This pipeline computes the correlation between significantly recurrent gene mutations and selected clinical features.
Testing the association between mutation status of 84 genes and 8 clinical features across 512 patients, 29 significant findings detected with Q value < 0.25.
-
IDH1 mutation correlated to 'Time to Death', 'YEARS_TO_BIRTH', 'RADIATION_THERAPY', 'KARNOFSKY_PERFORMANCE_SCORE', and 'HISTOLOGICAL_TYPE'.
-
TP53 mutation correlated to 'YEARS_TO_BIRTH', 'RADIATION_THERAPY', and 'HISTOLOGICAL_TYPE'.
-
ATRX mutation correlated to 'YEARS_TO_BIRTH' and 'HISTOLOGICAL_TYPE'.
-
CIC mutation correlated to 'Time to Death', 'RADIATION_THERAPY', and 'HISTOLOGICAL_TYPE'.
-
NOTCH1 mutation correlated to 'HISTOLOGICAL_TYPE'.
-
IDH2 mutation correlated to 'HISTOLOGICAL_TYPE'.
-
FUBP1 mutation correlated to 'YEARS_TO_BIRTH' and 'HISTOLOGICAL_TYPE'.
-
NF1 mutation correlated to 'Time to Death'.
-
PTEN mutation correlated to 'Time to Death', 'YEARS_TO_BIRTH', and 'HISTOLOGICAL_TYPE'.
-
EGFR mutation correlated to 'Time to Death', 'YEARS_TO_BIRTH', and 'KARNOFSKY_PERFORMANCE_SCORE'.
-
SMARCA4 mutation correlated to 'Time to Death'.
-
SETD2 mutation correlated to 'YEARS_TO_BIRTH'.
-
HTR3A mutation correlated to 'RACE'.
-
WRN mutation correlated to 'ETHNICITY'.
-
TMEM184A mutation correlated to 'Time to Death'.
Table 1. Get Full Table Overview of the association between mutation status of 84 genes and 8 clinical features. Shown in the table are P values (Q values). Thresholded by Q value < 0.25, 29 significant findings detected.
|
Clinical Features |
Time to Death |
YEARS TO BIRTH |
GENDER |
RADIATION THERAPY |
KARNOFSKY PERFORMANCE SCORE |
HISTOLOGICAL TYPE |
RACE | ETHNICITY | ||
| nMutated (%) | nWild-Type | logrank test | Wilcoxon-test | Fisher's exact test | Fisher's exact test | Wilcoxon-test | Fisher's exact test | Fisher's exact test | Fisher's exact test | |
| IDH1 | 397 (78%) | 115 |
1.11e-16 (7.46e-14) |
9.38e-09 (1.05e-06) |
1 (1.00) |
0.00895 (0.215) |
0.00252 (0.0769) |
0.00513 (0.138) |
0.0253 (0.437) |
1 (1.00) |
| TP53 | 247 (48%) | 265 |
0.054 (0.637) |
2.47e-12 (5.53e-10) |
0.286 (0.927) |
0.000168 (0.00708) |
0.279 (0.922) |
1e-05 (0.000611) |
0.0348 (0.557) |
0.715 (1.00) |
| CIC | 108 (21%) | 404 |
0.000136 (0.00608) |
0.107 (0.72) |
0.328 (0.93) |
3.46e-06 (0.000332) |
0.24 (0.888) |
1e-05 (0.000611) |
0.751 (1.00) |
0.656 (1.00) |
| PTEN | 24 (5%) | 488 |
0.00013 (0.00608) |
0.00176 (0.0623) |
0.297 (0.93) |
0.0671 (0.659) |
0.585 (1.00) |
0.00245 (0.0769) |
0.163 (0.787) |
1 (1.00) |
| EGFR | 35 (7%) | 477 |
1.23e-11 (2.07e-09) |
4.84e-11 (6.5e-09) |
1 (1.00) |
0.0148 (0.317) |
0.00495 (0.138) |
0.0484 (0.608) |
0.181 (0.81) |
0.718 (1.00) |
| ATRX | 194 (38%) | 318 |
0.0886 (0.659) |
1.97e-12 (5.53e-10) |
0.784 (1.00) |
0.0156 (0.317) |
0.797 (1.00) |
1e-05 (0.000611) |
0.436 (0.99) |
1 (1.00) |
| FUBP1 | 47 (9%) | 465 |
0.13 (0.751) |
0.00301 (0.0881) |
0.645 (1.00) |
0.0695 (0.659) |
0.236 (0.888) |
1e-05 (0.000611) |
1 (1.00) |
0.1 (0.702) |
| NOTCH1 | 42 (8%) | 470 |
0.12 (0.722) |
0.0424 (0.574) |
0.747 (1.00) |
0.392 (0.952) |
0.869 (1.00) |
5e-05 (0.0028) |
0.336 (0.931) |
1 (1.00) |
| IDH2 | 20 (4%) | 492 |
0.0594 (0.655) |
0.074 (0.659) |
1 (1.00) |
0.477 (1.00) |
