This pipeline computes the correlation between cancer subtypes identified by different molecular patterns and selected clinical features.
Testing the association between subtypes identified by 8 different clustering approaches and 43 clinical features across 200 patients, 9 significant findings detected with P value < 0.05 and Q value < 0.25.
-
3 subtypes identified in current cancer cohort by 'Copy Number Ratio CNMF subtypes'. These subtypes do not correlate to any clinical features.
-
6 subtypes identified in current cancer cohort by 'METHLYATION CNMF'. These subtypes correlate to 'HISTOLOGICAL.TYPE'.
-
CNMF clustering analysis on sequencing-based mRNA expression data identified 3 subtypes that correlate to 'HISTOLOGICAL.TYPE'.
-
Consensus hierarchical clustering analysis on sequencing-based mRNA expression data identified 4 subtypes that correlate to 'HISTOLOGICAL.TYPE'.
-
3 subtypes identified in current cancer cohort by 'MIRSEQ CNMF'. These subtypes correlate to 'HISTOLOGICAL.TYPE'.
-
3 subtypes identified in current cancer cohort by 'MIRSEQ CHIERARCHICAL'. These subtypes correlate to 'HISTOLOGICAL.TYPE'.
-
3 subtypes identified in current cancer cohort by 'MIRseq Mature CNMF subtypes'. These subtypes correlate to 'HISTOLOGICAL.TYPE'.
-
7 subtypes identified in current cancer cohort by 'MIRseq Mature cHierClus subtypes'. These subtypes correlate to 'HISTOLOGICAL.TYPE', 'NEOPLASMHISTOLOGICGRADE', and 'INITIAL_PATHOLOGIC_DX_YEAR'.
Table 1. Get Full Table Overview of the association between subtypes identified by 8 different clustering approaches and 43 clinical features. Shown in the table are P values (Q values). Thresholded by P value < 0.05 and Q value < 0.25, 9 significant findings detected.
|
Clinical Features |
Statistical Tests |
Copy Number Ratio CNMF subtypes |
METHLYATION CNMF |
RNAseq CNMF subtypes |
RNAseq cHierClus subtypes |
MIRSEQ CNMF |
MIRSEQ CHIERARCHICAL |
MIRseq Mature CNMF subtypes |
MIRseq Mature cHierClus subtypes |
| Time to Death | logrank test |
0.478 (1.00) |
0.241 (1.00) |
0.353 (1.00) |
0.0267 (1.00) |
0.499 (1.00) |
0.647 (1.00) |
0.829 (1.00) |
0.147 (1.00) |
| AGE | Kruskal-Wallis (anova) |
0.14 (1.00) |
0.0117 (1.00) |
0.11 (1.00) |
0.0489 (1.00) |
0.221 (1.00) |
0.166 (1.00) |
0.0366 (1.00) |
0.0118 (1.00) |
| PATHOLOGY T STAGE | Fisher's exact test |
0.945 (1.00) |
0.983 (1.00) |
0.723 (1.00) |
0.762 (1.00) |
0.89 (1.00) |
0.205 (1.00) |
0.785 (1.00) |
0.436 (1.00) |
| PATHOLOGY N STAGE | Fisher's exact test |
0.105 (1.00) |
0.331 (1.00) |
0.0153 (1.00) |
0.583 (1.00) |
0.587 (1.00) |
0.776 (1.00) |
0.727 (1.00) |
0.631 (1.00) |
| PATHOLOGY M STAGE | Fisher's exact test |
0.269 (1.00) |
0.716 (1.00) |
0.114 (1.00) |
0.242 (1.00) |
0.628 (1.00) |
0.389 (1.00) |
0.407 (1.00) |
0.643 (1.00) |
| HISTOLOGICAL TYPE | Fisher's exact test |
0.452 (1.00) |
1e-05 (0.00341) |
1e-05 (0.00341) |
1e-05 (0.00341) |
1e-05 (0.00341) |
1e-05 (0.00341) |
1e-05 (0.00341) |
1e-05 (0.00341) |
| RADIATIONS RADIATION REGIMENINDICATION | Fisher's exact test |
0.891 (1.00) |
0.429 (1.00) |
0.317 (1.00) |
0.536 (1.00) |
0.411 (1.00) |
0.628 (1.00) |
0.416 (1.00) |
0.0401 (1.00) |
| NUMBERPACKYEARSSMOKED | Kruskal-Wallis (anova) |
0.454 (1.00) |
0.544 (1.00) |
0.0633 (1.00) |
0.198 (1.00) |
0.0665 (1.00) |
0.213 (1.00) |
0.395 (1.00) |
0.491 (1.00) |
| NUMBER OF LYMPH NODES | Kruskal-Wallis (anova) |
0.166 (1.00) |
0.513 (1.00) |
0.0587 (1.00) |
0.557 (1.00) |
0.443 (1.00) |
0.667 (1.00) |
0.654 (1.00) |
0.485 (1.00) |
| RACE | Fisher's exact test |
0.536 (1.00) |
0.287 (1.00) |
0.238 (1.00) |
0.387 (1.00) |
0.386 (1.00) |
0.421 (1.00) |
0.156 (1.00) |
0.632 (1.00) |
| ETHNICITY | Fisher's exact test |
0.925 (1.00) |
0.164 (1.00) |
0.477 (1.00) |
0.436 (1.00) |
0.506 (1.00) |
0.0869 (1.00) |
0.335 (1.00) |
0.266 (1.00) |
| WEIGHT KG AT DIAGNOSIS | Kruskal-Wallis (anova) |
0.168 (1.00) |
0.56 (1.00) |
0.356 (1.00) |
0.517 (1.00) |
0.35 (1.00) |
0.634 (1.00) |
0.124 (1.00) |
0.407 (1.00) |
| TUMOR STATUS | Fisher's exact test |
0.578 (1.00) |
0.795 (1.00) |
0.945 (1.00) |
0.195 (1.00) |
0.724 (1.00) |
0.671 (1.00) |
0.566 (1.00) |
0.454 (1.00) |
| TUMOR SAMPLE PROCUREMENT COUNTRY | Fisher's exact test |
0.0209 (1.00) |
0.477 (1.00) |
0.662 (1.00) |
0.535 (1.00) |
0.579 (1.00) |
0.274 (1.00) |
0.311 (1.00) |
0.572 (1.00) |
| NEOPLASMHISTOLOGICGRADE | Fisher's exact test |
0.0515 (1.00) |
0.74 (1.00) |
0.131 (1.00) |
0.0169 (1.00) |
0.0596 (1.00) |
0.327 (1.00) |
0.0581 (1.00) |
0.00071 (0.236) |
| TOBACCO SMOKING YEAR STOPPED | Kruskal-Wallis (anova) |
0.426 (1.00) |
0.355 (1.00) |
0.799 (1.00) |
0.794 (1.00) |
0.317 (1.00) |
0.16 (1.00) |
0.737 (1.00) |
0.388 (1.00) |
| TOBACCO SMOKING PACK YEARS SMOKED | Kruskal-Wallis (anova) |
0.454 (1.00) |
0.544 (1.00) |
0.0633 (1.00) |
0.198 (1.00) |
0.0665 (1.00) |
0.213 (1.00) |
0.395 (1.00) |
0.491 (1.00) |
| TOBACCO SMOKING HISTORY | Fisher's exact test |
0.894 (1.00) |
0.539 (1.00) |
0.384 (1.00) |
0.413 (1.00) |
0.245 (1.00) |
0.38 (1.00) |
0.762 (1.00) |
0.899 (1.00) |
| PATIENT AGEBEGANSMOKINGINYEARS | Kruskal-Wallis (anova) |
0.386 (1.00) |
0.549 (1.00) |
0.812 (1.00) |
0.576 (1.00) |
0.239 (1.00) |
0.258 (1.00) |
0.0729 (1.00) |
0.105 (1.00) |
| RADIATION THERAPY TYPE | Fisher's exact test |
0.211 (1.00) |
0.366 (1.00) |
0.84 (1.00) |
0.458 (1.00) |
0.267 (1.00) |
0.724 (1.00) |
0.13 (1.00) |
0.0239 (1.00) |
| RADIATION ADJUVANT UNITS | Fisher's exact test |
1 (1.00) |
0.84 (1.00) |
1 (1.00) |
0.471 (1.00) |
0.137 (1.00) |
0.784 (1.00) |
0.417 (1.00) |
0.94 (1.00) |
| PREGNANCIES COUNT TOTAL | Kruskal-Wallis (anova) |
0.606 (1.00) |
0.417 (1.00) |
0.12 (1.00) |
0.274 (1.00) |
0.229 (1.00) |
0.0722 (1.00) |
0.464 (1.00) |
0.375 (1.00) |
| PREGNANCIES COUNT STILLBIRTH | Kruskal-Wallis (anova) |
0.221 (1.00) |
0.00866 (1.00) |
0.159 (1.00) |
0.468 (1.00) |
0.791 (1.00) |
0.939 (1.00) |
0.951 (1.00) |
0.26 (1.00) |
| PATIENT PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT | Kruskal-Wallis (anova) |
0.928 (1.00) |
0.206 (1.00) |
0.355 (1.00) |
0.347 (1.00) |
0.0365 (1.00) |
0.102 (1.00) |
0.217 (1.00) |
0.269 (1.00) |
| PREGNANCIES COUNT LIVE BIRTH | Kruskal-Wallis (anova) |
0.528 (1.00) |
0.258 (1.00) |
0.142 (1.00) |
0.155 (1.00) |
0.167 (1.00) |
0.0486 (1.00) |
0.618 (1.00) |
0.21 (1.00) |
| PATIENT PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT | Kruskal-Wallis (anova) |
0.883 (1.00) |
0.0424 (1.00) |
0.737 (1.00) |
0.911 (1.00) |
0.392 (1.00) |
0.512 (1.00) |
0.147 (1.00) |
0.737 (1.00) |
| PREGNANCIES COUNT ECTOPIC | Kruskal-Wallis (anova) |
0.266 (1.00) |
0.777 (1.00) |
0.96 (1.00) |
0.999 (1.00) |
0.831 (1.00) |
0.71 (1.00) |
0.911 (1.00) |
0.765 (1.00) |
| LYMPH NODE LOCATION | Fisher's exact test |
0.363 (1.00) |
0.593 (1.00) |
0.0318 (1.00) |
0.521 (1.00) |
0.343 (1.00) |
0.562 (1.00) |
0.131 (1.00) |
0.309 (1.00) |
| LOCATION OF POSITIVE MARGINS | Fisher's exact test |
0.933 (1.00) |
0.725 (1.00) |
0.839 (1.00) |
0.436 (1.00) |
0.706 (1.00) |
1 (1.00) |
0.0317 (1.00) |
0.984 (1.00) |
| MENOPAUSE STATUS | Fisher's exact test |
0.326 (1.00) |
0.0169 (1.00) |
0.275 (1.00) |
0.0133 (1.00) |
0.466 (1.00) |
0.493 (1.00) |
0.471 (1.00) |
0.025 (1.00) |
| LYMPHOVASCULAR INVOLVEMENT | Fisher's exact test |
0.245 (1.00) |
0.858 (1.00) |
0.459 (1.00) |
0.877 (1.00) |
0.325 (1.00) |
0.363 (1.00) |
0.665 (1.00) |
0.386 (1.00) |
| LYMPH NODES EXAMINED HE COUNT | Kruskal-Wallis (anova) |
0.166 (1.00) |
0.513 (1.00) |
0.0587 (1.00) |
0.557 (1.00) |
0.443 (1.00) |
0.667 (1.00) |
0.654 (1.00) |
0.485 (1.00) |
| LYMPH NODES EXAMINED | Kruskal-Wallis (anova) |
0.593 (1.00) |
0.829 (1.00) |
0.79 (1.00) |
0.78 (1.00) |
0.528 (1.00) |
0.856 (1.00) |
0.179 (1.00) |
0.24 (1.00) |
| KERATINIZATION SQUAMOUS CELL | Fisher's exact test |
0.628 (1.00) |
0.0149 (1.00) |
0.0222 (1.00) |
0.115 (1.00) |
0.031 (1.00) |
0.0822 (1.00) |
0.0205 (1.00) |
0.021 (1.00) |
| INITIAL PATHOLOGIC DX YEAR | Kruskal-Wallis (anova) |
0.213 (1.00) |
0.34 (1.00) |
0.147 (1.00) |
0.123 (1.00) |
0.173 (1.00) |
0.342 (1.00) |
0.222 (1.00) |
0.000681 (0.227) |
| HYSTERECTOMY TYPE | Fisher's exact test |
0.099 (1.00) |
0.285 (1.00) |
0.477 (1.00) |
0.169 (1.00) |
0.136 (1.00) |
0.132 (1.00) |
0.307 (1.00) |
0.028 (1.00) |
| HISTORY HORMONAL CONTRACEPTIVES USE | Fisher's exact test |
0.404 (1.00) |
0.647 (1.00) |
0.114 (1.00) |
0.0445 (1.00) |
0.944 (1.00) |
0.211 (1.00) |
0.931 (1.00) |
0.172 (1.00) |
| HEIGHT CM AT DIAGNOSIS | Kruskal-Wallis (anova) |
0.344 (1.00) |
0.158 (1.00) |
0.502 (1.00) |
0.41 (1.00) |
0.185 (1.00) |
0.678 (1.00) |
0.271 (1.00) |
0.482 (1.00) |
| CORPUS INVOLVEMENT | Fisher's exact test |
0.487 (1.00) |
0.536 (1.00) |
0.111 (1.00) |
0.877 (1.00) |
0.322 (1.00) |
0.659 (1.00) |
0.124 (1.00) |
0.899 (1.00) |
| CHEMO CONCURRENT TYPE | Fisher's exact test |
0.535 (1.00) |
0.296 (1.00) |
1 (1.00) |
1 (1.00) |
1 (1.00) |
0.103 (1.00) |
0.537 (1.00) |
0.89 (1.00) |
| CERVIX SUV RESULTS | Kruskal-Wallis (anova) |
0.0809 (1.00) |
0.456 (1.00) |
0.362 (1.00) |
0.0947 (1.00) |
0.0809 (1.00) |
|||
| AJCC TUMOR PATHOLOGIC PT | Fisher's exact test |
0.766 (1.00) |
0.402 (1.00) |
0.0624 (1.00) |
0.00179 (0.594) |
0.35 (1.00) |
0.113 (1.00) |
0.742 (1.00) |
0.304 (1.00) |
| AGE AT DIAGNOSIS | Kruskal-Wallis (anova) |
0.112 (1.00) |
0.0161 (1.00) |
0.128 (1.00) |
0.0454 (1.00) |
0.249 (1.00) |
0.156 (1.00) |
0.0504 (1.00) |
0.0105 (1.00) |
Table S1. Description of clustering approach #1: 'Copy Number Ratio CNMF subtypes'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 55 | 39 | 82 |
P value = 0.478 (logrank test), Q value = 1
Table S2. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 169 | 32 | 0.0 - 182.9 (13.6) |
| subtype1 | 53 | 8 | 0.0 - 173.3 (14.9) |
| subtype2 | 35 | 9 | 0.1 - 118.0 (13.1) |
| subtype3 | 81 | 15 | 0.0 - 182.9 (10.1) |
Figure S1. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #1: 'Time to Death'

P value = 0.14 (Kruskal-Wallis (anova)), Q value = 1
Table S3. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 175 | 47.5 (13.1) |
| subtype1 | 54 | 49.4 (10.4) |
| subtype2 | 39 | 46.7 (15.0) |
| subtype3 | 82 | 46.5 (13.7) |
Figure S2. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #2: 'AGE'

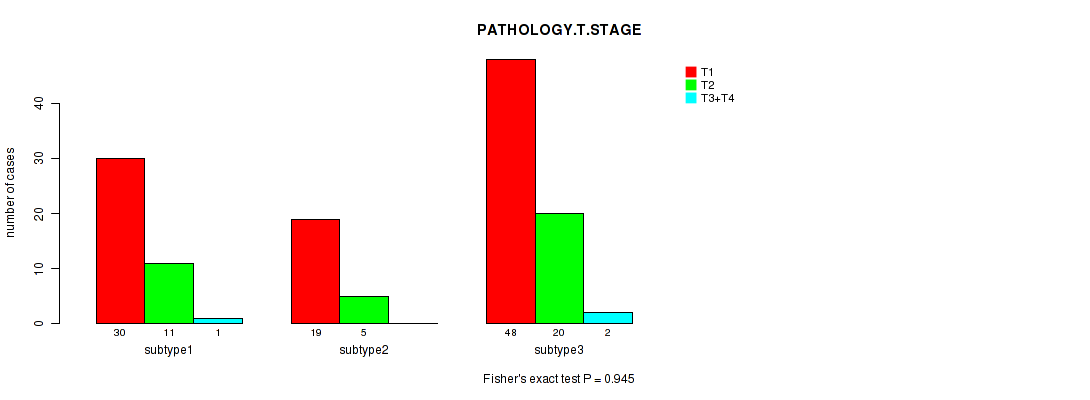
P value = 0.945 (Fisher's exact test), Q value = 1
Table S4. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
| nPatients | T1 | T2 | T3+T4 |
|---|---|---|---|
| ALL | 97 | 36 | 3 |
| subtype1 | 30 | 11 | 1 |
| subtype2 | 19 | 5 | 0 |
| subtype3 | 48 | 20 | 2 |
Figure S3. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
P value = 0.105 (Fisher's exact test), Q value = 1
Table S5. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 87 | 40 |
| subtype1 | 25 | 14 |
| subtype2 | 13 | 11 |
| subtype3 | 49 | 15 |
Figure S4. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'

P value = 0.269 (Fisher's exact test), Q value = 1
Table S6. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'
| nPatients | M0 | M1 | MX |
|---|---|---|---|
| ALL | 72 | 3 | 65 |
| subtype1 | 28 | 1 | 16 |
| subtype2 | 14 | 0 | 11 |
| subtype3 | 30 | 2 | 38 |
Figure S5. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'

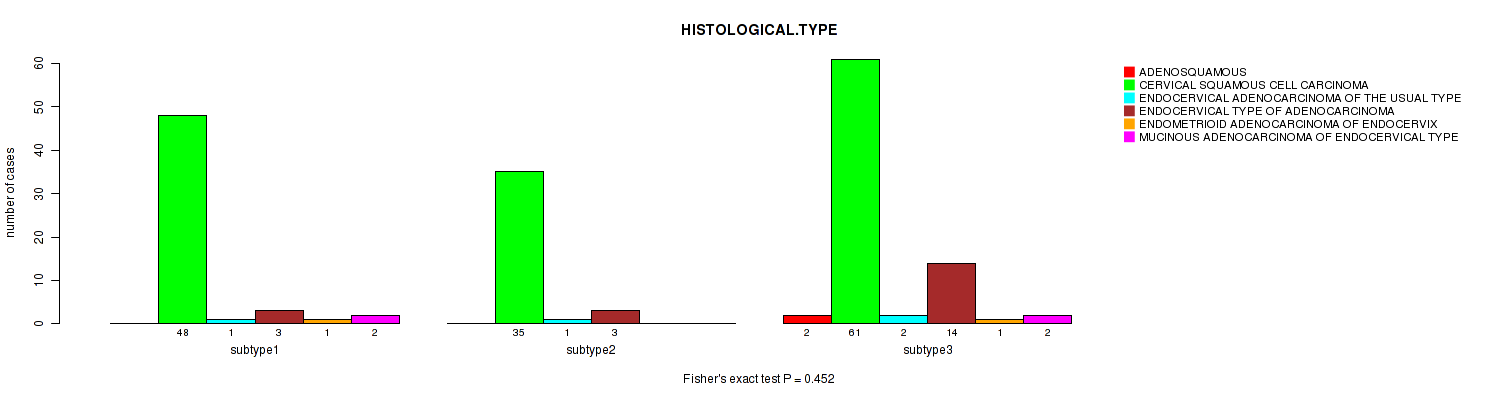
P value = 0.452 (Fisher's exact test), Q value = 1
Table S7. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
| nPatients | ADENOSQUAMOUS | CERVICAL SQUAMOUS CELL CARCINOMA | ENDOCERVICAL ADENOCARCINOMA OF THE USUAL TYPE | ENDOCERVICAL TYPE OF ADENOCARCINOMA | ENDOMETRIOID ADENOCARCINOMA OF ENDOCERVIX | MUCINOUS ADENOCARCINOMA OF ENDOCERVICAL TYPE |
|---|---|---|---|---|---|---|
| ALL | 2 | 144 | 4 | 20 | 2 | 4 |
| subtype1 | 0 | 48 | 1 | 3 | 1 | 2 |
| subtype2 | 0 | 35 | 1 | 3 | 0 | 0 |
| subtype3 | 2 | 61 | 2 | 14 | 1 | 2 |
Figure S6. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
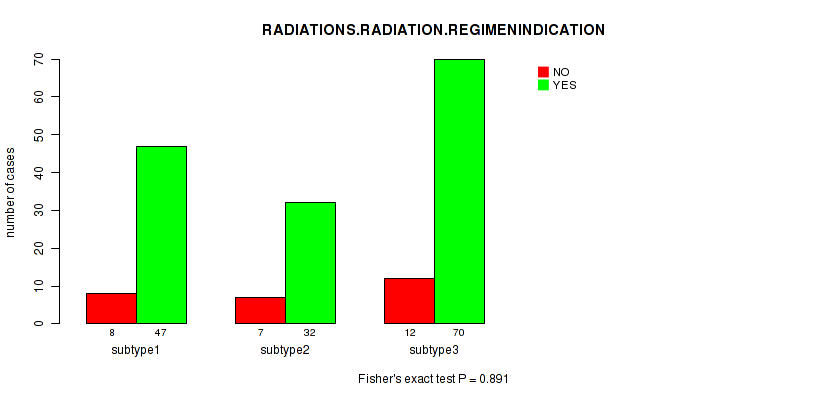
P value = 0.891 (Fisher's exact test), Q value = 1
Table S8. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 27 | 149 |
| subtype1 | 8 | 47 |
| subtype2 | 7 | 32 |
| subtype3 | 12 | 70 |
Figure S7. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'
P value = 0.454 (Kruskal-Wallis (anova)), Q value = 1
Table S9. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 53 | 18.6 (12.4) |
| subtype1 | 15 | 18.1 (15.0) |
| subtype2 | 11 | 21.0 (10.2) |
| subtype3 | 27 | 17.9 (11.9) |
Figure S8. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'

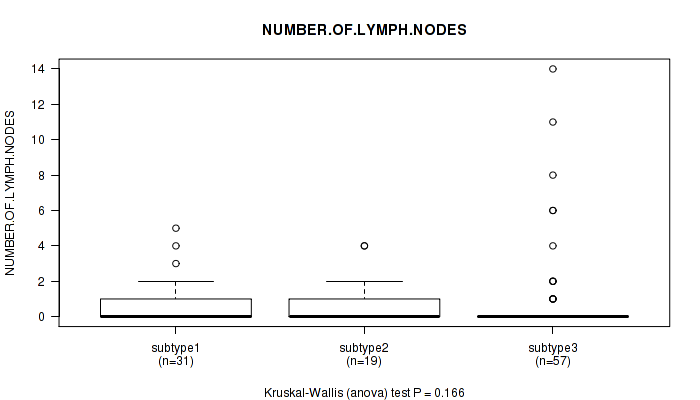
P value = 0.166 (Kruskal-Wallis (anova)), Q value = 1
Table S10. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 107 | 0.9 (2.2) |
| subtype1 | 31 | 0.8 (1.3) |
| subtype2 | 19 | 0.9 (1.3) |
| subtype3 | 57 | 1.0 (2.8) |
Figure S9. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
P value = 0.536 (Fisher's exact test), Q value = 1
Table S11. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #10: 'RACE'
| nPatients | AMERICAN INDIAN OR ALASKA NATIVE | ASIAN | BLACK OR AFRICAN AMERICAN | NATIVE HAWAIIAN OR OTHER PACIFIC ISLANDER | WHITE |
|---|---|---|---|---|---|
| ALL | 7 | 15 | 18 | 1 | 130 |
| subtype1 | 2 | 8 | 5 | 0 | 38 |
| subtype2 | 1 | 1 | 6 | 0 | 31 |
| subtype3 | 4 | 6 | 7 | 1 | 61 |
Figure S10. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #10: 'RACE'

P value = 0.925 (Fisher's exact test), Q value = 1
Table S12. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #11: 'ETHNICITY'
| nPatients | HISPANIC OR LATINO | NOT HISPANIC OR LATINO |
|---|---|---|
| ALL | 11 | 127 |
| subtype1 | 3 | 40 |
| subtype2 | 2 | 29 |
| subtype3 | 6 | 58 |
Figure S11. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #11: 'ETHNICITY'

P value = 0.168 (Kruskal-Wallis (anova)), Q value = 1
Table S13. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 162 | 75.6 (20.9) |
| subtype1 | 52 | 73.1 (25.4) |
| subtype2 | 37 | 76.9 (16.0) |
| subtype3 | 73 | 76.8 (19.7) |
Figure S12. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'

P value = 0.578 (Fisher's exact test), Q value = 1
Table S14. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #13: 'TUMOR_STATUS'
| nPatients | TUMOR FREE | WITH TUMOR |
|---|---|---|
| ALL | 48 | 19 |
| subtype1 | 17 | 5 |
| subtype2 | 10 | 6 |
| subtype3 | 21 | 8 |
Figure S13. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #13: 'TUMOR_STATUS'

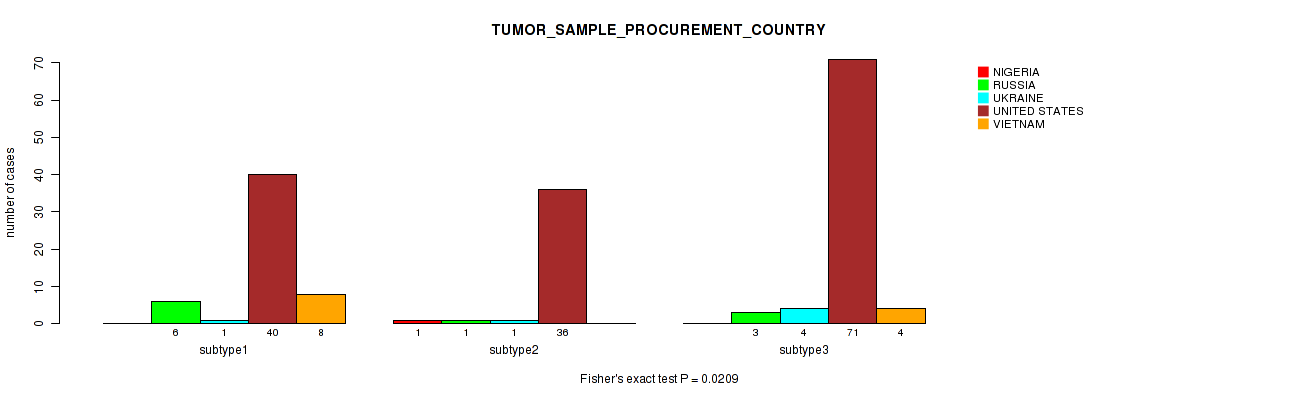
P value = 0.0209 (Fisher's exact test), Q value = 1
Table S15. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
| nPatients | NIGERIA | RUSSIA | UKRAINE | UNITED STATES | VIETNAM |
|---|---|---|---|---|---|
| ALL | 1 | 10 | 6 | 147 | 12 |
| subtype1 | 0 | 6 | 1 | 40 | 8 |
| subtype2 | 1 | 1 | 1 | 36 | 0 |
| subtype3 | 0 | 3 | 4 | 71 | 4 |
Figure S14. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
P value = 0.0515 (Fisher's exact test), Q value = 1
Table S16. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'
| nPatients | G1 | G2 | G3 | G4 | GX |
|---|---|---|---|---|---|
| ALL | 14 | 83 | 72 | 1 | 5 |
| subtype1 | 6 | 30 | 18 | 0 | 1 |
| subtype2 | 0 | 14 | 23 | 1 | 1 |
| subtype3 | 8 | 39 | 31 | 0 | 3 |
Figure S15. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'

P value = 0.426 (Kruskal-Wallis (anova)), Q value = 1
Table S17. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 25 | 1997.7 (11.7) |
| subtype1 | 8 | 1996.9 (11.1) |
| subtype2 | 4 | 1991.0 (12.8) |
| subtype3 | 13 | 2000.3 (11.8) |
Figure S16. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
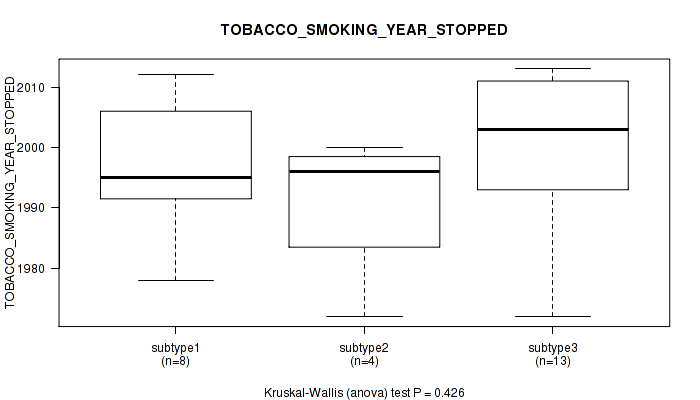
P value = 0.454 (Kruskal-Wallis (anova)), Q value = 1
Table S18. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 53 | 18.6 (12.4) |
| subtype1 | 15 | 18.1 (15.0) |
| subtype2 | 11 | 21.0 (10.2) |
| subtype3 | 27 | 17.9 (11.9) |
Figure S17. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'

P value = 0.894 (Fisher's exact test), Q value = 1
Table S19. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
| nPatients | CURRENT REFORMED SMOKER FOR < OR = 15 YEARS | CURRENT REFORMED SMOKER FOR > 15 YEARS | CURRENT REFORMED SMOKER, DURATION NOT SPECIFIED | CURRENT SMOKER | LIFELONG NON-SMOKER |
|---|---|---|---|---|---|
| ALL | 21 | 8 | 1 | 36 | 83 |
| subtype1 | 7 | 4 | 1 | 10 | 26 |
| subtype2 | 3 | 1 | 0 | 8 | 14 |
| subtype3 | 11 | 3 | 0 | 18 | 43 |
Figure S18. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'

P value = 0.386 (Kruskal-Wallis (anova)), Q value = 1
Table S20. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 48 | 20.5 (6.3) |
| subtype1 | 14 | 21.9 (7.1) |
| subtype2 | 10 | 18.6 (5.0) |
| subtype3 | 24 | 20.5 (6.3) |
Figure S19. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'

P value = 0.211 (Fisher's exact test), Q value = 1
Table S21. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
| nPatients | COMBINATION | EXTERNAL | EXTERNAL BEAM | INTERNAL |
|---|---|---|---|---|
| ALL | 17 | 37 | 11 | 5 |
| subtype1 | 6 | 12 | 3 | 0 |
| subtype2 | 4 | 6 | 3 | 4 |
| subtype3 | 7 | 19 | 5 | 1 |
Figure S20. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'

P value = 1 (Fisher's exact test), Q value = 1
Table S22. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
| nPatients | CGY | GY |
|---|---|---|
| ALL | 11 | 3 |
| subtype1 | 2 | 0 |
| subtype2 | 5 | 1 |
| subtype3 | 4 | 2 |
Figure S21. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'

P value = 0.606 (Kruskal-Wallis (anova)), Q value = 1
Table S23. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 152 | 3.4 (2.4) |
| subtype1 | 46 | 3.7 (2.6) |
| subtype2 | 34 | 3.5 (2.8) |
| subtype3 | 72 | 3.2 (2.1) |
Figure S22. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'

P value = 0.221 (Kruskal-Wallis (anova)), Q value = 1
Table S24. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 93 | 0.1 (0.4) |
| subtype1 | 28 | 0.1 (0.6) |
| subtype2 | 19 | 0.0 (0.0) |
| subtype3 | 46 | 0.1 (0.3) |
Figure S23. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'

P value = 0.928 (Kruskal-Wallis (anova)), Q value = 1
Table S25. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 105 | 0.3 (0.6) |
| subtype1 | 31 | 0.3 (0.5) |
| subtype2 | 21 | 0.3 (0.6) |
| subtype3 | 53 | 0.3 (0.6) |
Figure S24. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'

P value = 0.528 (Kruskal-Wallis (anova)), Q value = 1
Table S26. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 157 | 2.4 (1.7) |
| subtype1 | 48 | 2.4 (1.3) |
| subtype2 | 35 | 2.8 (2.4) |
| subtype3 | 74 | 2.2 (1.6) |
Figure S25. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'

P value = 0.883 (Kruskal-Wallis (anova)), Q value = 1
Table S27. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 100 | 0.8 (1.7) |
| subtype1 | 28 | 1.3 (2.8) |
| subtype2 | 22 | 0.6 (1.0) |
| subtype3 | 50 | 0.7 (1.1) |
Figure S26. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
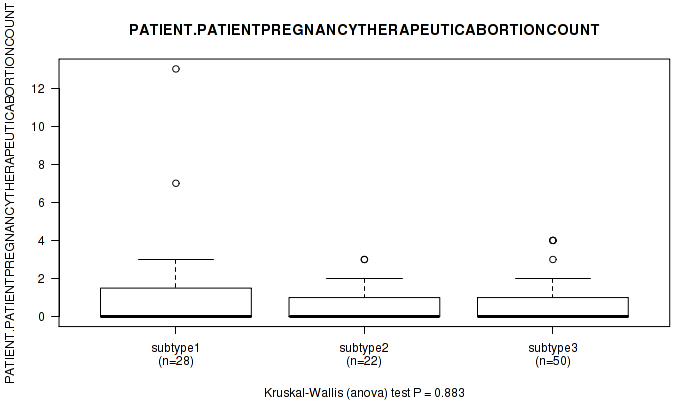
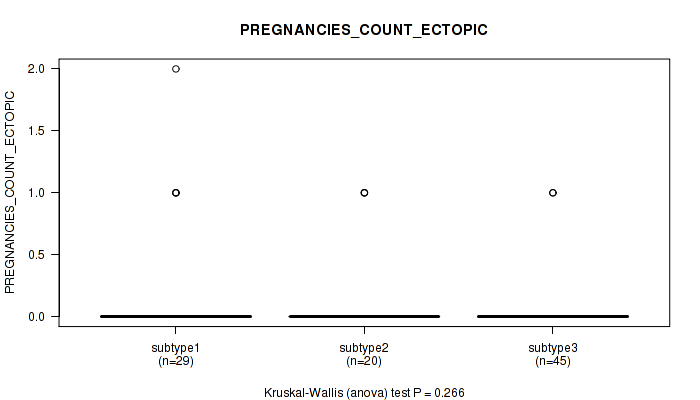
P value = 0.266 (Kruskal-Wallis (anova)), Q value = 1
Table S28. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 94 | 0.1 (0.3) |
| subtype1 | 29 | 0.2 (0.5) |
| subtype2 | 20 | 0.1 (0.4) |
| subtype3 | 45 | 0.0 (0.2) |
Figure S27. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'
P value = 0.363 (Fisher's exact test), Q value = 1
Table S29. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
| nPatients | 2003 | COMMON ILIAC | PELVIC (EXTERNAL ILIAC, INTERNAL ILIAC, OBTURATOR) |
|---|---|---|---|
| ALL | 1 | 1 | 38 |
| subtype1 | 0 | 0 | 15 |
| subtype2 | 1 | 0 | 7 |
| subtype3 | 0 | 1 | 16 |
Figure S28. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'

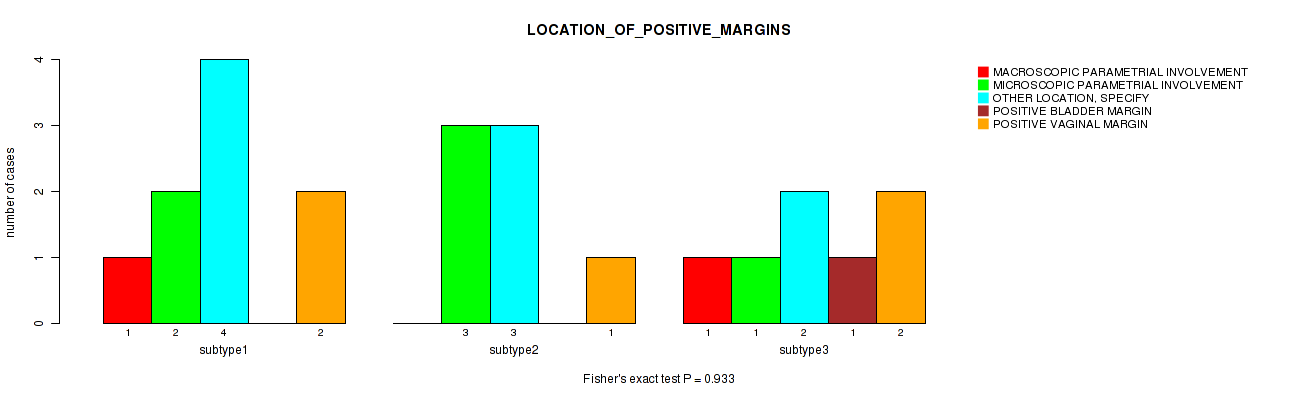
P value = 0.933 (Fisher's exact test), Q value = 1
Table S30. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'
| nPatients | MACROSCOPIC PARAMETRIAL INVOLVEMENT | MICROSCOPIC PARAMETRIAL INVOLVEMENT | OTHER LOCATION, SPECIFY | POSITIVE BLADDER MARGIN | POSITIVE VAGINAL MARGIN |
|---|---|---|---|---|---|
| ALL | 2 | 6 | 9 | 1 | 5 |
| subtype1 | 1 | 2 | 4 | 0 | 2 |
| subtype2 | 0 | 3 | 3 | 0 | 1 |
| subtype3 | 1 | 1 | 2 | 1 | 2 |
Figure S29. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'
P value = 0.326 (Fisher's exact test), Q value = 1
Table S31. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
| nPatients | INDETERMINATE (NEITHER PRE OR POSTMENOPAUSAL) | PERI (6-12 MONTHS SINCE LAST MENSTRUAL PERIOD) | POST (PRIOR BILATERAL OVARIECTOMY OR >12 MO SINCE LMP WITH NO PRIOR HYSTERECTOMY) | PRE (<6 MONTHS SINCE LMP AND NO PRIOR BILATERAL OVARIECTOMY AND NOT ON ESTROGEN REPLACEMENT) |
|---|---|---|---|---|
| ALL | 2 | 10 | 54 | 77 |
| subtype1 | 2 | 5 | 19 | 20 |
| subtype2 | 0 | 1 | 12 | 16 |
| subtype3 | 0 | 4 | 23 | 41 |
Figure S30. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #30: 'MENOPAUSE_STATUS'

