARHGEF5 | ARHGEF5 | ARHGEF5 | 6673 | 0.02 | 0.33 | NO |
| 12 | MYLK | MYLK | MYLK | 7163 | 0.014 | 0.31 | NO |
| 13 | CFL1 | CFL1 | CFL1 | 7406 | 0.012 | 0.3 | NO |
| 14 | ARHGAP1 | ARHGAP1 | ARHGAP1 | 7956 | 0.0062 | 0.27 | NO |
| 15 | ARHGEF11 | ARHGEF11 | ARHGEF11 | 7969 | 0.0061 | 0.28 | NO |
| 16 | ARPC1A | ARPC1A | ARPC1A | 8094 | 0.0051 | 0.27 | NO |
| 17 | PIP5K1A | PIP5K1A | PIP5K1A | 9120 | -0.0055 | 0.22 | NO |
| 18 | LIMK1 | LIMK1 | LIMK1 | 9521 | -0.0095 | 0.2 | NO |
| 19 | ACTR2 | ACTR2 | ACTR2 | 10211 | -0.016 | 0.17 | NO |
| 20 | BAIAP2 | BAIAP2 | BAIAP2 | 10248 | -0.017 | 0.18 | NO |
| 21 | ACTR3 | ACTR3 | ACTR3 | 10509 | -0.02 | 0.17 | NO |
| 22 | ARPC2 | ARPC2 | ARPC2 | 10643 | -0.021 | 0.18 | NO |
| 23 | ARPC3 | ARPC3 | ARPC3 | 10761 | -0.023 | 0.18 | NO |
| 24 | DIAPH1 | DIAPH1 | DIAPH1 | 11468 | -0.032 | 0.16 | NO |
| 25 | ARPC5 | ARPC5 | ARPC5 | 11502 | -0.032 | 0.17 | NO |
| 26 | PFN1 | PFN1 | PFN1 | 11981 | -0.04 | 0.16 | NO |
| 27 | SRC | SRC | SRC | 12472 | -0.048 | 0.16 | NO |
| 28 | ARPC1B | ARPC1B | ARPC1B | 13225 | -0.063 | 0.15 | NO |
| 29 | ARHGAP4 | ARHGAP4 | ARHGAP4 | 13538 | -0.07 | 0.16 | NO |
| 30 | MYL2 | MYL2 | MYL2 | 14368 | -0.094 | 0.16 | NO |
| 31 | ARHGEF1 | ARHGEF1 | ARHGEF1 | 14741 | -0.11 | 0.19 | NO |
Figure S7. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA GPCR PATHWAY.
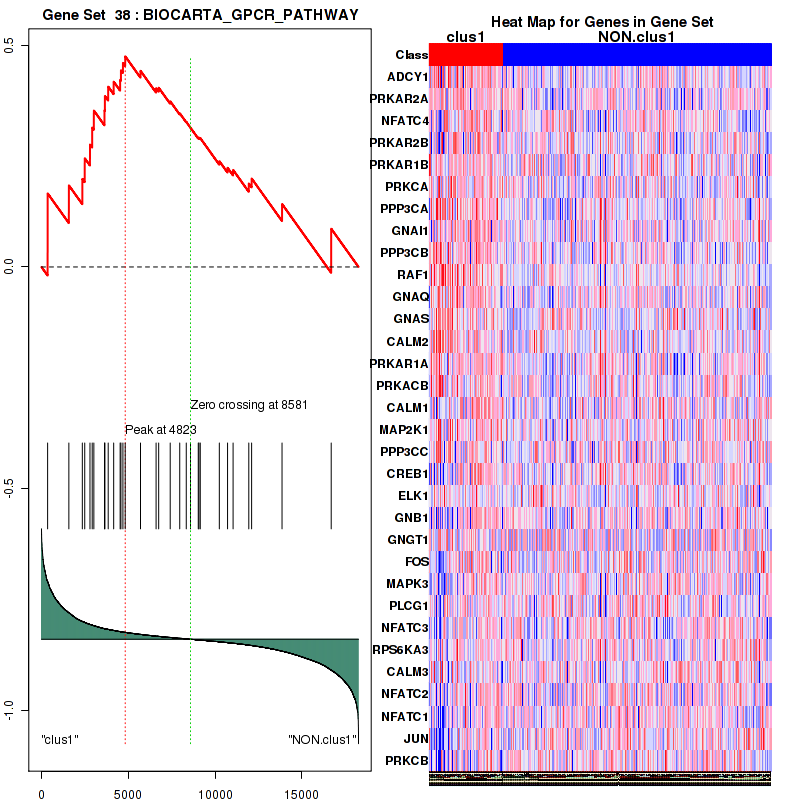
Figure S8. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA GPCR PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
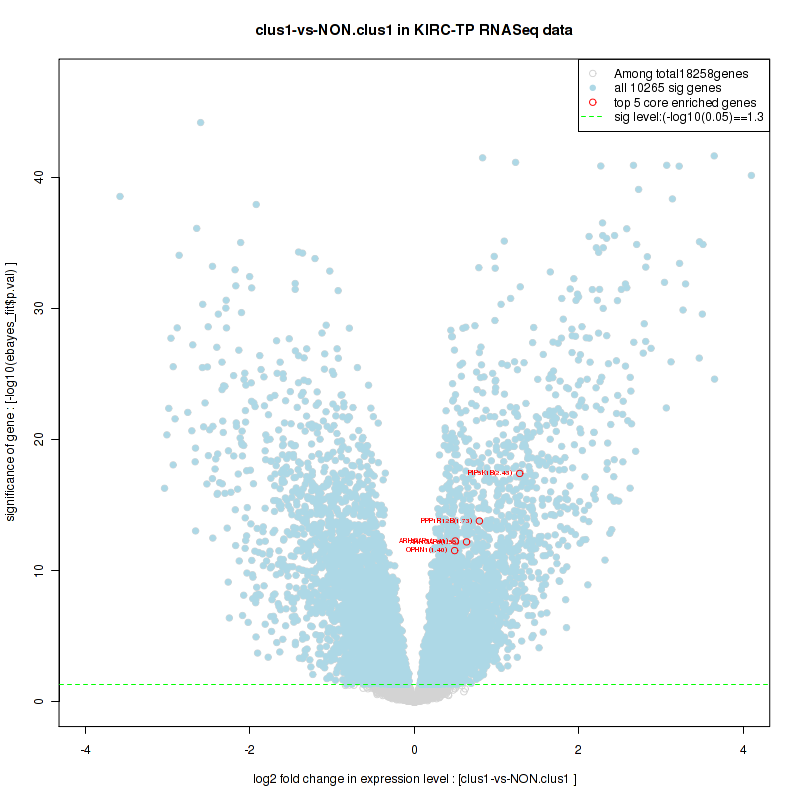
Table S5. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | SFRP1 | SFRP1 | SFRP1 | 82 | 0.54 | 0.034 | YES |
| 2 | NKD1 | NKD1 | NKD1 | 294 | 0.4 | 0.052 | YES |
| 3 | WNT4 | WNT4 | WNT4 | 374 | 0.37 | 0.074 | YES |
| 4 | AXIN2 | AXIN2 | AXIN2 | 434 | 0.35 | 0.097 | YES |
| 5 | WNT7A | WNT7A | WNT7A | 473 | 0.34 | 0.12 | YES |
| 6 | WNT11 | WNT11 | WNT11 | 479 | 0.34 | 0.14 | YES |
| 7 | WNT2 | WNT2 | WNT2 | 529 | 0.33 | 0.16 | YES |
| 8 | FZD9 | FZD9 | FZD9 | 576 | 0.32 | 0.18 | YES |
| 9 | CAMK2A | CAMK2A | CAMK2A | 697 | 0.29 | 0.2 | YES |
| 10 | DKK2 | DKK2 | DKK2 | 808 | 0.27 | 0.21 | YES |
| 11 | CAMK2B | CAMK2B | CAMK2B | 847 | 0.26 | 0.23 | YES |
| 12 | FZD7 | FZD7 | FZD7 | 869 | 0.26 | 0.25 | YES |
| 13 | WNT7B | WNT7B | WNT7B | 881 | 0.26 | 0.26 | YES |
| 14 | DAAM1 | DAAM1 | DAAM1 | 901 | 0.25 | 0.28 | YES |
| 15 | FZD10 | FZD10 | FZD10 | 923 | 0.25 | 0.3 | YES |
| 16 | WNT9B | WNT9B | WNT9B | 1213 | 0.21 | 0.3 | YES |
| 17 | FZD2 | FZD2 | FZD2 | 1235 | 0.21 | 0.31 | YES |
| 18 | SFRP2 | SFRP2 | SFRP2 | 1276 | 0.2 | 0.32 | YES |
| 19 | TBL1Y | TBL1Y | TBL1Y | 1387 | 0.19 | 0.33 | YES |
| 20 | VANGL2 | VANGL2 | VANGL2 | 1492 | 0.18 | 0.34 | YES |
| 21 | PRICKLE1 | PRICKLE1 | PRICKLE1 | 1548 | 0.18 | 0.35 | YES |
| 22 | MMP7 | MMP7 | MMP7 | 1698 | 0.16 | 0.35 | YES |
| 23 | WNT5A | WNT5A | WNT5A | 1750 | 0.16 | 0.36 | YES |
| 24 | CXXC4 | CXXC4 | CXXC4 | 1853 | 0.15 | 0.37 | YES |
| 25 | FZD3 | FZD3 | FZD3 | 1867 | 0.15 | 0.38 | YES |
| 26 | CHP2 | CHP2 | CHP2 | 1970 | 0.14 | 0.38 | YES |
| 27 | RAC3 | RAC3 | RAC3 | 2111 | 0.13 | 0.38 | YES |
| 28 | PORCN | PORCN | PORCN | 2124 | 0.13 | 0.39 | YES |
| 29 | FRAT2 | FRAT2 | FRAT2 | 2160 | 0.13 | 0.4 | YES |
| 30 | TCF7L1 | TCF7L1 | TCF7L1 | 2223 | 0.12 | 0.4 | YES |
| 31 | NFATC4 | NFATC4 | NFATC4 | 2354 | 0.12 | 0.4 | YES |
| 32 | WNT6 | WNT6 | WNT6 | 2428 | 0.11 | 0.41 | YES |
| 33 | FRAT1 | FRAT1 | FRAT1 | 2550 | 0.11 | 0.41 | YES |
| 34 | PPP2R5A | PPP2R5A | PPP2R5A | 2659 | 0.1 | 0.41 | YES |
| 35 | LEF1 | LEF1 | LEF1 | 2713 | 0.1 | 0.42 | YES |
| 36 | PRKCA | PRKCA | PRKCA | 2922 | 0.092 | 0.41 | YES |
| 37 | MAPK10 | MAPK10 | MAPK10 | 2972 | 0.09 | 0.42 | YES |
| 38 | PPP3CA | PPP3CA | PPP3CA | 3017 | 0.089 | 0.42 | YES |
| 39 | WIF1 | WIF1 | WIF1 | 3046 | 0.088 | 0.42 | YES |
| 40 | PPP2R5E | PPP2R5E | PPP2R5E | 3078 | 0.087 | 0.43 | YES |
| 41 | WNT5B | WNT5B | WNT5B | 3265 | 0.08 | 0.42 | YES |
| 42 | LRP6 | LRP6 | LRP6 | 3351 | 0.078 | 0.42 | YES |
| 43 | FZD6 | FZD6 | FZD6 | 3366 | 0.077 | 0.43 | YES |
| 44 | PSEN1 | PSEN1 | PSEN1 | 3392 | 0.076 | 0.43 | YES |
| 45 | WNT9A | WNT9A | WNT9A | 3451 | 0.074 | 0.44 | YES |
| 46 | DKK1 | DKK1 | DKK1 | 3532 | 0.072 | 0.44 | YES |
| 47 | PPP2R5C | PPP2R5C | PPP2R5C | 3539 | 0.072 | 0.44 | YES |
| 48 | RHOA | RHOA | RHOA | 3584 | 0.07 | 0.44 | YES |
| 49 | CTNNB1 | CTNNB1 | CTNNB1 | 3624 | 0.069 | 0.45 | YES |
| 50 | PPP3CB | PPP3CB | PPP3CB | 3652 | 0.069 | 0.45 | YES |
| 51 | TCF7 | TCF7 | TCF7 | 3722 | 0.067 | 0.45 | YES |
| 52 | SMAD4 | SMAD4 | SMAD4 | 3739 | 0.067 | 0.46 | YES |
| 53 | WNT3A | WNT3A | WNT3A | 3793 | 0.066 | 0.46 | YES |
| 54 | GSK3B | GSK3B | GSK3B | 3846 | 0.065 | 0.46 | YES |
| 55 | NFAT5 | NFAT5 | NFAT5 | 4102 | 0.059 | 0.45 | NO |
| 56 | SMAD2 | SMAD2 | SMAD2 | 4140 | 0.058 | 0.45 | NO |
| 57 | CHP | CHP | CHP | 4294 | 0.055 | 0.45 | NO |
| 58 | SFRP5 | SFRP5 | SFRP5 | 4448 | 0.052 | 0.44 | NO |
| 59 | FZD8 | FZD8 | FZD8 | 4475 | 0.051 | 0.44 | NO |
| 60 | MAPK8 | MAPK8 | MAPK8 | 4785 | 0.046 | 0.43 | NO |
| 61 | PRKACB | PRKACB | PRKACB | 4823 | 0.045 | 0.43 | NO |
| 62 | CTNNBIP1 | CTNNBIP1 | CTNNBIP1 | 4946 | 0.043 | 0.43 | NO |
| 63 | DAAM2 | DAAM2 | DAAM2 | 5227 | 0.039 | 0.42 | NO |
| 64 | SIAH1 | SIAH1 | SIAH1 | 5273 | 0.038 | 0.42 | NO |
| 65 | ROCK1 | ROCK1 | ROCK1 | 5280 | 0.038 | 0.42 | NO |
| 66 | PPP2R1B | PPP2R1B | PPP2R1B | 5432 | 0.036 | 0.41 | NO |
| 67 | CSNK1A1L | CSNK1A1L | CSNK1A1L | 5484 | 0.035 | 0.41 | NO |
| 68 | CHD8 | CHD8 | CHD8 | 5523 | 0.034 | 0.41 | NO |
| 69 | FBXW11 | FBXW11 | FBXW11 | 5534 | 0.034 | 0.42 | NO |
| 70 | MAP3K7 | MAP3K7 | MAP3K7 | 5657 | 0.032 | 0.41 | NO |
| 71 | ROCK2 | ROCK2 | ROCK2 | 5664 | 0.032 | 0.41 | NO |
| 72 | SKP1 | SKP1 | SKP1 | 5816 | 0.03 | 0.41 | NO |
| 73 | PRKACA | PRKACA | PRKACA | 5827 | 0.03 | 0.41 | NO |
| 74 | WNT8B | WNT8B | WNT8B | 5931 | 0.029 | 0.4 | NO |
| 75 | CSNK2A1 | CSNK2A1 | CSNK2A1 | 6036 | 0.027 | 0.4 | NO |
| 76 | CSNK2A2 | CSNK2A2 | CSNK2A2 | 6144 | 0.026 | 0.4 | NO |
| 77 | PPP2R5D | PPP2R5D | PPP2R5D | 6358 | 0.023 | 0.39 | NO |
| 78 | PRICKLE2 | PRICKLE2 | PRICKLE2 | 6561 | 0.021 | 0.38 | NO |
| 79 | RUVBL1 | RUVBL1 | RUVBL1 | 6686 | 0.02 | 0.37 | NO |
| 80 | PPP3CC | PPP3CC | PPP3CC | 6750 | 0.019 | 0.37 | NO |
| 81 | SOX17 | SOX17 | SOX17 | 6841 | 0.018 | 0.37 | NO |
| 82 | BTRC | BTRC | BTRC | 7082 | 0.015 | 0.35 | NO |
| 83 | CAMK2D | CAMK2D | CAMK2D | 7114 | 0.015 | 0.35 | NO |
| 84 | PPP2CB | PPP2CB | PPP2CB | 7143 | 0.014 | 0.35 | NO |
| 85 | CUL1 | CUL1 | CUL1 | 7320 | 0.012 | 0.34 | NO |
| 86 | DVL3 | DVL3 | DVL3 | 7351 | 0.012 | 0.34 | NO |
| 87 | EP300 | EP300 | EP300 | 7375 | 0.012 | 0.34 | NO |
| 88 | PPP2R1A | PPP2R1A | PPP2R1A | 7410 | 0.012 | 0.34 | NO |
| 89 | CREBBP | CREBBP | CREBBP | 7421 | 0.012 | 0.34 | NO |
| 90 | PPARD | PPARD | PPARD | 7532 | 0.01 | 0.34 | NO |
| 91 | DVL2 | DVL2 | DVL2 | 7756 | 0.008 | 0.32 | NO |
| 92 | LRP5 | LRP5 | LRP5 | 7838 | 0.0072 | 0.32 | NO |
| 93 | PPP2CA | PPP2CA | PPP2CA | 7844 | 0.0072 | 0.32 | NO |
| 94 | RAC1 | RAC1 | RAC1 | 7856 | 0.0071 | 0.32 | NO |
| 95 | TCF7L2 | TCF7L2 | TCF7L2 | 8131 | 0.0046 | 0.31 | NO |
| 96 | SENP2 | SENP2 | SENP2 | 8266 | 0.0032 | 0.3 | NO |
| 97 | APC | APC | APC | 8501 | 0.00067 | 0.29 | NO |
| 98 | PRKX | PRKX | PRKX | 8539 | 0.0004 | 0.28 | NO |
| 99 | SMAD3 | SMAD3 | SMAD3 | 8644 | -0.00078 | 0.28 | NO |
| 100 | CTBP1 | CTBP1 | CTBP1 | 8673 | -0.001 | 0.28 | NO |
| 101 | AXIN1 | AXIN1 | AXIN1 | 9215 | -0.0063 | 0.25 | NO |
| 102 | PPP3R1 | PPP3R1 | PPP3R1 | 9272 | -0.0068 | 0.24 | NO |
| 103 | CAMK2G | CAMK2G | CAMK2G | 9293 | -0.007 | 0.24 | NO |
| 104 | VANGL1 | VANGL1 | VANGL1 | 9352 | -0.0077 | 0.24 | NO |
| 105 | CTBP2 | CTBP2 | CTBP2 | 9452 | -0.0087 | 0.24 | NO |
| 106 | RBX1 | RBX1 | RBX1 | 9516 | -0.0094 | 0.23 | NO |
| 107 | CCND3 | CCND3 | CCND3 | 9559 | -0.0098 | 0.23 | NO |
| 108 | CSNK1E | CSNK1E | CSNK1E | 9786 | -0.012 | 0.22 | NO |
| 109 | CCND2 | CCND2 | CCND2 | 9809 | -0.012 | 0.22 | NO |
| 110 | NLK | NLK | NLK | 9827 | -0.012 | 0.22 | NO |
| 111 | DVL1 | DVL1 | DVL1 | 9928 | -0.013 | 0.22 | NO |
| 112 | CSNK2B | CSNK2B | CSNK2B | 10190 | -0.016 | 0.2 | NO |
| 113 | NFATC3 | NFATC3 | NFATC3 | 10243 | -0.017 | 0.2 | NO |
| 114 | TBL1X | TBL1X | TBL1X | 10295 | -0.017 | 0.2 | NO |
| 115 | CSNK1A1 | CSNK1A1 | CSNK1A1 | 10525 | -0.02 | 0.19 | NO |
| 116 | MAPK9 | MAPK9 | MAPK9 | 10740 | -0.023 | 0.18 | NO |
| 117 | TP53 | TP53 | TP53 | 10780 | -0.023 | 0.18 | NO |
| 118 | PLCB3 | PLCB3 | PLCB3 | 11190 | -0.028 | 0.16 | NO |
| 119 | CACYBP | CACYBP | CACYBP | 11742 | -0.036 | 0.13 | NO |
| 120 | SFRP4 | SFRP4 | SFRP4 | 11774 | -0.036 | 0.13 | NO |
| 121 | NFATC2 | NFATC2 | NFATC2 | 11938 | -0.039 | 0.12 | NO |
| 122 | NFATC1 | NFATC1 | NFATC1 | 12103 | -0.041 | 0.12 | NO |
| 123 | FZD4 | FZD4 | FZD4 | 12617 | -0.05 | 0.094 | NO |
| 124 | APC2 | APC2 | APC2 | 12767 | -0.054 | 0.089 | NO |
| 125 | FOSL1 | FOSL1 | FOSL1 | 13077 | -0.059 | 0.076 | NO |
| 126 | WNT16 | WNT16 | WNT16 | 13175 | -0.062 | 0.076 | NO |
| 127 | TBL1XR1 | TBL1XR1 | TBL1XR1 | 13218 | -0.062 | 0.078 | NO |
| 128 | WNT2B | WNT2B | WNT2B | 13310 | -0.064 | 0.077 | NO |
| 129 | PLCB4 | PLCB4 | PLCB4 | 13605 | -0.072 | 0.066 | NO |
| 130 | JUN | JUN | JUN | 13855 | -0.079 | 0.058 | NO |
| 131 | WNT10A | WNT10A | WNT10A | 13945 | -0.081 | 0.059 | NO |
| 132 | CCND1 | CCND1 | CCND1 | 14009 | -0.083 | 0.062 | NO |
| 133 | NKD2 | NKD2 | NKD2 | 14675 | -0.1 | 0.032 | NO |
| 134 | PRKCG | PRKCG | PRKCG | 14786 | -0.11 | 0.034 | NO |
| 135 | PPP3R2 | PPP3R2 | PPP3R2 | 15055 | -0.12 | 0.028 | NO |
| 136 | FZD1 | FZD1 | FZD1 | 15552 | -0.14 | 0.011 | NO |
| 137 | MYC | MYC | MYC | 15790 | -0.15 | 0.0083 | NO |
| 138 | WNT3 | WNT3 | WNT3 | 15824 | -0.15 | 0.017 | NO |
| 139 | WNT10B | WNT10B | WNT10B | 15843 | -0.15 | 0.027 | NO |
| 140 | PRKCB | PRKCB | PRKCB | 16685 | -0.2 | -0.0044 | NO |
| 141 | PLCB1 | PLCB1 | PLCB1 | 16818 | -0.21 | 0.0037 | NO |
| 142 | FZD5 | FZD5 | FZD5 | 16822 | -0.21 | 0.019 | NO |
| 143 | RAC2 | RAC2 | RAC2 | 17187 | -0.25 | 0.017 | NO |
| 144 | PLCB2 | PLCB2 | PLCB2 | 17454 | -0.28 | 0.022 | NO |
| 145 | WNT1 | WNT1 | WNT1 | 17674 | -0.31 | 0.032 | NO |
Figure S9. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA CREB PATHWAY.
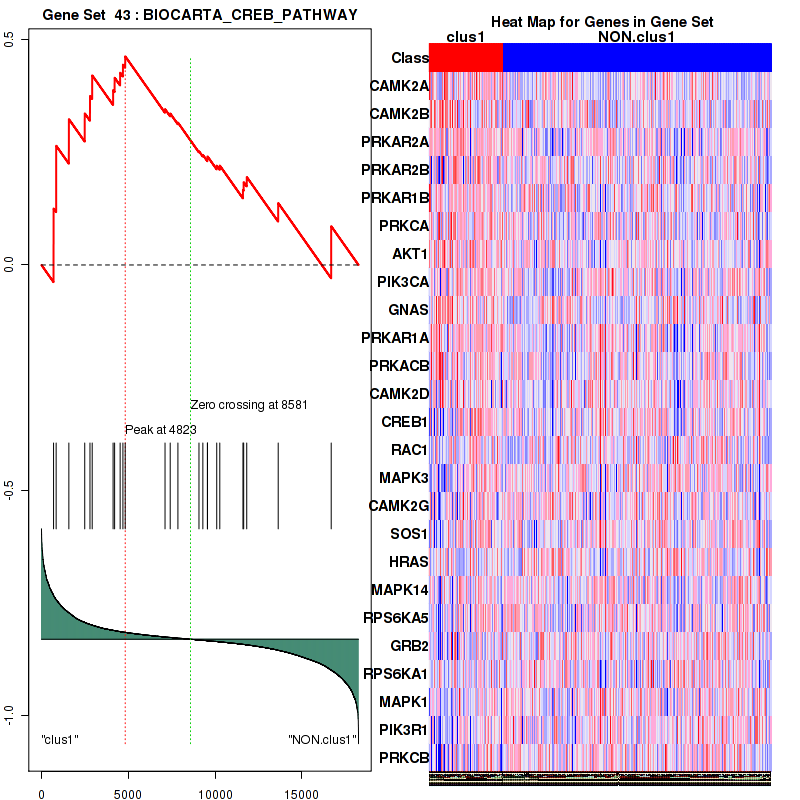
Figure S10. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA CREB PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
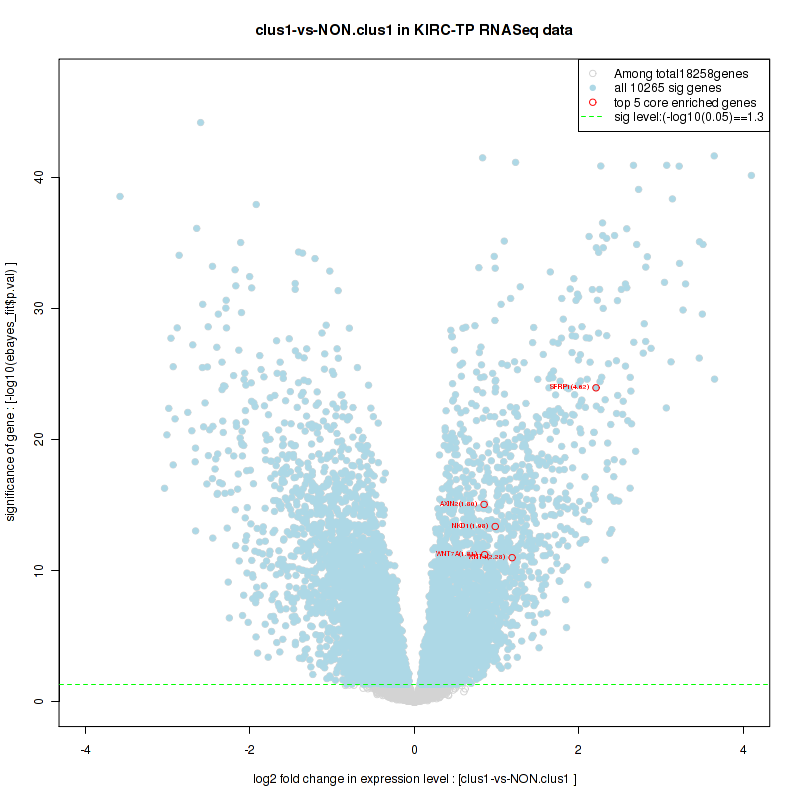
Table S6. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | FGF9 | FGF9 | FGF9 | 1 | 0.72 | 0.11 | YES |
| 2 | PDGFRA | PDGFRA | PDGFRA | 131 | 0.49 | 0.17 | YES |
| 3 | FGF7 | FGF7 | FGF7 | 189 | 0.45 | 0.24 | YES |
| 4 | FGF1 | FGF1 | FGF1 | 320 | 0.39 | 0.29 | YES |
| 5 | FGF13 | FGF13 | FGF13 | 389 | 0.37 | 0.34 | YES |
| 6 | FGF12 | FGF12 | FGF12 | 482 | 0.34 | 0.38 | YES |
| 7 | FGF14 | FGF14 | FGF14 | 775 | 0.27 | 0.41 | YES |
| 8 | IGF1 | IGF1 | IGF1 | 907 | 0.25 | 0.44 | YES |
| 9 | FGF18 | FGF18 | FGF18 | 987 | 0.24 | 0.47 | YES |
| 10 | EGF | EGF | EGF | 1856 | 0.15 | 0.44 | NO |
| 11 | BRAF | BRAF | BRAF | 3062 | 0.087 | 0.39 | NO |
| 12 | FGFR1 | FGFR1 | FGFR1 | 3154 | 0.084 | 0.4 | NO |
| 13 | MITF | MITF | MITF | 3266 | 0.08 | 0.4 | NO |
| 14 | IGF1R | IGF1R | IGF1R | 3519 | 0.072 | 0.4 | NO |
| 15 | RAF1 | RAF1 | RAF1 | 3843 | 0.065 | 0.39 | NO |
| 16 | PDGFC | PDGFC | PDGFC | 3908 | 0.063 | 0.4 | NO |
| 17 | AKT1 | AKT1 | AKT1 | 4126 | 0.059 | 0.39 | NO |
| 18 | PIK3CA | PIK3CA | PIK3CA | 4213 | 0.056 | 0.4 | NO |
| 19 | PIK3R3 | PIK3R3 | PIK3R3 | 4584 | 0.049 | 0.38 | NO |
| 20 | ARAF | ARAF | ARAF | 4791 | 0.046 | 0.38 | NO |
| 21 | CDH1 | CDH1 | CDH1 | 4809 | 0.045 | 0.39 | NO |
| 22 | AKT3 | AKT3 | AKT3 | 5114 | 0.04 | 0.38 | NO |
| 23 | KRAS | KRAS | KRAS | 5274 | 0.038 | 0.37 | NO |
| 24 | PTEN | PTEN | PTEN | 5552 | 0.034 | 0.36 | NO |
| 25 | AKT2 | AKT2 | AKT2 | 6078 | 0.027 | 0.34 | NO |
| 26 | CDK4 | CDK4 | CDK4 | 6104 | 0.026 | 0.34 | NO |
| 27 | PDGFRB | PDGFRB | PDGFRB | 6252 | 0.025 | 0.34 | NO |
| 28 | MAP2K1 | MAP2K1 | MAP2K1 | 6602 | 0.02 | 0.32 | NO |
| 29 | CDK6 | CDK6 | CDK6 | 6926 | 0.017 | 0.3 | NO |
| 30 | FGF23 | FGF23 | FGF23 | 7154 | 0.014 | 0.29 | NO |
| 31 | PDGFD | PDGFD | PDGFD | 8065 | 0.0053 | 0.24 | NO |
| 32 | CDKN1A | CDKN1A | CDKN1A | 8102 | 0.005 | 0.24 | NO |
| 33 | PIK3R2 | PIK3R2 | PIK3R2 | 8205 | 0.0037 | 0.24 | NO |
| 34 | MAP2K2 | MAP2K2 | MAP2K2 | 8354 | 0.0022 | 0.23 | NO |
| 35 | FGF2 | FGF2 | FGF2 | 8769 | -0.0019 | 0.21 | NO |
| 36 | MAPK3 | MAPK3 | MAPK3 | 9071 | -0.005 | 0.19 | NO |
| 37 | HRAS | HRAS | HRAS | 9562 | -0.0099 | 0.17 | NO |
| 38 | HGF | HGF | HGF | 9895 | -0.013 | 0.15 | NO |
| 39 | BAD | BAD | BAD | 10023 | -0.014 | 0.14 | NO |
| 40 | PDGFA | PDGFA | PDGFA | 10103 | -0.015 | 0.14 | NO |
| 41 | FGF8 | FGF8 | FGF8 | 10294 | -0.017 | 0.14 | NO |
| 42 | TP53 | TP53 | TP53 | 10780 | -0.023 | 0.11 | NO |
| 43 | PDGFB | PDGFB | PDGFB | 11761 | -0.036 | 0.063 | NO |
| 44 | MAPK1 | MAPK1 | MAPK1 | 11822 | -0.037 | 0.065 | NO |
| 45 | NRAS | NRAS | NRAS | 11870 | -0.038 | 0.068 | NO |
| 46 | RB1 | RB1 | RB1 | 12567 | -0.049 | 0.038 | NO |
| 47 | PIK3CB | PIK3CB | PIK3CB | 12586 | -0.05 | 0.044 | NO |
| 48 | E2F3 | E2F3 | E2F3 | 12723 | -0.052 | 0.044 | NO |
| 49 | MET | MET | MET | 12800 | -0.054 | 0.048 | NO |
| 50 | MDM2 | MDM2 | MDM2 | 13396 | -0.067 | 0.025 | NO |
| 51 | FGF17 | FGF17 | FGF17 | 13462 | -0.068 | 0.032 | NO |
| 52 | PIK3R1 | PIK3R1 | PIK3R1 | 13632 | -0.073 | 0.033 | NO |
| 53 | EGFR | EGFR | EGFR | 13768 | -0.076 | 0.037 | NO |
| 54 | CCND1 | CCND1 | CCND1 | 14009 | -0.083 | 0.036 | NO |
| 55 | PIK3CD | PIK3CD | PIK3CD | 14676 | -0.1 | 0.015 | NO |
| 56 | FGF5 | FGF5 | FGF5 | 14723 | -0.11 | 0.028 | NO |
| 57 | FGF20 | FGF20 | FGF20 | 15402 | -0.13 | 0.01 | NO |
| 58 | FGF11 | FGF11 | FGF11 | 15892 | -0.15 | 0.0062 | NO |
| 59 | PIK3CG | PIK3CG | PIK3CG | 16171 | -0.17 | 0.016 | NO |
| 60 | CDKN2A | CDKN2A | CDKN2A | 16564 | -0.19 | 0.023 | NO |
| 61 | PIK3R5 | PIK3R5 | PIK3R5 | 16760 | -0.21 | 0.043 | NO |
| 62 | E2F2 | E2F2 | E2F2 | 17347 | -0.26 | 0.05 | NO |
Figure S11. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG OXIDATIVE PHOSPHORYLATION.