0.621 (1.00) |
0.00065 (0.0243) |
0.45 (0.999) |
0.127 (0.744) |
| NF1 | 33 (6%) | 479 |
9.95e-05 (0.00514) |
0.0125 (0.28) |
0.59 (1.00) |
0.454 (1.00) |
0.549 (1.00) |
0.0155 (0.317) |
0.221 (0.883) |
0.258 (0.888) |
| SMARCA4 | 26 (5%) | 486 |
0.00715 (0.185) |
0.523 (1.00) |
0.318 (0.93) |
1 (1.00) |
0.432 (0.984) |
0.79 (1.00) |
0.115 (0.72) |
0.644 (1.00) |
| SETD2 | 10 (2%) | 502 |
0.56 (1.00) |
0.00794 (0.198) |
1 (1.00) |
0.747 (1.00) |
0.308 (0.93) |
0.362 (0.944) |
0.466 (1.00) |
1 (1.00) |
| HTR3A | 7 (1%) | 505 |
0.403 (0.952) |
0.54 (1.00) |
0.0489 (0.608) |
0.439 (0.99) |
0.0794 (0.659) |
0.9 (1.00) |
0.00927 (0.215) |
1 (1.00) |
| WRN | 5 (1%) | 507 |
0.0842 (0.659) |
0.381 (0.948) |
0.661 (1.00) |
1 (1.00) |
0.0379 (0.559) |
0.552 (1.00) |
0.266 (0.899) |
0.0025 (0.0769) |
| TMEM184A | 3 (1%) | 509 |
0.000525 (0.0207) |
0.151 (0.787) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.17 (0.787) |
1 (1.00) |
|
| PIK3R1 | 22 (4%) | 490 |
0.259 (0.888) |
0.0799 (0.659) |
0.275 (0.913) |
0.487 (1.00) |
0.522 (1.00) |
0.354 (0.931) |
0.0455 (0.598) |
0.623 (1.00) |
| ARID1A | 20 (4%) | 492 |
0.0379 (0.559) |
0.648 (1.00) |
0.493 (1.00) |
0.351 (0.931) |
0.964 (1.00) |
0.195 (0.836) |
1 (1.00) |
1 (1.00) |
| GAGE2A | 8 (2%) | 504 |
0.507 (1.00) |
0.733 (1.00) |
0.737 (1.00) |
1 (1.00) |
0.323 (0.93) |
1 (1.00) |
0.054 (0.637) |
0.428 (0.978) |
| NUDT11 | 11 (2%) | 501 |
0.525 (1.00) |
0.321 (0.93) |
0.36 (0.941) |
1 (1.00) |
0.641 (1.00) |
0.745 (1.00) |
1 (1.00) |
1 (1.00) |
| PIK3CA | 44 (9%) | 468 |
0.65 (1.00) |
0.0729 (0.659) |
0.875 (1.00) |
0.256 (0.888) |
0.102 (0.705) |
0.0182 (0.34) |
0.628 (1.00) |
0.743 (1.00) |
| STK19 | 10 (2%) | 502 |
0.307 (0.93) |
0.311 (0.93) |
0.12 (0.722) |
0.682 (1.00) |
0.23 (0.888) |
0.0754 (0.659) |
1 (1.00) |
0.14 (0.769) |
| EMG1 | 6 (1%) | 506 |
0.537 (1.00) |
0.0981 (0.694) |
0.415 (0.952) |
1 (1.00) |
0.366 (0.948) |
0.351 (0.931) |
1 (1.00) |
1 (1.00) |
| NIPBL | 18 (4%) | 494 |
0.0578 (0.654) |
0.176 (0.803) |
0.156 (0.787) |
0.311 (0.93) |
0.604 (1.00) |
0.0697 (0.659) |
0.0756 (0.659) |
0.112 (0.72) |
| TCF12 | 15 (3%) | 497 |
0.625 (1.00) |
0.225 (0.888) |
0.601 (1.00) |
1 (1.00) |
0.915 (1.00) |
0.895 (1.00) |
1 (1.00) |
0.613 (1.00) |
| DNMT3A | 10 (2%) | 502 |
0.0364 (0.559) |
0.479 (1.00) |
1 (1.00) |
0.747 (1.00) |
0.992 (1.00) |
0.337 (0.931) |
0.462 (1.00) |
0.503 (1.00) |
| TRERF1 | 6 (1%) | 506 |
0.482 (1.00) |
0.285 (0.927) |
0.0941 (0.672) |
0.682 (1.00) |
0.798 (1.00) |
0.167 (0.787) |
1 (1.00) |
1 (1.00) |
| CREBZF | 7 (1%) | 505 |
0.955 (1.00) |
0.171 (0.787) |
0.251 (0.888) |
1 (1.00) |
0.0699 (0.659) |
0.0314 (0.515) |
0.35 (0.931) |
0.386 (0.948) |
| FAM47C | 18 (4%) | 494 |
0.0775 (0.659) |
0.562 (1.00) |
0.47 (1.00) |
0.46 (1.00) |
0.649 (1.00) |
0.595 (1.00) |
1 (1.00) |
1 (1.00) |
| MYST4 | 11 (2%) | 501 |
0.0922 (0.666) |
0.108 (0.72) |
1 (1.00) |
1 (1.00) |
0.928 (1.00) |
0.189 (0.831) |
0.147 (0.787) |
0.503 (1.00) |
| MED9 | 3 (1%) | 509 |
0.23 (0.888) |
0.871 (1.00) |
0.257 (0.888) |
0.285 (0.927) |
0.0162 (0.32) |
1 (1.00) |
1 (1.00) |
|
| IRS4 | 7 (1%) | 505 |
0.16 (0.787) |
0.495 (1.00) |
1 (1.00) |
1 (1.00) |
0.664 (1.00) |
0.137 (0.769) |
1 (1.00) |
0.386 (0.948) |
| HTRA2 | 5 (1%) | 507 |