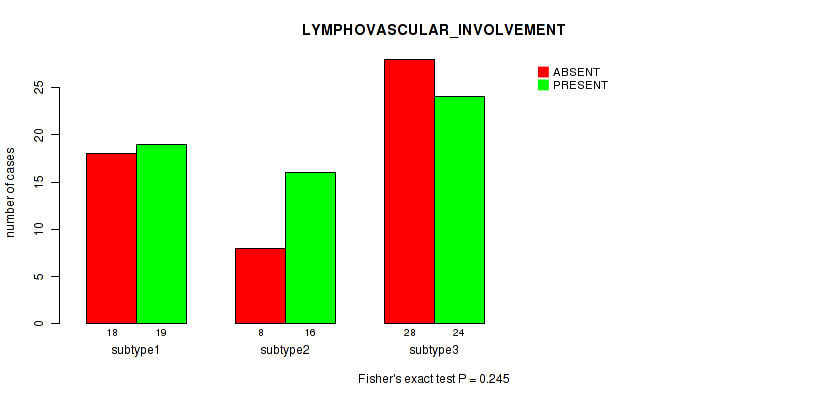
P value = 0.245 (Fisher's exact test), Q value = 1
Table S32. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 54 | 59 |
| subtype1 | 18 | 19 |
| subtype2 | 8 | 16 |
| subtype3 | 28 | 24 |
Figure S31. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'
P value = 0.166 (Kruskal-Wallis (anova)), Q value = 1
Table S33. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 107 | 0.9 (2.2) |
| subtype1 | 31 | 0.8 (1.3) |
| subtype2 | 19 | 0.9 (1.3) |
| subtype3 | 57 | 1.0 (2.8) |
Figure S32. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'

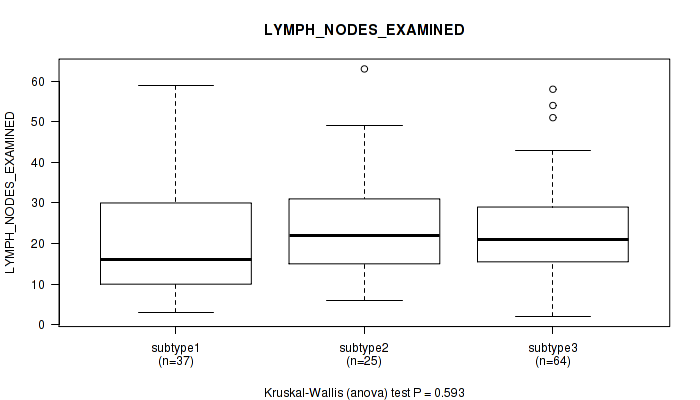
P value = 0.593 (Kruskal-Wallis (anova)), Q value = 1
Table S34. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 126 | 22.2 (13.0) |
| subtype1 | 37 | 21.1 (14.2) |
| subtype2 | 25 | 24.1 (14.0) |
| subtype3 | 64 | 22.2 (11.9) |
Figure S33. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
P value = 0.628 (Fisher's exact test), Q value = 1
Table S35. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'
| nPatients | KERATINIZING SQUAMOUS CELL CARCINOMA | NON-KERATINIZING SQUAMOUS CELL CARCINOMA |
|---|---|---|
| ALL | 35 | 81 |
| subtype1 | 12 | 26 |
| subtype2 | 9 | 16 |
| subtype3 | 14 | 39 |
Figure S34. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'

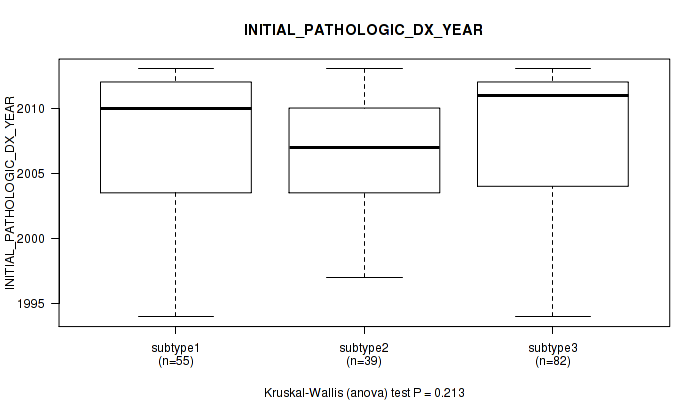
P value = 0.213 (Kruskal-Wallis (anova)), Q value = 1
Table S36. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 176 | 2007.6 (5.1) |
| subtype1 | 55 | 2007.5 (5.3) |
| subtype2 | 39 | 2006.9 (4.4) |
| subtype3 | 82 | 2008.0 (5.2) |
Figure S35. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
P value = 0.099 (Fisher's exact test), Q value = 1
Table S37. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
| nPatients | OTHER | RADICAL HYSTERECTOMY | SIMPLE HYSTERECTOMY |
|---|---|---|---|
| ALL | 3 | 113 | 5 |
| subtype1 | 0 | 37 | 0 |
| subtype2 | 0 | 24 | 0 |
| subtype3 | 3 | 52 | 5 |
Figure S36. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'

P value = 0.404 (Fisher's exact test), Q value = 1
Table S38. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'
| nPatients | CURRENT USER | FORMER USER | NEVER USED |
|---|---|---|---|
| ALL | 8 | 40 | 40 |
| subtype1 | 2 | 10 | 16 |
| subtype2 | 3 | 11 | 6 |
| subtype3 | 3 | 19 | 18 |
Figure S37. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'

P value = 0.344 (Kruskal-Wallis (anova)), Q value = 1
Table S39. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 156 | 162.3 (6.7) |
| subtype1 | 50 | 161.4 (6.3) |
| subtype2 | 35 | 163.2 (5.8) |
| subtype3 | 71 | 162.5 (7.2) |
Figure S38. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'

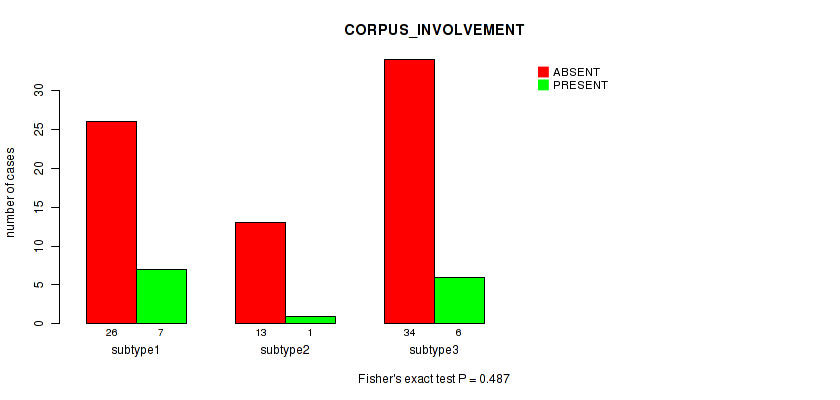
P value = 0.487 (Fisher's exact test), Q value = 1
Table S40. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 73 | 14 |
| subtype1 | 26 | 7 |
| subtype2 | 13 | 1 |
| subtype3 | 34 | 6 |
Figure S39. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
P value = 0.535 (Fisher's exact test), Q value = 1
Table S41. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'
| nPatients | CARBOPLATIN | CISPLATIN |
|---|---|---|
| ALL | 1 | 14 |
| subtype1 | 0 | 3 |
| subtype2 | 0 | 7 |
| subtype3 | 1 | 4 |
Figure S40. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'

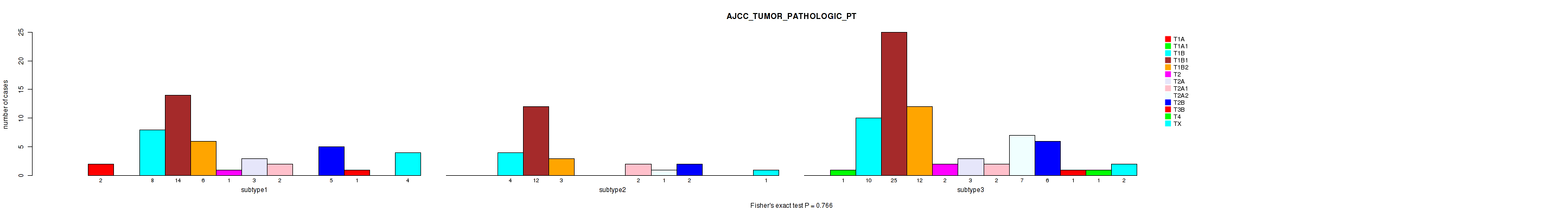
P value = 0.766 (Fisher's exact test), Q value = 1
Table S42. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'
| nPatients | T1A | T1A1 | T1B | T1B1 | T1B2 | T2 | T2A | T2A1 | T2A2 | T2B | T3B | T4 | TX |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 2 | 1 | 22 | 51 | 21 | 3 | 6 | 6 | 8 | 13 | 2 | 1 | 7 |
| subtype1 | 2 | 0 | 8 | 14 | 6 | 1 | 3 | 2 | 0 | 5 | 1 | 0 | 4 |
| subtype2 | 0 | 0 | 4 | 12 | 3 | 0 | 0 | 2 | 1 | 2 | 0 | 0 | 1 |
| subtype3 | 0 | 1 | 10 | 25 | 12 | 2 | 3 | 2 | 7 | 6 | 1 | 1 | 2 |
Figure S41. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'
P value = 0.112 (Kruskal-Wallis (anova)), Q value = 1
Table S43. Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 176 | 47.6 (13.1) |
| subtype1 | 55 | 49.7 (10.5) |
| subtype2 | 39 | 46.7 (15.0) |
| subtype3 | 82 | 46.5 (13.7) |
Figure S42. Get High-res Image Clustering Approach #1: 'Copy Number Ratio CNMF subtypes' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'

Table S44. Description of clustering approach #2: 'METHLYATION CNMF'
| Cluster Labels | 1 | 2 | 3 | 4 | 5 | 6 |
|---|---|---|---|---|---|---|
| Number of samples | 45 | 32 | 38 | 42 | 31 | 12 |
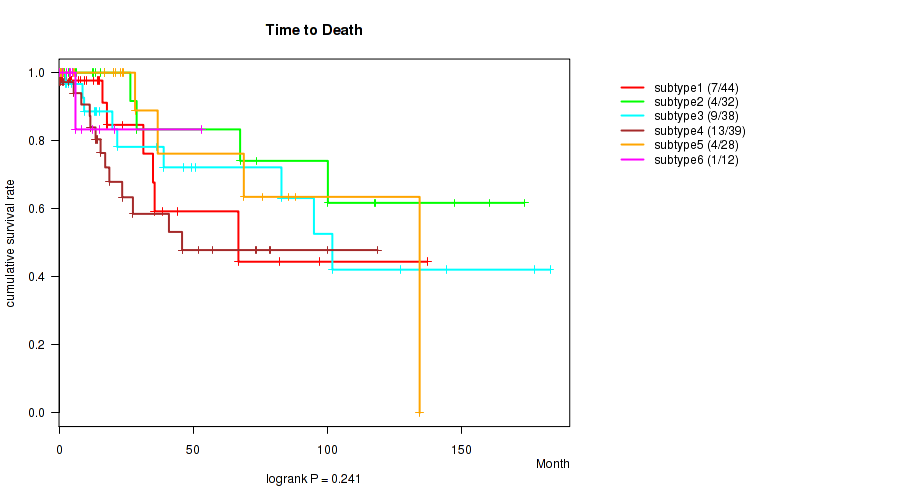
P value = 0.241 (logrank test), Q value = 1
Table S45. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 193 | 38 | 0.0 - 182.9 (13.9) |
| subtype1 | 44 | 7 | 0.1 - 137.2 (8.8) |
| subtype2 | 32 | 4 | 0.1 - 173.3 (13.1) |
| subtype3 | 38 | 9 | 0.0 - 182.9 (14.4) |
| subtype4 | 39 | 13 | 0.0 - 118.7 (15.5) |
| subtype5 | 28 | 4 | 0.0 - 134.3 (20.7) |
| subtype6 | 12 | 1 | 1.4 - 53.2 (6.0) |
Figure S43. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #1: 'Time to Death'
P value = 0.0117 (Kruskal-Wallis (anova)), Q value = 1
Table S46. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 199 | 47.5 (13.4) |
| subtype1 | 45 | 44.5 (10.5) |
| subtype2 | 32 | 48.7 (14.6) |
| subtype3 | 38 | 48.1 (14.1) |
| subtype4 | 41 | 42.7 (12.0) |
| subtype5 | 31 | 54.3 (13.5) |
| subtype6 | 12 | 52.1 (14.6) |
Figure S44. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #2: 'AGE'

P value = 0.983 (Fisher's exact test), Q value = 1
Table S47. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
| nPatients | T1 | T2 | T3+T4 |
|---|---|---|---|
| ALL | 108 | 39 | 4 |
| subtype1 | 25 | 12 | 1 |
| subtype2 | 18 | 5 | 0 |
| subtype3 | 23 | 9 | 1 |
| subtype4 | 18 | 7 | 1 |
| subtype5 | 18 | 4 | 1 |
| subtype6 | 6 | 2 | 0 |
Figure S45. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'

P value = 0.331 (Fisher's exact test), Q value = 1
Table S48. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 100 | 44 |
| subtype1 | 26 | 9 |
| subtype2 | 11 | 11 |
| subtype3 | 21 | 11 |
| subtype4 | 20 | 5 |
| subtype5 | 16 | 6 |
| subtype6 | 6 | 2 |
Figure S46. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'

P value = 0.716 (Fisher's exact test), Q value = 1
Table S49. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'
| nPatients | M0 | M1 | MX |
|---|---|---|---|
| ALL | 83 | 4 | 70 |
| subtype1 | 20 | 3 | 15 |
| subtype2 | 13 | 0 | 12 |
| subtype3 | 19 | 0 | 14 |
| subtype4 | 16 | 1 | 11 |
| subtype5 | 12 | 0 | 12 |
| subtype6 | 3 | 0 | 6 |
Figure S47. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'

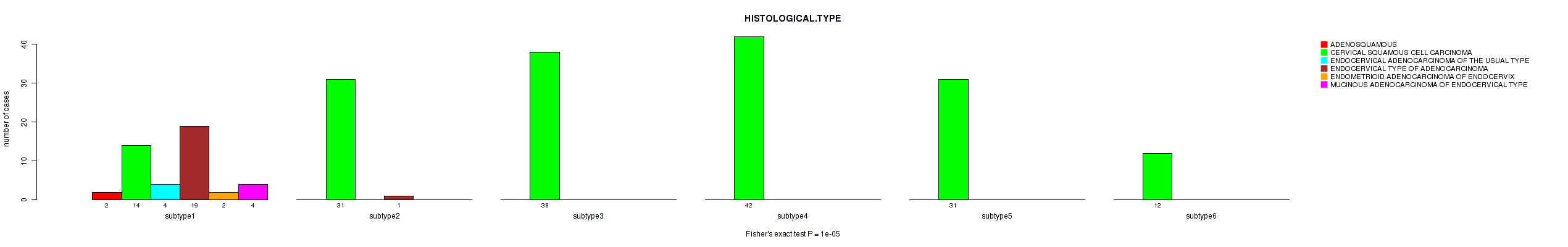
P value = 1e-05 (Fisher's exact test), Q value = 0.0034
Table S50. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
| nPatients | ADENOSQUAMOUS | CERVICAL SQUAMOUS CELL CARCINOMA | ENDOCERVICAL ADENOCARCINOMA OF THE USUAL TYPE | ENDOCERVICAL TYPE OF ADENOCARCINOMA | ENDOMETRIOID ADENOCARCINOMA OF ENDOCERVIX | MUCINOUS ADENOCARCINOMA OF ENDOCERVICAL TYPE |
|---|---|---|---|---|---|---|
| ALL | 2 | 168 | 4 | 20 | 2 | 4 |
| subtype1 | 2 | 14 | 4 | 19 | 2 | 4 |
| subtype2 | 0 | 31 | 0 | 1 | 0 | 0 |
| subtype3 | 0 | 38 | 0 | 0 | 0 | 0 |
| subtype4 | 0 | 42 | 0 | 0 | 0 | 0 |
| subtype5 | 0 | 31 | 0 | 0 | 0 | 0 |
| subtype6 | 0 | 12 | 0 | 0 | 0 | 0 |
Figure S48. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
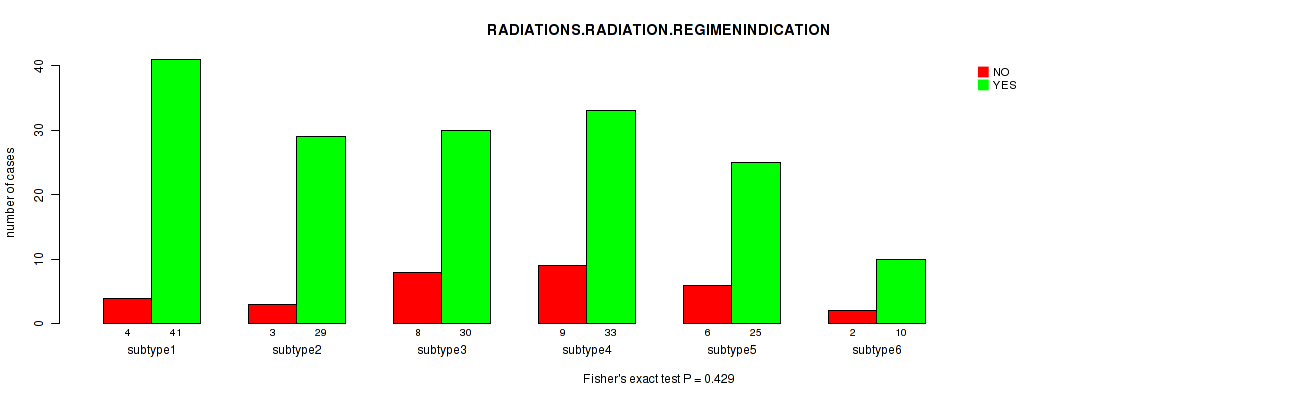
P value = 0.429 (Fisher's exact test), Q value = 1
Table S51. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 32 | 168 |
| subtype1 | 4 | 41 |
| subtype2 | 3 | 29 |
| subtype3 | 8 | 30 |
| subtype4 | 9 | 33 |
| subtype5 | 6 | 25 |
| subtype6 | 2 | 10 |
Figure S49. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'
P value = 0.544 (Kruskal-Wallis (anova)), Q value = 1
Table S52. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 62 | 19.4 (13.6) |
| subtype1 | 13 | 15.0 (11.8) |
| subtype2 | 11 | 21.5 (18.5) |
| subtype3 | 12 | 16.1 (7.6) |
| subtype4 | 12 | 21.3 (13.4) |
| subtype5 | 10 | 22.8 (12.4) |
| subtype6 | 4 | 24.5 (22.3) |
Figure S50. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'

P value = 0.513 (Kruskal-Wallis (anova)), Q value = 1
Table S53. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 123 | 1.1 (2.6) |
| subtype1 | 29 | 1.2 (3.3) |
| subtype2 | 19 | 1.4 (2.0) |
| subtype3 | 23 | 1.0 (2.1) |
| subtype4 | 21 | 0.7 (1.5) |
| subtype5 | 23 | 1.3 (3.5) |
| subtype6 | 8 | 0.4 (0.7) |
Figure S51. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'

P value = 0.287 (Fisher's exact test), Q value = 1
Table S54. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #10: 'RACE'
| nPatients | AMERICAN INDIAN OR ALASKA NATIVE | ASIAN | BLACK OR AFRICAN AMERICAN | NATIVE HAWAIIAN OR OTHER PACIFIC ISLANDER | WHITE |
|---|---|---|---|---|---|
| ALL | 7 | 16 | 23 | 1 | 148 |
| subtype1 | 0 | 3 | 3 | 0 | 38 |
| subtype2 | 1 | 3 | 3 | 0 | 24 |
| subtype3 | 1 | 6 | 6 | 0 | 25 |
| subtype4 | 4 | 3 | 6 | 1 | 28 |
| subtype5 | 1 | 0 | 2 | 0 | 25 |
| subtype6 | 0 | 1 | 3 | 0 | 8 |
Figure S52. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #10: 'RACE'

P value = 0.164 (Fisher's exact test), Q value = 1
Table S55. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #11: 'ETHNICITY'
| nPatients | HISPANIC OR LATINO | NOT HISPANIC OR LATINO |
|---|---|---|
| ALL | 13 | 144 |
| subtype1 | 5 | 32 |
| subtype2 | 2 | 22 |
| subtype3 | 0 | 33 |
| subtype4 | 5 | 28 |
| subtype5 | 1 | 20 |
| subtype6 | 0 | 9 |
Figure S53. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #11: 'ETHNICITY'

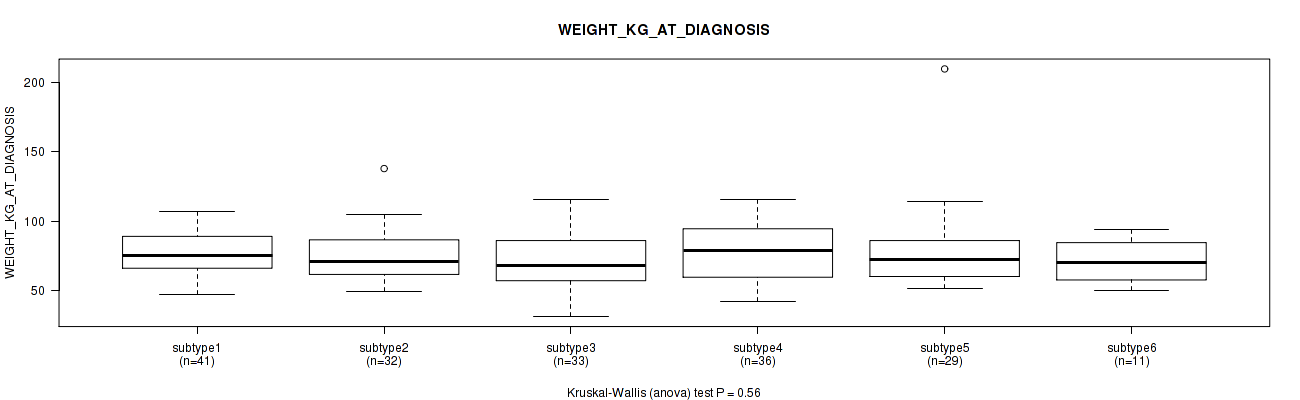
P value = 0.56 (Kruskal-Wallis (anova)), Q value = 1
Table S56. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 182 | 75.2 (20.9) |
| subtype1 | 41 | 75.7 (15.3) |
| subtype2 | 32 | 74.8 (19.3) |
| subtype3 | 33 | 71.3 (21.8) |
| subtype4 | 36 | 78.5 (19.5) |
| subtype5 | 29 | 77.0 (30.5) |
| subtype6 | 11 | 70.6 (15.0) |
Figure S54. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'
P value = 0.795 (Fisher's exact test), Q value = 1
Table S57. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #13: 'TUMOR_STATUS'
| nPatients | TUMOR FREE | WITH TUMOR |
|---|---|---|
| ALL | 51 | 22 |
| subtype1 | 9 | 6 |
| subtype2 | 8 | 4 |
| subtype3 | 12 | 4 |
| subtype4 | 12 | 6 |
| subtype5 | 6 | 2 |
| subtype6 | 4 | 0 |
Figure S55. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #13: 'TUMOR_STATUS'

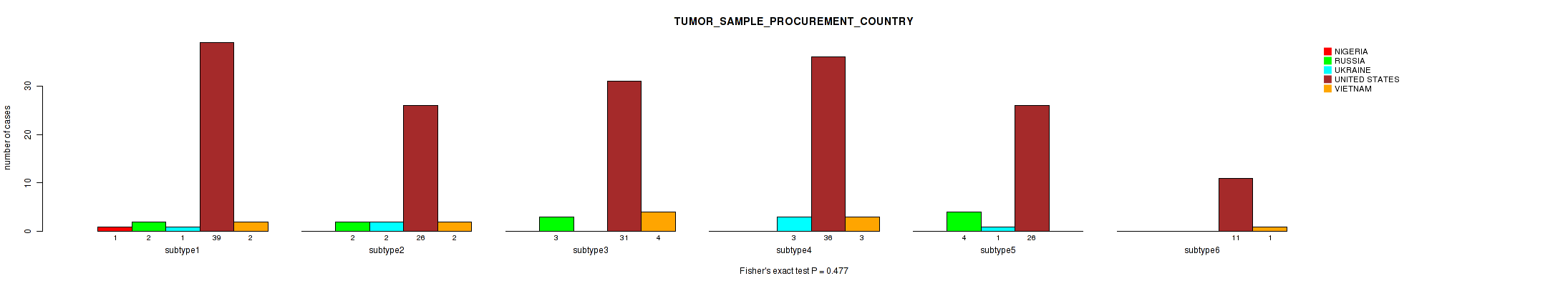
P value = 0.477 (Fisher's exact test), Q value = 1
Table S58. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
| nPatients | NIGERIA | RUSSIA | UKRAINE | UNITED STATES | VIETNAM |
|---|---|---|---|---|---|
| ALL | 1 | 11 | 7 | 169 | 12 |
| subtype1 | 1 | 2 | 1 | 39 | 2 |
| subtype2 | 0 | 2 | 2 | 26 | 2 |
| subtype3 | 0 | 3 | 0 | 31 | 4 |
| subtype4 | 0 | 0 | 3 | 36 | 3 |
| subtype5 | 0 | 4 | 1 | 26 | 0 |
| subtype6 | 0 | 0 | 0 | 11 | 1 |
Figure S56. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
P value = 0.74 (Fisher's exact test), Q value = 1
Table S59. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'
| nPatients | G1 | G2 | G3 | G4 | GX |
|---|---|---|---|---|---|
| ALL | 14 | 94 | 83 | 1 | 5 |
| subtype1 | 6 | 20 | 18 | 0 | 1 |
| subtype2 | 1 | 15 | 15 | 0 | 1 |
| subtype3 | 2 | 20 | 16 | 0 | 0 |
| subtype4 | 1 | 17 | 20 | 1 | 1 |
| subtype5 | 3 | 17 | 9 | 0 | 1 |
| subtype6 | 1 | 5 | 5 | 0 | 1 |
Figure S57. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'

P value = 0.355 (Kruskal-Wallis (anova)), Q value = 1
Table S60. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 27 | 1998.4 (11.5) |
| subtype1 | 4 | 1993.8 (14.8) |
| subtype2 | 4 | 1990.2 (16.0) |
| subtype3 | 5 | 1997.8 (3.3) |
| subtype4 | 4 | 2002.0 (5.6) |
| subtype5 | 6 | 1999.2 (14.0) |
| subtype6 | 4 | 2007.0 (8.3) |
Figure S58. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'

P value = 0.544 (Kruskal-Wallis (anova)), Q value = 1
Table S61. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 62 | 19.4 (13.6) |
| subtype1 | 13 | 15.0 (11.8) |
| subtype2 | 11 | 21.5 (18.5) |
| subtype3 | 12 | 16.1 (7.6) |
| subtype4 | 12 | 21.3 (13.4) |
| subtype5 | 10 | 22.8 (12.4) |
| subtype6 | 4 | 24.5 (22.3) |
Figure S59. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'

P value = 0.539 (Fisher's exact test), Q value = 1
Table S62. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
| nPatients | CURRENT REFORMED SMOKER FOR < OR = 15 YEARS | CURRENT REFORMED SMOKER FOR > 15 YEARS | CURRENT REFORMED SMOKER, DURATION NOT SPECIFIED | CURRENT SMOKER | LIFELONG NON-SMOKER |
|---|---|---|---|---|---|
| ALL | 25 | 8 | 3 | 43 | 89 |
| subtype1 | 3 | 1 | 0 | 10 | 25 |
| subtype2 | 3 | 2 | 1 | 6 | 16 |
| subtype3 | 5 | 2 | 0 | 8 | 19 |
| subtype4 | 5 | 0 | 1 | 10 | 16 |
| subtype5 | 5 | 3 | 1 | 6 | 9 |
| subtype6 | 4 | 0 | 0 | 3 | 4 |
Figure S60. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'

P value = 0.549 (Kruskal-Wallis (anova)), Q value = 1
Table S63. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 54 | 20.5 (6.2) |
| subtype1 | 12 | 18.7 (4.2) |
| subtype2 | 10 | 19.3 (6.7) |
| subtype3 | 9 | 19.9 (5.8) |
| subtype4 | 10 | 20.3 (5.2) |
| subtype5 | 9 | 24.7 (8.5) |
| subtype6 | 4 | 21.0 (6.2) |
Figure S61. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'

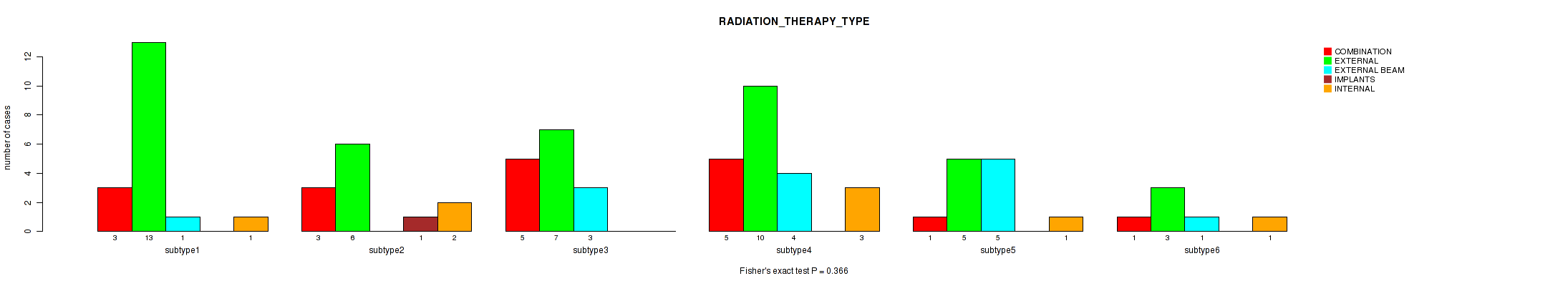
P value = 0.366 (Fisher's exact test), Q value = 1
Table S64. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
| nPatients | COMBINATION | EXTERNAL | EXTERNAL BEAM | IMPLANTS | INTERNAL |
|---|---|---|---|---|---|
| ALL | 18 | 44 | 14 | 1 | 8 |
| subtype1 | 3 | 13 | 1 | 0 | 1 |
| subtype2 | 3 | 6 | 0 | 1 | 2 |
| subtype3 | 5 | 7 | 3 | 0 | 0 |
| subtype4 | 5 | 10 | 4 | 0 | 3 |
| subtype5 | 1 | 5 | 5 | 0 | 1 |
| subtype6 | 1 | 3 | 1 | 0 | 1 |
Figure S62. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
P value = 0.84 (Fisher's exact test), Q value = 1
Table S65. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
| nPatients | CGY | GY |
|---|---|---|
| ALL | 16 | 4 |
| subtype1 | 4 | 0 |
| subtype2 | 1 | 0 |
| subtype3 | 3 | 1 |
| subtype4 | 5 | 2 |
| subtype5 | 2 | 0 |
| subtype6 | 1 | 1 |
Figure S63. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'

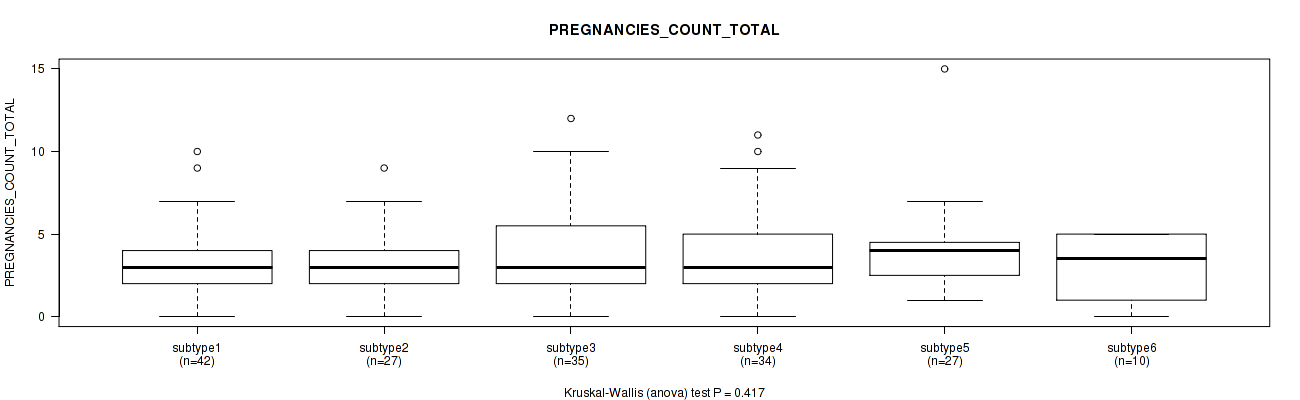
P value = 0.417 (Kruskal-Wallis (anova)), Q value = 1
Table S66. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 175 | 3.5 (2.4) |
| subtype1 | 42 | 3.0 (2.2) |
| subtype2 | 27 | 3.2 (2.1) |
| subtype3 | 35 | 3.8 (2.6) |
| subtype4 | 34 | 3.9 (2.7) |
| subtype5 | 27 | 3.9 (2.7) |
| subtype6 | 10 | 3.0 (2.1) |
Figure S64. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
P value = 0.00866 (Kruskal-Wallis (anova)), Q value = 1
Table S67. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 100 | 0.1 (0.4) |
| subtype1 | 22 | 0.0 (0.2) |
| subtype2 | 20 | 0.0 (0.0) |
| subtype3 | 21 | 0.3 (0.7) |
| subtype4 | 20 | 0.0 (0.0) |
| subtype5 | 13 | 0.0 (0.0) |
| subtype6 | 4 | 0.0 (0.0) |
Figure S65. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'

P value = 0.206 (Kruskal-Wallis (anova)), Q value = 1
Table S68. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 116 | 0.4 (0.6) |
| subtype1 | 23 | 0.5 (0.6) |
| subtype2 | 21 | 0.1 (0.3) |
| subtype3 | 27 | 0.5 (0.8) |
| subtype4 | 24 | 0.4 (0.6) |
| subtype5 | 16 | 0.2 (0.4) |
| subtype6 | 5 | 0.4 (0.5) |
Figure S66. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'

P value = 0.258 (Kruskal-Wallis (anova)), Q value = 1
Table S69. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 179 | 2.5 (1.8) |
| subtype1 | 41 | 2.1 (1.9) |
| subtype2 | 28 | 2.9 (1.9) |
| subtype3 | 37 | 2.5 (1.7) |
| subtype4 | 34 | 2.8 (1.9) |
| subtype5 | 28 | 2.3 (1.3) |
| subtype6 | 11 | 2.5 (1.6) |
Figure S67. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'

P value = 0.0424 (Kruskal-Wallis (anova)), Q value = 1
Table S70. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 109 | 0.9 (1.9) |
| subtype1 | 24 | 0.8 (1.0) |
| subtype2 | 21 | 0.3 (0.9) |
| subtype3 | 23 | 1.1 (2.3) |
| subtype4 | 22 | 0.9 (1.6) |
| subtype5 | 15 | 1.9 (3.3) |
| subtype6 | 4 | 0.0 (0.0) |
Figure S68. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'

P value = 0.777 (Kruskal-Wallis (anova)), Q value = 1
Table S71. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 101 | 0.1 (0.3) |
| subtype1 | 21 | 0.1 (0.3) |
| subtype2 | 21 | 0.0 (0.2) |
| subtype3 | 21 | 0.0 (0.2) |
| subtype4 | 20 | 0.1 (0.4) |
| subtype5 | 13 | 0.2 (0.6) |
| subtype6 | 5 | 0.2 (0.4) |
Figure S69. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'

P value = 0.593 (Fisher's exact test), Q value = 1
Table S72. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
| nPatients | 2003 | 2010 | COMMON ILIAC | PELVIC (EXTERNAL ILIAC, INTERNAL ILIAC, OBTURATOR) |
|---|---|---|---|---|
| ALL | 1 | 1 | 1 | 41 |
| subtype1 | 0 | 0 | 1 | 8 |
| subtype2 | 1 | 0 | 0 | 8 |
| subtype3 | 0 | 0 | 0 | 8 |
| subtype4 | 0 | 1 | 0 | 5 |
| subtype5 | 0 | 0 | 0 | 8 |
| subtype6 | 0 | 0 | 0 | 4 |
Figure S70. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'

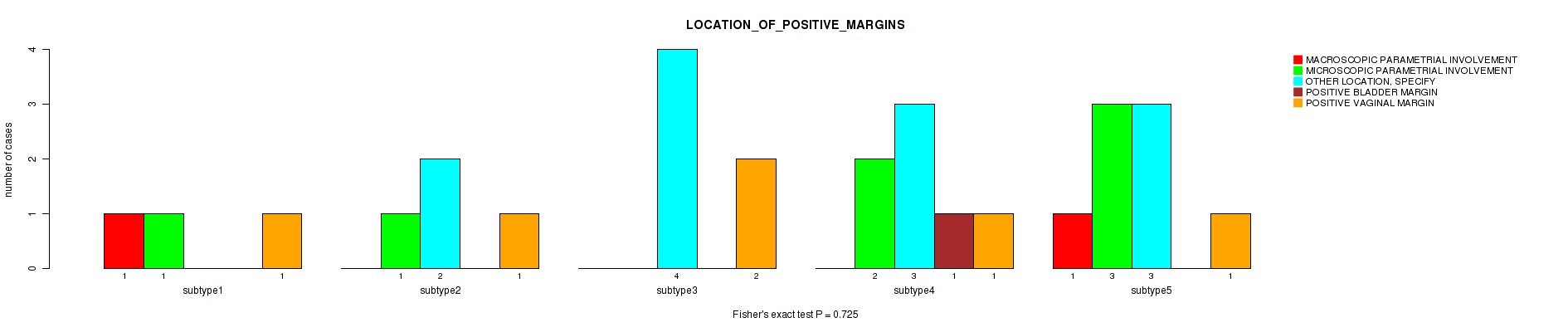
P value = 0.725 (Fisher's exact test), Q value = 1
Table S73. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'
| nPatients | MACROSCOPIC PARAMETRIAL INVOLVEMENT | MICROSCOPIC PARAMETRIAL INVOLVEMENT | OTHER LOCATION, SPECIFY | POSITIVE BLADDER MARGIN | POSITIVE VAGINAL MARGIN |
|---|---|---|---|---|---|
| ALL | 2 | 7 | 13 | 1 | 6 |
| subtype1 | 1 | 1 | 0 | 0 | 1 |
| subtype2 | 0 | 1 | 2 | 0 | 1 |
| subtype3 | 0 | 0 | 4 | 0 | 2 |
| subtype4 | 0 | 2 | 3 | 1 | 1 |
| subtype5 | 1 | 3 | 3 | 0 | 1 |
| subtype6 | 0 | 0 | 1 | 0 | 0 |
Figure S71. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'
P value = 0.0169 (Fisher's exact test), Q value = 1
Table S74. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
| nPatients | INDETERMINATE (NEITHER PRE OR POSTMENOPAUSAL) | PERI (6-12 MONTHS SINCE LAST MENSTRUAL PERIOD) | POST (PRIOR BILATERAL OVARIECTOMY OR >12 MO SINCE LMP WITH NO PRIOR HYSTERECTOMY) | PRE (<6 MONTHS SINCE LMP AND NO PRIOR BILATERAL OVARIECTOMY AND NOT ON ESTROGEN REPLACEMENT) |
|---|---|---|---|---|
| ALL | 2 | 10 | 58 | 85 |
| subtype1 | 0 | 3 | 10 | 24 |
| subtype2 | 1 | 2 | 12 | 11 |
| subtype3 | 0 | 3 | 14 | 16 |
| subtype4 | 0 | 1 | 3 | 21 |
| subtype5 | 1 | 0 | 13 | 9 |
| subtype6 | 0 | 1 | 6 | 4 |
Figure S72. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #30: 'MENOPAUSE_STATUS'