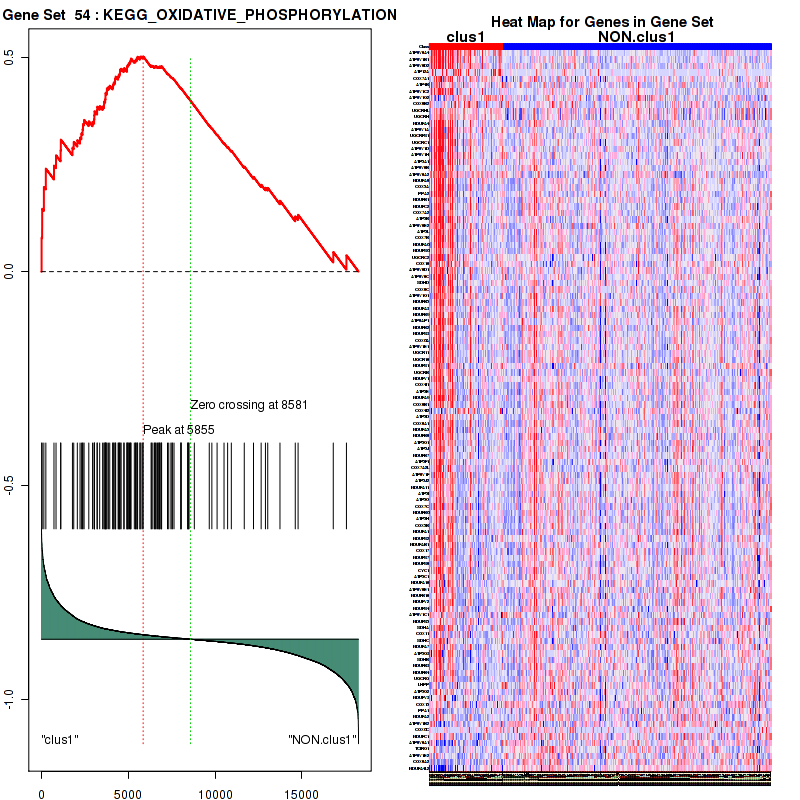
Figure S12. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG OXIDATIVE PHOSPHORYLATION, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
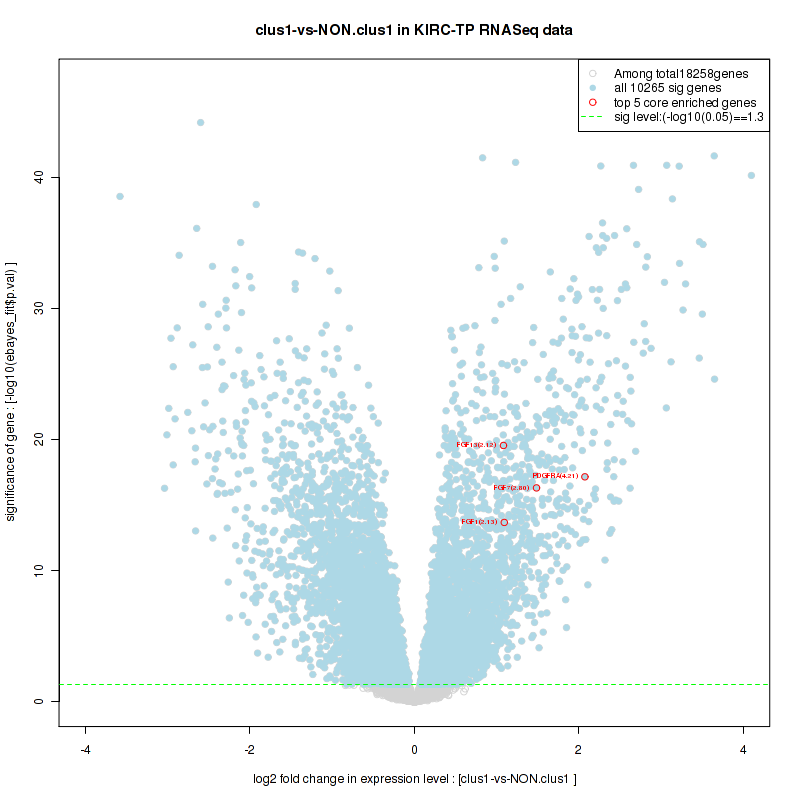
Table S7. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | KCNJ1 | KCNJ1 | KCNJ1 | 86 | 0.53 | 0.1 | YES |
| 2 | FXYD4 | FXYD4 | FXYD4 | 158 | 0.47 | 0.19 | YES |
| 3 | SCNN1A | SCNN1A | SCNN1A | 178 | 0.46 | 0.28 | YES |
| 4 | NR3C2 | NR3C2 | NR3C2 | 550 | 0.32 | 0.32 | YES |
| 5 | IGF1 | IGF1 | IGF1 | 907 | 0.25 | 0.35 | YES |
| 6 | HSD11B2 | HSD11B2 | HSD11B2 | 1183 | 0.21 | 0.38 | YES |
| 7 | IRS1 | IRS1 | IRS1 | 1223 | 0.21 | 0.42 | YES |
| 8 | HSD11B1 | HSD11B1 | HSD11B1 | 1256 | 0.2 | 0.46 | YES |
| 9 | SCNN1G | SCNN1G | SCNN1G | 1337 | 0.2 | 0.49 | YES |
| 10 | ATP1A2 | ATP1A2 | ATP1A2 | 1427 | 0.19 | 0.52 | YES |
| 11 | NEDD4L | NEDD4L | NEDD4L | 1887 | 0.15 | 0.52 | YES |
| 12 | IRS2 | IRS2 | IRS2 | 1940 | 0.14 | 0.55 | YES |
| 13 | PRKCA | PRKCA | PRKCA | 2922 | 0.092 | 0.51 | NO |
| 14 | ATP1A1 | ATP1A1 | ATP1A1 | 2947 | 0.092 | 0.53 | NO |
| 15 | SCNN1B | SCNN1B | SCNN1B | 3012 | 0.089 | 0.55 | NO |
| 16 | PIK3CA | PIK3CA | PIK3CA | 4213 | 0.056 | 0.49 | NO |
| 17 | PIK3R3 | PIK3R3 | PIK3R3 | 4584 | 0.049 | 0.48 | NO |
| 18 | ATP1B2 | ATP1B2 | ATP1B2 | 4940 | 0.043 | 0.47 | NO |
| 19 | SFN | SFN | SFN | 4995 | 0.042 | 0.47 | NO |
| 20 | KRAS | KRAS | KRAS | 5274 | 0.038 | 0.47 | NO |
| 21 | SLC9A3R2 | SLC9A3R2 | SLC9A3R2 | 5705 | 0.032 | 0.45 | NO |
| 22 | ATP1B3 | ATP1B3 | ATP1B3 | 5990 | 0.028 | 0.44 | NO |
| 23 | ATP1A4 | ATP1A4 | ATP1A4 | 6232 | 0.025 | 0.43 | NO |
| 24 | PDPK1 | PDPK1 | PDPK1 | 8176 | 0.0041 | 0.32 | NO |
| 25 | PIK3R2 | PIK3R2 | PIK3R2 | 8205 | 0.0037 | 0.32 | NO |
| 26 | MAPK3 | MAPK3 | MAPK3 | 9071 | -0.005 | 0.28 | NO |
| 27 | INSR | INSR | INSR | 11069 | -0.026 | 0.17 | NO |
| 28 | ATP1B1 | ATP1B1 | ATP1B1 | 11626 | -0.034 | 0.15 | NO |
| 29 | MAPK1 | MAPK1 | MAPK1 | 11822 | -0.037 | 0.15 | NO |
| 30 | PIK3CB | PIK3CB | PIK3CB | 12586 | -0.05 | 0.11 | NO |
| 31 | ATP1A3 | ATP1A3 | ATP1A3 | 13011 | -0.058 | 0.1 | NO |
| 32 | PIK3R1 | PIK3R1 | PIK3R1 | 13632 | -0.073 | 0.082 | NO |
| 33 | SGK1 | SGK1 | SGK1 | 13946 | -0.081 | 0.081 | NO |
| 34 | PIK3CD | PIK3CD | PIK3CD | 14676 | -0.1 | 0.061 | NO |
| 35 | PRKCG | PRKCG | PRKCG | 14786 | -0.11 | 0.076 | NO |
| 36 | PIK3CG | PIK3CG | PIK3CG | 16171 | -0.17 | 0.033 | NO |
| 37 | PRKCB | PRKCB | PRKCB | 16685 | -0.2 | 0.045 | NO |
| 38 | PIK3R5 | PIK3R5 | PIK3R5 | 16760 | -0.21 | 0.082 | NO |
Figure S13. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG O GLYCAN BIOSYNTHESIS.
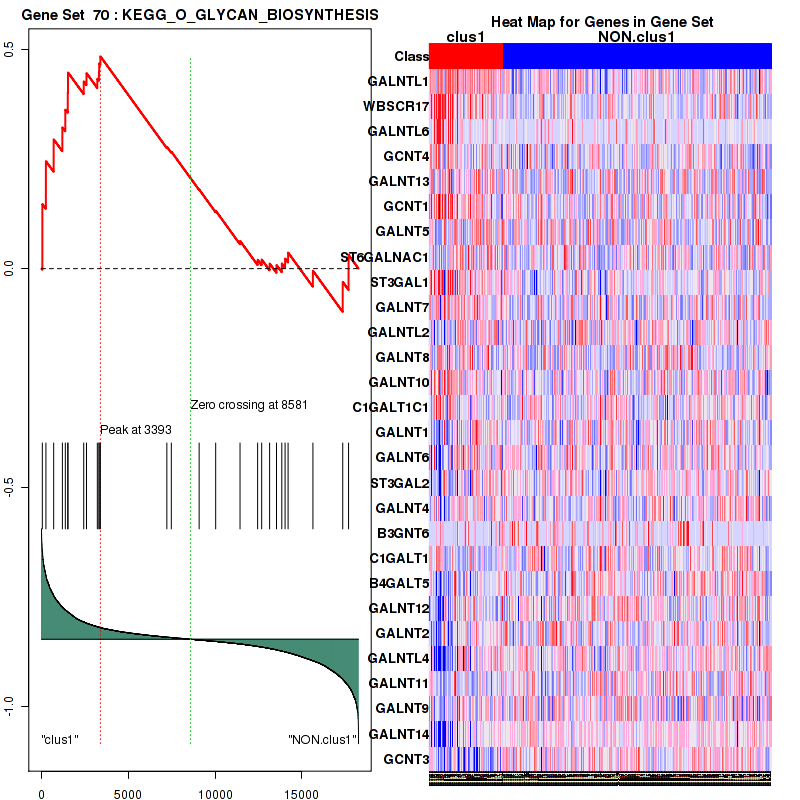
Figure S14. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG O GLYCAN BIOSYNTHESIS, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
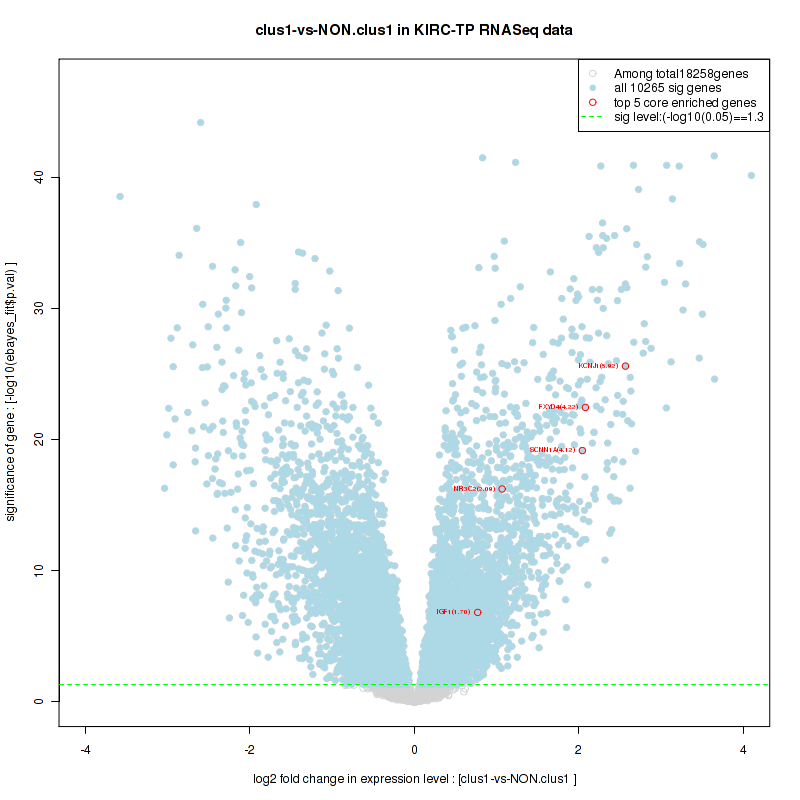
Table S8. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | ADCY1 | ADCY1 | ADCY1 | 355 | 0.38 | 0.17 | YES |
| 2 | PRKAR2A | PRKAR2A | PRKAR2A | 1578 | 0.17 | 0.18 | YES |
| 3 | NFATC4 | NFATC4 | NFATC4 | 2354 | 0.12 | 0.2 | YES |
| 4 | PRKAR2B | PRKAR2B | PRKAR2B | 2492 | 0.11 | 0.24 | YES |
| 5 | PRKAR1B | PRKAR1B | PRKAR1B | 2794 | 0.097 | 0.28 | YES |
| 6 | PRKCA | PRKCA | PRKCA | 2922 | 0.092 | 0.31 | YES |
| 7 | PPP3CA | PPP3CA | PPP3CA | 3017 | 0.089 | 0.35 | YES |
| 8 | GNAI1 | GNAI1 | GNAI1 | 3629 | 0.069 | 0.35 | YES |
| 9 | PPP3CB | PPP3CB | PPP3CB | 3652 | 0.069 | 0.39 | YES |
| 10 | RAF1 | RAF1 | RAF1 | 3843 | 0.065 | 0.41 | YES |
| 11 | GNAQ | GNAQ | GNAQ | 4157 | 0.058 | 0.42 | YES |
| 12 | GNAS | GNAS | GNAS | 4530 | 0.05 | 0.42 | YES |
| 13 | CALM2 | CALM2 | CALM2 | 4576 | 0.049 | 0.44 | YES |
| 14 | PRKAR1A | PRKAR1A | PRKAR1A | 4691 | 0.047 | 0.46 | YES |
| 15 | PRKACB | PRKACB | PRKACB | 4823 | 0.045 | 0.48 | YES |
| 16 | CALM1 | CALM1 | CALM1 | 5702 | 0.032 | 0.44 | NO |
| 17 | MAP2K1 | MAP2K1 | MAP2K1 | 6602 | 0.02 | 0.4 | NO |
| 18 | PPP3CC | PPP3CC | PPP3CC | 6750 | 0.019 | 0.4 | NO |
| 19 | CREB1 | CREB1 | CREB1 | 7413 | 0.012 | 0.37 | NO |
| 20 | ELK1 | ELK1 | ELK1 | 7962 | 0.0061 | 0.35 | NO |
| 21 | GNB1 | GNB1 | GNB1 | 8337 | 0.0025 | 0.33 | NO |
| 22 | GNGT1 | GNGT1 | GNGT1 | 8583 | -0.000015 | 0.31 | NO |
| 23 | FOS | FOS | FOS | 9031 | -0.0046 | 0.29 | NO |
| 24 | MAPK3 | MAPK3 | MAPK3 | 9071 | -0.005 | 0.29 | NO |
| 25 | PLCG1 | PLCG1 | PLCG1 | 9144 | -0.0056 | 0.29 | NO |
| 26 | NFATC3 | NFATC3 | NFATC3 | 10243 | -0.017 | 0.24 | NO |
| 27 | RPS6KA3 | RPS6KA3 | RPS6KA3 | 10715 | -0.022 | 0.22 | NO |
| 28 | CALM3 | CALM3 | CALM3 | 11035 | -0.026 | 0.22 | NO |
| 29 | NFATC2 | NFATC2 | NFATC2 | 11938 | -0.039 | 0.19 | NO |
| 30 | NFATC1 | NFATC1 | NFATC1 | 12103 | -0.041 | 0.2 | NO |
| 31 | JUN | JUN | JUN | 13855 | -0.079 | 0.14 | NO |
| 32 | PRKCB | PRKCB | PRKCB | 16685 | -0.2 | 0.086 | NO |
Figure S15. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG INOSITOL PHOSPHATE METABOLISM.
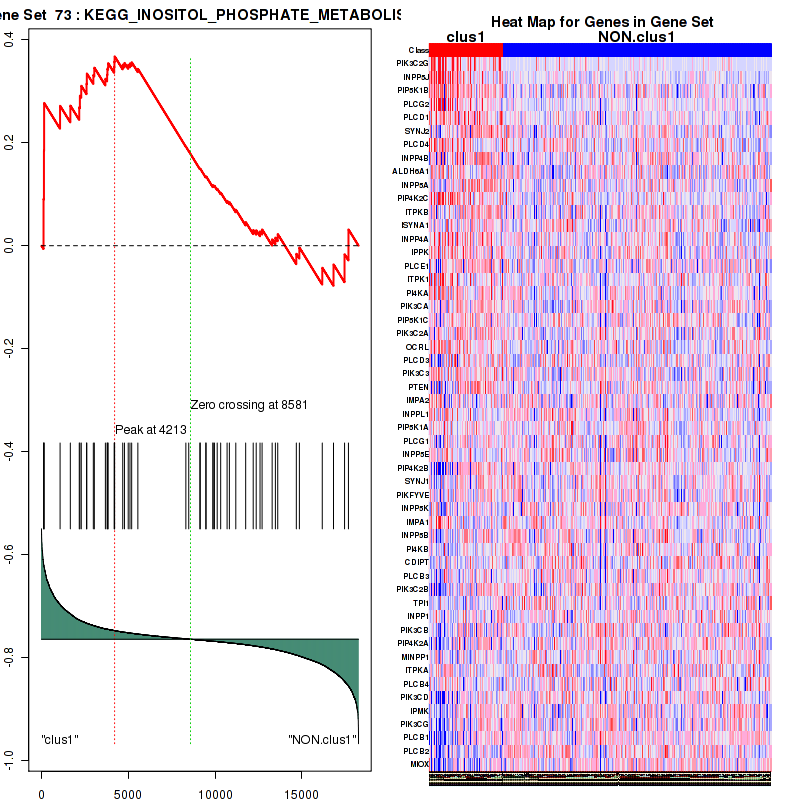
Figure S16. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG INOSITOL PHOSPHATE METABOLISM, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
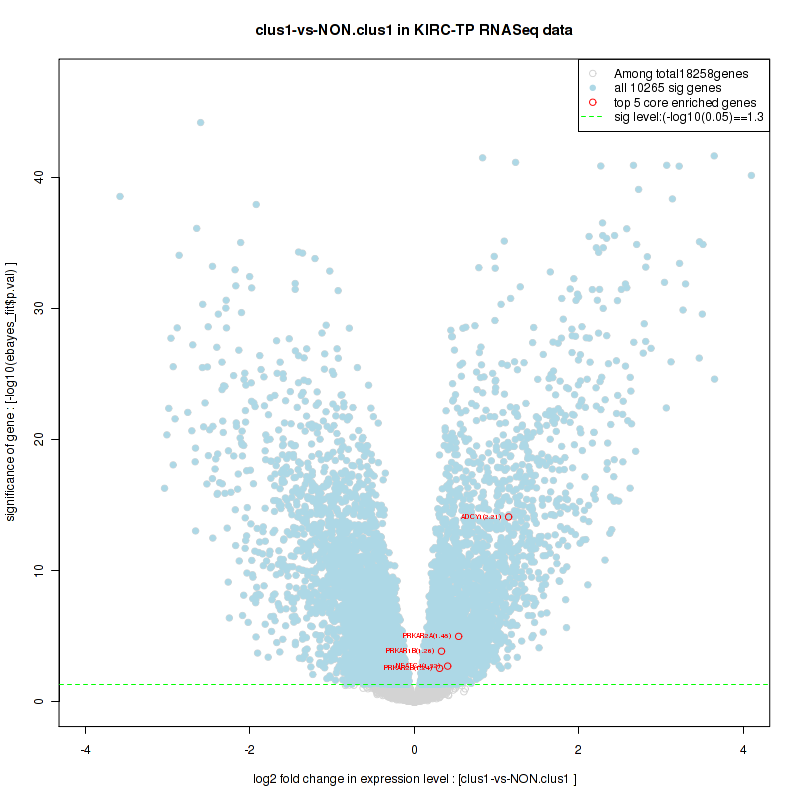
Table S9. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | CAMK2A | CAMK2A | CAMK2A | 697 | 0.29 | 0.13 | YES |
| 2 | CAMK2B | CAMK2B | CAMK2B | 847 | 0.26 | 0.26 | YES |
| 3 | PRKAR2A | PRKAR2A | PRKAR2A | 1578 | 0.17 | 0.32 | YES |
| 4 | PRKAR2B | PRKAR2B | PRKAR2B | 2492 | 0.11 | 0.34 | YES |
| 5 | PRKAR1B | PRKAR1B | PRKAR1B | 2794 | 0.097 | 0.38 | YES |
| 6 | PRKCA | PRKCA | PRKCA | 2922 | 0.092 | 0.42 | YES |
| 7 | AKT1 | AKT1 | AKT1 | 4126 | 0.059 | 0.39 | YES |
| 8 | PIK3CA | PIK3CA | PIK3CA | 4213 | 0.056 | 0.42 | YES |
| 9 | GNAS | GNAS | GNAS | 4530 | 0.05 | 0.43 | YES |
| 10 | PRKAR1A | PRKAR1A | PRKAR1A | 4691 | 0.047 | 0.44 | YES |
| 11 | PRKACB | PRKACB | PRKACB | 4823 | 0.045 | 0.46 | YES |
| 12 | CAMK2D | CAMK2D | CAMK2D | 7114 | 0.015 | 0.35 | NO |
| 13 | CREB1 | CREB1 | CREB1 | 7413 | 0.012 | 0.34 | NO |
| 14 | RAC1 | RAC1 | RAC1 | 7856 | 0.0071 | 0.32 | NO |
| 15 | MAPK3 | MAPK3 | MAPK3 | 9071 | -0.005 | 0.25 | NO |
| 16 | CAMK2G | CAMK2G | CAMK2G | 9293 | -0.007 | 0.24 | NO |
| 17 | SOS1 | SOS1 | SOS1 | 9555 | -0.0098 | 0.24 | NO |
| 18 | HRAS | HRAS | HRAS | 9562 | -0.0099 | 0.24 | NO |
| 19 | MAPK14 | MAPK14 | MAPK14 | 10093 | -0.015 | 0.22 | NO |
| 20 | RPS6KA5 | RPS6KA5 | RPS6KA5 | 10268 | -0.017 | 0.22 | NO |
| 21 | GRB2 | GRB2 | GRB2 | 11603 | -0.034 | 0.17 | NO |
| 22 | RPS6KA1 | RPS6KA1 | RPS6KA1 | 11644 | -0.034 | 0.18 | NO |
| 23 | MAPK1 | MAPK1 | MAPK1 | 11822 | -0.037 | 0.2 | NO |
| 24 | PIK3R1 | PIK3R1 | PIK3R1 | 13632 | -0.073 | 0.14 | NO |
| 25 | PRKCB | PRKCB | PRKCB | 16685 | -0.2 | 0.086 | NO |
Figure S17. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG CARDIAC MUSCLE CONTRACTION.
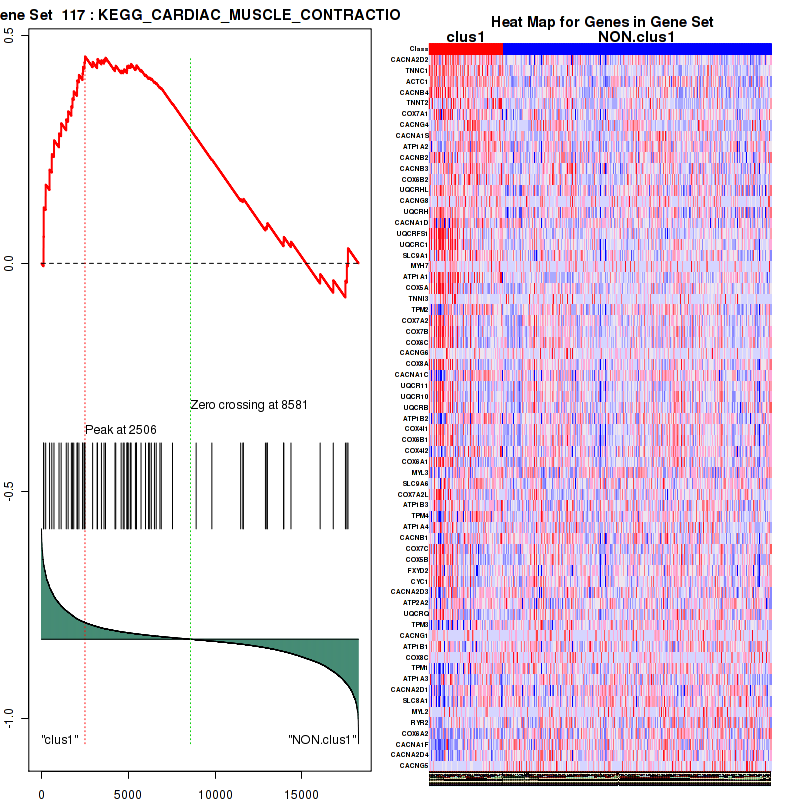
Figure S18. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG CARDIAC MUSCLE CONTRACTION, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
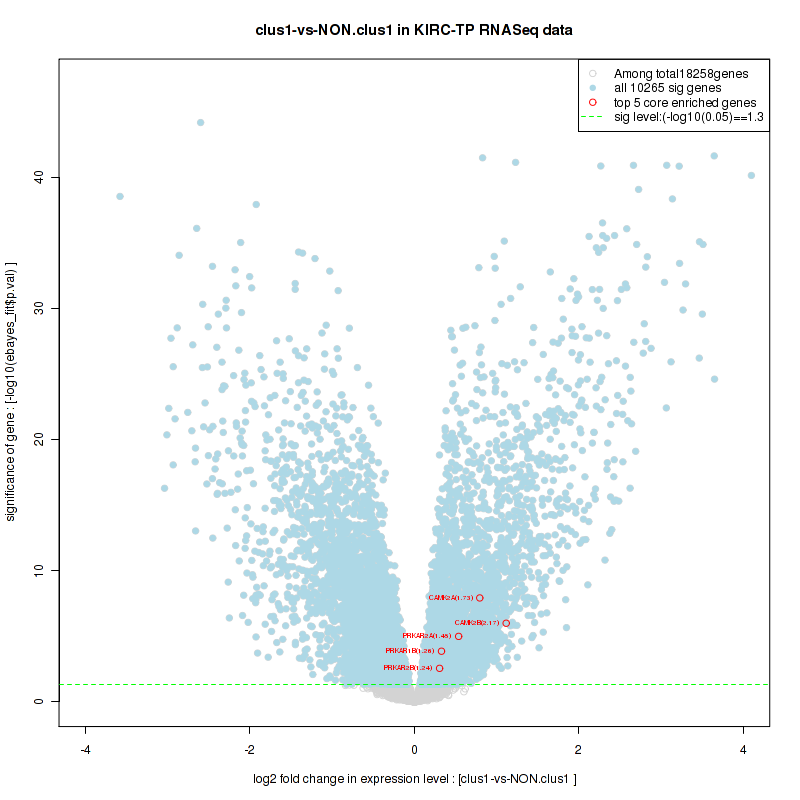
Table S10. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | ATP6V0A4 | ATP6V0A4 | ATP6V0A4 | 8 | 0.69 | 0.08 | YES |
| 2 | ATP6V1B1 | ATP6V1B1 | ATP6V1B1 | 40 | 0.59 | 0.15 | YES |
| 3 | ATP6V0D2 | ATP6V0D2 | ATP6V0D2 | 139 | 0.48 | 0.2 | YES |
| 4 | ATP12A | ATP12A | ATP12A | 251 | 0.42 | 0.24 | YES |
| 5 | COX7A1 | COX7A1 | COX7A1 | 721 | 0.28 | 0.25 | YES |
| 6 | ATP4B | ATP4B | ATP4B | 838 | 0.26 | 0.27 | YES |
| 7 | ATP6V1C2 | ATP6V1C2 | ATP6V1C2 | 1102 | 0.22 | 0.28 | YES |
| 8 | ATP6V1G2 | ATP6V1G2 | ATP6V1G2 | 1115 | 0.22 | 0.31 | YES |
| 9 | COX6B2 | COX6B2 | COX6B2 | 1785 | 0.16 | 0.29 | YES |
| 10 | UQCRHL | UQCRHL | UQCRHL | 1858 | 0.15 | 0.3 | YES |
| 11 | UQCRH | UQCRH | UQCRH | 2066 | 0.13 | 0.31 | YES |
| 12 | NDUFA4 | NDUFA4 | NDUFA4 | 2221 | 0.12 | 0.31 | YES |
| 13 | ATP6V1A | ATP6V1A | ATP6V1A | 2299 | 0.12 | 0.32 | YES |
| 14 | UQCRFS1 | UQCRFS1 | UQCRFS1 | 2367 | 0.12 | 0.33 | YES |
| 15 | UQCRC1 | UQCRC1 | UQCRC1 | 2391 | 0.12 | 0.34 | YES |
| 16 | ATP6V1D | ATP6V1D | ATP6V1D | 2454 | 0.11 | 0.36 | YES |
| 17 | ATP6V1H | ATP6V1H | ATP6V1H | 2724 | 0.1 | 0.35 | YES |
| 18 | ATP5A1 | ATP5A1 | ATP5A1 | 2943 | 0.092 | 0.35 | YES |
| 19 | ATP6V0B | ATP6V0B | ATP6V0B | 3032 | 0.088 | 0.36 | YES |
| 20 | ATP6V0A2 | ATP6V0A2 | ATP6V0A2 | 3047 | 0.088 | 0.36 | YES |
| 21 | NDUFA6 | NDUFA6 | NDUFA6 | 3055 | 0.088 | 0.38 | YES |
| 22 | COX5A | COX5A | COX5A | 3205 | 0.082 | 0.38 | YES |
| 23 | PPA2 | PPA2 | PPA2 | 3211 | 0.082 | 0.39 | YES |
| 24 | NDUFB1 | NDUFB1 | NDUFB1 | 3353 | 0.078 | 0.39 | YES |
| 25 | NDUFC2 | NDUFC2 | NDUFC2 | 3523 | 0.072 | 0.39 | YES |
| 26 | COX7A2 | COX7A2 | COX7A2 | 3603 | 0.07 | 0.39 | YES |
| 27 | ATP5B | ATP5B | ATP5B | 3610 | 0.07 | 0.4 | YES |
| 28 | ATP6V0E2 | ATP6V0E2 | ATP6V0E2 | 3670 | 0.068 | 0.4 | YES |
| 29 | ATP5L | ATP5L | ATP5L | 3686 | 0.068 | 0.41 | YES |
| 30 | COX7B | COX7B | COX7B | 3687 | 0.068 | 0.42 | YES |
| 31 | NDUFA8 | NDUFA8 | NDUFA8 | 3758 | 0.067 | 0.42 | YES |
| 32 | NDUFS8 | NDUFS8 | NDUFS8 | 3769 | 0.066 | 0.43 | YES |
| 33 | UQCRC2 | UQCRC2 | UQCRC2 | 3852 | 0.064 | 0.43 | YES |
| 34 | COX10 | COX10 | COX10 | 3930 | 0.063 | 0.44 | YES |
| 35 | ATP6V0D1 | ATP6V0D1 | ATP6V0D1 | 4097 | 0.059 | 0.43 | YES |
| 36 | ATP6V0C | ATP6V0C | ATP6V0C | 4127 | 0.058 | 0.44 | YES |
| 37 | SDHD | SDHD | SDHD | 4164 | 0.058 | 0.44 | YES |
| 38 | COX6C | COX6C | COX6C | 4235 | 0.056 | 0.44 | YES |
| 39 | ATP6V1G1 | ATP6V1G1 | ATP6V1G1 | 4243 | 0.056 | 0.45 | YES |
| 40 | NDUFB5 | NDUFB5 | NDUFB5 | 4274 | 0.055 | 0.46 | YES |
| 41 | NDUFA3 | NDUFA3 | NDUFA3 | 4413 | 0.052 | 0.46 | YES |
| 42 | NDUFB9 | NDUFB9 | NDUFB9 | 4441 | 0.052 | 0.46 | YES |
| 43 | ATP6AP1 | ATP6AP1 | ATP6AP1 | 4491 | 0.051 | 0.46 | YES |
| 44 | NDUFB2 | NDUFB2 | NDUFB2 | 4521 | 0.05 | 0.47 | YES |
| 45 | NDUFS3 | NDUFS3 | NDUFS3 | 4524 | 0.05 | 0.47 | YES |
| 46 | COX8A | COX8A | COX8A | 4590 | 0.049 | 0.47 | YES |
| 47 | ATP6V1E1 | ATP6V1E1 | ATP6V1E1 | 4740 | 0.046 | 0.47 | YES |
| 48 | UQCR11 | UQCR11 | UQCR11 | 4761 | 0.046 | 0.48 | YES |
| 49 | UQCR10 | UQCR10 | UQCR10 | 4926 | 0.043 | 0.47 | YES |
| 50 | NDUFS1 | NDUFS1 | NDUFS1 | 4927 | 0.043 | 0.48 | YES |
| 51 | UQCRB | UQCRB | UQCRB | 4937 | 0.043 | 0.48 | YES |
| 52 | NDUFV1 | NDUFV1 | NDUFV1 | 4985 | 0.042 | 0.48 | YES |
| 53 | COX4I1 | COX4I1 | COX4I1 | 5027 | 0.042 | 0.49 | YES |
| 54 | ATP5E | ATP5E | ATP5E | 5071 | 0.041 | 0.49 | YES |
| 55 | NDUFA9 | NDUFA9 | NDUFA9 | 5112 | 0.04 | 0.49 | YES |
| 56 | COX6B1 | COX6B1 | COX6B1 | 5135 | 0.04 | 0.5 | YES |
| 57 | COX4I2 | COX4I2 | COX4I2 | 5157 | 0.04 | 0.5 | YES |
| 58 | ATP5D | ATP5D | ATP5D | 5346 | 0.037 | 0.49 | YES |
| 59 | COX6A1 | COX6A1 | COX6A1 | 5413 | 0.036 | 0.49 | YES |
| 60 | NDUFA5 | NDUFA5 | NDUFA5 | 5430 | 0.036 | 0.5 | YES |
| 61 | NDUFB6 | NDUFB6 | NDUFB6 | 5498 | 0.035 | 0.5 | YES |
| 62 | ATP5G1 | ATP5G1 | ATP5G1 | 5504 | 0.035 | 0.5 | YES |
| 63 | ATP5J | ATP5J | ATP5J | 5536 | 0.034 | 0.5 | YES |
| 64 | NDUFB7 | NDUFB7 | NDUFB7 | 5668 | 0.032 | 0.5 | YES |
| 65 | ATP5F1 | ATP5F1 | ATP5F1 | 5679 | 0.032 | 0.5 | YES |
| 66 | COX7A2L | COX7A2L | COX7A2L | 5732 | 0.032 | 0.5 | YES |
| 67 | ATP6V1F | ATP6V1F | ATP6V1F | 5830 | 0.03 | 0.5 | YES |
| 68 | ATP5J2 | ATP5J2 | ATP5J2 | 5855 | 0.03 | 0.5 | YES |
| 69 | NDUFA11 | NDUFA11 | NDUFA11 | 6313 | 0.024 | 0.48 | NO |
| 70 | ATP5I | ATP5I | ATP5I | 6346 | 0.023 | 0.48 | NO |
| 71 | ATP5O | ATP5O | ATP5O | 6414 | 0.022 | 0.48 | NO |
| 72 | COX7C | COX7C | COX7C | 6505 | 0.021 | 0.48 | NO |
| 73 | NDUFB8 | NDUFB8 | NDUFB8 | 6532 | 0.021 | 0.48 | NO |
| 74 | ATP5H | ATP5H | ATP5H | 6560 | 0.021 | 0.48 | NO |
| 75 | COX5B | COX5B | COX5B | 6606 | 0.02 | 0.48 | NO |
| 76 | NDUFA1 | NDUFA1 | NDUFA1 | 6685 | 0.02 | 0.48 | NO |
| 77 | NDUFS2 | NDUFS2 | NDUFS2 | 6717 | 0.019 | 0.48 | NO |
| 78 | NDUFAB1 | NDUFAB1 | NDUFAB1 | 6761 | 0.019 | 0.48 | NO |
| 79 | COX17 | COX17 | COX17 | 6800 | 0.018 | 0.48 | NO |
| 80 | NDUFS7 | NDUFS7 | NDUFS7 | 6813 | 0.018 | 0.48 | NO |
| 81 | NDUFS6 | NDUFS6 | NDUFS6 | 6883 | 0.017 | 0.48 | NO |
| 82 | CYC1 | CYC1 | CYC1 | 6898 | 0.017 | 0.48 | NO |
| 83 | ATP5C1 | ATP5C1 | ATP5C1 | 6920 | 0.017 | 0.48 | NO |
| 84 | NDUFA10 | NDUFA10 | NDUFA10 | 7257 | 0.013 | 0.46 | NO |
| 85 | ATP6V0E1 | ATP6V0E1 | ATP6V0E1 | 7328 | 0.012 | 0.46 | NO |
| 86 | NDUFB10 | NDUFB10 | NDUFB10 | 7453 | 0.011 | 0.46 | NO |
| 87 | NDUFV2 | NDUFV2 | NDUFV2 | 7512 | 0.011 | 0.45 | NO |
| 88 | NDUFS4 | NDUFS4 | NDUFS4 | 7551 | 0.01 | 0.45 | NO |
| 89 | ATP6V1C1 | ATP6V1C1 | ATP6V1C1 | 7633 | 0.0092 | 0.45 | NO |
| 90 | NDUFS5 | NDUFS5 | NDUFS5 | 8024 | 0.0056 | 0.43 | NO |
| 91 | SDHA | SDHA | SDHA | 8032 | 0.0056 | 0.43 | NO |
| 92 | COX11 | COX11 | COX11 | 8046 | 0.0055 | 0.43 | NO |
| 93 | SDHC | SDHC | SDHC | 8418 | 0.0016 | 0.41 | NO |
| 94 | NDUFA7 | NDUFA7 | NDUFA7 | 8438 | 0.0014 | 0.41 | NO |
| 95 | ATP5G3 | ATP5G3 | ATP5G3 | 8476 | 0.00098 | 0.4 | NO |
| 96 | SDHB | SDHB | SDHB | 8493 | 0.00078 | 0.4 | NO |
| 97 | NDUFB3 | NDUFB3 | NDUFB3 | 8804 | -0.0022 | 0.39 | NO |
| 98 | NDUFB4 | NDUFB4 | NDUFB4 | 9664 | -0.011 | 0.34 | NO |
| 99 | UQCRQ | UQCRQ | UQCRQ | 9813 | -0.012 | 0.34 | NO |
| 100 | LHPP | LHPP | LHPP | 10114 | -0.015 | 0.32 | NO |
| 101 | ATP5G2 | ATP5G2 | ATP5G2 | 10512 | -0.02 | 0.3 | NO |
| 102 | NDUFV3 | NDUFV3 | NDUFV3 | 10721 | -0.022 | 0.29 | NO |
| 103 | COX15 | COX15 | COX15 | 10933 | -0.025 | 0.28 | NO |
| 104 | PPA1 | PPA1 | PPA1 | 11679 | -0.035 | 0.25 | NO |
| 105 | NDUFA2 | NDUFA2 | NDUFA2 | 12209 | -0.043 | 0.22 | NO |
| 106 | ATP6V1B2 | ATP6V1B2 | ATP6V1B2 | 12650 | -0.051 | 0.2 | NO |
| 107 | COX8C | COX8C | COX8C | 12898 | -0.056 | 0.2 | NO |
| 108 | NDUFC1 | NDUFC1 | NDUFC1 | 13031 | -0.058 | 0.2 | NO |
| 109 | ATP6V0A1 | ATP6V0A1 | ATP6V0A1 | 13734 | -0.075 | 0.17 | NO |
| 110 | TCIRG1 | TCIRG1 | TCIRG1 | 14615 | -0.1 | 0.13 | NO |
| 111 | ATP6V1E2 | ATP6V1E2 | ATP6V1E2 | 14777 | -0.11 | 0.13 | NO |
| 112 | COX6A2 | COX6A2 | COX6A2 | 16805 | -0.21 | 0.046 | NO |
| 113 | NDUFA4L2 | NDUFA4L2 | NDUFA4L2 | 17561 | -0.29 | 0.038 | NO |
Figure S19. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG WNT SIGNALING PATHWAY.