0.339 (0.931) |
0.0279 (0.469) |
1 (1.00) |
0.39 (0.952) |
0.451 (0.999) |
1 (1.00) |
0.294 (0.927) |
|
| NRAS | 4 (1%) | 508 |
0.28 (0.922) |
0.883 (1.00) |
1 (1.00) |
0.645 (1.00) |
0.471 (1.00) |
0.023 (0.406) |
1 (1.00) |
1 (1.00) |
| TNRC18 | 9 (2%) | 503 |
0.321 (0.93) |
0.948 (1.00) |
0.198 (0.841) |
0.493 (1.00) |
0.915 (1.00) |
0.151 (0.787) |
1 (1.00) |
1 (1.00) |
| ZNF709 | 4 (1%) | 508 |
0.0842 (0.659) |
0.595 (1.00) |
0.132 (0.751) |
1 (1.00) |
0.139 (0.769) |
0.22 (0.883) |
0.243 (0.888) |
|
| PLXNA3 | 9 (2%) | 503 |
0.0907 (0.663) |
0.249 (0.888) |
0.311 (0.93) |
0.158 (0.787) |
0.588 (1.00) |
0.834 (1.00) |
0.109 (0.72) |
1 (1.00) |
| ROBO3 | 5 (1%) | 507 |
0.49 (1.00) |
0.533 (1.00) |
0.661 (1.00) |
0.645 (1.00) |
0.664 (1.00) |
0.378 (0.948) |
1 (1.00) |
0.294 (0.927) |
| RB1 | 6 (1%) | 506 |
0.829 (1.00) |
0.165 (0.787) |
0.415 (0.952) |
1 (1.00) |
0.0807 (0.659) |
0.315 (0.93) |
1 (1.00) |
|
| SRPX | 4 (1%) | 508 |
0.151 (0.787) |
0.0427 (0.574) |
0.0394 (0.561) |
1 (1.00) |
0.555 (1.00) |
0.377 (0.948) |
0.22 (0.883) |
1 (1.00) |
| PDGFRA | 10 (2%) | 502 |
0.0741 (0.659) |
0.707 (1.00) |
0.354 (0.931) |
0.747 (1.00) |
0.142 (0.775) |
0.115 (0.72) |
1 (1.00) |
0.14 (0.769) |
| ZNF512B | 5 (1%) | 507 |
0.08 (0.659) |
0.301 (0.93) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.294 (0.927) |
|
| ZMIZ1 | 9 (2%) | 503 |
0.0712 (0.659) |
0.862 (1.00) |
0.522 (1.00) |
1 (1.00) |
0.149 (0.787) |
0.44 (0.99) |
0.0661 (0.659) |
0.0174 (0.334) |
| TMEM216 | 3 (1%) | 509 |
0.771 (1.00) |
0.817 (1.00) |
0.0888 (0.659) |
1 (1.00) |
0.367 (0.948) |
1 (1.00) |
1 (1.00) |
|
| ARID2 | 11 (2%) | 501 |
0.51 (1.00) |
0.109 (0.72) |
1 (1.00) |
0.215 (0.883) |
0.807 (1.00) |
0.206 (0.864) |
0.244 (0.888) |
1 (1.00) |
| RBPJ | 7 (1%) | 505 |
0.873 (1.00) |
0.0538 (0.637) |
1 (1.00) |
0.682 (1.00) |
0.335 (0.931) |
0.274 (0.913) |
1 (1.00) |
1 (1.00) |
| CUL4B | 10 (2%) | 502 |
0.211 (0.882) |
0.824 (1.00) |
0.12 (0.722) |
0.329 (0.93) |
0.641 (1.00) |
0.335 (0.931) |
0.219 (0.883) |
1 (1.00) |
| SLC6A3 | 9 (2%) | 503 |
0.642 (1.00) |
0.377 (0.948) |
0.522 (1.00) |
1 (1.00) |
0.761 (1.00) |
1 (1.00) |
0.386 (0.948) |
|
| KRT3 | 4 (1%) | 508 |
0.0883 (0.659) |
0.856 (1.00) |
0.632 (1.00) |
1 (1.00) |
0.105 (0.72) |
0.684 (1.00) |
1 (1.00) |
0.243 (0.888) |
| MYT1 | 6 (1%) | 506 |
0.268 (0.899) |
0.408 (0.952) |
1 (1.00) |
0.0861 (0.659) |
0.734 (1.00) |
0.17 (0.787) |
1 (1.00) |
1 (1.00) |
| SMOC1 | 3 (1%) | 509 |
0.489 (1.00) |
0.443 (0.993) |
1 (1.00) |
0.523 (1.00) |
0.978 (1.00) |
0.475 (1.00) |
1 (1.00) |
1 (1.00) |
| ZBTB20 | 21 (4%) | 491 |
0.229 (0.888) |
0.495 (1.00) |
0.371 (0.948) |
0.161 (0.787) |
0.798 (1.00) |
0.182 (0.81) |
0.127 (0.744) |
0.636 (1.00) |
| PPL | 6 (1%) | 506 |
0.0401 (0.561) |
0.727 (1.00) |
1 (1.00) |
1 (1.00) |
0.798 (1.00) |
0.401 (0.952) |
1 (1.00) |
0.294 (0.927) |
| PTPN11 | 7 (1%) | 505 |
0.253 (0.888) |
0.342 (0.931) |
1 (1.00) |
1 (1.00) |
0.755 (1.00) |
0.309 (0.93) |
0.354 (0.931) |
1 (1.00) |
| RET | 7 (1%) | 505 |
0.651 (1.00) |
0.5 (1.00) |
0.251 (0.888) |
0.439 (0.99) |
0.6 (1.00) |
0.0724 (0.659) |
1 (1.00) |
1 (1.00) |
| ANKRD36 | 7 (1%) | 505 |
0.0892 (0.659) |
0.593 (1.00) |
0.706 (1.00) |
0.0463 (0.598) |
0.0699 (0.659) |