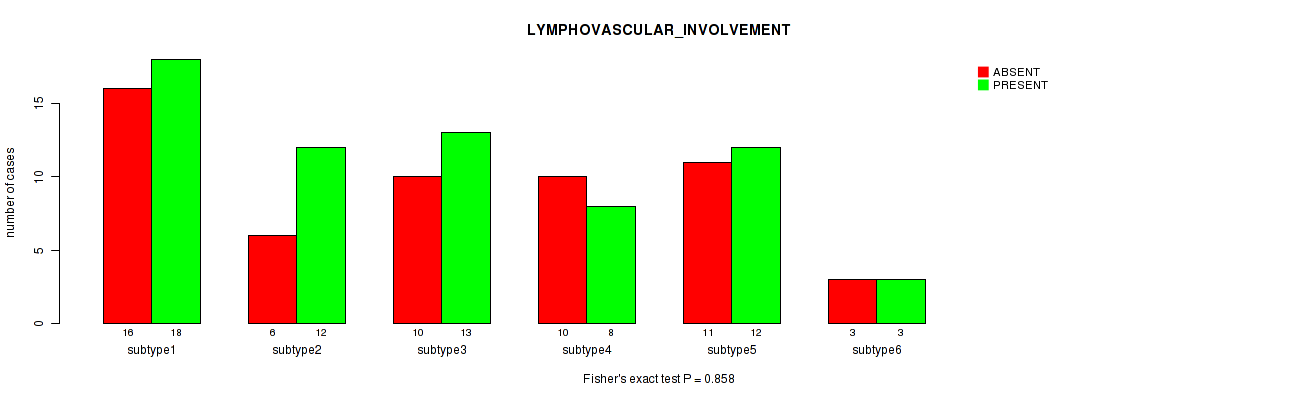
P value = 0.858 (Fisher's exact test), Q value = 1
Table S75. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 56 | 66 |
| subtype1 | 16 | 18 |
| subtype2 | 6 | 12 |
| subtype3 | 10 | 13 |
| subtype4 | 10 | 8 |
| subtype5 | 11 | 12 |
| subtype6 | 3 | 3 |
Figure S73. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'
P value = 0.513 (Kruskal-Wallis (anova)), Q value = 1
Table S76. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 123 | 1.1 (2.6) |
| subtype1 | 29 | 1.2 (3.3) |
| subtype2 | 19 | 1.4 (2.0) |
| subtype3 | 23 | 1.0 (2.1) |
| subtype4 | 21 | 0.7 (1.5) |
| subtype5 | 23 | 1.3 (3.5) |
| subtype6 | 8 | 0.4 (0.7) |
Figure S74. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'

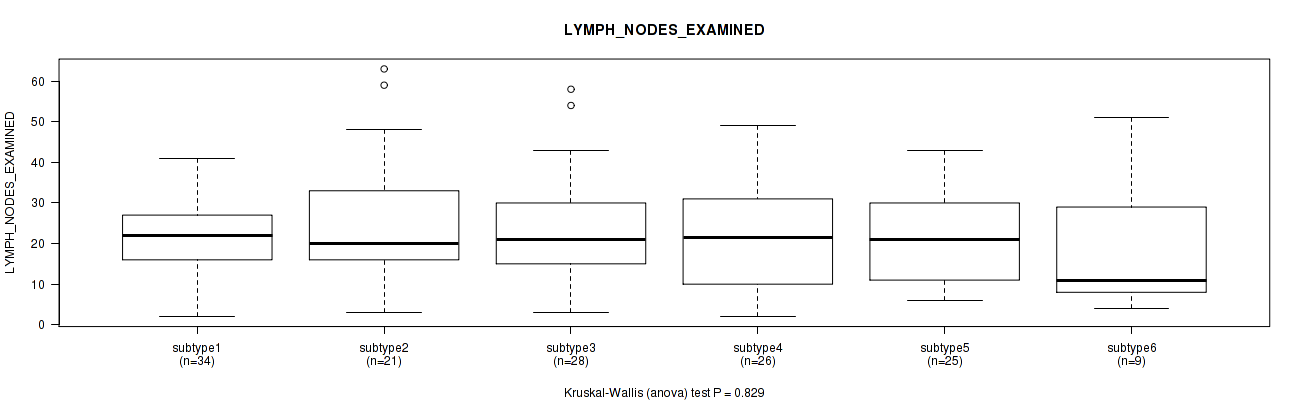
P value = 0.829 (Kruskal-Wallis (anova)), Q value = 1
Table S77. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 143 | 22.4 (12.7) |
| subtype1 | 34 | 21.2 (9.6) |
| subtype2 | 21 | 25.4 (16.2) |
| subtype3 | 28 | 24.3 (13.7) |
| subtype4 | 26 | 21.0 (12.2) |
| subtype5 | 25 | 21.8 (11.0) |
| subtype6 | 9 | 19.2 (16.8) |
Figure S75. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
P value = 0.0149 (Fisher's exact test), Q value = 1
Table S78. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'
| nPatients | KERATINIZING SQUAMOUS CELL CARCINOMA | NON-KERATINIZING SQUAMOUS CELL CARCINOMA |
|---|---|---|
| ALL | 43 | 88 |
| subtype1 | 0 | 13 |
| subtype2 | 5 | 18 |
| subtype3 | 10 | 20 |
| subtype4 | 14 | 17 |
| subtype5 | 12 | 13 |
| subtype6 | 2 | 7 |
Figure S76. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'

P value = 0.34 (Kruskal-Wallis (anova)), Q value = 1
Table S79. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 200 | 2007.3 (5.2) |
| subtype1 | 45 | 2008.4 (4.4) |
| subtype2 | 32 | 2006.8 (6.0) |
| subtype3 | 38 | 2006.3 (5.6) |
| subtype4 | 42 | 2006.6 (5.2) |
| subtype5 | 31 | 2007.5 (5.2) |
| subtype6 | 12 | 2009.2 (3.7) |
Figure S77. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'

P value = 0.285 (Fisher's exact test), Q value = 1
Table S80. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
| nPatients | OTHER | RADICAL HYSTERECTOMY | SIMPLE HYSTERECTOMY |
|---|---|---|---|
| ALL | 3 | 129 | 5 |
| subtype1 | 2 | 31 | 2 |
| subtype2 | 0 | 21 | 0 |
| subtype3 | 0 | 28 | 1 |
| subtype4 | 0 | 20 | 2 |
| subtype5 | 0 | 23 | 0 |
| subtype6 | 1 | 6 | 0 |
Figure S78. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
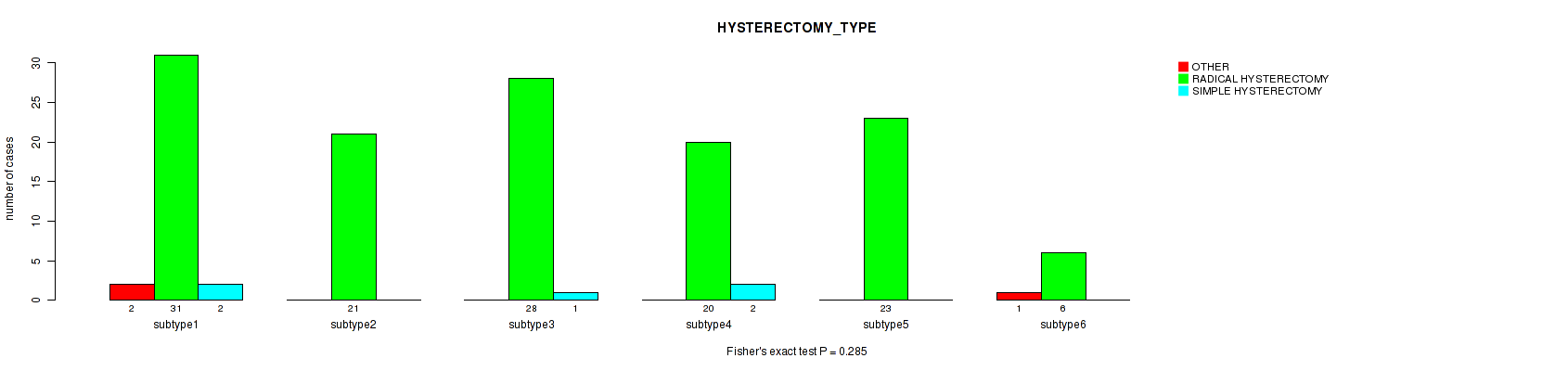
P value = 0.647 (Fisher's exact test), Q value = 1
Table S81. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'
| nPatients | CURRENT USER | FORMER USER | NEVER USED |
|---|---|---|---|
| ALL | 8 | 40 | 47 |
| subtype1 | 5 | 13 | 10 |
| subtype2 | 1 | 7 | 8 |
| subtype3 | 1 | 6 | 10 |
| subtype4 | 1 | 8 | 7 |
| subtype5 | 0 | 3 | 9 |
| subtype6 | 0 | 3 | 3 |
Figure S79. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'

P value = 0.158 (Kruskal-Wallis (anova)), Q value = 1
Table S82. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 173 | 162.1 (7.2) |
| subtype1 | 40 | 163.3 (7.0) |
| subtype2 | 30 | 163.3 (6.9) |
| subtype3 | 32 | 159.1 (8.2) |
| subtype4 | 33 | 161.7 (6.4) |
| subtype5 | 27 | 162.2 (7.2) |
| subtype6 | 11 | 164.1 (6.5) |
Figure S80. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'

P value = 0.536 (Fisher's exact test), Q value = 1
Table S83. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 81 | 15 |
| subtype1 | 24 | 4 |
| subtype2 | 14 | 1 |
| subtype3 | 13 | 4 |
| subtype4 | 13 | 1 |
| subtype5 | 14 | 5 |
| subtype6 | 3 | 0 |
Figure S81. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'

P value = 0.296 (Fisher's exact test), Q value = 1
Table S84. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'
| nPatients | CARBOPLATIN | CISPLATIN |
|---|---|---|
| ALL | 3 | 17 |
| subtype1 | 0 | 2 |
| subtype2 | 1 | 1 |
| subtype3 | 1 | 5 |
| subtype4 | 0 | 7 |
| subtype5 | 1 | 2 |
| subtype6 | 0 | 0 |
Figure S82. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'

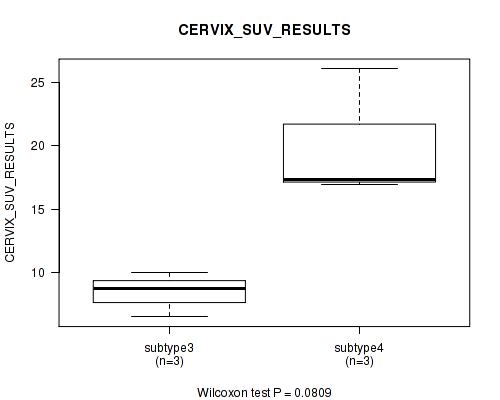
P value = 0.0809 (Kruskal-Wallis (anova)), Q value = 1
Table S85. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #41: 'CERVIX_SUV_RESULTS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 10 | 12.8 (6.1) |
| subtype2 | 1 | 11.1 (NA) |
| subtype3 | 3 | 8.4 (1.8) |
| subtype4 | 3 | 20.1 (5.2) |
| subtype5 | 2 | 9.0 (4.2) |
| subtype6 | 1 | 13.8 (NA) |
Figure S83. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #41: 'CERVIX_SUV_RESULTS'
P value = 0.402 (Fisher's exact test), Q value = 1
Table S86. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'
| nPatients | T1A | T1A1 | T1B | T1B1 | T1B2 | T2 | T2A | T2A1 | T2A2 | T2B | T3B | T4 | TIS | TX |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 2 | 1 | 26 | 55 | 24 | 3 | 6 | 7 | 9 | 14 | 2 | 2 | 1 | 8 |
| subtype1 | 0 | 0 | 1 | 18 | 6 | 0 | 2 | 3 | 3 | 4 | 1 | 0 | 0 | 0 |
| subtype2 | 1 | 0 | 4 | 8 | 5 | 1 | 1 | 0 | 1 | 2 | 0 | 0 | 0 | 3 |
| subtype3 | 1 | 0 | 7 | 13 | 2 | 2 | 1 | 2 | 1 | 3 | 0 | 1 | 0 | 0 |
| subtype4 | 0 | 0 | 8 | 7 | 3 | 0 | 1 | 1 | 2 | 3 | 0 | 1 | 1 | 3 |
| subtype5 | 0 | 1 | 6 | 6 | 5 | 0 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | 1 |
| subtype6 | 0 | 0 | 0 | 3 | 3 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 1 |
Figure S84. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'

P value = 0.0161 (Kruskal-Wallis (anova)), Q value = 1
Table S87. Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 200 | 47.6 (13.4) |
| subtype1 | 45 | 44.5 (10.5) |
| subtype2 | 32 | 48.7 (14.6) |
| subtype3 | 38 | 48.1 (14.1) |
| subtype4 | 42 | 43.2 (12.3) |
| subtype5 | 31 | 54.3 (13.5) |
| subtype6 | 12 | 52.1 (14.6) |
Figure S85. Get High-res Image Clustering Approach #2: 'METHLYATION CNMF' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
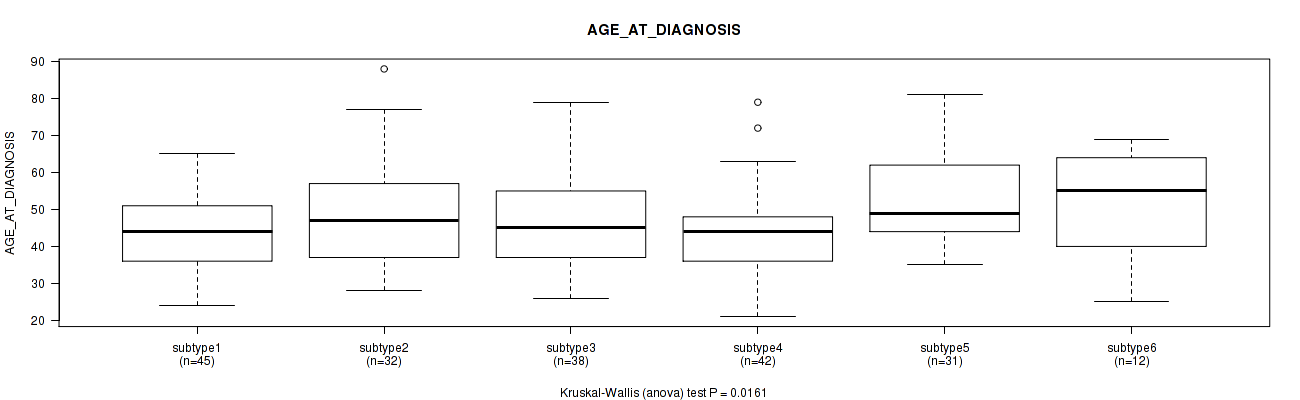
Table S88. Description of clustering approach #3: 'RNAseq CNMF subtypes'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 77 | 47 | 56 |
P value = 0.353 (logrank test), Q value = 1
Table S89. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 173 | 33 | 0.0 - 182.9 (13.6) |
| subtype1 | 75 | 13 | 0.0 - 182.9 (13.9) |
| subtype2 | 44 | 12 | 0.0 - 118.7 (14.5) |
| subtype3 | 54 | 8 | 0.1 - 147.4 (9.9) |
Figure S86. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #1: 'Time to Death'

P value = 0.11 (Kruskal-Wallis (anova)), Q value = 1
Table S90. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 179 | 47.4 (12.9) |
| subtype1 | 77 | 50.1 (13.7) |
| subtype2 | 46 | 45.6 (13.9) |
| subtype3 | 56 | 45.2 (10.1) |
Figure S87. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #2: 'AGE'

P value = 0.723 (Fisher's exact test), Q value = 1
Table S91. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
| nPatients | T1 | T2 | T3+T4 |
|---|---|---|---|
| ALL | 100 | 37 | 3 |
| subtype1 | 40 | 19 | 2 |
| subtype2 | 25 | 7 | 0 |
| subtype3 | 35 | 11 | 1 |
Figure S88. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
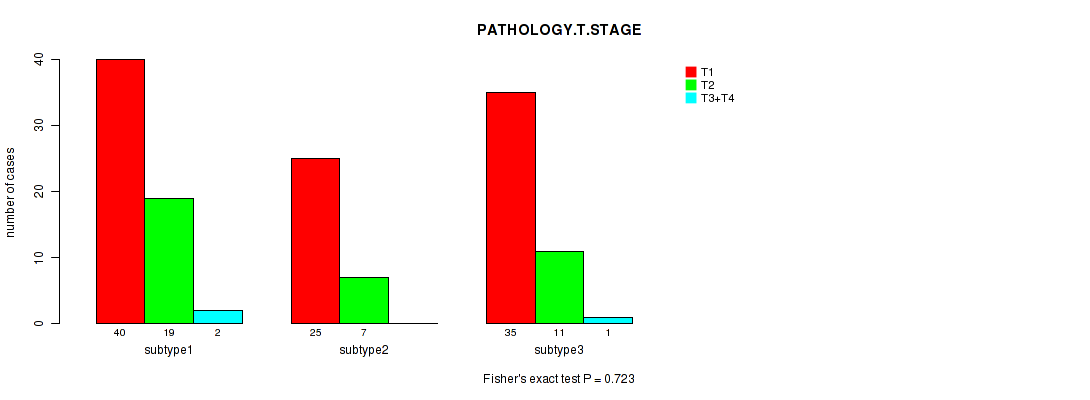
P value = 0.0153 (Fisher's exact test), Q value = 1
Table S92. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 89 | 42 |
| subtype1 | 31 | 26 |
| subtype2 | 23 | 7 |
| subtype3 | 35 | 9 |
Figure S89. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'
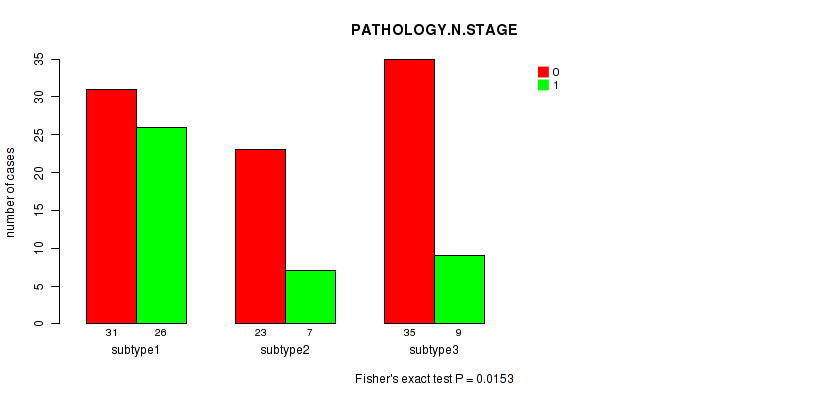
P value = 0.114 (Fisher's exact test), Q value = 1
Table S93. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'
| nPatients | M0 | M1 | MX |
|---|---|---|---|
| ALL | 76 | 3 | 65 |
| subtype1 | 29 | 0 | 33 |
| subtype2 | 20 | 0 | 14 |
| subtype3 | 27 | 3 | 18 |
Figure S90. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'

P value = 1e-05 (Fisher's exact test), Q value = 0.0034
Table S94. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
| nPatients | ADENOSQUAMOUS | CERVICAL SQUAMOUS CELL CARCINOMA | ENDOCERVICAL ADENOCARCINOMA OF THE USUAL TYPE | ENDOCERVICAL TYPE OF ADENOCARCINOMA | ENDOMETRIOID ADENOCARCINOMA OF ENDOCERVIX | MUCINOUS ADENOCARCINOMA OF ENDOCERVICAL TYPE |
|---|---|---|---|---|---|---|
| ALL | 2 | 148 | 4 | 20 | 2 | 4 |
| subtype1 | 0 | 77 | 0 | 0 | 0 | 0 |
| subtype2 | 0 | 47 | 0 | 0 | 0 | 0 |
| subtype3 | 2 | 24 | 4 | 20 | 2 | 4 |
Figure S91. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'

P value = 0.317 (Fisher's exact test), Q value = 1
Table S95. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 30 | 150 |
| subtype1 | 14 | 63 |
| subtype2 | 10 | 37 |
| subtype3 | 6 | 50 |
Figure S92. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'

P value = 0.0633 (Kruskal-Wallis (anova)), Q value = 1
Table S96. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 54 | 18.6 (12.3) |
| subtype1 | 28 | 21.2 (13.3) |
| subtype2 | 11 | 18.8 (9.9) |
| subtype3 | 15 | 13.6 (11.0) |
Figure S93. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'
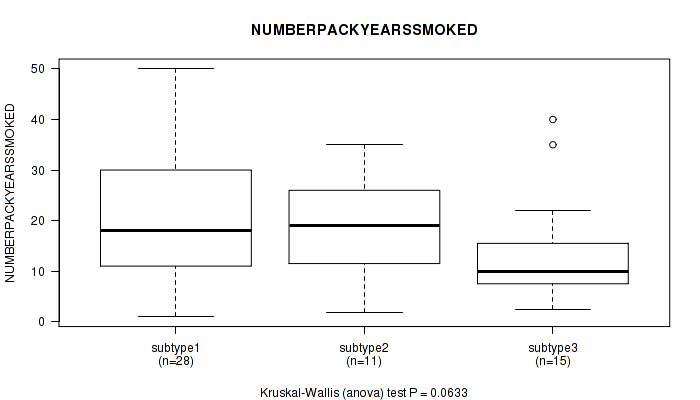
P value = 0.0587 (Kruskal-Wallis (anova)), Q value = 1
Table S97. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 111 | 1.1 (2.6) |
| subtype1 | 49 | 1.2 (1.9) |
| subtype2 | 25 | 1.0 (3.2) |
| subtype3 | 37 | 0.9 (2.9) |
Figure S94. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
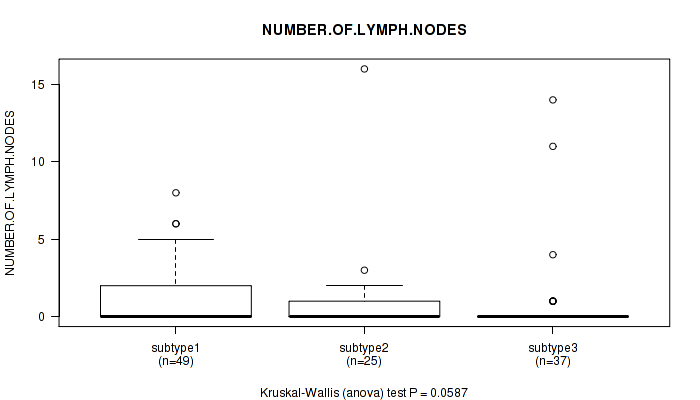
P value = 0.238 (Fisher's exact test), Q value = 1
Table S98. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #10: 'RACE'
| nPatients | AMERICAN INDIAN OR ALASKA NATIVE | ASIAN | BLACK OR AFRICAN AMERICAN | NATIVE HAWAIIAN OR OTHER PACIFIC ISLANDER | WHITE |
|---|---|---|---|---|---|
| ALL | 6 | 15 | 18 | 1 | 135 |
| subtype1 | 2 | 8 | 8 | 0 | 56 |
| subtype2 | 4 | 3 | 6 | 1 | 32 |
| subtype3 | 0 | 4 | 4 | 0 | 47 |
Figure S95. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #10: 'RACE'

P value = 0.477 (Fisher's exact test), Q value = 1
Table S99. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #11: 'ETHNICITY'
| nPatients | HISPANIC OR LATINO | NOT HISPANIC OR LATINO |
|---|---|---|
| ALL | 12 | 129 |
| subtype1 | 3 | 56 |
| subtype2 | 4 | 32 |
| subtype3 | 5 | 41 |
Figure S96. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #11: 'ETHNICITY'

P value = 0.356 (Kruskal-Wallis (anova)), Q value = 1
Table S100. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 166 | 75.3 (20.5) |
| subtype1 | 72 | 74.7 (25.0) |
| subtype2 | 41 | 74.9 (18.0) |
| subtype3 | 53 | 76.5 (15.2) |
Figure S97. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'

P value = 0.945 (Fisher's exact test), Q value = 1
Table S101. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #13: 'TUMOR_STATUS'
| nPatients | TUMOR FREE | WITH TUMOR |
|---|---|---|
| ALL | 50 | 20 |
| subtype1 | 22 | 8 |
| subtype2 | 13 | 6 |
| subtype3 | 15 | 6 |
Figure S98. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #13: 'TUMOR_STATUS'
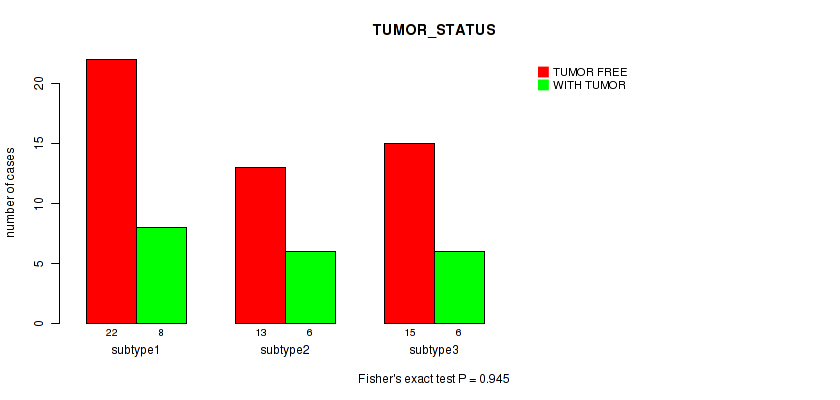
P value = 0.662 (Fisher's exact test), Q value = 1
Table S102. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
| nPatients | NIGERIA | RUSSIA | UKRAINE | UNITED STATES | VIETNAM |
|---|---|---|---|---|---|
| ALL | 1 | 11 | 6 | 150 | 12 |
| subtype1 | 0 | 7 | 2 | 61 | 7 |
| subtype2 | 0 | 1 | 2 | 42 | 2 |
| subtype3 | 1 | 3 | 2 | 47 | 3 |
Figure S99. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'

P value = 0.131 (Fisher's exact test), Q value = 1
Table S103. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'
| nPatients | G1 | G2 | G3 | G4 | GX |
|---|---|---|---|---|---|
| ALL | 14 | 85 | 74 | 1 | 5 |
| subtype1 | 6 | 44 | 26 | 0 | 1 |
| subtype2 | 2 | 17 | 25 | 1 | 1 |
| subtype3 | 6 | 24 | 23 | 0 | 3 |
Figure S100. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'

P value = 0.799 (Kruskal-Wallis (anova)), Q value = 1
Table S104. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 25 | 1997.7 (11.7) |
| subtype1 | 14 | 1997.9 (12.6) |
| subtype2 | 3 | 1996.0 (3.6) |
| subtype3 | 8 | 1998.1 (13.2) |
Figure S101. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'

P value = 0.0633 (Kruskal-Wallis (anova)), Q value = 1
Table S105. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 54 | 18.6 (12.3) |
| subtype1 | 28 | 21.2 (13.3) |
| subtype2 | 11 | 18.8 (9.9) |
| subtype3 | 15 | 13.6 (11.0) |
Figure S102. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'

P value = 0.384 (Fisher's exact test), Q value = 1
Table S106. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
| nPatients | CURRENT REFORMED SMOKER FOR < OR = 15 YEARS | CURRENT REFORMED SMOKER FOR > 15 YEARS | CURRENT REFORMED SMOKER, DURATION NOT SPECIFIED | CURRENT SMOKER | LIFELONG NON-SMOKER |
|---|---|---|---|---|---|
| ALL | 23 | 8 | 1 | 36 | 83 |
| subtype1 | 11 | 7 | 1 | 16 | 33 |
| subtype2 | 4 | 0 | 0 | 10 | 21 |
| subtype3 | 8 | 1 | 0 | 10 | 29 |
Figure S103. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'

P value = 0.812 (Kruskal-Wallis (anova)), Q value = 1
Table S107. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 48 | 20.5 (6.3) |
| subtype1 | 25 | 21.9 (7.7) |
| subtype2 | 9 | 19.1 (4.2) |
| subtype3 | 14 | 19.0 (3.9) |
Figure S104. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'

P value = 0.84 (Fisher's exact test), Q value = 1
Table S108. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
| nPatients | COMBINATION | EXTERNAL | EXTERNAL BEAM | INTERNAL |
|---|---|---|---|---|
| ALL | 17 | 37 | 14 | 5 |
| subtype1 | 8 | 13 | 7 | 2 |
| subtype2 | 5 | 11 | 5 | 2 |
| subtype3 | 4 | 13 | 2 | 1 |
Figure S105. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
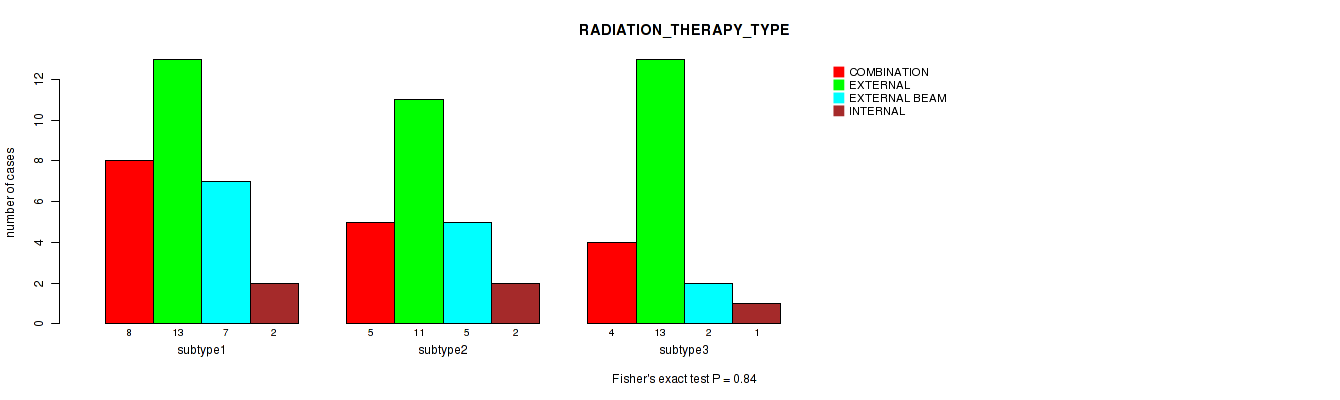
P value = 1 (Fisher's exact test), Q value = 1
Table S109. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
| nPatients | CGY | GY |
|---|---|---|
| ALL | 12 | 3 |
| subtype1 | 4 | 1 |
| subtype2 | 5 | 1 |
| subtype3 | 3 | 1 |
Figure S106. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
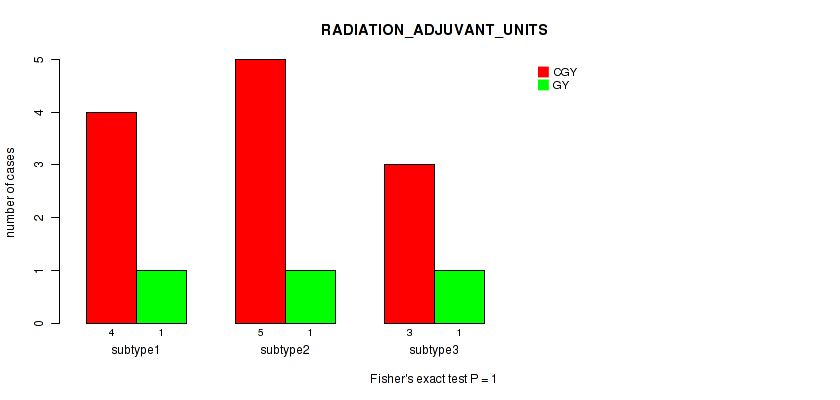
P value = 0.12 (Kruskal-Wallis (anova)), Q value = 1
Table S110. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 156 | 3.5 (2.5) |
| subtype1 | 67 | 3.8 (2.7) |
| subtype2 | 40 | 3.6 (2.3) |
| subtype3 | 49 | 2.9 (2.1) |
Figure S107. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'

P value = 0.159 (Kruskal-Wallis (anova)), Q value = 1
Table S111. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 96 | 0.1 (0.4) |
| subtype1 | 45 | 0.2 (0.5) |
| subtype2 | 23 | 0.0 (0.0) |
| subtype3 | 28 | 0.0 (0.2) |
Figure S108. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'

P value = 0.355 (Kruskal-Wallis (anova)), Q value = 1
Table S112. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 108 | 0.3 (0.6) |
| subtype1 | 52 | 0.3 (0.6) |
| subtype2 | 27 | 0.2 (0.4) |
| subtype3 | 29 | 0.4 (0.6) |
Figure S109. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'

P value = 0.142 (Kruskal-Wallis (anova)), Q value = 1
Table S113. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 161 | 2.4 (1.7) |
| subtype1 | 71 | 2.4 (1.5) |
| subtype2 | 41 | 2.7 (1.9) |
| subtype3 | 49 | 2.1 (1.8) |
Figure S110. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'

P value = 0.737 (Kruskal-Wallis (anova)), Q value = 1
Table S114. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 103 | 0.9 (1.9) |
| subtype1 | 49 | 1.2 (2.6) |
| subtype2 | 24 | 0.6 (1.1) |
| subtype3 | 30 | 0.7 (1.1) |
Figure S111. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'

P value = 0.96 (Kruskal-Wallis (anova)), Q value = 1
Table S115. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 97 | 0.1 (0.3) |
| subtype1 | 46 | 0.1 (0.4) |
| subtype2 | 23 | 0.1 (0.3) |
| subtype3 | 28 | 0.1 (0.3) |
Figure S112. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'

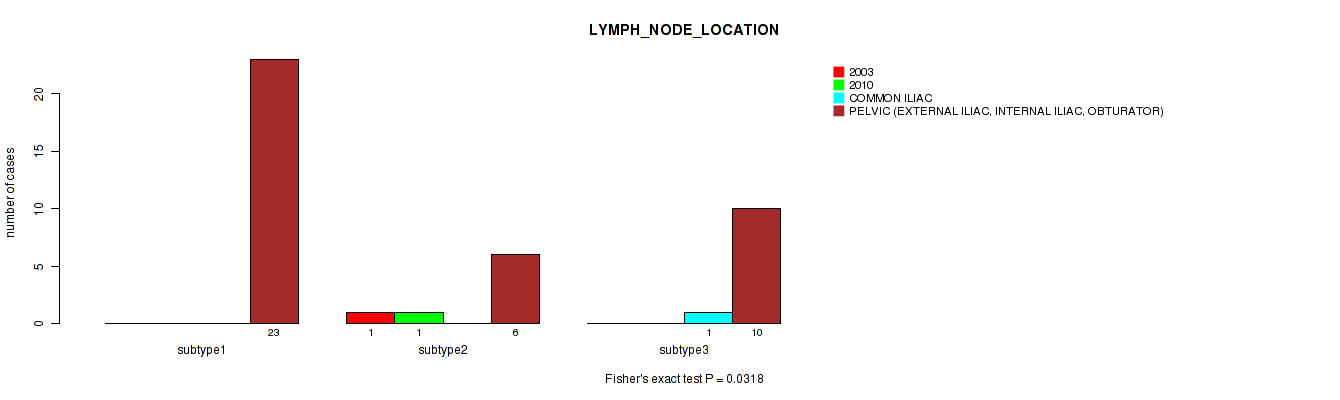
P value = 0.0318 (Fisher's exact test), Q value = 1
Table S116. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
| nPatients | 2003 | 2010 | COMMON ILIAC | PELVIC (EXTERNAL ILIAC, INTERNAL ILIAC, OBTURATOR) |
|---|---|---|---|---|
| ALL | 1 | 1 | 1 | 39 |
| subtype1 | 0 | 0 | 0 | 23 |
| subtype2 | 1 | 1 | 0 | 6 |
| subtype3 | 0 | 0 | 1 | 10 |
Figure S113. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
P value = 0.839 (Fisher's exact test), Q value = 1
Table S117. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'
| nPatients | MACROSCOPIC PARAMETRIAL INVOLVEMENT | MICROSCOPIC PARAMETRIAL INVOLVEMENT | OTHER LOCATION, SPECIFY | POSITIVE BLADDER MARGIN | POSITIVE VAGINAL MARGIN |
|---|---|---|---|---|---|
| ALL | 2 | 6 | 12 | 1 | 5 |
| subtype1 | 1 | 2 | 6 | 1 | 2 |
| subtype2 | 0 | 2 | 5 | 0 | 2 |
| subtype3 | 1 | 2 | 1 | 0 | 1 |
Figure S114. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'

P value = 0.275 (Fisher's exact test), Q value = 1
Table S118. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
| nPatients | INDETERMINATE (NEITHER PRE OR POSTMENOPAUSAL) | PERI (6-12 MONTHS SINCE LAST MENSTRUAL PERIOD) | POST (PRIOR BILATERAL OVARIECTOMY OR >12 MO SINCE LMP WITH NO PRIOR HYSTERECTOMY) | PRE (<6 MONTHS SINCE LMP AND NO PRIOR BILATERAL OVARIECTOMY AND NOT ON ESTROGEN REPLACEMENT) |
|---|---|---|---|---|
| ALL | 2 | 10 | 53 | 81 |
| subtype1 | 2 | 4 | 30 | 30 |
| subtype2 | 0 | 2 | 9 | 22 |
| subtype3 | 0 | 4 | 14 | 29 |
Figure S115. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
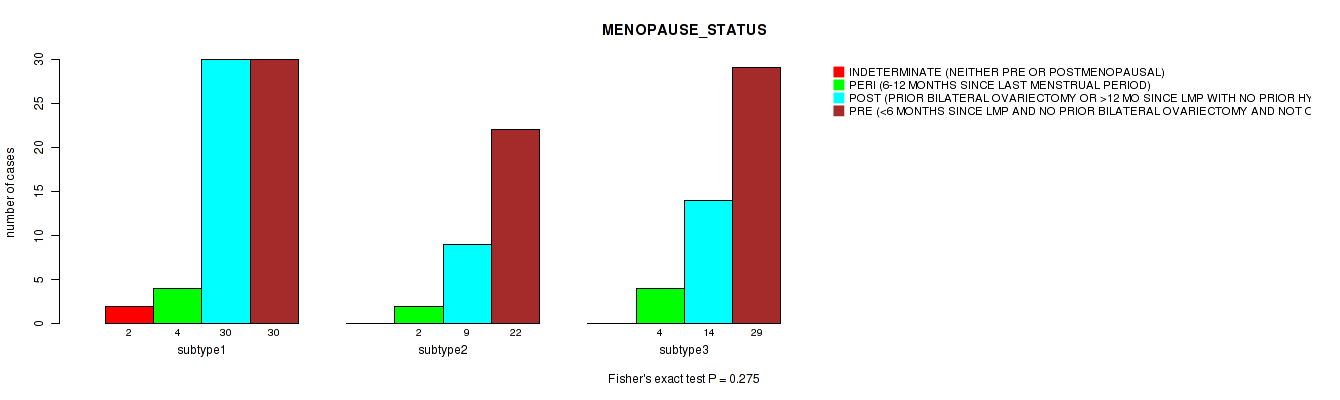
P value = 0.459 (Fisher's exact test), Q value = 1
Table S119. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 54 | 62 |
| subtype1 | 21 | 31 |
| subtype2 | 12 | 10 |
| subtype3 | 21 | 21 |
Figure S116. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'