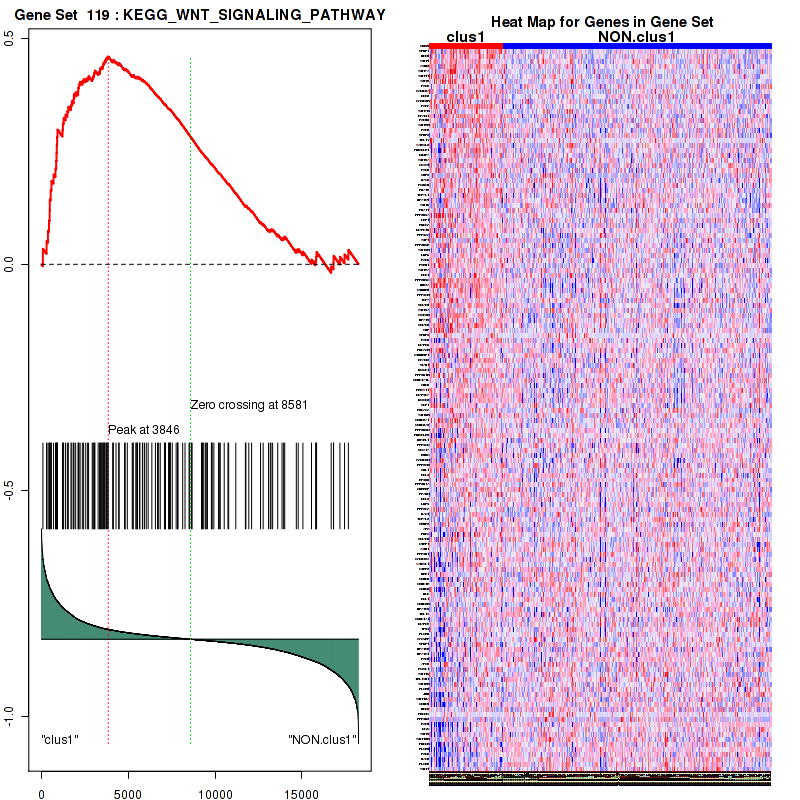
Figure S20. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG WNT SIGNALING PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
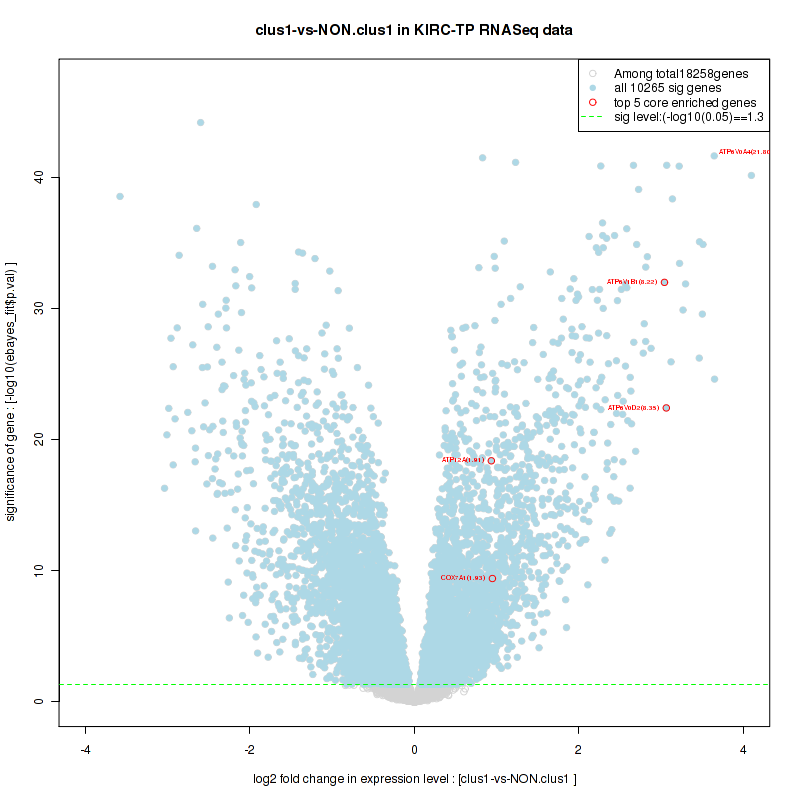
For the top enriched genes, if you want to check whether they are
-
up-regulated, please check the list of up-regulated genes
-
down-regulated, please check the list of down-regulated genes
For the top enriched genes, if you want to check whether they are
-
highly expressed genes, please check the list of high (top 30%) expressed genes
-
low expressed genes, please check the list of low (bottom 30%) expressed genes
An expression pattern of top(30%)/middle(30%)/low(30%) in this subtype against other subtypes is available in a heatmap
For the top enriched genes, if you want to check whether they are
-
significantly differently expressed genes by eBayes lm fit, please check the list of significant genes
Table 4. Get Full Table This table shows top 10 pathways which are significantly enriched in cluster clus2. It displays only significant gene sets satisfying nom.p.val.threshold (-1), fwer.p.val.threshold (-1) , fdr.q.val.threshold (0.25) and the default table is sorted by Normalized Enrichment Score (NES). Further details on NES statistics, please visit The Broad GSEA website.
| GeneSet(GS) | Size(#genes) | genes.ES.table | ES | NES | NOM.p.val | FDR.q.val | FWER.p.val | Tag.. | Gene.. | Signal | FDR..median. | glob.p.val |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BIOCARTA AT1R PATHWAY | 32 | genes.ES.table | 0.34 | 1.3 | 0.18 | 0.72 | 0.98 | 0.094 | 0.1 | 0.084 | 0.6 | 0.23 |
| BIOCARTA DEATH PATHWAY | 32 | genes.ES.table | 0.32 | 1.2 | 0.29 | 0.82 | 1 | 0.56 | 0.37 | 0.36 | 0.74 | 0.28 |
| BIOCARTA VEGF PATHWAY | 28 | genes.ES.table | 0.41 | 1.3 | 0.23 | 0.71 | 0.99 | 0.18 | 0.14 | 0.15 | 0.62 | 0.22 |
| KEGG GLYCOLYSIS GLUCONEOGENESIS | 57 | genes.ES.table | 0.39 | 1.2 | 0.21 | 0.78 | 1 | 0.14 | 0.047 | 0.13 | 0.69 | 0.26 |
| KEGG PENTOSE AND GLUCURONATE INTERCONVERSIONS | 26 | genes.ES.table | 0.67 | 1.4 | 0.11 | 0.99 | 0.96 | 0.42 | 0.1 | 0.38 | 0.8 | 0.38 |
| KEGG GALACTOSE METABOLISM | 25 | genes.ES.table | 0.41 | 1.3 | 0.18 | 0.72 | 0.99 | 0.12 | 0.025 | 0.12 | 0.63 | 0.22 |
| KEGG FATTY ACID METABOLISM | 40 | genes.ES.table | 0.52 | 1.4 | 0.14 | 0.87 | 0.96 | 0.32 | 0.12 | 0.29 | 0.7 | 0.32 |
| KEGG GLYCINE SERINE AND THREONINE METABOLISM | 30 | genes.ES.table | 0.52 | 1.2 | 0.23 | 0.79 | 1 | 0.4 | 0.12 | 0.35 | 0.7 | 0.26 |
| KEGG VALINE LEUCINE AND ISOLEUCINE DEGRADATION | 43 | genes.ES.table | 0.59 | 1.6 | 0.074 | 1 | 0.75 | 0.7 | 0.28 | 0.5 | 0.68 | 0.38 |
| KEGG LYSINE DEGRADATION | 43 | genes.ES.table | 0.34 | 1.3 | 0.16 | 0.67 | 0.98 | 0.33 | 0.26 | 0.24 | 0.57 | 0.2 |
Table S11. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | CYP4A11 | CYP4A11 | CYP4A11 | 2 | 0.83 | 0.082 | YES |
| 2 | CYP4A22 | CYP4A22 | CYP4A22 | 40 | 0.59 | 0.14 | YES |
| 3 | SLC27A2 | SLC27A2 | SLC27A2 | 48 | 0.57 | 0.2 | YES |
| 4 | PCK1 | PCK1 | PCK1 | 60 | 0.55 | 0.25 | YES |
| 5 | AQP7 | AQP7 | AQP7 | 104 | 0.48 | 0.29 | YES |
| 6 | FABP4 | FABP4 | FABP4 | 140 | 0.45 | 0.34 | YES |
| 7 | ACADL | ACADL | ACADL | 192 | 0.42 | 0.38 | YES |
| 8 | HMGCS2 | HMGCS2 | HMGCS2 | 208 | 0.41 | 0.42 | YES |
| 9 | CD36 | CD36 | CD36 | 416 | 0.32 | 0.44 | YES |
| 10 | EHHADH | EHHADH | EHHADH | 589 | 0.28 | 0.45 | YES |
| 11 | RXRG | RXRG | RXRG | 850 | 0.22 | 0.46 | YES |
| 12 | CYP27A1 | CYP27A1 | CYP27A1 | 930 | 0.21 | 0.48 | YES |
| 13 | PLIN1 | PLIN1 | PLIN1 | 1158 | 0.18 | 0.48 | YES |
| 14 | LPL | LPL | LPL | 1170 | 0.18 | 0.5 | YES |
| 15 | GK | GK | GK | 1189 | 0.18 | 0.52 | YES |
| 16 | FABP1 | FABP1 | FABP1 | 1290 | 0.17 | 0.53 | YES |
| 17 | ANGPTL4 | ANGPTL4 | ANGPTL4 | 1521 | 0.15 | 0.53 | YES |
| 18 | ACADM | ACADM | ACADM | 1565 | 0.15 | 0.55 | YES |
| 19 | CPT2 | CPT2 | CPT2 | 1600 | 0.15 | 0.56 | YES |
| 20 | PCK2 | PCK2 | PCK2 | 1805 | 0.14 | 0.56 | YES |
| 21 | ACSL1 | ACSL1 | ACSL1 | 1830 | 0.13 | 0.57 | YES |
| 22 | FABP7 | FABP7 | FABP7 | 1967 | 0.13 | 0.58 | YES |
| 23 | PPARG | PPARG | PPARG | 2095 | 0.12 | 0.58 | YES |
| 24 | CPT1A | CPT1A | CPT1A | 2171 | 0.12 | 0.59 | YES |
| 25 | PPARA | PPARA | PPARA | 2804 | 0.092 | 0.56 | NO |
| 26 | FABP3 | FABP3 | FABP3 | 3888 | 0.062 | 0.51 | NO |
| 27 | ACOX1 | ACOX1 | ACOX1 | 4519 | 0.05 | 0.48 | NO |
| 28 | FABP2 | FABP2 | FABP2 | 4549 | 0.049 | 0.48 | NO |
| 29 | PDPK1 | PDPK1 | PDPK1 | 4977 | 0.042 | 0.46 | NO |
| 30 | SCP2 | SCP2 | SCP2 | 5236 | 0.038 | 0.45 | NO |
| 31 | SCD5 | SCD5 | SCD5 | 5326 | 0.037 | 0.45 | NO |
| 32 | CPT1B | CPT1B | CPT1B | 5677 | 0.032 | 0.44 | NO |
| 33 | RXRB | RXRB | RXRB | 5814 | 0.03 | 0.43 | NO |
| 34 | SCD | SCD | SCD | 6237 | 0.024 | 0.41 | NO |
| 35 | ILK | ILK | ILK | 6403 | 0.023 | 0.4 | NO |
| 36 | ACAA1 | ACAA1 | ACAA1 | 6952 | 0.016 | 0.38 | NO |
| 37 | SORBS1 | SORBS1 | SORBS1 | 7859 | 0.0057 | 0.33 | NO |
| 38 | PPARD | PPARD | PPARD | 8237 | 0.0011 | 0.31 | NO |
| 39 | NR1H3 | NR1H3 | NR1H3 | 8541 | -0.0023 | 0.29 | NO |
| 40 | UBC | UBC | UBC | 8864 | -0.006 | 0.27 | NO |
| 41 | FADS2 | FADS2 | FADS2 | 8886 | -0.0064 | 0.27 | NO |
| 42 | RXRA | RXRA | RXRA | 9230 | -0.01 | 0.25 | NO |
| 43 | ACSL3 | ACSL3 | ACSL3 | 9963 | -0.019 | 0.22 | NO |
| 44 | ACSL6 | ACSL6 | ACSL6 | 10111 | -0.021 | 0.21 | NO |
| 45 | SLC27A1 | SLC27A1 | SLC27A1 | 10185 | -0.022 | 0.21 | NO |
| 46 | CYP7A1 | CYP7A1 | CYP7A1 | 10407 | -0.024 | 0.2 | NO |
| 47 | CPT1C | CPT1C | CPT1C | 11021 | -0.032 | 0.17 | NO |
| 48 | OLR1 | OLR1 | OLR1 | 11762 | -0.043 | 0.13 | NO |
| 49 | FABP6 | FABP6 | FABP6 | 12675 | -0.058 | 0.087 | NO |
| 50 | FABP5 | FABP5 | FABP5 | 12948 | -0.063 | 0.078 | NO |
| 51 | ACSL5 | ACSL5 | ACSL5 | 13195 | -0.069 | 0.072 | NO |
| 52 | APOA2 | APOA2 | APOA2 | 13380 | -0.074 | 0.069 | NO |
| 53 | CYP8B1 | CYP8B1 | CYP8B1 | 13455 | -0.076 | 0.072 | NO |
| 54 | SLC27A4 | SLC27A4 | SLC27A4 | 13578 | -0.079 | 0.073 | NO |
| 55 | DBI | DBI | DBI | 13651 | -0.081 | 0.077 | NO |
| 56 | ADIPOQ | ADIPOQ | ADIPOQ | 13792 | -0.084 | 0.078 | NO |
| 57 | ACSL4 | ACSL4 | ACSL4 | 14134 | -0.095 | 0.069 | NO |
| 58 | MMP1 | MMP1 | MMP1 | 14280 | -0.1 | 0.07 | NO |
| 59 | ACOX3 | ACOX3 | ACOX3 | 14632 | -0.11 | 0.062 | NO |
| 60 | SLC27A5 | SLC27A5 | SLC27A5 | 15757 | -0.18 | 0.018 | NO |
| 61 | APOA1 | APOA1 | APOA1 | 15978 | -0.19 | 0.025 | NO |
| 62 | APOC3 | APOC3 | APOC3 | 16695 | -0.25 | 0.011 | NO |
| 63 | ME1 | ME1 | ME1 | 17052 | -0.29 | 0.02 | NO |
| 64 | PLTP | PLTP | PLTP | 17916 | -0.46 | 0.019 | NO |
Figure S21. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA AT1R PATHWAY.
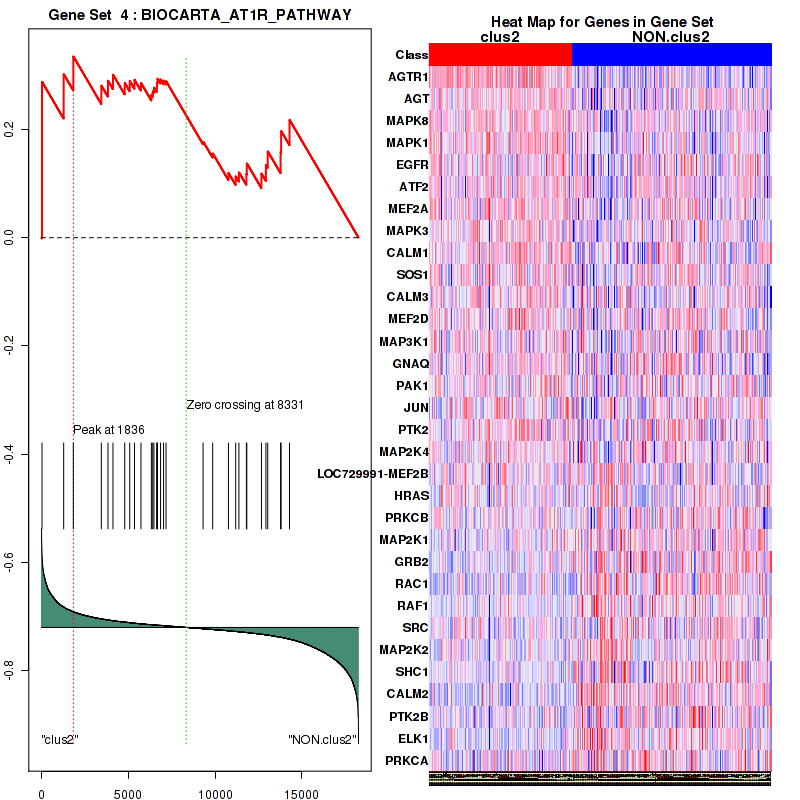
Figure S22. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA AT1R PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
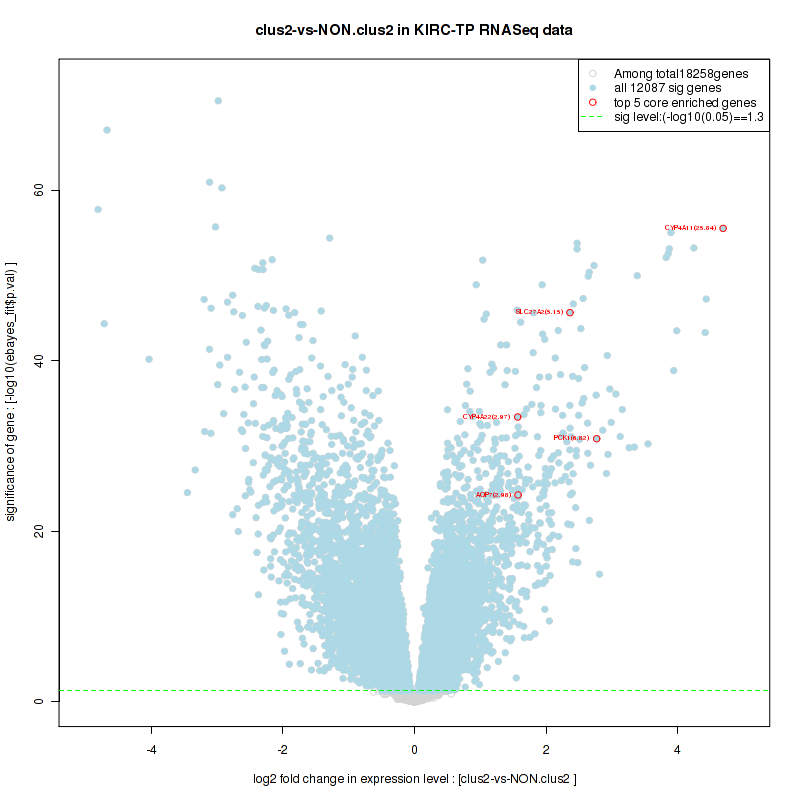
Table S12. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | HMGCS2 | HMGCS2 | HMGCS2 | 208 | 0.41 | 0.08 | YES |
| 2 | ALDH6A1 | ALDH6A1 | ALDH6A1 | 346 | 0.34 | 0.15 | YES |
| 3 | AOX1 | AOX1 | AOX1 | 456 | 0.31 | 0.21 | YES |
| 4 | EHHADH | EHHADH | EHHADH | 589 | 0.28 | 0.26 | YES |
| 5 | ALDH3A2 | ALDH3A2 | ALDH3A2 | 862 | 0.22 | 0.3 | YES |
| 6 | ACAA2 | ACAA2 | ACAA2 | 906 | 0.22 | 0.35 | YES |
| 7 | HIBCH | HIBCH | HIBCH | 982 | 0.2 | 0.39 | YES |
| 8 | ABAT | ABAT | ABAT | 1040 | 0.2 | 0.43 | YES |
| 9 | ACAT1 | ACAT1 | ACAT1 | 1266 | 0.18 | 0.46 | YES |
| 10 | ACADM | ACADM | ACADM | 1565 | 0.15 | 0.47 | YES |
| 11 | PCCA | PCCA | PCCA | 1576 | 0.15 | 0.51 | YES |
| 12 | AUH | AUH | AUH | 1675 | 0.14 | 0.53 | YES |
| 13 | ACADSB | ACADSB | ACADSB | 1806 | 0.14 | 0.56 | YES |
| 14 | MUT | MUT | MUT | 2315 | 0.11 | 0.55 | YES |
| 15 | ALDH9A1 | ALDH9A1 | ALDH9A1 | 2638 | 0.097 | 0.56 | YES |
| 16 | ALDH7A1 | ALDH7A1 | ALDH7A1 | 3122 | 0.082 | 0.55 | YES |
| 17 | IVD | IVD | IVD | 3749 | 0.065 | 0.53 | YES |
| 18 | ALDH2 | ALDH2 | ALDH2 | 3788 | 0.064 | 0.54 | YES |
| 19 | ACADS | ACADS | ACADS | 3860 | 0.063 | 0.55 | YES |
| 20 | MCEE | MCEE | MCEE | 4359 | 0.052 | 0.54 | YES |
| 21 | BCKDHB | BCKDHB | BCKDHB | 4392 | 0.052 | 0.55 | YES |
| 22 | MCCC1 | MCCC1 | MCCC1 | 4544 | 0.049 | 0.55 | YES |
| 23 | DBT | DBT | DBT | 4632 | 0.048 | 0.56 | YES |
| 24 | HMGCL | HMGCL | HMGCL | 4708 | 0.047 | 0.56 | YES |
| 25 | HADHB | HADHB | HADHB | 4720 | 0.047 | 0.57 | YES |
| 26 | HADHA | HADHA | HADHA | 4730 | 0.047 | 0.58 | YES |
| 27 | HIBADH | HIBADH | HIBADH | 4886 | 0.044 | 0.58 | YES |
| 28 | HMGCS1 | HMGCS1 | HMGCS1 | 4899 | 0.044 | 0.59 | YES |
| 29 | ACAD8 | ACAD8 | ACAD8 | 5148 | 0.04 | 0.59 | YES |
| 30 | BCKDHA | BCKDHA | BCKDHA | 5197 | 0.039 | 0.59 | YES |
| 31 | ALDH1B1 | ALDH1B1 | ALDH1B1 | 5616 | 0.032 | 0.58 | NO |
| 32 | OXCT1 | OXCT1 | OXCT1 | 6260 | 0.024 | 0.55 | NO |
| 33 | ECHS1 | ECHS1 | ECHS1 | 6506 | 0.022 | 0.54 | NO |
| 34 | ACAA1 | ACAA1 | ACAA1 | 6952 | 0.016 | 0.52 | NO |
| 35 | MCCC2 | MCCC2 | MCCC2 | 7312 | 0.012 | 0.5 | NO |
| 36 | DLD | DLD | DLD | 8567 | -0.0026 | 0.43 | NO |
| 37 | HADH | HADH | HADH | 8673 | -0.0038 | 0.43 | NO |
| 38 | IL4I1 | IL4I1 | IL4I1 | 9171 | -0.0095 | 0.4 | NO |
| 39 | BCAT2 | BCAT2 | BCAT2 | 10122 | -0.021 | 0.36 | NO |
| 40 | HSD17B10 | HSD17B10 | HSD17B10 | 12118 | -0.049 | 0.26 | NO |
| 41 | ACAT2 | ACAT2 | ACAT2 | 12516 | -0.055 | 0.25 | NO |
| 42 | PCCB | PCCB | PCCB | 13086 | -0.067 | 0.23 | NO |
| 43 | OXCT2 | OXCT2 | OXCT2 | 16486 | -0.24 | 0.097 | NO |
Figure S23. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA DEATH PATHWAY.
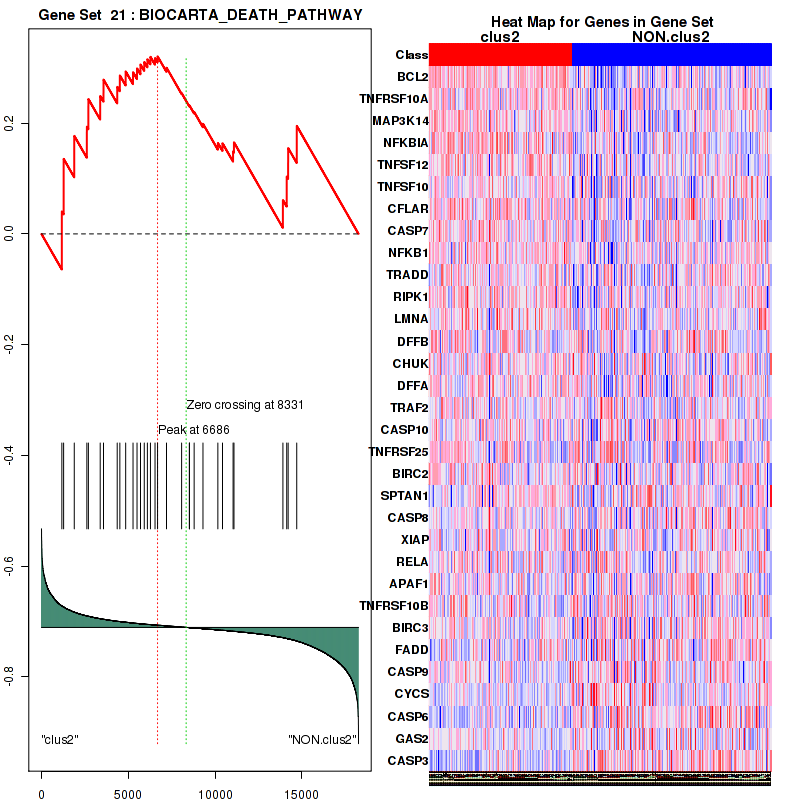
Figure S24. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA DEATH PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
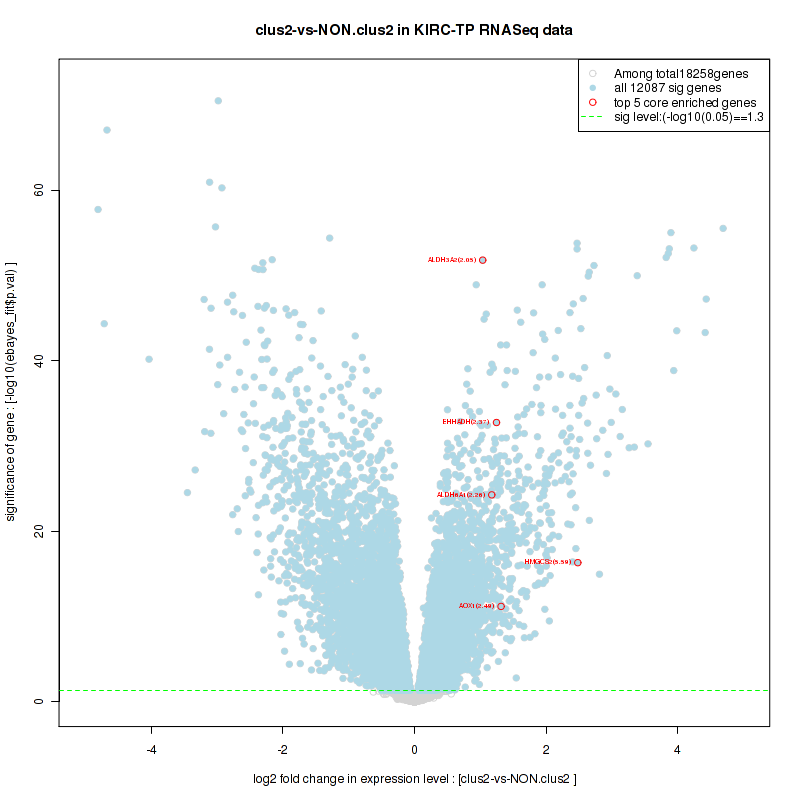
Table S13. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | AMDHD1 | AMDHD1 | AMDHD1 | 28 | 0.62 | 0.13 | YES |
| 2 | DDC | DDC | DDC | 82 | 0.51 | 0.24 | YES |
| 3 | ASPA | ASPA | ASPA | 168 | 0.43 | 0.32 | YES |
| 4 | FTCD | FTCD | FTCD | 170 | 0.43 | 0.42 | YES |
| 5 | ACY3 | ACY3 | ACY3 | 567 | 0.28 | 0.45 | YES |
| 6 | ABP1 | ABP1 | ABP1 | 689 | 0.26 | 0.5 | YES |
| 7 | HDC | HDC | HDC | 726 | 0.25 | 0.55 | YES |
| 8 | ALDH3A2 | ALDH3A2 | ALDH3A2 | 862 | 0.22 | 0.59 | YES |
| 9 | MAOB | MAOB | MAOB | 1080 | 0.19 | 0.62 | YES |
| 10 | HNMT | HNMT | HNMT | 1716 | 0.14 | 0.62 | NO |
| 11 | ALDH9A1 | ALDH9A1 | ALDH9A1 | 2638 | 0.097 | 0.59 | NO |
| 12 | ALDH7A1 | ALDH7A1 | ALDH7A1 | 3122 | 0.082 | 0.58 | NO |
| 13 | LCMT2 | LCMT2 | LCMT2 | 3714 | 0.066 | 0.56 | NO |
| 14 | ALDH2 | ALDH2 | ALDH2 | 3788 | 0.064 | 0.57 | NO |
| 15 | HAL | HAL | HAL | 4303 | 0.054 | 0.55 | NO |
| 16 | ALDH1B1 | ALDH1B1 | ALDH1B1 | 5616 | 0.032 | 0.49 | NO |
| 17 | ALDH3A1 | ALDH3A1 | ALDH3A1 | 5855 | 0.029 | 0.48 | NO |
| 18 | TRMT11 | TRMT11 | TRMT11 | 6152 | 0.025 | 0.47 | NO |
| 19 | HEMK1 | HEMK1 | HEMK1 | 7412 | 0.011 | 0.4 | NO |
| 20 | ALDH3B1 | ALDH3B1 | ALDH3B1 | 7707 | 0.0076 | 0.39 | NO |
| 21 | METTL2B | METTL2B | METTL2B | 8232 | 0.0012 | 0.36 | NO |
| 22 | UROC1 | UROC1 | UROC1 | 10124 | -0.021 | 0.26 | NO |
| 23 | MAOA | MAOA | MAOA | 10573 | -0.026 | 0.24 | NO |
| 24 | LCMT1 | LCMT1 | LCMT1 | 11018 | -0.032 | 0.22 | NO |
| 25 | WBSCR22 | WBSCR22 | WBSCR22 | 13246 | -0.07 | 0.12 | NO |
| 26 | METTL6 | METTL6 | METTL6 | 13876 | -0.087 | 0.1 | NO |
| 27 | ALDH3B2 | ALDH3B2 | ALDH3B2 | 15603 | -0.17 | 0.041 | NO |
| 28 | ALDH1A3 | ALDH1A3 | ALDH1A3 | 17979 | -0.49 | 0.015 | NO |
Figure S25. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA VEGF PATHWAY.