0.237 (0.888) |
0.0698 (0.659) |
1 (1.00) |
| NEU2 | 5 (1%) | 507 |
0.227 (0.888) |
0.168 (0.787) |
1 (1.00) |
1 (1.00) |
0.38 (0.948) |
1 (1.00) |
0.294 (0.927) |
|
| PRX | 10 (2%) | 502 |
0.761 (1.00) |
0.463 (1.00) |
0.354 (0.931) |
0.329 (0.93) |
0.33 (0.93) |
0.781 (1.00) |
1 (1.00) |
0.503 (1.00) |
| RBBP6 | 6 (1%) | 506 |
0.195 (0.836) |
0.321 (0.93) |
1 (1.00) |
0.682 (1.00) |
0.773 (1.00) |
0.125 (0.74) |
1 (1.00) |
|
| CIB1 | 4 (1%) | 508 |
0.407 (0.952) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.755 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
| DDX5 | 5 (1%) | 507 |
0.114 (0.72) |
0.854 (1.00) |
1 (1.00) |
1 (1.00) |
0.289 (0.927) |
0.852 (1.00) |
1 (1.00) |
0.243 (0.888) |
| NAP1L2 | 4 (1%) | 508 |
0.267 (0.899) |
0.178 (0.803) |
1 (1.00) |
0.645 (1.00) |
0.195 (0.836) |
1 (1.00) |
1 (1.00) |
|
| SCN4A | 6 (1%) | 506 |
0.187 (0.827) |
0.26 (0.888) |
0.415 (0.952) |
0.682 (1.00) |
0.333 (0.931) |
0.775 (1.00) |
1 (1.00) |
1 (1.00) |
| AGBL1 | 5 (1%) | 507 |
0.254 (0.888) |
0.203 (0.856) |
0.386 (0.948) |
1 (1.00) |
0.192 (0.836) |
1 (1.00) |
1 (1.00) |
|
| TPX2 | 5 (1%) | 507 |
0.233 (0.888) |
0.983 (1.00) |
0.661 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.0383 (0.559) |
|
| KRT15 | 6 (1%) | 506 |
0.751 (1.00) |
0.116 (0.72) |
0.696 (1.00) |
0.215 (0.883) |
0.41 (0.952) |
0.886 (1.00) |
0.313 (0.93) |
1 (1.00) |
| C14ORF4 | 3 (1%) | 509 |
0.467 (1.00) |
0.739 (1.00) |
0.589 (1.00) |
1 (1.00) |
0.786 (1.00) |
0.17 (0.787) |
1 (1.00) |
|
| R3HDM1 | 7 (1%) | 505 |
0.0584 (0.654) |
0.872 (1.00) |
1 (1.00) |
1 (1.00) |
0.755 (1.00) |
0.494 (1.00) |
1 (1.00) |
0.386 (0.948) |
| G6PC | 6 (1%) | 506 |
0.878 (1.00) |
0.401 (0.952) |
0.696 (1.00) |
0.412 (0.952) |
0.51 (1.00) |
0.403 (0.952) |
1 (1.00) |
0.055 (0.638) |
| DLC1 | 6 (1%) | 506 |
0.561 (1.00) |
0.553 (1.00) |
0.415 (0.952) |
0.0861 (0.659) |
0.666 (1.00) |
1 (1.00) |
0.342 (0.931) |
|
| SLC12A7 | 7 (1%) | 505 |
0.178 (0.803) |
0.54 (1.00) |
0.706 (1.00) |
0.412 (0.952) |
0.52 (1.00) |
1 (1.00) |
0.35 (0.931) |
1 (1.00) |
| SMARCB1 | 4 (1%) | 508 |
0.312 (0.93) |
0.811 (1.00) |
0.632 (1.00) |
0.16 (0.787) |
0.755 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
| LAMA4 | 4 (1%) | 508 |
0.796 (1.00) |
0.354 (0.931) |
0.132 (0.751) |
1 (1.00) |
0.755 (1.00) |
0.0667 (0.659) |
0.0795 (0.659) |
0.243 (0.888) |
| MAST2 | 5 (1%) | 507 |
0.472 (1.00) |
0.705 (1.00) |
0.661 (1.00) |
0.162 (0.787) |
0.45 (0.999) |
1 (1.00) |
1 (1.00) |
|
| ZMYM2 | 7 (1%) | 505 |
0.153 (0.787) |
0.7 (1.00) |
1 (1.00) |
0.255 (0.888) |
0.664 (1.00) |
1 (1.00) |
1 (1.00) |
0.386 (0.948) |
| SLFN11 | 4 (1%) | 508 |
0.114 (0.72) |
0.0862 (0.659) |
0.632 (1.00) |
0.645 (1.00) |
0.567 (1.00) |
1 (1.00) |
0.243 (0.888) |
|
| C4BPA | 6 (1%) | 506 |
0.528 (1.00) |
0.64 (1.00) |
0.415 (0.952) |
0.412 (0.952) |
0.166 (0.787) |
0.885 (1.00) |
0.307 (0.93) |
0.342 (0.931) |
| KTELC1 | 5 (1%) | 507 |
0.312 (0.93) |
0.66 (1.00) |
0.386 (0.948) |
0.653 (1.00) |
0.852 (1.00) |
1 (1.00) |
1 (1.00) |
|
| ASXL2 | 3 (1%) | 509 |
0.12 (0.722) |
0.413 (0.952) |
0.589 (1.00) |
1 (1.00) |
0.627 (1.00) |
1 (1.00) |
1 (1.00) |
|
| CCDC135 | 7 (1%) | 505 |
0.236 (0.888) |
0.757 (1.00) |
0.251 (0.888) |
1 (1.00) |
0.706 (1.00) |