P value = 0.0587 (Kruskal-Wallis (anova)), Q value = 1
Table S120. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 111 | 1.1 (2.6) |
| subtype1 | 49 | 1.2 (1.9) |
| subtype2 | 25 | 1.0 (3.2) |
| subtype3 | 37 | 0.9 (2.9) |
Figure S117. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'
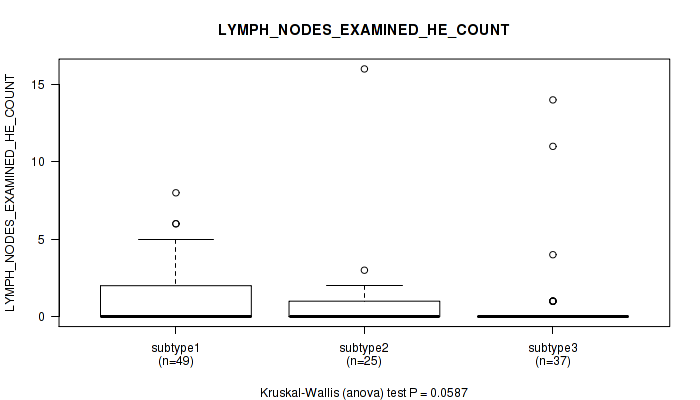
P value = 0.79 (Kruskal-Wallis (anova)), Q value = 1
Table S121. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 130 | 22.3 (12.8) |
| subtype1 | 56 | 24.3 (15.5) |
| subtype2 | 31 | 21.2 (10.7) |
| subtype3 | 43 | 20.5 (9.9) |
Figure S118. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'

P value = 0.0222 (Fisher's exact test), Q value = 1
Table S122. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'
| nPatients | KERATINIZING SQUAMOUS CELL CARCINOMA | NON-KERATINIZING SQUAMOUS CELL CARCINOMA |
|---|---|---|
| ALL | 39 | 81 |
| subtype1 | 21 | 38 |
| subtype2 | 16 | 23 |
| subtype3 | 2 | 20 |
Figure S119. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'
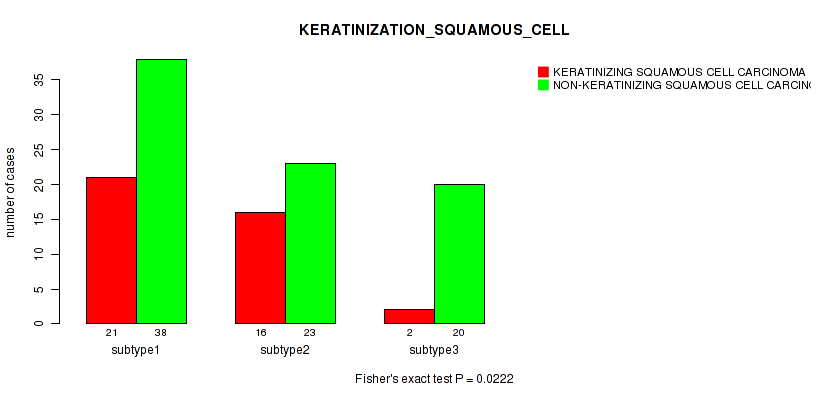
P value = 0.147 (Kruskal-Wallis (anova)), Q value = 1
Table S123. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 180 | 2007.5 (5.1) |
| subtype1 | 77 | 2007.2 (5.7) |
| subtype2 | 47 | 2006.9 (4.7) |
| subtype3 | 56 | 2008.5 (4.4) |
Figure S120. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'

P value = 0.477 (Fisher's exact test), Q value = 1
Table S124. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
| nPatients | OTHER | RADICAL HYSTERECTOMY | SIMPLE HYSTERECTOMY |
|---|---|---|---|
| ALL | 3 | 117 | 5 |
| subtype1 | 1 | 55 | 1 |
| subtype2 | 0 | 23 | 2 |
| subtype3 | 2 | 39 | 2 |
Figure S121. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'

P value = 0.114 (Fisher's exact test), Q value = 1
Table S125. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'
| nPatients | CURRENT USER | FORMER USER | NEVER USED |
|---|---|---|---|
| ALL | 8 | 40 | 43 |
| subtype1 | 1 | 12 | 22 |
| subtype2 | 2 | 11 | 10 |
| subtype3 | 5 | 17 | 11 |
Figure S122. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'

P value = 0.502 (Kruskal-Wallis (anova)), Q value = 1
Table S126. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 160 | 162.1 (6.9) |
| subtype1 | 70 | 161.7 (7.3) |
| subtype2 | 38 | 161.3 (5.6) |
| subtype3 | 52 | 163.1 (7.1) |
Figure S123. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'
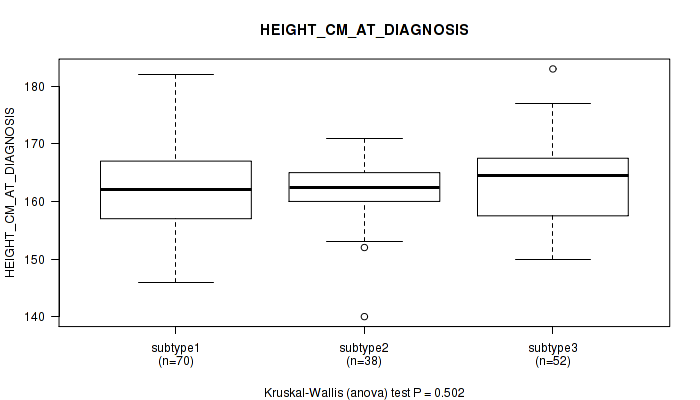
P value = 0.111 (Fisher's exact test), Q value = 1
Table S127. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 78 | 14 |
| subtype1 | 30 | 8 |
| subtype2 | 18 | 0 |
| subtype3 | 30 | 6 |
Figure S124. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'

P value = 1 (Fisher's exact test), Q value = 1
Table S128. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'
| nPatients | CARBOPLATIN | CISPLATIN |
|---|---|---|
| ALL | 2 | 16 |
| subtype1 | 1 | 6 |
| subtype2 | 1 | 7 |
| subtype3 | 0 | 3 |
Figure S125. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'
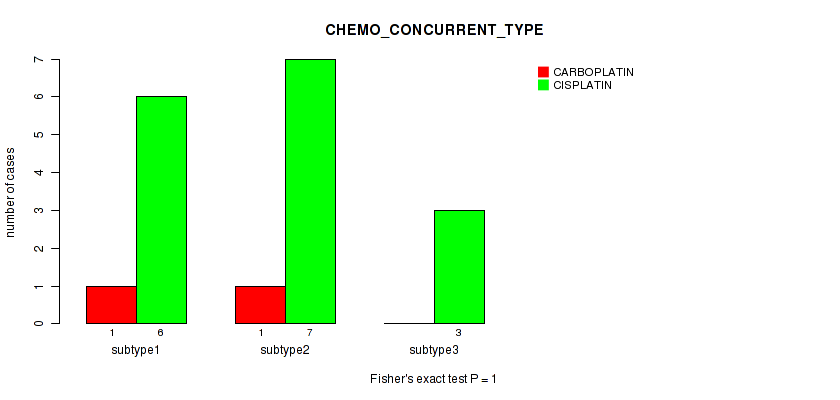
P value = 0.0624 (Fisher's exact test), Q value = 1
Table S129. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'
| nPatients | T1A | T1A1 | T1B | T1B1 | T1B2 | T2 | T2A | T2A1 | T2A2 | T2B | T3B | T4 | TX |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 2 | 1 | 23 | 52 | 22 | 3 | 6 | 6 | 9 | 13 | 2 | 1 | 7 |
| subtype1 | 2 | 1 | 14 | 16 | 7 | 2 | 3 | 2 | 4 | 8 | 1 | 1 | 2 |
| subtype2 | 0 | 0 | 8 | 11 | 6 | 1 | 1 | 2 | 2 | 1 | 0 | 0 | 4 |
| subtype3 | 0 | 0 | 1 | 25 | 9 | 0 | 2 | 2 | 3 | 4 | 1 | 0 | 1 |
Figure S126. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'
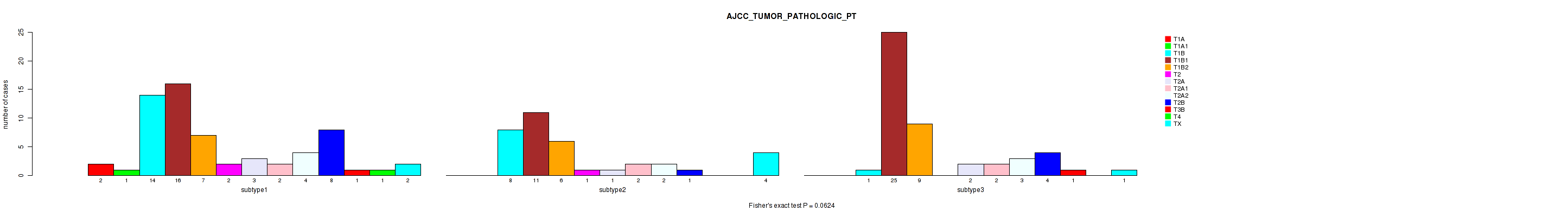
P value = 0.128 (Kruskal-Wallis (anova)), Q value = 1
Table S130. Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 180 | 47.5 (12.9) |
| subtype1 | 77 | 50.1 (13.7) |
| subtype2 | 47 | 46.0 (14.0) |
| subtype3 | 56 | 45.2 (10.1) |
Figure S127. Get High-res Image Clustering Approach #3: 'RNAseq CNMF subtypes' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'

Table S131. Description of clustering approach #4: 'RNAseq cHierClus subtypes'
| Cluster Labels | 1 | 2 | 3 | 4 |
|---|---|---|---|---|
| Number of samples | 91 | 25 | 17 | 47 |
P value = 0.0267 (logrank test), Q value = 1
Table S132. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 173 | 33 | 0.0 - 182.9 (13.6) |
| subtype1 | 86 | 16 | 0.1 - 182.9 (15.5) |
| subtype2 | 25 | 3 | 0.0 - 147.4 (13.1) |
| subtype3 | 17 | 7 | 0.1 - 78.7 (11.7) |
| subtype4 | 45 | 7 | 0.1 - 137.2 (9.7) |
Figure S128. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'

P value = 0.0489 (Kruskal-Wallis (anova)), Q value = 1
Table S133. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 179 | 47.4 (12.9) |
| subtype1 | 90 | 49.0 (14.4) |
| subtype2 | 25 | 50.6 (9.8) |
| subtype3 | 17 | 40.9 (12.2) |
| subtype4 | 47 | 45.1 (10.4) |
Figure S129. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #2: 'AGE'

P value = 0.762 (Fisher's exact test), Q value = 1
Table S134. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
| nPatients | T1 | T2 | T3+T4 |
|---|---|---|---|
| ALL | 100 | 37 | 3 |
| subtype1 | 45 | 19 | 2 |
| subtype2 | 19 | 3 | 0 |
| subtype3 | 9 | 4 | 0 |
| subtype4 | 27 | 11 | 1 |
Figure S130. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'

P value = 0.583 (Fisher's exact test), Q value = 1
Table S135. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 89 | 42 |
| subtype1 | 39 | 22 |
| subtype2 | 13 | 8 |
| subtype3 | 10 | 3 |
| subtype4 | 27 | 9 |
Figure S131. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'

P value = 0.242 (Fisher's exact test), Q value = 1
Table S136. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'
| nPatients | M0 | M1 | MX |
|---|---|---|---|
| ALL | 76 | 3 | 65 |
| subtype1 | 33 | 0 | 34 |
| subtype2 | 15 | 0 | 8 |
| subtype3 | 8 | 0 | 6 |
| subtype4 | 20 | 3 | 17 |
Figure S132. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'

P value = 1e-05 (Fisher's exact test), Q value = 0.0034
Table S137. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
| nPatients | ADENOSQUAMOUS | CERVICAL SQUAMOUS CELL CARCINOMA | ENDOCERVICAL ADENOCARCINOMA OF THE USUAL TYPE | ENDOCERVICAL TYPE OF ADENOCARCINOMA | ENDOMETRIOID ADENOCARCINOMA OF ENDOCERVIX | MUCINOUS ADENOCARCINOMA OF ENDOCERVICAL TYPE |
|---|---|---|---|---|---|---|
| ALL | 2 | 148 | 4 | 20 | 2 | 4 |
| subtype1 | 0 | 91 | 0 | 0 | 0 | 0 |
| subtype2 | 0 | 25 | 0 | 0 | 0 | 0 |
| subtype3 | 0 | 17 | 0 | 0 | 0 | 0 |
| subtype4 | 2 | 15 | 4 | 20 | 2 | 4 |
Figure S133. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'

P value = 0.536 (Fisher's exact test), Q value = 1
Table S138. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 30 | 150 |
| subtype1 | 17 | 74 |
| subtype2 | 4 | 21 |
| subtype3 | 4 | 13 |
| subtype4 | 5 | 42 |
Figure S134. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'

P value = 0.198 (Kruskal-Wallis (anova)), Q value = 1
Table S139. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 54 | 18.6 (12.3) |
| subtype1 | 30 | 21.0 (12.9) |
| subtype2 | 4 | 16.2 (12.5) |
| subtype3 | 7 | 17.8 (9.5) |
| subtype4 | 13 | 14.2 (11.7) |
Figure S135. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'

P value = 0.557 (Kruskal-Wallis (anova)), Q value = 1
Table S140. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 111 | 1.1 (2.6) |
| subtype1 | 49 | 1.3 (2.8) |
| subtype2 | 20 | 0.8 (1.2) |
| subtype3 | 12 | 0.3 (0.7) |
| subtype4 | 30 | 1.1 (3.2) |
Figure S136. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'

P value = 0.387 (Fisher's exact test), Q value = 1
Table S141. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #10: 'RACE'
| nPatients | AMERICAN INDIAN OR ALASKA NATIVE | ASIAN | BLACK OR AFRICAN AMERICAN | NATIVE HAWAIIAN OR OTHER PACIFIC ISLANDER | WHITE |
|---|---|---|---|---|---|
| ALL | 6 | 15 | 18 | 1 | 135 |
| subtype1 | 4 | 7 | 9 | 1 | 67 |
| subtype2 | 0 | 4 | 3 | 0 | 17 |
| subtype3 | 2 | 1 | 3 | 0 | 11 |
| subtype4 | 0 | 3 | 3 | 0 | 40 |
Figure S137. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #10: 'RACE'
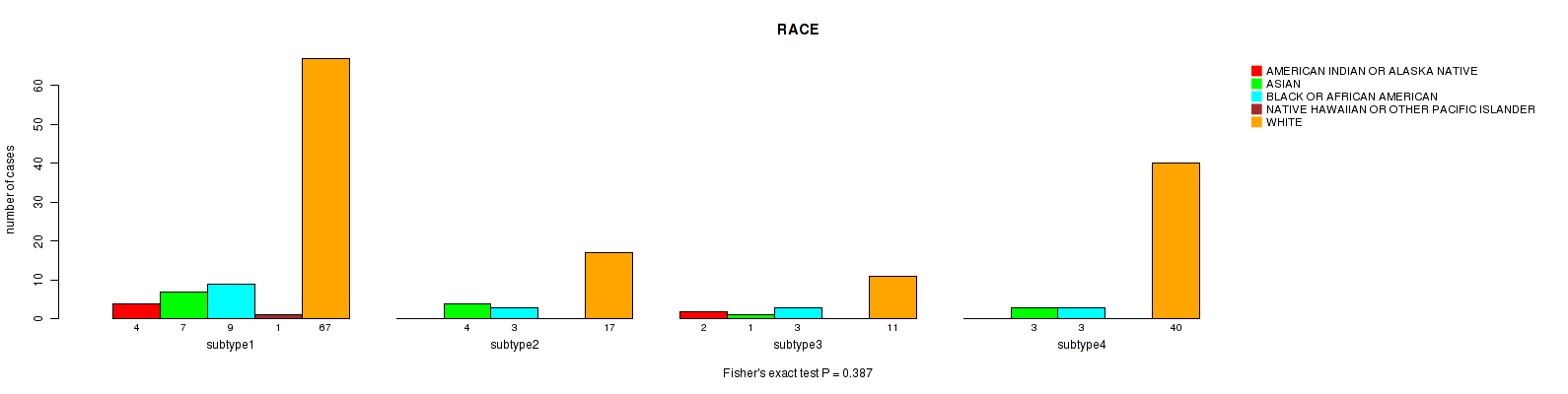
P value = 0.436 (Fisher's exact test), Q value = 1
Table S142. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #11: 'ETHNICITY'
| nPatients | HISPANIC OR LATINO | NOT HISPANIC OR LATINO |
|---|---|---|
| ALL | 12 | 129 |
| subtype1 | 6 | 63 |
| subtype2 | 0 | 20 |
| subtype3 | 1 | 12 |
| subtype4 | 5 | 34 |
Figure S138. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #11: 'ETHNICITY'

P value = 0.517 (Kruskal-Wallis (anova)), Q value = 1
Table S143. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 166 | 75.3 (20.5) |
| subtype1 | 84 | 75.9 (24.3) |
| subtype2 | 24 | 71.0 (16.8) |
| subtype3 | 14 | 75.6 (16.2) |
| subtype4 | 44 | 76.5 (15.3) |
Figure S139. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'

P value = 0.195 (Fisher's exact test), Q value = 1
Table S144. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #13: 'TUMOR_STATUS'
| nPatients | TUMOR FREE | WITH TUMOR |
|---|---|---|
| ALL | 50 | 20 |
| subtype1 | 23 | 9 |
| subtype2 | 12 | 1 |
| subtype3 | 5 | 4 |
| subtype4 | 10 | 6 |
Figure S140. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #13: 'TUMOR_STATUS'

P value = 0.535 (Fisher's exact test), Q value = 1
Table S145. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
| nPatients | NIGERIA | RUSSIA | UKRAINE | UNITED STATES | VIETNAM |
|---|---|---|---|---|---|
| ALL | 1 | 11 | 6 | 150 | 12 |
| subtype1 | 0 | 7 | 2 | 76 | 6 |
| subtype2 | 0 | 2 | 0 | 20 | 3 |
| subtype3 | 0 | 0 | 2 | 14 | 1 |
| subtype4 | 1 | 2 | 2 | 40 | 2 |
Figure S141. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'

P value = 0.0169 (Fisher's exact test), Q value = 1
Table S146. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'
| nPatients | G1 | G2 | G3 | G4 | GX |
|---|---|---|---|---|---|
| ALL | 14 | 85 | 74 | 1 | 5 |
| subtype1 | 6 | 52 | 30 | 0 | 2 |
| subtype2 | 1 | 6 | 17 | 0 | 1 |
| subtype3 | 1 | 5 | 10 | 1 | 0 |
| subtype4 | 6 | 22 | 17 | 0 | 2 |
Figure S142. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'

P value = 0.794 (Kruskal-Wallis (anova)), Q value = 1
Table S147. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 25 | 1997.7 (11.7) |
| subtype1 | 12 | 1997.6 (13.1) |
| subtype2 | 6 | 2001.2 (10.9) |
| subtype3 | 2 | 1997.5 (3.5) |
| subtype4 | 5 | 1994.0 (12.8) |
Figure S143. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'

P value = 0.198 (Kruskal-Wallis (anova)), Q value = 1
Table S148. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 54 | 18.6 (12.3) |
| subtype1 | 30 | 21.0 (12.9) |
| subtype2 | 4 | 16.2 (12.5) |
| subtype3 | 7 | 17.8 (9.5) |
| subtype4 | 13 | 14.2 (11.7) |
Figure S144. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'

P value = 0.413 (Fisher's exact test), Q value = 1
Table S149. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
| nPatients | CURRENT REFORMED SMOKER FOR < OR = 15 YEARS | CURRENT REFORMED SMOKER FOR > 15 YEARS | CURRENT REFORMED SMOKER, DURATION NOT SPECIFIED | CURRENT SMOKER | LIFELONG NON-SMOKER |
|---|---|---|---|---|---|
| ALL | 23 | 8 | 1 | 36 | 83 |
| subtype1 | 10 | 6 | 1 | 20 | 37 |
| subtype2 | 7 | 1 | 0 | 2 | 12 |
| subtype3 | 2 | 0 | 0 | 5 | 8 |
| subtype4 | 4 | 1 | 0 | 9 | 26 |
Figure S145. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
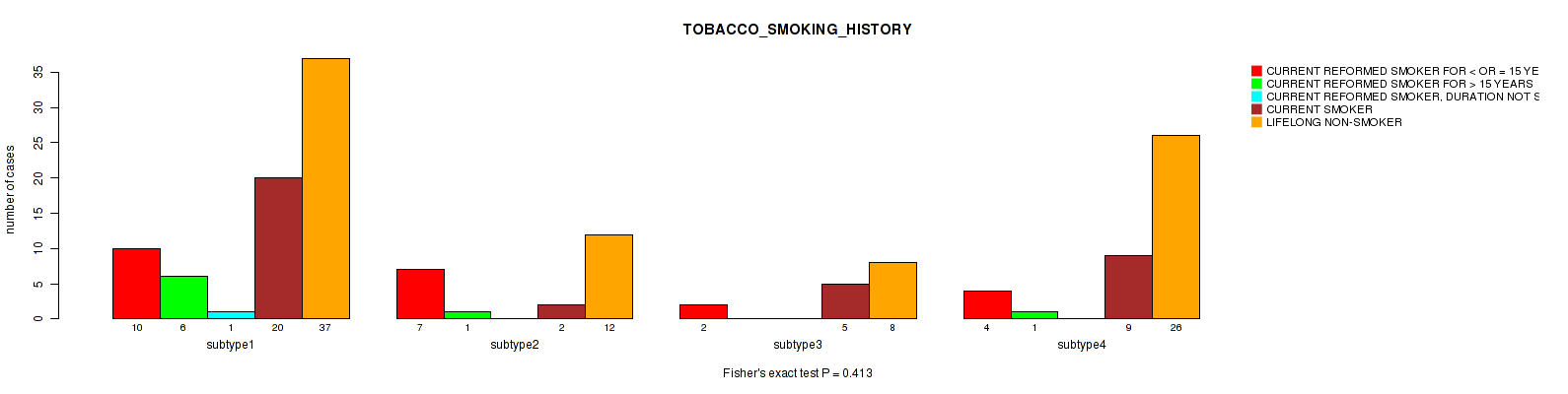
P value = 0.576 (Kruskal-Wallis (anova)), Q value = 1
Table S150. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 48 | 20.5 (6.3) |
| subtype1 | 25 | 21.4 (7.1) |
| subtype2 | 5 | 22.6 (8.2) |
| subtype3 | 6 | 18.0 (4.0) |
| subtype4 | 12 | 19.0 (4.1) |
Figure S146. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'
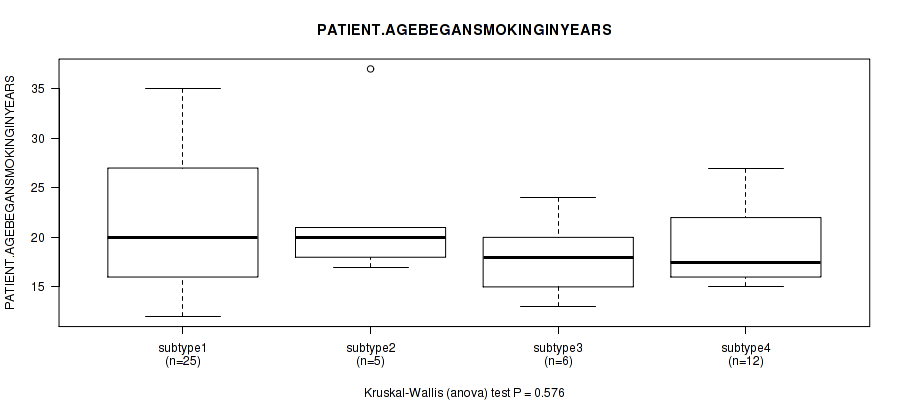
P value = 0.458 (Fisher's exact test), Q value = 1
Table S151. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
| nPatients | COMBINATION | EXTERNAL | EXTERNAL BEAM | INTERNAL |
|---|---|---|---|---|
| ALL | 17 | 37 | 14 | 5 |
| subtype1 | 10 | 14 | 7 | 3 |
| subtype2 | 1 | 5 | 4 | 1 |
| subtype3 | 2 | 6 | 2 | 1 |
| subtype4 | 4 | 12 | 1 | 0 |
Figure S147. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'

P value = 0.471 (Fisher's exact test), Q value = 1
Table S152. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
| nPatients | CGY | GY |
|---|---|---|
| ALL | 12 | 3 |
| subtype1 | 3 | 2 |
| subtype2 | 2 | 1 |
| subtype3 | 4 | 0 |
| subtype4 | 3 | 0 |
Figure S148. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
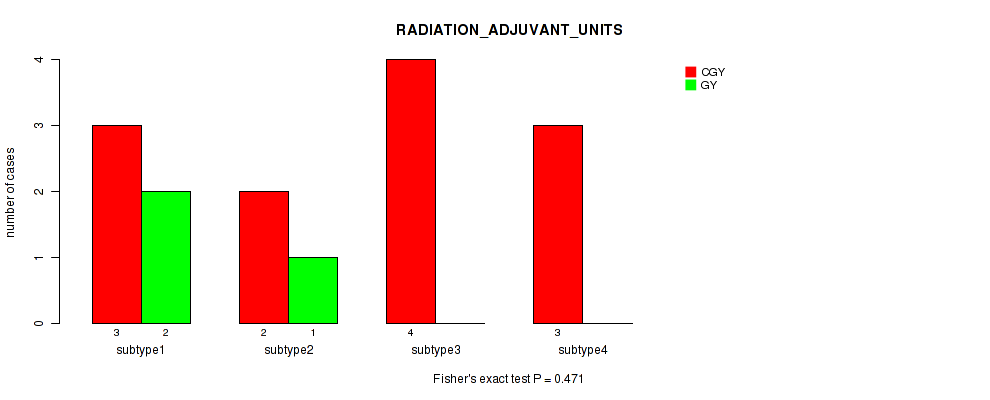
P value = 0.274 (Kruskal-Wallis (anova)), Q value = 1
Table S153. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 156 | 3.5 (2.5) |
| subtype1 | 79 | 3.7 (2.7) |
| subtype2 | 20 | 3.2 (1.8) |
| subtype3 | 16 | 3.7 (2.1) |
| subtype4 | 41 | 2.9 (2.2) |
Figure S149. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
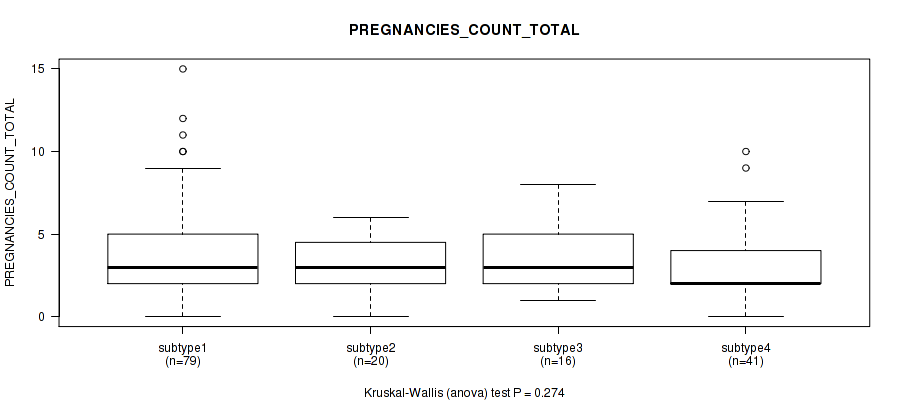
P value = 0.468 (Kruskal-Wallis (anova)), Q value = 1
Table S154. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 96 | 0.1 (0.4) |
| subtype1 | 52 | 0.1 (0.5) |
| subtype2 | 11 | 0.0 (0.0) |
| subtype3 | 10 | 0.0 (0.0) |
| subtype4 | 23 | 0.0 (0.2) |
Figure S150. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'

P value = 0.347 (Kruskal-Wallis (anova)), Q value = 1
Table S155. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 108 | 0.3 (0.6) |
| subtype1 | 61 | 0.3 (0.6) |
| subtype2 | 12 | 0.2 (0.4) |
| subtype3 | 11 | 0.3 (0.5) |
| subtype4 | 24 | 0.5 (0.6) |
Figure S151. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'

P value = 0.155 (Kruskal-Wallis (anova)), Q value = 1
Table S156. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 161 | 2.4 (1.7) |
| subtype1 | 82 | 2.4 (1.6) |
| subtype2 | 22 | 2.4 (1.4) |
| subtype3 | 16 | 2.9 (2.0) |
| subtype4 | 41 | 2.0 (1.9) |
Figure S152. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'
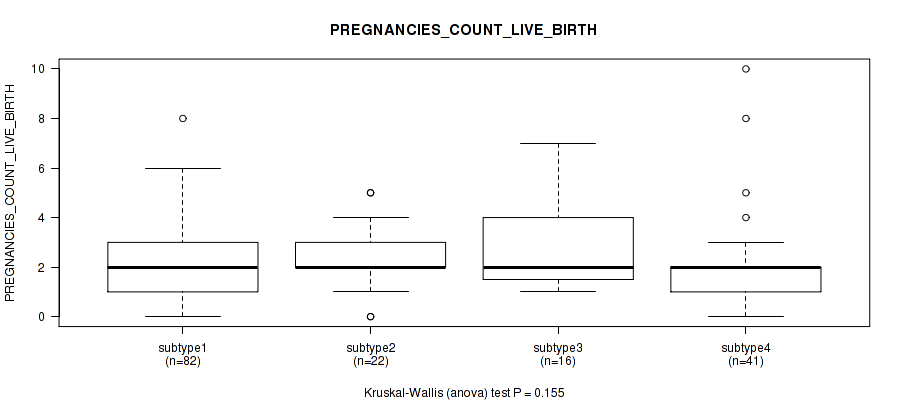
P value = 0.911 (Kruskal-Wallis (anova)), Q value = 1
Table S157. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 103 | 0.9 (1.9) |
| subtype1 | 54 | 1.0 (2.4) |
| subtype2 | 13 | 1.0 (1.5) |
| subtype3 | 11 | 0.7 (1.1) |
| subtype4 | 25 | 0.7 (1.0) |
Figure S153. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
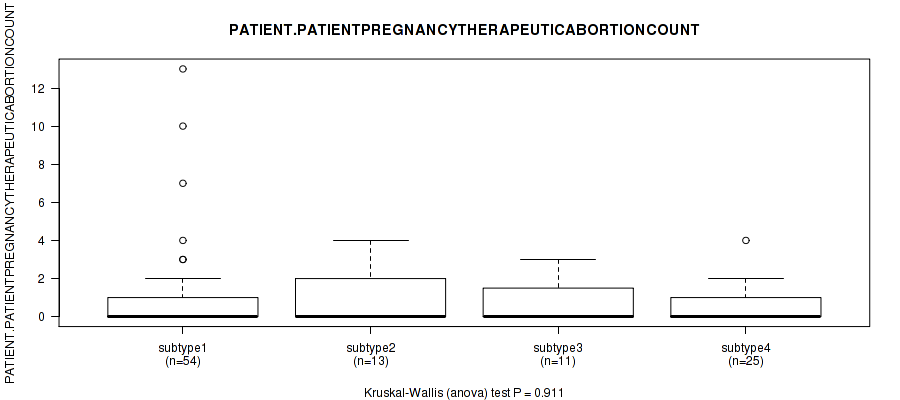
P value = 0.999 (Kruskal-Wallis (anova)), Q value = 1
Table S158. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 97 | 0.1 (0.3) |
| subtype1 | 53 | 0.1 (0.4) |
| subtype2 | 12 | 0.1 (0.3) |
| subtype3 | 10 | 0.1 (0.3) |
| subtype4 | 22 | 0.1 (0.3) |
Figure S154. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'

P value = 0.521 (Fisher's exact test), Q value = 1
Table S159. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
| nPatients | 2003 | 2010 | COMMON ILIAC | PELVIC (EXTERNAL ILIAC, INTERNAL ILIAC, OBTURATOR) |
|---|---|---|---|---|
| ALL | 1 | 1 | 1 | 39 |
| subtype1 | 0 | 1 | 0 | 21 |
| subtype2 | 1 | 0 | 0 | 7 |
| subtype3 | 0 | 0 | 0 | 3 |
| subtype4 | 0 | 0 | 1 | 8 |
Figure S155. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
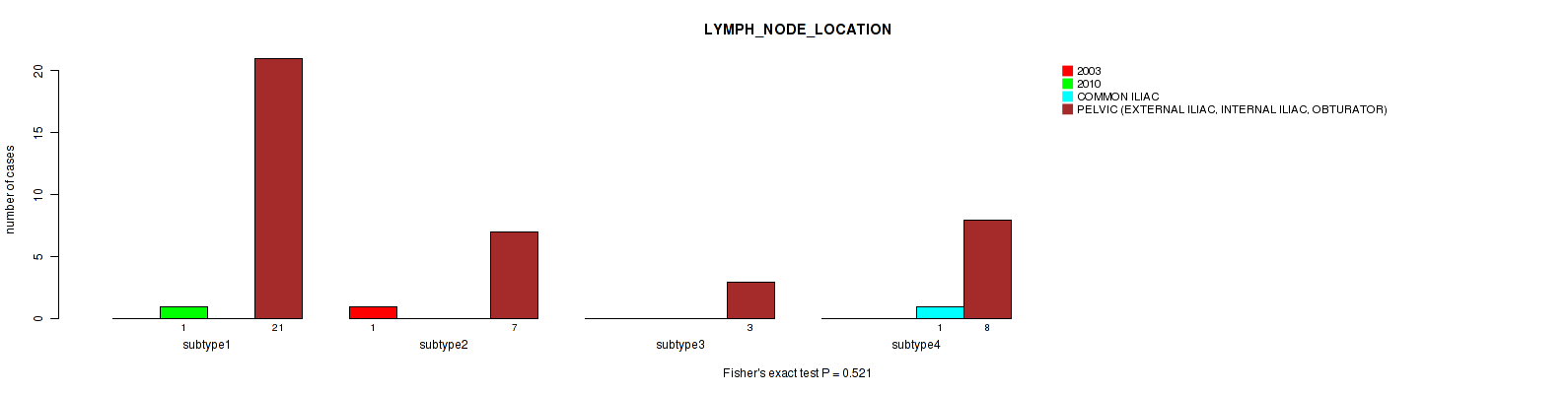
P value = 0.436 (Fisher's exact test), Q value = 1
Table S160. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'
| nPatients | MACROSCOPIC PARAMETRIAL INVOLVEMENT | MICROSCOPIC PARAMETRIAL INVOLVEMENT | OTHER LOCATION, SPECIFY | POSITIVE BLADDER MARGIN | POSITIVE VAGINAL MARGIN |
|---|---|---|---|---|---|
| ALL | 2 | 6 | 12 | 1 | 5 |
| subtype1 | 1 | 3 | 9 | 1 | 1 |
| subtype2 | 0 | 1 | 2 | 0 | 2 |
| subtype3 | 0 | 1 | 1 | 0 | 1 |
| subtype4 | 1 | 1 | 0 | 0 | 1 |
Figure S156. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'

P value = 0.0133 (Fisher's exact test), Q value = 1
Table S161. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
| nPatients | INDETERMINATE (NEITHER PRE OR POSTMENOPAUSAL) | PERI (6-12 MONTHS SINCE LAST MENSTRUAL PERIOD) | POST (PRIOR BILATERAL OVARIECTOMY OR >12 MO SINCE LMP WITH NO PRIOR HYSTERECTOMY) | PRE (<6 MONTHS SINCE LMP AND NO PRIOR BILATERAL OVARIECTOMY AND NOT ON ESTROGEN REPLACEMENT) |
|---|---|---|---|---|
| ALL | 2 | 10 | 53 | 81 |
| subtype1 | 2 | 3 | 30 | 37 |
| subtype2 | 0 | 3 | 12 | 8 |
| subtype3 | 0 | 1 | 0 | 11 |
| subtype4 | 0 | 3 | 11 | 25 |
Figure S157. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #30: 'MENOPAUSE_STATUS'

P value = 0.877 (Fisher's exact test), Q value = 1
Table S162. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 54 | 62 |
| subtype1 | 23 | 28 |
| subtype2 | 8 | 12 |
| subtype3 | 5 | 5 |
| subtype4 | 18 | 17 |
Figure S158. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'

P value = 0.557 (Kruskal-Wallis (anova)), Q value = 1
Table S163. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 111 | 1.1 (2.6) |
| subtype1 | 49 | 1.3 (2.8) |
| subtype2 | 20 | 0.8 (1.2) |
| subtype3 | 12 | 0.3 (0.7) |
| subtype4 | 30 | 1.1 (3.2) |
Figure S159. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'

P value = 0.78 (Kruskal-Wallis (anova)), Q value = 1
Table S164. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 130 | 22.3 (12.8) |
| subtype1 | 60 | 23.1 (13.9) |
| subtype2 | 21 | 20.5 (14.6) |
| subtype3 | 13 | 22.9 (13.2) |
| subtype4 | 36 | 21.7 (9.8) |
Figure S160. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'

P value = 0.115 (Fisher's exact test), Q value = 1
Table S165. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'
| nPatients | KERATINIZING SQUAMOUS CELL CARCINOMA | NON-KERATINIZING SQUAMOUS CELL CARCINOMA |
|---|---|---|
| ALL | 39 | 81 |
| subtype1 | 27 | 46 |
| subtype2 | 6 | 13 |
| subtype3 | 5 | 8 |
| subtype4 | 1 | 14 |
Figure S161. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'

P value = 0.123 (Kruskal-Wallis (anova)), Q value = 1
Table S166. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 180 | 2007.5 (5.1) |
| subtype1 | 91 | 2007.1 (5.2) |
| subtype2 | 25 | 2008.8 (4.5) |
| subtype3 | 17 | 2005.3 (6.1) |
| subtype4 | 47 | 2008.3 (4.4) |
Figure S162. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
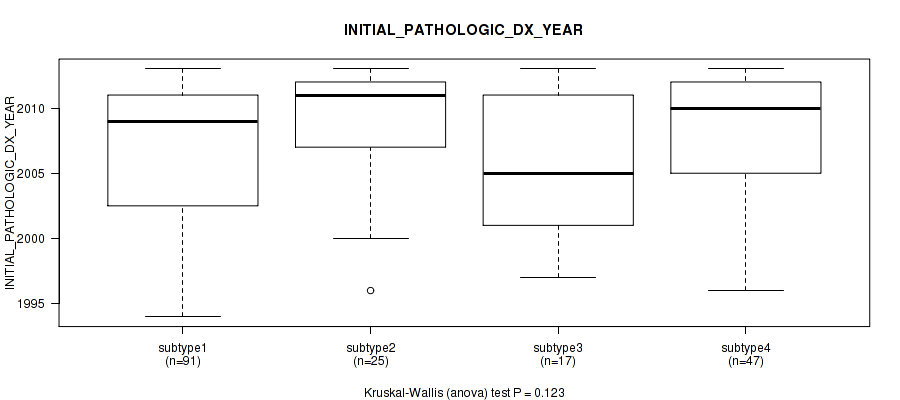
P value = 0.169 (Fisher's exact test), Q value = 1
Table S167. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
| nPatients | OTHER | RADICAL HYSTERECTOMY | SIMPLE HYSTERECTOMY |
|---|---|---|---|
| ALL | 3 | 117 | 5 |
| subtype1 | 1 | 56 | 1 |
| subtype2 | 0 | 19 | 0 |
| subtype3 | 0 | 9 | 2 |
| subtype4 | 2 | 33 | 2 |
Figure S163. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'