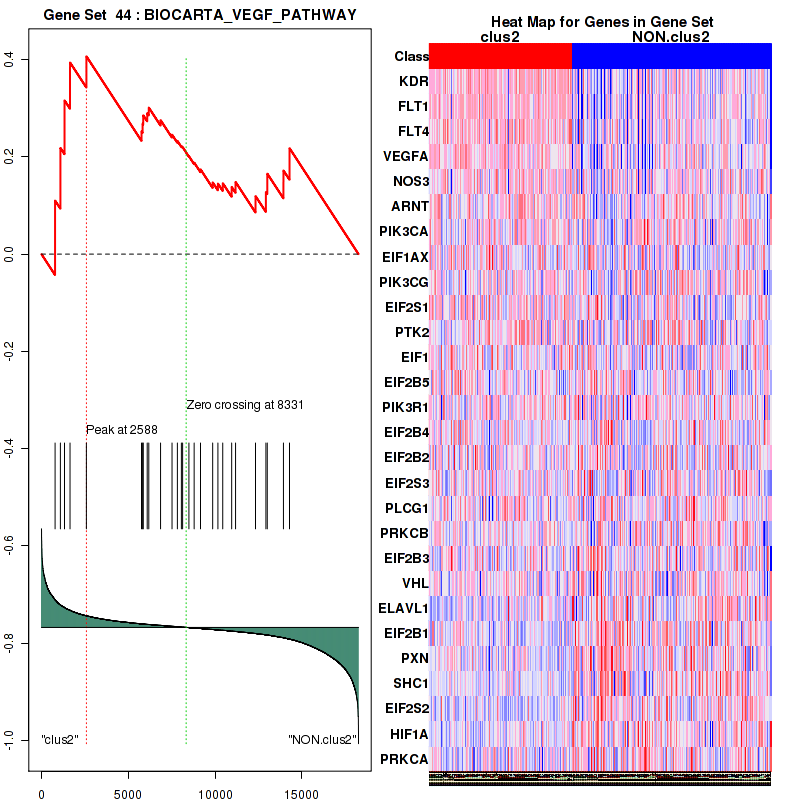
Figure S26. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA VEGF PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
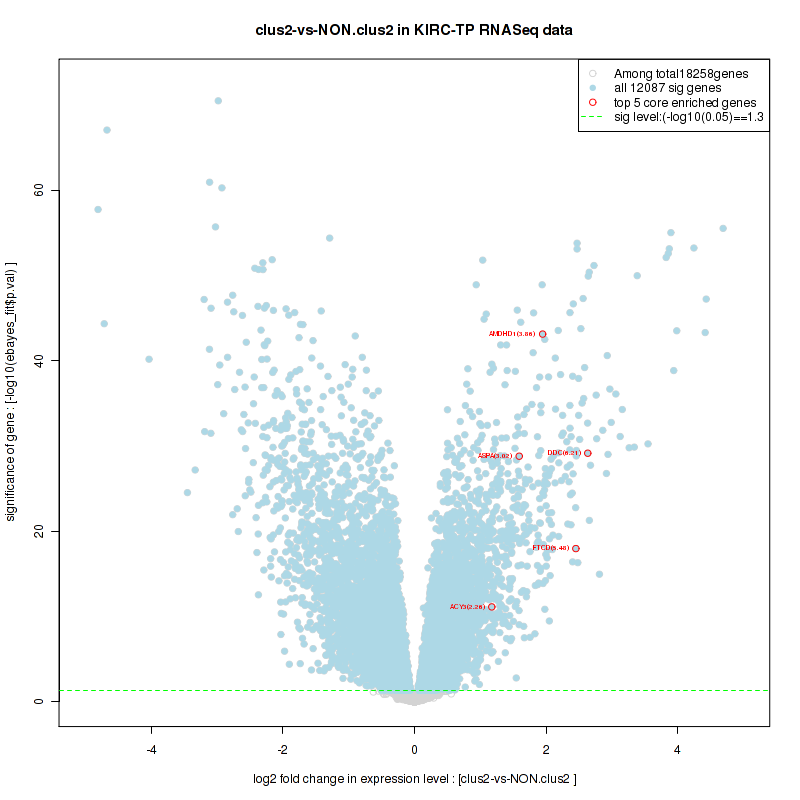
Table S14. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | ALDH6A1 | ALDH6A1 | ALDH6A1 | 346 | 0.34 | 0.085 | YES |
| 2 | EHHADH | EHHADH | EHHADH | 589 | 0.28 | 0.16 | YES |
| 3 | ALDH3A2 | ALDH3A2 | ALDH3A2 | 862 | 0.22 | 0.21 | YES |
| 4 | HIBCH | HIBCH | HIBCH | 982 | 0.2 | 0.27 | YES |
| 5 | ABAT | ABAT | ABAT | 1040 | 0.2 | 0.33 | YES |
| 6 | ACAT1 | ACAT1 | ACAT1 | 1266 | 0.18 | 0.37 | YES |
| 7 | ACADM | ACADM | ACADM | 1565 | 0.15 | 0.4 | YES |
| 8 | PCCA | PCCA | PCCA | 1576 | 0.15 | 0.44 | YES |
| 9 | MLYCD | MLYCD | MLYCD | 2254 | 0.11 | 0.44 | YES |
| 10 | MUT | MUT | MUT | 2315 | 0.11 | 0.47 | YES |
| 11 | ALDH9A1 | ALDH9A1 | ALDH9A1 | 2638 | 0.097 | 0.48 | YES |
| 12 | LDHAL6A | LDHAL6A | LDHAL6A | 2745 | 0.094 | 0.51 | YES |
| 13 | ALDH7A1 | ALDH7A1 | ALDH7A1 | 3122 | 0.082 | 0.51 | YES |
| 14 | ACSS3 | ACSS3 | ACSS3 | 3136 | 0.081 | 0.54 | YES |
| 15 | ACSS1 | ACSS1 | ACSS1 | 3287 | 0.077 | 0.55 | YES |
| 16 | SUCLA2 | SUCLA2 | SUCLA2 | 3371 | 0.074 | 0.57 | YES |
| 17 | ALDH2 | ALDH2 | ALDH2 | 3788 | 0.064 | 0.57 | NO |
| 18 | MCEE | MCEE | MCEE | 4359 | 0.052 | 0.55 | NO |
| 19 | HADHA | HADHA | HADHA | 4730 | 0.047 | 0.55 | NO |
| 20 | ACACB | ACACB | ACACB | 5401 | 0.035 | 0.52 | NO |
| 21 | ALDH1B1 | ALDH1B1 | ALDH1B1 | 5616 | 0.032 | 0.52 | NO |
| 22 | LDHA | LDHA | LDHA | 5933 | 0.028 | 0.51 | NO |
| 23 | ECHS1 | ECHS1 | ECHS1 | 6506 | 0.022 | 0.49 | NO |
| 24 | SUCLG1 | SUCLG1 | SUCLG1 | 8368 | -0.00036 | 0.38 | NO |
| 25 | ACSS2 | ACSS2 | ACSS2 | 9894 | -0.018 | 0.31 | NO |
| 26 | SUCLG2 | SUCLG2 | SUCLG2 | 11049 | -0.033 | 0.25 | NO |
| 27 | LDHB | LDHB | LDHB | 12499 | -0.055 | 0.19 | NO |
| 28 | ACAT2 | ACAT2 | ACAT2 | 12516 | -0.055 | 0.21 | NO |
| 29 | PCCB | PCCB | PCCB | 13086 | -0.067 | 0.2 | NO |
| 30 | ACACA | ACACA | ACACA | 14505 | -0.11 | 0.15 | NO |
| 31 | LDHAL6B | LDHAL6B | LDHAL6B | 15780 | -0.18 | 0.14 | NO |
Figure S27. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG GLYCOLYSIS GLUCONEOGENESIS.
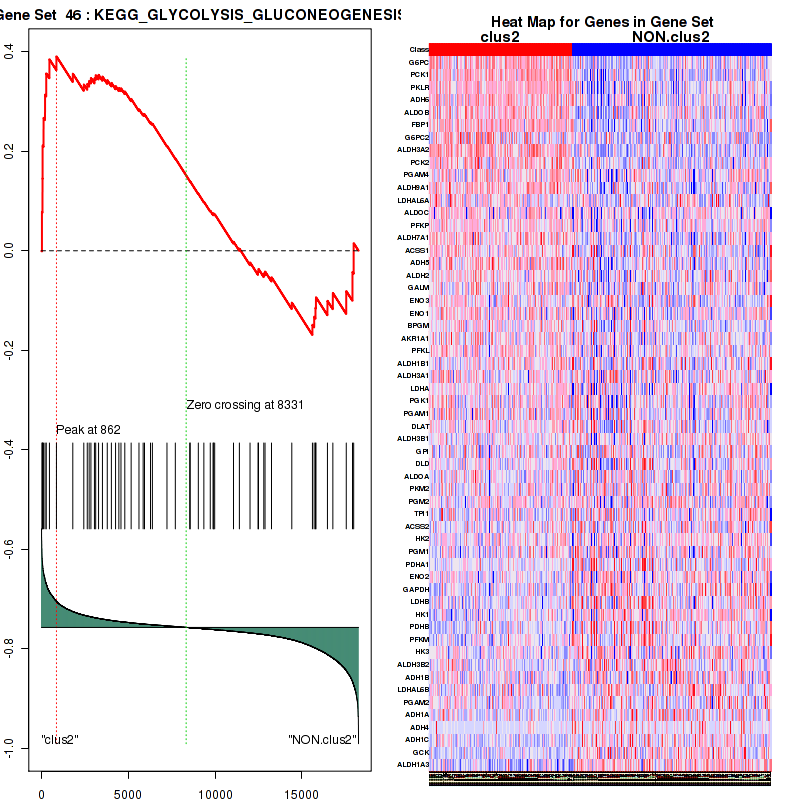
Figure S28. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG GLYCOLYSIS GLUCONEOGENESIS, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
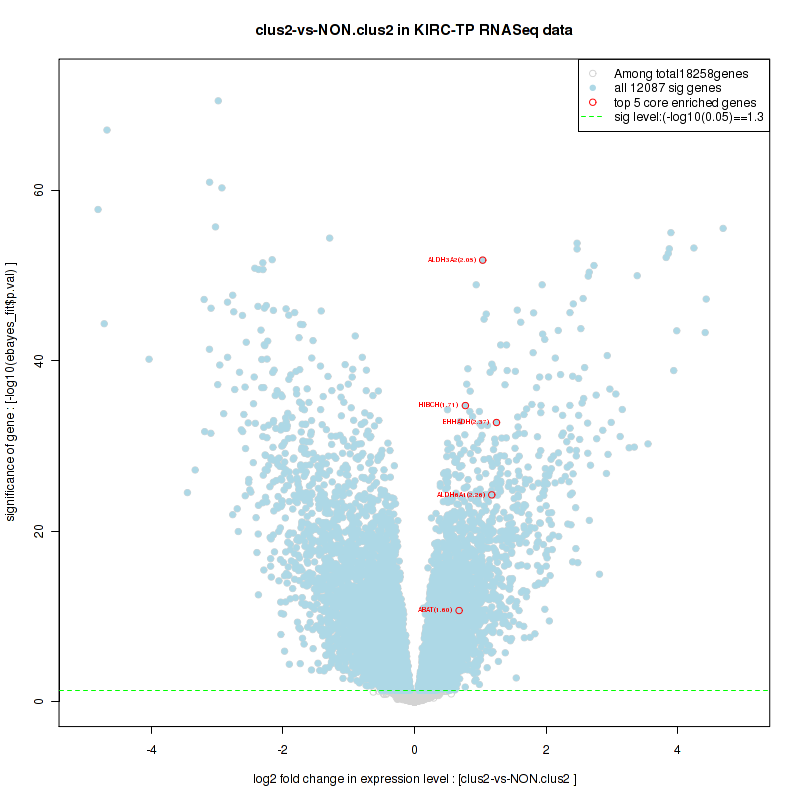
Table S15. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | HAO2 | HAO2 | HAO2 | 15 | 0.74 | 0.095 | YES |
| 2 | SLC27A2 | SLC27A2 | SLC27A2 | 48 | 0.57 | 0.17 | YES |
| 3 | DAO | DAO | DAO | 297 | 0.36 | 0.2 | YES |
| 4 | EHHADH | EHHADH | EHHADH | 589 | 0.28 | 0.22 | YES |
| 5 | PXMP2 | PXMP2 | PXMP2 | 723 | 0.25 | 0.25 | YES |
| 6 | AGXT | AGXT | AGXT | 738 | 0.24 | 0.28 | YES |
| 7 | EPHX2 | EPHX2 | EPHX2 | 878 | 0.22 | 0.3 | YES |
| 8 | NOS2 | NOS2 | NOS2 | 1031 | 0.2 | 0.32 | YES |
| 9 | DDO | DDO | DDO | 1131 | 0.19 | 0.34 | YES |
| 10 | FAR2 | FAR2 | FAR2 | 1293 | 0.17 | 0.35 | YES |
| 11 | PECR | PECR | PECR | 1299 | 0.17 | 0.37 | YES |
| 12 | PEX11A | PEX11A | PEX11A | 1327 | 0.17 | 0.39 | YES |
| 13 | ACSL1 | ACSL1 | ACSL1 | 1830 | 0.13 | 0.38 | YES |
| 14 | DECR2 | DECR2 | DECR2 | 2015 | 0.12 | 0.39 | YES |
| 15 | NUDT12 | NUDT12 | NUDT12 | 2131 | 0.12 | 0.4 | YES |
| 16 | MLYCD | MLYCD | MLYCD | 2254 | 0.11 | 0.4 | YES |
| 17 | PEX11G | PEX11G | PEX11G | 2329 | 0.11 | 0.42 | YES |
| 18 | PAOX | PAOX | PAOX | 2420 | 0.1 | 0.42 | YES |
| 19 | PHYH | PHYH | PHYH | 2555 | 0.1 | 0.43 | YES |
| 20 | DHRS4 | DHRS4 | DHRS4 | 2654 | 0.097 | 0.44 | YES |
| 21 | CRAT | CRAT | CRAT | 2665 | 0.096 | 0.45 | YES |
| 22 | AGPS | AGPS | AGPS | 2714 | 0.095 | 0.46 | YES |
| 23 | CAT | CAT | CAT | 2786 | 0.092 | 0.47 | YES |
| 24 | ECH1 | ECH1 | ECH1 | 2840 | 0.091 | 0.48 | YES |
| 25 | PEX1 | PEX1 | PEX1 | 2970 | 0.086 | 0.48 | YES |
| 26 | ABCD3 | ABCD3 | ABCD3 | 3084 | 0.083 | 0.48 | YES |
| 27 | PECI | PECI | PECI | 3432 | 0.073 | 0.47 | NO |
| 28 | MPV17L | MPV17L | MPV17L | 3601 | 0.069 | 0.47 | NO |
| 29 | ABCD4 | ABCD4 | ABCD4 | 3751 | 0.065 | 0.47 | NO |
| 30 | AMACR | AMACR | AMACR | 3844 | 0.063 | 0.48 | NO |
| 31 | PEX3 | PEX3 | PEX3 | 3949 | 0.061 | 0.48 | NO |
| 32 | PEX7 | PEX7 | PEX7 | 4410 | 0.052 | 0.46 | NO |
| 33 | PEX10 | PEX10 | PEX10 | 4467 | 0.051 | 0.46 | NO |
| 34 | ACOX1 | ACOX1 | ACOX1 | 4519 | 0.05 | 0.47 | NO |
| 35 | PMVK | PMVK | PMVK | 4567 | 0.049 | 0.47 | NO |
| 36 | HMGCL | HMGCL | HMGCL | 4708 | 0.047 | 0.47 | NO |
| 37 | ABCD2 | ABCD2 | ABCD2 | 5027 | 0.042 | 0.46 | NO |
| 38 | PEX5 | PEX5 | PEX5 | 5052 | 0.041 | 0.46 | NO |
| 39 | SCP2 | SCP2 | SCP2 | 5236 | 0.038 | 0.46 | NO |
| 40 | PEX11B | PEX11B | PEX11B | 5460 | 0.035 | 0.45 | NO |
| 41 | PEX19 | PEX19 | PEX19 | 6569 | 0.021 | 0.39 | NO |
| 42 | PEX26 | PEX26 | PEX26 | 6590 | 0.021 | 0.39 | NO |
| 43 | PEX12 | PEX12 | PEX12 | 6891 | 0.017 | 0.38 | NO |
| 44 | ACAA1 | ACAA1 | ACAA1 | 6952 | 0.016 | 0.38 | NO |
| 45 | PIPOX | PIPOX | PIPOX | 7085 | 0.015 | 0.37 | NO |
| 46 | PEX13 | PEX13 | PEX13 | 7090 | 0.015 | 0.37 | NO |
| 47 | SLC25A17 | SLC25A17 | SLC25A17 | 7477 | 0.01 | 0.35 | NO |
| 48 | SOD1 | SOD1 | SOD1 | 8555 | -0.0025 | 0.3 | NO |
| 49 | CROT | CROT | CROT | 8832 | -0.0056 | 0.28 | NO |
| 50 | GNPAT | GNPAT | GNPAT | 9056 | -0.0082 | 0.27 | NO |
| 51 | HSD17B4 | HSD17B4 | HSD17B4 | 9175 | -0.0096 | 0.26 | NO |
| 52 | PEX14 | PEX14 | PEX14 | 9435 | -0.013 | 0.25 | NO |
| 53 | PEX6 | PEX6 | PEX6 | 9912 | -0.018 | 0.23 | NO |
| 54 | ACSL3 | ACSL3 | ACSL3 | 9963 | -0.019 | 0.23 | NO |
| 55 | ACSL6 | ACSL6 | ACSL6 | 10111 | -0.021 | 0.22 | NO |
| 56 | GSTK1 | GSTK1 | GSTK1 | 10609 | -0.027 | 0.2 | NO |
| 57 | IDH2 | IDH2 | IDH2 | 10659 | -0.027 | 0.2 | NO |
| 58 | ABCD1 | ABCD1 | ABCD1 | 10747 | -0.029 | 0.2 | NO |
| 59 | PEX2 | PEX2 | PEX2 | 11255 | -0.036 | 0.18 | NO |
| 60 | FAR1 | FAR1 | FAR1 | 11319 | -0.036 | 0.18 | NO |
| 61 | MPV17 | MPV17 | MPV17 | 11841 | -0.044 | 0.15 | NO |
| 62 | PRDX5 | PRDX5 | PRDX5 | 12146 | -0.049 | 0.14 | NO |
| 63 | PRDX1 | PRDX1 | PRDX1 | 12312 | -0.052 | 0.14 | NO |
| 64 | PEX16 | PEX16 | PEX16 | 12334 | -0.052 | 0.15 | NO |
| 65 | HACL1 | HACL1 | HACL1 | 12355 | -0.053 | 0.15 | NO |
| 66 | ACSL5 | ACSL5 | ACSL5 | 13195 | -0.069 | 0.12 | NO |
| 67 | NUDT19 | NUDT19 | NUDT19 | 13396 | -0.074 | 0.11 | NO |
| 68 | SOD2 | SOD2 | SOD2 | 13549 | -0.078 | 0.12 | NO |
| 69 | MVK | MVK | MVK | 13559 | -0.078 | 0.12 | NO |
| 70 | ACOT8 | ACOT8 | ACOT8 | 13789 | -0.084 | 0.12 | NO |
| 71 | PXMP4 | PXMP4 | PXMP4 | 13805 | -0.085 | 0.13 | NO |
| 72 | ACSL4 | ACSL4 | ACSL4 | 14134 | -0.095 | 0.13 | NO |
| 73 | ACOX3 | ACOX3 | ACOX3 | 14632 | -0.11 | 0.12 | NO |
| 74 | IDH1 | IDH1 | IDH1 | 14782 | -0.12 | 0.12 | NO |
| 75 | BAAT | BAAT | BAAT | 14874 | -0.12 | 0.13 | NO |
| 76 | XDH | XDH | XDH | 17674 | -0.4 | 0.032 | NO |
Figure S29. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG PENTOSE AND GLUCURONATE INTERCONVERSIONS.
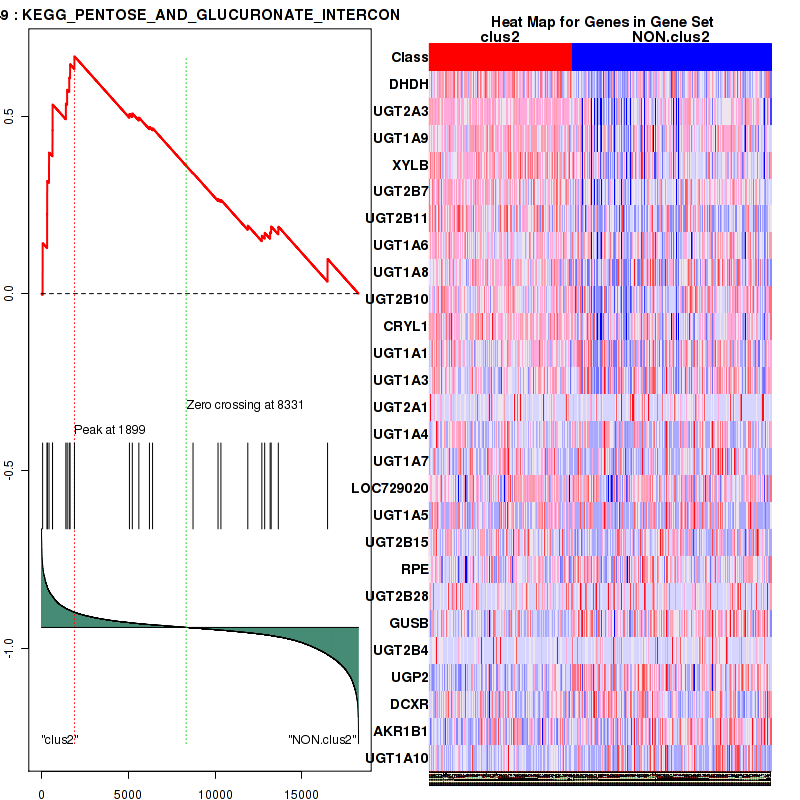
Figure S30. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG PENTOSE AND GLUCURONATE INTERCONVERSIONS, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
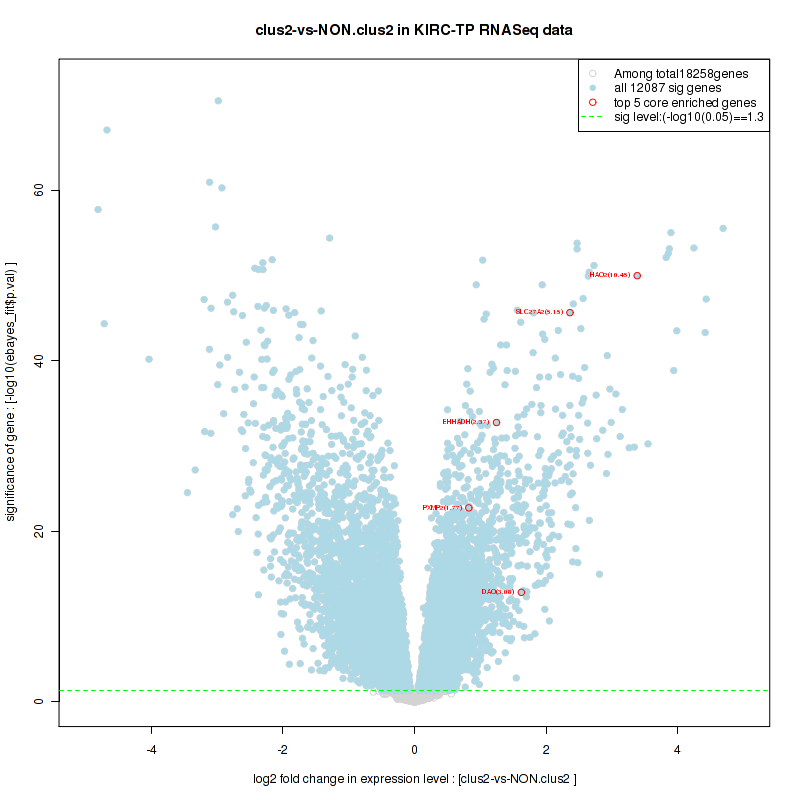
Table S16. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | G6PC | G6PC | G6PC | 27 | 0.64 | 0.1 | YES |
| 2 | PCK1 | PCK1 | PCK1 | 60 | 0.55 | 0.19 | YES |
| 3 | CD36 | CD36 | CD36 | 416 | 0.32 | 0.22 | YES |
| 4 | G6PC2 | G6PC2 | G6PC2 | 458 | 0.31 | 0.26 | YES |
| 5 | RXRG | RXRG | RXRG | 850 | 0.22 | 0.28 | YES |
| 6 | PRKAA2 | PRKAA2 | PRKAA2 | 1485 | 0.16 | 0.27 | YES |
| 7 | SLC2A4 | SLC2A4 | SLC2A4 | 1712 | 0.14 | 0.28 | YES |
| 8 | PCK2 | PCK2 | PCK2 | 1805 | 0.14 | 0.3 | YES |
| 9 | ACSL1 | ACSL1 | ACSL1 | 1830 | 0.13 | 0.32 | YES |
| 10 | MAPK8 | MAPK8 | MAPK8 | 1836 | 0.13 | 0.34 | YES |
| 11 | PPARGC1A | PPARGC1A | PPARGC1A | 1847 | 0.13 | 0.36 | YES |
| 12 | AKT3 | AKT3 | AKT3 | 2165 | 0.12 | 0.36 | YES |
| 13 | CPT1A | CPT1A | CPT1A | 2171 | 0.12 | 0.38 | YES |
| 14 | MAPK10 | MAPK10 | MAPK10 | 2413 | 0.1 | 0.38 | YES |
| 15 | NFKBIA | NFKBIA | NFKBIA | 2614 | 0.098 | 0.39 | YES |
| 16 | PPARA | PPARA | PPARA | 2804 | 0.092 | 0.39 | YES |
| 17 | PRKAB1 | PRKAB1 | PRKAB1 | 3576 | 0.07 | 0.36 | NO |
| 18 | LEPR | LEPR | LEPR | 4069 | 0.058 | 0.34 | NO |
| 19 | CAMKK1 | CAMKK1 | CAMKK1 | 4122 | 0.057 | 0.35 | NO |
| 20 | NFKB1 | NFKB1 | NFKB1 | 4513 | 0.05 | 0.34 | NO |
| 21 | NFKBIE | NFKBIE | NFKBIE | 4780 | 0.046 | 0.33 | NO |
| 22 | TRADD | TRADD | TRADD | 4848 | 0.045 | 0.33 | NO |
| 23 | PRKAA1 | PRKAA1 | PRKAA1 | 5315 | 0.037 | 0.31 | NO |
| 24 | JAK2 | JAK2 | JAK2 | 5316 | 0.037 | 0.32 | NO |
| 25 | ACACB | ACACB | ACACB | 5401 | 0.035 | 0.32 | NO |
| 26 | CPT1B | CPT1B | CPT1B | 5677 | 0.032 | 0.31 | NO |
| 27 | RXRB | RXRB | RXRB | 5814 | 0.03 | 0.31 | NO |
| 28 | CHUK | CHUK | CHUK | 5920 | 0.028 | 0.3 | NO |
| 29 | TRAF2 | TRAF2 | TRAF2 | 6275 | 0.024 | 0.29 | NO |
| 30 | PRKAB2 | PRKAB2 | PRKAB2 | 7185 | 0.014 | 0.24 | NO |
| 31 | IKBKB | IKBKB | IKBKB | 7991 | 0.0042 | 0.2 | NO |
| 32 | NFKBIB | NFKBIB | NFKBIB | 8133 | 0.0023 | 0.19 | NO |
| 33 | MAPK9 | MAPK9 | MAPK9 | 8630 | -0.0034 | 0.16 | NO |
| 34 | ADIPOR1 | ADIPOR1 | ADIPOR1 | 8663 | -0.0037 | 0.16 | NO |
| 35 | RELA | RELA | RELA | 8793 | -0.0052 | 0.16 | NO |
| 36 | STAT3 | STAT3 | STAT3 | 8849 | -0.0058 | 0.16 | NO |
| 37 | RXRA | RXRA | RXRA | 9230 | -0.01 | 0.14 | NO |
| 38 | ADIPOR2 | ADIPOR2 | ADIPOR2 | 9590 | -0.015 | 0.12 | NO |
| 39 | AKT2 | AKT2 | AKT2 | 9906 | -0.018 | 0.1 | NO |
| 40 | ACSL3 | ACSL3 | ACSL3 | 9963 | -0.019 | 0.1 | NO |
| 41 | STK11 | STK11 | STK11 | 10042 | -0.02 | 0.1 | NO |
| 42 | ACSL6 | ACSL6 | ACSL6 | 10111 | -0.021 | 0.1 | NO |
| 43 | PRKAG1 | PRKAG1 | PRKAG1 | 10133 | -0.021 | 0.1 | NO |
| 44 | TNFRSF1B | TNFRSF1B | TNFRSF1B | 10148 | -0.021 | 0.11 | NO |
| 45 | MTOR | MTOR | MTOR | 10232 | -0.022 | 0.11 | NO |
| 46 | CAMKK2 | CAMKK2 | CAMKK2 | 10283 | -0.023 | 0.11 | NO |
| 47 | AKT1 | AKT1 | AKT1 | 10352 | -0.024 | 0.11 | NO |
| 48 | CPT1C | CPT1C | CPT1C | 11021 | -0.032 | 0.076 | NO |
| 49 | PTPN11 | PTPN11 | PTPN11 | 11033 | -0.032 | 0.08 | NO |
| 50 | TNFRSF1A | TNFRSF1A | TNFRSF1A | 11406 | -0.038 | 0.066 | NO |
| 51 | PRKAG2 | PRKAG2 | PRKAG2 | 11691 | -0.042 | 0.057 | NO |
| 52 | PRKCQ | PRKCQ | PRKCQ | 12116 | -0.049 | 0.042 | NO |
| 53 | SLC2A1 | SLC2A1 | SLC2A1 | 12418 | -0.054 | 0.034 | NO |
| 54 | TNF | TNF | TNF | 13078 | -0.066 | 0.0081 | NO |
| 55 | IKBKG | IKBKG | IKBKG | 13157 | -0.068 | 0.015 | NO |
| 56 | ACSL5 | ACSL5 | ACSL5 | 13195 | -0.069 | 0.024 | NO |
| 57 | ADIPOQ | ADIPOQ | ADIPOQ | 13792 | -0.084 | 0.0047 | NO |
| 58 | ACSL4 | ACSL4 | ACSL4 | 14134 | -0.095 | 0.0012 | NO |
| 59 | SOCS3 | SOCS3 | SOCS3 | 14708 | -0.12 | -0.012 | NO |
| 60 | IRS1 | IRS1 | IRS1 | 15689 | -0.17 | -0.038 | NO |
| 61 | POMC | POMC | POMC | 15694 | -0.17 | -0.01 | NO |
| 62 | AGRP | AGRP | AGRP | 16498 | -0.24 | -0.017 | NO |
| 63 | IRS2 | IRS2 | IRS2 | 17433 | -0.35 | -0.012 | NO |
| 64 | LEP | LEP | LEP | 17496 | -0.36 | 0.042 | NO |
Figure S31. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG GALACTOSE METABOLISM.