1 (1.00) |
0.386 (0.948) |
|
| SLC25A5 | 4 (1%) | 508 |
0.0229 (0.406) |
0.579 (1.00) |
0.329 (0.93) |
1 (1.00) |
0.25 (0.888) |
0.837 (1.00) |
0.221 (0.883) |
1 (1.00) |
| ABCA7 | 9 (2%) | 503 |
0.752 (1.00) |
0.163 (0.787) |
0.738 (1.00) |
1 (1.00) |
0.559 (1.00) |
0.534 (1.00) |
1 (1.00) |
1 (1.00) |
| NKD2 | 4 (1%) | 508 |
0.989 (1.00) |
0.687 (1.00) |
1 (1.00) |
0.304 (0.93) |
0.456 (1.00) |
1 (1.00) |
1 (1.00) |
|
| MAX | 4 (1%) | 508 |
0.801 (1.00) |
0.166 (0.787) |
0.329 (0.93) |
0.563 (1.00) |
0.978 (1.00) |
0.838 (1.00) |
1 (1.00) |
1 (1.00) |
| VSIG4 | 6 (1%) | 506 |
0.915 (1.00) |
0.946 (1.00) |
0.415 (0.952) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
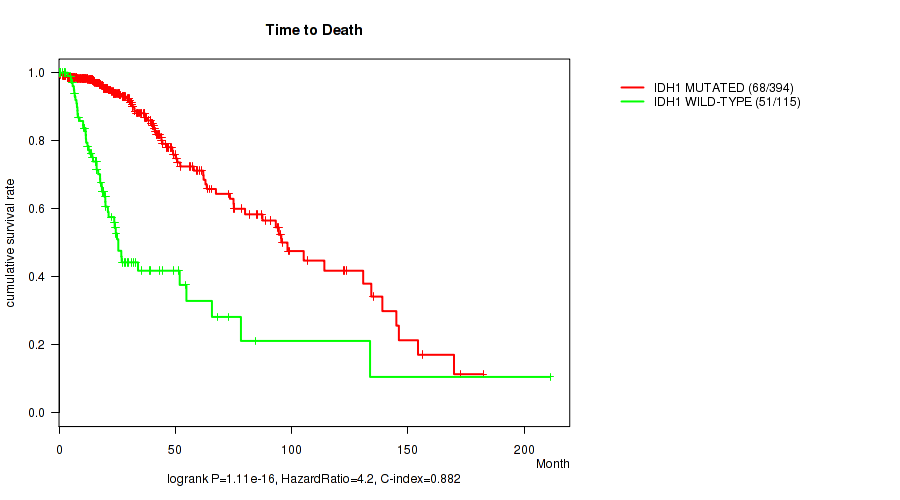
P value = 1.11e-16 (logrank test), Q value = 7.5e-14
Table S1. Gene #1: 'IDH1 MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 509 | 119 | 0.0 - 211.2 (20.7) |
| IDH1 MUTATED | 394 | 68 | 0.0 - 182.3 (23.3) |
| IDH1 WILD-TYPE | 115 | 51 | 0.1 - 211.2 (16.2) |
Figure S1. Get High-res Image Gene #1: 'IDH1 MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
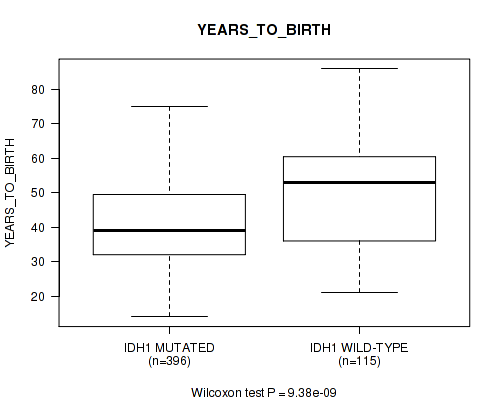
P value = 9.38e-09 (Wilcoxon-test), Q value = 1.1e-06
Table S2. Gene #1: 'IDH1 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 511 | 42.9 (13.4) |
| IDH1 MUTATED | 396 | 41.0 (12.4) |
| IDH1 WILD-TYPE | 115 | 49.6 (14.6) |
Figure S2. Get High-res Image Gene #1: 'IDH1 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
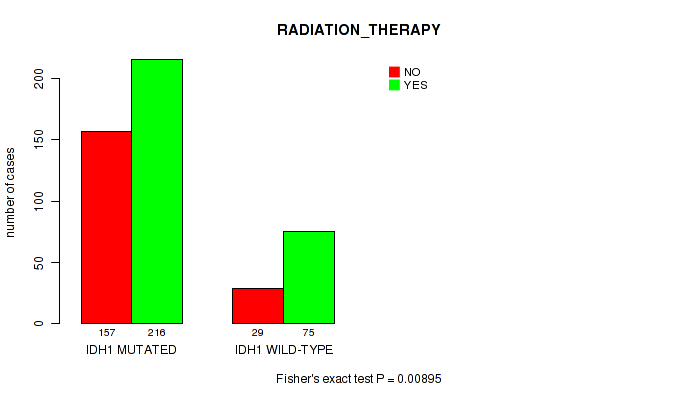
P value = 0.00895 (Fisher's exact test), Q value = 0.21
Table S3. Gene #1: 'IDH1 MUTATION STATUS' versus Clinical Feature #4: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 186 | 291 |
| IDH1 MUTATED | 157 | 216 |
| IDH1 WILD-TYPE | 29 | 75 |
Figure S3. Get High-res Image Gene #1: 'IDH1 MUTATION STATUS' versus Clinical Feature #4: 'RADIATION_THERAPY'
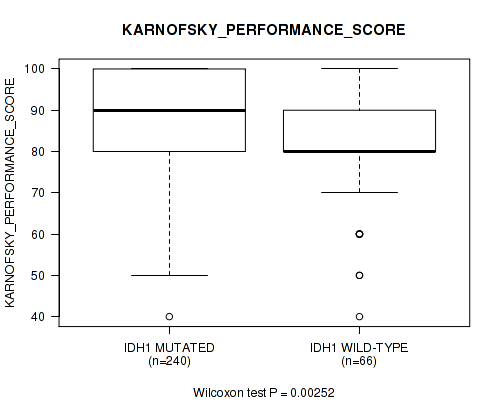
P value = 0.00252 (Wilcoxon-test), Q value = 0.077
Table S4. Gene #1: 'IDH1 MUTATION STATUS' versus Clinical Feature #5: 'KARNOFSKY_PERFORMANCE_SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 306 | 86.7 (12.4) |
| IDH1 MUTATED | 240 | 87.8 (11.8) |
| IDH1 WILD-TYPE | 66 | 82.7 (13.8) |
Figure S4. Get High-res Image Gene #1: 'IDH1 MUTATION STATUS' versus Clinical Feature #5: 'KARNOFSKY_PERFORMANCE_SCORE'
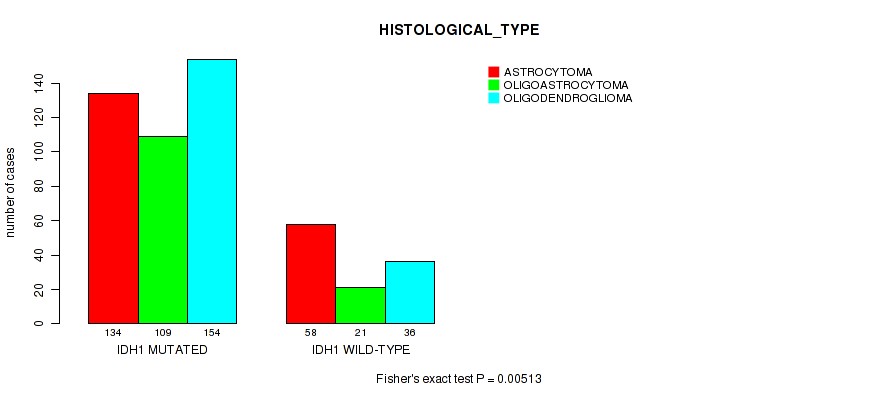
P value = 0.00513 (Fisher's exact test), Q value = 0.14
Table S5. Gene #1: 'IDH1 MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 192 | 130 | 190 |
| IDH1 MUTATED | 134 | 109 | 154 |
| IDH1 WILD-TYPE | 58 | 21 | 36 |
Figure S5. Get High-res Image Gene #1: 'IDH1 MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
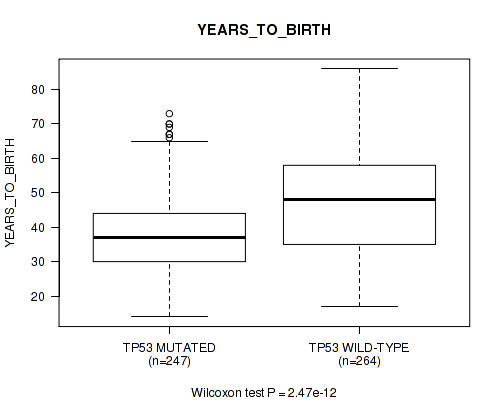
P value = 2.47e-12 (Wilcoxon-test), Q value = 5.5e-10
Table S6. Gene #2: 'TP53 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 511 | 42.9 (13.4) |
| TP53 MUTATED | 247 | 38.6 (11.3) |
| TP53 WILD-TYPE | 264 | 47.0 (13.9) |
Figure S6. Get High-res Image Gene #2: 'TP53 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
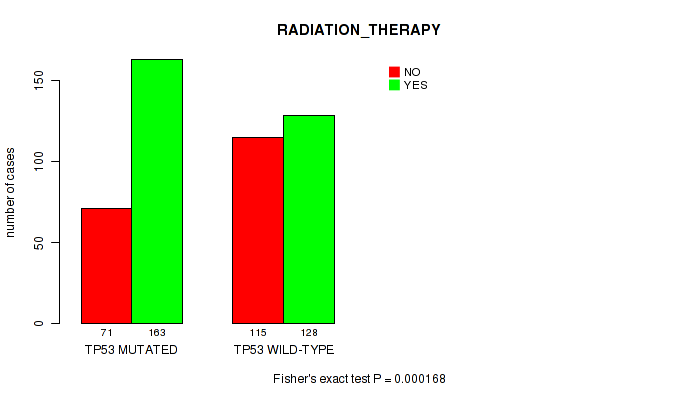
P value = 0.000168 (Fisher's exact test), Q value = 0.0071
Table S7. Gene #2: 'TP53 MUTATION STATUS' versus Clinical Feature #4: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 186 | 291 |