P value = 0.0445 (Fisher's exact test), Q value = 1
Table S168. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'
| nPatients | CURRENT USER | FORMER USER | NEVER USED |
|---|---|---|---|
| ALL | 8 | 40 | 43 |
| subtype1 | 0 | 15 | 22 |
| subtype2 | 1 | 8 | 6 |
| subtype3 | 2 | 3 | 6 |
| subtype4 | 5 | 14 | 9 |
Figure S164. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'

P value = 0.41 (Kruskal-Wallis (anova)), Q value = 1
Table S169. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 160 | 162.1 (6.9) |
| subtype1 | 81 | 161.0 (7.2) |
| subtype2 | 24 | 163.5 (6.6) |
| subtype3 | 12 | 162.6 (4.2) |
| subtype4 | 43 | 163.0 (6.8) |
Figure S165. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'

P value = 0.877 (Fisher's exact test), Q value = 1
Table S170. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 78 | 14 |
| subtype1 | 35 | 8 |
| subtype2 | 11 | 2 |
| subtype3 | 6 | 0 |
| subtype4 | 26 | 4 |
Figure S166. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'

P value = 1 (Fisher's exact test), Q value = 1
Table S171. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'
| nPatients | CARBOPLATIN | CISPLATIN |
|---|---|---|
| ALL | 2 | 16 |
| subtype1 | 2 | 8 |
| subtype2 | 0 | 3 |
| subtype3 | 0 | 3 |
| subtype4 | 0 | 2 |
Figure S167. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'

P value = 0.00179 (Fisher's exact test), Q value = 0.59
Table S172. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'
| nPatients | T1A | T1A1 | T1B | T1B1 | T1B2 | T2 | T2A | T2A1 | T2A2 | T2B | T3B | T4 | TX |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 2 | 1 | 23 | 52 | 22 | 3 | 6 | 6 | 9 | 13 | 2 | 1 | 7 |
| subtype1 | 1 | 0 | 16 | 16 | 12 | 2 | 3 | 1 | 5 | 8 | 1 | 1 | 3 |
| subtype2 | 1 | 1 | 0 | 15 | 2 | 1 | 0 | 1 | 0 | 1 | 0 | 0 | 2 |
| subtype3 | 0 | 0 | 6 | 2 | 1 | 0 | 1 | 2 | 1 | 0 | 0 | 0 | 1 |
| subtype4 | 0 | 0 | 1 | 19 | 7 | 0 | 2 | 2 | 3 | 4 | 1 | 0 | 1 |
Figure S168. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'

P value = 0.0454 (Kruskal-Wallis (anova)), Q value = 1
Table S173. Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 180 | 47.5 (12.9) |
| subtype1 | 91 | 49.2 (14.4) |
| subtype2 | 25 | 50.6 (9.8) |
| subtype3 | 17 | 40.9 (12.2) |
| subtype4 | 47 | 45.1 (10.4) |
Figure S169. Get High-res Image Clustering Approach #4: 'RNAseq cHierClus subtypes' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
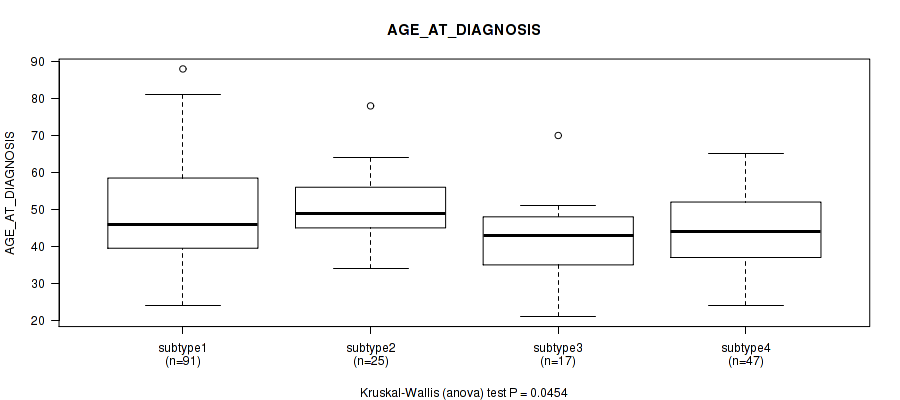
Table S174. Description of clustering approach #5: 'MIRSEQ CNMF'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 62 | 60 | 78 |
P value = 0.499 (logrank test), Q value = 1
Table S175. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 193 | 38 | 0.0 - 182.9 (13.9) |
| subtype1 | 61 | 10 | 0.1 - 182.9 (13.1) |
| subtype2 | 57 | 9 | 0.0 - 147.4 (14.9) |
| subtype3 | 75 | 19 | 0.0 - 177.0 (14.4) |
Figure S170. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #1: 'Time to Death'
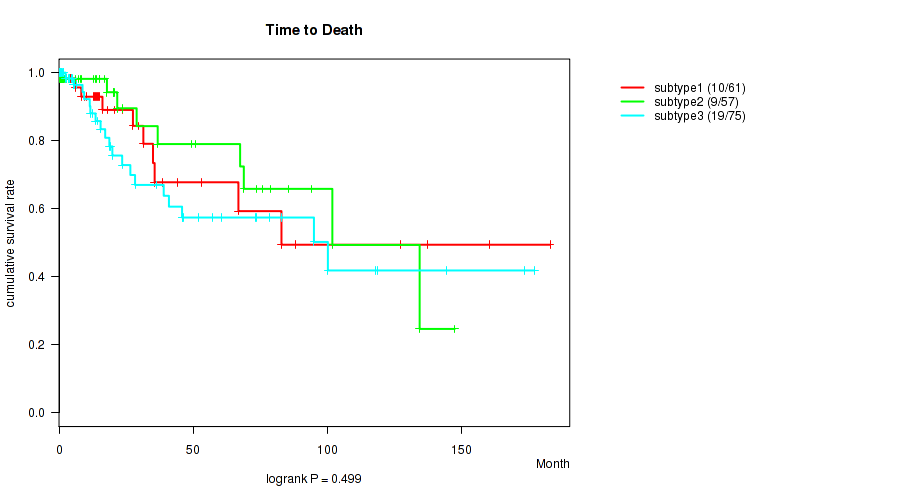
P value = 0.221 (Kruskal-Wallis (anova)), Q value = 1
Table S176. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 199 | 47.5 (13.4) |
| subtype1 | 62 | 46.4 (10.2) |
| subtype2 | 60 | 50.1 (14.4) |
| subtype3 | 77 | 46.3 (14.7) |
Figure S171. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #2: 'AGE'
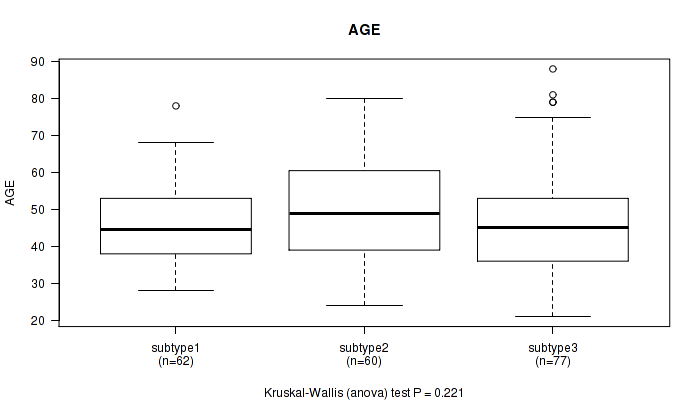
P value = 0.89 (Fisher's exact test), Q value = 1
Table S177. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
| nPatients | T1 | T2 | T3+T4 |
|---|---|---|---|
| ALL | 108 | 39 | 4 |
| subtype1 | 38 | 16 | 1 |
| subtype2 | 34 | 12 | 1 |
| subtype3 | 36 | 11 | 2 |
Figure S172. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'

P value = 0.587 (Fisher's exact test), Q value = 1
Table S178. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 100 | 44 |
| subtype1 | 38 | 13 |
| subtype2 | 30 | 16 |
| subtype3 | 32 | 15 |
Figure S173. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'

P value = 0.628 (Fisher's exact test), Q value = 1
Table S179. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'
| nPatients | M0 | M1 | MX |
|---|---|---|---|
| ALL | 83 | 4 | 70 |
| subtype1 | 28 | 3 | 25 |
| subtype2 | 28 | 0 | 21 |
| subtype3 | 27 | 1 | 24 |
Figure S174. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'

P value = 1e-05 (Fisher's exact test), Q value = 0.0034
Table S180. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
| nPatients | ADENOSQUAMOUS | CERVICAL SQUAMOUS CELL CARCINOMA | ENDOCERVICAL ADENOCARCINOMA OF THE USUAL TYPE | ENDOCERVICAL TYPE OF ADENOCARCINOMA | ENDOMETRIOID ADENOCARCINOMA OF ENDOCERVIX | MUCINOUS ADENOCARCINOMA OF ENDOCERVICAL TYPE |
|---|---|---|---|---|---|---|
| ALL | 2 | 168 | 4 | 20 | 2 | 4 |
| subtype1 | 2 | 36 | 3 | 16 | 2 | 3 |
| subtype2 | 0 | 54 | 1 | 4 | 0 | 1 |
| subtype3 | 0 | 78 | 0 | 0 | 0 | 0 |
Figure S175. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'

P value = 0.411 (Fisher's exact test), Q value = 1
Table S181. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 32 | 168 |
| subtype1 | 8 | 54 |
| subtype2 | 8 | 52 |
| subtype3 | 16 | 62 |
Figure S176. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'

P value = 0.0665 (Kruskal-Wallis (anova)), Q value = 1
Table S182. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 62 | 19.4 (13.6) |
| subtype1 | 14 | 12.5 (9.4) |
| subtype2 | 21 | 22.0 (14.4) |
| subtype3 | 27 | 21.0 (14.0) |
Figure S177. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'

P value = 0.443 (Kruskal-Wallis (anova)), Q value = 1
Table S183. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 123 | 1.1 (2.6) |
| subtype1 | 40 | 0.9 (2.6) |
| subtype2 | 46 | 1.3 (2.5) |
| subtype3 | 37 | 0.9 (2.7) |
Figure S178. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'

P value = 0.386 (Fisher's exact test), Q value = 1
Table S184. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #10: 'RACE'
| nPatients | AMERICAN INDIAN OR ALASKA NATIVE | ASIAN | BLACK OR AFRICAN AMERICAN | NATIVE HAWAIIAN OR OTHER PACIFIC ISLANDER | WHITE |
|---|---|---|---|---|---|
| ALL | 7 | 16 | 23 | 1 | 148 |
| subtype1 | 0 | 5 | 8 | 0 | 48 |
| subtype2 | 1 | 6 | 6 | 0 | 44 |
| subtype3 | 6 | 5 | 9 | 1 | 56 |
Figure S179. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #10: 'RACE'

P value = 0.506 (Fisher's exact test), Q value = 1
Table S185. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #11: 'ETHNICITY'
| nPatients | HISPANIC OR LATINO | NOT HISPANIC OR LATINO |
|---|---|---|
| ALL | 13 | 144 |
| subtype1 | 4 | 47 |
| subtype2 | 2 | 42 |
| subtype3 | 7 | 55 |
Figure S180. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #11: 'ETHNICITY'

P value = 0.35 (Kruskal-Wallis (anova)), Q value = 1
Table S186. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 182 | 75.2 (20.9) |
| subtype1 | 60 | 76.7 (16.3) |
| subtype2 | 52 | 74.1 (26.3) |
| subtype3 | 70 | 74.8 (20.0) |
Figure S181. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'

P value = 0.724 (Fisher's exact test), Q value = 1
Table S187. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #13: 'TUMOR_STATUS'
| nPatients | TUMOR FREE | WITH TUMOR |
|---|---|---|
| ALL | 51 | 22 |
| subtype1 | 19 | 6 |
| subtype2 | 12 | 6 |
| subtype3 | 20 | 10 |
Figure S182. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #13: 'TUMOR_STATUS'
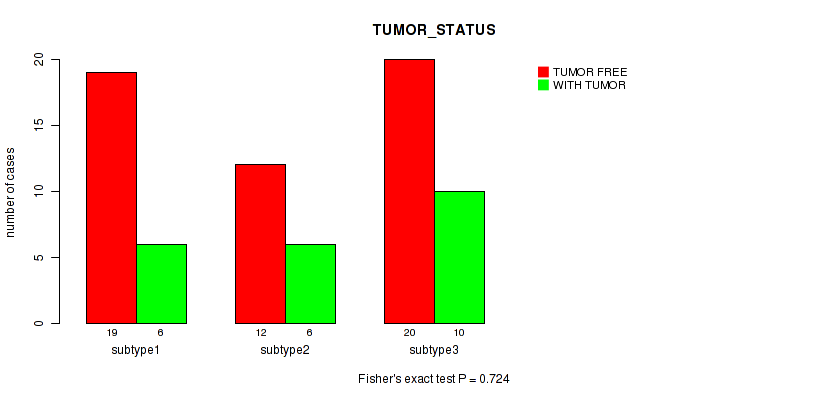
P value = 0.579 (Fisher's exact test), Q value = 1
Table S188. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
| nPatients | NIGERIA | RUSSIA | UKRAINE | UNITED STATES | VIETNAM |
|---|---|---|---|---|---|
| ALL | 1 | 11 | 7 | 169 | 12 |
| subtype1 | 1 | 2 | 3 | 53 | 3 |
| subtype2 | 0 | 6 | 2 | 47 | 5 |
| subtype3 | 0 | 3 | 2 | 69 | 4 |
Figure S183. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'

P value = 0.0596 (Fisher's exact test), Q value = 1
Table S189. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'
| nPatients | G1 | G2 | G3 | G4 | GX |
|---|---|---|---|---|---|
| ALL | 14 | 94 | 83 | 1 | 5 |
| subtype1 | 6 | 24 | 30 | 0 | 2 |
| subtype2 | 6 | 34 | 18 | 1 | 0 |
| subtype3 | 2 | 36 | 35 | 0 | 3 |
Figure S184. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'

P value = 0.317 (Kruskal-Wallis (anova)), Q value = 1
Table S190. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 27 | 1998.4 (11.5) |
| subtype1 | 6 | 1997.0 (13.6) |
| subtype2 | 9 | 1995.1 (11.5) |
| subtype3 | 12 | 2001.5 (10.6) |
Figure S185. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
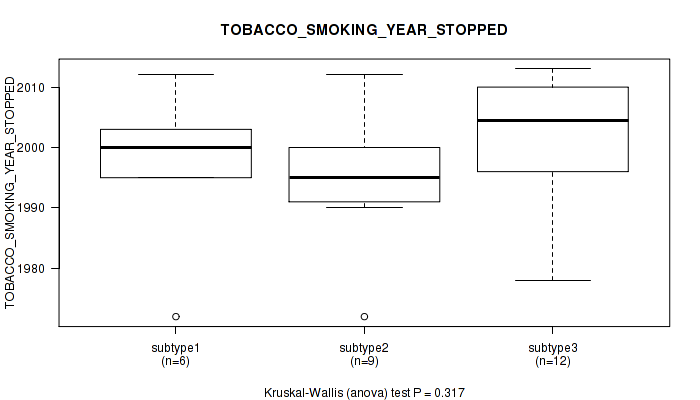
P value = 0.0665 (Kruskal-Wallis (anova)), Q value = 1
Table S191. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 62 | 19.4 (13.6) |
| subtype1 | 14 | 12.5 (9.4) |
| subtype2 | 21 | 22.0 (14.4) |
| subtype3 | 27 | 21.0 (14.0) |
Figure S186. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'
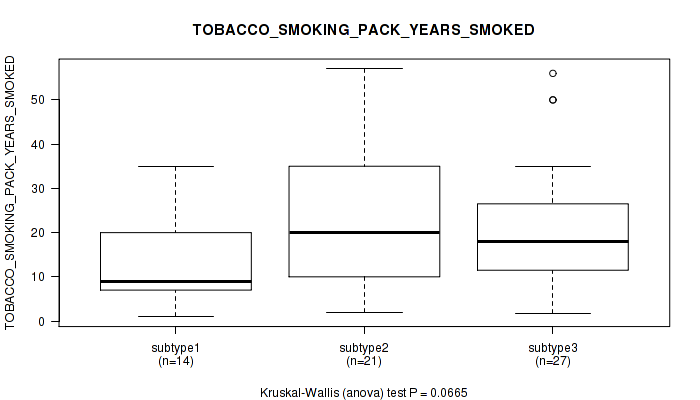
P value = 0.245 (Fisher's exact test), Q value = 1
Table S192. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
| nPatients | CURRENT REFORMED SMOKER FOR < OR = 15 YEARS | CURRENT REFORMED SMOKER FOR > 15 YEARS | CURRENT REFORMED SMOKER, DURATION NOT SPECIFIED | CURRENT SMOKER | LIFELONG NON-SMOKER |
|---|---|---|---|---|---|
| ALL | 25 | 8 | 3 | 43 | 89 |
| subtype1 | 7 | 2 | 0 | 8 | 35 |
| subtype2 | 7 | 4 | 2 | 15 | 26 |
| subtype3 | 11 | 2 | 1 | 20 | 28 |
Figure S187. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'

P value = 0.239 (Kruskal-Wallis (anova)), Q value = 1
Table S193. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 54 | 20.5 (6.2) |
| subtype1 | 12 | 20.3 (5.7) |
| subtype2 | 19 | 22.1 (6.5) |
| subtype3 | 23 | 19.2 (6.0) |
Figure S188. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'

P value = 0.267 (Fisher's exact test), Q value = 1
Table S194. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
| nPatients | COMBINATION | EXTERNAL | EXTERNAL BEAM | IMPLANTS | INTERNAL |
|---|---|---|---|---|---|
| ALL | 18 | 44 | 14 | 1 | 8 |
| subtype1 | 5 | 16 | 3 | 0 | 0 |
| subtype2 | 3 | 13 | 5 | 1 | 3 |
| subtype3 | 10 | 15 | 6 | 0 | 5 |
Figure S189. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
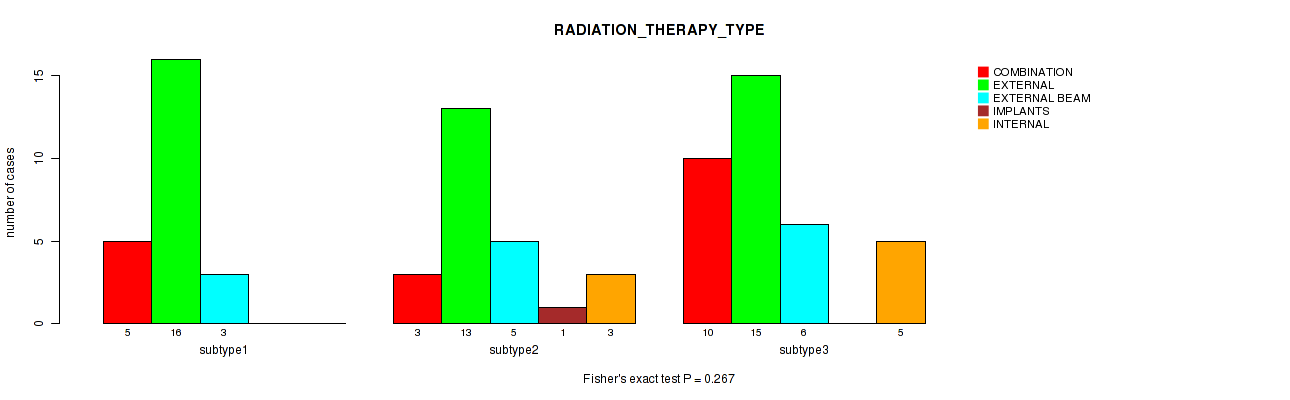
P value = 0.137 (Fisher's exact test), Q value = 1
Table S195. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
| nPatients | CGY | GY |
|---|---|---|
| ALL | 16 | 4 |
| subtype1 | 4 | 0 |
| subtype2 | 6 | 0 |
| subtype3 | 6 | 4 |
Figure S190. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
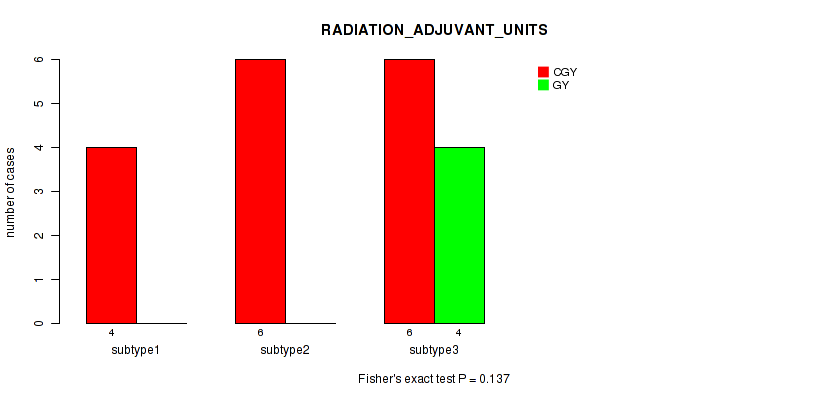
P value = 0.229 (Kruskal-Wallis (anova)), Q value = 1
Table S196. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 175 | 3.5 (2.4) |
| subtype1 | 53 | 3.2 (2.5) |
| subtype2 | 55 | 3.7 (2.4) |
| subtype3 | 67 | 3.5 (2.5) |
Figure S191. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'

P value = 0.791 (Kruskal-Wallis (anova)), Q value = 1
Table S197. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 100 | 0.1 (0.4) |
| subtype1 | 28 | 0.1 (0.3) |
| subtype2 | 30 | 0.1 (0.5) |
| subtype3 | 42 | 0.1 (0.3) |
Figure S192. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'

P value = 0.0365 (Kruskal-Wallis (anova)), Q value = 1
Table S198. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 116 | 0.4 (0.6) |
| subtype1 | 31 | 0.5 (0.7) |
| subtype2 | 37 | 0.4 (0.5) |
| subtype3 | 48 | 0.2 (0.6) |
Figure S193. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'

P value = 0.167 (Kruskal-Wallis (anova)), Q value = 1
Table S199. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 179 | 2.5 (1.8) |
| subtype1 | 54 | 2.3 (1.9) |
| subtype2 | 56 | 2.7 (1.7) |
| subtype3 | 69 | 2.5 (1.7) |
Figure S194. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'

P value = 0.392 (Kruskal-Wallis (anova)), Q value = 1
Table S200. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 109 | 0.9 (1.9) |
| subtype1 | 30 | 0.9 (1.5) |
| subtype2 | 34 | 1.2 (2.4) |
| subtype3 | 45 | 0.7 (1.7) |
Figure S195. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
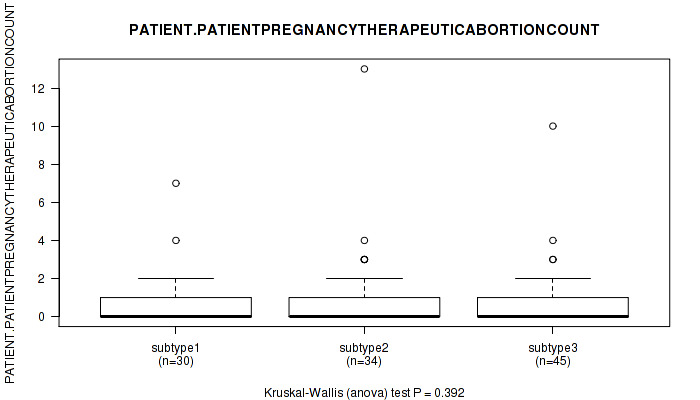
P value = 0.831 (Kruskal-Wallis (anova)), Q value = 1
Table S201. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 101 | 0.1 (0.3) |
| subtype1 | 28 | 0.1 (0.3) |
| subtype2 | 30 | 0.1 (0.4) |
| subtype3 | 43 | 0.1 (0.3) |
Figure S196. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'

P value = 0.343 (Fisher's exact test), Q value = 1
Table S202. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
| nPatients | 2003 | 2010 | COMMON ILIAC | PELVIC (EXTERNAL ILIAC, INTERNAL ILIAC, OBTURATOR) |
|---|---|---|---|---|
| ALL | 1 | 1 | 1 | 41 |
| subtype1 | 0 | 0 | 1 | 12 |
| subtype2 | 0 | 0 | 0 | 16 |
| subtype3 | 1 | 1 | 0 | 13 |
Figure S197. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
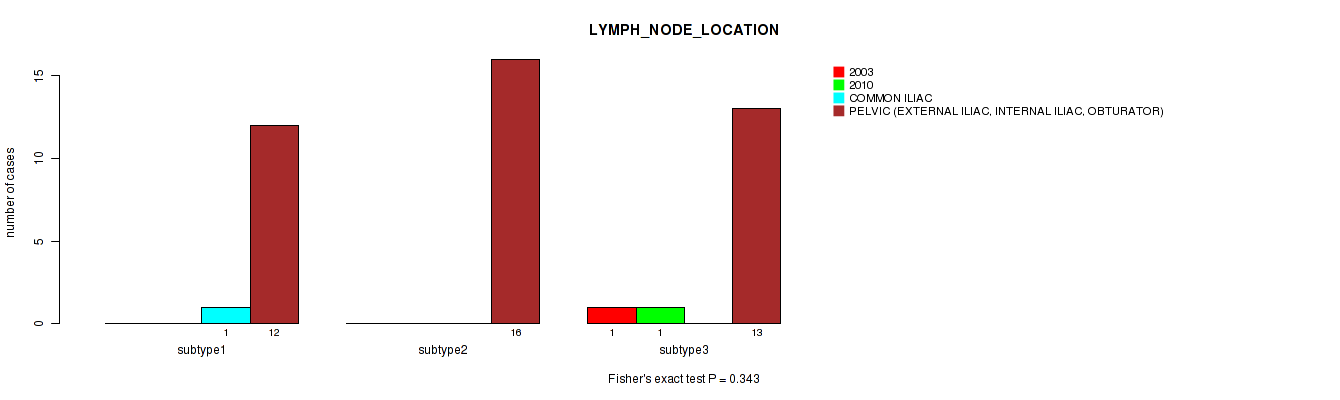
P value = 0.706 (Fisher's exact test), Q value = 1
Table S203. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'
| nPatients | MACROSCOPIC PARAMETRIAL INVOLVEMENT | MICROSCOPIC PARAMETRIAL INVOLVEMENT | OTHER LOCATION, SPECIFY | POSITIVE BLADDER MARGIN | POSITIVE VAGINAL MARGIN |
|---|---|---|---|---|---|
| ALL | 2 | 7 | 13 | 1 | 6 |
| subtype1 | 1 | 2 | 1 | 0 | 1 |
| subtype2 | 1 | 2 | 5 | 1 | 3 |
| subtype3 | 0 | 3 | 7 | 0 | 2 |
Figure S198. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'

P value = 0.466 (Fisher's exact test), Q value = 1
Table S204. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
| nPatients | INDETERMINATE (NEITHER PRE OR POSTMENOPAUSAL) | PERI (6-12 MONTHS SINCE LAST MENSTRUAL PERIOD) | POST (PRIOR BILATERAL OVARIECTOMY OR >12 MO SINCE LMP WITH NO PRIOR HYSTERECTOMY) | PRE (<6 MONTHS SINCE LMP AND NO PRIOR BILATERAL OVARIECTOMY AND NOT ON ESTROGEN REPLACEMENT) |
|---|---|---|---|---|
| ALL | 2 | 10 | 58 | 85 |
| subtype1 | 1 | 4 | 14 | 28 |
| subtype2 | 1 | 3 | 23 | 22 |
| subtype3 | 0 | 3 | 21 | 35 |
Figure S199. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #30: 'MENOPAUSE_STATUS'

P value = 0.325 (Fisher's exact test), Q value = 1
Table S205. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 56 | 66 |
| subtype1 | 17 | 28 |
| subtype2 | 20 | 22 |
| subtype3 | 19 | 16 |
Figure S200. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'

P value = 0.443 (Kruskal-Wallis (anova)), Q value = 1
Table S206. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 123 | 1.1 (2.6) |
| subtype1 | 40 | 0.9 (2.6) |
| subtype2 | 46 | 1.3 (2.5) |
| subtype3 | 37 | 0.9 (2.7) |
Figure S201. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'

P value = 0.528 (Kruskal-Wallis (anova)), Q value = 1
Table S207. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 143 | 22.4 (12.7) |
| subtype1 | 49 | 21.0 (10.7) |
| subtype2 | 49 | 25.0 (14.8) |
| subtype3 | 45 | 21.2 (12.0) |
Figure S202. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
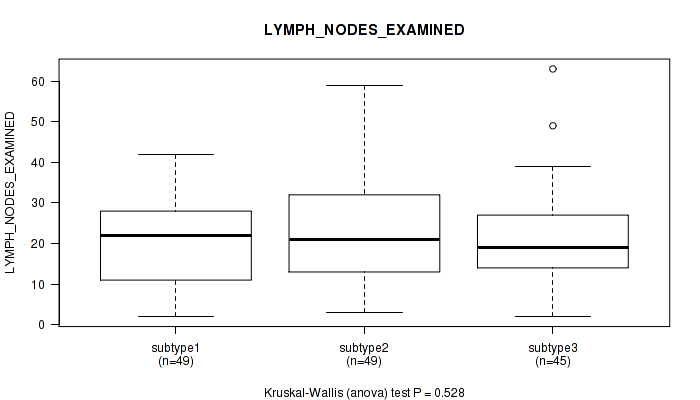
P value = 0.031 (Fisher's exact test), Q value = 1
Table S208. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'
| nPatients | KERATINIZING SQUAMOUS CELL CARCINOMA | NON-KERATINIZING SQUAMOUS CELL CARCINOMA |
|---|---|---|
| ALL | 43 | 88 |
| subtype1 | 4 | 25 |
| subtype2 | 14 | 27 |
| subtype3 | 25 | 36 |
Figure S203. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'

P value = 0.173 (Kruskal-Wallis (anova)), Q value = 1
Table S209. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 200 | 2007.3 (5.2) |
| subtype1 | 62 | 2008.0 (5.3) |
| subtype2 | 60 | 2007.4 (5.0) |
| subtype3 | 78 | 2006.6 (5.3) |
Figure S204. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
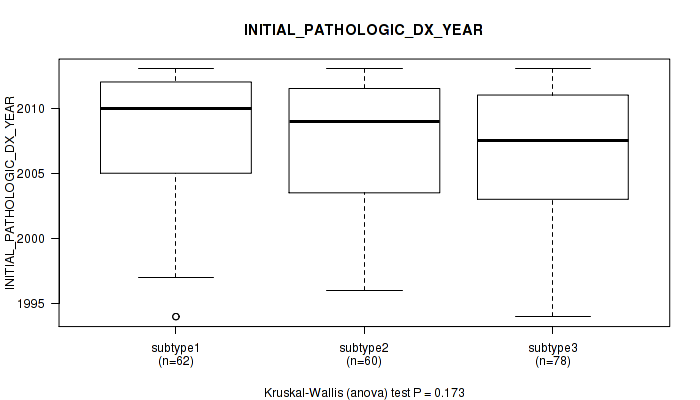
P value = 0.136 (Fisher's exact test), Q value = 1
Table S210. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
| nPatients | OTHER | RADICAL HYSTERECTOMY | SIMPLE HYSTERECTOMY |
|---|---|---|---|
| ALL | 3 | 129 | 5 |
| subtype1 | 3 | 46 | 1 |
| subtype2 | 0 | 47 | 1 |
| subtype3 | 0 | 36 | 3 |
Figure S205. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'

P value = 0.944 (Fisher's exact test), Q value = 1
Table S211. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'
| nPatients | CURRENT USER | FORMER USER | NEVER USED |
|---|---|---|---|
| ALL | 8 | 40 | 47 |
| subtype1 | 4 | 13 | 16 |
| subtype2 | 2 | 13 | 15 |
| subtype3 | 2 | 14 | 16 |
Figure S206. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'

P value = 0.185 (Kruskal-Wallis (anova)), Q value = 1
Table S212. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 173 | 162.1 (7.2) |
| subtype1 | 57 | 163.6 (6.6) |
| subtype2 | 48 | 162.1 (7.8) |
| subtype3 | 68 | 160.8 (7.1) |
Figure S207. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'

P value = 0.322 (Fisher's exact test), Q value = 1
Table S213. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 81 | 15 |
| subtype1 | 28 | 6 |
| subtype2 | 27 | 7 |
| subtype3 | 26 | 2 |
Figure S208. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
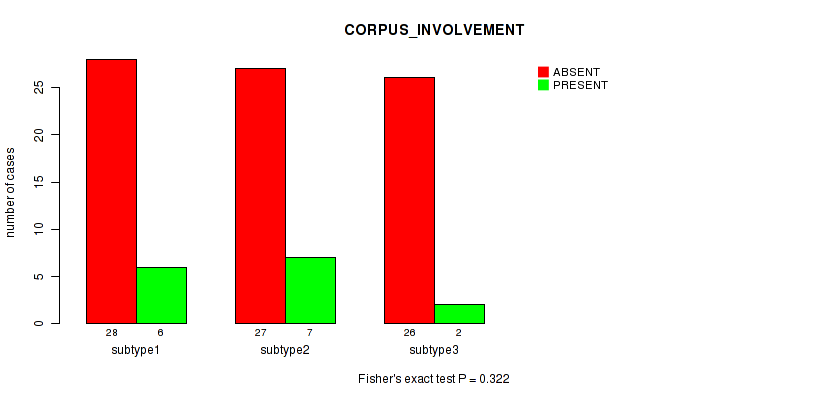
P value = 1 (Fisher's exact test), Q value = 1
Table S214. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'
| nPatients | CARBOPLATIN | CISPLATIN |
|---|---|---|
| ALL | 3 | 17 |
| subtype1 | 0 | 4 |
| subtype2 | 1 | 3 |
| subtype3 | 2 | 10 |
Figure S209. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'

P value = 0.456 (Kruskal-Wallis (anova)), Q value = 1
Table S215. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #41: 'CERVIX_SUV_RESULTS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 10 | 12.8 (6.1) |
| subtype2 | 4 | 10.2 (3.3) |
| subtype3 | 6 | 14.6 (7.1) |
Figure S210. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #41: 'CERVIX_SUV_RESULTS'
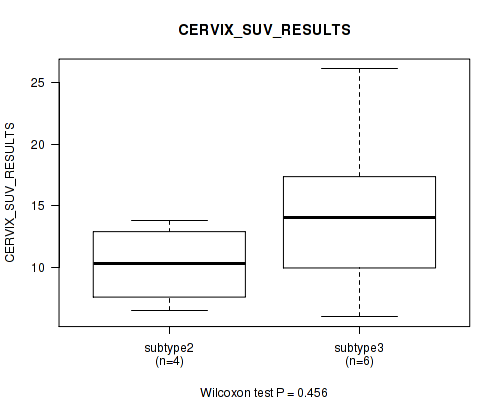
P value = 0.35 (Fisher's exact test), Q value = 1
Table S216. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'
| nPatients | T1A | T1A1 | T1B | T1B1 | T1B2 | T2 | T2A | T2A1 | T2A2 | T2B | T3B | T4 | TIS | TX |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 2 | 1 | 26 | 55 | 24 | 3 | 6 | 7 | 9 | 14 | 2 | 2 | 1 | 8 |
| subtype1 | 0 | 0 | 6 | 21 | 11 | 1 | 2 | 4 | 5 | 4 | 1 | 0 | 0 | 1 |
| subtype2 | 1 | 1 | 5 | 20 | 7 | 0 | 3 | 2 | 2 | 5 | 0 | 1 | 0 | 2 |
| subtype3 | 1 | 0 | 15 | 14 | 6 | 2 | 1 | 1 | 2 | 5 | 1 | 1 | 1 | 5 |
Figure S211. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'

P value = 0.249 (Kruskal-Wallis (anova)), Q value = 1
Table S217. Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 200 | 47.6 (13.4) |
| subtype1 | 62 | 46.4 (10.2) |
| subtype2 | 60 | 50.1 (14.4) |
| subtype3 | 78 | 46.5 (14.7) |
Figure S212. Get High-res Image Clustering Approach #5: 'MIRSEQ CNMF' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'

Table S218. Description of clustering approach #6: 'MIRSEQ CHIERARCHICAL'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 37 | 123 | 40 |
P value = 0.647 (logrank test), Q value = 1
Table S219. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 193 | 38 | 0.0 - 182.9 (13.9) |
| subtype1 | 36 | 7 | 0.1 - 137.2 (7.6) |
| subtype2 | 118 | 23 | 0.0 - 182.9 (14.5) |
| subtype3 | 39 | 8 | 0.0 - 177.0 (15.0) |
Figure S213. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #1: 'Time to Death'

P value = 0.166 (Kruskal-Wallis (anova)), Q value = 1
Table S220. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 199 | 47.5 (13.4) |
| subtype1 | 37 | 43.6 (10.6) |
| subtype2 | 122 | 48.0 (13.6) |
| subtype3 | 40 | 49.4 (14.6) |
Figure S214. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #2: 'AGE'

P value = 0.205 (Fisher's exact test), Q value = 1
Table S221. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
| nPatients | T1 | T2 | T3+T4 |
|---|---|---|---|
| ALL | 108 | 39 | 4 |
| subtype1 | 22 | 7 | 1 |
| subtype2 | 56 | 27 | 3 |
| subtype3 | 30 | 5 | 0 |
Figure S215. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
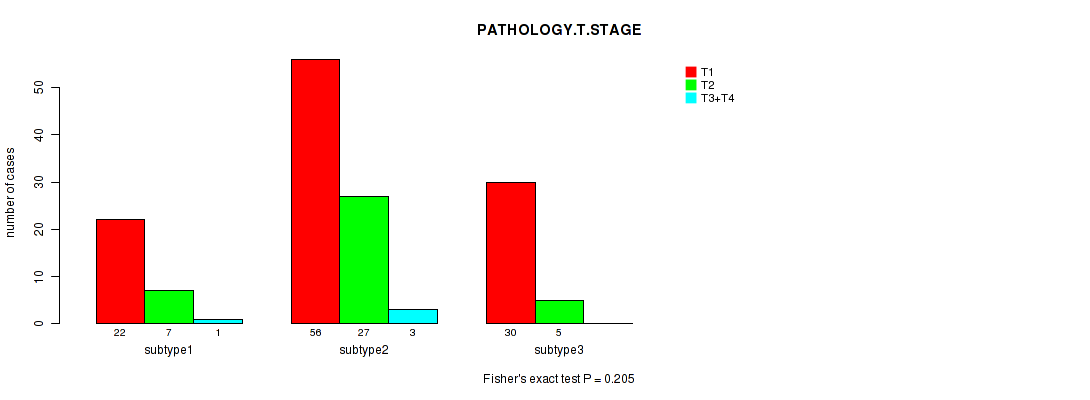
P value = 0.776 (Fisher's exact test), Q value = 1
Table S222. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 100 | 44 |
| subtype1 | 21 | 7 |
| subtype2 | 57 | 26 |
| subtype3 | 22 | 11 |
Figure S216. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'