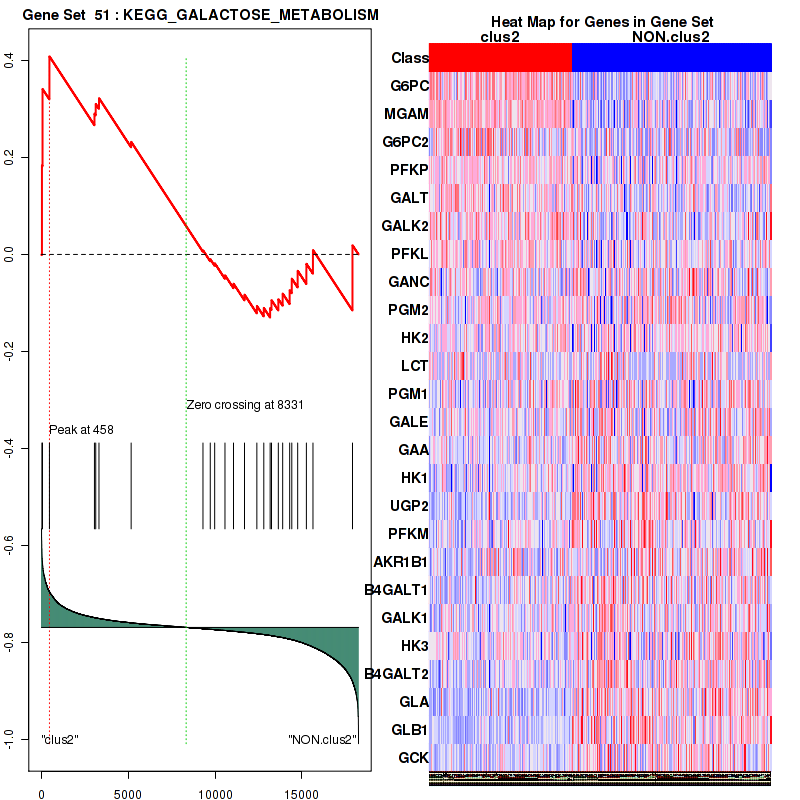
Figure S32. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG GALACTOSE METABOLISM, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
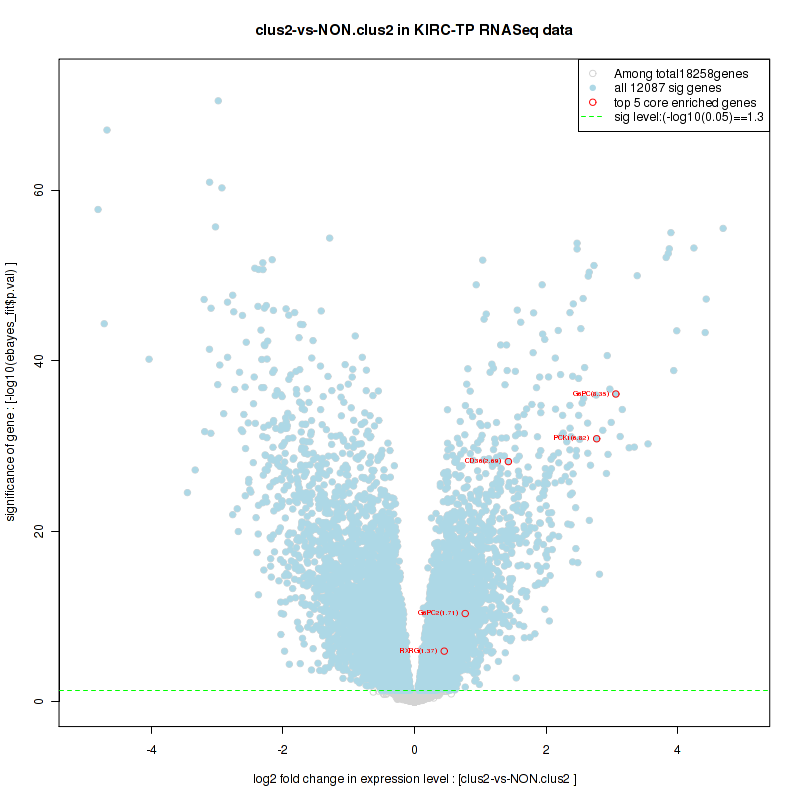
Table S17. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | DHDH | DHDH | DHDH | 67 | 0.53 | 0.14 | YES |
| 2 | UGT2A3 | UGT2A3 | UGT2A3 | 327 | 0.35 | 0.22 | YES |
| 3 | UGT1A9 | UGT1A9 | UGT1A9 | 331 | 0.34 | 0.32 | YES |
| 4 | XYLB | XYLB | XYLB | 436 | 0.31 | 0.4 | YES |
| 5 | UGT2B7 | UGT2B7 | UGT2B7 | 636 | 0.27 | 0.46 | YES |
| 6 | UGT2B11 | UGT2B11 | UGT2B11 | 640 | 0.27 | 0.53 | YES |
| 7 | UGT1A6 | UGT1A6 | UGT1A6 | 1406 | 0.16 | 0.54 | YES |
| 8 | UGT1A8 | UGT1A8 | UGT1A8 | 1479 | 0.16 | 0.57 | YES |
| 9 | UGT2B10 | UGT2B10 | UGT2B10 | 1599 | 0.15 | 0.61 | YES |
| 10 | CRYL1 | CRYL1 | CRYL1 | 1649 | 0.15 | 0.65 | YES |
| 11 | UGT1A1 | UGT1A1 | UGT1A1 | 1899 | 0.13 | 0.67 | YES |
| 12 | UGT1A3 | UGT1A3 | UGT1A3 | 5066 | 0.041 | 0.51 | NO |
| 13 | UGT2A1 | UGT2A1 | UGT2A1 | 5231 | 0.038 | 0.51 | NO |
| 14 | UGT1A4 | UGT1A4 | UGT1A4 | 5609 | 0.032 | 0.5 | NO |
| 15 | UGT1A7 | UGT1A7 | UGT1A7 | 6219 | 0.024 | 0.47 | NO |
| 16 | LOC729020 | LOC729020 | LOC729020 | 6398 | 0.023 | 0.47 | NO |
| 17 | UGT1A5 | UGT1A5 | UGT1A5 | 8729 | -0.0045 | 0.34 | NO |
| 18 | UGT2B15 | UGT2B15 | UGT2B15 | 10169 | -0.022 | 0.27 | NO |
| 19 | RPE | RPE | RPE | 10333 | -0.024 | 0.26 | NO |
| 20 | UGT2B28 | UGT2B28 | UGT2B28 | 11882 | -0.045 | 0.19 | NO |
| 21 | GUSB | GUSB | GUSB | 12691 | -0.058 | 0.16 | NO |
| 22 | UGT2B4 | UGT2B4 | UGT2B4 | 12854 | -0.062 | 0.17 | NO |
| 23 | UGP2 | UGP2 | UGP2 | 13169 | -0.068 | 0.17 | NO |
| 24 | DCXR | DCXR | DCXR | 13226 | -0.07 | 0.19 | NO |
| 25 | AKR1B1 | AKR1B1 | AKR1B1 | 13641 | -0.08 | 0.19 | NO |
| 26 | UGT1A10 | UGT1A10 | UGT1A10 | 16475 | -0.23 | 0.098 | NO |
Figure S33. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG FATTY ACID METABOLISM.
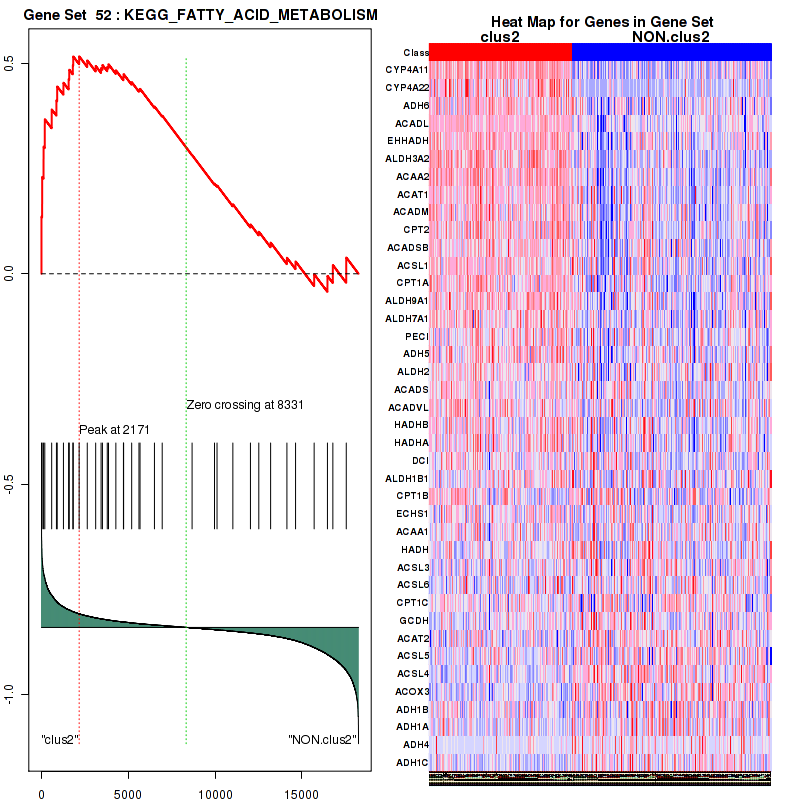
Figure S34. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG FATTY ACID METABOLISM, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
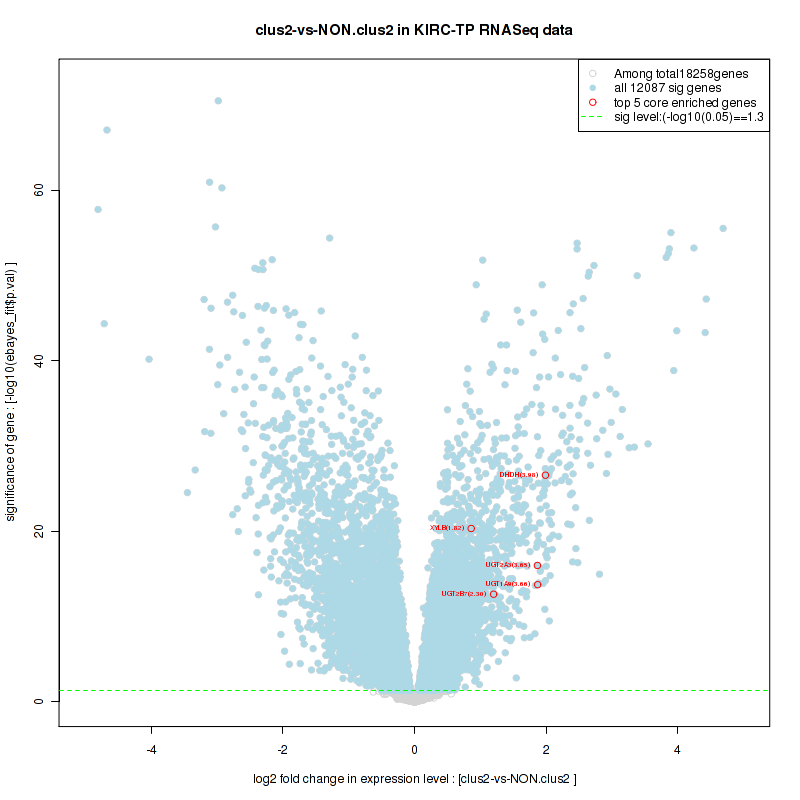
Table S18. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | CYP4A11 | CYP4A11 | CYP4A11 | 2 | 0.83 | 0.14 | YES |
| 2 | CYP4A22 | CYP4A22 | CYP4A22 | 40 | 0.59 | 0.23 | YES |
| 3 | ADH6 | ADH6 | ADH6 | 121 | 0.47 | 0.3 | YES |
| 4 | ACADL | ACADL | ACADL | 192 | 0.42 | 0.37 | YES |
| 5 | EHHADH | EHHADH | EHHADH | 589 | 0.28 | 0.39 | YES |
| 6 | ALDH3A2 | ALDH3A2 | ALDH3A2 | 862 | 0.22 | 0.41 | YES |
| 7 | ACAA2 | ACAA2 | ACAA2 | 906 | 0.22 | 0.44 | YES |
| 8 | ACAT1 | ACAT1 | ACAT1 | 1266 | 0.18 | 0.45 | YES |
| 9 | ACADM | ACADM | ACADM | 1565 | 0.15 | 0.46 | YES |
| 10 | CPT2 | CPT2 | CPT2 | 1600 | 0.15 | 0.48 | YES |
| 11 | ACADSB | ACADSB | ACADSB | 1806 | 0.14 | 0.5 | YES |
| 12 | ACSL1 | ACSL1 | ACSL1 | 1830 | 0.13 | 0.52 | YES |
| 13 | CPT1A | CPT1A | CPT1A | 2171 | 0.12 | 0.52 | YES |
| 14 | ALDH9A1 | ALDH9A1 | ALDH9A1 | 2638 | 0.097 | 0.51 | NO |
| 15 | ALDH7A1 | ALDH7A1 | ALDH7A1 | 3122 | 0.082 | 0.49 | NO |
| 16 | PECI | PECI | PECI | 3432 | 0.073 | 0.49 | NO |
| 17 | ADH5 | ADH5 | ADH5 | 3512 | 0.071 | 0.5 | NO |
| 18 | ALDH2 | ALDH2 | ALDH2 | 3788 | 0.064 | 0.49 | NO |
| 19 | ACADS | ACADS | ACADS | 3860 | 0.063 | 0.5 | NO |
| 20 | ACADVL | ACADVL | ACADVL | 4284 | 0.054 | 0.48 | NO |
| 21 | HADHB | HADHB | HADHB | 4720 | 0.047 | 0.47 | NO |
| 22 | HADHA | HADHA | HADHA | 4730 | 0.047 | 0.48 | NO |
| 23 | DCI | DCI | DCI | 5194 | 0.039 | 0.46 | NO |
| 24 | ALDH1B1 | ALDH1B1 | ALDH1B1 | 5616 | 0.032 | 0.44 | NO |
| 25 | CPT1B | CPT1B | CPT1B | 5677 | 0.032 | 0.44 | NO |
| 26 | ECHS1 | ECHS1 | ECHS1 | 6506 | 0.022 | 0.4 | NO |
| 27 | ACAA1 | ACAA1 | ACAA1 | 6952 | 0.016 | 0.38 | NO |
| 28 | HADH | HADH | HADH | 8673 | -0.0038 | 0.28 | NO |
| 29 | ACSL3 | ACSL3 | ACSL3 | 9963 | -0.019 | 0.22 | NO |
| 30 | ACSL6 | ACSL6 | ACSL6 | 10111 | -0.021 | 0.21 | NO |
| 31 | CPT1C | CPT1C | CPT1C | 11021 | -0.032 | 0.16 | NO |
| 32 | GCDH | GCDH | GCDH | 12035 | -0.047 | 0.12 | NO |
| 33 | ACAT2 | ACAT2 | ACAT2 | 12516 | -0.055 | 0.1 | NO |
| 34 | ACSL5 | ACSL5 | ACSL5 | 13195 | -0.069 | 0.074 | NO |
| 35 | ACSL4 | ACSL4 | ACSL4 | 14134 | -0.095 | 0.038 | NO |
| 36 | ACOX3 | ACOX3 | ACOX3 | 14632 | -0.11 | 0.03 | NO |
| 37 | ADH1B | ADH1B | ADH1B | 15704 | -0.17 | -0.0011 | NO |
| 38 | ADH1A | ADH1A | ADH1A | 16469 | -0.23 | -0.005 | NO |
| 39 | ADH4 | ADH4 | ADH4 | 16779 | -0.26 | 0.021 | NO |
| 40 | ADH1C | ADH1C | ADH1C | 17552 | -0.37 | 0.039 | NO |
Figure S35. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG GLYCINE SERINE AND THREONINE METABOLISM.
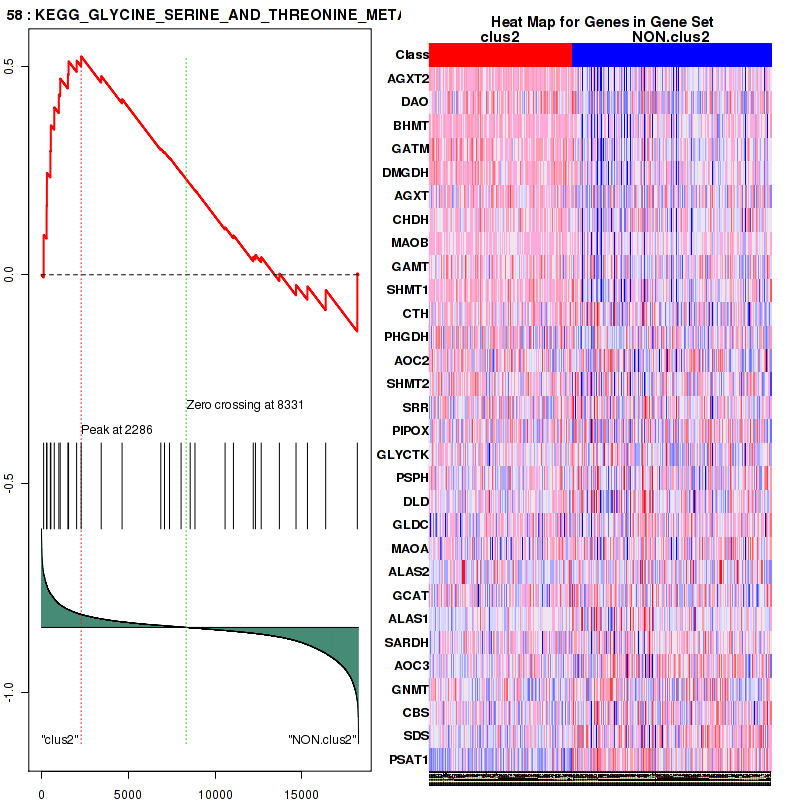
Figure S36. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG GLYCINE SERINE AND THREONINE METABOLISM, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
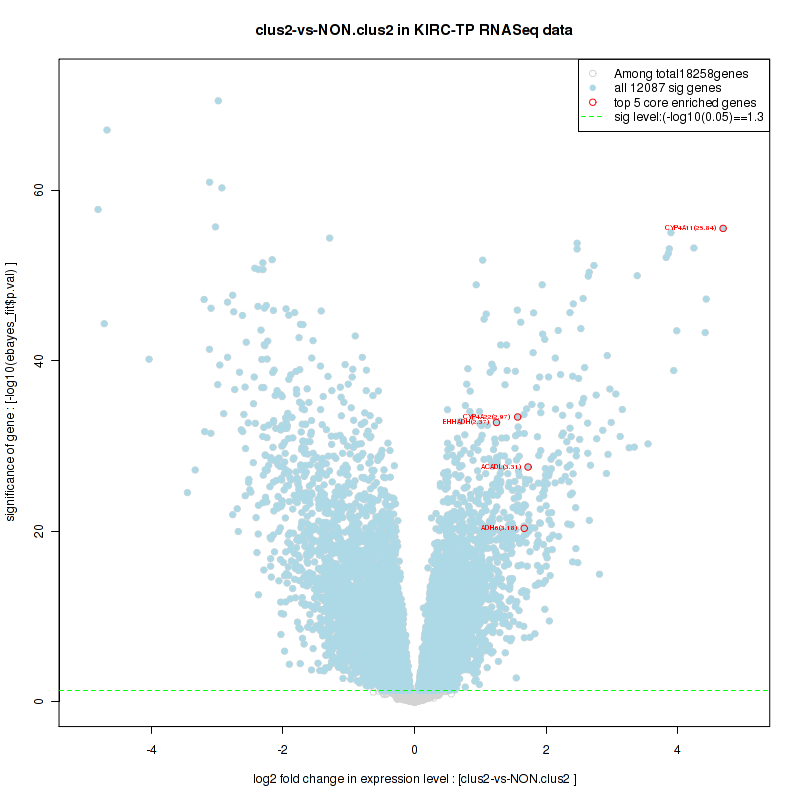
Table S19. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | AGPAT9 | AGPAT9 | AGPAT9 | 469 | 0.3 | 0.031 | YES |
| 2 | DGKB | DGKB | DGKB | 662 | 0.26 | 0.069 | YES |
| 3 | DGKI | DGKI | DGKI | 692 | 0.25 | 0.11 | YES |
| 4 | DGKH | DGKH | DGKH | 755 | 0.24 | 0.16 | YES |
| 5 | ALDH3A2 | ALDH3A2 | ALDH3A2 | 862 | 0.22 | 0.19 | YES |
| 6 | LIPC | LIPC | LIPC | 904 | 0.22 | 0.23 | YES |
| 7 | LPL | LPL | LPL | 1170 | 0.18 | 0.25 | YES |
| 8 | GK | GK | GK | 1189 | 0.18 | 0.28 | YES |
| 9 | PPAP2B | PPAP2B | PPAP2B | 1206 | 0.18 | 0.31 | YES |
| 10 | DGKE | DGKE | DGKE | 1267 | 0.18 | 0.34 | YES |
| 11 | PPAP2A | PPAP2A | PPAP2A | 1313 | 0.17 | 0.37 | YES |
| 12 | PNPLA3 | PNPLA3 | PNPLA3 | 1461 | 0.16 | 0.39 | YES |
| 13 | ALDH9A1 | ALDH9A1 | ALDH9A1 | 2638 | 0.097 | 0.35 | NO |
| 14 | ALDH7A1 | ALDH7A1 | ALDH7A1 | 3122 | 0.082 | 0.34 | NO |
| 15 | DGKD | DGKD | DGKD | 3207 | 0.079 | 0.35 | NO |
| 16 | DAK | DAK | DAK | 3399 | 0.074 | 0.35 | NO |
| 17 | AGPAT3 | AGPAT3 | AGPAT3 | 3579 | 0.07 | 0.35 | NO |
| 18 | ALDH2 | ALDH2 | ALDH2 | 3788 | 0.064 | 0.35 | NO |
| 19 | LCLAT1 | LCLAT1 | LCLAT1 | 4147 | 0.056 | 0.34 | NO |
| 20 | AKR1A1 | AKR1A1 | AKR1A1 | 4798 | 0.046 | 0.32 | NO |
| 21 | ALDH1B1 | ALDH1B1 | ALDH1B1 | 5616 | 0.032 | 0.28 | NO |
| 22 | DGKZ | DGKZ | DGKZ | 6086 | 0.026 | 0.26 | NO |
| 23 | AGPAT1 | AGPAT1 | AGPAT1 | 6452 | 0.022 | 0.24 | NO |
| 24 | GLYCTK | GLYCTK | GLYCTK | 7373 | 0.011 | 0.19 | NO |
| 25 | MGLL | MGLL | MGLL | 7583 | 0.0091 | 0.18 | NO |
| 26 | AGPAT4 | AGPAT4 | AGPAT4 | 9319 | -0.012 | 0.09 | NO |
| 27 | AGPAT6 | AGPAT6 | AGPAT6 | 9404 | -0.012 | 0.088 | NO |
| 28 | MBOAT2 | MBOAT2 | MBOAT2 | 10902 | -0.031 | 0.011 | NO |
| 29 | GPAM | GPAM | GPAM | 11594 | -0.04 | -0.019 | NO |
| 30 | DGKQ | DGKQ | DGKQ | 12113 | -0.048 | -0.038 | NO |
| 31 | LIPG | LIPG | LIPG | 12190 | -0.05 | -0.033 | NO |
| 32 | AGK | AGK | AGK | 12543 | -0.056 | -0.042 | NO |
| 33 | MBOAT1 | MBOAT1 | MBOAT1 | 12586 | -0.057 | -0.034 | NO |
| 34 | DGAT1 | DGAT1 | DGAT1 | 13040 | -0.066 | -0.047 | NO |
| 35 | AGPAT2 | AGPAT2 | AGPAT2 | 13453 | -0.076 | -0.055 | NO |
| 36 | AKR1B1 | AKR1B1 | AKR1B1 | 13641 | -0.08 | -0.051 | NO |
| 37 | LIPF | LIPF | LIPF | 13645 | -0.08 | -0.036 | NO |
| 38 | GPAT2 | GPAT2 | GPAT2 | 13679 | -0.081 | -0.022 | NO |
| 39 | DGKA | DGKA | DGKA | 14218 | -0.098 | -0.034 | NO |
| 40 | AWAT2 | AWAT2 | AWAT2 | 14612 | -0.11 | -0.034 | NO |
| 41 | GLA | GLA | GLA | 15256 | -0.14 | -0.043 | NO |
| 42 | PPAP2C | PPAP2C | PPAP2C | 16140 | -0.2 | -0.054 | NO |
| 43 | CEL | CEL | CEL | 16371 | -0.22 | -0.025 | NO |
| 44 | DGAT2 | DGAT2 | DGAT2 | 16902 | -0.28 | -0.0031 | NO |
| 45 | DGKG | DGKG | DGKG | 17764 | -0.42 | 0.027 | NO |
Figure S37. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG VALINE LEUCINE AND ISOLEUCINE DEGRADATION.
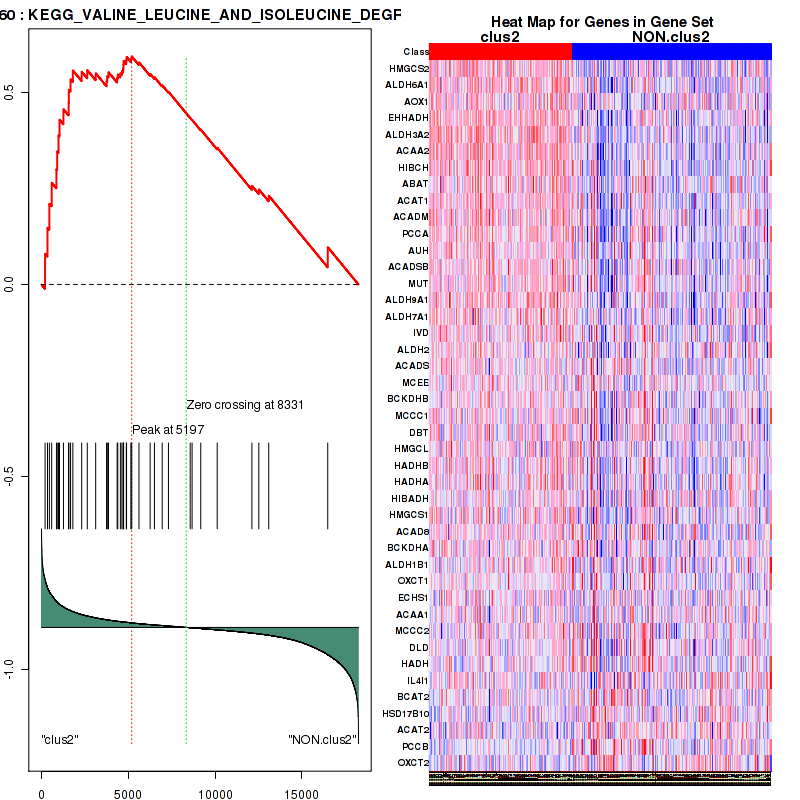
Figure S38. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG VALINE LEUCINE AND ISOLEUCINE DEGRADATION, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
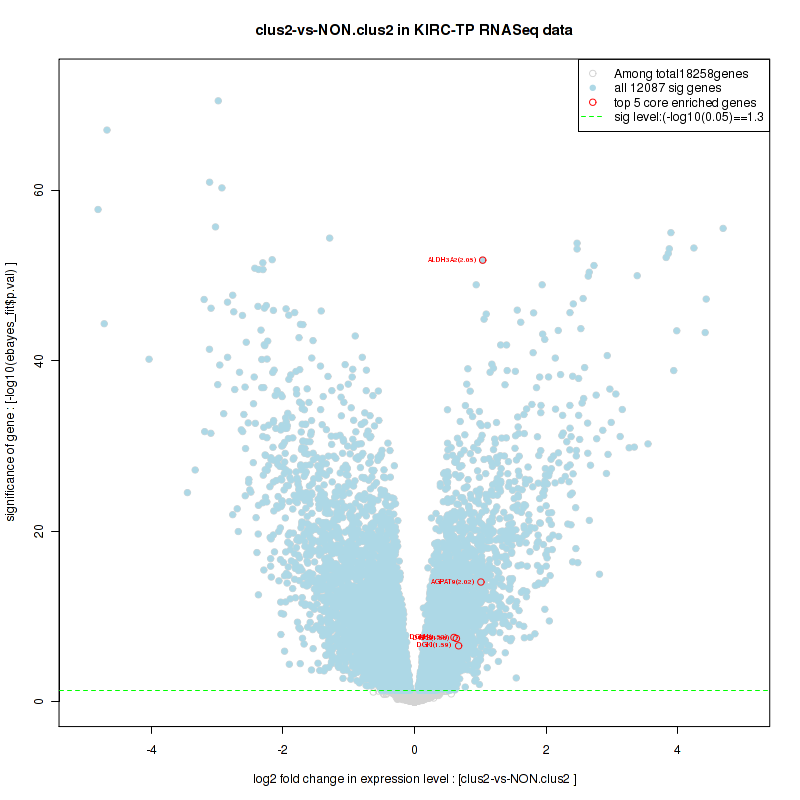
Table S20. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | CREB3L3 | CREB3L3 | CREB3L3 | 258 | 0.38 | 0.1 | YES |
| 2 | AQP4 | AQP4 | AQP4 | 417 | 0.32 | 0.19 | YES |
| 3 | AVPR2 | AVPR2 | AVPR2 | 1028 | 0.2 | 0.21 | YES |
| 4 | DYNC2H1 | DYNC2H1 | DYNC2H1 | 1309 | 0.17 | 0.25 | YES |
| 5 | ADCY9 | ADCY9 | ADCY9 | 1441 | 0.16 | 0.29 | YES |
| 6 | CREB5 | CREB5 | CREB5 | 1941 | 0.13 | 0.3 | YES |
| 7 | DCTN6 | DCTN6 | DCTN6 | 2627 | 0.097 | 0.29 | YES |
| 8 | DYNC2LI1 | DYNC2LI1 | DYNC2LI1 | 2712 | 0.095 | 0.32 | YES |
| 9 | AQP3 | AQP3 | AQP3 | 2964 | 0.086 | 0.33 | YES |
| 10 | CREB3L4 | CREB3L4 | CREB3L4 | 3166 | 0.08 | 0.34 | YES |
| 11 | VAMP2 | VAMP2 | VAMP2 | 3196 | 0.079 | 0.36 | YES |
| 12 | DYNLL2 | DYNLL2 | DYNLL2 | 3722 | 0.066 | 0.35 | NO |
| 13 | DCTN1 | DCTN1 | DCTN1 | 4380 | 0.052 | 0.33 | NO |
| 14 | CREB3 | CREB3 | CREB3 | 5007 | 0.042 | 0.31 | NO |
| 15 | DYNC1I2 | DYNC1I2 | DYNC1I2 | 5256 | 0.038 | 0.31 | NO |
| 16 | CREB1 | CREB1 | CREB1 | 5992 | 0.027 | 0.28 | NO |
| 17 | PRKACB | PRKACB | PRKACB | 7170 | 0.014 | 0.22 | NO |
| 18 | DYNC1LI2 | DYNC1LI2 | DYNC1LI2 | 7906 | 0.0051 | 0.18 | NO |
| 19 | ADCY6 | ADCY6 | ADCY6 | 8034 | 0.0035 | 0.17 | NO |
| 20 | ARHGDIB | ARHGDIB | ARHGDIB | 8045 | 0.0034 | 0.17 | NO |
| 21 | DYNC1H1 | DYNC1H1 | DYNC1H1 | 8266 | 0.00072 | 0.16 | NO |
| 22 | RAB5C | RAB5C | RAB5C | 8527 | -0.0021 | 0.15 | NO |
| 23 | DCTN4 | DCTN4 | DCTN4 | 8569 | -0.0026 | 0.14 | NO |
| 24 | ARHGDIA | ARHGDIA | ARHGDIA | 8808 | -0.0054 | 0.13 | NO |
| 25 | RAB5B | RAB5B | RAB5B | 8983 | -0.0074 | 0.12 | NO |
| 26 | RAB11A | RAB11A | RAB11A | 9140 | -0.0092 | 0.12 | NO |
| 27 | STX4 | STX4 | STX4 | 9267 | -0.011 | 0.12 | NO |
| 28 | CREB3L2 | CREB3L2 | CREB3L2 | 9469 | -0.013 | 0.11 | NO |
| 29 | DCTN5 | DCTN5 | DCTN5 | 10096 | -0.021 | 0.081 | NO |
| 30 | RAB11B | RAB11B | RAB11B | 10453 | -0.025 | 0.068 | NO |
| 31 | PRKACA | PRKACA | PRKACA | 10539 | -0.026 | 0.072 | NO |
| 32 | DYNC1LI1 | DYNC1LI1 | DYNC1LI1 | 10756 | -0.029 | 0.068 | NO |
| 33 | NSF | NSF | NSF | 10888 | -0.03 | 0.07 | NO |
| 34 | DCTN2 | DCTN2 | DCTN2 | 11600 | -0.04 | 0.043 | NO |
| 35 | RAB5A | RAB5A | RAB5A | 11834 | -0.044 | 0.044 | NO |
| 36 | PRKX | PRKX | PRKX | 11958 | -0.046 | 0.05 | NO |
| 37 | DYNLL1 | DYNLL1 | DYNLL1 | 12254 | -0.051 | 0.05 | NO |
| 38 | DYNC1I1 | DYNC1I1 | DYNC1I1 | 13525 | -0.078 | 0.0031 | NO |
| 39 | GNAS | GNAS | GNAS | 14553 | -0.11 | -0.021 | NO |
| 40 | AQP2 | AQP2 | AQP2 | 15604 | -0.17 | -0.028 | NO |
| 41 | CREB3L1 | CREB3L1 | CREB3L1 | 18136 | -0.58 | 0.0067 | NO |
Figure S39. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG LYSINE DEGRADATION.