| TP53 MUTATED | 71 | 163 |
| TP53 WILD-TYPE | 115 | 128 |
Figure S7. Get High-res Image Gene #2: 'TP53 MUTATION STATUS' versus Clinical Feature #4: 'RADIATION_THERAPY'
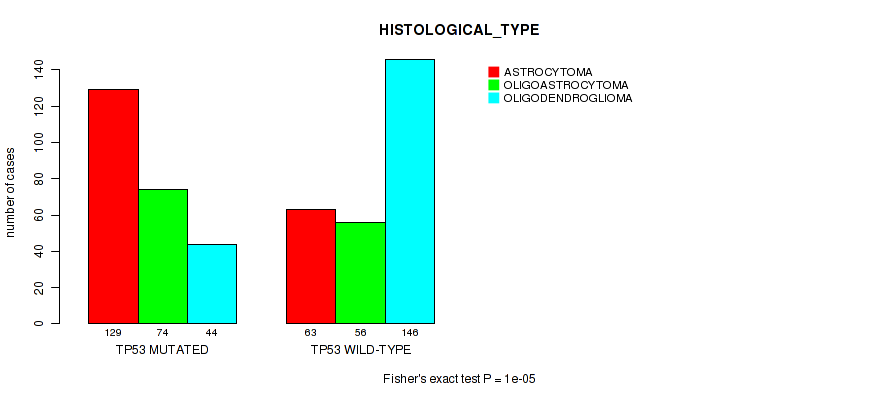
P value = 1e-05 (Fisher's exact test), Q value = 0.00061
Table S8. Gene #2: 'TP53 MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 192 | 130 | 190 |
| TP53 MUTATED | 129 | 74 | 44 |
| TP53 WILD-TYPE | 63 | 56 | 146 |
Figure S8. Get High-res Image Gene #2: 'TP53 MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
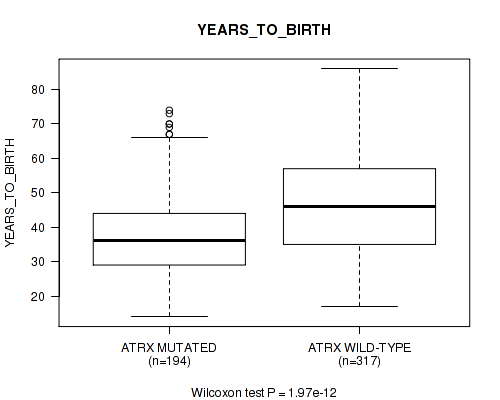
P value = 1.97e-12 (Wilcoxon-test), Q value = 5.5e-10
Table S9. Gene #3: 'ATRX MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 511 | 42.9 (13.4) |
| ATRX MUTATED | 194 | 37.8 (11.9) |
| ATRX WILD-TYPE | 317 | 46.1 (13.3) |
Figure S9. Get High-res Image Gene #3: 'ATRX MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
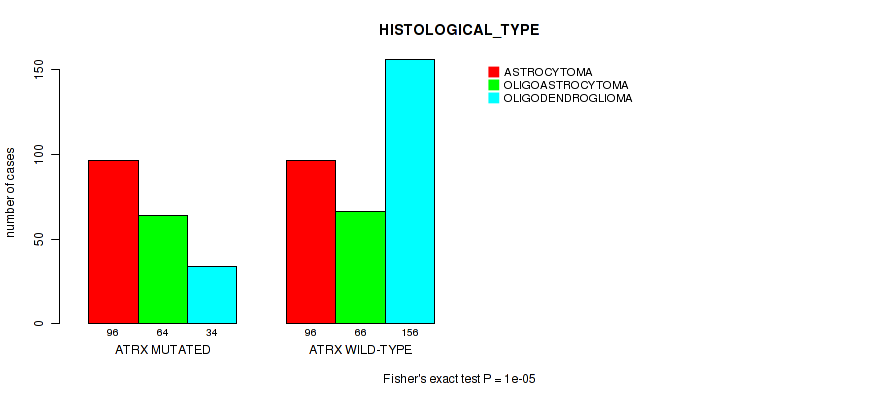
P value = 1e-05 (Fisher's exact test), Q value = 0.00061
Table S10. Gene #3: 'ATRX MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 192 | 130 | 190 |
| ATRX MUTATED | 96 | 64 | 34 |
| ATRX WILD-TYPE | 96 | 66 | 156 |
Figure S10. Get High-res Image Gene #3: 'ATRX MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
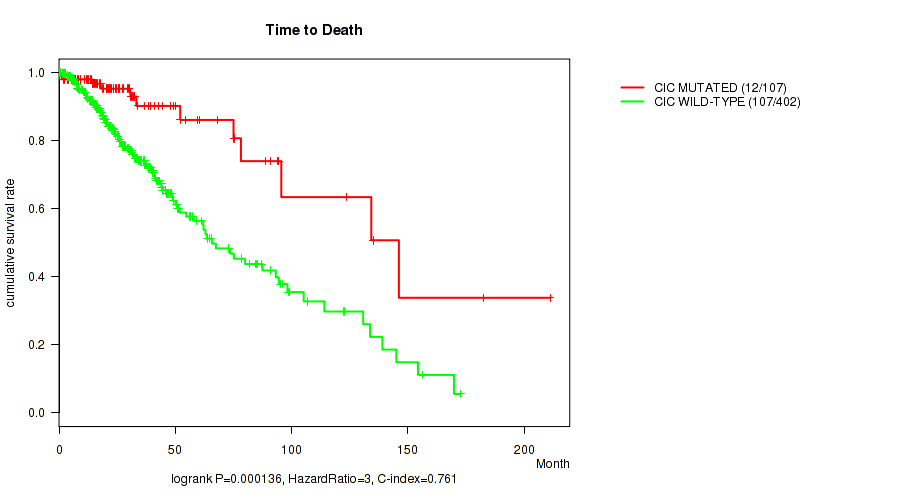
P value = 0.000136 (logrank test), Q value = 0.0061
Table S11. Gene #4: 'CIC MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 509 | 119 | 0.0 - 211.2 (20.7) |
| CIC MUTATED | 107 | 12 | 0.1 - 211.2 (23.2) |
| CIC WILD-TYPE | 402 | 107 | 0.0 - 172.8 (20.1) |
Figure S11. Get High-res Image Gene #4: 'CIC MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
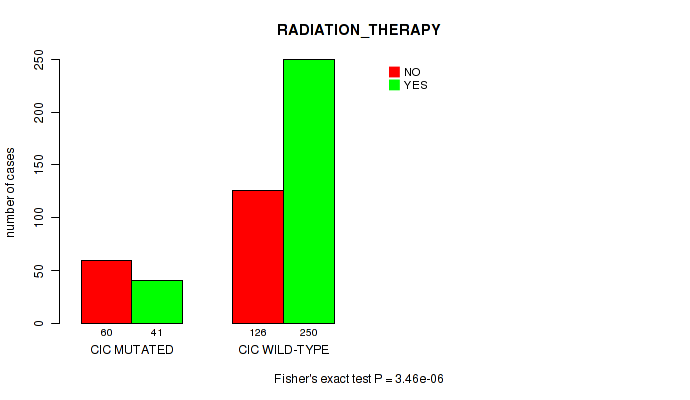
P value = 3.46e-06 (Fisher's exact test), Q value = 0.00033
Table S12. Gene #4: 'CIC MUTATION STATUS' versus Clinical Feature #4: 'RADIATION_THERAPY'
| nPatients | NO | YES |
|---|---|---|
| ALL | 186 | 291 |
| CIC MUTATED | 60 | 41 |
| CIC WILD-TYPE | 126 | 250 |
Figure S12. Get High-res Image Gene #4: 'CIC MUTATION STATUS' versus Clinical Feature #4: 'RADIATION_THERAPY'
P value = 1e-05 (Fisher's exact test), Q value = 0.00061
Table S13. Gene #4: 'CIC MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 192 | 130 | 190 |
| CIC MUTATED | 3 | 23 | 82 |
| CIC WILD-TYPE | 189 | 107 | 108 |
Figure S13. Get High-res Image Gene #4: 'CIC MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
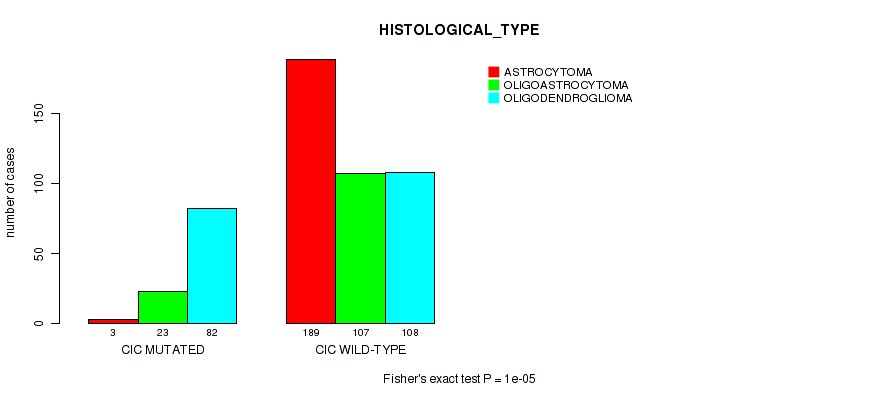
P value = 5e-05 (Fisher's exact test), Q value = 0.0028
Table S14. Gene #5: 'NOTCH1 MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 192 | 130 | 190 |
| NOTCH1 MUTATED | 6 | 7 | 29 |
| NOTCH1 WILD-TYPE | 186 | 123 | 161 |
Figure S14. Get High-res Image Gene #5: 'NOTCH1 MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
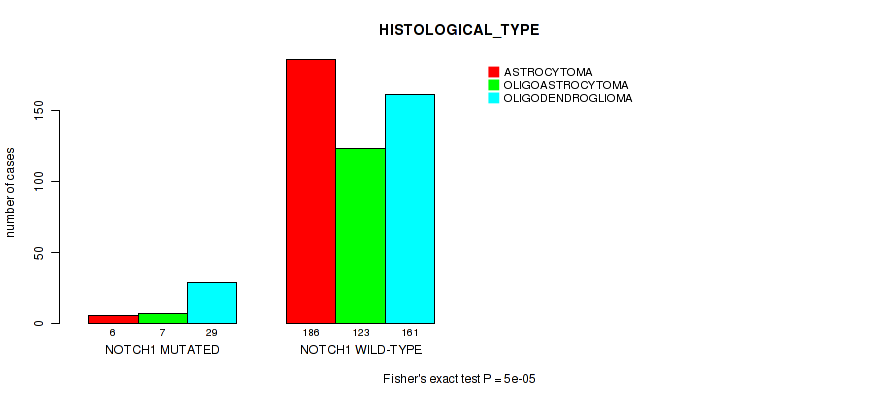
P value = 0.00065 (Fisher's exact test), Q value = 0.024
Table S15. Gene #6: 'IDH2 MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 192 | 130 | 190 |
| IDH2 MUTATED | 1 | 5 | 14 |
| IDH2 WILD-TYPE | 191 | 125 | 176 |
Figure S15. Get High-res Image Gene #6: 'IDH2 MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
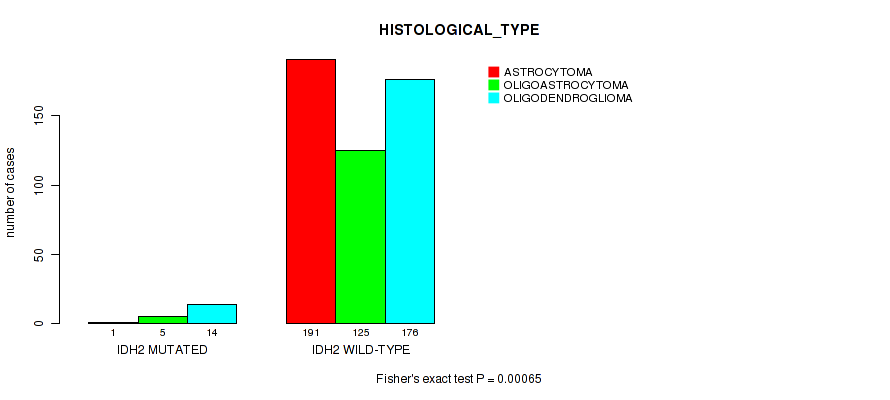