P value = 0.389 (Fisher's exact test), Q value = 1
Table S223. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'
| nPatients | M0 | M1 | MX |
|---|---|---|---|
| ALL | 83 | 4 | 70 |
| subtype1 | 13 | 2 | 16 |
| subtype2 | 49 | 2 | 39 |
| subtype3 | 21 | 0 | 15 |
Figure S217. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'

P value = 1e-05 (Fisher's exact test), Q value = 0.0034
Table S224. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
| nPatients | ADENOSQUAMOUS | CERVICAL SQUAMOUS CELL CARCINOMA | ENDOCERVICAL ADENOCARCINOMA OF THE USUAL TYPE | ENDOCERVICAL TYPE OF ADENOCARCINOMA | ENDOMETRIOID ADENOCARCINOMA OF ENDOCERVIX | MUCINOUS ADENOCARCINOMA OF ENDOCERVICAL TYPE |
|---|---|---|---|---|---|---|
| ALL | 2 | 168 | 4 | 20 | 2 | 4 |
| subtype1 | 2 | 7 | 4 | 19 | 1 | 4 |
| subtype2 | 0 | 121 | 0 | 1 | 1 | 0 |
| subtype3 | 0 | 40 | 0 | 0 | 0 | 0 |
Figure S218. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
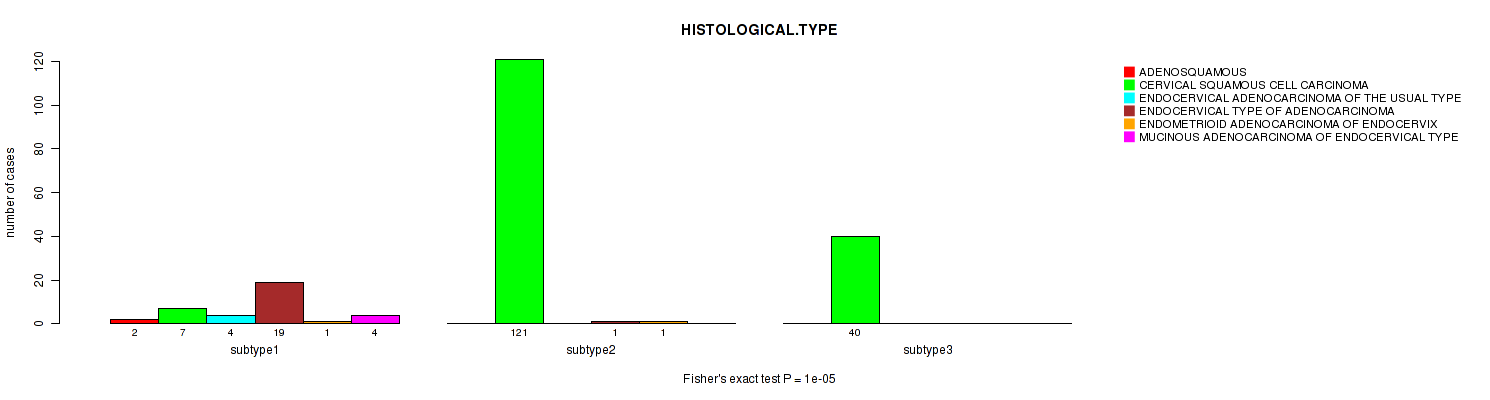
P value = 0.628 (Fisher's exact test), Q value = 1
Table S225. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 32 | 168 |
| subtype1 | 4 | 33 |
| subtype2 | 22 | 101 |
| subtype3 | 6 | 34 |
Figure S219. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'

P value = 0.213 (Kruskal-Wallis (anova)), Q value = 1
Table S226. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 62 | 19.4 (13.6) |
| subtype1 | 12 | 14.5 (12.2) |
| subtype2 | 40 | 20.8 (13.9) |
| subtype3 | 10 | 20.0 (13.9) |
Figure S220. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'

P value = 0.667 (Kruskal-Wallis (anova)), Q value = 1
Table S227. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 123 | 1.1 (2.6) |
| subtype1 | 25 | 1.3 (3.5) |
| subtype2 | 68 | 0.9 (2.3) |
| subtype3 | 30 | 1.2 (2.1) |
Figure S221. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'

P value = 0.421 (Fisher's exact test), Q value = 1
Table S228. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #10: 'RACE'
| nPatients | AMERICAN INDIAN OR ALASKA NATIVE | ASIAN | BLACK OR AFRICAN AMERICAN | NATIVE HAWAIIAN OR OTHER PACIFIC ISLANDER | WHITE |
|---|---|---|---|---|---|
| ALL | 7 | 16 | 23 | 1 | 148 |
| subtype1 | 0 | 2 | 2 | 0 | 32 |
| subtype2 | 7 | 9 | 16 | 1 | 88 |
| subtype3 | 0 | 5 | 5 | 0 | 28 |
Figure S222. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #10: 'RACE'

P value = 0.0869 (Fisher's exact test), Q value = 1
Table S229. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #11: 'ETHNICITY'
| nPatients | HISPANIC OR LATINO | NOT HISPANIC OR LATINO |
|---|---|---|
| ALL | 13 | 144 |
| subtype1 | 4 | 25 |
| subtype2 | 9 | 87 |
| subtype3 | 0 | 32 |
Figure S223. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #11: 'ETHNICITY'

P value = 0.634 (Kruskal-Wallis (anova)), Q value = 1
Table S230. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 182 | 75.2 (20.9) |
| subtype1 | 34 | 76.1 (15.6) |
| subtype2 | 110 | 74.7 (18.8) |
| subtype3 | 38 | 75.9 (29.6) |
Figure S224. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'

P value = 0.671 (Fisher's exact test), Q value = 1
Table S231. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #13: 'TUMOR_STATUS'
| nPatients | TUMOR FREE | WITH TUMOR |
|---|---|---|
| ALL | 51 | 22 |
| subtype1 | 7 | 5 |
| subtype2 | 32 | 12 |
| subtype3 | 12 | 5 |
Figure S225. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #13: 'TUMOR_STATUS'

P value = 0.274 (Fisher's exact test), Q value = 1
Table S232. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
| nPatients | NIGERIA | RUSSIA | UKRAINE | UNITED STATES | VIETNAM |
|---|---|---|---|---|---|
| ALL | 1 | 11 | 7 | 169 | 12 |
| subtype1 | 0 | 0 | 2 | 34 | 1 |
| subtype2 | 1 | 6 | 4 | 105 | 7 |
| subtype3 | 0 | 5 | 1 | 30 | 4 |
Figure S226. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'

P value = 0.327 (Fisher's exact test), Q value = 1
Table S233. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'
| nPatients | G1 | G2 | G3 | G4 | GX |
|---|---|---|---|---|---|
| ALL | 14 | 94 | 83 | 1 | 5 |
| subtype1 | 5 | 18 | 12 | 0 | 2 |
| subtype2 | 5 | 56 | 55 | 1 | 3 |
| subtype3 | 4 | 20 | 16 | 0 | 0 |
Figure S227. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'
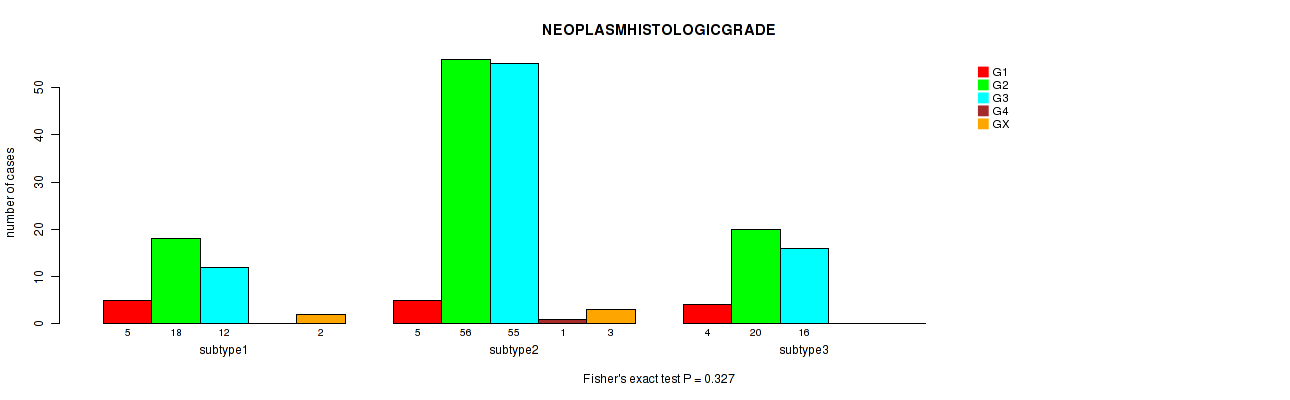
P value = 0.16 (Kruskal-Wallis (anova)), Q value = 1
Table S234. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 27 | 1998.4 (11.5) |
| subtype1 | 4 | 1993.8 (14.8) |
| subtype2 | 18 | 2001.4 (9.8) |
| subtype3 | 5 | 1991.2 (12.9) |
Figure S228. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'

P value = 0.213 (Kruskal-Wallis (anova)), Q value = 1
Table S235. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 62 | 19.4 (13.6) |
| subtype1 | 12 | 14.5 (12.2) |
| subtype2 | 40 | 20.8 (13.9) |
| subtype3 | 10 | 20.0 (13.9) |
Figure S229. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'
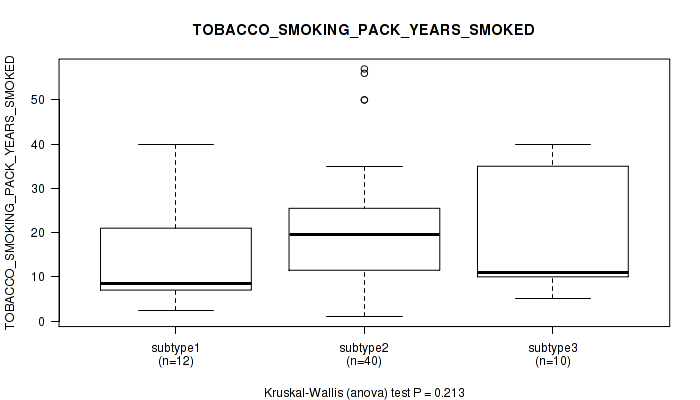
P value = 0.38 (Fisher's exact test), Q value = 1
Table S236. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
| nPatients | CURRENT REFORMED SMOKER FOR < OR = 15 YEARS | CURRENT REFORMED SMOKER FOR > 15 YEARS | CURRENT REFORMED SMOKER, DURATION NOT SPECIFIED | CURRENT SMOKER | LIFELONG NON-SMOKER |
|---|---|---|---|---|---|
| ALL | 25 | 8 | 3 | 43 | 89 |
| subtype1 | 3 | 1 | 0 | 9 | 21 |
| subtype2 | 19 | 3 | 2 | 26 | 48 |
| subtype3 | 3 | 4 | 1 | 8 | 20 |
Figure S230. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'

P value = 0.258 (Kruskal-Wallis (anova)), Q value = 1
Table S237. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 54 | 20.5 (6.2) |
| subtype1 | 11 | 18.7 (4.4) |
| subtype2 | 35 | 20.2 (6.1) |
| subtype3 | 8 | 24.1 (7.8) |
Figure S231. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'
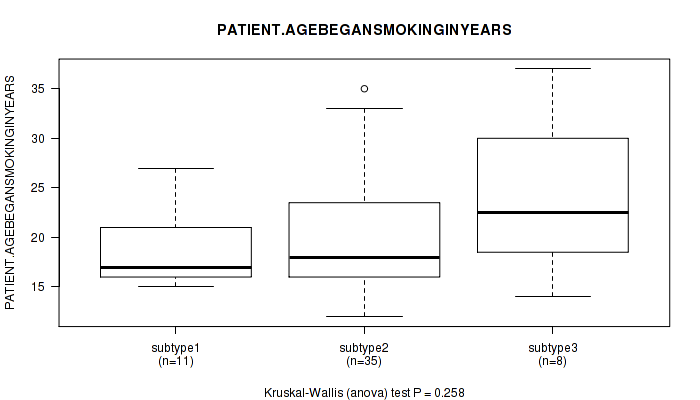
P value = 0.724 (Fisher's exact test), Q value = 1
Table S238. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
| nPatients | COMBINATION | EXTERNAL | EXTERNAL BEAM | IMPLANTS | INTERNAL |
|---|---|---|---|---|---|
| ALL | 18 | 44 | 14 | 1 | 8 |
| subtype1 | 3 | 10 | 1 | 0 | 1 |
| subtype2 | 12 | 27 | 10 | 0 | 6 |
| subtype3 | 3 | 7 | 3 | 1 | 1 |
Figure S232. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'

P value = 0.784 (Fisher's exact test), Q value = 1
Table S239. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
| nPatients | CGY | GY |
|---|---|---|
| ALL | 16 | 4 |
| subtype1 | 4 | 0 |
| subtype2 | 9 | 3 |
| subtype3 | 3 | 1 |
Figure S233. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'

P value = 0.0722 (Kruskal-Wallis (anova)), Q value = 1
Table S240. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 175 | 3.5 (2.4) |
| subtype1 | 33 | 2.8 (2.0) |
| subtype2 | 106 | 3.6 (2.8) |
| subtype3 | 36 | 3.7 (1.6) |
Figure S234. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'

P value = 0.939 (Kruskal-Wallis (anova)), Q value = 1
Table S241. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 100 | 0.1 (0.4) |
| subtype1 | 17 | 0.1 (0.2) |
| subtype2 | 61 | 0.1 (0.4) |
| subtype3 | 22 | 0.0 (0.2) |
Figure S235. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'

P value = 0.102 (Kruskal-Wallis (anova)), Q value = 1
Table S242. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 116 | 0.4 (0.6) |
| subtype1 | 17 | 0.6 (0.6) |
| subtype2 | 73 | 0.3 (0.6) |
| subtype3 | 26 | 0.3 (0.5) |
Figure S236. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'

P value = 0.0486 (Kruskal-Wallis (anova)), Q value = 1
Table S243. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 179 | 2.5 (1.8) |
| subtype1 | 33 | 2.0 (1.8) |
| subtype2 | 107 | 2.6 (1.9) |
| subtype3 | 39 | 2.7 (1.4) |
Figure S237. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'

P value = 0.512 (Kruskal-Wallis (anova)), Q value = 1
Table S244. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 109 | 0.9 (1.9) |
| subtype1 | 19 | 0.6 (0.8) |
| subtype2 | 65 | 1.0 (2.3) |
| subtype3 | 25 | 1.0 (1.3) |
Figure S238. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'

P value = 0.71 (Kruskal-Wallis (anova)), Q value = 1
Table S245. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 101 | 0.1 (0.3) |
| subtype1 | 16 | 0.1 (0.3) |
| subtype2 | 63 | 0.1 (0.3) |
| subtype3 | 22 | 0.1 (0.4) |
Figure S239. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'

P value = 0.562 (Fisher's exact test), Q value = 1
Table S246. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
| nPatients | 2003 | 2010 | COMMON ILIAC | PELVIC (EXTERNAL ILIAC, INTERNAL ILIAC, OBTURATOR) |
|---|---|---|---|---|
| ALL | 1 | 1 | 1 | 41 |
| subtype1 | 0 | 0 | 1 | 7 |
| subtype2 | 1 | 1 | 0 | 24 |
| subtype3 | 0 | 0 | 0 | 10 |
Figure S240. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'

P value = 1 (Fisher's exact test), Q value = 1
Table S247. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'
| nPatients | MACROSCOPIC PARAMETRIAL INVOLVEMENT | MICROSCOPIC PARAMETRIAL INVOLVEMENT | OTHER LOCATION, SPECIFY | POSITIVE BLADDER MARGIN | POSITIVE VAGINAL MARGIN |
|---|---|---|---|---|---|
| ALL | 2 | 7 | 13 | 1 | 6 |
| subtype1 | 1 | 0 | 0 | 0 | 1 |
| subtype2 | 1 | 5 | 9 | 1 | 4 |
| subtype3 | 0 | 2 | 4 | 0 | 1 |
Figure S241. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'

P value = 0.493 (Fisher's exact test), Q value = 1
Table S248. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
| nPatients | INDETERMINATE (NEITHER PRE OR POSTMENOPAUSAL) | PERI (6-12 MONTHS SINCE LAST MENSTRUAL PERIOD) | POST (PRIOR BILATERAL OVARIECTOMY OR >12 MO SINCE LMP WITH NO PRIOR HYSTERECTOMY) | PRE (<6 MONTHS SINCE LMP AND NO PRIOR BILATERAL OVARIECTOMY AND NOT ON ESTROGEN REPLACEMENT) |
|---|---|---|---|---|
| ALL | 2 | 10 | 58 | 85 |
| subtype1 | 0 | 1 | 9 | 21 |
| subtype2 | 1 | 5 | 35 | 48 |
| subtype3 | 1 | 4 | 14 | 16 |
Figure S242. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
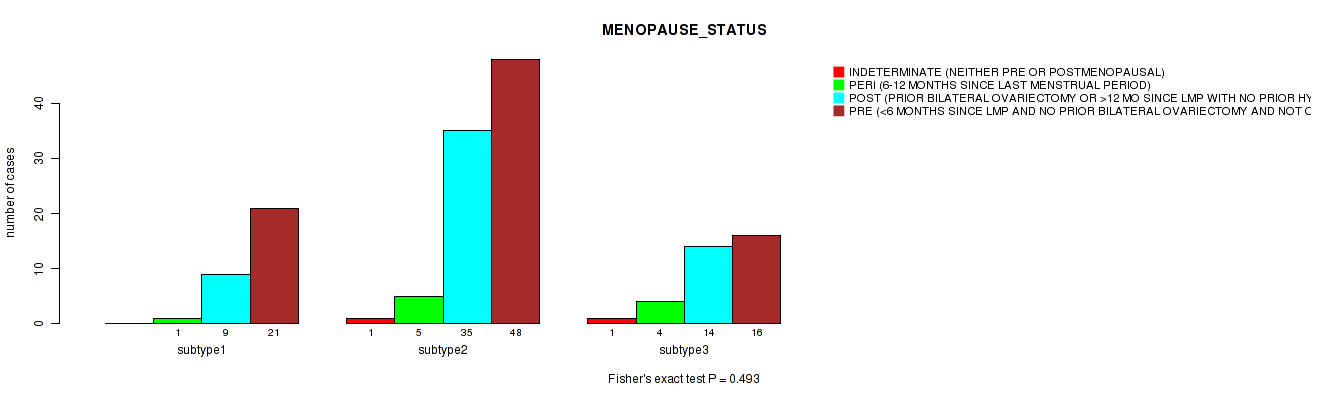
P value = 0.363 (Fisher's exact test), Q value = 1
Table S249. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 56 | 66 |
| subtype1 | 15 | 13 |
| subtype2 | 30 | 33 |
| subtype3 | 11 | 20 |
Figure S243. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'

P value = 0.667 (Kruskal-Wallis (anova)), Q value = 1
Table S250. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 123 | 1.1 (2.6) |
| subtype1 | 25 | 1.3 (3.5) |
| subtype2 | 68 | 0.9 (2.3) |
| subtype3 | 30 | 1.2 (2.1) |
Figure S244. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'

P value = 0.856 (Kruskal-Wallis (anova)), Q value = 1
Table S251. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 143 | 22.4 (12.7) |
| subtype1 | 28 | 21.6 (9.4) |
| subtype2 | 82 | 21.8 (12.7) |
| subtype3 | 33 | 24.5 (15.0) |
Figure S245. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'

P value = 0.0822 (Fisher's exact test), Q value = 1
Table S252. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'
| nPatients | KERATINIZING SQUAMOUS CELL CARCINOMA | NON-KERATINIZING SQUAMOUS CELL CARCINOMA |
|---|---|---|
| ALL | 43 | 88 |
| subtype1 | 0 | 9 |
| subtype2 | 32 | 59 |
| subtype3 | 11 | 20 |
Figure S246. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'

P value = 0.342 (Kruskal-Wallis (anova)), Q value = 1
Table S253. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 200 | 2007.3 (5.2) |
| subtype1 | 37 | 2008.2 (4.7) |
| subtype2 | 123 | 2007.0 (5.2) |
| subtype3 | 40 | 2007.2 (5.5) |
Figure S247. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'

P value = 0.132 (Fisher's exact test), Q value = 1
Table S254. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
| nPatients | OTHER | RADICAL HYSTERECTOMY | SIMPLE HYSTERECTOMY |
|---|---|---|---|
| ALL | 3 | 129 | 5 |
| subtype1 | 2 | 24 | 2 |
| subtype2 | 1 | 69 | 3 |
| subtype3 | 0 | 36 | 0 |
Figure S248. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
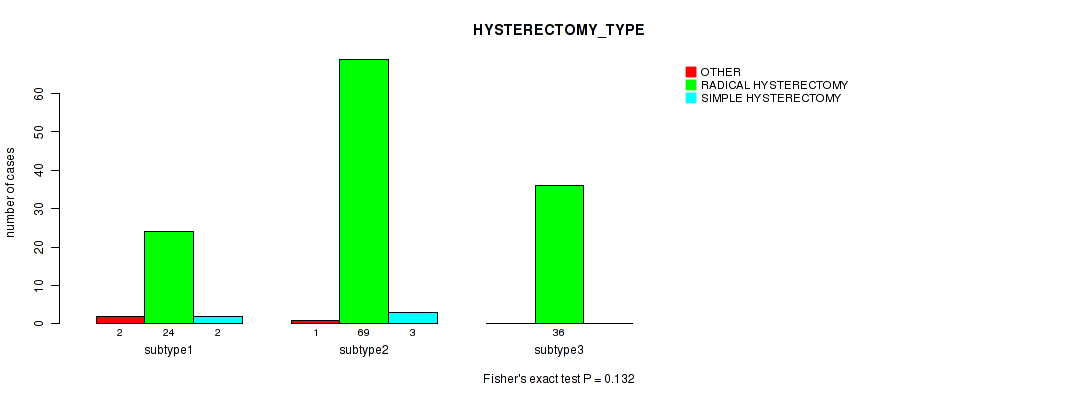
P value = 0.211 (Fisher's exact test), Q value = 1
Table S255. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'
| nPatients | CURRENT USER | FORMER USER | NEVER USED |
|---|---|---|---|
| ALL | 8 | 40 | 47 |
| subtype1 | 4 | 11 | 7 |
| subtype2 | 3 | 23 | 29 |
| subtype3 | 1 | 6 | 11 |
Figure S249. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'

P value = 0.678 (Kruskal-Wallis (anova)), Q value = 1
Table S256. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 173 | 162.1 (7.2) |
| subtype1 | 33 | 163.2 (7.3) |
| subtype2 | 105 | 161.8 (7.3) |
| subtype3 | 35 | 161.8 (7.0) |
Figure S250. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'

P value = 0.659 (Fisher's exact test), Q value = 1
Table S257. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 81 | 15 |
| subtype1 | 21 | 2 |
| subtype2 | 40 | 9 |
| subtype3 | 20 | 4 |
Figure S251. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'

P value = 0.103 (Fisher's exact test), Q value = 1
Table S258. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'
| nPatients | CARBOPLATIN | CISPLATIN |
|---|---|---|
| ALL | 3 | 17 |
| subtype1 | 0 | 2 |
| subtype2 | 1 | 13 |
| subtype3 | 2 | 2 |
Figure S252. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'

P value = 0.362 (Kruskal-Wallis (anova)), Q value = 1
Table S259. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #41: 'CERVIX_SUV_RESULTS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 10 | 12.8 (6.1) |
| subtype2 | 7 | 14.2 (6.6) |
| subtype3 | 3 | 9.7 (3.8) |
Figure S253. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #41: 'CERVIX_SUV_RESULTS'

P value = 0.113 (Fisher's exact test), Q value = 1
Table S260. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'
| nPatients | T1A | T1A1 | T1B | T1B1 | T1B2 | T2 | T2A | T2A1 | T2A2 | T2B | T3B | T4 | TIS | TX |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 2 | 1 | 26 | 55 | 24 | 3 | 6 | 7 | 9 | 14 | 2 | 2 | 1 | 8 |
| subtype1 | 0 | 0 | 1 | 16 | 5 | 0 | 2 | 3 | 1 | 1 | 1 | 0 | 0 | 1 |
| subtype2 | 1 | 0 | 17 | 23 | 15 | 3 | 2 | 3 | 8 | 11 | 1 | 2 | 1 | 6 |
| subtype3 | 1 | 1 | 8 | 16 | 4 | 0 | 2 | 1 | 0 | 2 | 0 | 0 | 0 | 1 |
Figure S254. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'

P value = 0.156 (Kruskal-Wallis (anova)), Q value = 1
Table S261. Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 200 | 47.6 (13.4) |
| subtype1 | 37 | 43.6 (10.6) |
| subtype2 | 123 | 48.1 (13.6) |
| subtype3 | 40 | 49.4 (14.6) |
Figure S255. Get High-res Image Clustering Approach #6: 'MIRSEQ CHIERARCHICAL' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
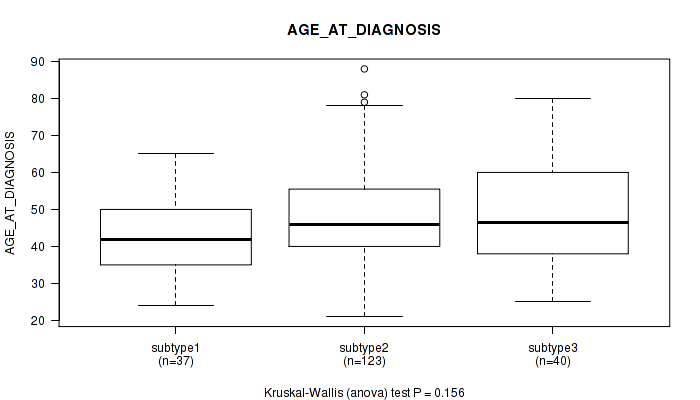
Table S262. Description of clustering approach #7: 'MIRseq Mature CNMF subtypes'
| Cluster Labels | 1 | 2 | 3 |
|---|---|---|---|
| Number of samples | 64 | 65 | 71 |
P value = 0.829 (logrank test), Q value = 1
Table S263. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 193 | 38 | 0.0 - 182.9 (13.9) |
| subtype1 | 63 | 10 | 0.1 - 182.9 (13.1) |
| subtype2 | 61 | 13 | 0.0 - 147.4 (13.9) |
| subtype3 | 69 | 15 | 0.0 - 177.0 (14.5) |
Figure S256. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #1: 'Time to Death'

P value = 0.0366 (Kruskal-Wallis (anova)), Q value = 1
Table S264. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 199 | 47.5 (13.4) |
| subtype1 | 64 | 47.8 (11.1) |
| subtype2 | 65 | 50.4 (14.3) |
| subtype3 | 70 | 44.5 (14.0) |
Figure S257. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #2: 'AGE'
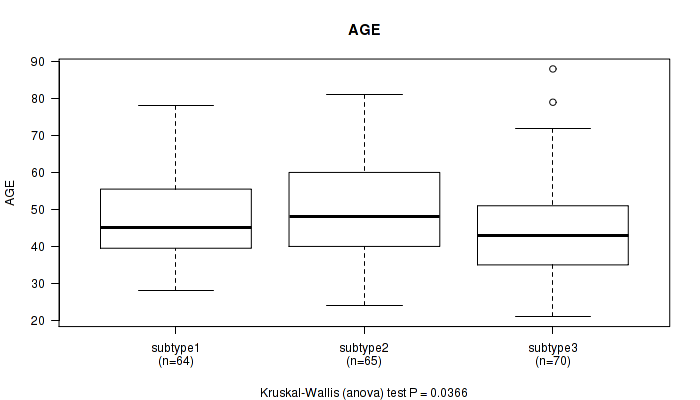
P value = 0.785 (Fisher's exact test), Q value = 1
Table S265. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
| nPatients | T1 | T2 | T3+T4 |
|---|---|---|---|
| ALL | 108 | 39 | 4 |
| subtype1 | 37 | 17 | 1 |
| subtype2 | 35 | 12 | 2 |
| subtype3 | 36 | 10 | 1 |
Figure S258. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
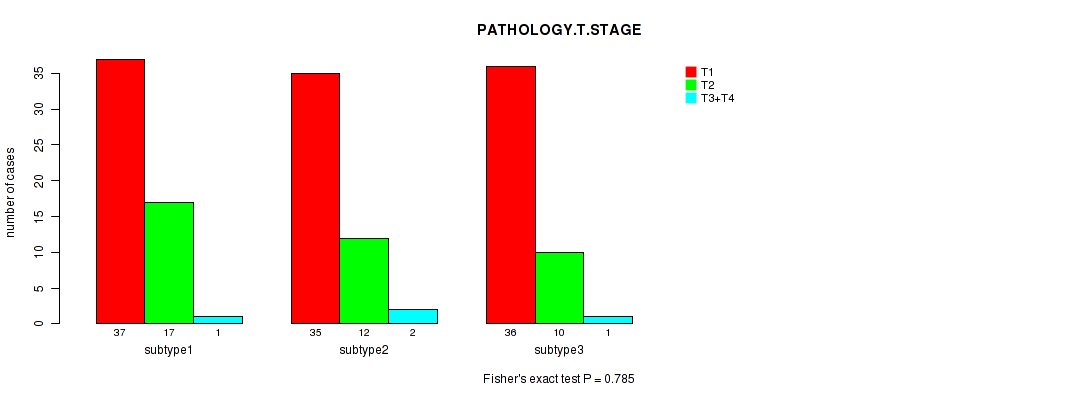
P value = 0.727 (Fisher's exact test), Q value = 1
Table S266. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 100 | 44 |
| subtype1 | 37 | 14 |
| subtype2 | 32 | 17 |
| subtype3 | 31 | 13 |
Figure S259. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'
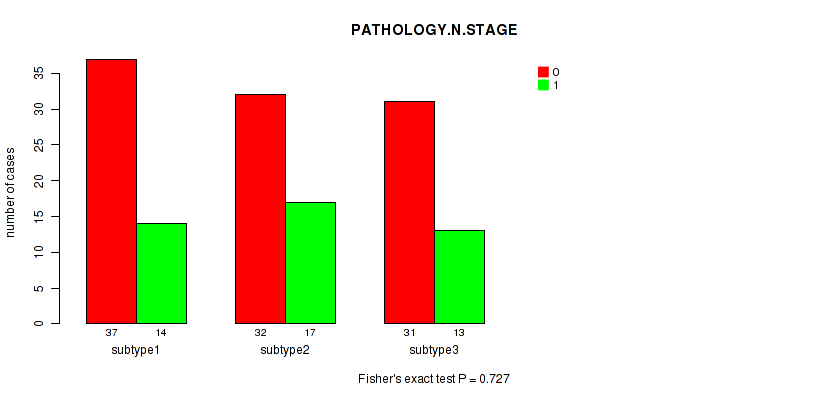
P value = 0.407 (Fisher's exact test), Q value = 1
Table S267. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'
| nPatients | M0 | M1 | MX |
|---|---|---|---|
| ALL | 83 | 4 | 70 |
| subtype1 | 26 | 3 | 28 |
| subtype2 | 30 | 0 | 22 |
| subtype3 | 27 | 1 | 20 |
Figure S260. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'

P value = 1e-05 (Fisher's exact test), Q value = 0.0034
Table S268. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
| nPatients | ADENOSQUAMOUS | CERVICAL SQUAMOUS CELL CARCINOMA | ENDOCERVICAL ADENOCARCINOMA OF THE USUAL TYPE | ENDOCERVICAL TYPE OF ADENOCARCINOMA | ENDOMETRIOID ADENOCARCINOMA OF ENDOCERVIX | MUCINOUS ADENOCARCINOMA OF ENDOCERVICAL TYPE |
|---|---|---|---|---|---|---|
| ALL | 2 | 168 | 4 | 20 | 2 | 4 |
| subtype1 | 2 | 37 | 4 | 16 | 2 | 3 |
| subtype2 | 0 | 60 | 0 | 4 | 0 | 1 |
| subtype3 | 0 | 71 | 0 | 0 | 0 | 0 |
Figure S261. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
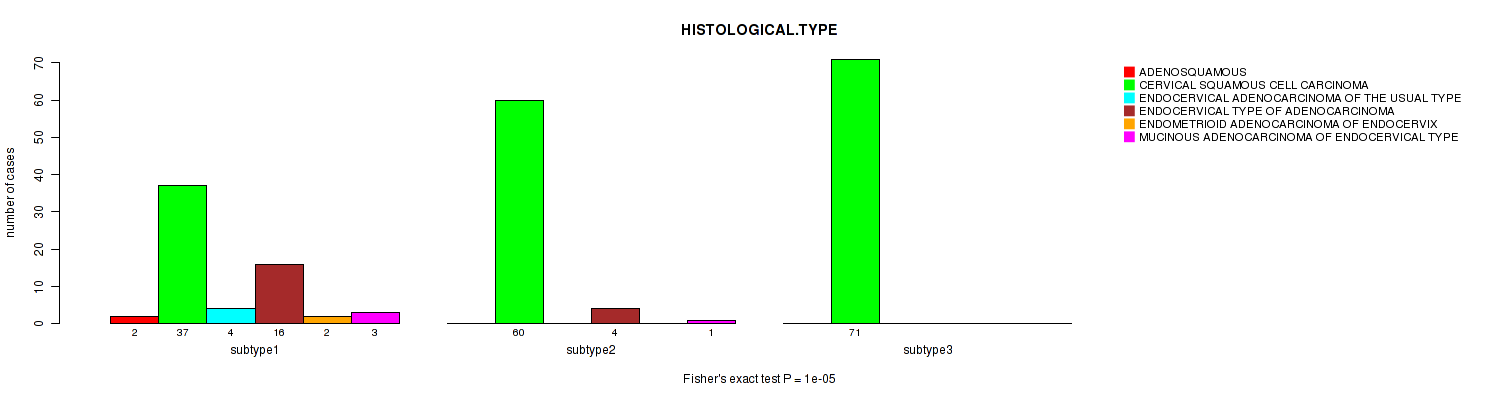
P value = 0.416 (Fisher's exact test), Q value = 1
Table S269. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 32 | 168 |
| subtype1 | 7 | 57 |
| subtype2 | 11 | 54 |
| subtype3 | 14 | 57 |
Figure S262. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'

P value = 0.395 (Kruskal-Wallis (anova)), Q value = 1
Table S270. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 62 | 19.4 (13.6) |
| subtype1 | 18 | 17.2 (15.5) |
| subtype2 | 22 | 20.3 (11.2) |
| subtype3 | 22 | 20.4 (14.5) |
Figure S263. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'

P value = 0.654 (Kruskal-Wallis (anova)), Q value = 1
Table S271. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 123 | 1.1 (2.6) |
| subtype1 | 40 | 0.9 (2.6) |
| subtype2 | 48 | 1.3 (2.4) |
| subtype3 | 35 | 0.9 (2.7) |
Figure S264. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'

P value = 0.156 (Fisher's exact test), Q value = 1
Table S272. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #10: 'RACE'
| nPatients | AMERICAN INDIAN OR ALASKA NATIVE | ASIAN | BLACK OR AFRICAN AMERICAN | NATIVE HAWAIIAN OR OTHER PACIFIC ISLANDER | WHITE |
|---|---|---|---|---|---|
| ALL | 7 | 16 | 23 | 1 | 148 |
| subtype1 | 0 | 4 | 9 | 0 | 50 |
| subtype2 | 1 | 7 | 5 | 0 | 49 |
| subtype3 | 6 | 5 | 9 | 1 | 49 |
Figure S265. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #10: 'RACE'

P value = 0.335 (Fisher's exact test), Q value = 1
Table S273. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #11: 'ETHNICITY'
| nPatients | HISPANIC OR LATINO | NOT HISPANIC OR LATINO |
|---|---|---|
| ALL | 13 | 144 |
| subtype1 | 4 | 47 |
| subtype2 | 2 | 47 |
| subtype3 | 7 | 50 |
Figure S266. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #11: 'ETHNICITY'

P value = 0.124 (Kruskal-Wallis (anova)), Q value = 1
Table S274. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 182 | 75.2 (20.9) |
| subtype1 | 61 | 76.3 (16.1) |
| subtype2 | 58 | 72.0 (25.7) |
| subtype3 | 63 | 77.1 (19.9) |
Figure S267. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'

P value = 0.566 (Fisher's exact test), Q value = 1
Table S275. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #13: 'TUMOR_STATUS'
| nPatients | TUMOR FREE | WITH TUMOR |
|---|---|---|
| ALL | 51 | 22 |
| subtype1 | 18 | 6 |
| subtype2 | 14 | 9 |
| subtype3 | 19 | 7 |
Figure S268. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #13: 'TUMOR_STATUS'

P value = 0.311 (Fisher's exact test), Q value = 1
Table S276. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
| nPatients | NIGERIA | RUSSIA | UKRAINE | UNITED STATES | VIETNAM |
|---|---|---|---|---|---|
| ALL | 1 | 11 | 7 | 169 | 12 |
| subtype1 | 1 | 2 | 3 | 55 | 3 |
| subtype2 | 0 | 7 | 2 | 50 | 6 |
| subtype3 | 0 | 2 | 2 | 64 | 3 |
Figure S269. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'