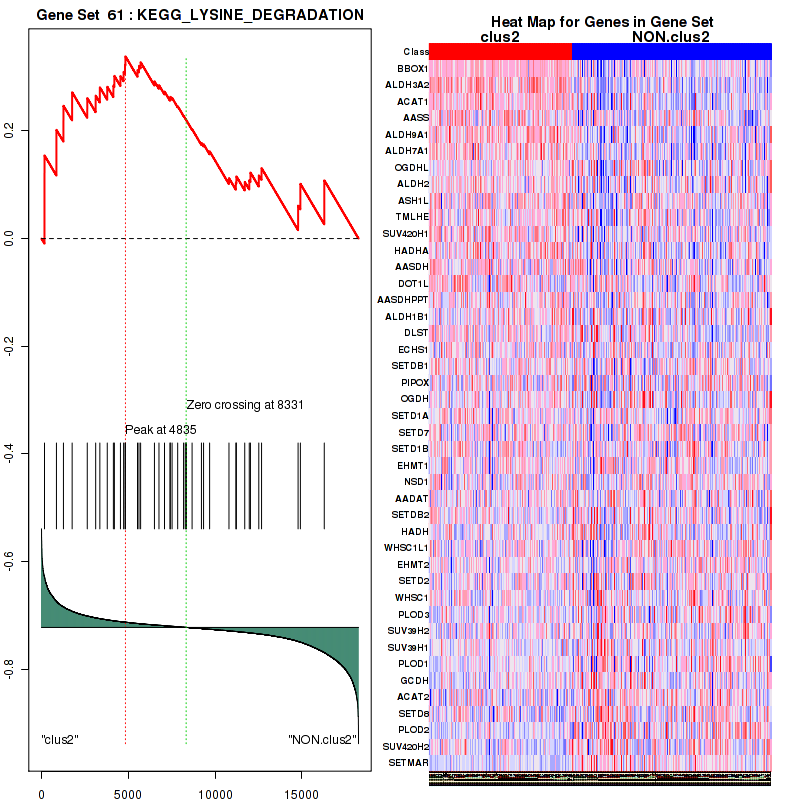
Figure S40. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG LYSINE DEGRADATION, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
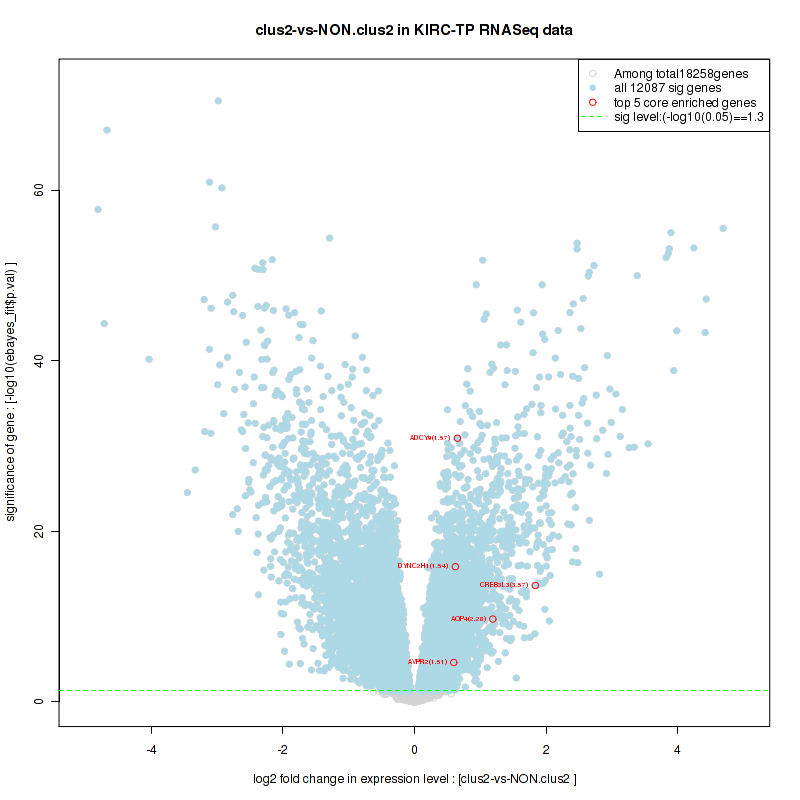
For the top enriched genes, if you want to check whether they are
-
up-regulated, please check the list of up-regulated genes
-
down-regulated, please check the list of down-regulated genes
For the top enriched genes, if you want to check whether they are
-
highly expressed genes, please check the list of high (top 30%) expressed genes
-
low expressed genes, please check the list of low (bottom 30%) expressed genes
An expression pattern of top(30%)/middle(30%)/low(30%) in this subtype against other subtypes is available in a heatmap
For the top enriched genes, if you want to check whether they are
-
significantly differently expressed genes by eBayes lm fit, please check the list of significant genes
Table 5. Get Full Table This table shows top 10 pathways which are significantly enriched in cluster clus3. It displays only significant gene sets satisfying nom.p.val.threshold (-1), fwer.p.val.threshold (-1) , fdr.q.val.threshold (0.25) and the default table is sorted by Normalized Enrichment Score (NES). Further details on NES statistics, please visit The Broad GSEA website.
| GeneSet(GS) | Size(#genes) | genes.ES.table | ES | NES | NOM.p.val | FDR.q.val | FWER.p.val | Tag.. | Gene.. | Signal | FDR..median. | glob.p.val |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BIOCARTA INFLAM PATHWAY | 25 | genes.ES.table | 0.66 | 1.5 | 0.063 | 0.6 | 0.92 | 0.76 | 0.28 | 0.55 | 0.47 | 0.18 |
| BIOCARTA GSK3 PATHWAY | 26 | genes.ES.table | 0.53 | 1.6 | 0.028 | 0.56 | 0.7 | 0.15 | 0.054 | 0.15 | 0.34 | 0.18 |
| BIOCARTA WNT PATHWAY | 25 | genes.ES.table | 0.41 | 1.4 | 0.084 | 0.58 | 0.93 | 0.08 | 0.054 | 0.076 | 0.46 | 0.17 |
| KEGG AMINO SUGAR AND NUCLEOTIDE SUGAR METABOLISM | 43 | genes.ES.table | 0.47 | 1.6 | 0.076 | 0.54 | 0.73 | 0.51 | 0.35 | 0.33 | 0.33 | 0.17 |
| KEGG DNA REPLICATION | 35 | genes.ES.table | 0.53 | 1.6 | 0.075 | 0.51 | 0.74 | 0.66 | 0.38 | 0.41 | 0.33 | 0.15 |
| KEGG BASE EXCISION REPAIR | 32 | genes.ES.table | 0.44 | 1.4 | 0.14 | 0.59 | 0.93 | 0.47 | 0.36 | 0.3 | 0.46 | 0.17 |
| KEGG HOMOLOGOUS RECOMBINATION | 27 | genes.ES.table | 0.62 | 1.6 | 0.061 | 0.55 | 0.82 | 0.41 | 0.23 | 0.31 | 0.38 | 0.18 |
| KEGG CHEMOKINE SIGNALING PATHWAY | 181 | genes.ES.table | 0.51 | 1.5 | 0.089 | 0.68 | 0.9 | 0.44 | 0.25 | 0.34 | 0.52 | 0.23 |
| KEGG CELL CYCLE | 123 | genes.ES.table | 0.53 | 1.7 | 0.043 | 0.67 | 0.56 | 0.23 | 0.12 | 0.2 | 0.34 | 0.23 |
| KEGG OOCYTE MEIOSIS | 106 | genes.ES.table | 0.42 | 1.5 | 0.094 | 0.68 | 0.91 | 0.21 | 0.14 | 0.18 | 0.52 | 0.23 |
Table S21. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | IL6 | IL6 | IL6 | 260 | 0.45 | 0.094 | YES |
| 2 | C8G | C8G | C8G | 557 | 0.38 | 0.17 | YES |
| 3 | NCAM1 | NCAM1 | NCAM1 | 824 | 0.33 | 0.23 | YES |
| 4 | NCAM2 | NCAM2 | NCAM2 | 949 | 0.32 | 0.3 | YES |
| 5 | C9 | C9 | C9 | 1062 | 0.3 | 0.37 | YES |
| 6 | IL1A | IL1A | IL1A | 1186 | 0.28 | 0.43 | YES |
| 7 | CCL5 | CCL5 | CCL5 | 1522 | 0.25 | 0.47 | YES |
| 8 | C1QB | C1QB | C1QB | 1894 | 0.22 | 0.5 | YES |
| 9 | C1QC | C1QC | C1QC | 2119 | 0.2 | 0.54 | YES |
| 10 | IL1B | IL1B | IL1B | 2148 | 0.2 | 0.58 | YES |
| 11 | C1QA | C1QA | C1QA | 2164 | 0.2 | 0.63 | YES |
| 12 | BAX | BAX | BAX | 4091 | 0.11 | 0.55 | NO |
| 13 | ELK1 | ELK1 | ELK1 | 4856 | 0.084 | 0.53 | NO |
| 14 | MAP2K2 | MAP2K2 | MAP2K2 | 6021 | 0.059 | 0.48 | NO |
| 15 | PRKX | PRKX | PRKX | 6689 | 0.047 | 0.46 | NO |
| 16 | PRNP | PRNP | PRNP | 7173 | 0.04 | 0.44 | NO |
| 17 | C5 | C5 | C5 | 7278 | 0.038 | 0.44 | NO |
| 18 | HSPA5 | HSPA5 | HSPA5 | 7317 | 0.038 | 0.45 | NO |
| 19 | SOD1 | SOD1 | SOD1 | 7875 | 0.029 | 0.42 | NO |
| 20 | STIP1 | STIP1 | STIP1 | 8061 | 0.027 | 0.42 | NO |
| 21 | MAP2K1 | MAP2K1 | MAP2K1 | 8955 | 0.015 | 0.38 | NO |
| 22 | HSPA1A | HSPA1A | HSPA1A | 9585 | 0.0069 | 0.34 | NO |
| 23 | PRKACA | PRKACA | PRKACA | 9732 | 0.0048 | 0.34 | NO |
| 24 | LAMC1 | LAMC1 | LAMC1 | 10575 | -0.0057 | 0.29 | NO |
| 25 | MAPK3 | MAPK3 | MAPK3 | 13069 | -0.039 | 0.16 | NO |
| 26 | PRKACB | PRKACB | PRKACB | 13598 | -0.048 | 0.15 | NO |
| 27 | NOTCH1 | NOTCH1 | NOTCH1 | 13625 | -0.048 | 0.16 | NO |
| 28 | MAPK1 | MAPK1 | MAPK1 | 13695 | -0.05 | 0.16 | NO |
| 29 | FYN | FYN | FYN | 15334 | -0.087 | 0.096 | NO |
| 30 | EGR1 | EGR1 | EGR1 | 15866 | -0.1 | 0.093 | NO |
| 31 | C6 | C6 | C6 | 16822 | -0.16 | 0.079 | NO |
Figure S41. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA INFLAM PATHWAY.
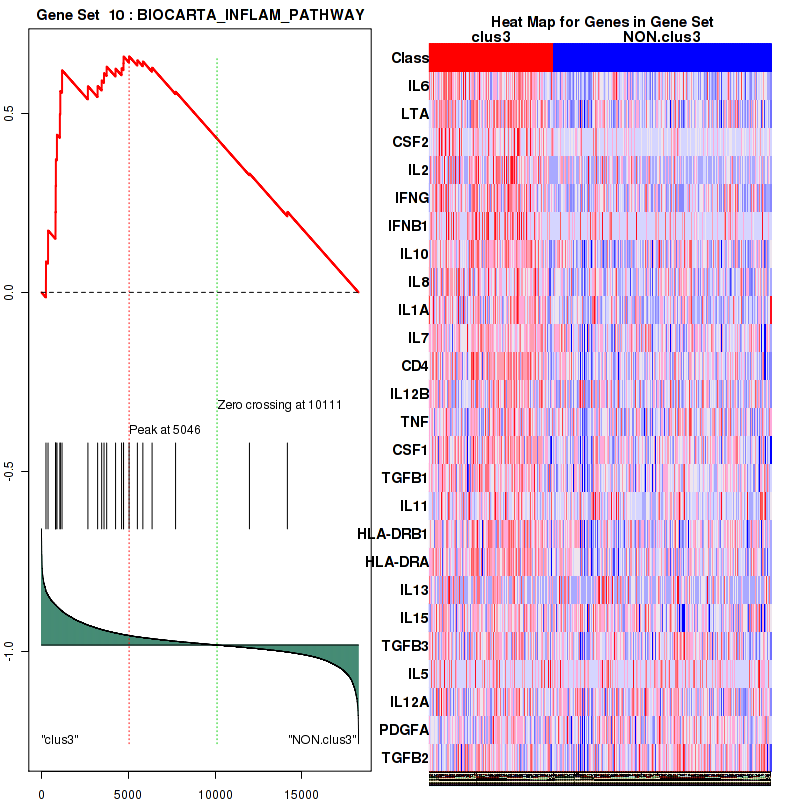
Figure S42. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA INFLAM PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
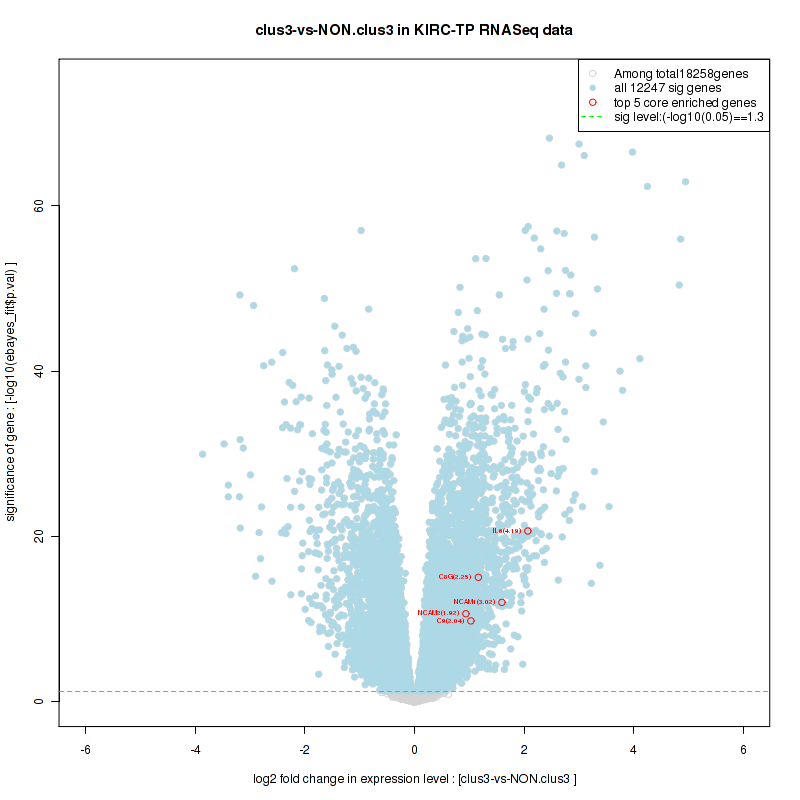
Table S22. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | AIM2 | AIM2 | AIM2 | 96 | 0.55 | 0.099 | YES |
| 2 | IL6 | IL6 | IL6 | 260 | 0.45 | 0.18 | YES |
| 3 | ZBP1 | ZBP1 | ZBP1 | 370 | 0.42 | 0.25 | YES |
| 4 | IKBKE | IKBKE | IKBKE | 390 | 0.41 | 0.32 | YES |
| 5 | PYCARD | PYCARD | PYCARD | 762 | 0.34 | 0.37 | YES |
| 6 | IFNB1 | IFNB1 | IFNB1 | 892 | 0.32 | 0.42 | YES |
| 7 | CCL5 | CCL5 | CCL5 | 1522 | 0.25 | 0.43 | YES |
| 8 | CCL4 | CCL4 | CCL4 | 1702 | 0.23 | 0.47 | YES |
| 9 | IRF7 | IRF7 | IRF7 | 2050 | 0.2 | 0.49 | YES |
| 10 | IL1B | IL1B | IL1B | 2148 | 0.2 | 0.52 | YES |
| 11 | POLR3G | POLR3G | POLR3G | 2152 | 0.2 | 0.56 | YES |
| 12 | CCL4L2 | CCL4L2 | CCL4L2 | 2187 | 0.2 | 0.59 | YES |
| 13 | IL18 | IL18 | IL18 | 2297 | 0.19 | 0.62 | YES |
| 14 | CASP1 | CASP1 | CASP1 | 3214 | 0.14 | 0.6 | NO |
| 15 | RIPK3 | RIPK3 | RIPK3 | 3573 | 0.12 | 0.6 | NO |
| 16 | TMEM173 | TMEM173 | TMEM173 | 4181 | 0.1 | 0.59 | NO |
| 17 | CXCL10 | CXCL10 | CXCL10 | 5000 | 0.08 | 0.56 | NO |
| 18 | IKBKG | IKBKG | IKBKG | 5016 | 0.08 | 0.57 | NO |
| 19 | IRF3 | IRF3 | IRF3 | 5240 | 0.074 | 0.57 | NO |
| 20 | POLR3K | POLR3K | POLR3K | 5243 | 0.074 | 0.59 | NO |
| 21 | TBK1 | TBK1 | TBK1 | 6725 | 0.047 | 0.51 | NO |
| 22 | POLR3D | POLR3D | POLR3D | 8349 | 0.023 | 0.43 | NO |
| 23 | TREX1 | TREX1 | TREX1 | 8395 | 0.022 | 0.43 | NO |
| 24 | NFKBIB | NFKBIB | NFKBIB | 8541 | 0.021 | 0.43 | NO |
| 25 | POLR3H | POLR3H | POLR3H | 8832 | 0.016 | 0.41 | NO |
| 26 | POLR3C | POLR3C | POLR3C | 9033 | 0.014 | 0.41 | NO |
| 27 | IKBKB | IKBKB | IKBKB | 9331 | 0.01 | 0.39 | NO |
| 28 | ADAR | ADAR | ADAR | 9448 | 0.0086 | 0.39 | NO |
| 29 | POLR1C | POLR1C | POLR1C | 9589 | 0.0069 | 0.38 | NO |
| 30 | RELA | RELA | RELA | 9833 | 0.0034 | 0.37 | NO |
| 31 | POLR3A | POLR3A | POLR3A | 9898 | 0.0025 | 0.36 | NO |
| 32 | DDX58 | DDX58 | DDX58 | 9989 | 0.0015 | 0.36 | NO |
| 33 | RIPK1 | RIPK1 | RIPK1 | 10704 | -0.0071 | 0.32 | NO |
| 34 | POLR3GL | POLR3GL | POLR3GL | 10808 | -0.0084 | 0.32 | NO |
| 35 | POLR3F | POLR3F | POLR3F | 11188 | -0.013 | 0.3 | NO |
| 36 | MAVS | MAVS | MAVS | 11263 | -0.014 | 0.3 | NO |
| 37 | POLR1D | POLR1D | POLR1D | 11268 | -0.014 | 0.3 | NO |
| 38 | NFKB1 | NFKB1 | NFKB1 | 12805 | -0.035 | 0.22 | NO |
| 39 | IL33 | IL33 | IL33 | 13032 | -0.039 | 0.22 | NO |
| 40 | POLR3B | POLR3B | POLR3B | 13137 | -0.041 | 0.22 | NO |
| 41 | CHUK | CHUK | CHUK | 13895 | -0.053 | 0.19 | NO |
| 42 | NFKBIA | NFKBIA | NFKBIA | 14643 | -0.069 | 0.16 | NO |
| 43 | IFNA13 | IFNA13 | IFNA13 | 15751 | -0.1 | 0.12 | NO |
| 44 | IFNA5 | IFNA5 | IFNA5 | 15835 | -0.1 | 0.13 | NO |
Figure S43. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA GSK3 PATHWAY.
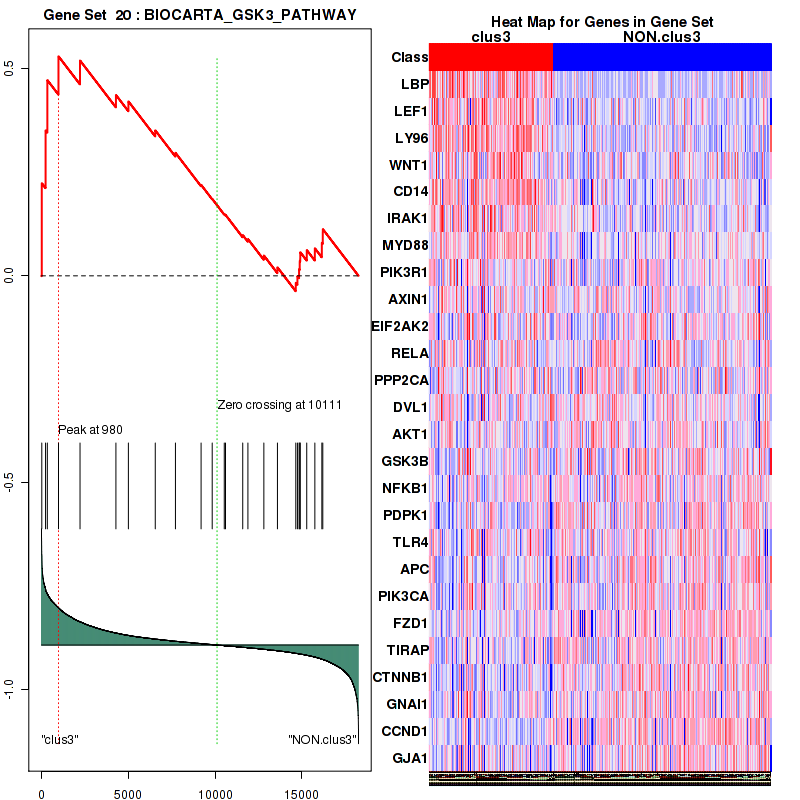
Figure S44. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA GSK3 PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
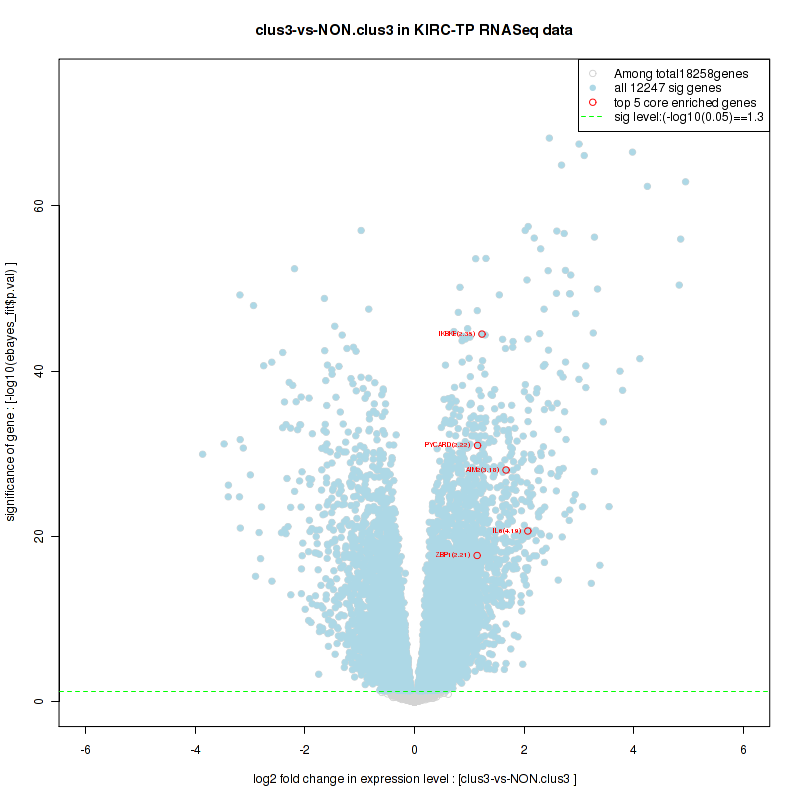
Table S23. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | CDC25C | CDC25C | CDC25C | 156 | 0.51 | 0.028 | YES |
| 2 | BUB1 | BUB1 | BUB1 | 190 | 0.49 | 0.06 | YES |
| 3 | CCNE1 | CCNE1 | CCNE1 | 265 | 0.45 | 0.088 | YES |
| 4 | PTTG1 | PTTG1 | PTTG1 | 277 | 0.44 | 0.12 | YES |
| 5 | PLK1 | PLK1 | PLK1 | 300 | 0.44 | 0.15 | YES |
| 6 | CCNB2 | CCNB2 | CCNB2 | 377 | 0.42 | 0.17 | YES |
| 7 | CDC20 | CDC20 | CDC20 | 407 | 0.41 | 0.2 | YES |
| 8 | PKMYT1 | PKMYT1 | PKMYT1 | 459 | 0.4 | 0.23 | YES |
| 9 | ORC6L | ORC6L | ORC6L | 536 | 0.38 | 0.25 | YES |
| 10 | BUB1B | BUB1B | BUB1B | 566 | 0.37 | 0.27 | YES |
| 11 | TTK | TTK | TTK | 603 | 0.37 | 0.3 | YES |
| 12 | SFN | SFN | SFN | 769 | 0.34 | 0.31 | YES |
| 13 | CDC45 | CDC45 | CDC45 | 771 | 0.34 | 0.34 | YES |
| 14 | E2F2 | E2F2 | E2F2 | 831 | 0.33 | 0.36 | YES |
| 15 | ORC1L | ORC1L | ORC1L | 1000 | 0.31 | 0.37 | YES |
| 16 | CCNA1 | CCNA1 | CCNA1 | 1011 | 0.3 | 0.39 | YES |
| 17 | CHEK2 | CHEK2 | CHEK2 | 1145 | 0.29 | 0.4 | YES |
| 18 | CCNA2 | CCNA2 | CCNA2 | 1149 | 0.29 | 0.42 | YES |
| 19 | ESPL1 | ESPL1 | ESPL1 | 1228 | 0.28 | 0.44 | YES |
| 20 | CDKN2A | CDKN2A | CDKN2A | 1278 | 0.27 | 0.46 | YES |
| 21 | CCNB1 | CCNB1 | CCNB1 | 1380 | 0.26 | 0.47 | YES |
| 22 | CDK1 | CDK1 | CDK1 | 1429 | 0.26 | 0.48 | YES |
| 23 | CDC6 | CDC6 | CDC6 | 1595 | 0.24 | 0.49 | YES |
| 24 | SMC1B | SMC1B | SMC1B | 2010 | 0.21 | 0.48 | YES |
| 25 | CCNE2 | CCNE2 | CCNE2 | 2026 | 0.21 | 0.5 | YES |
| 26 | CDC25A | CDC25A | CDC25A | 2053 | 0.2 | 0.51 | YES |
| 27 | CDC7 | CDC7 | CDC7 | 2185 | 0.2 | 0.52 | YES |
| 28 | MAD2L1 | MAD2L1 | MAD2L1 | 2284 | 0.19 | 0.53 | YES |
| 29 | E2F1 | E2F1 | E2F1 | 2905 | 0.16 | 0.5 | NO |
| 30 | CDC25B | CDC25B | CDC25B | 2988 | 0.15 | 0.51 | NO |
| 31 | MAD2L2 | MAD2L2 | MAD2L2 | 3709 | 0.12 | 0.48 | NO |
| 32 | E2F3 | E2F3 | E2F3 | 4130 | 0.1 | 0.46 | NO |
| 33 | CHEK1 | CHEK1 | CHEK1 | 4160 | 0.1 | 0.47 | NO |
| 34 | MCM6 | MCM6 | MCM6 | 4264 | 0.1 | 0.47 | NO |
| 35 | TGFB1 | TGFB1 | TGFB1 | 4266 | 0.1 | 0.48 | NO |
| 36 | MCM2 | MCM2 | MCM2 | 4479 | 0.094 | 0.47 | NO |
| 37 | CDK7 | CDK7 | CDK7 | 4744 | 0.086 | 0.46 | NO |
| 38 | MAD1L1 | MAD1L1 | MAD1L1 | 4864 | 0.084 | 0.46 | NO |
| 39 | PTTG2 | PTTG2 | PTTG2 | 4873 | 0.083 | 0.47 | NO |
| 40 | ANAPC11 | ANAPC11 | ANAPC11 | 5130 | 0.077 | 0.46 | NO |
| 41 | MCM4 | MCM4 | MCM4 | 5162 | 0.076 | 0.46 | NO |
| 42 | ANAPC5 | ANAPC5 | ANAPC5 | 5238 | 0.074 | 0.46 | NO |
| 43 | YWHAG | YWHAG | YWHAG | 5383 | 0.071 | 0.46 | NO |
| 44 | RBX1 | RBX1 | RBX1 | 5560 | 0.068 | 0.46 | NO |
| 45 | PCNA | PCNA | PCNA | 5756 | 0.064 | 0.45 | NO |
| 46 | CDK4 | CDK4 | CDK4 | 5814 | 0.063 | 0.45 | NO |
| 47 | TGFB3 | TGFB3 | TGFB3 | 5839 | 0.062 | 0.45 | NO |
| 48 | ORC5L | ORC5L | ORC5L | 5900 | 0.061 | 0.46 | NO |
| 49 | MDM2 | MDM2 | MDM2 | 5987 | 0.059 | 0.45 | NO |
| 50 | TP53 | TP53 | TP53 | 6107 | 0.057 | 0.45 | NO |
| 51 | MCM5 | MCM5 | MCM5 | 6226 | 0.055 | 0.45 | NO |
| 52 | ATR | ATR | ATR | 6267 | 0.055 | 0.45 | NO |
| 53 | CDK2 | CDK2 | CDK2 | 6339 | 0.054 | 0.45 | NO |
| 54 | CCNH | CCNH | CCNH | 6559 | 0.05 | 0.44 | NO |
| 55 | MYC | MYC | MYC | 6757 | 0.046 | 0.44 | NO |
| 56 | GADD45G | GADD45G | GADD45G | 6915 | 0.044 | 0.43 | NO |
| 57 | CUL1 | CUL1 | CUL1 | 7041 | 0.042 | 0.42 | NO |
| 58 | YWHAZ | YWHAZ | YWHAZ | 7119 | 0.04 | 0.42 | NO |
| 59 | ANAPC7 | ANAPC7 | ANAPC7 | 7298 | 0.038 | 0.42 | NO |
| 60 | CDKN2D | CDKN2D | CDKN2D | 7395 | 0.036 | 0.41 | NO |
| 61 | CDK6 | CDK6 | CDK6 | 7442 | 0.036 | 0.41 | NO |
| 62 | CCNB3 | CCNB3 | CCNB3 | 7579 | 0.034 | 0.41 | NO |
| 63 | E2F4 | E2F4 | E2F4 | 7651 | 0.032 | 0.41 | NO |
| 64 | CCND2 | CCND2 | CCND2 | 7685 | 0.032 | 0.41 | NO |
| 65 | BUB3 | BUB3 | BUB3 | 7688 | 0.032 | 0.41 | NO |
| 66 | ANAPC4 | ANAPC4 | ANAPC4 | 7798 | 0.03 | 0.41 | NO |
| 67 | GADD45B | GADD45B | GADD45B | 7840 | 0.03 | 0.41 | NO |
| 68 | YWHAQ | YWHAQ | YWHAQ | 8196 | 0.025 | 0.39 | NO |
| 69 | E2F5 | E2F5 | E2F5 | 8213 | 0.025 | 0.39 | NO |
| 70 | CCND3 | CCND3 | CCND3 | 8237 | 0.024 | 0.39 | NO |
| 71 | SKP2 | SKP2 | SKP2 | 8303 | 0.024 | 0.39 | NO |
| 72 | PRKDC | PRKDC | PRKDC | 8462 | 0.022 | 0.38 | NO |
| 73 | CDKN2C | CDKN2C | CDKN2C | 8572 | 0.02 | 0.38 | NO |
| 74 | RBL1 | RBL1 | RBL1 | 8592 | 0.02 | 0.38 | NO |
| 75 | TFDP2 | TFDP2 | TFDP2 | 8599 | 0.02 | 0.38 | NO |
| 76 | ZBTB17 | ZBTB17 | ZBTB17 | 8614 | 0.02 | 0.38 | NO |
| 77 | CDC16 | CDC16 | CDC16 | 8631 | 0.019 | 0.38 | NO |
| 78 | MCM7 | MCM7 | MCM7 | 9239 | 0.011 | 0.34 | NO |
| 79 | FZR1 | FZR1 | FZR1 | 9531 | 0.0075 | 0.33 | NO |
| 80 | ANAPC1 | ANAPC1 | ANAPC1 | 10004 | 0.0014 | 0.3 | NO |
| 81 | HDAC1 | HDAC1 | HDAC1 | 10082 | 0.00028 | 0.3 | NO |
| 82 | ANAPC10 | ANAPC10 | ANAPC10 | 10482 | -0.0046 | 0.28 | NO |
| 83 | GADD45A | GADD45A | GADD45A | 10559 | -0.0056 | 0.27 | NO |
| 84 | YWHAB | YWHAB | YWHAB | 10613 | -0.0062 | 0.27 | NO |
| 85 | MCM3 | MCM3 | MCM3 | 10624 | -0.0063 | 0.27 | NO |
| 86 | RB1 | RB1 | RB1 | 10739 | -0.0076 | 0.27 | NO |
| 87 | CDKN1A | CDKN1A | CDKN1A | 11031 | -0.011 | 0.25 | NO |
| 88 | CDC27 | CDC27 | CDC27 | 11202 | -0.013 | 0.24 | NO |
| 89 | YWHAH | YWHAH | YWHAH | 11315 | -0.015 | 0.24 | NO |
| 90 | ATM | ATM | ATM | 11407 | -0.016 | 0.23 | NO |
| 91 | RAD21 | RAD21 | RAD21 | 11652 | -0.019 | 0.22 | NO |
| 92 | SKP1 | SKP1 | SKP1 | 11804 | -0.021 | 0.21 | NO |
| 93 | GSK3B | GSK3B | GSK3B | 11889 | -0.022 | 0.21 | NO |
| 94 | ORC3L | ORC3L | ORC3L | 12250 | -0.027 | 0.19 | NO |
| 95 | YWHAE | YWHAE | YWHAE | 12383 | -0.029 | 0.19 | NO |
| 96 | SMC1A | SMC1A | SMC1A | 12430 | -0.03 | 0.19 | NO |
| 97 | CDKN1B | CDKN1B | CDKN1B | 12524 | -0.031 | 0.18 | NO |
| 98 | ORC2L | ORC2L | ORC2L | 12553 | -0.031 | 0.18 | NO |
| 99 | ABL1 | ABL1 | ABL1 | 12580 | -0.032 | 0.19 | NO |
| 100 | SMAD2 | SMAD2 | SMAD2 | 12777 | -0.035 | 0.18 | NO |
| 101 | SMAD3 | SMAD3 | SMAD3 | 12834 | -0.036 | 0.18 | NO |
| 102 | CDC23 | CDC23 | CDC23 | 12903 | -0.037 | 0.18 | NO |
| 103 | HDAC2 | HDAC2 | HDAC2 | 12909 | -0.037 | 0.18 | NO |
| 104 | CREBBP | CREBBP | CREBBP | 13220 | -0.042 | 0.16 | NO |
| 105 | RBL2 | RBL2 | RBL2 | 13384 | -0.044 | 0.16 | NO |
| 106 | TFDP1 | TFDP1 | TFDP1 | 13675 | -0.049 | 0.15 | NO |
| 107 | ANAPC13 | ANAPC13 | ANAPC13 | 13907 | -0.053 | 0.14 | NO |
| 108 | SMC3 | SMC3 | SMC3 | 13973 | -0.054 | 0.14 | NO |
| 109 | ORC4L | ORC4L | ORC4L | 13997 | -0.055 | 0.14 | NO |
| 110 | EP300 | EP300 | EP300 | 14084 | -0.056 | 0.14 | NO |
| 111 | TGFB2 | TGFB2 | TGFB2 | 14162 | -0.058 | 0.14 | NO |
| 112 | STAG1 | STAG1 | STAG1 | 14331 | -0.062 | 0.13 | NO |
| 113 | STAG2 | STAG2 | STAG2 | 14392 | -0.063 | 0.14 | NO |
| 114 | WEE1 | WEE1 | WEE1 | 14886 | -0.075 | 0.11 | NO |
| 115 | ANAPC2 | ANAPC2 | ANAPC2 | 14900 | -0.075 | 0.12 | NO |
| 116 | WEE2 | WEE2 | WEE2 | 15232 | -0.084 | 0.11 | NO |
| 117 | CDKN2B | CDKN2B | CDKN2B | 15336 | -0.087 | 0.11 | NO |
| 118 | SMAD4 | SMAD4 | SMAD4 | 15618 | -0.096 | 0.097 | NO |
| 119 | CDC26 | CDC26 | CDC26 | 15759 | -0.1 | 0.097 | NO |
| 120 | CDKN1C | CDKN1C | CDKN1C | 15977 | -0.11 | 0.093 | NO |
| 121 | CCND1 | CCND1 | CCND1 | 16145 | -0.12 | 0.092 | NO |
| 122 | CDC14A | CDC14A | CDC14A | 16669 | -0.15 | 0.073 | NO |
| 123 | CDC14B | CDC14B | CDC14B | 17287 | -0.2 | 0.054 | NO |
Figure S45. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: BIOCARTA WNT PATHWAY.