P value = 0.00301 (Wilcoxon-test), Q value = 0.088
Table S16. Gene #8: 'FUBP1 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 511 | 42.9 (13.4) |
| FUBP1 MUTATED | 47 | 47.4 (9.5) |
| FUBP1 WILD-TYPE | 464 | 42.5 (13.6) |
Figure S16. Get High-res Image Gene #8: 'FUBP1 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
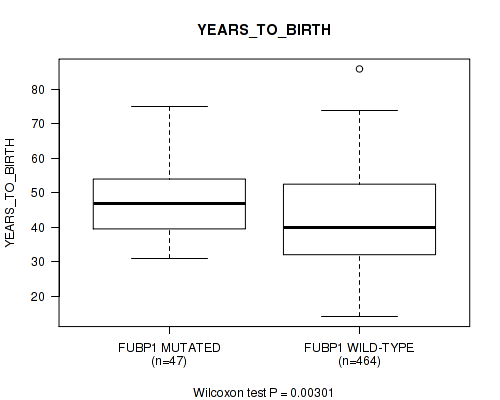
P value = 1e-05 (Fisher's exact test), Q value = 0.00061
Table S17. Gene #8: 'FUBP1 MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 192 | 130 | 190 |
| FUBP1 MUTATED | 2 | 9 | 36 |
| FUBP1 WILD-TYPE | 190 | 121 | 154 |
Figure S17. Get High-res Image Gene #8: 'FUBP1 MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
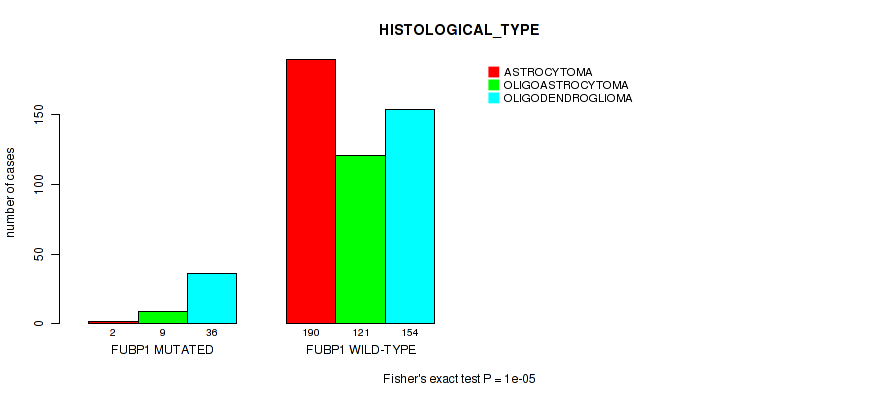
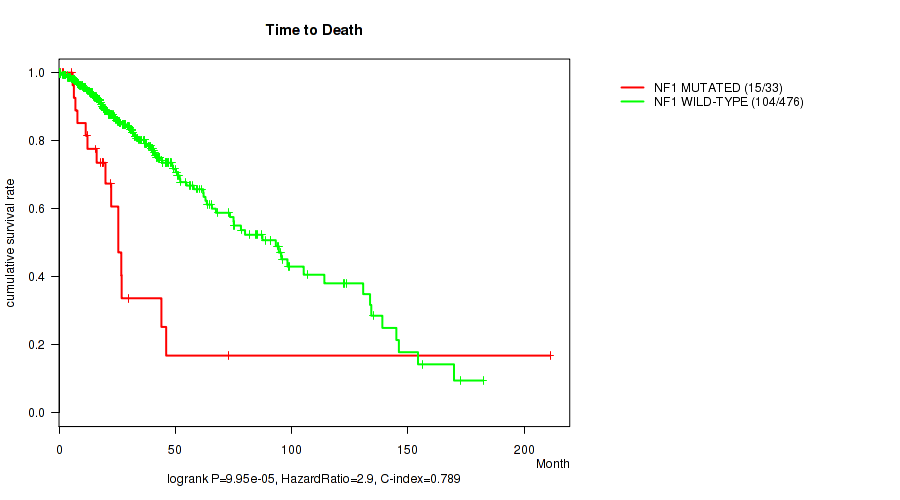
P value = 9.95e-05 (logrank test), Q value = 0.0051
Table S18. Gene #9: 'NF1 MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 509 | 119 | 0.0 - 211.2 (20.7) |
| NF1 MUTATED | 33 | 15 | 0.2 - 211.2 (18.6) |
| NF1 WILD-TYPE | 476 | 104 | 0.0 - 182.3 (21.3) |
Figure S18. Get High-res Image Gene #9: 'NF1 MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
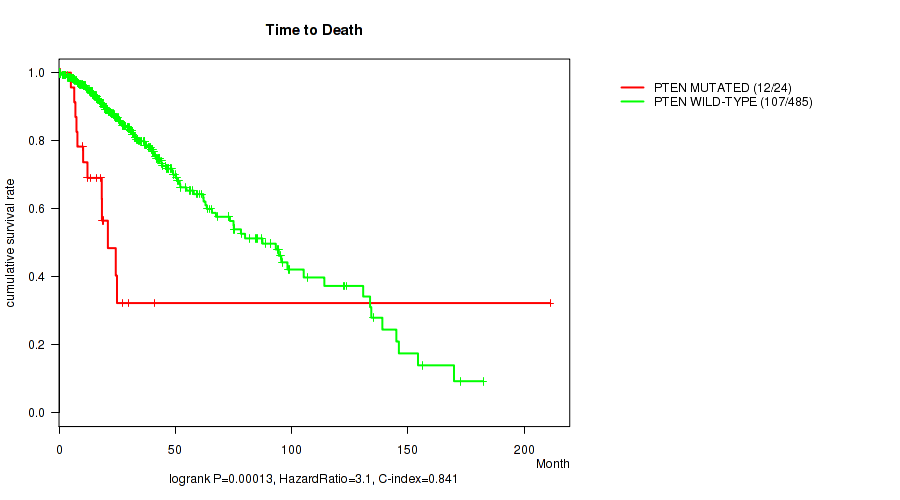
P value = 0.00013 (logrank test), Q value = 0.0061
Table S19. Gene #11: 'PTEN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 509 | 119 | 0.0 - 211.2 (20.7) |
| PTEN MUTATED | 24 | 12 | 0.5 - 211.2 (17.1) |
| PTEN WILD-TYPE | 485 | 107 | 0.0 - 182.3 (21.3) |
Figure S19. Get High-res Image Gene #11: 'PTEN MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
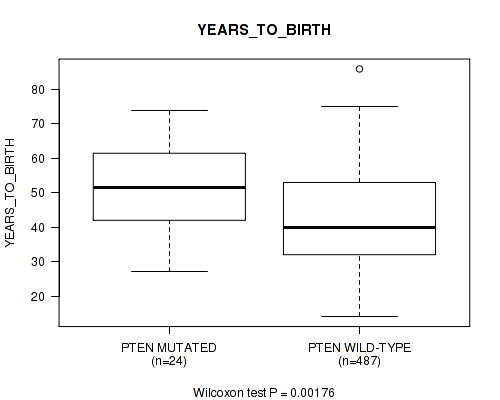
P value = 0.00176 (Wilcoxon-test), Q value = 0.062
Table S20. Gene #11: 'PTEN MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 511 | 42.9 (13.4) |
| PTEN MUTATED | 24 | 51.0 (12.0) |
| PTEN WILD-TYPE | 487 | 42.5 (13.3) |
Figure S20. Get High-res Image Gene #11: 'PTEN MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
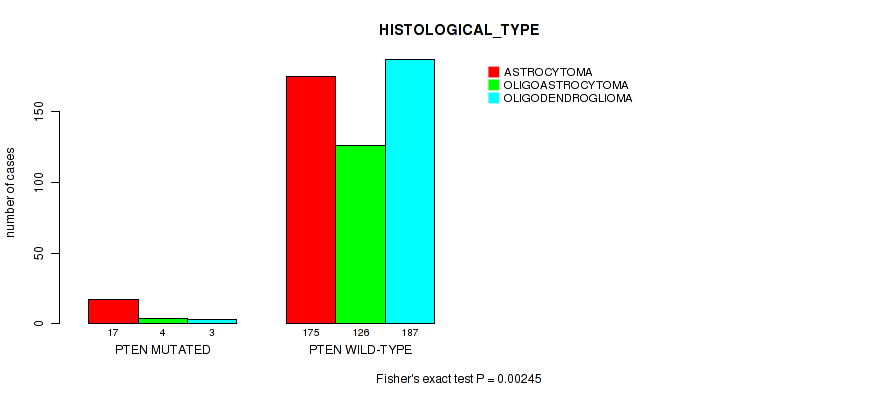
P value = 0.00245 (Fisher's exact test), Q value = 0.077
Table S21. Gene #11: 'PTEN MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
| nPatients | ASTROCYTOMA | OLIGOASTROCYTOMA | OLIGODENDROGLIOMA |
|---|---|---|---|
| ALL | 192 | 130 | 190 |
| PTEN MUTATED | 17 | 4 | 3 |
| PTEN WILD-TYPE | 175 | 126 | 187 |
Figure S21. Get High-res Image Gene #11: 'PTEN MUTATION STATUS' versus Clinical Feature #6: 'HISTOLOGICAL_TYPE'
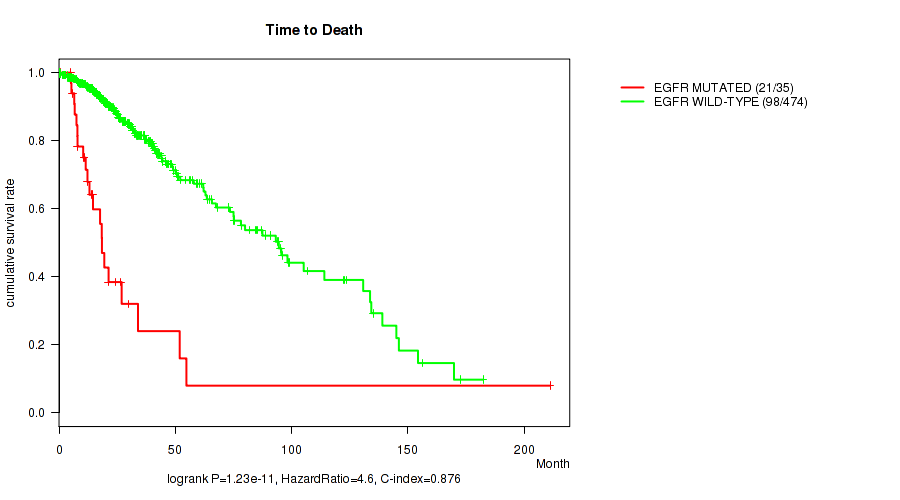
P value = 1.23e-11 (logrank test), Q value = 2.1e-09
Table S22. Gene #12: 'EGFR MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 509 | 119 | 0.0 - 211.2 (20.7) |
| EGFR MUTATED | 35 | 21 | 0.5 - 211.2 (13.1) |
| EGFR WILD-TYPE | 474 | 98 | 0.0 - 182.3 (21.5) |
Figure S22. Get High-res Image Gene #12: 'EGFR MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
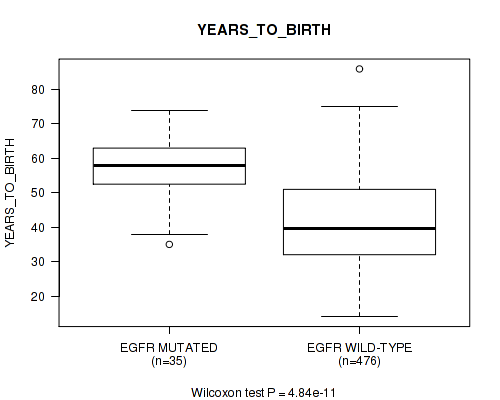
P value = 4.84e-11 (Wilcoxon-test), Q value = 6.5e-09
Table S23. Gene #12: 'EGFR MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 511 | 42.9 (13.4) |
| EGFR MUTATED | 35 | 57.7 (9.1) |
| EGFR WILD-TYPE | 476 | 41.8 (13.0) |
Figure S23. Get High-res Image Gene #12: 'EGFR MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
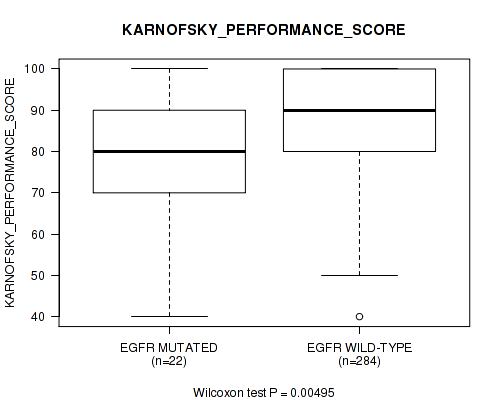
P value = 0.00495 (Wilcoxon-test), Q value = 0.14