P value = 0.0581 (Fisher's exact test), Q value = 1
Table S277. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'
| nPatients | G1 | G2 | G3 | G4 | GX |
|---|---|---|---|---|---|
| ALL | 14 | 94 | 83 | 1 | 5 |
| subtype1 | 6 | 25 | 29 | 0 | 3 |
| subtype2 | 7 | 35 | 22 | 1 | 0 |
| subtype3 | 1 | 34 | 32 | 0 | 2 |
Figure S270. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'
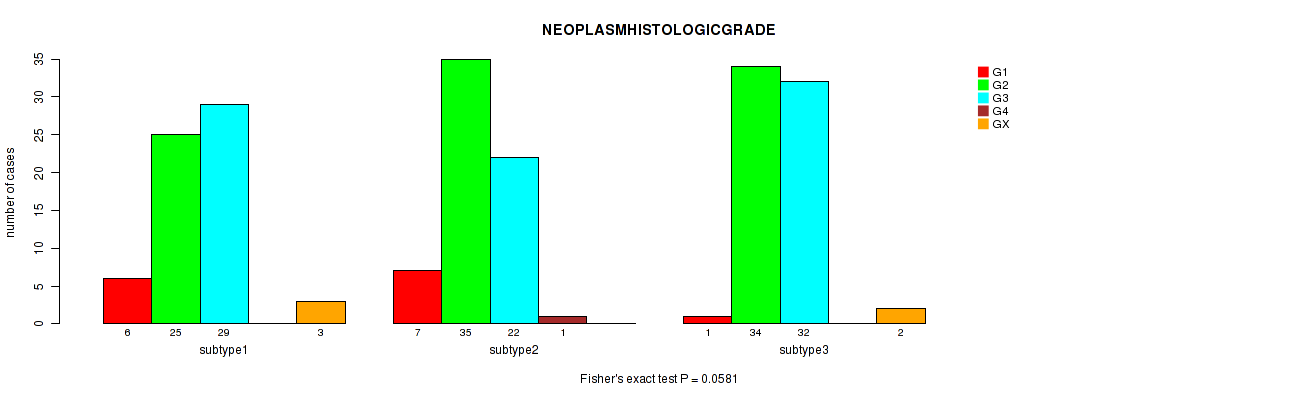
P value = 0.737 (Kruskal-Wallis (anova)), Q value = 1
Table S278. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 27 | 1998.4 (11.5) |
| subtype1 | 7 | 1998.7 (13.2) |
| subtype2 | 10 | 1996.8 (12.1) |
| subtype3 | 10 | 1999.7 (10.8) |
Figure S271. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
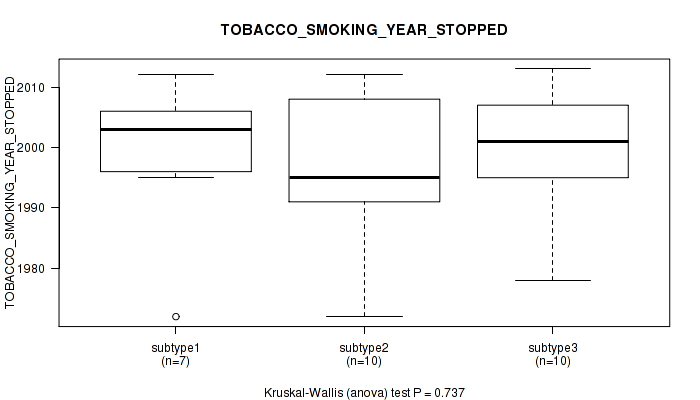
P value = 0.395 (Kruskal-Wallis (anova)), Q value = 1
Table S279. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 62 | 19.4 (13.6) |
| subtype1 | 18 | 17.2 (15.5) |
| subtype2 | 22 | 20.3 (11.2) |
| subtype3 | 22 | 20.4 (14.5) |
Figure S272. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'

P value = 0.762 (Fisher's exact test), Q value = 1
Table S280. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
| nPatients | CURRENT REFORMED SMOKER FOR < OR = 15 YEARS | CURRENT REFORMED SMOKER FOR > 15 YEARS | CURRENT REFORMED SMOKER, DURATION NOT SPECIFIED | CURRENT SMOKER | LIFELONG NON-SMOKER |
|---|---|---|---|---|---|
| ALL | 25 | 8 | 3 | 43 | 89 |
| subtype1 | 7 | 2 | 0 | 11 | 34 |
| subtype2 | 9 | 4 | 2 | 16 | 27 |
| subtype3 | 9 | 2 | 1 | 16 | 28 |
Figure S273. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
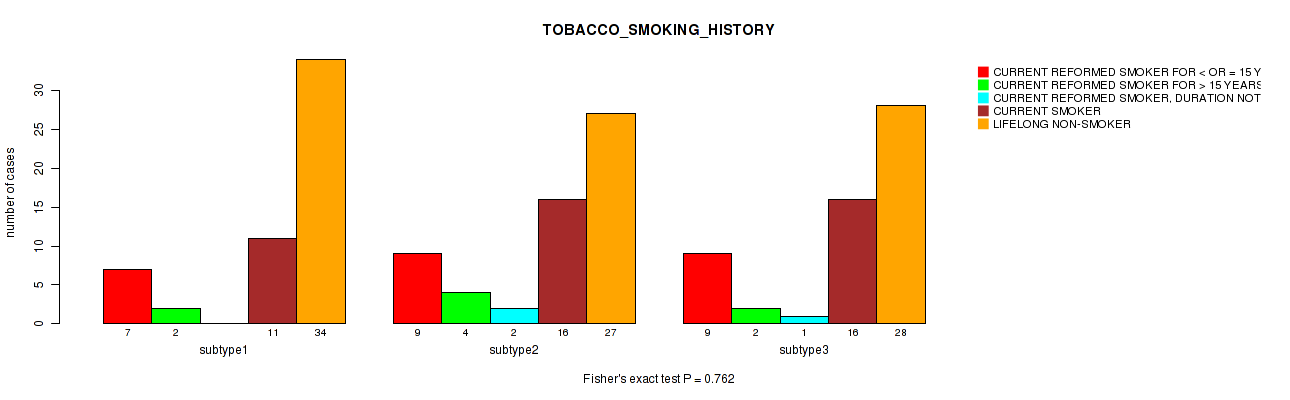
P value = 0.0729 (Kruskal-Wallis (anova)), Q value = 1
Table S281. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 54 | 20.5 (6.2) |
| subtype1 | 16 | 21.3 (6.7) |
| subtype2 | 19 | 22.1 (6.5) |
| subtype3 | 19 | 18.1 (4.8) |
Figure S274. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'

P value = 0.13 (Fisher's exact test), Q value = 1
Table S282. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
| nPatients | COMBINATION | EXTERNAL | EXTERNAL BEAM | IMPLANTS | INTERNAL |
|---|---|---|---|---|---|
| ALL | 18 | 44 | 14 | 1 | 8 |
| subtype1 | 5 | 19 | 2 | 0 | 0 |
| subtype2 | 5 | 10 | 6 | 1 | 4 |
| subtype3 | 8 | 15 | 6 | 0 | 4 |
Figure S275. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
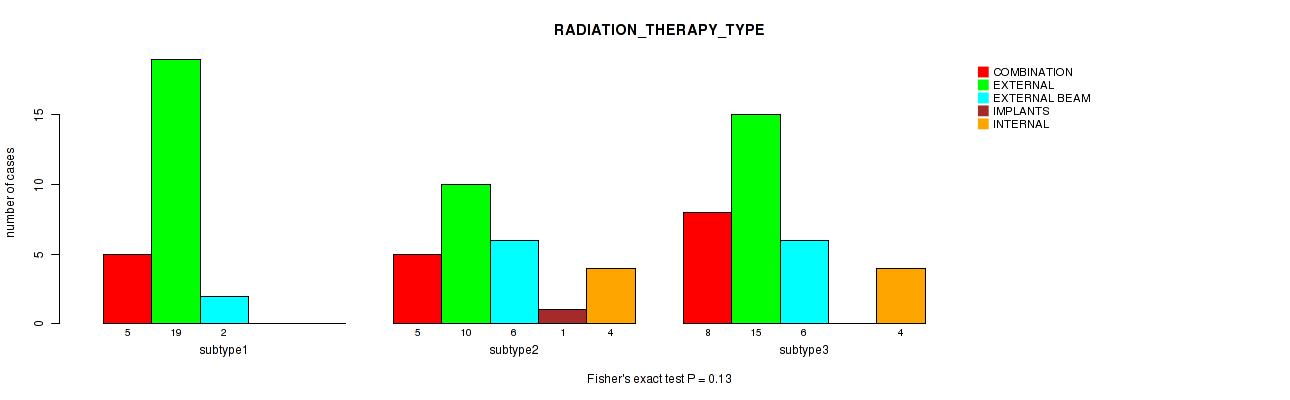
P value = 0.417 (Fisher's exact test), Q value = 1
Table S283. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
| nPatients | CGY | GY |
|---|---|---|
| ALL | 16 | 4 |
| subtype1 | 5 | 0 |
| subtype2 | 5 | 1 |
| subtype3 | 6 | 3 |
Figure S276. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'

P value = 0.464 (Kruskal-Wallis (anova)), Q value = 1
Table S284. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 175 | 3.5 (2.4) |
| subtype1 | 55 | 3.3 (2.5) |
| subtype2 | 60 | 3.6 (2.3) |
| subtype3 | 60 | 3.6 (2.6) |
Figure S277. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
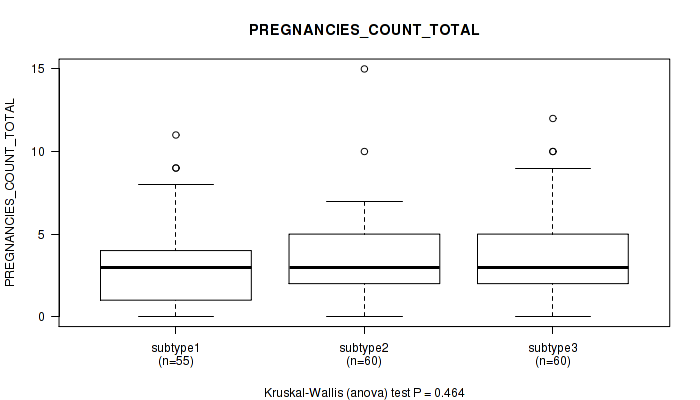
P value = 0.951 (Kruskal-Wallis (anova)), Q value = 1
Table S285. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 100 | 0.1 (0.4) |
| subtype1 | 29 | 0.1 (0.3) |
| subtype2 | 32 | 0.1 (0.6) |
| subtype3 | 39 | 0.1 (0.2) |
Figure S278. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'

P value = 0.217 (Kruskal-Wallis (anova)), Q value = 1
Table S286. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 116 | 0.4 (0.6) |
| subtype1 | 31 | 0.5 (0.6) |
| subtype2 | 41 | 0.3 (0.5) |
| subtype3 | 44 | 0.3 (0.6) |
Figure S279. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'

P value = 0.618 (Kruskal-Wallis (anova)), Q value = 1
Table S287. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 179 | 2.5 (1.8) |
| subtype1 | 56 | 2.4 (1.9) |
| subtype2 | 61 | 2.5 (1.7) |
| subtype3 | 62 | 2.5 (1.8) |
Figure S280. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'

P value = 0.147 (Kruskal-Wallis (anova)), Q value = 1
Table S288. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 109 | 0.9 (1.9) |
| subtype1 | 31 | 0.8 (1.5) |
| subtype2 | 36 | 1.2 (2.3) |
| subtype3 | 42 | 0.7 (1.7) |
Figure S281. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
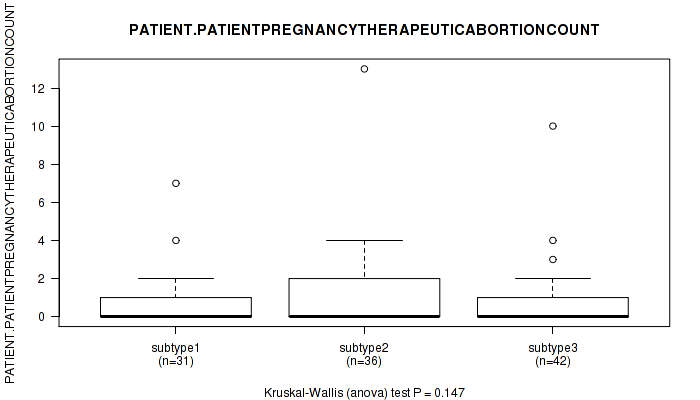
P value = 0.911 (Kruskal-Wallis (anova)), Q value = 1
Table S289. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 101 | 0.1 (0.3) |
| subtype1 | 29 | 0.1 (0.3) |
| subtype2 | 32 | 0.1 (0.4) |
| subtype3 | 40 | 0.1 (0.3) |
Figure S282. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'

P value = 0.131 (Fisher's exact test), Q value = 1
Table S290. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
| nPatients | 2003 | 2010 | COMMON ILIAC | PELVIC (EXTERNAL ILIAC, INTERNAL ILIAC, OBTURATOR) |
|---|---|---|---|---|
| ALL | 1 | 1 | 1 | 41 |
| subtype1 | 0 | 0 | 1 | 13 |
| subtype2 | 0 | 0 | 0 | 17 |
| subtype3 | 1 | 1 | 0 | 11 |
Figure S283. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'

P value = 0.0317 (Fisher's exact test), Q value = 1
Table S291. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'
| nPatients | MACROSCOPIC PARAMETRIAL INVOLVEMENT | MICROSCOPIC PARAMETRIAL INVOLVEMENT | OTHER LOCATION, SPECIFY | POSITIVE BLADDER MARGIN | POSITIVE VAGINAL MARGIN |
|---|---|---|---|---|---|
| ALL | 2 | 7 | 13 | 1 | 6 |
| subtype1 | 1 | 1 | 0 | 0 | 3 |
| subtype2 | 1 | 3 | 5 | 1 | 3 |
| subtype3 | 0 | 3 | 8 | 0 | 0 |
Figure S284. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'

P value = 0.471 (Fisher's exact test), Q value = 1
Table S292. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
| nPatients | INDETERMINATE (NEITHER PRE OR POSTMENOPAUSAL) | PERI (6-12 MONTHS SINCE LAST MENSTRUAL PERIOD) | POST (PRIOR BILATERAL OVARIECTOMY OR >12 MO SINCE LMP WITH NO PRIOR HYSTERECTOMY) | PRE (<6 MONTHS SINCE LMP AND NO PRIOR BILATERAL OVARIECTOMY AND NOT ON ESTROGEN REPLACEMENT) |
|---|---|---|---|---|
| ALL | 2 | 10 | 58 | 85 |
| subtype1 | 1 | 4 | 18 | 26 |
| subtype2 | 1 | 3 | 24 | 24 |
| subtype3 | 0 | 3 | 16 | 35 |
Figure S285. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
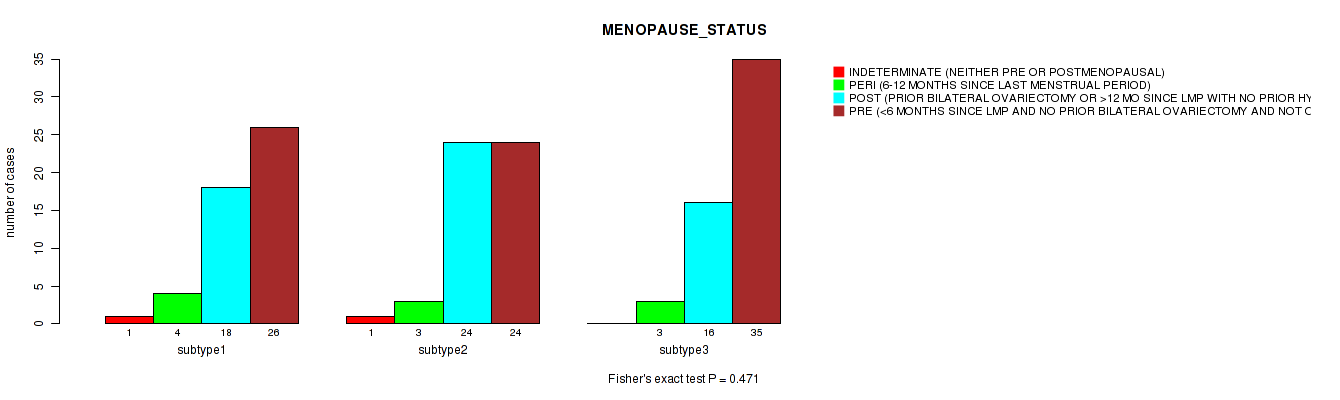
P value = 0.665 (Fisher's exact test), Q value = 1
Table S293. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 56 | 66 |
| subtype1 | 19 | 27 |
| subtype2 | 20 | 23 |
| subtype3 | 17 | 16 |
Figure S286. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'

P value = 0.654 (Kruskal-Wallis (anova)), Q value = 1
Table S294. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 123 | 1.1 (2.6) |
| subtype1 | 40 | 0.9 (2.6) |
| subtype2 | 48 | 1.3 (2.4) |
| subtype3 | 35 | 0.9 (2.7) |
Figure S287. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'
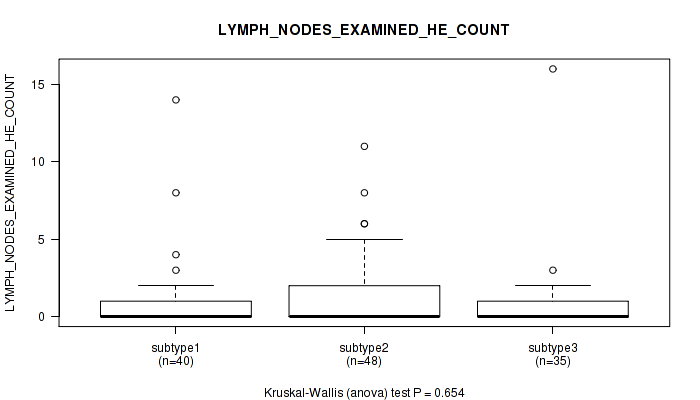
P value = 0.179 (Kruskal-Wallis (anova)), Q value = 1
Table S295. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 143 | 22.4 (12.7) |
| subtype1 | 49 | 20.7 (11.0) |
| subtype2 | 51 | 25.6 (14.2) |
| subtype3 | 43 | 20.5 (12.0) |
Figure S288. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'

P value = 0.0205 (Fisher's exact test), Q value = 1
Table S296. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'
| nPatients | KERATINIZING SQUAMOUS CELL CARCINOMA | NON-KERATINIZING SQUAMOUS CELL CARCINOMA |
|---|---|---|
| ALL | 43 | 88 |
| subtype1 | 4 | 26 |
| subtype2 | 20 | 27 |
| subtype3 | 19 | 35 |
Figure S289. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'

P value = 0.222 (Kruskal-Wallis (anova)), Q value = 1
Table S297. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 200 | 2007.3 (5.2) |
| subtype1 | 64 | 2007.9 (5.2) |
| subtype2 | 65 | 2007.4 (5.1) |
| subtype3 | 71 | 2006.7 (5.3) |
Figure S290. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
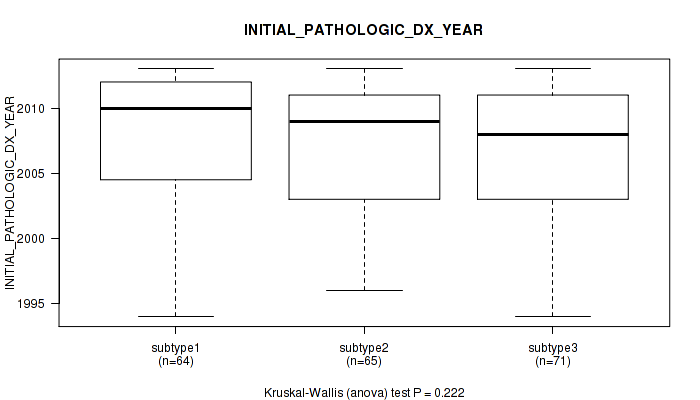
P value = 0.307 (Fisher's exact test), Q value = 1
Table S298. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
| nPatients | OTHER | RADICAL HYSTERECTOMY | SIMPLE HYSTERECTOMY |
|---|---|---|---|
| ALL | 3 | 129 | 5 |
| subtype1 | 3 | 46 | 2 |
| subtype2 | 0 | 49 | 1 |
| subtype3 | 0 | 34 | 2 |
Figure S291. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
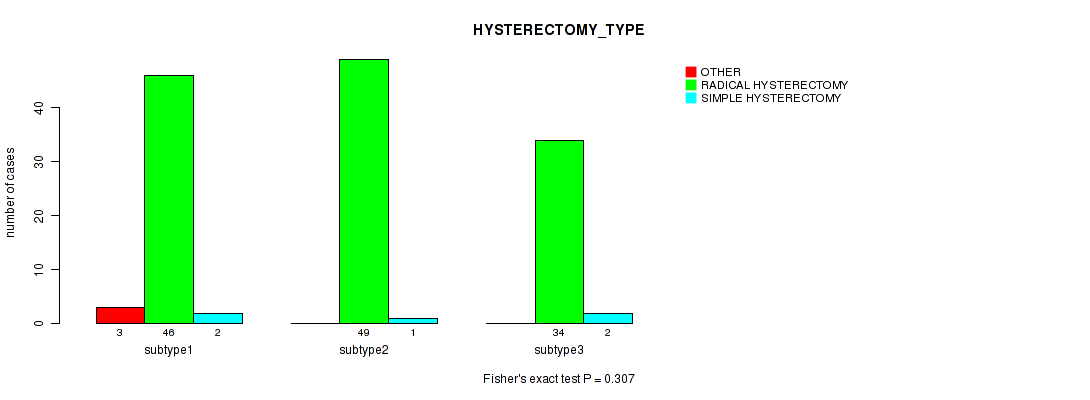
P value = 0.931 (Fisher's exact test), Q value = 1
Table S299. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'
| nPatients | CURRENT USER | FORMER USER | NEVER USED |
|---|---|---|---|
| ALL | 8 | 40 | 47 |
| subtype1 | 4 | 14 | 15 |
| subtype2 | 2 | 13 | 16 |
| subtype3 | 2 | 13 | 16 |
Figure S292. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'

P value = 0.271 (Kruskal-Wallis (anova)), Q value = 1
Table S300. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 173 | 162.1 (7.2) |
| subtype1 | 58 | 163.4 (6.4) |
| subtype2 | 53 | 161.1 (8.5) |
| subtype3 | 62 | 161.7 (6.6) |
Figure S293. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'

P value = 0.124 (Fisher's exact test), Q value = 1
Table S301. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 81 | 15 |
| subtype1 | 29 | 6 |
| subtype2 | 27 | 8 |
| subtype3 | 25 | 1 |
Figure S294. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
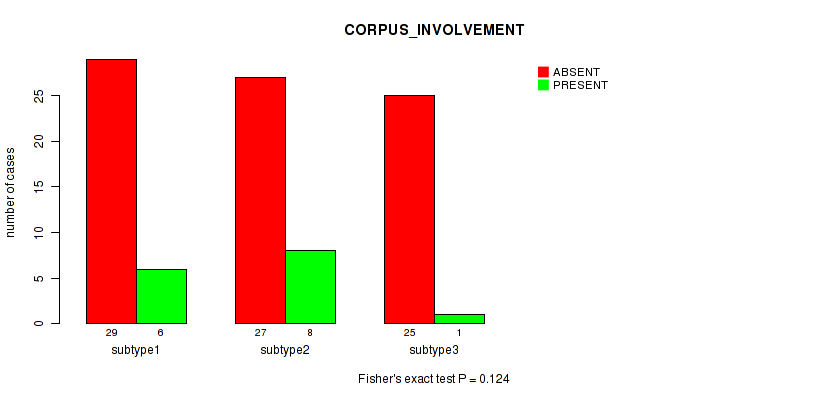
P value = 0.537 (Fisher's exact test), Q value = 1
Table S302. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'
| nPatients | CARBOPLATIN | CISPLATIN |
|---|---|---|
| ALL | 3 | 17 |
| subtype1 | 0 | 3 |
| subtype2 | 2 | 4 |
| subtype3 | 1 | 10 |
Figure S295. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'

P value = 0.0947 (Kruskal-Wallis (anova)), Q value = 1
Table S303. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #41: 'CERVIX_SUV_RESULTS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 10 | 12.8 (6.1) |
| subtype2 | 5 | 9.4 (3.4) |
| subtype3 | 5 | 16.3 (6.4) |
Figure S296. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #41: 'CERVIX_SUV_RESULTS'

P value = 0.742 (Fisher's exact test), Q value = 1
Table S304. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'
| nPatients | T1A | T1A1 | T1B | T1B1 | T1B2 | T2 | T2A | T2A1 | T2A2 | T2B | T3B | T4 | TIS | TX |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 2 | 1 | 26 | 55 | 24 | 3 | 6 | 7 | 9 | 14 | 2 | 2 | 1 | 8 |
| subtype1 | 0 | 0 | 6 | 20 | 11 | 2 | 2 | 4 | 5 | 4 | 1 | 0 | 0 | 2 |
| subtype2 | 1 | 1 | 7 | 19 | 7 | 1 | 3 | 2 | 2 | 4 | 0 | 2 | 0 | 3 |
| subtype3 | 1 | 0 | 13 | 16 | 6 | 0 | 1 | 1 | 2 | 6 | 1 | 0 | 1 | 3 |
Figure S297. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'

P value = 0.0504 (Kruskal-Wallis (anova)), Q value = 1
Table S305. Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 200 | 47.6 (13.4) |
| subtype1 | 64 | 47.8 (11.1) |
| subtype2 | 65 | 50.4 (14.3) |
| subtype3 | 71 | 44.7 (14.0) |
Figure S298. Get High-res Image Clustering Approach #7: 'MIRseq Mature CNMF subtypes' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'

Table S306. Description of clustering approach #8: 'MIRseq Mature cHierClus subtypes'
| Cluster Labels | 1 | 2 | 3 | 4 | 5 | 6 | 7 |
|---|---|---|---|---|---|---|---|
| Number of samples | 23 | 40 | 17 | 31 | 29 | 27 | 33 |
P value = 0.147 (logrank test), Q value = 1
Table S307. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'
| nPatients | nDeath | Duration Range (Median), Month | |
|---|---|---|---|
| ALL | 193 | 38 | 0.0 - 182.9 (13.9) |
| subtype1 | 22 | 3 | 0.5 - 97.0 (7.6) |
| subtype2 | 38 | 7 | 0.1 - 182.9 (20.0) |
| subtype3 | 17 | 6 | 2.4 - 177.0 (36.8) |
| subtype4 | 30 | 11 | 0.0 - 118.0 (14.0) |
| subtype5 | 29 | 4 | 0.0 - 173.3 (8.3) |
| subtype6 | 26 | 4 | 0.1 - 137.2 (4.3) |
| subtype7 | 31 | 3 | 0.0 - 147.4 (15.0) |
Figure S299. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #1: 'Time to Death'

P value = 0.0118 (Kruskal-Wallis (anova)), Q value = 1
Table S308. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #2: 'AGE'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 199 | 47.5 (13.4) |
| subtype1 | 23 | 42.6 (11.2) |
| subtype2 | 39 | 48.9 (11.2) |
| subtype3 | 17 | 49.8 (16.2) |
| subtype4 | 31 | 41.1 (13.7) |
| subtype5 | 29 | 51.4 (13.9) |
| subtype6 | 27 | 46.1 (9.0) |
| subtype7 | 33 | 51.7 (15.3) |
Figure S300. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #2: 'AGE'
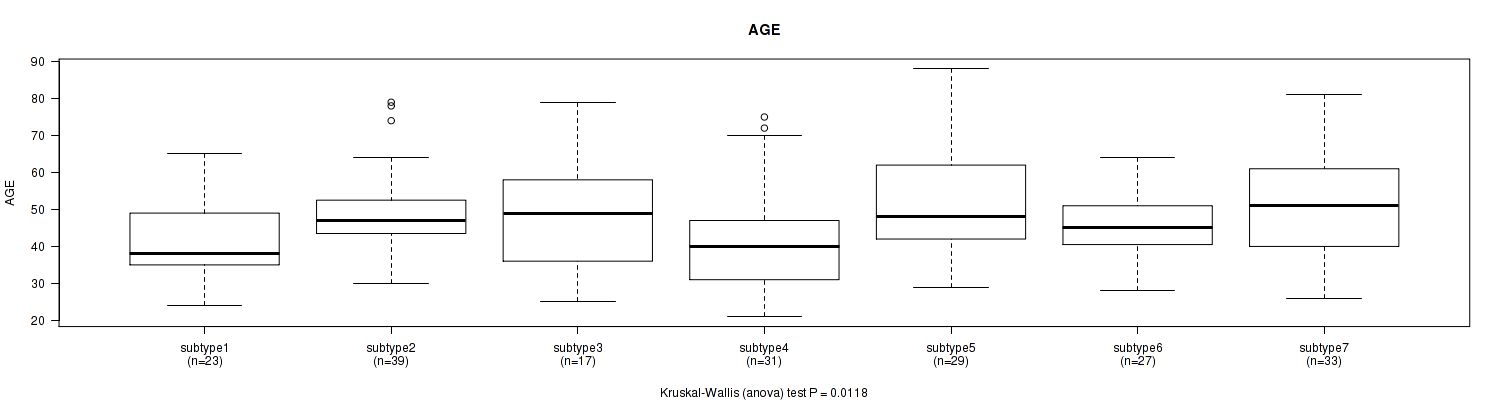
P value = 0.436 (Fisher's exact test), Q value = 1
Table S309. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
| nPatients | T1 | T2 | T3+T4 |
|---|---|---|---|
| ALL | 108 | 39 | 4 |
| subtype1 | 14 | 5 | 0 |
| subtype2 | 18 | 9 | 2 |
| subtype3 | 12 | 2 | 0 |
| subtype4 | 14 | 3 | 2 |
| subtype5 | 14 | 8 | 0 |
| subtype6 | 14 | 7 | 0 |
| subtype7 | 22 | 5 | 0 |
Figure S301. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #3: 'PATHOLOGY.T.STAGE'
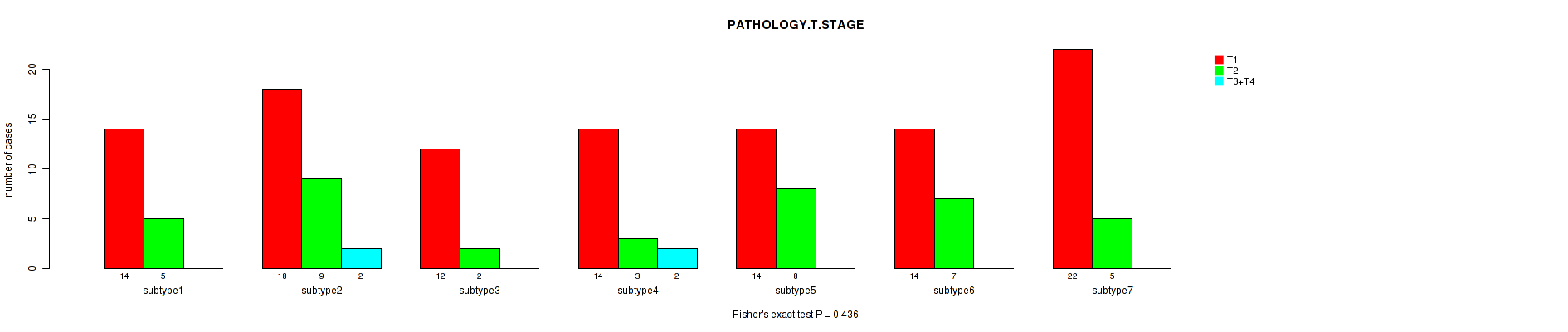
P value = 0.631 (Fisher's exact test), Q value = 1
Table S310. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'
| nPatients | 0 | 1 |
|---|---|---|
| ALL | 100 | 44 |
| subtype1 | 13 | 4 |
| subtype2 | 19 | 10 |
| subtype3 | 7 | 7 |
| subtype4 | 14 | 5 |
| subtype5 | 13 | 7 |
| subtype6 | 16 | 4 |
| subtype7 | 18 | 7 |
Figure S302. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #4: 'PATHOLOGY.N.STAGE'

P value = 0.643 (Fisher's exact test), Q value = 1
Table S311. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'
| nPatients | M0 | M1 | MX |
|---|---|---|---|
| ALL | 83 | 4 | 70 |
| subtype1 | 7 | 1 | 11 |
| subtype2 | 18 | 1 | 14 |
| subtype3 | 9 | 0 | 5 |
| subtype4 | 10 | 1 | 9 |
| subtype5 | 9 | 0 | 13 |
| subtype6 | 14 | 1 | 7 |
| subtype7 | 16 | 0 | 11 |
Figure S303. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #5: 'PATHOLOGY.M.STAGE'

P value = 1e-05 (Fisher's exact test), Q value = 0.0034
Table S312. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'
| nPatients | ADENOSQUAMOUS | CERVICAL SQUAMOUS CELL CARCINOMA | ENDOCERVICAL ADENOCARCINOMA OF THE USUAL TYPE | ENDOCERVICAL TYPE OF ADENOCARCINOMA | ENDOMETRIOID ADENOCARCINOMA OF ENDOCERVIX | MUCINOUS ADENOCARCINOMA OF ENDOCERVICAL TYPE |
|---|---|---|---|---|---|---|
| ALL | 2 | 168 | 4 | 20 | 2 | 4 |
| subtype1 | 0 | 1 | 4 | 14 | 0 | 4 |
| subtype2 | 0 | 39 | 0 | 1 | 0 | 0 |
| subtype3 | 0 | 17 | 0 | 0 | 0 | 0 |
| subtype4 | 0 | 30 | 0 | 1 | 0 | 0 |
| subtype5 | 0 | 29 | 0 | 0 | 0 | 0 |
| subtype6 | 2 | 19 | 0 | 4 | 2 | 0 |
| subtype7 | 0 | 33 | 0 | 0 | 0 | 0 |
Figure S304. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #6: 'HISTOLOGICAL.TYPE'

P value = 0.0401 (Fisher's exact test), Q value = 1
Table S313. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'
| nPatients | NO | YES |
|---|---|---|
| ALL | 32 | 168 |
| subtype1 | 1 | 22 |
| subtype2 | 9 | 31 |
| subtype3 | 7 | 10 |
| subtype4 | 6 | 25 |
| subtype5 | 3 | 26 |
| subtype6 | 2 | 25 |
| subtype7 | 4 | 29 |
Figure S305. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #7: 'RADIATIONS.RADIATION.REGIMENINDICATION'

P value = 0.491 (Kruskal-Wallis (anova)), Q value = 1
Table S314. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 62 | 19.4 (13.6) |
| subtype1 | 8 | 13.1 (12.3) |
| subtype2 | 11 | 18.9 (16.1) |
| subtype3 | 6 | 16.3 (5.8) |
| subtype4 | 9 | 17.2 (9.7) |
| subtype5 | 11 | 27.5 (18.1) |
| subtype6 | 6 | 19.2 (10.6) |
| subtype7 | 11 | 20.3 (13.0) |
Figure S306. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #8: 'NUMBERPACKYEARSSMOKED'

P value = 0.485 (Kruskal-Wallis (anova)), Q value = 1
Table S315. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 123 | 1.1 (2.6) |
| subtype1 | 16 | 1.1 (2.8) |
| subtype2 | 29 | 1.6 (3.4) |
| subtype3 | 13 | 1.7 (2.7) |
| subtype4 | 15 | 1.3 (3.6) |
| subtype5 | 14 | 0.7 (1.1) |
| subtype6 | 13 | 0.2 (0.4) |
| subtype7 | 23 | 0.7 (1.4) |
Figure S307. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #9: 'NUMBER.OF.LYMPH.NODES'
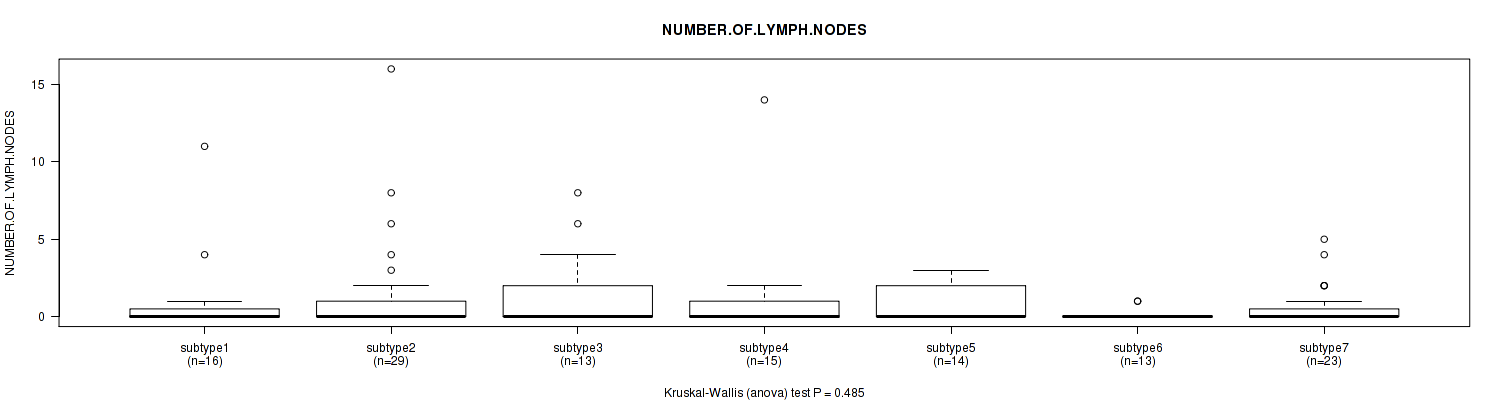
P value = 0.632 (Fisher's exact test), Q value = 1
Table S316. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #10: 'RACE'
| nPatients | AMERICAN INDIAN OR ALASKA NATIVE | ASIAN | BLACK OR AFRICAN AMERICAN | NATIVE HAWAIIAN OR OTHER PACIFIC ISLANDER | WHITE |
|---|---|---|---|---|---|
| ALL | 7 | 16 | 23 | 1 | 148 |
| subtype1 | 0 | 1 | 1 | 0 | 21 |
| subtype2 | 1 | 4 | 4 | 0 | 30 |
| subtype3 | 0 | 2 | 3 | 0 | 12 |
| subtype4 | 3 | 1 | 5 | 1 | 21 |
| subtype5 | 2 | 1 | 6 | 0 | 20 |
| subtype6 | 0 | 3 | 2 | 0 | 21 |
| subtype7 | 1 | 4 | 2 | 0 | 23 |
Figure S308. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #10: 'RACE'
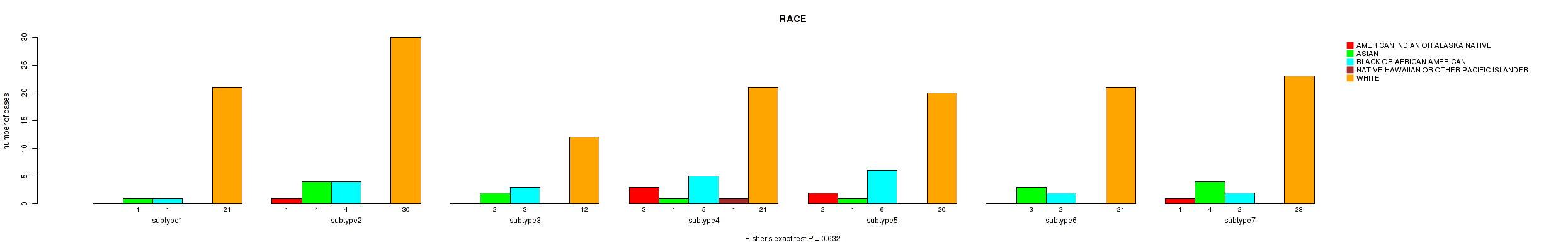
P value = 0.266 (Fisher's exact test), Q value = 1
Table S317. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #11: 'ETHNICITY'
| nPatients | HISPANIC OR LATINO | NOT HISPANIC OR LATINO |
|---|---|---|
| ALL | 13 | 144 |
| subtype1 | 3 | 16 |
| subtype2 | 1 | 29 |
| subtype3 | 0 | 14 |
| subtype4 | 3 | 20 |
| subtype5 | 4 | 19 |
| subtype6 | 1 | 22 |
| subtype7 | 1 | 24 |
Figure S309. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #11: 'ETHNICITY'