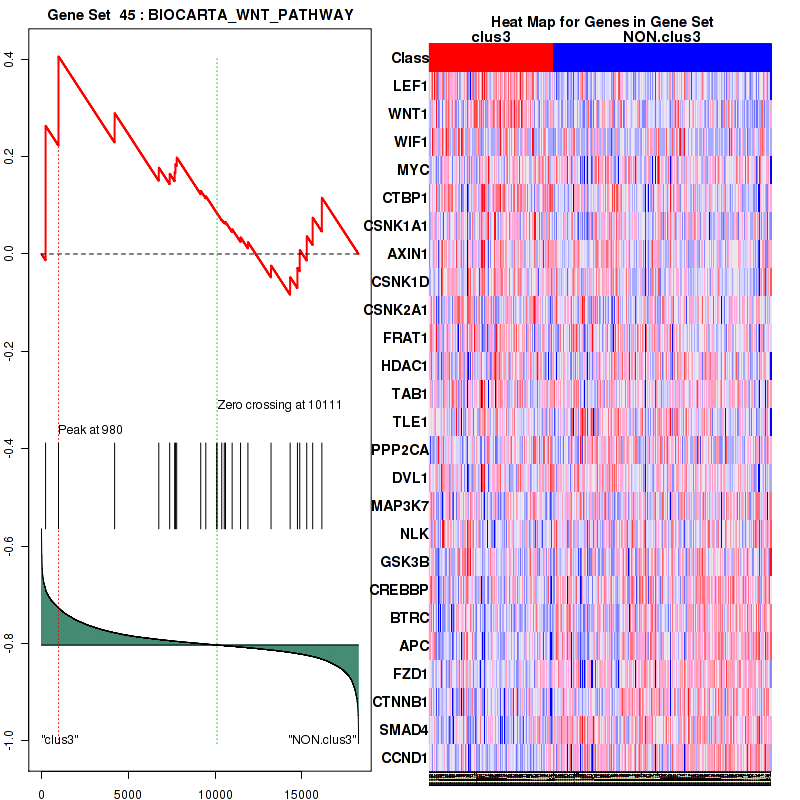
Figure S46. Get High-res Image For the top 5 core enriched genes in the pathway: BIOCARTA WNT PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
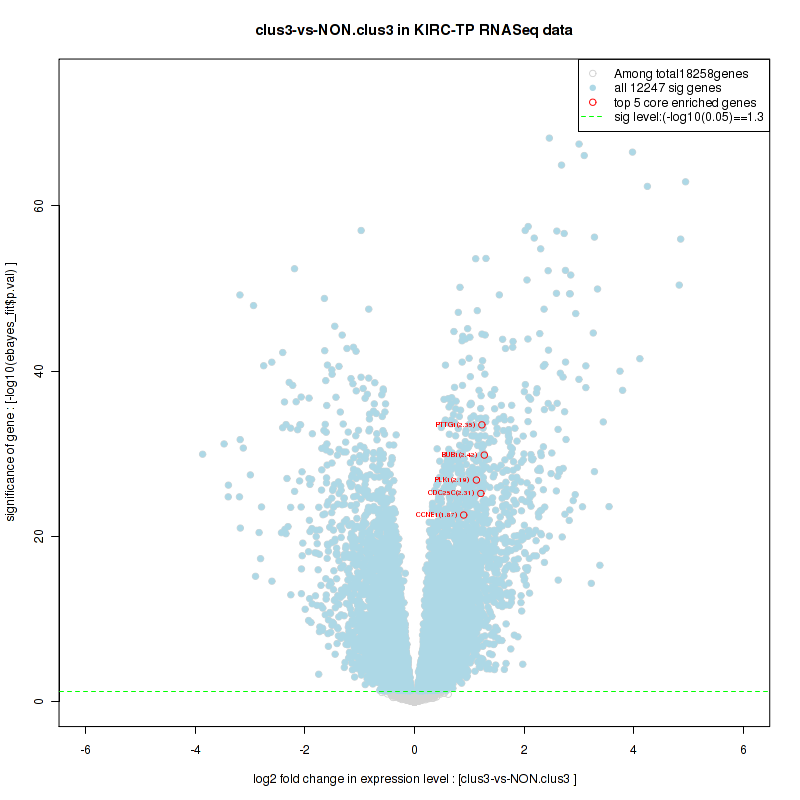
Table S24. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | CCNE1 | CCNE1 | CCNE1 | 265 | 0.45 | 0.043 | YES |
| 2 | GTSE1 | GTSE1 | GTSE1 | 350 | 0.42 | 0.092 | YES |
| 3 | CCNB2 | CCNB2 | CCNB2 | 377 | 0.42 | 0.14 | YES |
| 4 | BAI1 | BAI1 | BAI1 | 462 | 0.4 | 0.19 | YES |
| 5 | RRM2 | RRM2 | RRM2 | 575 | 0.37 | 0.23 | YES |
| 6 | SFN | SFN | SFN | 769 | 0.34 | 0.26 | YES |
| 7 | TP73 | TP73 | TP73 | 987 | 0.31 | 0.29 | YES |
| 8 | CHEK2 | CHEK2 | CHEK2 | 1145 | 0.29 | 0.32 | YES |
| 9 | SERPINE1 | SERPINE1 | SERPINE1 | 1183 | 0.28 | 0.35 | YES |
| 10 | CD82 | CD82 | CD82 | 1231 | 0.28 | 0.39 | YES |
| 11 | PMAIP1 | PMAIP1 | PMAIP1 | 1262 | 0.27 | 0.42 | YES |
| 12 | CDKN2A | CDKN2A | CDKN2A | 1278 | 0.27 | 0.45 | YES |
| 13 | CCNB1 | CCNB1 | CCNB1 | 1380 | 0.26 | 0.48 | YES |
| 14 | CDK1 | CDK1 | CDK1 | 1429 | 0.26 | 0.51 | YES |
| 15 | CCNE2 | CCNE2 | CCNE2 | 2026 | 0.21 | 0.51 | YES |
| 16 | BID | BID | BID | 2506 | 0.18 | 0.5 | YES |
| 17 | BBC3 | BBC3 | BBC3 | 3565 | 0.12 | 0.46 | YES |
| 18 | LRDD | LRDD | LRDD | 3619 | 0.12 | 0.47 | YES |
| 19 | CASP3 | CASP3 | CASP3 | 3639 | 0.12 | 0.49 | YES |
| 20 | CASP8 | CASP8 | CASP8 | 4019 | 0.11 | 0.48 | YES |
| 21 | BAX | BAX | BAX | 4091 | 0.11 | 0.49 | YES |
| 22 | IGFBP3 | IGFBP3 | IGFBP3 | 4112 | 0.11 | 0.5 | YES |
| 23 | CHEK1 | CHEK1 | CHEK1 | 4160 | 0.1 | 0.51 | YES |
| 24 | TP53I3 | TP53I3 | TP53I3 | 5458 | 0.07 | 0.45 | NO |
| 25 | TNFRSF10B | TNFRSF10B | TNFRSF10B | 5476 | 0.069 | 0.46 | NO |
| 26 | IGF1 | IGF1 | IGF1 | 5598 | 0.067 | 0.46 | NO |
| 27 | FAS | FAS | FAS | 5615 | 0.067 | 0.47 | NO |
| 28 | PERP | PERP | PERP | 5721 | 0.065 | 0.47 | NO |
| 29 | CDK4 | CDK4 | CDK4 | 5814 | 0.063 | 0.47 | NO |
| 30 | MDM2 | MDM2 | MDM2 | 5987 | 0.059 | 0.47 | NO |
| 31 | APAF1 | APAF1 | APAF1 | 5991 | 0.059 | 0.48 | NO |
| 32 | TP53 | TP53 | TP53 | 6107 | 0.057 | 0.48 | NO |
| 33 | SHISA5 | SHISA5 | SHISA5 | 6197 | 0.056 | 0.48 | NO |
| 34 | DDB2 | DDB2 | DDB2 | 6263 | 0.055 | 0.49 | NO |
| 35 | ATR | ATR | ATR | 6267 | 0.055 | 0.49 | NO |
| 36 | CDK2 | CDK2 | CDK2 | 6339 | 0.054 | 0.5 | NO |
| 37 | RPRM | RPRM | RPRM | 6420 | 0.052 | 0.5 | NO |
| 38 | ZMAT3 | ZMAT3 | ZMAT3 | 6595 | 0.049 | 0.49 | NO |
| 39 | SESN3 | SESN3 | SESN3 | 6721 | 0.047 | 0.49 | NO |
| 40 | CASP9 | CASP9 | CASP9 | 6841 | 0.045 | 0.49 | NO |
| 41 | GADD45G | GADD45G | GADD45G | 6915 | 0.044 | 0.49 | NO |
| 42 | RFWD2 | RFWD2 | RFWD2 | 6960 | 0.043 | 0.5 | NO |
| 43 | SESN2 | SESN2 | SESN2 | 7141 | 0.04 | 0.49 | NO |
| 44 | CDK6 | CDK6 | CDK6 | 7442 | 0.036 | 0.48 | NO |
| 45 | CCNB3 | CCNB3 | CCNB3 | 7579 | 0.034 | 0.48 | NO |
| 46 | CCND2 | CCND2 | CCND2 | 7685 | 0.032 | 0.48 | NO |
| 47 | GADD45B | GADD45B | GADD45B | 7840 | 0.03 | 0.47 | NO |
| 48 | CCND3 | CCND3 | CCND3 | 8237 | 0.024 | 0.45 | NO |
| 49 | RRM2B | RRM2B | RRM2B | 8318 | 0.023 | 0.45 | NO |
| 50 | CYCS | CYCS | CYCS | 8517 | 0.021 | 0.44 | NO |
| 51 | EI24 | EI24 | EI24 | 9586 | 0.0069 | 0.38 | NO |
| 52 | SERPINB5 | SERPINB5 | SERPINB5 | 9793 | 0.0039 | 0.37 | NO |
| 53 | MDM4 | MDM4 | MDM4 | 10234 | -0.0015 | 0.35 | NO |
| 54 | GADD45A | GADD45A | GADD45A | 10559 | -0.0056 | 0.33 | NO |
| 55 | TSC2 | TSC2 | TSC2 | 11004 | -0.011 | 0.31 | NO |
| 56 | CDKN1A | CDKN1A | CDKN1A | 11031 | -0.011 | 0.31 | NO |
| 57 | ATM | ATM | ATM | 11407 | -0.016 | 0.29 | NO |
| 58 | SIAH1 | SIAH1 | SIAH1 | 12035 | -0.024 | 0.26 | NO |
| 59 | SESN1 | SESN1 | SESN1 | 12461 | -0.03 | 0.24 | NO |
| 60 | PTEN | PTEN | PTEN | 12629 | -0.032 | 0.24 | NO |
| 61 | THBS1 | THBS1 | THBS1 | 13841 | -0.052 | 0.18 | NO |
| 62 | CCNG2 | CCNG2 | CCNG2 | 14301 | -0.061 | 0.16 | NO |
| 63 | CCNG1 | CCNG1 | CCNG1 | 14816 | -0.073 | 0.14 | NO |
| 64 | TP53AIP1 | TP53AIP1 | TP53AIP1 | 14974 | -0.077 | 0.14 | NO |
| 65 | PPM1D | PPM1D | PPM1D | 15479 | -0.091 | 0.12 | NO |
| 66 | RCHY1 | RCHY1 | RCHY1 | 15787 | -0.1 | 0.12 | NO |
| 67 | CCND1 | CCND1 | CCND1 | 16145 | -0.12 | 0.12 | NO |
Figure S47. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG AMINO SUGAR AND NUCLEOTIDE SUGAR METABOLISM.
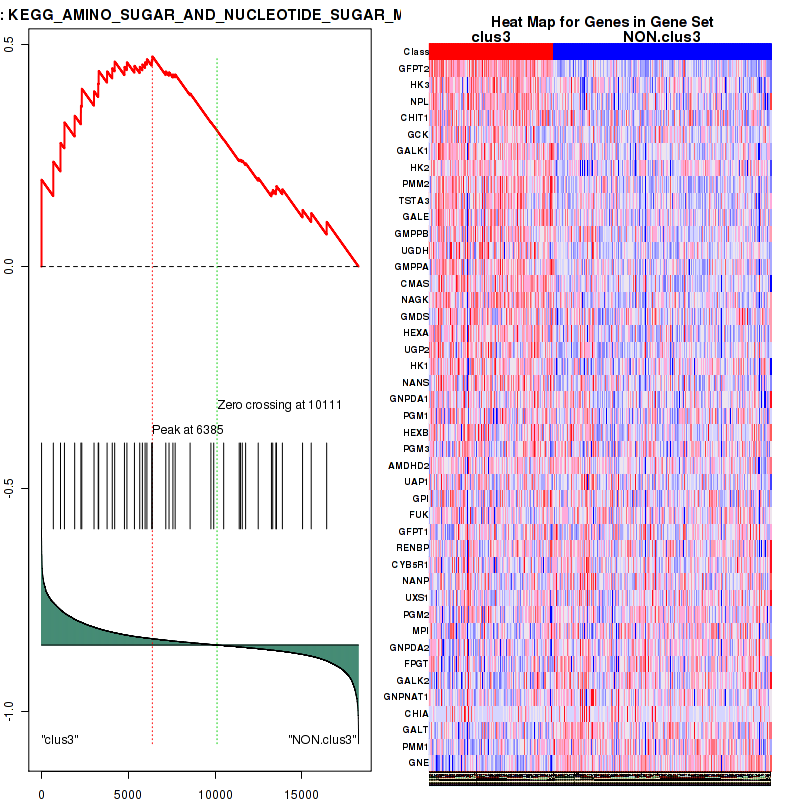
Figure S48. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG AMINO SUGAR AND NUCLEOTIDE SUGAR METABOLISM, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
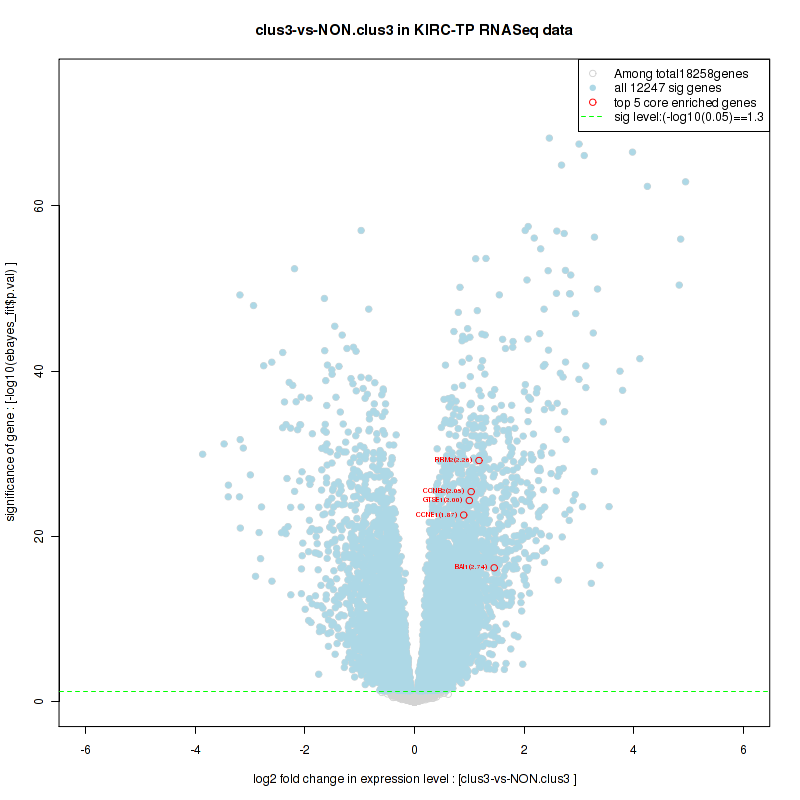
Table S25. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | IL6 | IL6 | IL6 | 260 | 0.45 | 0.04 | YES |
| 2 | CASP5 | CASP5 | CASP5 | 293 | 0.44 | 0.09 | YES |
| 3 | NOD2 | NOD2 | NOD2 | 402 | 0.41 | 0.13 | YES |
| 4 | MEFV | MEFV | MEFV | 591 | 0.37 | 0.17 | YES |
| 5 | CARD9 | CARD9 | CARD9 | 672 | 0.36 | 0.2 | YES |
| 6 | PYCARD | PYCARD | PYCARD | 762 | 0.34 | 0.24 | YES |
| 7 | PSTPIP1 | PSTPIP1 | PSTPIP1 | 956 | 0.31 | 0.27 | YES |
| 8 | CCL11 | CCL11 | CCL11 | 1057 | 0.3 | 0.3 | YES |
| 9 | IL8 | IL8 | IL8 | 1086 | 0.3 | 0.33 | YES |
| 10 | PYDC1 | PYDC1 | PYDC1 | 1230 | 0.28 | 0.36 | YES |
| 11 | CCL13 | CCL13 | CCL13 | 1268 | 0.27 | 0.39 | YES |
| 12 | NLRC4 | NLRC4 | NLRC4 | 1342 | 0.27 | 0.41 | YES |
| 13 | CCL7 | CCL7 | CCL7 | 1366 | 0.26 | 0.44 | YES |
| 14 | CXCL1 | CXCL1 | CXCL1 | 1427 | 0.26 | 0.47 | YES |
| 15 | BIRC3 | BIRC3 | BIRC3 | 1435 | 0.26 | 0.5 | YES |
| 16 | CXCL2 | CXCL2 | CXCL2 | 1503 | 0.25 | 0.53 | YES |
| 17 | CCL5 | CCL5 | CCL5 | 1522 | 0.25 | 0.56 | YES |
| 18 | NAIP | NAIP | NAIP | 1880 | 0.22 | 0.56 | YES |
| 19 | IL1B | IL1B | IL1B | 2148 | 0.2 | 0.57 | YES |
| 20 | IL18 | IL18 | IL18 | 2297 | 0.19 | 0.58 | YES |
| 21 | CCL8 | CCL8 | CCL8 | 2434 | 0.18 | 0.6 | YES |
| 22 | RIPK2 | RIPK2 | RIPK2 | 2488 | 0.18 | 0.62 | YES |
| 23 | NLRP3 | NLRP3 | NLRP3 | 2928 | 0.15 | 0.61 | NO |
| 24 | CASP1 | CASP1 | CASP1 | 3214 | 0.14 | 0.61 | NO |
| 25 | TNF | TNF | TNF | 3603 | 0.12 | 0.61 | NO |
| 26 | CASP8 | CASP8 | CASP8 | 4019 | 0.11 | 0.6 | NO |
| 27 | HSP90B1 | HSP90B1 | HSP90B1 | 4638 | 0.09 | 0.57 | NO |
| 28 | IKBKG | IKBKG | IKBKG | 5016 | 0.08 | 0.56 | NO |
| 29 | TRIP6 | TRIP6 | TRIP6 | 5290 | 0.073 | 0.56 | NO |
| 30 | MAPK11 | MAPK11 | MAPK11 | 5548 | 0.068 | 0.55 | NO |
| 31 | TNFAIP3 | TNFAIP3 | TNFAIP3 | 6285 | 0.055 | 0.52 | NO |
| 32 | MAPK13 | MAPK13 | MAPK13 | 6640 | 0.048 | 0.5 | NO |
| 33 | NLRP1 | NLRP1 | NLRP1 | 7749 | 0.031 | 0.44 | NO |
| 34 | NFKBIB | NFKBIB | NFKBIB | 8541 | 0.021 | 0.4 | NO |
| 35 | MAPK9 | MAPK9 | MAPK9 | 8589 | 0.02 | 0.4 | NO |
| 36 | NOD1 | NOD1 | NOD1 | 8680 | 0.019 | 0.4 | NO |
| 37 | BIRC2 | BIRC2 | BIRC2 | 9091 | 0.013 | 0.38 | NO |
| 38 | MAPK14 | MAPK14 | MAPK14 | 9234 | 0.011 | 0.37 | NO |
| 39 | IKBKB | IKBKB | IKBKB | 9331 | 0.01 | 0.37 | NO |
| 40 | ERBB2IP | ERBB2IP | ERBB2IP | 9405 | 0.0092 | 0.37 | NO |
| 41 | CARD8 | CARD8 | CARD8 | 9574 | 0.0071 | 0.36 | NO |
| 42 | RELA | RELA | RELA | 9833 | 0.0034 | 0.34 | NO |
| 43 | SUGT1 | SUGT1 | SUGT1 | 10073 | 0.0004 | 0.33 | NO |
| 44 | TAB1 | TAB1 | TAB1 | 10118 | -0.000062 | 0.33 | NO |
| 45 | XIAP | XIAP | XIAP | 10159 | -0.00064 | 0.33 | NO |
| 46 | MAP3K7 | MAP3K7 | MAP3K7 | 10980 | -0.01 | 0.28 | NO |
| 47 | TAB3 | TAB3 | TAB3 | 10983 | -0.011 | 0.28 | NO |
| 48 | TAB2 | TAB2 | TAB2 | 11362 | -0.016 | 0.26 | NO |
| 49 | HSP90AA1 | HSP90AA1 | HSP90AA1 | 12550 | -0.031 | 0.2 | NO |
| 50 | NFKB1 | NFKB1 | NFKB1 | 12805 | -0.035 | 0.19 | NO |
| 51 | MAPK3 | MAPK3 | MAPK3 | 13069 | -0.039 | 0.18 | NO |
| 52 | CARD6 | CARD6 | CARD6 | 13331 | -0.043 | 0.18 | NO |
| 53 | MAPK1 | MAPK1 | MAPK1 | 13695 | -0.05 | 0.16 | NO |
| 54 | CHUK | CHUK | CHUK | 13895 | -0.053 | 0.16 | NO |
| 55 | CCL2 | CCL2 | CCL2 | 14285 | -0.061 | 0.14 | NO |
| 56 | MAPK12 | MAPK12 | MAPK12 | 14300 | -0.061 | 0.15 | NO |
| 57 | NFKBIA | NFKBIA | NFKBIA | 14643 | -0.069 | 0.14 | NO |
| 58 | TRAF6 | TRAF6 | TRAF6 | 16737 | -0.15 | 0.041 | NO |
| 59 | MAPK8 | MAPK8 | MAPK8 | 17022 | -0.18 | 0.046 | NO |
| 60 | MAPK10 | MAPK10 | MAPK10 | 17066 | -0.18 | 0.066 | NO |
Figure S49. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG DNA REPLICATION.
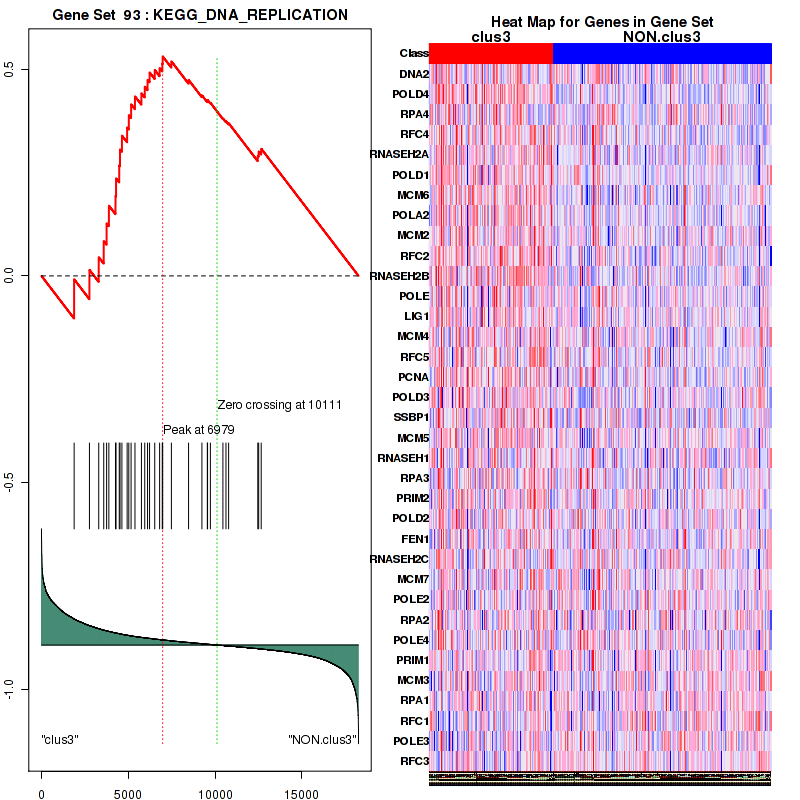
Figure S50. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG DNA REPLICATION, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
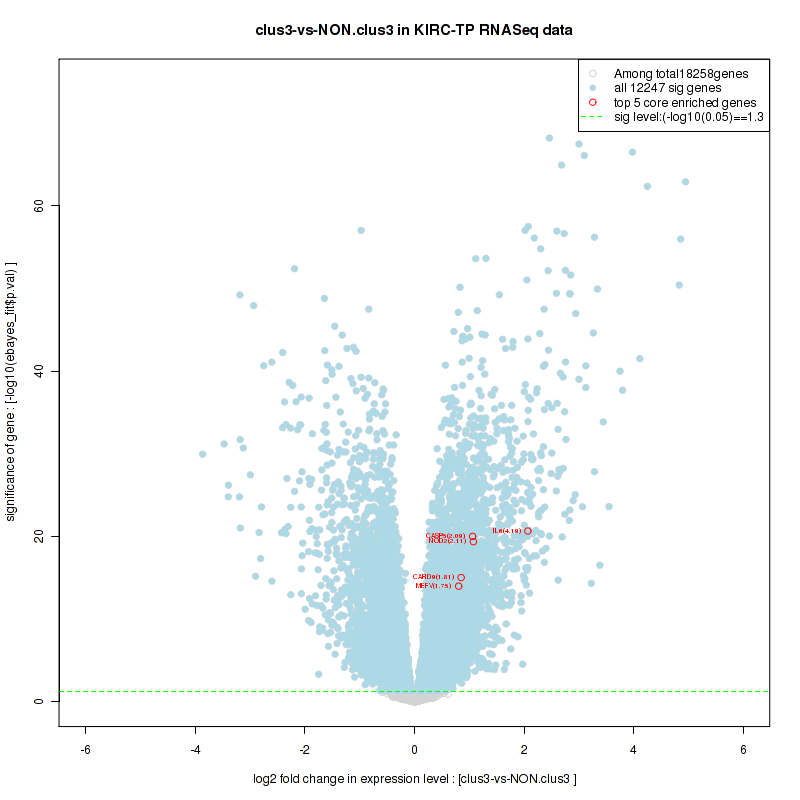
Table S26. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | LBP | LBP | LBP | 18 | 0.75 | 0.22 | YES |
| 2 | LEF1 | LEF1 | LEF1 | 237 | 0.47 | 0.35 | YES |
| 3 | LY96 | LY96 | LY96 | 339 | 0.42 | 0.47 | YES |
| 4 | WNT1 | WNT1 | WNT1 | 980 | 0.31 | 0.53 | YES |
| 5 | CD14 | CD14 | CD14 | 2220 | 0.19 | 0.52 | NO |
| 6 | IRAK1 | IRAK1 | IRAK1 | 4288 | 0.1 | 0.44 | NO |
| 7 | MYD88 | MYD88 | MYD88 | 5007 | 0.08 | 0.42 | NO |
| 8 | PIK3R1 | PIK3R1 | PIK3R1 | 6554 | 0.05 | 0.35 | NO |
| 9 | AXIN1 | AXIN1 | AXIN1 | 7714 | 0.032 | 0.3 | NO |
| 10 | EIF2AK2 | EIF2AK2 | EIF2AK2 | 9192 | 0.012 | 0.22 | NO |
| 11 | RELA | RELA | RELA | 9833 | 0.0034 | 0.19 | NO |
| 12 | PPP2CA | PPP2CA | PPP2CA | 10526 | -0.0052 | 0.15 | NO |
| 13 | DVL1 | DVL1 | DVL1 | 10599 | -0.006 | 0.15 | NO |
| 14 | AKT1 | AKT1 | AKT1 | 11589 | -0.018 | 0.098 | NO |
| 15 | GSK3B | GSK3B | GSK3B | 11889 | -0.022 | 0.089 | NO |
| 16 | NFKB1 | NFKB1 | NFKB1 | 12805 | -0.035 | 0.049 | NO |
| 17 | PDPK1 | PDPK1 | PDPK1 | 13588 | -0.048 | 0.021 | NO |
| 18 | TLR4 | TLR4 | TLR4 | 14645 | -0.069 | -0.017 | NO |
| 19 | APC | APC | APC | 14740 | -0.071 | -0.00076 | NO |
| 20 | PIK3CA | PIK3CA | PIK3CA | 14835 | -0.073 | 0.016 | NO |
| 21 | FZD1 | FZD1 | FZD1 | 14873 | -0.074 | 0.036 | NO |
| 22 | TIRAP | TIRAP | TIRAP | 14904 | -0.075 | 0.057 | NO |
| 23 | CTNNB1 | CTNNB1 | CTNNB1 | 15277 | -0.086 | 0.062 | NO |
| 24 | GNAI1 | GNAI1 | GNAI1 | 15742 | -0.1 | 0.067 | NO |
| 25 | CCND1 | CCND1 | CCND1 | 16145 | -0.12 | 0.08 | NO |
| 26 | GJA1 | GJA1 | GJA1 | 16201 | -0.12 | 0.11 | NO |
Figure S51. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG BASE EXCISION REPAIR.
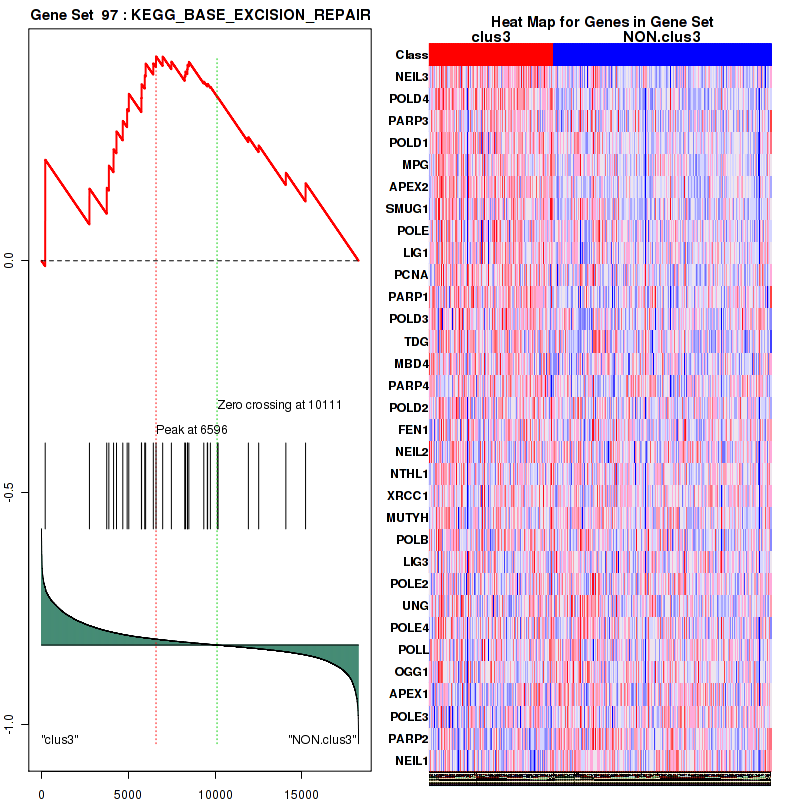
Figure S52. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG BASE EXCISION REPAIR, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
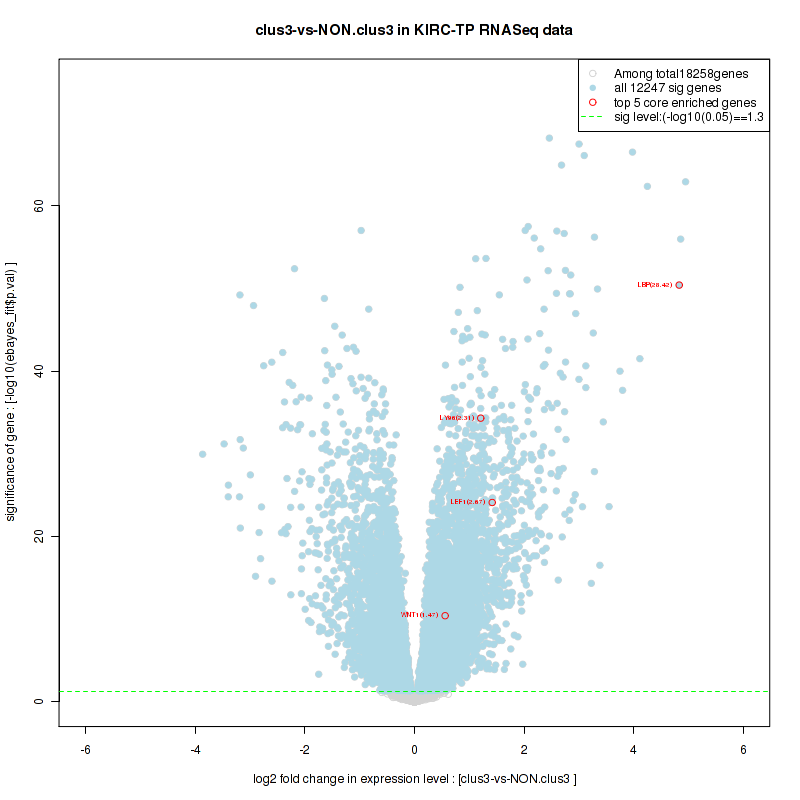
Table S27. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | GFPT2 | GFPT2 | GFPT2 | 4 | 0.89 | 0.2 | YES |
| 2 | HK3 | HK3 | HK3 | 683 | 0.35 | 0.24 | YES |
| 3 | NPL | NPL | NPL | 1096 | 0.29 | 0.28 | YES |
| 4 | CHIT1 | CHIT1 | CHIT1 | 1324 | 0.27 | 0.32 | YES |
| 5 | GCK | GCK | GCK | 1912 | 0.22 | 0.34 | YES |
| 6 | GALK1 | GALK1 | GALK1 | 2272 | 0.19 | 0.36 | YES |
| 7 | HK2 | HK2 | HK2 | 2323 | 0.19 | 0.4 | YES |
| 8 | PMM2 | PMM2 | PMM2 | 3026 | 0.15 | 0.39 | YES |
| 9 | TSTA3 | TSTA3 | TSTA3 | 3269 | 0.14 | 0.41 | YES |
| 10 | GALE | GALE | GALE | 3295 | 0.14 | 0.44 | YES |
| 11 | GMPPB | GMPPB | GMPPB | 3779 | 0.12 | 0.44 | YES |
| 12 | UGDH | UGDH | UGDH | 4077 | 0.11 | 0.45 | YES |
| 13 | GMPPA | GMPPA | GMPPA | 4216 | 0.1 | 0.46 | YES |
| 14 | CMAS | CMAS | CMAS | 4780 | 0.086 | 0.45 | YES |
| 15 | NAGK | NAGK | NAGK | 4920 | 0.082 | 0.46 | YES |
| 16 | GMDS | GMDS | GMDS | 5354 | 0.072 | 0.45 | YES |
| 17 | HEXA | HEXA | HEXA | 5660 | 0.066 | 0.45 | YES |
| 18 | UGP2 | UGP2 | UGP2 | 5812 | 0.063 | 0.46 | YES |
| 19 | HK1 | HK1 | HK1 | 5967 | 0.06 | 0.46 | YES |
| 20 | NANS | NANS | NANS | 6072 | 0.058 | 0.47 | YES |
| 21 | GNPDA1 | GNPDA1 | GNPDA1 | 6347 | 0.053 | 0.46 | YES |
| 22 | PGM1 | PGM1 | PGM1 | 6385 | 0.052 | 0.47 | YES |
| 23 | HEXB | HEXB | HEXB | 7160 | 0.04 | 0.44 | NO |
| 24 | PGM3 | PGM3 | PGM3 | 7346 | 0.037 | 0.44 | NO |
| 25 | AMDHD2 | AMDHD2 | AMDHD2 | 7570 | 0.034 | 0.43 | NO |
| 26 | UAP1 | UAP1 | UAP1 | 7697 | 0.032 | 0.43 | NO |
| 27 | GPI | GPI | GPI | 8560 | 0.02 | 0.39 | NO |
| 28 | FUK | FUK | FUK | 9758 | 0.0045 | 0.32 | NO |
| 29 | GFPT1 | GFPT1 | GFPT1 | 9922 | 0.0022 | 0.32 | NO |
| 30 | RENBP | RENBP | RENBP | 10493 | -0.0047 | 0.29 | NO |
| 31 | CYB5R1 | CYB5R1 | CYB5R1 | 11387 | -0.016 | 0.24 | NO |
| 32 | NANP | NANP | NANP | 11458 | -0.017 | 0.24 | NO |
| 33 | UXS1 | UXS1 | UXS1 | 11562 | -0.018 | 0.24 | NO |
| 34 | PGM2 | PGM2 | PGM2 | 11756 | -0.021 | 0.23 | NO |
| 35 | MPI | MPI | MPI | 12483 | -0.03 | 0.2 | NO |
| 36 | GNPDA2 | GNPDA2 | GNPDA2 | 13244 | -0.042 | 0.17 | NO |
| 37 | FPGT | FPGT | FPGT | 13319 | -0.043 | 0.17 | NO |
| 38 | GALK2 | GALK2 | GALK2 | 13500 | -0.046 | 0.17 | NO |
| 39 | GNPNAT1 | GNPNAT1 | GNPNAT1 | 13530 | -0.047 | 0.18 | NO |
| 40 | CHIA | CHIA | CHIA | 13872 | -0.053 | 0.17 | NO |
| 41 | GALT | GALT | GALT | 15035 | -0.078 | 0.13 | NO |
| 42 | PMM1 | PMM1 | PMM1 | 15535 | -0.093 | 0.12 | NO |
| 43 | GNE | GNE | GNE | 16424 | -0.13 | 0.1 | NO |
Figure S53. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG HOMOLOGOUS RECOMBINATION.