Table S24. Gene #12: 'EGFR MUTATION STATUS' versus Clinical Feature #5: 'KARNOFSKY_PERFORMANCE_SCORE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 306 | 86.7 (12.4) |
| EGFR MUTATED | 22 | 78.2 (16.8) |
| EGFR WILD-TYPE | 284 | 87.4 (11.8) |
Figure S24. Get High-res Image Gene #12: 'EGFR MUTATION STATUS' versus Clinical Feature #5: 'KARNOFSKY_PERFORMANCE_SCORE'
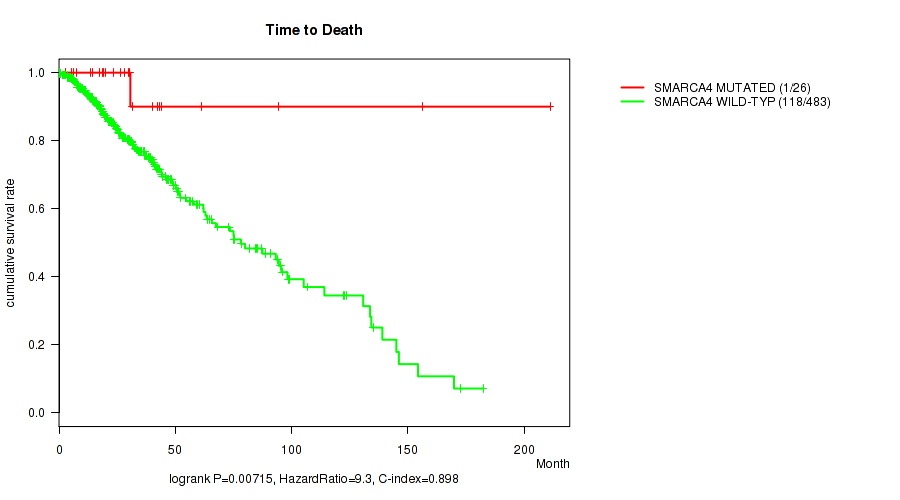
P value = 0.00715 (logrank test), Q value = 0.18
Table S25. Gene #18: 'SMARCA4 MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 509 | 119 | 0.0 - 211.2 (20.7) |
| SMARCA4 MUTATED | 26 | 1 | 0.2 - 211.2 (27.2) |
| SMARCA4 WILD-TYPE | 483 | 118 | 0.0 - 182.3 (20.7) |
Figure S25. Get High-res Image Gene #18: 'SMARCA4 MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
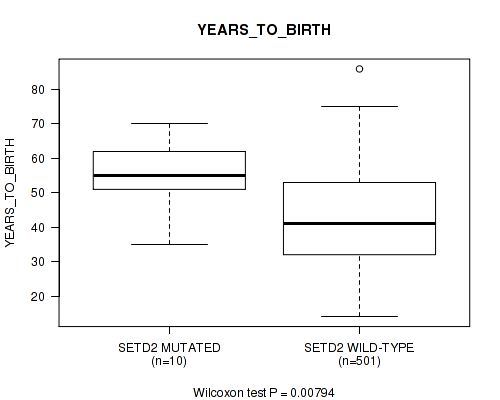
P value = 0.00794 (Wilcoxon-test), Q value = 0.2
Table S26. Gene #44: 'SETD2 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 511 | 42.9 (13.4) |
| SETD2 MUTATED | 10 | 54.0 (10.9) |
| SETD2 WILD-TYPE | 501 | 42.7 (13.3) |
Figure S26. Get High-res Image Gene #44: 'SETD2 MUTATION STATUS' versus Clinical Feature #2: 'YEARS_TO_BIRTH'
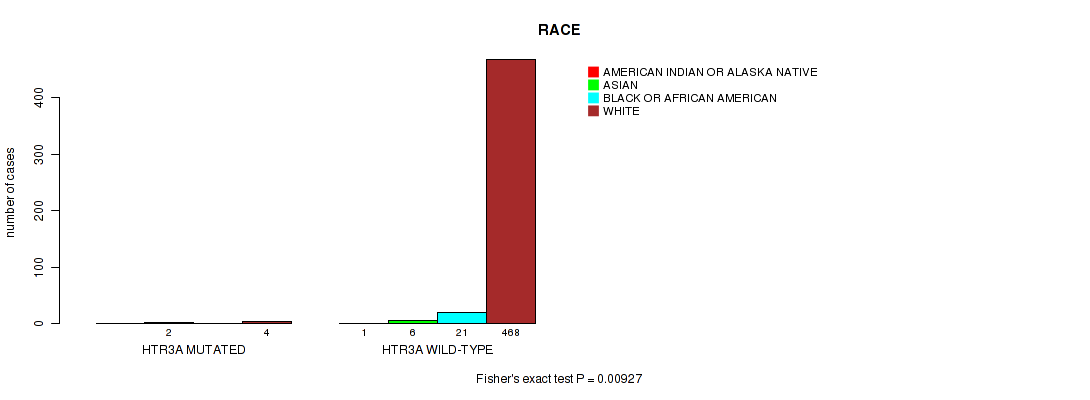
P value = 0.00927 (Fisher's exact test), Q value = 0.21
Table S27. Gene #63: 'HTR3A MUTATION STATUS' versus Clinical Feature #7: 'RACE'
| nPatients | AMERICAN INDIAN OR ALASKA NATIVE | ASIAN | BLACK OR AFRICAN AMERICAN | WHITE |
|---|---|---|---|---|
| ALL | 1 | 8 | 21 | 472 |
| HTR3A MUTATED | 0 | 2 | 0 | 4 |
| HTR3A WILD-TYPE | 1 | 6 | 21 | 468 |
Figure S27. Get High-res Image Gene #63: 'HTR3A MUTATION STATUS' versus Clinical Feature #7: 'RACE'
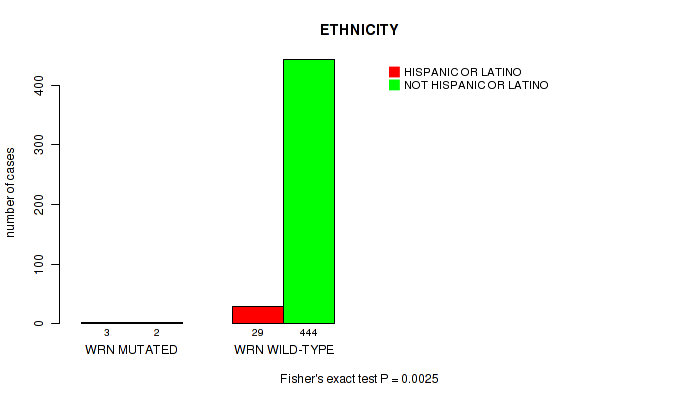
P value = 0.0025 (Fisher's exact test), Q value = 0.077
Table S28. Gene #70: 'WRN MUTATION STATUS' versus Clinical Feature #8: 'ETHNICITY'
| nPatients | HISPANIC OR LATINO | NOT HISPANIC OR LATINO |
|---|---|---|
| ALL | 32 | 446 |
| WRN MUTATED | 3 | 2 |
| WRN WILD-TYPE | 29 | 444 |
Figure S28. Get High-res Image Gene #70: 'WRN MUTATION STATUS' versus Clinical Feature #8: 'ETHNICITY'
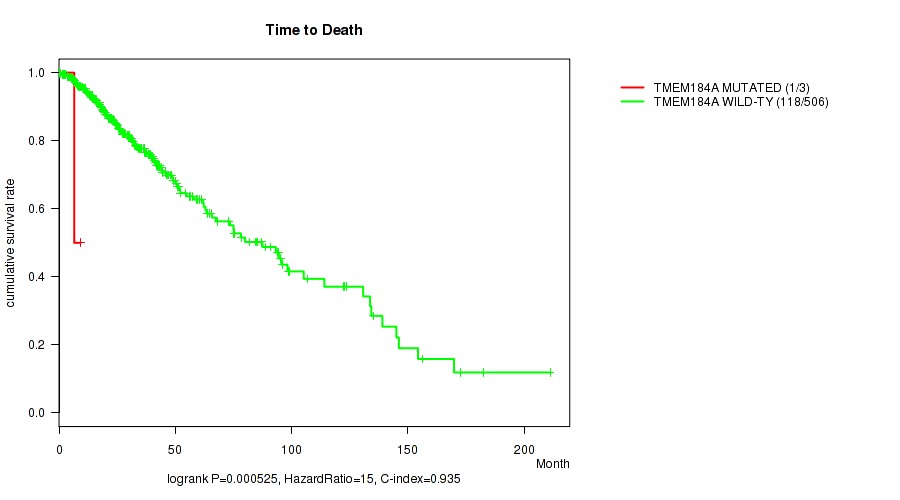
P value = 0.000525 (logrank test), Q value = 0.021
Table S29. Gene #76: 'TMEM184A MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 509 | 119 | 0.0 - 211.2 (20.7) |
| TMEM184A MUTATED | 3 | 1 | 0.3 - 9.2 (6.5) |
| TMEM184A WILD-TYPE | 506 | 118 | 0.0 - 211.2 (20.8) |
Figure S29. Get High-res Image Gene #76: 'TMEM184A MUTATION STATUS' versus Clinical Feature #1: 'Time to Death'
-
Mutation data file = sample_sig_gene_table.txt from Mutsig_2CV pipeline
-
Processed Mutation data file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/GDAC_Correlate_Genomic_Events_Preprocess/LGG-TP/20067756/transformed.cor.cli.txt
-
Clinical data file = /xchip/cga/gdac-prod/tcga-gdac/jobResults/Append_Data/LGG-TP/19775312/LGG-TP.merged_data.txt
-
Number of patients = 512
-
Number of significantly mutated genes = 84
-
Number of selected clinical features = 8
-
Exclude genes that fewer than K tumors have mutations, K = 3
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For binary or multi-class clinical features (nominal or ordinal), two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.