P value = 0.407 (Kruskal-Wallis (anova)), Q value = 1
Table S318. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 182 | 75.2 (20.9) |
| subtype1 | 21 | 76.8 (13.7) |
| subtype2 | 38 | 75.7 (18.9) |
| subtype3 | 14 | 64.9 (11.3) |
| subtype4 | 27 | 74.2 (19.2) |
| subtype5 | 27 | 75.0 (20.7) |
| subtype6 | 25 | 77.0 (17.0) |
| subtype7 | 30 | 77.9 (32.5) |
Figure S310. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #12: 'WEIGHT_KG_AT_DIAGNOSIS'
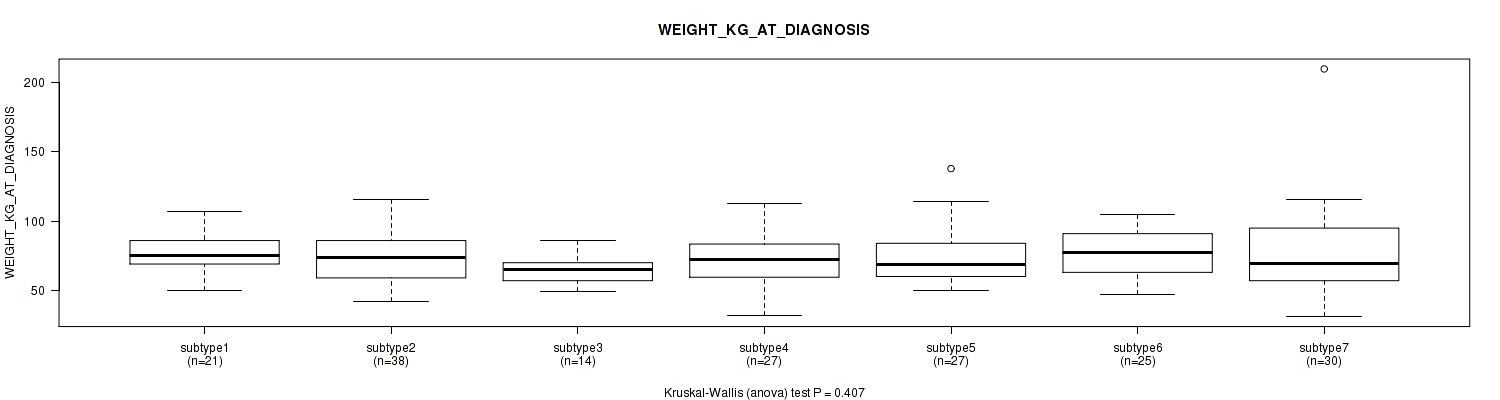
P value = 0.454 (Fisher's exact test), Q value = 1
Table S319. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #13: 'TUMOR_STATUS'
| nPatients | TUMOR FREE | WITH TUMOR |
|---|---|---|
| ALL | 51 | 22 |
| subtype1 | 4 | 1 |
| subtype2 | 14 | 3 |
| subtype3 | 8 | 5 |
| subtype4 | 7 | 5 |
| subtype5 | 6 | 3 |
| subtype6 | 4 | 4 |
| subtype7 | 8 | 1 |
Figure S311. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #13: 'TUMOR_STATUS'

P value = 0.572 (Fisher's exact test), Q value = 1
Table S320. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
| nPatients | NIGERIA | RUSSIA | UKRAINE | UNITED STATES | VIETNAM |
|---|---|---|---|---|---|
| ALL | 1 | 11 | 7 | 169 | 12 |
| subtype1 | 0 | 0 | 0 | 23 | 0 |
| subtype2 | 0 | 3 | 1 | 33 | 3 |
| subtype3 | 0 | 1 | 0 | 16 | 0 |
| subtype4 | 0 | 0 | 2 | 28 | 1 |
| subtype5 | 0 | 2 | 1 | 25 | 1 |
| subtype6 | 1 | 2 | 2 | 19 | 3 |
| subtype7 | 0 | 3 | 1 | 25 | 4 |
Figure S312. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #14: 'TUMOR_SAMPLE_PROCUREMENT_COUNTRY'
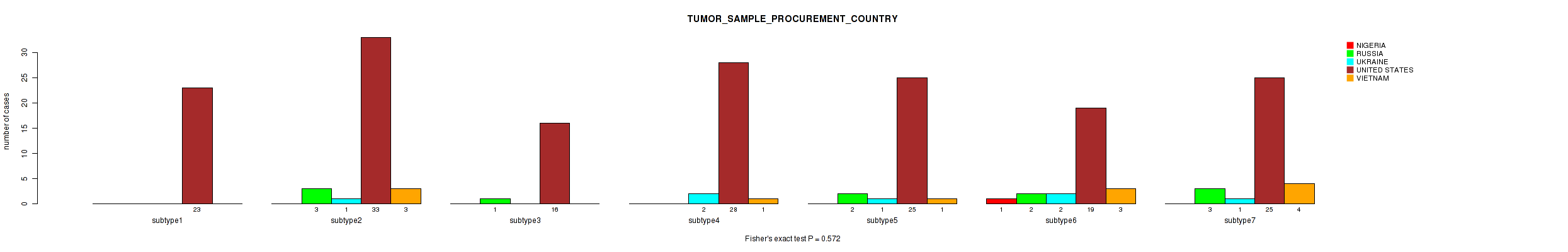
P value = 0.00071 (Fisher's exact test), Q value = 0.24
Table S321. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'
| nPatients | G1 | G2 | G3 | G4 | GX |
|---|---|---|---|---|---|
| ALL | 14 | 94 | 83 | 1 | 5 |
| subtype1 | 5 | 14 | 3 | 0 | 1 |
| subtype2 | 1 | 22 | 12 | 1 | 3 |
| subtype3 | 0 | 8 | 9 | 0 | 0 |
| subtype4 | 2 | 8 | 19 | 0 | 0 |
| subtype5 | 2 | 16 | 11 | 0 | 0 |
| subtype6 | 0 | 8 | 18 | 0 | 1 |
| subtype7 | 4 | 18 | 11 | 0 | 0 |
Figure S313. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #15: 'NEOPLASMHISTOLOGICGRADE'

P value = 0.388 (Kruskal-Wallis (anova)), Q value = 1
Table S322. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 27 | 1998.4 (11.5) |
| subtype1 | 4 | 1993.8 (14.8) |
| subtype2 | 6 | 2001.2 (13.4) |
| subtype3 | 3 | 1998.3 (4.2) |
| subtype4 | 2 | 2001.0 (8.5) |
| subtype5 | 4 | 2002.5 (11.3) |
| subtype6 | 2 | 2009.0 (4.2) |
| subtype7 | 6 | 1991.5 (11.6) |
Figure S314. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #16: 'TOBACCO_SMOKING_YEAR_STOPPED'

P value = 0.491 (Kruskal-Wallis (anova)), Q value = 1
Table S323. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 62 | 19.4 (13.6) |
| subtype1 | 8 | 13.1 (12.3) |
| subtype2 | 11 | 18.9 (16.1) |
| subtype3 | 6 | 16.3 (5.8) |
| subtype4 | 9 | 17.2 (9.7) |
| subtype5 | 11 | 27.5 (18.1) |
| subtype6 | 6 | 19.2 (10.6) |
| subtype7 | 11 | 20.3 (13.0) |
Figure S315. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #17: 'TOBACCO_SMOKING_PACK_YEARS_SMOKED'

P value = 0.899 (Fisher's exact test), Q value = 1
Table S324. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
| nPatients | CURRENT REFORMED SMOKER FOR < OR = 15 YEARS | CURRENT REFORMED SMOKER FOR > 15 YEARS | CURRENT REFORMED SMOKER, DURATION NOT SPECIFIED | CURRENT SMOKER | LIFELONG NON-SMOKER |
|---|---|---|---|---|---|
| ALL | 25 | 8 | 3 | 43 | 89 |
| subtype1 | 3 | 1 | 0 | 5 | 12 |
| subtype2 | 7 | 1 | 1 | 7 | 22 |
| subtype3 | 3 | 2 | 0 | 3 | 8 |
| subtype4 | 2 | 0 | 1 | 8 | 11 |
| subtype5 | 5 | 1 | 0 | 6 | 10 |
| subtype6 | 2 | 0 | 0 | 5 | 14 |
| subtype7 | 3 | 3 | 1 | 9 | 12 |
Figure S316. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #18: 'TOBACCO_SMOKING_HISTORY'
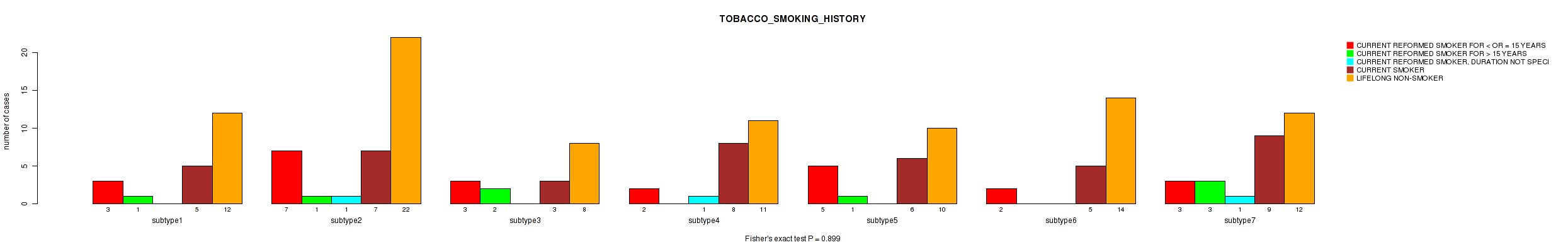
P value = 0.105 (Kruskal-Wallis (anova)), Q value = 1
Table S325. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 54 | 20.5 (6.2) |
| subtype1 | 8 | 18.5 (3.8) |
| subtype2 | 9 | 23.4 (6.0) |
| subtype3 | 5 | 22.2 (7.0) |
| subtype4 | 6 | 16.7 (2.8) |
| subtype5 | 10 | 19.5 (7.9) |
| subtype6 | 6 | 18.2 (4.4) |
| subtype7 | 10 | 23.1 (6.7) |
Figure S317. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #19: 'PATIENT.AGEBEGANSMOKINGINYEARS'

P value = 0.0239 (Fisher's exact test), Q value = 1
Table S326. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'
| nPatients | COMBINATION | EXTERNAL | EXTERNAL BEAM | IMPLANTS | INTERNAL |
|---|---|---|---|---|---|
| ALL | 18 | 44 | 14 | 1 | 8 |
| subtype1 | 1 | 7 | 0 | 0 | 0 |
| subtype2 | 4 | 12 | 5 | 0 | 2 |
| subtype3 | 6 | 0 | 1 | 1 | 0 |
| subtype4 | 3 | 6 | 3 | 0 | 2 |
| subtype5 | 2 | 7 | 1 | 0 | 1 |
| subtype6 | 2 | 6 | 0 | 0 | 1 |
| subtype7 | 0 | 6 | 4 | 0 | 2 |
Figure S318. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #20: 'RADIATION_THERAPY_TYPE'

P value = 0.94 (Fisher's exact test), Q value = 1
Table S327. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
| nPatients | CGY | GY |
|---|---|---|
| ALL | 16 | 4 |
| subtype1 | 2 | 0 |
| subtype2 | 4 | 2 |
| subtype3 | 0 | 0 |
| subtype4 | 3 | 1 |
| subtype5 | 1 | 0 |
| subtype6 | 3 | 0 |
| subtype7 | 3 | 1 |
Figure S319. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #21: 'RADIATION_ADJUVANT_UNITS'
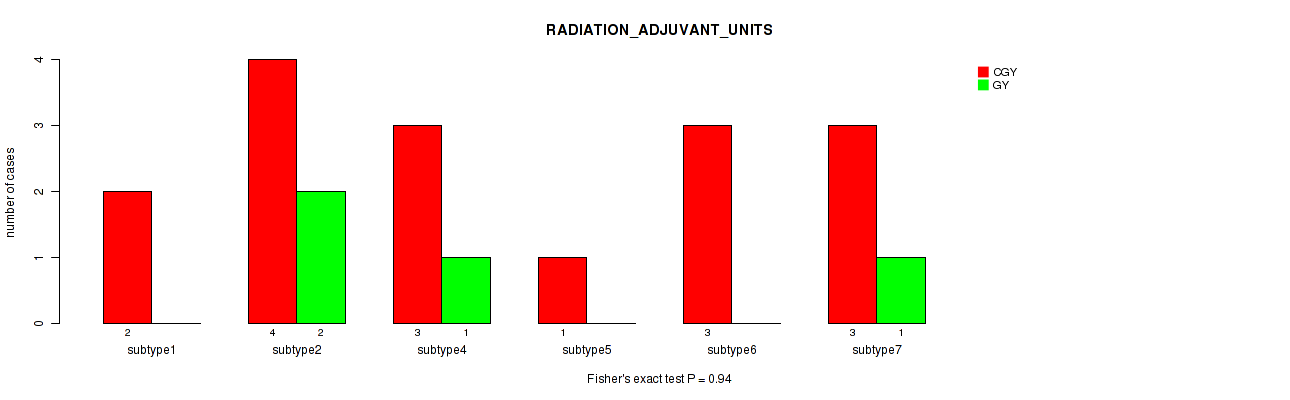
P value = 0.375 (Kruskal-Wallis (anova)), Q value = 1
Table S328. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 175 | 3.5 (2.4) |
| subtype1 | 21 | 3.0 (2.2) |
| subtype2 | 34 | 3.4 (2.8) |
| subtype3 | 17 | 3.9 (1.9) |
| subtype4 | 26 | 3.7 (2.6) |
| subtype5 | 25 | 4.0 (3.3) |
| subtype6 | 24 | 2.9 (2.2) |
| subtype7 | 28 | 3.5 (1.5) |
Figure S320. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #22: 'PREGNANCIES_COUNT_TOTAL'

P value = 0.26 (Kruskal-Wallis (anova)), Q value = 1
Table S329. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 100 | 0.1 (0.4) |
| subtype1 | 13 | 0.1 (0.3) |
| subtype2 | 21 | 0.0 (0.2) |
| subtype3 | 15 | 0.3 (0.8) |
| subtype4 | 15 | 0.0 (0.0) |
| subtype5 | 15 | 0.1 (0.3) |
| subtype6 | 7 | 0.0 (0.0) |
| subtype7 | 14 | 0.0 (0.0) |
Figure S321. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #23: 'PREGNANCIES_COUNT_STILLBIRTH'
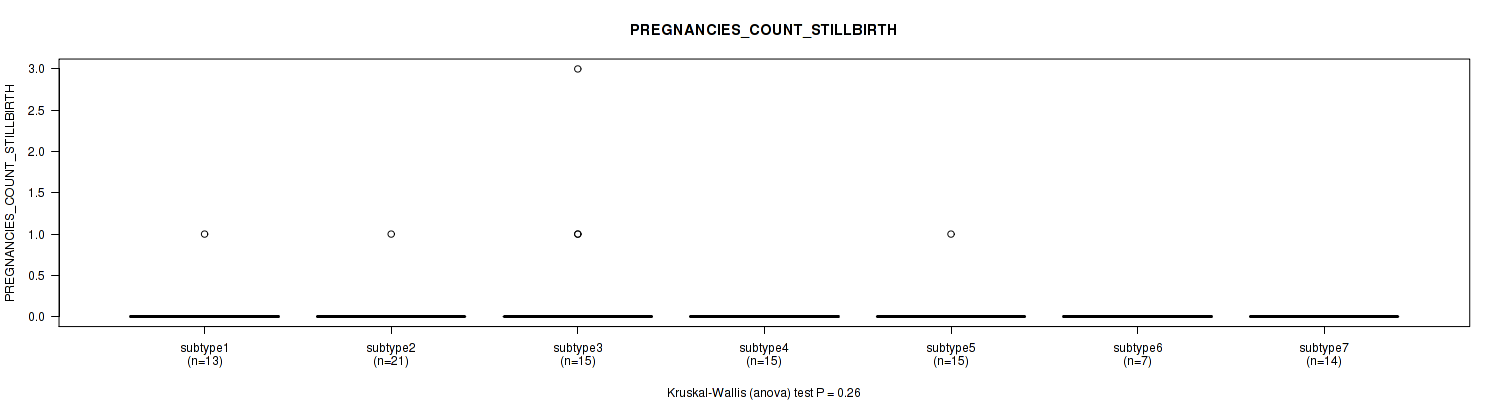
P value = 0.269 (Kruskal-Wallis (anova)), Q value = 1
Table S330. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 116 | 0.4 (0.6) |
| subtype1 | 12 | 0.5 (0.5) |
| subtype2 | 25 | 0.3 (0.6) |
| subtype3 | 15 | 0.3 (0.6) |
| subtype4 | 18 | 0.2 (0.4) |
| subtype5 | 16 | 0.3 (0.8) |
| subtype6 | 11 | 0.7 (0.8) |
| subtype7 | 19 | 0.3 (0.5) |
Figure S322. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #24: 'PATIENT.PATIENTPREGNANCYSPONTANEOUSABORTIONCOUNT'
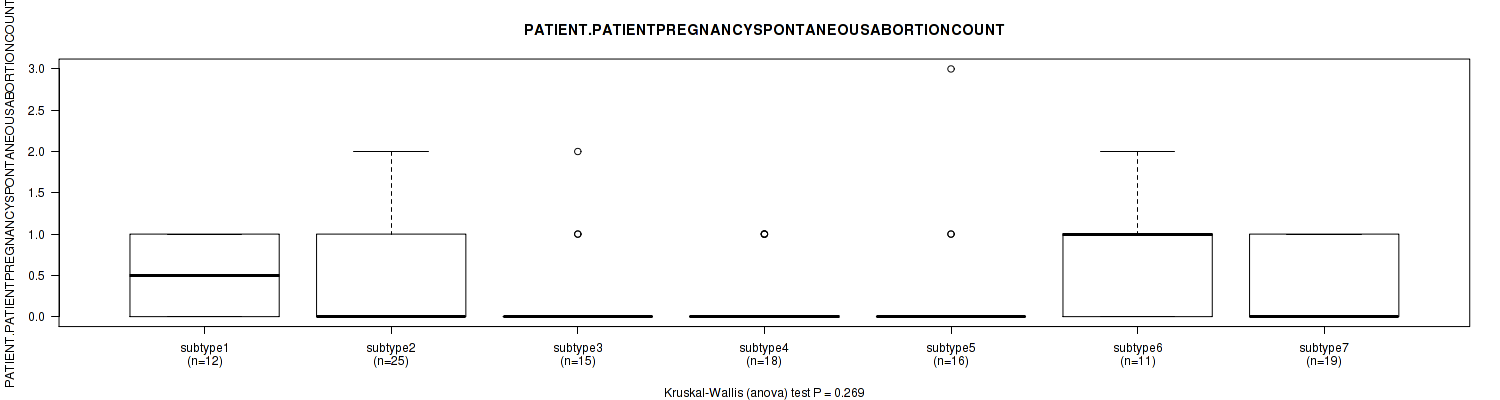
P value = 0.21 (Kruskal-Wallis (anova)), Q value = 1
Table S331. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 179 | 2.5 (1.8) |
| subtype1 | 20 | 2.3 (2.1) |
| subtype2 | 34 | 2.2 (1.5) |
| subtype3 | 17 | 2.7 (1.8) |
| subtype4 | 26 | 2.9 (2.2) |
| subtype5 | 26 | 2.7 (1.9) |
| subtype6 | 25 | 2.0 (1.7) |
| subtype7 | 31 | 2.6 (1.3) |
Figure S323. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #25: 'PREGNANCIES_COUNT_LIVE_BIRTH'

P value = 0.737 (Kruskal-Wallis (anova)), Q value = 1
Table S332. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 109 | 0.9 (1.9) |
| subtype1 | 14 | 0.4 (0.6) |
| subtype2 | 23 | 1.1 (2.7) |
| subtype3 | 14 | 0.7 (1.1) |
| subtype4 | 17 | 0.6 (0.9) |
| subtype5 | 16 | 1.4 (3.0) |
| subtype6 | 8 | 1.2 (1.4) |
| subtype7 | 17 | 0.9 (1.3) |
Figure S324. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #26: 'PATIENT.PATIENTPREGNANCYTHERAPEUTICABORTIONCOUNT'

P value = 0.765 (Kruskal-Wallis (anova)), Q value = 1
Table S333. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 101 | 0.1 (0.3) |
| subtype1 | 12 | 0.1 (0.3) |
| subtype2 | 21 | 0.1 (0.3) |
| subtype3 | 15 | 0.1 (0.3) |
| subtype4 | 15 | 0.0 (0.0) |
| subtype5 | 17 | 0.2 (0.4) |
| subtype6 | 7 | 0.1 (0.4) |
| subtype7 | 14 | 0.1 (0.5) |
Figure S325. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #27: 'PREGNANCIES_COUNT_ECTOPIC'

P value = 0.309 (Fisher's exact test), Q value = 1
Table S334. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'
| nPatients | 2003 | 2010 | COMMON ILIAC | PELVIC (EXTERNAL ILIAC, INTERNAL ILIAC, OBTURATOR) |
|---|---|---|---|---|
| ALL | 1 | 1 | 1 | 41 |
| subtype1 | 0 | 0 | 0 | 5 |
| subtype2 | 0 | 0 | 0 | 13 |
| subtype3 | 0 | 0 | 0 | 6 |
| subtype4 | 1 | 0 | 0 | 4 |
| subtype5 | 0 | 1 | 0 | 6 |
| subtype6 | 0 | 0 | 1 | 1 |
| subtype7 | 0 | 0 | 0 | 6 |
Figure S326. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #28: 'LYMPH_NODE_LOCATION'

P value = 0.984 (Fisher's exact test), Q value = 1
Table S335. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'
| nPatients | MACROSCOPIC PARAMETRIAL INVOLVEMENT | MICROSCOPIC PARAMETRIAL INVOLVEMENT | OTHER LOCATION, SPECIFY | POSITIVE BLADDER MARGIN | POSITIVE VAGINAL MARGIN |
|---|---|---|---|---|---|
| ALL | 2 | 7 | 13 | 1 | 6 |
| subtype1 | 1 | 0 | 0 | 0 | 1 |
| subtype2 | 0 | 3 | 4 | 1 | 2 |
| subtype3 | 0 | 2 | 3 | 0 | 1 |
| subtype4 | 0 | 1 | 2 | 0 | 1 |
| subtype5 | 0 | 1 | 2 | 0 | 1 |
| subtype6 | 0 | 0 | 0 | 0 | 0 |
| subtype7 | 1 | 0 | 2 | 0 | 0 |
Figure S327. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #29: 'LOCATION_OF_POSITIVE_MARGINS'

P value = 0.025 (Fisher's exact test), Q value = 1
Table S336. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #30: 'MENOPAUSE_STATUS'
| nPatients | INDETERMINATE (NEITHER PRE OR POSTMENOPAUSAL) | PERI (6-12 MONTHS SINCE LAST MENSTRUAL PERIOD) | POST (PRIOR BILATERAL OVARIECTOMY OR >12 MO SINCE LMP WITH NO PRIOR HYSTERECTOMY) | PRE (<6 MONTHS SINCE LMP AND NO PRIOR BILATERAL OVARIECTOMY AND NOT ON ESTROGEN REPLACEMENT) |
|---|---|---|---|---|
| ALL | 2 | 10 | 58 | 85 |
| subtype1 | 0 | 0 | 6 | 13 |
| subtype2 | 1 | 2 | 10 | 16 |
| subtype3 | 1 | 0 | 8 | 7 |
| subtype4 | 0 | 1 | 2 | 16 |
| subtype5 | 0 | 1 | 15 | 9 |
| subtype6 | 0 | 2 | 4 | 12 |
| subtype7 | 0 | 4 | 13 | 12 |
Figure S328. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #30: 'MENOPAUSE_STATUS'

P value = 0.386 (Fisher's exact test), Q value = 1
Table S337. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 56 | 66 |
| subtype1 | 12 | 5 |
| subtype2 | 12 | 12 |
| subtype3 | 4 | 9 |
| subtype4 | 6 | 8 |
| subtype5 | 6 | 8 |
| subtype6 | 6 | 11 |
| subtype7 | 10 | 13 |
Figure S329. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #31: 'LYMPHOVASCULAR_INVOLVEMENT'

P value = 0.485 (Kruskal-Wallis (anova)), Q value = 1
Table S338. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 123 | 1.1 (2.6) |
| subtype1 | 16 | 1.1 (2.8) |
| subtype2 | 29 | 1.6 (3.4) |
| subtype3 | 13 | 1.7 (2.7) |
| subtype4 | 15 | 1.3 (3.6) |
| subtype5 | 14 | 0.7 (1.1) |
| subtype6 | 13 | 0.2 (0.4) |
| subtype7 | 23 | 0.7 (1.4) |
Figure S330. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #32: 'LYMPH_NODES_EXAMINED_HE_COUNT'

P value = 0.24 (Kruskal-Wallis (anova)), Q value = 1
Table S339. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 143 | 22.4 (12.7) |
| subtype1 | 18 | 23.2 (10.7) |
| subtype2 | 31 | 18.0 (11.3) |
| subtype3 | 13 | 26.5 (17.3) |
| subtype4 | 18 | 23.1 (11.3) |
| subtype5 | 19 | 20.7 (13.7) |
| subtype6 | 18 | 21.3 (10.1) |
| subtype7 | 26 | 26.5 (13.9) |
Figure S331. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #33: 'LYMPH_NODES_EXAMINED'

P value = 0.021 (Fisher's exact test), Q value = 1
Table S340. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'
| nPatients | KERATINIZING SQUAMOUS CELL CARCINOMA | NON-KERATINIZING SQUAMOUS CELL CARCINOMA |
|---|---|---|
| ALL | 43 | 88 |
| subtype1 | 0 | 5 |
| subtype2 | 8 | 19 |
| subtype3 | 5 | 9 |
| subtype4 | 12 | 12 |
| subtype5 | 7 | 16 |
| subtype6 | 0 | 13 |
| subtype7 | 11 | 14 |
Figure S332. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #34: 'KERATINIZATION_SQUAMOUS_CELL'

P value = 0.000681 (Kruskal-Wallis (anova)), Q value = 0.23
Table S341. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 200 | 2007.3 (5.2) |
| subtype1 | 23 | 2008.3 (4.7) |
| subtype2 | 40 | 2007.8 (4.9) |
| subtype3 | 17 | 2002.6 (5.4) |
| subtype4 | 31 | 2005.4 (5.2) |
| subtype5 | 29 | 2007.7 (5.6) |
| subtype6 | 27 | 2008.9 (4.7) |
| subtype7 | 33 | 2008.5 (4.2) |
Figure S333. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #35: 'INITIAL_PATHOLOGIC_DX_YEAR'
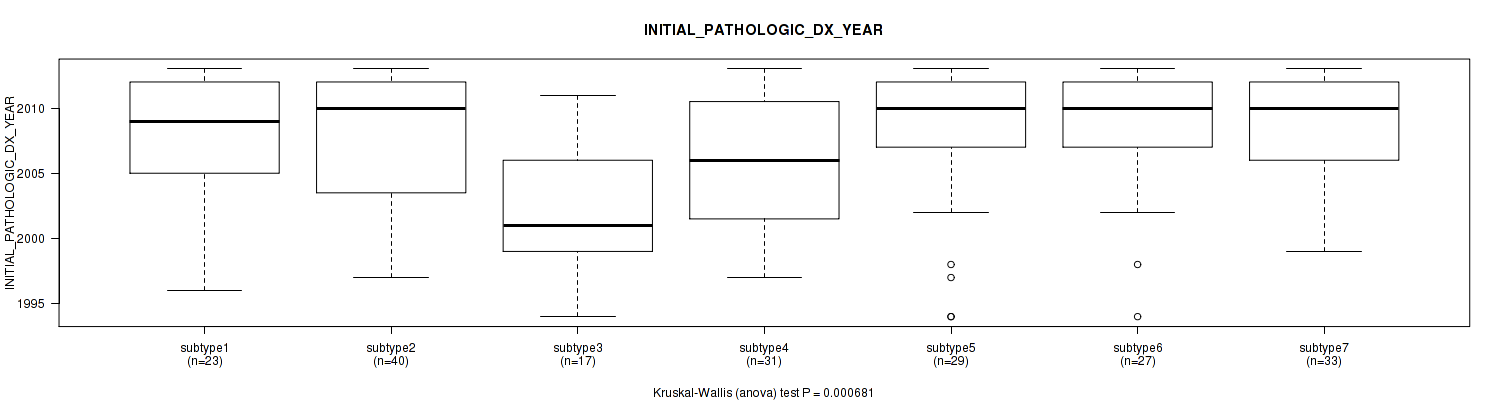
P value = 0.028 (Fisher's exact test), Q value = 1
Table S342. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'
| nPatients | OTHER | RADICAL HYSTERECTOMY | SIMPLE HYSTERECTOMY |
|---|---|---|---|
| ALL | 3 | 129 | 5 |
| subtype1 | 2 | 14 | 1 |
| subtype2 | 0 | 26 | 0 |
| subtype3 | 0 | 13 | 0 |
| subtype4 | 0 | 13 | 2 |
| subtype5 | 1 | 15 | 0 |
| subtype6 | 0 | 20 | 2 |
| subtype7 | 0 | 28 | 0 |
Figure S334. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #36: 'HYSTERECTOMY_TYPE'

P value = 0.172 (Fisher's exact test), Q value = 1
Table S343. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'
| nPatients | CURRENT USER | FORMER USER | NEVER USED |
|---|---|---|---|
| ALL | 8 | 40 | 47 |
| subtype1 | 3 | 7 | 3 |
| subtype2 | 0 | 10 | 10 |
| subtype3 | 0 | 1 | 7 |
| subtype4 | 2 | 6 | 6 |
| subtype5 | 0 | 4 | 9 |
| subtype6 | 2 | 6 | 6 |
| subtype7 | 1 | 6 | 6 |
Figure S335. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #37: 'HISTORY_HORMONAL_CONTRACEPTIVES_USE'

P value = 0.482 (Kruskal-Wallis (anova)), Q value = 1
Table S344. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 173 | 162.1 (7.2) |
| subtype1 | 21 | 164.8 (8.4) |
| subtype2 | 37 | 162.6 (7.5) |
| subtype3 | 13 | 161.0 (7.0) |
| subtype4 | 25 | 161.9 (7.4) |
| subtype5 | 27 | 160.6 (7.9) |
| subtype6 | 23 | 161.7 (4.5) |
| subtype7 | 27 | 161.8 (6.8) |
Figure S336. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #38: 'HEIGHT_CM_AT_DIAGNOSIS'

P value = 0.899 (Fisher's exact test), Q value = 1
Table S345. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'
| nPatients | ABSENT | PRESENT |
|---|---|---|
| ALL | 81 | 15 |
| subtype1 | 14 | 1 |
| subtype2 | 16 | 3 |
| subtype3 | 9 | 3 |
| subtype4 | 10 | 1 |
| subtype5 | 9 | 2 |
| subtype6 | 10 | 2 |
| subtype7 | 13 | 3 |
Figure S337. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #39: 'CORPUS_INVOLVEMENT'

P value = 0.89 (Fisher's exact test), Q value = 1
Table S346. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'
| nPatients | CARBOPLATIN | CISPLATIN |
|---|---|---|
| ALL | 3 | 17 |
| subtype1 | 0 | 0 |
| subtype2 | 1 | 4 |
| subtype3 | 2 | 3 |
| subtype4 | 0 | 5 |
| subtype5 | 0 | 2 |
| subtype6 | 0 | 1 |
| subtype7 | 0 | 2 |
Figure S338. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #40: 'CHEMO_CONCURRENT_TYPE'
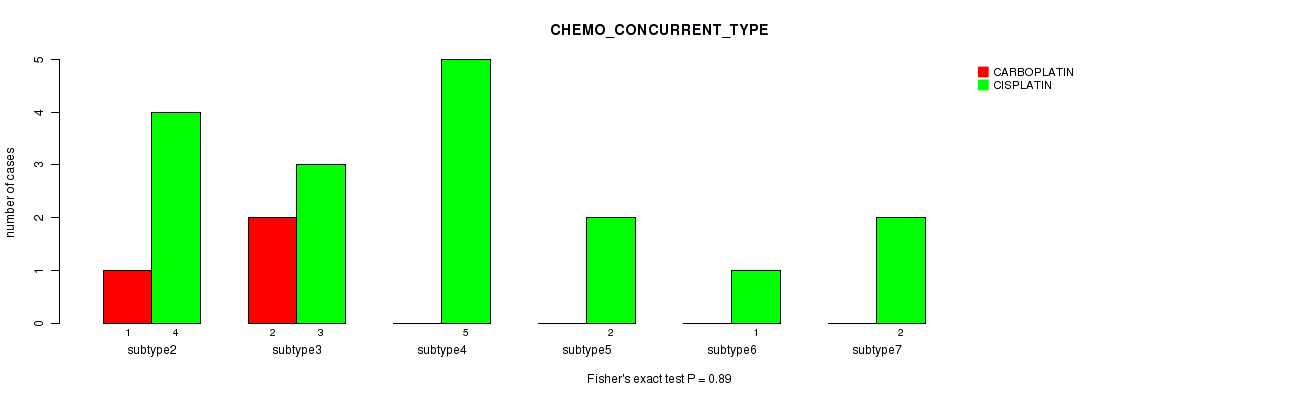
P value = 0.0809 (Kruskal-Wallis (anova)), Q value = 1
Table S347. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #41: 'CERVIX_SUV_RESULTS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 10 | 12.8 (6.1) |
| subtype2 | 1 | 6.0 (NA) |
| subtype3 | 2 | 9.3 (0.9) |
| subtype4 | 3 | 20.1 (5.2) |
| subtype5 | 1 | 11.1 (NA) |
| subtype7 | 3 | 10.8 (3.8) |
Figure S339. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #41: 'CERVIX_SUV_RESULTS'
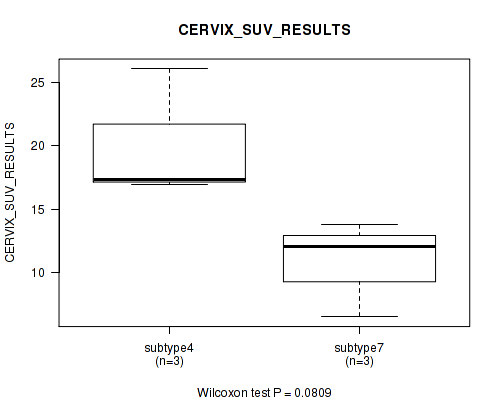
P value = 0.304 (Fisher's exact test), Q value = 1
Table S348. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'
| nPatients | T1A | T1A1 | T1B | T1B1 | T1B2 | T2 | T2A | T2A1 | T2A2 | T2B | T3B | T4 | TIS | TX |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ALL | 2 | 1 | 26 | 55 | 24 | 3 | 6 | 7 | 9 | 14 | 2 | 2 | 1 | 8 |
| subtype1 | 0 | 0 | 1 | 11 | 2 | 0 | 1 | 2 | 1 | 1 | 0 | 0 | 0 | 0 |
| subtype2 | 0 | 0 | 5 | 10 | 3 | 1 | 1 | 2 | 2 | 3 | 1 | 1 | 0 | 4 |
| subtype3 | 0 | 0 | 6 | 5 | 1 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 0 |
| subtype4 | 0 | 0 | 6 | 6 | 2 | 0 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 2 |
| subtype5 | 1 | 0 | 4 | 3 | 6 | 1 | 0 | 0 | 2 | 5 | 0 | 0 | 0 | 1 |
| subtype6 | 0 | 0 | 1 | 8 | 5 | 1 | 1 | 1 | 3 | 1 | 0 | 0 | 0 | 1 |
| subtype7 | 1 | 1 | 3 | 12 | 5 | 0 | 2 | 1 | 0 | 2 | 0 | 0 | 0 | 0 |
Figure S340. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #42: 'AJCC_TUMOR_PATHOLOGIC_PT'

P value = 0.0105 (Kruskal-Wallis (anova)), Q value = 1
Table S349. Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
| nPatients | Mean (Std.Dev) | |
|---|---|---|
| ALL | 200 | 47.6 (13.4) |
| subtype1 | 23 | 42.6 (11.2) |
| subtype2 | 40 | 49.2 (11.3) |
| subtype3 | 17 | 49.8 (16.2) |
| subtype4 | 31 | 41.1 (13.7) |
| subtype5 | 29 | 51.4 (13.9) |
| subtype6 | 27 | 46.1 (9.0) |
| subtype7 | 33 | 51.7 (15.3) |
Figure S341. Get High-res Image Clustering Approach #8: 'MIRseq Mature cHierClus subtypes' versus Clinical Feature #43: 'AGE_AT_DIAGNOSIS'
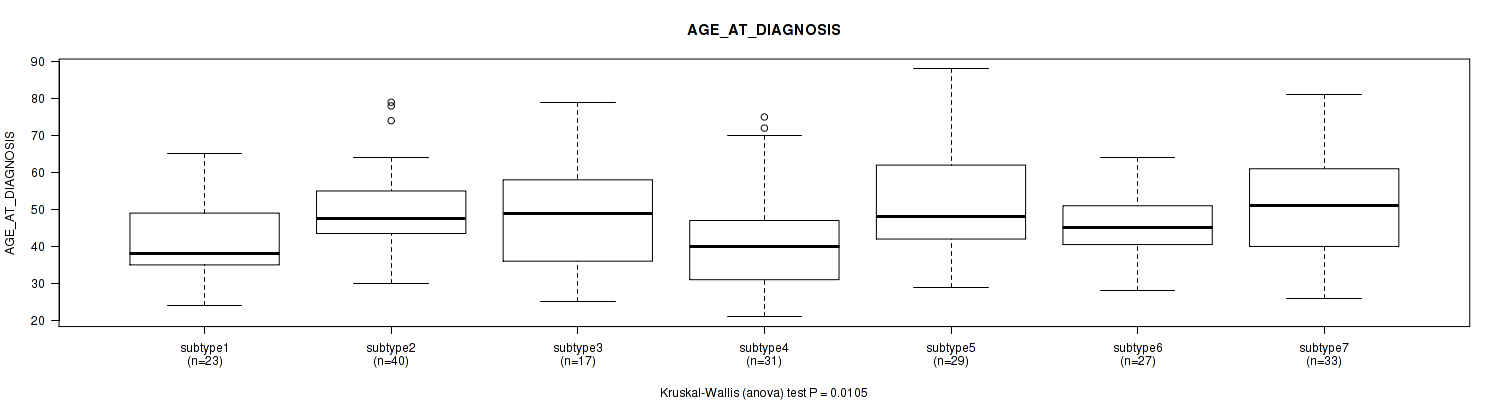
-
Cluster data file = CESC-TP.mergedcluster.txt
-
Clinical data file = CESC-TP.merged_data.txt
-
Number of patients = 200
-
Number of clustering approaches = 8
-
Number of selected clinical features = 43
-
Exclude small clusters that include fewer than K patients, K = 3
consensus non-negative matrix factorization clustering approach (Brunet et al. 2004)
Resampling-based clustering method (Monti et al. 2003)
For survival clinical features, the Kaplan-Meier survival curves of tumors with and without gene mutations were plotted and the statistical significance P values were estimated by logrank test (Bland and Altman 2004) using the 'survdiff' function in R
For binary clinical features, two-tailed Fisher's exact tests (Fisher 1922) were used to estimate the P values using the 'fisher.test' function in R
For multiple hypothesis correction, Q value is the False Discovery Rate (FDR) analogue of the P value (Benjamini and Hochberg 1995), defined as the minimum FDR at which the test may be called significant. We used the 'Benjamini and Hochberg' method of 'p.adjust' function in R to convert P values into Q values.
In addition to the links below, the full results of the analysis summarized in this report can also be downloaded programmatically using firehose_get, or interactively from either the Broad GDAC website or TCGA Data Coordination Center Portal.