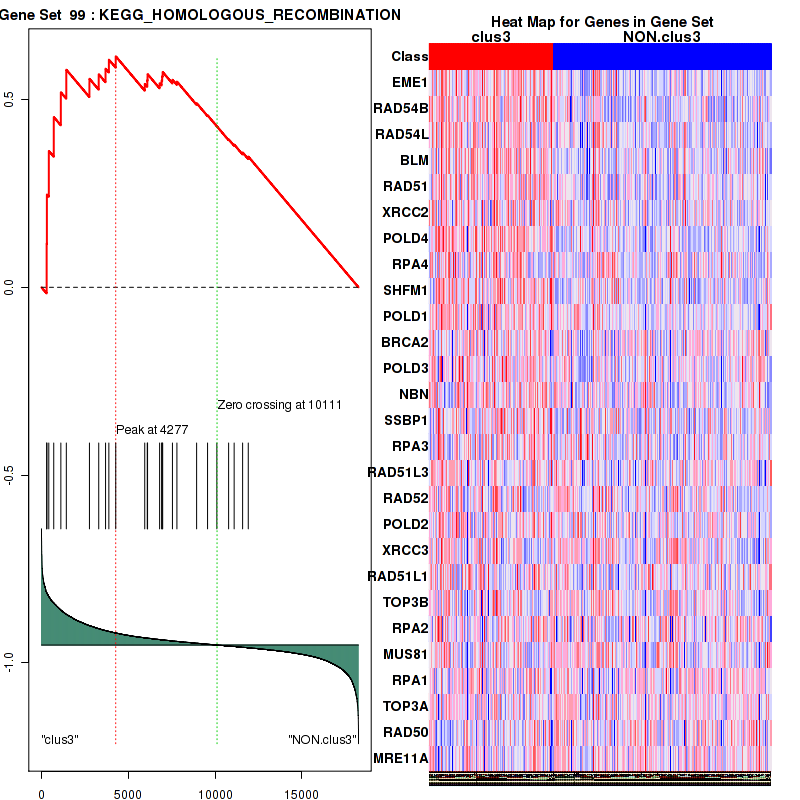
Figure S54. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG HOMOLOGOUS RECOMBINATION, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
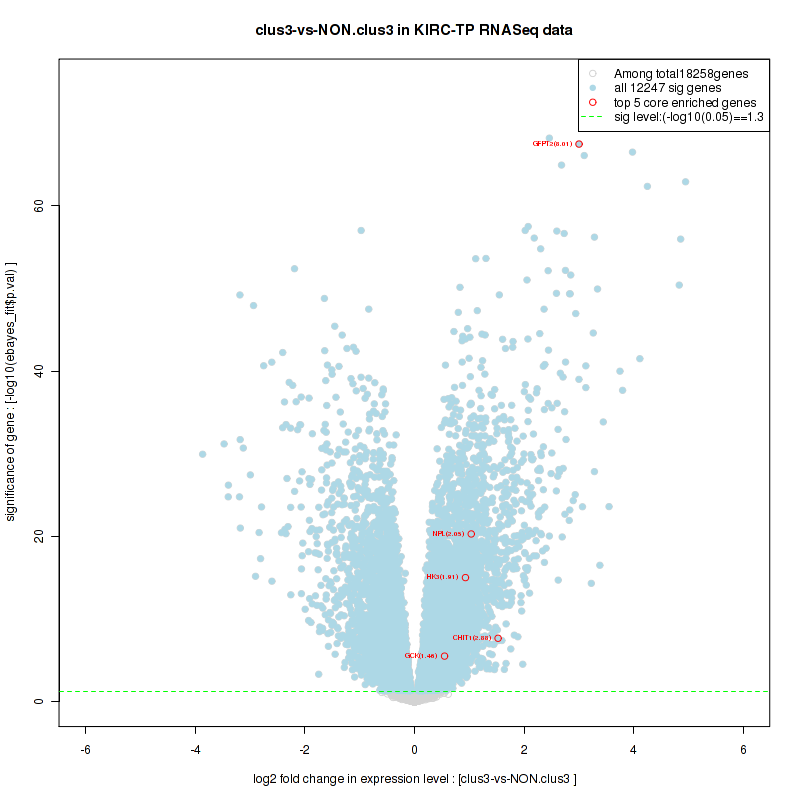
Table S28. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | DNA2 | DNA2 | DNA2 | 1881 | 0.22 | -0.0081 | YES |
| 2 | POLD4 | POLD4 | POLD4 | 2763 | 0.16 | 0.015 | YES |
| 3 | RPA4 | RPA4 | RPA4 | 3298 | 0.14 | 0.045 | YES |
| 4 | RFC4 | RFC4 | RFC4 | 3586 | 0.12 | 0.084 | YES |
| 5 | RNASEH2A | RNASEH2A | RNASEH2A | 3752 | 0.12 | 0.13 | YES |
| 6 | POLD1 | POLD1 | POLD1 | 3882 | 0.11 | 0.17 | YES |
| 7 | MCM6 | MCM6 | MCM6 | 4264 | 0.1 | 0.19 | YES |
| 8 | POLA2 | POLA2 | POLA2 | 4295 | 0.1 | 0.24 | YES |
| 9 | MCM2 | MCM2 | MCM2 | 4479 | 0.094 | 0.27 | YES |
| 10 | RFC2 | RFC2 | RFC2 | 4524 | 0.093 | 0.31 | YES |
| 11 | RNASEH2B | RNASEH2B | RNASEH2B | 4634 | 0.09 | 0.34 | YES |
| 12 | POLE | POLE | POLE | 4934 | 0.082 | 0.36 | YES |
| 13 | LIG1 | LIG1 | LIG1 | 5030 | 0.08 | 0.39 | YES |
| 14 | MCM4 | MCM4 | MCM4 | 5162 | 0.076 | 0.42 | YES |
| 15 | RFC5 | RFC5 | RFC5 | 5384 | 0.071 | 0.43 | YES |
| 16 | PCNA | PCNA | PCNA | 5756 | 0.064 | 0.44 | YES |
| 17 | POLD3 | POLD3 | POLD3 | 5948 | 0.06 | 0.46 | YES |
| 18 | SSBP1 | SSBP1 | SSBP1 | 6110 | 0.057 | 0.47 | YES |
| 19 | MCM5 | MCM5 | MCM5 | 6226 | 0.055 | 0.49 | YES |
| 20 | RNASEH1 | RNASEH1 | RNASEH1 | 6522 | 0.05 | 0.5 | YES |
| 21 | RPA3 | RPA3 | RPA3 | 6804 | 0.046 | 0.5 | YES |
| 22 | PRIM2 | PRIM2 | PRIM2 | 6958 | 0.043 | 0.51 | YES |
| 23 | POLD2 | POLD2 | POLD2 | 6979 | 0.043 | 0.53 | YES |
| 24 | FEN1 | FEN1 | FEN1 | 7482 | 0.035 | 0.52 | NO |
| 25 | RNASEH2C | RNASEH2C | RNASEH2C | 8472 | 0.021 | 0.47 | NO |
| 26 | MCM7 | MCM7 | MCM7 | 9239 | 0.011 | 0.44 | NO |
| 27 | POLE2 | POLE2 | POLE2 | 9550 | 0.0073 | 0.42 | NO |
| 28 | RPA2 | RPA2 | RPA2 | 9564 | 0.0072 | 0.42 | NO |
| 29 | POLE4 | POLE4 | POLE4 | 9729 | 0.0049 | 0.42 | NO |
| 30 | PRIM1 | PRIM1 | PRIM1 | 10450 | -0.0042 | 0.38 | NO |
| 31 | MCM3 | MCM3 | MCM3 | 10624 | -0.0063 | 0.37 | NO |
| 32 | RPA1 | RPA1 | RPA1 | 10779 | -0.0081 | 0.37 | NO |
| 33 | RFC1 | RFC1 | RFC1 | 12456 | -0.03 | 0.29 | NO |
| 34 | POLE3 | POLE3 | POLE3 | 12508 | -0.031 | 0.3 | NO |
| 35 | RFC3 | RFC3 | RFC3 | 12655 | -0.033 | 0.31 | NO |
Figure S55. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG CHEMOKINE SIGNALING PATHWAY.
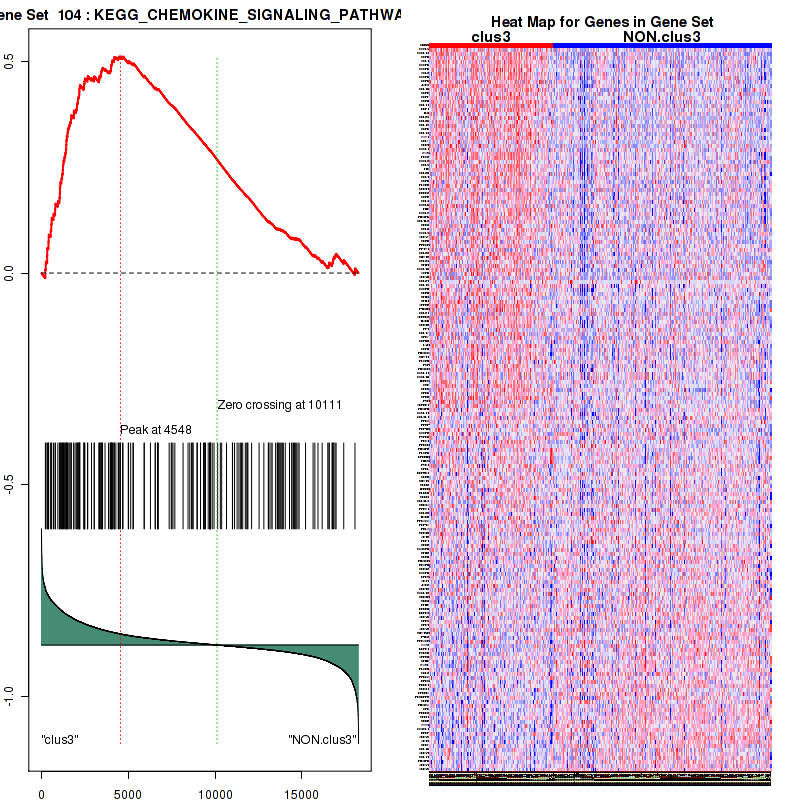
Figure S56. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG CHEMOKINE SIGNALING PATHWAY, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
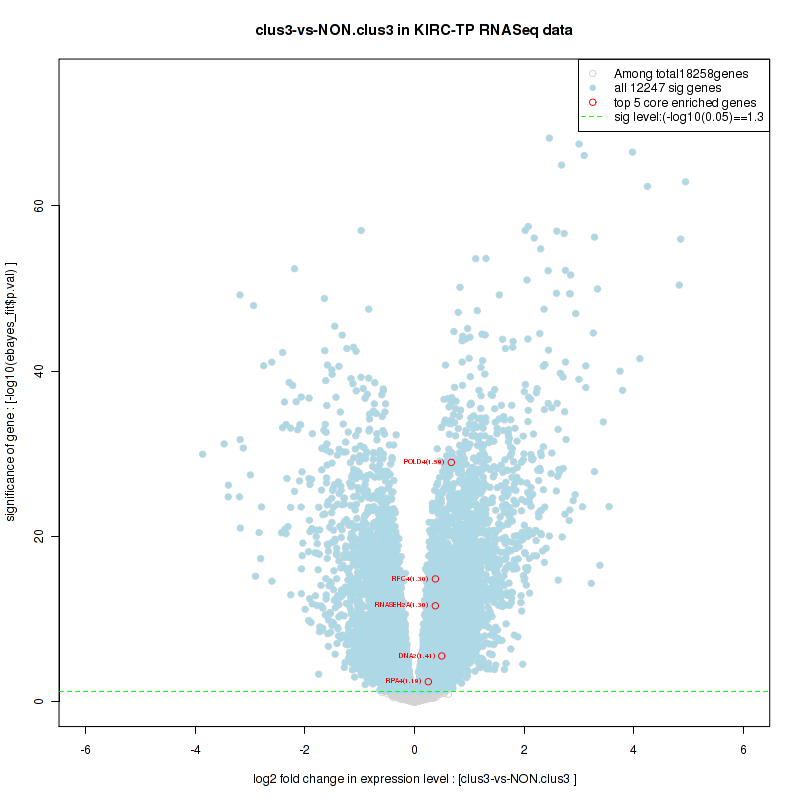
Table S29. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | F2 | F2 | F2 | 6 | 0.87 | 0.063 | YES |
| 2 | FGA | FGA | FGA | 200 | 0.48 | 0.088 | YES |
| 3 | C1S | C1S | C1S | 251 | 0.46 | 0.12 | YES |
| 4 | C1R | C1R | C1R | 276 | 0.44 | 0.15 | YES |
| 5 | C2 | C2 | C2 | 384 | 0.41 | 0.17 | YES |
| 6 | FGG | FGG | FGG | 453 | 0.4 | 0.2 | YES |
| 7 | C8G | C8G | C8G | 557 | 0.38 | 0.22 | YES |
| 8 | FGB | FGB | FGB | 571 | 0.37 | 0.25 | YES |
| 9 | MBL2 | MBL2 | MBL2 | 674 | 0.36 | 0.27 | YES |
| 10 | PLAUR | PLAUR | PLAUR | 710 | 0.35 | 0.29 | YES |
| 11 | C4BPA | C4BPA | C4BPA | 852 | 0.33 | 0.31 | YES |
| 12 | CR1 | CR1 | CR1 | 858 | 0.33 | 0.33 | YES |
| 13 | CFH | CFH | CFH | 917 | 0.32 | 0.35 | YES |
| 14 | BDKRB1 | BDKRB1 | BDKRB1 | 990 | 0.31 | 0.37 | YES |
| 15 | CFB | CFB | CFB | 1001 | 0.31 | 0.39 | YES |
| 16 | C9 | C9 | C9 | 1062 | 0.3 | 0.41 | YES |
| 17 | CFD | CFD | CFD | 1067 | 0.3 | 0.43 | YES |
| 18 | C3 | C3 | C3 | 1118 | 0.29 | 0.45 | YES |
| 19 | SERPINE1 | SERPINE1 | SERPINE1 | 1183 | 0.28 | 0.47 | YES |
| 20 | F3 | F3 | F3 | 1218 | 0.28 | 0.48 | YES |
| 21 | PLAU | PLAU | PLAU | 1321 | 0.27 | 0.5 | YES |
| 22 | CR2 | CR2 | CR2 | 1714 | 0.23 | 0.49 | YES |
| 23 | F12 | F12 | F12 | 1782 | 0.23 | 0.51 | YES |
| 24 | SERPINA5 | SERPINA5 | SERPINA5 | 1817 | 0.22 | 0.52 | YES |
| 25 | C1QB | C1QB | C1QB | 1894 | 0.22 | 0.53 | YES |
| 26 | BDKRB2 | BDKRB2 | BDKRB2 | 1968 | 0.21 | 0.54 | YES |
| 27 | C1QC | C1QC | C1QC | 2119 | 0.2 | 0.55 | YES |
| 28 | C1QA | C1QA | C1QA | 2164 | 0.2 | 0.56 | YES |
| 29 | SERPINF2 | SERPINF2 | SERPINF2 | 2215 | 0.19 | 0.57 | YES |
| 30 | C5AR1 | C5AR1 | C5AR1 | 2240 | 0.19 | 0.59 | YES |
| 31 | SERPINA1 | SERPINA1 | SERPINA1 | 2288 | 0.19 | 0.6 | YES |
| 32 | SERPING1 | SERPING1 | SERPING1 | 2365 | 0.18 | 0.61 | YES |
| 33 | CPB2 | CPB2 | CPB2 | 2414 | 0.18 | 0.62 | YES |
| 34 | C3AR1 | C3AR1 | C3AR1 | 2694 | 0.16 | 0.61 | YES |
| 35 | F13A1 | F13A1 | F13A1 | 2752 | 0.16 | 0.62 | YES |
| 36 | F5 | F5 | F5 | 2782 | 0.16 | 0.63 | YES |
| 37 | CFI | CFI | CFI | 3178 | 0.14 | 0.62 | NO |
| 38 | PROS1 | PROS1 | PROS1 | 4411 | 0.096 | 0.56 | NO |
| 39 | SERPINC1 | SERPINC1 | SERPINC1 | 4879 | 0.083 | 0.54 | NO |
| 40 | C4A | C4A | C4A | 5042 | 0.079 | 0.54 | NO |
| 41 | F10 | F10 | F10 | 6029 | 0.059 | 0.49 | NO |
| 42 | SERPIND1 | SERPIND1 | SERPIND1 | 6447 | 0.051 | 0.47 | NO |
| 43 | CD55 | CD55 | CD55 | 6949 | 0.044 | 0.44 | NO |
| 44 | THBD | THBD | THBD | 7272 | 0.038 | 0.43 | NO |
| 45 | C5 | C5 | C5 | 7278 | 0.038 | 0.43 | NO |
| 46 | F2R | F2R | F2R | 9771 | 0.0043 | 0.3 | NO |
| 47 | TFPI | TFPI | TFPI | 10658 | -0.0067 | 0.25 | NO |
| 48 | KNG1 | KNG1 | KNG1 | 12147 | -0.026 | 0.17 | NO |
| 49 | C7 | C7 | C7 | 12739 | -0.034 | 0.14 | NO |
| 50 | F8 | F8 | F8 | 13945 | -0.054 | 0.075 | NO |
| 51 | CD46 | CD46 | CD46 | 14015 | -0.055 | 0.075 | NO |
| 52 | CD59 | CD59 | CD59 | 14089 | -0.056 | 0.076 | NO |
| 53 | C4BPB | C4BPB | C4BPB | 14533 | -0.066 | 0.056 | NO |
| 54 | PROC | PROC | PROC | 16111 | -0.12 | -0.022 | NO |
| 55 | VWF | VWF | VWF | 16181 | -0.12 | -0.017 | NO |
| 56 | F7 | F7 | F7 | 16320 | -0.13 | -0.016 | NO |
| 57 | KLKB1 | KLKB1 | KLKB1 | 16420 | -0.13 | -0.012 | NO |
| 58 | C6 | C6 | C6 | 16822 | -0.16 | -0.022 | NO |
| 59 | PLAT | PLAT | PLAT | 16936 | -0.17 | -0.016 | NO |
| 60 | MASP2 | MASP2 | MASP2 | 17564 | -0.24 | -0.033 | NO |
| 61 | MASP1 | MASP1 | MASP1 | 17783 | -0.28 | -0.025 | NO |
| 62 | F11 | F11 | F11 | 17821 | -0.28 | -0.0064 | NO |
| 63 | PLG | PLG | PLG | 18121 | -0.42 | 0.0075 | NO |
Figure S57. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG CELL CYCLE.
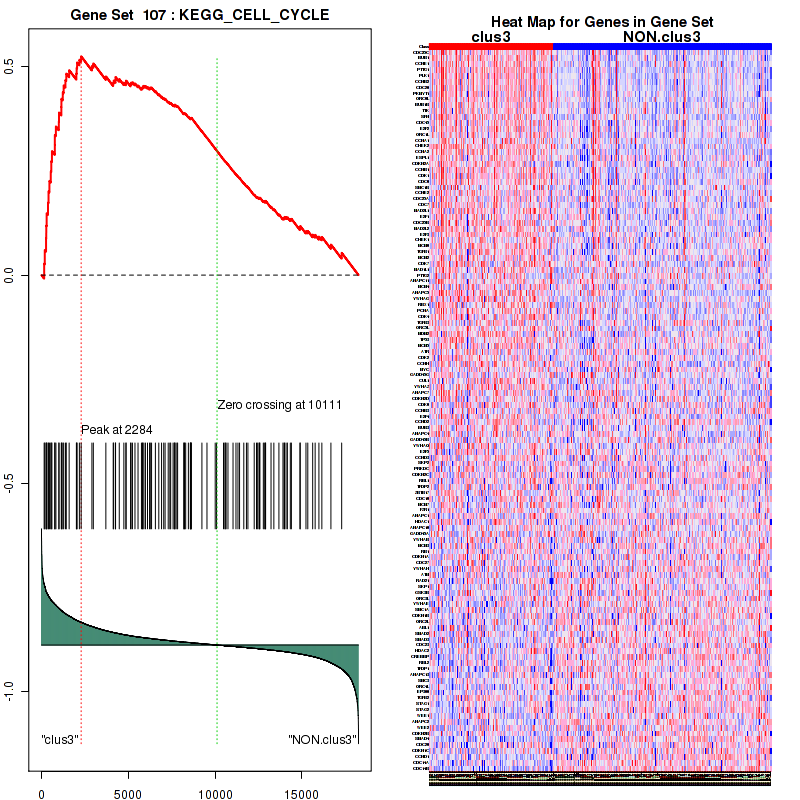
Figure S58. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG CELL CYCLE, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
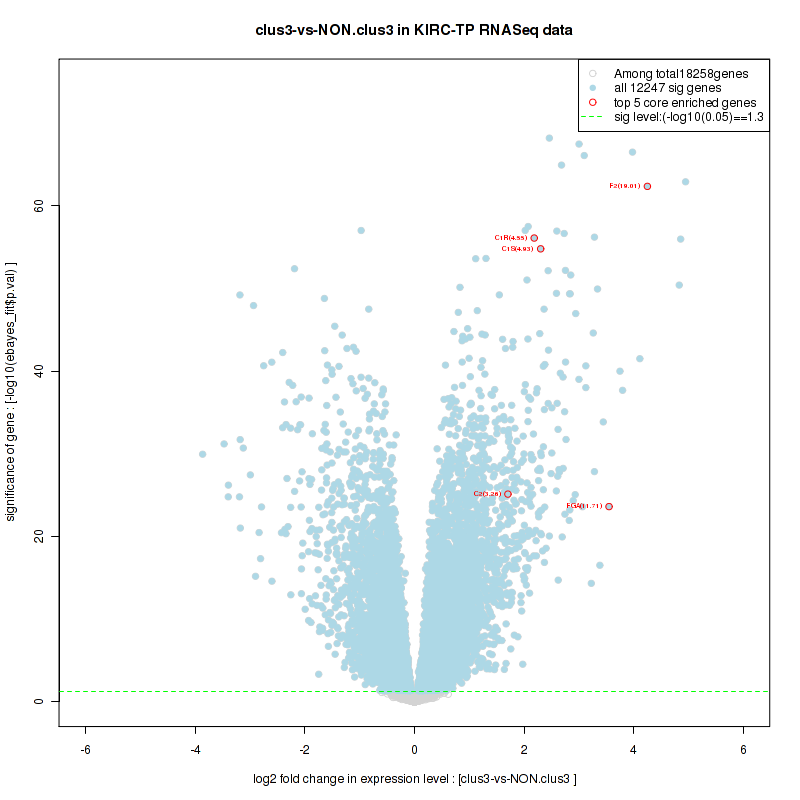
Table S30. Get Full Table This table shows a Running Enrichment Score (RES) of each gene in this pathway, that is, the enrichment score at this point in the ranked list of genes. All genes are ranked by Signal-to-Noise (S2N), a measure of similarity as default and are used to obtain ES matrix of all genes. In this way, GSEA tool uses expression pattern of not only overlapped genes but also not-overlapped genes to produce ES matrix.
| Rank | GENE | SYMBOL | DESC | LIST.LOC | S2N | RES | CORE_ENRICHMENT |
|---|---|---|---|---|---|---|---|
| 1 | EME1 | EME1 | EME1 | 299 | 0.44 | 0.12 | YES |
| 2 | RAD54B | RAD54B | RAD54B | 312 | 0.44 | 0.25 | YES |
| 3 | RAD54L | RAD54L | RAD54L | 423 | 0.4 | 0.36 | YES |
| 4 | BLM | BLM | BLM | 708 | 0.35 | 0.45 | YES |
| 5 | RAD51 | RAD51 | RAD51 | 1115 | 0.29 | 0.52 | YES |
| 6 | XRCC2 | XRCC2 | XRCC2 | 1430 | 0.26 | 0.58 | YES |
| 7 | POLD4 | POLD4 | POLD4 | 2763 | 0.16 | 0.56 | YES |
| 8 | RPA4 | RPA4 | RPA4 | 3298 | 0.14 | 0.57 | YES |
| 9 | SHFM1 | SHFM1 | SHFM1 | 3690 | 0.12 | 0.58 | YES |
| 10 | POLD1 | POLD1 | POLD1 | 3882 | 0.11 | 0.61 | YES |
| 11 | BRCA2 | BRCA2 | BRCA2 | 4277 | 0.1 | 0.62 | YES |
| 12 | POLD3 | POLD3 | POLD3 | 5948 | 0.06 | 0.54 | NO |
| 13 | NBN | NBN | NBN | 6094 | 0.058 | 0.55 | NO |
| 14 | SSBP1 | SSBP1 | SSBP1 | 6110 | 0.057 | 0.57 | NO |
| 15 | RPA3 | RPA3 | RPA3 | 6804 | 0.046 | 0.54 | NO |
| 16 | RAD51L3 | RAD51L3 | RAD51L3 | 6918 | 0.044 | 0.55 | NO |
| 17 | RAD52 | RAD52 | RAD52 | 6953 | 0.044 | 0.56 | NO |
| 18 | POLD2 | POLD2 | POLD2 | 6979 | 0.043 | 0.57 | NO |
| 19 | XRCC3 | XRCC3 | XRCC3 | 7543 | 0.034 | 0.55 | NO |
| 20 | RAD51L1 | RAD51L1 | RAD51L1 | 7800 | 0.03 | 0.55 | NO |
| 21 | TOP3B | TOP3B | TOP3B | 8936 | 0.015 | 0.49 | NO |
| 22 | RPA2 | RPA2 | RPA2 | 9564 | 0.0072 | 0.46 | NO |
| 23 | MUS81 | MUS81 | MUS81 | 10101 | 0.00014 | 0.43 | NO |
| 24 | RPA1 | RPA1 | RPA1 | 10779 | -0.0081 | 0.39 | NO |
| 25 | TOP3A | TOP3A | TOP3A | 11096 | -0.012 | 0.38 | NO |
| 26 | RAD50 | RAD50 | RAD50 | 11583 | -0.018 | 0.36 | NO |
| 27 | MRE11A | MRE11A | MRE11A | 11908 | -0.023 | 0.35 | NO |
Figure S59. Get High-res Image This plot shows mRNAseq_cNMF expression data heatmap (on the left) a RunningEnrichmentScore(RES) plot (on the top right) and a Signal2Noise(S2N) plot (on the bottom right) of genes in the pathway: KEGG OOCYTE MEIOSIS.
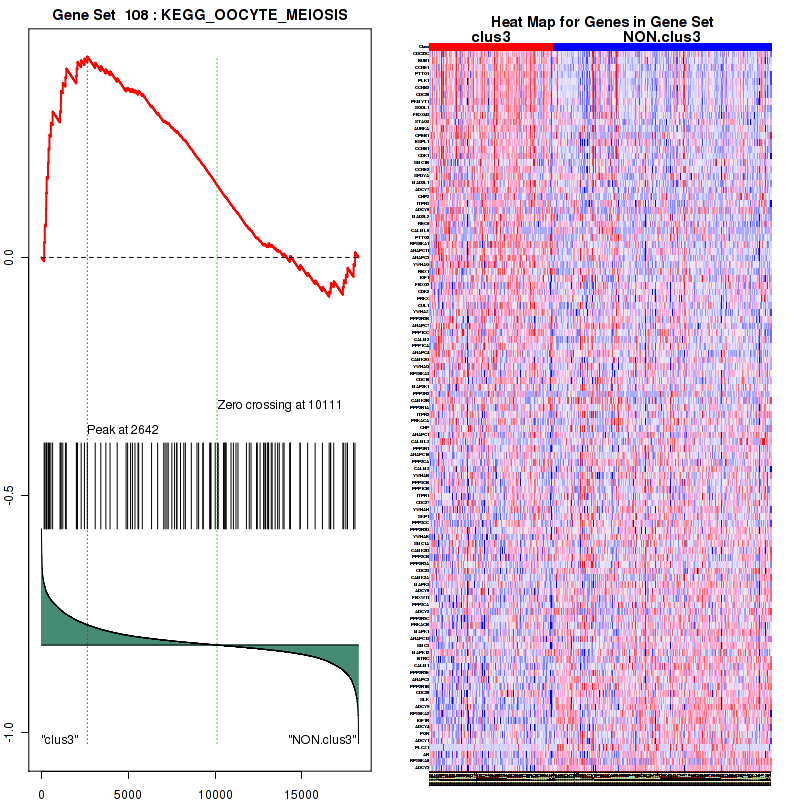
Figure S60. Get High-res Image For the top 5 core enriched genes in the pathway: KEGG OOCYTE MEIOSIS, this volcano plot shows how much they are up/down-regulated and significant. The significance was calculated by empirical bayesian fit
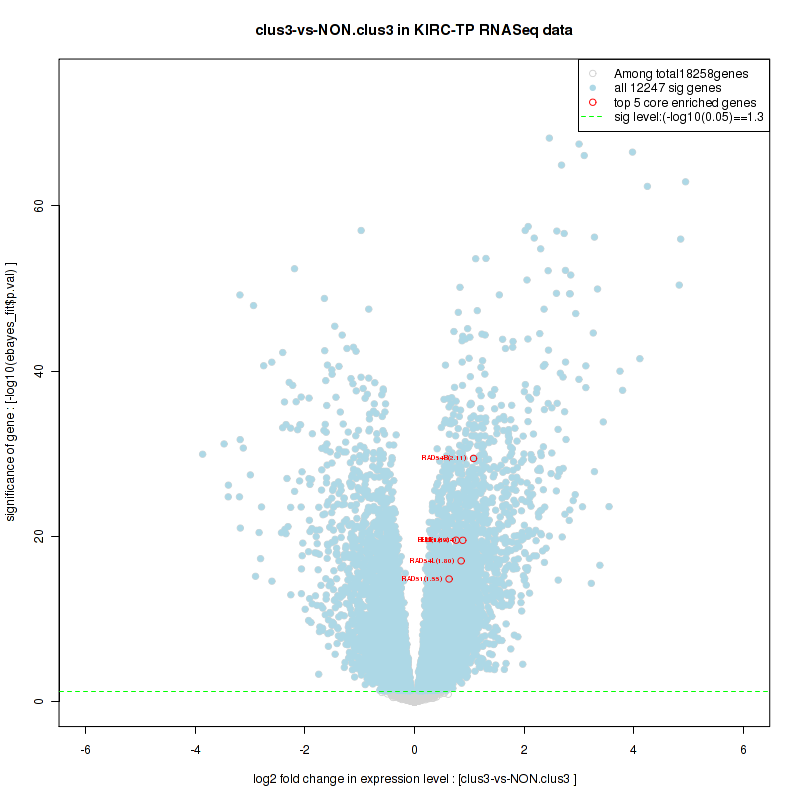
For the top enriched genes, if you want to check whether they are
-
up-regulated, please check the list of up-regulated genes
-
down-regulated, please check the list of down-regulated genes
For the top enriched genes, if you want to check whether they are
-
highly expressed genes, please check the list of high (top 30%) expressed genes
-
low expressed genes, please check the list of low (bottom 30%) expressed genes
An expression pattern of top(30%)/middle(30%)/low(30%) in this subtype against other subtypes is available in a heatmap
For the top enriched genes, if you want to check whether they are
-
significantly differently expressed genes by eBayes lm fit, please check the list of significant genes
-
Gene set database = c2.cp.v3.0-2.symbols.gmt
-
Expression data file = KIRC-TP.uncv2.mRNAseq_RSEM_normalized_log2.txt
-
Phenotype data file = KIRC-TP.mergedcluster.txt
For the Gene Set Enrichment Analysis (GSEA), Broad GSEA-P-R.1.0 version is used with class2: canonical pathways geneses from MSigDB. Further details about statistics are available inThe Broad GSEA website.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.